Advances in Bioscience and Biotechnology
Vol.4 No.5A(2013), Article ID:31986,9 pages DOI:10.4236/abb.2013.45A002
Novel aspects of tRNA-derived small RNAs with potential impact in infectious diseases
![]()
1Laboratory of Functional Genomics, Institute Pasteur of Montevideo, Montevideo, Uruguay
2Department of Medicine, Hospital de Clínicas, Montevideo, Uruguay
Email: #rgarcia@pasteur.edu.uy
Copyright © 2013 Maria Rosa Garcia-Silva et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Received 28 March 2013; revised 4 May 2013; accepted 21 May 2013
Keywords: Non-Coding RNAs; Small tRNAs; tRFs; tRNA Halves; Infectious Diseases
ABSTRACT
The complete understanding of the growing catalog of regulatory non-coding RNAs is going to shed light in different aspects of a wide range of pathogenic mechanisms in human diseases. This review was aimed to highlight recent advances in the small noncoding RNA world that could have implications in the development of new strategies in medical sciences. Among the diverse group of small non-coding RNAs, we highlight the group of tRNA-derived fragments as molecules known for a long time which have recently emerged as novel regulators influencing several aspects of cell biology. We describe here recent advances in the field of tRNA-derived fragments playing key roles in the biology of some infectious agents, including E. coli, A. fumigatus, G. lamblia, Ascaris, T. cruzi, Virus, Prions and a brief overview linking them to cancer biology. Additionally, we focus on the potential implications of these molecules in future biotechnological applications in the development of new biomarkers and as new therapeutic targets.
1. INTRODUCTION
In the last years the comprehension of the small noncoding RNAs (sncRNA) world has been growing thanks to the advance in next-generation sequencing technologies and the discovery of RNA interference (RNAi). Effectors functions of these sncRNAs are achieved through small RNA-associated ribonucleoprotein (RNP) complexes. Based on their origin, the proteins to which they are associated, their biogenesis and the biological process in which they participate, sncRNAs have been grouped into different classes, at present the best known are microRNAs (miRNAs), small-interfering RNAs (siRNAs) and PIWI-related RNAs (piRNAs) [1,2] . Additionally, novel members of this family of sncRNAs originated from cleavage or processing of already known ncRNAs such as snRNAs, snoRNAs, tRNAs and rRNAs have recently emerged. The tRNA-derived small RNAs, are originated from tRNAs by enzymatic regulated processes [3-8] . Although tRNA cleavage is not a novelty [9], it was not until recently that they were defined and classified as new members of the sncRNA family.
1.1. Classification and Biogenesis
The classification of each sncRNA molecule inside some family or group is usually established by its biogenesis pathways, structural characteristics and its biological function. In particular, for the small tRNA family, a defined nomenclature for their sub-families or group does not exist yet. This could be due to the fact that they have been recently described and the mechanisms in which they are involved seem to be different from that described for other sncRNAs like miRNAs and siRNAs, making their classification even more difficult.
However, at present it is possible to distinguish two main groups in the small tRNAs family: tRNA halves and tRFs.
1.1.1. tRNA Halves
As depicted in Figure 1, tRNA halves or tsRNAs are those small tRNAs derived from the cleavage of tRNAs at the anticodon loop, generating both 3’ and 5’ fragments of 30 - 35 nt length [6,9,10] . This phenomenon had been described earlier in prokaryotes as consequence of bacteriophage infection [9], but today it is known to take place in a wide range of eukaryotic cells as consequence of a diverse spectra of stress conditions. For this
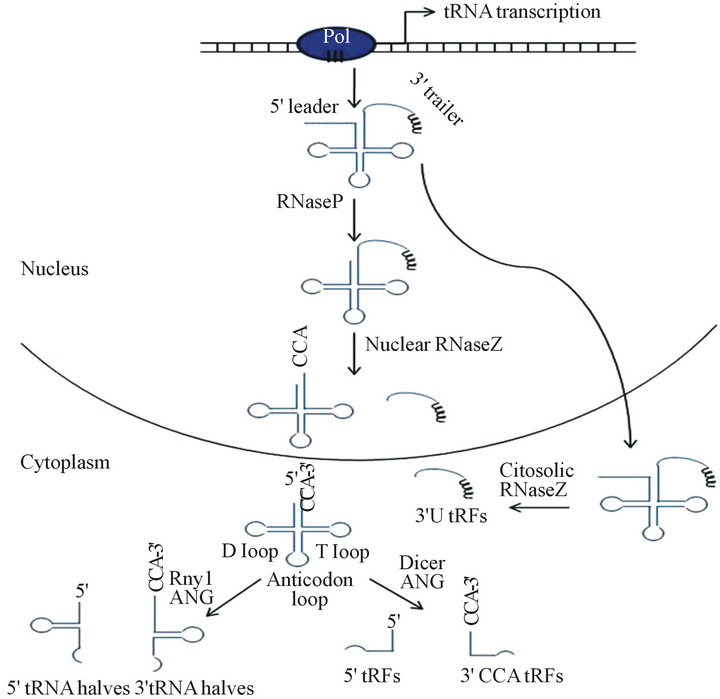
Figure 1. small tRNAs generation pathways from mature tRNAs through the action of anticodon-cleaving enzymes as RNaseZ, Angiogenin, Rny1 or Dicer and today accepted classification and nomenclature: 3’ U tRFs, 3’CCA tRFs, 5’ tRFs and tRNA halves.
reason, in some studies these tRNA fragments are also named tiRNAs for tRNA derived stress-induced fragments [5,7-8,11-14] . This is the case of the first report concerning these small tRNAs found in the ciliate Tetrahymena thermophila as an early adaptive response to starvation caused by essential amino acid deprivation with a possible cell cycle regulation effect [14]. After that, several articles were published describing the cleavage of tRNAs under certain stress conditions, mainly oxidative stress [7,9,12,13,15-17] .
The biogenesis of these molecules is still unclear, the position at the anticodon loop in which tRNAs are cleaved differ among organisms and tRNA species, indicating the involvement of some kind of specific endonuclease according to the organism and tRNA structure [14]. It is known that anticodon nucleases are responsible for tRNA cleavage in bacteria [10] , while Angiogenin and Rny1 cause the stress-induced tRNA cleavage in higher eukaryotes cells and fission yeast respectively [16]. Moreover, cleavage seems to occur also on mature tRNAs, since they usually have mature 5’ ends, and 3’CCA sequence [16]. Even though certain stress stimuli increase tRNA cleavage, in most cases, the pool of full-length tRNAs levels is not affected and the amount of tRNA halves is lower than that of full-length tRNA, indicating that only a small portion of the mature tRNAs pool is involved in this specific and regulated process [9,16] .
1.1.2. tRFs
This class of small tRNAs comprises tRNA fragments derived from both the 5’ and 3’ end of the tRNA, similar to tRNA halves, but in contrast to them, tRFs have a smaller size (~20 nt) as a consequence of different cleavage points in the tRNA molecule. The tRFs originated from 3’ end of immature tRNA molecule or pre-tRNA are also included in this class [3,4,18,19] Figure 1. Even though the three mentioned groups are easily distinguishable, there is no concordance in the literature about tRFs nomenclature. Some authors make reference to the group of tRFs derived of the 5’end through a cleavage in the D loop of mature or immature tRNA as 5’tRFs. Those derived from the cleavage in the T loop of mature tRNA are referred as 3’CCA tRFs (including the 3’end CCA sequence of the mature tRNA), and those derived from the 3’ end of immature or pretRNAs that maintain the polyuridines tract, are referred as 3’ U tRFs, [19]. Other authors like Lee et al., referred to them as tRF-5, tRF-3 and tRF-1 series, respectively and Haussecker et al., called them Type I and Type II groups in reference to 3’ CCA tRFs and 3’ U tRFs [3,4] . As we previously proposed, we will use the terms of 5’tRFs, 3’ CCA tRFs and 3’U tRFs for the three reported classes of tRFs [33].
Ultra-high-throughput sequencing and biological studies made by Lee et al., [4] revealed a precise sequence conservation of the ends of all sequences analyzed, suggesting that a precise cleavage event is necessary to produce each group of small tRFs. For instance, the 3’ CCA tRF cleavage site is preferentially located between A/U and U/A and 5’tRF are preferentially cleaved after A [4]. Despite the fact that biogenesis of tRFs has not been fully elucidated and it varies among organisms, it is known that molecules belonging to the tRNA maturation machinery, like the endonuclease RNAse Z, are responsible for the generation of the 3’U tRFs as well as molecules of the miRNA/siRNA pathways, like Dicer, are involved in 5’ tRFs and 3’CCA tRFs production [3,4,16,18,20,22] . In line with this, some observations were recently made about tRFs biogenesis, on the cleavage of tRNAs and the way in which they are processed in certain kind of cells:
• Given the absence of 5’ leader sequences in the 5’tRFs as consequence of the RNAse P cleavage in the early process of tRNA maturation, they could be generated at any point of later tRNA processing [3].
• As it was described, their biogenesis could be carried out by Dicer in mammalian cells, and in a Dicerindependent way as was described in Schizosaccharomyces pombe. The differences among 5’ tRFs generated in these two organisms, mainly in their size, support the idea that an enzyme other than Dicer is responsible for their production in yeast [19].
• The generation of 3’CAA tRFs also seems to be Dicer-dependent, and it should take place after the cleavage by both RNAse P and tRNAse Z, and the addition of CCA; considering that they include the CCA sequence at their 3’ ends [3].
• As it was mentioned, 3’ U tRFs are formed by the action of tRNAse Z from the pre-tRNA. This process occurs in the nucleus, where tRNA maturation takes place and then tRNA is transported to the cytoplasm. Since 3’ U tRFs are found almost exclusively in the cytoplasm it means that after their generation they need to be exported from the nucleus [3,18] or a cytoplasmic pool of tRNAse Z could be responsible for the generation of 3’ U tRFs from tRNAs that exited the nucleus prematurely [23] . Besides these observations, Li et al. have recently described a variety of 5’and 3’tRNA fragments of ~20 nt length Dicer-independent [24]. These results suggest that tRFs are produced by the Angiogenin cleavage (which was mentioned to be involved in the generation of tRNA halves) in human and mouse cells [20] .
1.2. Biological Functions
The descriptions mentioned above regarding the regulated biogenesis process of small tRNAs, the stimuli that induce their intracellular increase, and the fact that they are associated with proteins belonging to specific cellular machineries, encouraged authors to suggest that these tRNAs species should have some specific functions in diverse biological process. In this respect, several published works associate small tRNAs with cellular process as translational regulation, cell cycle and cellular proliferation, cell differentiation and virulence Table 1. These functions could be carried out through either canonical or non-canonical miRNA/siRNA pathways considering that small tRNAs molecules were incorporated to Argonaute proteins or not.
Currently, no specific targets and/or biological effects have been determined in most studies performed. Some of the most discussed articles in this respect are the ones published by Haussecker et al. and Lee et al. [3,4] . Lee et al. described a possible influence of one of this tRFs (called tRF-1001) on cell proliferation in mammalian cells through a mechanism different from miRNA/siRNA. Haussecker et al. performed experiments with a type I tRF (called cand45, which is the same tRF that Lee et al. called tRF-1001) that showed a silencing activity termed sense-induced trans silencing (SITS) in a similar manner to miRNA/siRNA ADDIN EN.CITE ADDIN EN.CITE.DATA [3,4] . In agreement with the argument made by Pederson about Haussecker’s results, the relevance of that work relies on the non canonical miRNA/ siRNA mode of action of these tRFs molecules [8]. Ivanov et al. [21] have reported that tRNA halves contribute to the dissociation of different forms of the translation initiation factor eIF4G from capped and uncapped mRNA. In addition, they proposed that tRNA halves cooperate with a translational repressor known to displace eIF4G from RNA to negatively regulate translation in stressed cells.
The search of alternative sncRNA pathways has been the focus of different works that found the presence of small tRNAs in organisms like Trypanosoma cruzi, for which is well known the lack of canonical RNAi machinery [12,25] . These findings showed these tRNAs species are interesting players of diverse cellular proesses which involve novel modes of action, independent of their association or not with RNAi mechanisms.
Moreover, if we take into account the list of organisms in which both tRNA-halves and tRFs have been found Table 1, we can note that they are present throughout the evolutionary scale and more, since they have been associated to exosomes content from prion-infected neuronal cells [26]. In each cell type, they have their specific and novel modes of action, even though they could share
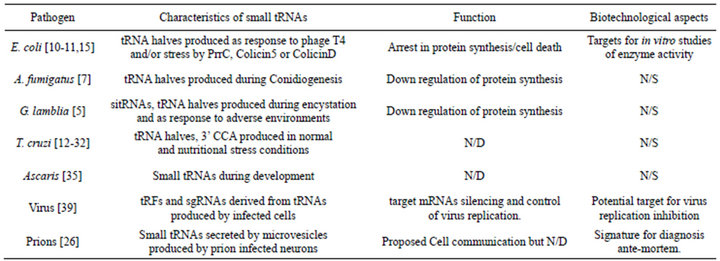
Table 1. Summary of main descriptions made up to date of tRNA cleavage phenomena in potential human pathogen organisms and its relation with future biotechnological implications. N/D: Not determined; N/S: Not Studied.
similar biological activities.
2. SMALL TRNAS AND INFECTIOUS DISEASES
Recent advances in sequencing technologies and bioinformatics applied to causative agents of human diseases have stimulated several groups to study the putative role of small tRNAs in a wide range of human pathogens. In a recent review [33] we have highlighted the relevance of these molecules in RNA interference phenomena; here we focus in the description of some examples of pathogens that produce small tRNAs under different conditions (Table 1). A deeper knowledge of these novel molecules would give us new tools to manage and to cope with unsolved issues in the pathogenesis of several human infectious diseases.
2.1. Escherichia coli
E. coli is a Gram-negative bacterium that is commonly found in the lower intestine of different organisms. Most E. coli strains are harmless, but some strains can cause serious food poisoning in humans, they can cause diarrhea, while others cause urinary tract infections, respiratory illness and pneumonia among other pathological states. E. coli infections are globally increasing in number and severity. It is estimated that this year there will be approximately 76 million illnesses and 5,200 deaths in the United States only attributed to food poisoning outbreaks caused by E. coli O157:H7 and other strains with a mean cost of 50 billion US dollars [27]. In 1990 Levitz and coworkers [11] presented one of the first evidences of specific tRNA cleavage in E. coli by the endoribonuclease Prrc in response to phage T4 infection. In this study the authors adventure the possibility that this tRNA processing pathways could be viewed as a host response to phage division. Later, in 1999, Ogawa et al. [10] found other enzyme, Colicin E5, a special ribonuclease, responsible for the processing of 3’ half fragments from tRNAHis, tRNAAsn, and tRNAAsp. They demonstrated that the functional role for this RNase is to inhibit protein synthesis of E. coli as a defensive mechanism to cope with phage infection. This bacterial suicide mechanism could be exploited as a tool to eliminate bacterial replication during host infection. A variety of molecules that inhibit protein synthesis have been used longtime to elucidate complicated cell mechanisms and as antibacterial agents. In this respect, it is important to determine whether the activity of Colicin E5 toward the tRNAs reflects the physiological action of this RNase leading to cell death.
2.2. Aspargillus fumigatus
A. fumigatus is a filamentous fungi and facultative pathogen for plants and animals with special risk for immunosuppressed patients. The aspergillosis is considered the most common airborne fungal pathogen. The mortality rate of invasive Aspergillosis could reach 90% of cases, depending on the immune status of the host, due to the difficulty of specific diagnostics and therapeutic possibilities [28]. In 2008, Jochl et al. [7] reported the cleavage of tRNA molecules during cell differentiation in A. fumigatus. They didn’t find cleavage specificity for certain tRNAs, since all nuclear encoded tRNAs except cytoplasmic tRNAMet, tRNATrp and tRNAAsp were identified in the small transcriptome of this fungi. These small tRNA (tRNA halves) were generated by an endonucleolytic cleavage of tRNAs at a position 3’ adjacent to the anti-codon loop. The nuclease responsible for this cleavage is unknown but the possible biological function for this process is to downregulate protein synthesis during differentiation [7] . Since conidia represent the resting state of A. fumigatus in the host it is possible that conidial tRNA depletion might be a novel mechanism to down-regulate protein synthesis in this fungus inhibiting the evolution of the disease.
2.3. Giardia lamblia
At the same time of the identification of tRNA fragments in A. fumigatus, Li et al. [29] described a particular class of small tRNAs in G. lamblia, named stress-induced tRNA-derived RNA (sitRNAs). The authors revealed a global tRNA cleavage at the anticodon left arm in response to encystations and under different conditions of physicochemical stress. G. lamblia is considered a successful unicellular intestinal parasite due to its ability to reproduce in different and hostile environments by differentiation from a vegetative trophozoite to a cyst form, being a common causative agent of diarrhea and malnutrition [30]. The sitRNAs of G. lamblia are different from those previously described for other organisms since they are a little longer, being about 46 nt in length, and derived from the 3’ end of mature tRNAs with a significant fraction having 3’ terminal CCA nucleotides. These differences serve as clues for the elucidation of biogenesis pathways for these sitRNAs. Even though the authors didn’t find any enzyme responsible for this cleavage in this work, they argue that the main consequence of this phenomenon is down regulation of global gene expression in the parasite [29]. These results may provide new insight into a novel mechanism for G. lamblia to regulate gene expression therefore is a possible new therapeutic target for control of parasite replication in the host.
2.4. Trypanosoma cruzi
At present, the families of small no-coding regulatory RNAs (i.e. miRNAs and siRNAs) are recognized as key players in post-transcriptional gene regulation in most eukaryotes. Nonetheless, T. cruzi lacks canonical sRNA pathways [31]. Furthermore, this protozoan has a complex life cycle including diverse cellular forms which alternate between invertebrate and mammalian hosts. To cope with these changes T. cruzi undergoes rapid modifications in gene expression which are achieved essentially at a post-transcriptional level. However the precise mechanisms of this fine tune regulation remain to be elucidated. In a recent effort of our group aimed to identify the presence of alternate sncRNA pathways in T. cruzi clone Dm28c, we reported the presence of a homogeneous population of tRNA halves. In addition we showed that this class of small tRNAs is recruited to particular cytoplasmic organelles in stressed T. cruzi epimastigotes. The main classes of small tRNAs found for T. cruzi were 5’ halves of three specific tRNAs [12]. A year later Franzen et al. performed high throughput sequencing of T. cruzi CL Brener epimastigotes in optimal growth conditions and showed that tRNA halves were originated predominantly from the 3’ arms and not restricted to particular tRNAs [32]. Despite the wide confirmation for the expression of these small tRNAs in T. cruzi, its biological role remains uncertain. Recent unpublished data from our laboratory showed that small tRNAs are being released as vesicle cargo shed by T. cruzi. Additionally, this cargo is being transferred among parasites and from parasite to host cells, leading to metaciclogenesis transformation and to an increased susceptibility to infection (Submitted manuscript). These results drive us to propose these molecules as possible virulence factors which could be potential drug targets for the control of the disease.
2.5. Ascaris
As we have discussed previously [33] it was not until 2010 that the use of deep sequencing techniques allowed researchers to systematically find small tRNAs molecules in eukaryotic pathogens. That is the case of A. suum, a nematode parasite closely directed to A. lumbricoides. A. lumbricoides affects 1 billion people worldwide and is considered an important neglected disease [34] . Wang et al. in 2011 performed deep small RNA transcriptome sequencing of this parasite and identified a diversity of sRNA classes including RNAs of 32 nt with a 5’ monophosphate. Many of these sRNAs were derived from snoRNAs and tRNAs, among others. They verified the presence of small tRNAs in the small transcriptome of Ascaris but these molecules were not the focus of this study. The work of these authors sheds a vast amount of data that needs to be analyzed deeper in respect to small tRNAs, for example to systematically catalog the expression of small tRNAs through male and female reproducetive systems, during zygote maturation, and all stages of development [8] . Although there is no direct evidence relating these molecules to Ascaris pathogenesis, they should not be dismissed considering their potential relevance in related infectious agents.
2.6. Virus: HIV, Respiratory Syncytial Virus and Herpes Simplex Virus
The etiology and worldwide pandemic significance of viral diseases such as AIDS and human respiratory Syncytial virus (RSV) (the most common cause of bronchiolitis and pneumonia in babies) go beyond the scope of this review. We intended simply to highlight the most relevant knowledge on small tRNAs that would allow future biomedical applications. It is known today that viral reverse transcriptases use tRNAs (commonly tRNALys, tRNAPro, or tRNATrp) as primers for the initiation of reverse transcription and DNA viral synthesis [36]. Yeung et al. [37] have previously reported the identification of a 18 nt small RNA, originated from the double-stranded RNA hybrid formed by the HIV RNA and the 3’ end of the human cellular tRNALys. The authors used luciferase reporter assays to show silencing activity produced by the union of this tRNALys, considered as a 3’tRF, with Dicer and AGO2. These results allowed the authors to propose a function for tRFs in RNAi machinery, targeting HIV and viral defense. We would like to point here a finding by Reese et al. [38] in 2010, not specific for tRFs but for tRNA molecules of interest for this review. In this study the authors reported that mammalian herpes viruses could exploit tRNA biogenesis pathways to produce fully functional viral miRNAs. In this system, molecules of 50 nts derived from tRNAs produced by RNA Pol III are aligned with a pre-miRNA hairpin of 30 nts and cleaved by RNase Z to liberate the pre-miRNA hairpins. These pre-miRNAs are further processed to mature viral miRNAs by Dicer [38]. This is an original report in which tRNA molecules different of tRFs could act as gene control factors. Recently Wang et al. [39] have used high throughput sequencing to identify sncRNAs in cells infected with RSV. They found that RSV infection generates a large production of 5’-derived tRFs of about 30 nts long and the processing of this tRFs appeared to be specific of the endonuclease Angiogenin. Moreover at least for tRFs from tRNA-Glu-CTC they showed a possible role suppressing target mRNA and supporting RSV replication. We could conclude that this work provides original information on the possibility of future uses for tRFs as inhibitors of viral infections.
2.7. Prions
In the last few years, numerous reports have highlighted the role of exosomes in diverse biological processes such as antigen presentation, immune response, cell to cell communication and burden of infectious pathogens [40]. In addition, exosomes carry as cargo mRNA and miRNAs that could be delivered to other cells and be totally functional in receptor cells [41]. Prion diseases constitute a group of transmissible neurodegenerative disorders, invariably fatal, that include bovine spongiform encephalopathy among others, where the main symptoms are dementia, neuronal loss, spongiform change and death. One model of prion propagation proposes that these diseases are associated with the conformational conversion of the host cellular prion protein into an abnormal pathogenic isoform by a protein guide directed mechanism. Diagnosis and treatment of prion disease is almost impracticable by the absence of effecttive pre-mortem diagnostic methods. The identification of non-invasive, sensitive and specific diagnostic markers during the pre-clinical phase is of major importance [42]. Recently, Bellingham et al. [26] analyzed the small RNA cargo of uninfected and prion-infected exosomes by deep sequencing. Surprisingly, these authors demonstrated that exosomes released from prion-infected neuronal cells contain a variety of small RNA where 90 % corresponded to small tRNAs. Additionally, they reported a specific miRNA signature in exosomes released from prion-infected neuronal cells. They discuss the possible role of exosomes in miRNAs delivery during prion infection disease that could be of interest for diagnostics [26].
2.8. Cancer
We are not going to dwell here since it is out of the scope of this review, but a description of small tRNAs implications in human pathogenesis would be incomplete without pointing out the advances in the field of cancer biology. The relation of small tRNAs and cancer are described deeper in a recent review by Martens-Uzunova et al. [43]. The authors compile evidence since the first isolation of a small tRNA molecule almost 30 years ago from human urinary bladder carcinoma to nowadays with data provided by next generation sequencing. These data led to the identification of small tRNAs of different size, class and origin on a genome wide scale. Studies of Cole et al. [6] and Kawaji et al. [44], revealed that small tRNAs are one of the main populations of sRNAs found in cancer cell lines. These authors provided experimental evidences showing different expression patterns of these molecules between normal and cancer cells. These findings are closing to complete the molecular overview of this disease.
3. BIOTECHNOLOGICAL ASPECTS OF SMALL TRNAS
Similarly to miRNAs and siRNAs, small tRNAs are becoming as valuable research and biotechnological tools in cell biology and medicine.
3.1. Microarrays
Saikia and co-workers [45] have made use of microarray technology to evaluate patterns of tRNA cleavage in response to different stress sources in murine cells. They designed customized microarray chips containing sequences of mouse nuclear-encoded tRNAs, as well as other yeast and Escherichia coli tRNA probes for hybridization and normalization controls. To ensure that only tRNA fragments were detected and to avoid interference from mature tRNAs, total RNA from treated cells was size-fractioned (retaining only RNAs 30 - 40 nt in length) and stringent hybridization conditions were applied. This approach allowed them to identify cleavage patterns characteristic of certain types of stress and to further understand the processes underlying tRNA fragments generation. Such genome-wide strategies could provide insights into the role of tRNA fragments in pathways involved in numerous disorders and conditions if applied adequately.
3.2. Gene Silencing
A remarkable example of the biotechnological application of tRFs is that of the TRUE gene silencing technology, developed by Nashimoto and collaborators. TRUE gene silencing stands for tRNase ZL-utilizing efficacious gene silencing [23,46-47] . tRNaseZL is the long form of tRNaseZ, the enzyme responsible for removing the 3’ trailer from precursor tRNA (pre-tRNA), a key step in tRNAs maturation. tRNaseZ has also been implied in the generation of certain classes of tRFs [19]. In a series of studies, Nashimoto’s group demonstrated the ability of tRNaseZL to cleave any target RNA at any desired site by recognizing a pre-tRNA-like or micropre-tRNA-like complex formed between the target RNA and an artificially-designed small guide RNA (sgRNA). They describe several classes of sgRNA, including 5’-tRNA-halves.
In their initial work, they report that 5’-tRNA-halves were able to act as sgRNAs to cleave at specific sites two HIV mRNAs (ENV and GAG) [48]. Ten years later, they were able to downregulate the expression of two endogenous human genes (Bcl-2 and GSK-3β), using tRNaseZL and sgRNAs, and propose the term TRUE gene silencing for this technology [47] . Later on, they were successful at inhibiting the endogenous human vascular endothelial growth factor (VEGF) gene expression using this methodology, with tRNA halves (and others) as sgRNAs ADDIN EN.CITE ADDIN EN.CITE.DATA [23] . Finally, they showed that miRNAs can also act as sgRNAs, directing the cleavage of target mRNAs by tRNaseZL in human cells ADDIN EN.CITE ADDIN EN.CITE.DATA [23] . In the last years they expanded the possibilities of TRUE gene silencing by demonstrating that a certain kind of sgRNAs can readily be incorporated in human cells and efficiently eliminate target miRNAs [49].
Interestingly, the silencing effect using sgRNAs greatly surpassed that of siRNAs in some cases, both for endogenous genes and in luciferase assays. The authors also suggested that this technology should be safer than siRNAs, since potential off-target effects of sgRNAs are expected to be less serious than those induced by siRNAs because sgRNAs target a smaller region in the mRNA than siRNAs. These are two desirable characteristics of potential therapeutic agents for several diseases.
In line with these findings, Scherer et al., in search for more efficient and controlled short hairpin RNA (shRNA) expression systems, designed a tRNA-shRNA chimera expression construct under the control of tRNA promoters. Their results indicate that tRNaseZL cleaves the tRNA-shRNA chimera, since these chimeras resemble the normal substrate of the enzyme, therefore releasing the hairpin which is then incorporated in the RNA interference machinery, regulating the expression of the targeted genes [50] .
3.3. Biomarker Potential
Much remains to be understood regarding tRNA fragments’ role in diseases. However, growing evidence indicates that they are implicated in a range of disorders, including cancer [51], inflammatory diseases [52] and viral infection [39]. As was the case of miRNAs, research in the field of tRNA fragments is likely to increase rapidly, and information about their involvement in certain diseases will soon become available. Interestingly, small tRNAs patterns in cancer cell lines were reported profoundly altered when compared to normal counterparts [6,44] . Additionally, a line of active current research is the study of circulating small RNAs in serum, plasma, and other body fluids, contained in extracellular vesicles, which may represent an interesting niche for the search of noninvasive biomarkers in cancer and other complex diseases [53,54] . Certain tRNA fragments have been identified in \microvesicles from immune cells [55] and prion-infected neuronal cells [26], and further research in this direction should provide useful tools for diagnosis.
4. CONCLUDING REMARKS
The complete understanding of a sncRNA scenario in cell biology will provide valuable information for their future applications in biomedicine especially in the field of biomarker development and new therapeutic approaches. One of these promising molecules are small tRNAs which were intensively studied in the last decade as specific products of regulated pathways and not merely results of nonspecific degradative processes. This review was aimed to point out the relevance of these findings for some pathogens and diseases that have been done in this area.
We presented as examples the findings achieved in tRFs field for: Bacteria as E. coli [10,11,15] , Fungus as A. fumigatus ADDIN EN.CITE ADDIN EN.CITE.DATA [7] , Protozoas as G. lamblia [29] and parasites as T. cruzi [12-32] and Ascaris ADDIN EN.CITE ADDIN EN.CITE.DATA [35] ; Viruses as HIV [37,46,48,56] , Herpes [38] and RSV [39]; and neurodegenerative agents as Prions [26]; last we wanted to briefly note, not in depth, the case for cancer [6,44] . Briefly all the authors observe the production of these small tRNAs under different stimuli that could be stress, cell cycle progression and/or infection by a pathogen agent. The consequences or biological roles of the expression of these molecules is not completely defined but could lead cells to down-regulate protein synthesis and mRNA silencing, as the cases for small tRNAs of G. lamblia [29], A. fumigatus ADDIN EN.CITE ADDIN EN.CITE.DATA [7] and virus [39].
Efforts should be focused on the complete description of small tRNAs and its role as virulence factors, drug targets, biomarkers or as relevant molecules for infection.
With further research effort it is likely that technologies based on small tRNAs will develop and more applications will arise. It is of particular interest their use as non-invasive biomarkers for the diagnosis of certain diseases and as sgRNAs in gene therapeutic approaches.
REFERENCES
- Farazi, T.A., Juranek, S.A. and Tuschl, T. (2008) The growing catalogue of small RNAs and their association with distinct Argonaute/Piwi family members. Development, 135, 1201-1214.
- Rother, S. and Meister, G. (2011) Small RNAs derived from longer non-coding RNAs. Biochimie, 93, 1905- 1915.
- Haussecker, D., Huang, Y., Lau, A., Parameswaran, P., Fire, A.Z. and Kay, M.A. (2010) Human tRNA-derived small RNAs in the global regulation of RNA silencing. RNA, 16, 673-695. doi:10.1261/rna.2000810
- Lee, Y.S., Shibata, Y., Malhotra, A. and Dutta, A. (2009) A novel class of small RNAs: tRNA-derived RNA fragments (tRFs). Genes & Development, 23, 2639-2649
- Babiarz, J.E., Ruby, J.G. Wang, Y. Bartel, D.P. and Blelloch, R. (2008) Mouse ES cells express endogenous shRNAs, siRNAs, and other Microprocessor-independent, Dicer-dependent small RNAs. Genes & Development, 22, 2773-2785.
- Cole, C., Sobala, A., Lu, C., Thatcher, S.R., Bowman, A., Brown, J.W., Green, P.J., Barton, G.J. and Hutvagner, G. (2009) Filtering of deep sequencing data reveals the existence of abundant Dicer-dependent small RNAs derived from tRNAs. RNA, 15, 2147-2160.
- Jochl, C., Rederstorff, M., Hertel, J., Stadler, P.F., Hofacker, I.L., Schrettl, M., Haas, H. and Huttenhofer, A. (2008) Small ncRNA transcriptome analysis from Aspergillus fumigatus suggests a novel mechanism for regulation of protein synthesis. Nucleic Acids Research, 36, 2677-2689. doi:10.1093/nar/gkn123
- Pederson, T. (2010) Regulatory RNAs derived from transfer RNA? RNA, 16, 1865-1869. doi:10.1261/rna.2266510
- Thompson, D.M. and Parker, R. (2009) Stressing out over tRNA cleavage. Cell, 138, 215-219.
- Ogawa, T., Tomita, K., Ueda, T., Watanabe, K., Uozumi, T. and Masaki, H. (1999) A cytotoxic ribonuclease targeting specific transfer RNA anticodons. Science, 283, 2097-2100. doi:10.1126/science.283.5410.2097
- Levitz, R., Chapman, D., Amitsur, M., Green, R., Snyder, L. and Kaufmann, G. (1990) The optional E. coli prr locus encodes a latent form of phage T4-induced anticodon nuclease. The EMBO Journal, 9, 1383-1389.
- Garcia-Silva, M.R., Frugier, M., Tosar, J.P., CorreaDominguez, A., Ronalte-Alves, L., Parodi-Talice, A., Rovira, C., Robello, C., Goldenberg, S. and Cayota, A. (2010) A population of tRNA-derived small RNAs is actively produced in Trypanosoma cruzi and recruited to specific cytoplasmic granules. Molecular and Biochemical Parasitology, 171, 64-73. doi:10.1016/j.molbiopara.2010.02.003
- Haiser, H.J., Karginov, F.V., Hannon, G.J. and Elliot, M.A. (2008) Developmentally regulated cleavage of tRNAs in the bacterium Streptomyces coelicolor. Nucleic Acids Research, 36, 732-741. doi:10.1093/nar/gkm1096
- Lee, S.R. and Collins, K. (2005) Starvation-induced cleavage of the tRNA anticodon loop in Tetrahymena thermophila. The Journal of Biological Chemistry, 280, 42744-42749. doi:10.1074/jbc.M510356200
- Tomita, K., Ogawa, T., Uozumi, T., Watanabe, K. and Masaki, H. (2000) A cytotoxic ribonuclease which specifically cleaves four isoaccepting arginine tRNAs at their anticodon loops. Proceedings of the National Academy of Sciences of the United State of America, 97, 8278-8283. doi:10.1073/pnas.140213797
- Thompson, D.M., Lu, C., Green, P.J. and Parker, R. (2008) tRNA cleavage is a conserved response to oxidative stress in eukaryotes. RNA, 14, 2095-2103. doi:10.1261/rna.1232808
- Zhang, S., Sun, L. and Kragler, F. (2009) The phloemdelivered RNA pool contains small noncoding RNAs and interferes with translation. Plant Physiology, 150, 378- 387.
- Liao, J.Y., Ma, L.M., Guo, Y.H., Zhang, Y.C., Zhou, H., Shao, P., Chen, Y.Q. and Qu, L.H. (2010) Deep sequencing of human nuclear and cytoplasmic small RNAs reveals an unexpectedly complex subcellular distribution of miRNAs and tRNA 3’ trailers. PLoS One, 5, e10563.
- Sobala, A. and Hutvagner, G. (2011) Transfer RNAderived fragments: Origins, processing, and functions. Wiley Interdisciplinary Reviews: RNA, 2, 853-862.
- Emara, M.M., Ivanov, P., Hickman, T., Dawra, N., Tisdale, S., Kedersha, N., Hu, G.F. and Anderson, P. (2010) Angiogenin-induced tRNA-derived stress-induced RNAs promote stress-induced stress granule assembly. The Journal of Biological Chemistry, 285, 10959-10968. doi:10.1074/jbc.M109.077560
- Ivanov, P., Emara, M.M., Villen, J., Gygi, S.P. and Anderson, P. (2011) Angiogenin-induced tRNA fragments inhibit translation initiation. Molecular Cell, 43, 613-623.
- Yamasaki, S., P. Ivanov, G.F. Hu, and P. Anderson, (2009) Angiogenin cleaves tRNA and promotes stressinduced translational repression. The Journal of Cell Biology, 185, 35-42. doi:10.1083/jcb.200811106
- Elbarbary, R.A., Takaku, H., Tamura, M. and Nashimoto, M. (2009) Inhibition of vascular endothelial growth factor expression by TRUE gene silencing. Biochemical and Biophysical Research Communications, 379, 924-927.
- Li, Z., Ender, C., Meister, G., Moore, P.S., Chang, Y., and John, B. (2012) Extensive terminal and asymmetric processing of small RNAs from rRNAs, snoRNAs, snRNAs, and tRNAs. Nucleic Acids Research, 40, 6787- 6799.
- Garcia Silva, M.R., Tosar, J.P., Frugier, M., Pantano, S., Bonilla, B., Esteban, L., Serra, E., Rovira, C., Robello, C. and Cayota, A. (2010) Cloning, characterization and subcellular localization of a Trypanosoma cruzi argonaute protein defining a new subfamily distinctive of trypanosomatids. Gene, 466, 26-35. doi:10.1016/j.gene.2010.06.012
- Bellingham, S.A., Coleman, B.M. and Hill, A.F. (2012) Small RNA deep sequencing reveals a distinct miRNA signature released in exosomes from prion-infected neuronal cells. Nucleic Acids Research, 40, 10937-10949.
- Bavaro, M.F. (2012) E. coli O157:H7 and other toxigenic strains: The curse of global food distribution. Current Gastroenterology Reports, 14, 317-323.
- Latge, J.P. (1999) Aspergillus fumigatus and aspergillosis. Clinical Microbiology Reviews, 12, 310-350.
- Li, Y., Luo, J., Zhou, H., Liao, J.Y., Ma, L.M., Chen, Y.Q. and Qu, L.H. (2008) Stress-induced tRNA-derived RNAs: A novel class of small RNAs in the primitive eukaryote Giardia lamblia. Nucleic Acids Research, 36, 6048-6055.
- Adam, R.D. (2001) Biology of Giardia lamblia. Clinical Microbiology Reviews, 14, 447-475.
- Ullu, E., Tschudi, C. and Chakraborty, T. (2004) RNA interference in protozoan parasites. Cellular Microbiology, 6, 509-519.
- Franzen, O., Arner, E., Ferella, M., Nilsson, D., Respuela, P., Carninci, P., Hayashizaki, Y., Aslund, L., Andersson, B. and Daub, C.O. (2011) The short non-coding transcriptome of the protozoan parasite Trypanosoma cruzi. PLOS Neglected Tropical Diseases, 5, e1283. doi:10.1371/journal.pntd.0001283
- Garcia-Silva, M.R., Cabrera-Cabrera, F., Güida, M.C., and Cayota, A. (2012) Hints of tRNA-Derived Small RNAs Role in RNA Silencing Mechanisms. Genes, 3, 603-614. doi:10.3390/genes3040603
- Bethony, J., Brooker, S., Albonico, M., Geiger, S.M., Loukas, A., Diemert, D. and Hotez, P.J. (2006) Soiltransmitted helminth infections: Ascariasis, trichuriasis, and hookworm. The Lancet, 367, 1521-1532.
- Wang, J., Czech, B., Crunk, A., Wallace, A., Mitreva, M., Hannon, G.J. and Davis, R.E. (2011) Deep small RNA sequencing from the nematode Ascaris reveals conservation, functional diversification, and novel developmental profiles. Genome Research, 21, 1462-1477.
- Marquet, R., Isel, C., Ehresmann, C. and Ehresmann, B. (1995) tRNAs as primer of reverse transcriptases. Biochimie, 77, 113-124. doi:10.1016/0300-9084(96)88114-4
- Yeung, M.L., Bennasser, Y., Watashi, K., Le, S.Y., Houzet, L. and Jeang, K.T. (2009) Pyrosequencing of small non-coding RNAs in HIV-1 infected cells: Evidence for the processing of a viral-cellular doublestranded RNA hybrid. Nucleic Acids Research, 37, 6575- 6586. doi:10.1093/nar/gkp707
- Reese, T.A., Xia, J., Johnson, L.S., Zhou, X., Zhang, W. and Virgin, H.W. (2010) Identification of novel microRNA-like molecules generated from herpesvirus and host tRNA transcripts. Journal of Virology, 84, 10344-10353. doi:10.1128/JVI.00707-10
- Wang, Q., Lee, I., Ren, J., Ajay, S.S., Lee, Y.S. and Bao, X. (2013) Identification and functional characterization of tRNA-derived RNA fragments (tRFs) in respiratory syncytial virus infection. Molecular Therapy, 21, 368- 379. doi:10.1038/mt.2012.237
- Thery, C., Zitvogel, L. and Amigorena, S. (2002) Exosomes: Composition, biogenesis and function. Nature Reviews Immunology, 2, 569-579.
- Valadi, H., Ekstrom, K., Bossios, A., Sjostrand, M., Lee, J.J. and Lotvall, J.O. (2007) Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nature Cell Biology, 9, 654-659. doi:10.1038/ncb1596
- Aguzzi, A. and Heikenwalder, M. (2006) Pathogenesis of prion diseases: Current status and future outlook. Nature Reviews Microbiology, 4, 765-775. doi:10.1038/nrmicro1492
- Martens-Uzunova, E.S., Olvedy, M. and Jenster, G. (2013) Beyond microRNA—Novel RNAs derived from small non-coding RNA and their implication in cancer. Cancer Letters, (in Press). doi:10.1016/j.canlet.2012.11.058
- Kawaji, H., Nakamura, M., Takahashi, Y., Sandelin, A., Katayama, S., Fukuda, S., Daub, C.O., Kai, C., Kawai, J., Yasuda, J., Carninci, P. and Hayashizaki, Y. (2008) Hidden layers of human small RNAs. BMC Genomics, 9, 157. doi:10.1186/1471-2164-9-157
- Saikia, M., Krokowski, D., Guan, B.J., Ivanov, P., Parisien, M., Hu, G.F., Anderson, P., Pan, T. and Hatzoglou, M. (2012) Genome-wide identification and quantitative analysis of cleaved tRNA fragments induced by cellular stress. The Journal of Biological Chemistry, 287, 42708- 42725. doi:10.1074/jbc.M112.371799
- Habu, Y., Miyano-Kurosaki, N., Kitano, M., Endo, Y., Yukita, M., Ohira, S., Takaku, H. and Nashimoto M. (2005) Inhibition of HIV-1 gene expression by retroviral vector-mediated small-guide RNAs that direct specific RNA cleavage by tRNase ZL. Nucleic Acids Research, 33, 235-243. doi:10.1093/nar/gki164
- Nakashima, A., Takaku, H., Shibata, H.S., Negishi, Y., Takagi, M., Tamura, M. and Nashimoto, M. (2007) Gene silencing by the tRNA maturase tRNase ZL under the direction of small-guide RNA. Gene Therapy, 14, 78-85. doi:10.1038/sj.gt.3302841
- Nashimoto, M. (1996) Specific cleavage of target RNAs from HIV-1 with 5’ half tRNA by mammalian tRNA 3’ processing endoribonuclease. RNA, 2, 523-524.
- Takahashi, M., Elbarbary, R.A., Abe, M., Sato, M., Yoshida, T., Yamada, Y., Tamura, M. and Nashimoto, M. (2012) Elimination of specific miRNAs by naked 14-nt sgRNAs. PLoS ONE, 7, e38496. doi:10.1371/journal.pone.0038496
- Scherer, L.J., Frank, R. and Rossi, J.J. (2007) Optimization and characterization of tRNA-shRNA expression constructs. Nucleic Acids Research, 35, 2620-2628. doi:10.1093/nar/gkm103
- Maute, R.L., Schneider, C., Sumazin, P., Holmes, A., Califano, A., Basso, K. and Dalla-Favera, R. (2013) tRNA-derived microRNA modulates proliferation and the DNA damage response and is down-regulated in B cell lymphoma. Proceedings of the National Academy of Sciences of the United State of America, 110, 1404-1409.
- Xia, J., Joyce, C.E., Bowcock, A.M. and Zhang, W. (2013) Noncanonical microRNAs and endogenous siRNAs in normal and psoriatic human skin. Human Molecular Genetics, 22, 737-748. doi:10.1093/hmg/dds481
- Cortez, M.A., Bueso-Ramos, C., Ferdin, J., Lopez-Berestein, G., Sood, A.K. and Calin, G.A. (2011) MicroRNAs in body fluids—The mix of hormones and biomarkers. Nature Reviews Clinical Oncology, 8, 467-477. doi:10.1038/nrclinonc.2011.76
- Shah, M.Y. and Calin, G.A. (2013) The mix of two worlds: Non-coding RNAs and hormones. Nucleic Acid Therapeutics, 23, 2-8.
- Nolte-’t Hoen, E.N.M., Buermans, H.P.J., Waasdorp, M., Stoorvogel, W., Wauben, M.H.M. and ’t Hoen, P.A.C. (2012) Deep sequencing of RNA from immune cell-derived vesicles uncovers the selective incorporation of small non-coding RNA biotypes with potential regulatory functions. Nucleic Acids Research, 40, 9272-9285.
- Nashimoto, M. and Kaspar, R. (1997) Specific cleavage of a target RNA from HIV-1 by mammalian tRNA 3’ processing endoribonuclease directed by an RNA heptamer. Nucleic Acids Symposium Series, 36, 22-25. doi:10.1093/nar/gks658
NOTES
*These authors contributed equally to this work.
#Corresponding author.

