Advances in Bioscience and Biotechnology
Vol.4 No.7(2013), Article ID:33889,7 pages DOI:10.4236/abb.2013.47101
Shared and discrete interacting partners of ELL1 and ELL2 by yeast two-hybrid assay
![]()
Department of Immunology, School of Medicine, University of Pittsburgh, Pittsburgh, USA
Email: Milcarek@pitt.edu
Copyright © 2013 Fortuna Arumemi et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Received 1 May 2013; revised 30 May 2013; accepted 10 June 2013
Keywords: Transcription Elongation; Immunoglobulin Synthesis; Yeast Two-Hybrid System
ABSTRACT
ELL2 (eleven-nineteen lysine-rich leukemia transcription elongation factor), a component of a larger complex with pTEFb (cyclin T and CDK9) and AF4, is up-regulated in plasma cells where it influences mRNA processing by increasing exon skipping and enhancing proximal poly (A) site use. ELL2 is needed to produce the secretory-specific Ig heavy chain mRNA while ELL1 mRNA does not change in abundance with B cell stages. To investigate the potential interactions of other proteins with the ELL1 and ELL2 proteins, we preformed yeast two-hybrid studies. HSP40 and Testin were found to bind to ELL2 in its amino-terminal half. PCNA binds to ELL2 in a region encompassing amino acids 186 - 344. The potent transcription factors HIF1 α and ZNF622 interact with both ELL1 and 2 in the central, proline rich region. Meanwhile, BBS2 and ING3 interact with ELL1 but not ELL2 in this central proline-rich region. Many of the ELL-interacting-proteins uncovered in the two-hybrid screen are tumour suppressors that may work through the ELL: pTEFb complex to suppress or activate sets of genes in plasma cells.
1. INTRODUCTION
The RNA polymerase II on the Ig heavy chain gene (Igh) in plasma cells is associated with high levels of serine phosphorylation of the carboxyl-terminal domain, addition of polyadenylation factors, and the transcription elongation factors ELL2 and PC4 [1]. This sets the polymerase apart from the RNAP-II in B cells, where it is factor poor [2]. ELL2 has been shown to drive Igh alternative mRNA processing [3]. ELL2 (Eleven-nineteen Lysine-rich Leukemia gene 2) is part of a large “super elongation complex” (SEC) that contains pTEFb and other elongation factors including AF4/aff1 [4]. While ELL1, 2, and 3 proteins, encoded by separate genes, were all shown to be in SECs [5], subsequent analysis of the SECs indicated sub-types with specificity for different genes [6]. Only ELL2 was found in the SEC with pTEFb and HIV-1 Tat and TAR in T-lymphocytes [7,8]. As shown in Figure 1, other investigators found a distinct ELL2 sub-complex [9]. ELL2 also links to AF4 and to the PAF complex (mPAF or polymerase associated factor); this is crucial for regulation since PAF recruits the mRNA polyadenylation factors, linking SEC to productive elongation and promoter proximal 3’ end processing [8,10].
As shown in Table 1, the ELLs are very similar from aa 1 - 180. This NH2-proximal region is responsible for elongation enhancement and PAF binding [11]. EAF1 was found to interact with ELL in the elongation and Cterminal domains [12]. EAF2 binds to the ELL elongation domain but not the C-terminal domain [13].
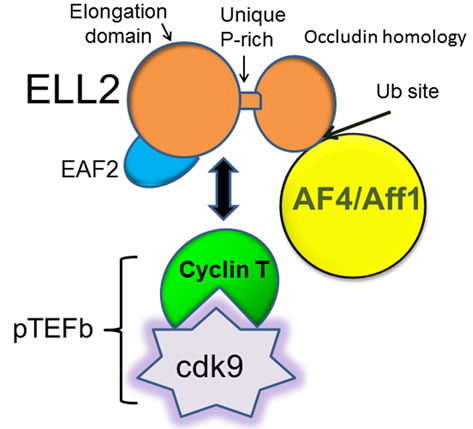
Figure 1. Interactions of ELL2 with pTEFb composed of cyclin T and Cdk9, a serine kinase. The mRNA for Aff1 and ELL2 are both up-regulated in plasma cells [2]. Adapted from [9].
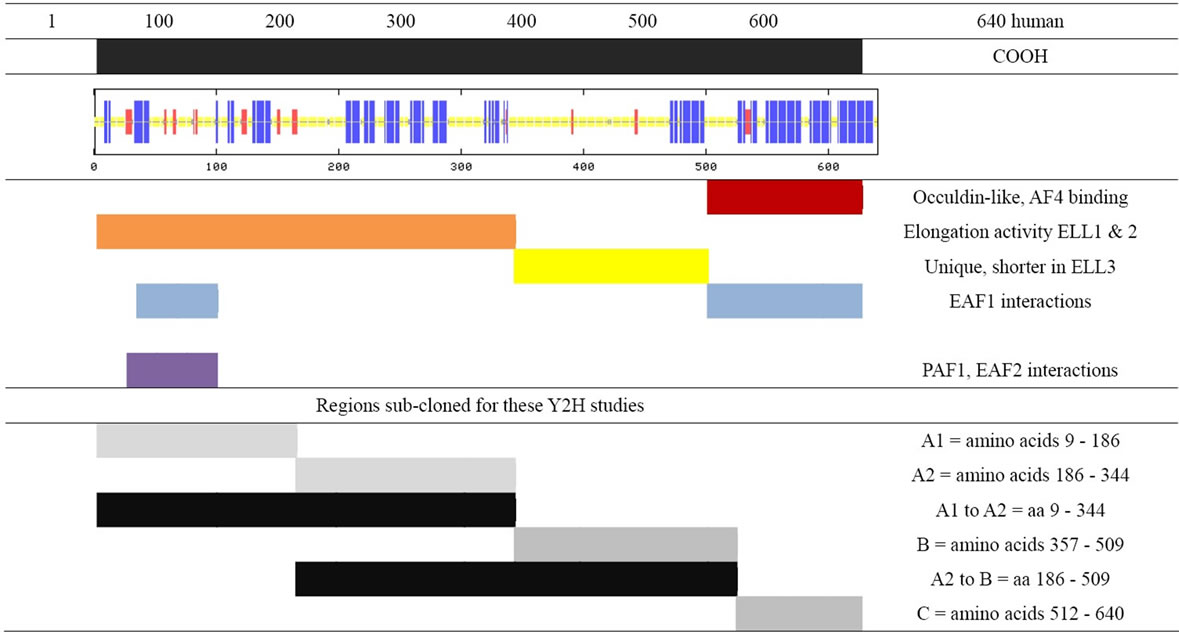
Table 1. Map of human ELL2 indicating important regions discussed in text. In the predicted structure, blue represents regions of alpha helices; red, the beta sheets; and yellow, the unstructured loops.
ELL1 and 2 differ from each other at aa ~352 - 443 and 473 - 516, mapping in exon 8. The mid-portion of mammalian ELL3 is much shorter because of an extensive deletion in this same region and from 180 - 248. The differences in the region covering 352 - 516 may drive unique protein associations of ELL 1 & 2. This portion of ELL1 and 2 is rich in hydrophobic amino acids like proline and leucine but also contains hydrophilic charged residues like lysine and glutamate. No known protein motif has been identified in this domain. It has, however, been determined to be the region that is required for dELL association with heat shock puffs in Drosophila [14].
The C-terminal domain is conserved in phylogeny in all the members of the ELL-family including Drosophila; it shares homology with occludin, an integral plasma membrane protein located in tight junctions. The C-terminal occludin-like domain of all the ELLs forms a unique structure with a highly positively-charged surface; these have been proposed to facilitate protein: protein interactions [15]. This region of ELL2 also contains a ubiquitination site acted on by Siah1 which controls its degradation; binding of AF4 blocks Siah1 binding and enhances stability [16]. The C-terminal domain of dELL is required for viability in Drosophila [14]. The C-terminal domain of mammalian ELL1 is sufficient for immortalization of myeloid progenitors most likely through its interactions with p53 [17]. By analogy, the interaction of ELL3 with p53 in mouse stem cells is most likely through this conserved C-terminal domain [18].
To uncover proteins that might interact with the ELL proteins, a yeast two-hybrid screen was performed. Since the C-terminal domain trans-activated transcription in the yeast two hybrid system on its own, our search was confined to the first ~509 amino acids. Unique protein: protein interactions were uncovered in our screens; ELL1 and ELL2 had some shared and some discrete partners.
2. MATERIALS AND METHODS
2.1. Plasmid Constructs
Portions of the human ELL2 protein coding region, as shown in Table 1, were cloned and then transformed into the pGBKT7 vector construct using the YeastmakerTM Yeast Transformation System (Clonetech). The bait vector used was pGBKT7 and pGADT7 RecAB, the prey vector (Clontech). We cloned human ELL2 divided into 3 parts designated A1-A2, A2-B, and B (Tables 1 and 2) into the multi-cloning site of pGBKT7. All pGADT7 RecAB and pGBKT7-derived vectors were transformed into Y187 and Y2HGold yeast strains, respectively (Clontech Yeastmaker Yeast Transformation System 2). The transformants were, respectively, selected on Leuand Trpminimal media plates after growth at 30˚C for 3 to 5 days.
2.2. Yeast Two-Hybrid Screening
To test for new interactions with the ELL proteins, a yeast two-hybrid screen was performed as described pre-

Table 2. Interactions found through yeast two hybrid assays between segments of human ELL bait and cDNA prey segments in a HeLa activation domain library. The Genebank identifiers and nucleotide numbers encoding the interacting cDNAs are indicated within the boxes.
viously [19,20] using the Matchmaker Gold Yeast TwoHybrid System (Clonetech Catalog # 630489). The bait plasmids described in section 2.1 were transformed into the yeast strain Y2H Gold. The resulting strains were then mated with the yeast strain Y187 that was pretransformed with the Mate & Plate Normalized HeLa Library (Clonetech Catalog # 630479). Colonies were selected on synthetic dropout plates plus 40 ug/ml X-aGal, lacking adenine, histidine, leucine, and tryptophan (SD/-Ade/-His/-Leu/-Trp). Blue colonies (indicating potential protein interaction) were then restreaked on the same dropout plates containing 40 ug/ml X-a-Gal and 125 ng/ml Aureobasidin A to reduce the possibility of false positives. Plasmids from the positive colonies were then isolated and transformed back into Stellar competent E. coli cells. The transformed plasmids were isolated again and sequenced using T7 universal primers to identify the cDNA inserts from the HeLa library. For final conformation, isolated plasmids were transformed individually into Y187 yeast and mated again with Y2HGold yeast containing ELL2 fragments in the method described above. The positive plasmids obtained from these subsequent interactions were isolated and transformed into Stellar competent E. coli cells and were finally sequenced using T7 primers to confirm the interactions found in the library screen.
2.3. mRNA Analyses
The total RNA from B220+ splenic B-cells was isolated from fresh mouse spleens as previously described [3]. Briefly RNA was isolated using the RNeasy mini kit (Qiagen # 74106). The mRNA was copied with reverse transcriptase using Superscript first strand synthesis (Invitrogen # 11904-018) and a dT-primer, and then quantitative polymerase chain reaction was performed using the Sybergreen PCR master mix (Applied Bio # 4309155) and the indicated primers. The Ct values of each sample were normalized relative to HPRT. PCR primers were as follows:
HPRT/ ref 16083680/1
Forward CCTAAGATGAGCGCAAGTTGAA Reverse CCACAGGACTAGAACACCTGCTAA
ELL1/ ref 87855970/1
Forward GGAGTTACGGGTTGTCGTGT Reverse AGAGATGTGCCCTTGGCTTC
ELL2/ ref 23545157/8
Forward: CTGGCCAGGTTACAGTGAGA Reverse AGCGTCTTTGCTGGAACAC
ELL3/ ref 85776881/2
Forward: AGAGCCTCTCAGCTCCATCAGCC Reverse GCCCCAACTCGAGCATGCAG
3. RESULTS AND DISCUSSION
3.1. ELL mRNA Expression Pattern in Spleen Cells
We measured the expression of the mouse ELL 1, 2 and 3 mRNAs in the differentiation of B cells to plasmablasts and plasma cells, induced ex vivo by Lipopolysaccharide (LPS), by dT-primed reverse transcriptase followed by real-time quantitative PCR. In these experiments the amount of ELL message was normalized to HPRT set as 1; we have previously shown that HPRT levels do not change in the B to PC transition [21]. We see a shift in the ELL expression patterns, Figure 2. The level of ELL2 mRNA increases at least 6-fold after LPS induction and comprises a substantial portion of the total ELL mRNA population in those cells. Meanwhile ELL3 mRNA is the most abundant ELL in the resting B cells but its level falls 3 - 10 fold after LPS stimulation such that then it comprises less than 10% of the total ELL mRNAs. It is more difficult to estimate how much of each protein is present at the various stages based on the affinities of the antibodies used in westerns, but we have shown previously that ELL2 protein increases substantially versus a control following LPS stimulation [3].
An analysis of B cell stages by mRNA micro-array reported in the http://www.immgen.org web site came to a similar conclusion. As we report here, the amount of ELL1 mRNA does not vary significantly among B cell stages. Meanwhile, ELL2 was highest in germinal centers, where the most mature plasma cells reside, while ELL3 expression was highest in marginal zone B cells, representing less mature B cell stages.
The magnitude of the changeover to ELL2 expression in mature cells following LPS stimulation leads us to ask what factors might be interacting with ELL1 and 2 and how these could modulate IgH secretory mRNA processing. The unique regions between aa 357 to 509 both in mouse and human ELL1 and 2 are the likely targets for interactions with distinct regulatory molecules.
3.2. Yeast Two Hybrid Interactions
We conducted yeast two hybrid interaction studies using regions of human ELL2 and ELL1 that we cloned in the Gal4-binding domain (BD) as “bait”. We omitted ELL3 from our analyses since its level drops rather than rises after LPS stimulation. We screened our bait against a cDNA library (segments encoding ~200 amino acids), using the normalized library of HeLa S3; here the cDNAs are cloned into a plasmid containing the transcriptional activation domain (aka “prey” in an AD library). The HeLa “normalized” library was chosen because it was pre-hybridized to eliminate highly abundant genes like rRNA proteins and HeLa expresses numerous plasma cell mRNAs and proteins, for example blimp-1 and IRF4 (data not shown).
Regions of human ELL2 were cloned in the BD vector as outlined in Table 1 and tested for whether the AD HeLa library cells were required for transcriptional activation. Theoretically, in the yeast two-hybrid system only when the two domains (BD and AD) interact, transcription of a reporter gene ensues. We found that the Cregion of ELL2 in the BD trans-activated all by itself, that is, in the absence of any added mammalian activa-
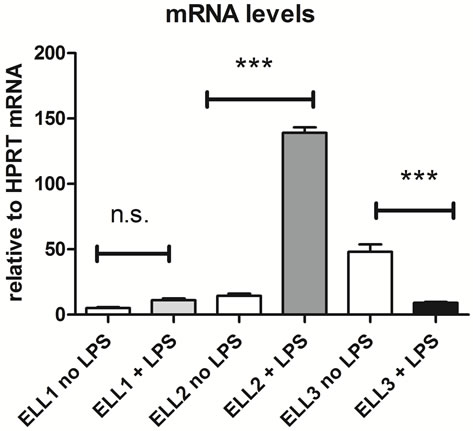
Figure 2. Expression of ELL mRNAs in mouse splenic B-cells relative to HPRT mRNA standard. ***indicates P < 0.001, error bars show SEM.
tion domain partners; this excluded it from further analyses here. This transcriptional activity could be because the C-terminal fragment interacts with basal yeast factors or with DNA through the high positive charge. The cloned ELL2 segments A1 or A2 alone failed to produce any AD partners, perhaps because they were too small. On the other hand, ELL2 fragments encompassing A1 to A2 and A2 to B and B alone interacted with proteins cloned in the AD HeLa library, and yielded multiple isolations of segments of several proteins, as indicated in Table 2.
Segment ELL1 B, but not ELL2 B, in the BD vector interacted with BBS2 in the AD. Mutations in the BBS2 gene have been observed in patients with Bardet-Biedl syndrome type 2, an autosomal recessive disorder characterized by severe retinopathy, obesity, polydactyly, renal malformation, and mental retardation. The mouse knockout of BBS2 shows kidney abnormalities [22]. The significance of this ELL1 interaction is not known.
Repeatedly we found that a protein made from a portion of the HIF1 α mRNA clone bound to ELL2 in the middle/B region and to ELL1 in the same area, amino acids 357-509, corresponding roughly to exon 8 (Tables 1 and 2). The regions in common between the two proteins are at the beginning and end of exon 8 but not the middle ~79 amino acids; one or both of these may be responsible for the interactions. ELL1 had previously been reported to interact with HIF1 α only by co-IP studies [23]. HIF1 α has also been shown to interact with EAF2 [24]. EAF2 interacts with ELL [25] so those coIPs may have been detecting complex formation; here we show a direct ELL1 or ELL2:HIF1 α interaction. Our mapped interactions occur in sequences where a putative exon resides in the 3’ UTR of the HIF α mRNA, a region that is not part of the main HIF1 α open reading frame. But transcript variant 3 (Table 2) was found to use an alternate in-frame acceptor splice site at the next exon compared to variant 1 [26]. In that paper, a novel mRNA isoform of human HIF-1 α was found to be up-regulated following T cell receptor stimulation. This activationinducible isoform was expressed using the alternative first exon I.3 and would encode a protein 24 amino acids longer than the ubiquitous HIF-1 α isoform. This isoform I.3 shows its highest expression in peripheral blood leukocytes (neutrophils, B and T cells) and the thymus [26]. This suggests that ELL2 may play a role in the exon skipping and alternative splicing of HIF-1α in lymphoid cells. Hypoxia inducible factor (HIF-1) is a transcription factor with two subunits, a constitutive β subunit and an α subunit; its expression is increased in mammalian cells cultured under reduced oxygen and HIF-1 induces numerous genes [27]. Those induced proteins play essential roles in responses to hypoxia and may also link inflammatory mediators to ion channel expression [28]. Plasma cells produce immunoglobulins under hypoxic conditions. Mice with a conditional HIF-1 α deficiency in the B cell lineage still produced secreted autoantibodies but peritoneal B1 cells and B2 cell maturation were impaired [29]. Thus the HIF-1 α interactions we see here with ELL2 may be biologically significant for plasma cell development.
Hsp40 interacts in the yeast two-hybrid study with the A1 to A2 protein fragment of ELL2, encompassing the elongation region of ELL2. Hsp40 proteins are conserved from Escherichia coli to humans; they help to degrade proteins to which they bind [30]. We previously showed that in the elongation domain of ELL2, around amino acids 133 - 186, there are numerous potential proteolytic cleavage sites, conserved from zebrafish to humans, on which proteases could act [3]. These potential cleavage sites coupled with the associations with Hsp40 may account for the apparent rapid turnover of ELL2 [7].
A portion of the ING3, inhibitor of growth protein 3, well within the protein coding region of both spliced variants, was found in our yeast two hybrid studies to interact with the unique (B) region of ELL1. The ING genes encode a conserved homeo-domain that can facilitate binding to the trimethylated lysine of histone H3, H3K4me3, [31]. ING3 contains a PHD-finger, a common motif in proteins involved in chromatin remodeling. ING3 can act as a tumor suppressor and interfering with its degradation may aid in control of tumors, especially melanoma [32]. ING3 interactions with ELL1 could account for a suppression of transcription.
PCNA (proliferating cell nuclear antigen) has many functions in DNA replication and repair; proteins that bind to PCNA have been shown to be subject to ubiquitination and more rapid turnover [33]. In systemic lupus erythematous (where many autoantibodies are secreted) the clinical significance of the anti-PCNA antibodies remains enigmatic [34]. PCNA is expressed throughout the differentiation from B cell to plasma cell but the specific interactions with ELL2 in the A2-B region that we see here may be biologically significant. PCNA and Hsp40 interactions may contribute to the degradation of ELL2 in plasma cells.
TES, the so called testis derived transcript (3 LIM domains) or Testin, is, in fact, ubiquitously expressed and is involved in adhesion, motility, cell-to-cell interactions and cell signaling. Testin was found in many tissues but is differentially expressed in mammary versus coronary arteries [35]. Testin is also silenced in childhood blood tumors, like Acute Lymphocytic Leukemia [36]. In our analyses we found that a portion of the open reading frame of Testin interacts with the A1 to A2 region of ELL2. Testin appears to be a tumor suppressor [37]. The significance of its interaction with ELL2 for plasma cell differentiation remains obscure but intriguing based on its role in hematopoietic tumors.
In our studies, ZNF622 interacts with both ELL1 and ELL2. ZNF622 is a positive regulator of the MAPK cascade and behaves as an activator of the ubiquitous, DNA bound transcription factor MYBL2 [38].
Thus it seems clear that while ELL1 and 2 each interact with HIF-1α and ZNF622, each has unique interactions with other transcription factors. The significance of this for the genes that each transcription factor targets remains to be explained. The interaction of the tumor suppressor TP53 with the conserved C-terminal portion of ELL and the interactions of the tumor suppressors ING3 and TES with other portions of ELL1 or 2 suggest that they may all act through the pTEFb super elongation complex to suppress activation of certain genes whose expression would lead to uncontrolled growth.
4. CONCLUSION
Some have suggested that the role of individual ELL factors may be redundant but the observation that the timing of their expression, primary sequence differences, and their interactions with other proteins as shown here in the yeast two hybrid system permit us to consider that each factor may play a unique role in gene expression. Once we fully understand how polyadenylation is linked to transcription through the elongation process, and specifically through ELL2, we may then be able to more easily see how the competition between splicing and polyadenylation can be accomplished and if the factors we have identified play a role.
5. ACKNOWLEDGEMENTS
This work was supported by MCB-0842725 to CM, training support to F.A. from 3R01CA056103-16S1 to O. J. Finn, and UPCI shared resources, supported in part by award P30CA047904.
![]()
![]()
REFERENCES
- Shell, S. A., Martincic, K., Tran, J. and Milcarek, C. (2007) Increased phosphorylation of the carboxyl terminal domain of RNA polymerase II and loading of polyadenylation and co-transcriptional factors contribute to regulation of the Ig heavy chain mRNA in plasma cells. The Journal of Immunology, 179, 7663-7673.
- Milcarek, C., Albring, M., Langer, C. and Park, K.S. (2011) The eleven-nineteen lysine-rich leukemia gene (ELL2) influences the histone H3 modifications accompanying the shift to secretory Immunoglobulin heavy chain mRNA production. Journal of Biological Chemistry, 286, 33795-33803. doi:10.1074/jbc.M111.272096
- Martincic, K., Alkan, S.A., Cheatle, A., Borghesi, L. and Milcarek, C. (2009) Transcription elongation factor ELL2 directs immunoglobulin secretion in plasma cells by stimulating altered RNA processing. Nature Immunology, 10, 1102-1109. doi:10.1038/ni.1786
- Lin, C., Garrett, A.S., De Kumar, B., Smith, E.R., Gogol, M., Seidel, C., Krumlauf, R. and Shilatifard, A. (2011) Dynamic transcriptional events in embryonic stem cells mediated by the super elongation complex (SEC). Genes & Development, 25, 1486-1498. doi:10.1101/gad.2059211
- Smith, E., Lin, C. and Shilatifard, A. (2011) The super elongation complex (SEC) and MLL in development and disease. Genes & Development, 25, 661-672. doi:10.1101/gad.2015411
- Luo, Z., Lin, C., Guest, E., Garrett, A.S., Mohaghegh, N., Swanson, S., Marshall, S., Florens, L., Washburn, M.P. and Shilatifard, A. (2012) The super elongation complex family of RNA polymerase II elongation factors: Gene target specificity and transcriptional output. Molecular and Cellular Biology, 32, 2608-2617. doi:10.1128/MCB.00182-12
- He, N., Liu, M., Hsu, J., Xue, Y., Chou, S., Burlingame, A., Krogan, N.J., Alber, T. and Zhou, Q. (2010) HIV-1 tat and host AFF4 recruit two transcription elongation factors into a bifunctional complex for coordinated activation of HIV-1 transcription. Molecular Cell, 38, 428- 438. doi:10.1016/j.molcel.2010.04.013
- He, N., Chan, C.K., Sobhian, B., Chou, S., Xue, Y., Liu, M., Alber, T., Benkirane, M. and Zhou, Q. (2011) Human polymerase-associated factor complex (PAFc) connects the super elongation complex (SEC) to RNA polymerase II on chromatin. Proceedings of the National Academy of Sciences, 108, E636-E645. doi:10.1073/pnas.1107107108
- Biswas, D., Milne, T.A., Basrur, V., Kim, J., ElenitobaJohnson, K.S.J., Allis, C.D. and Roeder, R.G. (2011) Function of leukemogenic mixed lineage leukemia 1 (MLL) fusion proteins through distinct partner protein complexes. Proceedings of the National Academy of Sciences, 108, 15751-15756. doi:10.1073/pnas.1111498108
- Barboric, M., Lenasi, T., Chen, H., Johansen, E.B., Guo, S. and Peterlin, B.M. (2009) 7SK snRNP/pTEFb couples transcription elongation with alternative splicing and is essential for vertebrate development. Proceedings of the National Academy of Sciences of the USA, 106, 7798- 7803. doi:10.1073/pnas.0903188106
- Chou, S., Upton, H., Bao, K., Schulze-Gahmen, U., Samelson, A.J., He, N., Nowak, A., Lu, H., Krogan, N.J., Zhou, Q. and Alber, T. (2013) HIV-1 tat recruits transcription elongation factors dispersed along a flexible AFF4 scaffold. Proceedings of the National Academy of Sciences, 110, E123-E131. doi:10.1073/pnas.1216971110
- Luo, R.T., Lavau, C., Du, C.C., Simone, F., Polak, P.E., Kawamata, S. and Thirman, M.J. (2001) The elongation domain of ELL is dispensable but its ELL-associated factor 1 interaction domain is essential for MLL-ELLinduced leukemogenesis. Molecular and Cellular Biology, 21, 5678-5687. doi:10.1128/MCB.21.16.5678-5687.2001
- Simone, F., Luo, R.T., Polak, P.E., Kaberlein, J.J. and Thirman, M.J. (2003) ELL-associated factor 2 (EAF2), a functional homolog of EAF1 with alternative ELL binding properties. Blood, 101, 2355-2362. doi:10.1182/blood-2002-06-1664
- Gerber, M.A., Shilatifard, A. and Eissenberg, J.C. (2005) Mutational analysis of an RNA polymerase II elongation factor in drosophila melanogaster. Molecular and Cellular Biology, 25, 7803-7811. doi:10.1128/MCB.25.17.7803-7811.2005
- Li, Y., Fanning, A.S., Anderson, J.M. and Lavie, A. (2005) Structure of the conserved cytoplasmic C-terminal domain of occludin: Identification of the ZO-1 binding surface. Journal of Molecular Biology, 352, 151-164. doi:10.1016/j.jmb.2005.07.017
- Liu, M., Hsu, J., Chan, C., Li, Z. and Zhou, Q. (2012) The ubiquitin ligase Siah1 controls ELL2 stability and formation of super elongation complexes to modulate gene transcription. Molecular Cell, 46, 325-334. doi:10.1016/j.molcel.2012.03.007
- Wiederschain, D., Kawai, H., Gu, J., Shilatifard, A. and Yuan, Z.-M. (2003) Molecular basis of p53 functional inactivation by the leukemic protein MLL-ELL. Molecular and Cellular Biology, 23, 4230-4246. doi:10.1128/MCB.23.12.4230-4246.2003
- Ahn, H.-J., Cha, Y., Moon, S.-H., Jung, J.-E. and Park, K.-S. (2012) ELL3 enhances differentiation of mouse embryonic stem cells by regulating epithelial-mesenchymal transition and apoptosis. PLoS ONE, 7, e40293. doi:10.1371/journal.pone.0040293
- Du, J.X., Hagos, E.G., Nandan, M.O., Bialkowska, A.B., Yu, B. and Yang, V.W. (2011) The E3 ubiquitin ligase SMAD ubiquitination regulatory factor 2 negatively regulates Krüppel-like factor 5 protein. Journal of Biological Chemistry, 286, 40354-40364. doi:10.1074/jbc.M111.258707
- Du, J.X., Yun, C.C., Bialkowska, A. and Yang, V.W. (2007) Protein inhibitor of activated STAT1 interacts with and up-regulates activities of the pro-proliferative transcription factor Krüppel-like factor 5. Journal of Biological Chemistry, 282, 4782-4793. doi:10.1074/jbc.M603413200
- Harrold, S., Genovese, C., Kobrin, B., Morrison, S.L. and Milcarek, C. (1991) A comparison of apparent mRNA half-life using kinetic labeling techniques vs decay following administration of transcriptional inhibitors. Analytical Biochemistry, 198, 19-29. doi:10.1016/0003-2697(91)90500-S
- Guo, D.-F., Beyer, A.M., Yang, B., Nishimura, D.Y., Sheffield, V.C. and Rahmouni, K. (2011) Inactivation of Bardet-Biedl syndrome genes causes kidney defects. American Journal of Physiology—Renal Physiology, 300, F574-F580. doi:10.1152/ajprenal.00150.2010
- Liu, L., Ai, J., Xiao, W., Liu, J., Wang, Y., Xin, D., He, Z., Guo, Y. and Wang, Z. (2010) ELL is an HIF-1α partner that regulates and responds to hypoxia response in PC3 cells. Prostate, 70, 797-805.
- Xiao, W., Ai, J., Habermacher, G., Volpert, O., Yang, X., Zhang, A.-Y., Hahn, J., Cai, X. and Wang, Z. (2009) U19/Eaf2 binds to and stabilizes von Hippel-Lindau protein. Cancer Research, 69, 2599-2606. doi:10.1158/0008-5472.CAN-08-2595
- Xiao, W., Jiang, F. and Wang, Z. (2006) ELL binding regulates U19/EAf2 intracellular localization, stability, and transactivation. Prostate, 66, 1-12. doi:10.1002/pros.20309
- Lukashev, D. and Sitkovsky, M. (2008) Preferential expression of the novel alternative isoform I.3 of hypoxiainducible factor 1α in activated human T lymphocytes. Human Immunology, 69, 421-425. doi:10.1016/j.humimm.2008.05.004
- Semenza, G.L. (1999) Regulation of mammalian O2 homeostasis by hypoxia-inducible factor 1. Annual Review of Cell and Developmental Biology, 15, 551-578. doi:10.1146/annurev.cellbio.15.1.551
- Hatano, N., Itoh, Y., Suzuki, H., Muraki, Y., Hayashi, H., Onozaki, K., Wood, I.C., Beech, D.J. and Muraki, K. (2012) Hypoxia-inducible factor-1α (HIF1α) switches on transient receptor potential ankyrin repeat 1 (TRPA1) gene expression via a hypoxia response element-like Motif to modulate cytokine release. Journal of Biological Chemistry, 287, 31962-31972. doi:10.1074/jbc.M112.361139
- Sitkovsky, M. and Lukashev, D. (2005) Regulation of immune cells by local-tissue oxygen tension: HIF1[alpha] and adenosine receptors. Nature Reviews Immunology, 5, 712-721. doi:10.1038/nri1685
- Stolz, A. and Wolf, D.H. (2010) Endoplasmic reticulum associated protein degradation: A chaperone assisted journey to hell. Biochimica et Biophysica Acta (BBA)— Molecular Cell Research, 1803, 694-705. doi:10.1016/j.bbamcr.2010.02.005
- Ludwig, S., Klitzsch, A. and Baniahmad, A. (2011) The ING tumor suppressors in cellular senescence and chromatin. Cell & Bioscience, 1, 25. doi:10.1186/2045-3701-1-25
- Chen, G., Wang, Y., Garate, M., Zhou, J. and Li, G. (2010) The tumor suppressor ING3 is degraded by SCFSkp2- mediated ubiquitin-proteasome system. Oncogene, 29, 1498-1508. doi:10.1038/onc.2009.424
- Kirchmaier, A.L. (2011) Ub-family modifications at the replication fork: Regulating PCNA-interacting components. FEBS Letters, 585, 2920-2928. doi:10.1016/j.febslet.2011.08.008
- Mahler, M., Miyachi, K., Peebles, C. and Fritzler, M.J. (2012) The clinical significance of autoantibodies to the proliferating cell nuclear antigen (PCNA). Autoimmunity Reviews, 11, 771-775. doi:10.1016/j.autrev.2012.02.012
- Archacki, S.R., Angheloiu, G., Moravec, C.S., Liu, H., Topol, E.J. and Wang, Q.K. (2012) Comparative gene expression analysis between coronary arteries and internal mammary arteries identifies a role for the TES gene in endothelial cell functions relevant to coronary artery disease. Human Molecular Genetics, 21, 1364-1373. doi:10.1093/hmg/ddr574
- Weeks, R., Kees, U., Song, S. and Morison, I. (2010) Silencing of TESTIN by dense biallelic promoter methylation is the most common molecular event in childhood acute lymphoblastic leukaemia. Molecular Cancer, 9, 163. doi:10.1186/1476-4598-9-163
- Ma, H., Weng, D., Chen, Y., Huang, W., Pan, K., Wang, H., Sun, J., Wang, Q., Zhou, Z., Wang, H. and Xia, J. (2010) Extensive analysis of D7S486 in primary gastric cancer supports TESTIN as a candidate tumor suppressor gene. Molecular Cancer, 9, 190. doi:10.1186/1476-4598-9-190
- Seong, H.-A., Kim, K.-T. and Ha, H. (2003) Enhancement of B-MYB transcriptional activity by ZPR9, a novel zinc finger protein. Journal of Biological Chemistry, 278, 9655-9662. doi:10.1074/jbc.M207478200

