American Journal of Plant Sciences
Vol.3 No.2(2012), Article ID:17537,5 pages DOI:10.4236/ajps.2012.32022
Meiotic Studies in Withania somnifera (L.) Dunal.: A Threatened Medicinal Herb of Indian Thar Desert
![]()
1Cytogenetics and Molecular Biology Laboratory, Department of Botany, Jai Narain Vyas University, Jodhpur, India; 2Directorate of Rapeseed-Mustard Research, Indian Council of Agriculture Research, Bharatpur, India; 3Department of Biotechnology and Bioinformatics, North-Eastern Hill University, Shillong, India.
Email: *srrao22@yahoo.com, srrao@mail.com
Received July 20th, 2011; revised August 25th, 2011; accepted October 11th, 2011
Keywords: Withania; Medicinal; Meiosis; Chromosome Behavior
ABSTRACT
Withania somnifera recently categorized as threatened plant of Rajasthan and known for its high medicinal value has been studied for existing genetic variations in four accessions collected from four different locations. From the present studies it is apparent that the gametic number of W. somnifera is n = 24 and 2n = 48. From the combined information generated by present author and earlier workers, it is tentatively believed that W. somnifera might be of allopolyploid origin with basic number of x = 12. In this case also precocious separation of rod bivalents but not structural hetrozygosity might be the reason for occurrence of univalents. From the detailed meiotic analysis, it is concluded that the species diversification is brought out by polyploidy in W. somnifera. Multidisciplinary approach for genome diagnostics with use of various marker systems might through more light on existing intra-specific genetic variation in W. somnifera.
1. Introduction
Withania somnifera (L.) Dunal. belonging to family Solanaceae and commonly known as “Ashwagandha” or “Asgandh” is considered a wonder herb having assorted Ayurvedic, Unani and indigenous medicinal properties [1]. “Ashwaganda” roots are compared with ginseng roots for their restorative properties of vigor and vitality, hence been given the name “Indian ginseng” [1,2]. The whole plant, specially the leaves and the root bark are abortifacient, adaptogen, antibiotic, aphrodisiac, deobstruent, diuretic, narcotic, strongly sedative and tonic [3,4]. The fruit and seeds are diuretic while latter are hypotonic too [4]. The roots of W. somnifera contain bioactive compounds with sedative, anti-inflammatory, anti-tumour, antibacterial and anti-spasmodic properties [2]. W. somnifera acts mainly on the reproductive and nervous systems having a rejuvenative effect on the body and is used to improve vitality and aid recovery after chronic illness [5].
In India, the plant grows widely in drier areas in the sub-tropical and semi-temperate regions from plains to a height of 1700 m [2]. The plant is mainly propagated by seeds [3,6]. Its introduction, distribution, taxonomy and medicinal importance were studied by many authors [3,6,7]. Significant levels of genotypic and phenotypic variation, especially between the wild and cultivated morphotypes are also reported [8-11]. Variation with respect to growth habit, chemical profiling and sequence divergence using nrITS region of rDNA are recently published [1]. Earlier, Negi et al. [12] grouped the Indian germplasm into Nagori and Kashmiri groups based on AFLP dataset. The reported chromosome numbers for W. somnifera are 24, 48 and 72. Mohan Ram and Kamini [13] had reported 2n = 24 while 2n = 48 was reported by many authors [1,2,5,11,14-16]. Bir and Neelam [17] reported 2n = 72 presumably a polyploid number reported for this species. In view of such contradictory reports, the present investigation was conducted in W. somnifera accessions with sole objective to comprehend chromosome behavior during meiosis.
2. Materials and Methods
For locating populations of this species extensive field trips were conducted in various districts of Rajasthan. The particular plants were marked and labeled properly before collecting materials for study. For meiotic studies flower buds were used as basic material. Herbarium specimens along with phenotypic data of all the collected plants have been prepared and deposited in local herbaria of Botanical Survey of India, Jodhpur to obtain accession numbers.
Flower buds of appropriate size were collected from mandate mature plants and fixed in freshly prepared 1:3 glacial acetic acid: 95% ethanol mixture for a minimum 24 hours at room temperature and later stored in 70% ethanol at 10˚C. Anthers were squashed in 1% acetocarmine solution for meiotic studies. On average 25 PMCs were analyzed at diplotene/diakinesis/metaphase I. Pollen grains were stained in 1:1 glycerine:acetocarmine mixtures for estimating the percentage pollen stainability and on average 10 slides were scored for stainable pollen grains.
3. Results and Discussion
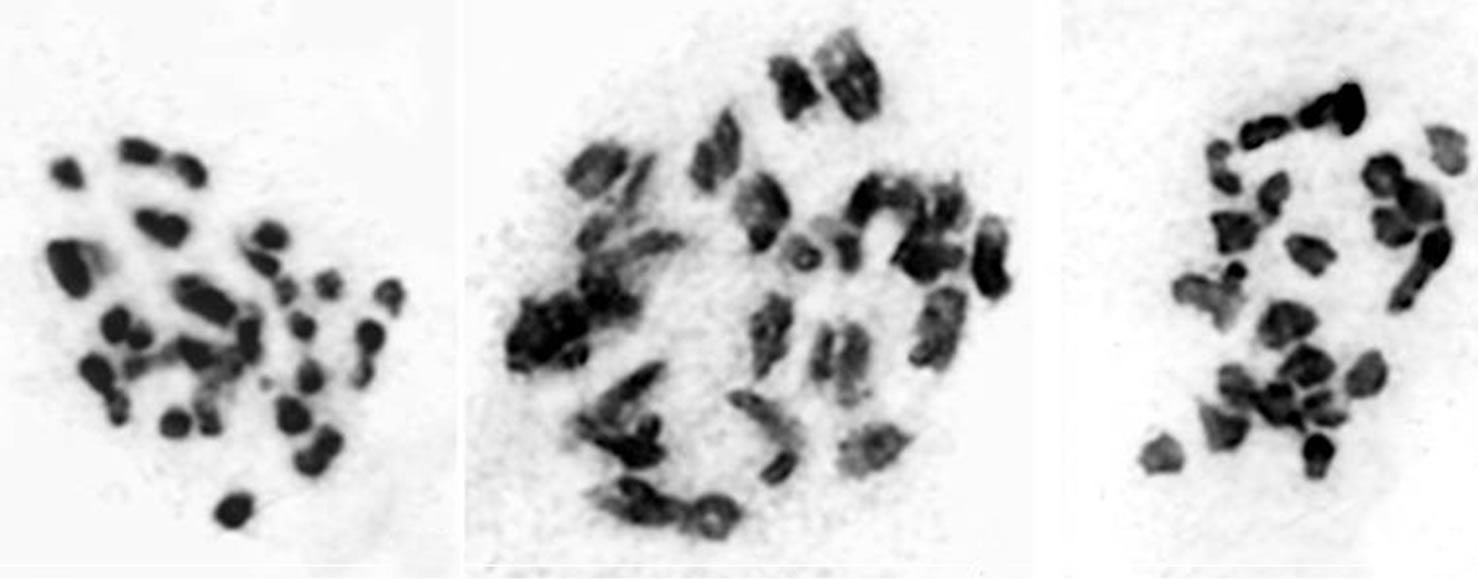
Four accessions of Withania somnifera from different locations belonging to family Solanaceae have been studied presently and illustrated in Table 1 and Figure 1(a)-(i). All these four were characterized by the gametic number of n = 24 (Figure 1). It was interesting to note that in two accessions Withania somnifera BSJO-26497 “A” and Withania somnifera BSJO-26497 “B”, of few cells with quadrivalent associations were recorded (Figures 1(a) and (b)) while the other two accessions were
 (a)
(a)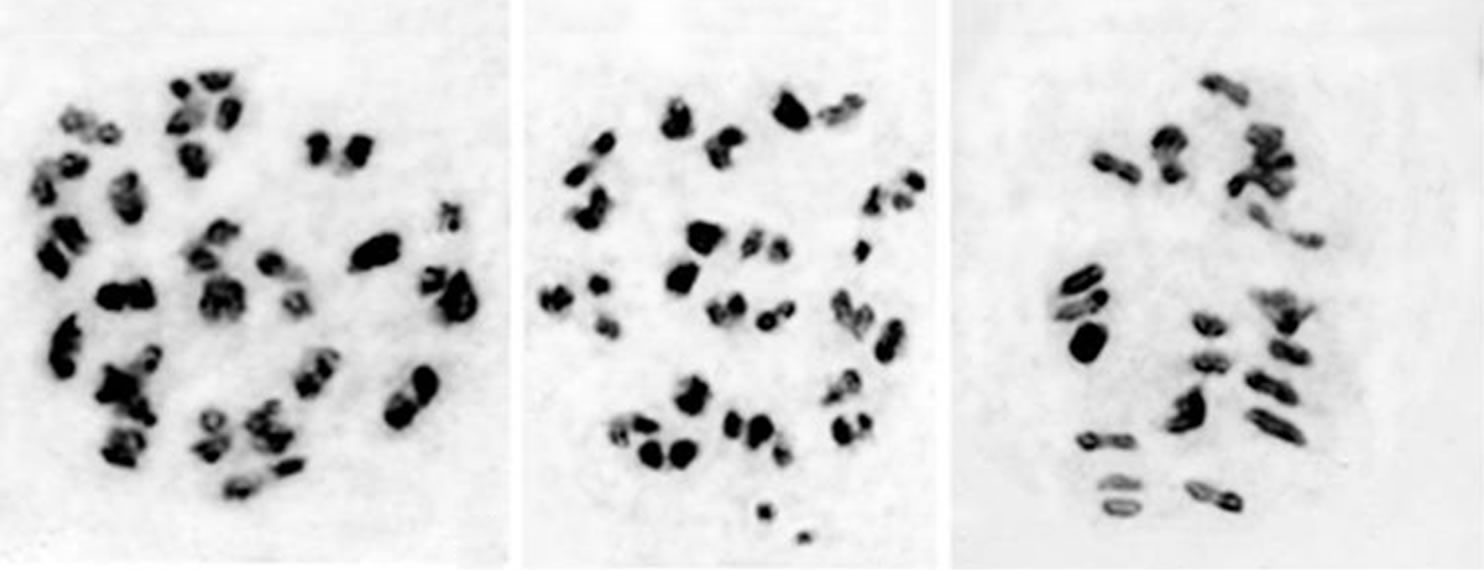 (b)
(b) (c)
(c)
Figure 1. Male meiosis in Withania somnifera 2n = 48; (a) and (i) W. somnifera BSJO-26497 “A”; (a) Metaphase I 20 II + 1 IV + 4 I; (i) Anaphase II; (b) and (c) W. somnifera BSJO-26497 “B”; (b) 23 II + 2 I; (c) Metaphase I 24 II; (d) and (e) W. somnifera BSJO-26497 “C”; (d) Metaphase I 20 II + 8 I; (e) Metaphase I 17 II + 14 I; (f)-(h) W. somnifera BSJO-26497 “D”; (f) Diakinesis 22 II + 4 I; (g) Metaphase I 24 II; (h) Anaphase I.
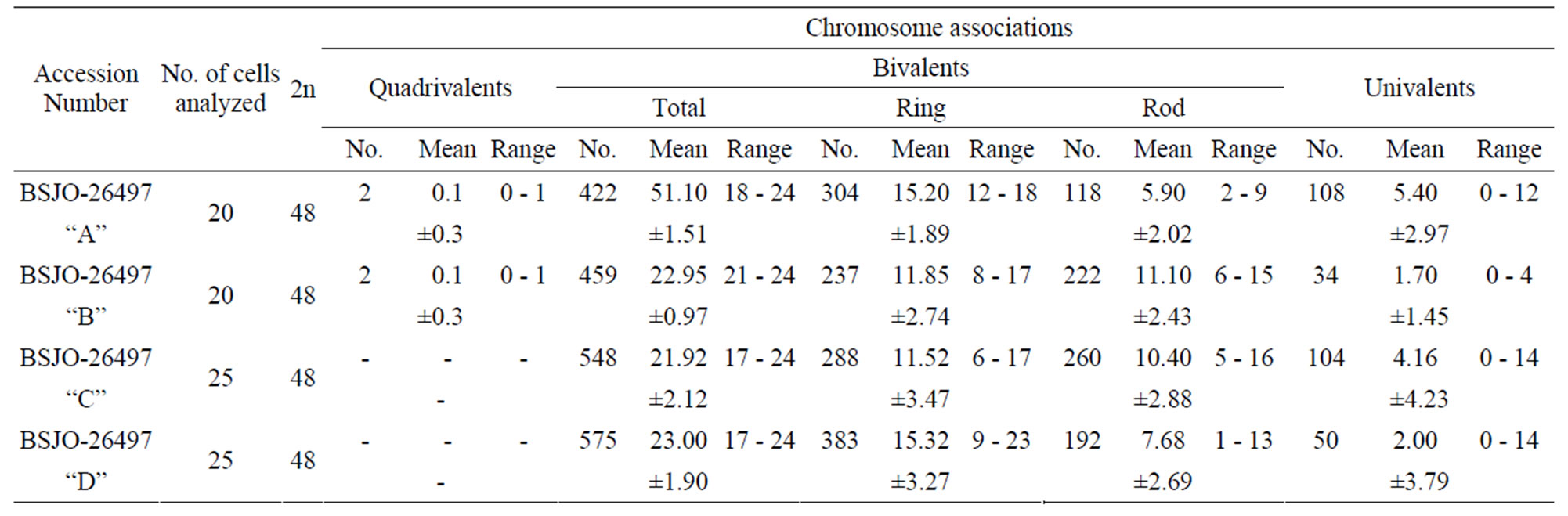
Table 1. Mean number and range of associations at diplotene/diakinesis/metaphase I in Withania somnifera accessions.
characteristic in having only bivalents and univalents (Table 1). Among the bivalents ring types have always dominated the rod types. Another remarkable feature of present study was regular occurrence of univalents. From the present observations combined with earlier published information [1,2,5,11,14-16], it is a clear that the genus Withania consist of a single basic number of x = 12. All the somatic numbers (24, 48 and 72) are the derivatives of x = 12 only. No deviant numbers were recorded in any of the PMCs analyzed. From these observations, Withania somnifera seems to be a tetraploid species of an unknown (allo/auto) nature. Complete lack of multivalent associations coupled with maximum number of bivalents indicate the probability of Withania somnifera might be of allopolyploid origin with basic number of x = 12.
In three accessions (A, B, C) most of the PMCs analyzed had shown a mixture of both the bivalents and univalents, while some of them had shown the presence of only bivalents. In remaining fourth accession (D) the majority of PMCs had shown presence of only bivalents, while some PMCs had shown the mixture of both the bivalents and univalents. The mean value of total bivalents is ranged between 21.10 and 23.0. Out of total bivalents the mean value of ring bivalents was ranged from 11.52 to 15.32 and of rod bivalents was ranged from 5.90 to 11.10 (Table 1).
The four different accessions of Withania somnifera were characteristics in having univalents with a mean value ranging from 2.00 - 5.40. Structural heterozygosity in bivalents rather than precautious separation of rod bivalents might be responsible for regular occurrence of univalents in the PMCs analyzed. A salient feature observed in the present investigation is that one bivalent was found consistently being associated with nucleolus during diakinesis/prometaphase I, this is indicative of nucleolar nature of bivalents. Such information through association analysis of bivalents were reported in an important medicinally important desert plant i.e. Dipcadi erythraeum recently, which is also an amphidiploids origin as revealed by cytogenetical investigation recently [18]. However, in the absence of karyological data it is difficult to confirm the apparent amphidiploids origin of the species.
The number of chiasmata per PMC ranged from 24 to 47 with a mean chiasmata value of 33.44 to 38.32 indifferent accessions. The mean value of chiasma terminalized also ranged between 28.56 and 35.64 to give rise terminalization coefficient value of about 0.85 to 0.93. The unterminalized chiasmata were ranged from 2.68 to 5.25. Generally 2 - 3 chiasmata were recorded and apparently the chiasmata were localized at distal region. The smaller size of the bivalents with availability of short segments for recombination may be the possible reason [6,19] for the varying degree of chiasma frequency. Similar observations were reported in case of certain crop plants like Eleusine [20-21], Vigna [22], Crotalaria [23], and hard wood tree species viz. Anogeissus, Salvadora, Capparis [24-26].
The distribution of chromosomes and chromatids at anaphase I and II was found to be regular with equal distribution in majority of the PMCs. Although presence of lagging univalents and bivalents was encountered in three out of four accessions studied presently, however, their number never exceeded more than 15 percent of the total PMCs analyzed. The percentage pollen stainability was ranged between 83.61 and 92.54. It suggests that the univalents encountered regularly at diakinesis/metaphase— I might be the reason for occurrence of laggards. However, they do not directly affect the viability of gametes and/or fertility [27,28] of the concerned taxon. From the perusal of literature it is apparent that the two species of Withania reported from India, W. somnifera and W. coagulans are polyploid derivatives with basic number x = 12. Such trends of polyploidy are noticeable in various genera belonging to Solanaceae for example Solanum tuberosum [12,19,21,29,31,32], Solanum trilobatum [13, 15] and Solanum nigrum [33,34] are the classical examples of polyploidy, though the exact nature of polyploidy (autopolyploidy/allopolyploidy/segmental allopolyploidy) is at variance. Therefore it is concluded that the speciation and species diversification in the genus Withania somnifera was mainly brought out by polyploidy.
Unfortunately not many details on either quantification or molecular composition of 2C/4C DNA are available for the genus Withania. Recently several workers [1,2] have assessed the genetic variation and intra-specific relationships and within Withania somnifera using highly defined molecular markers i.e. AFLPs, nrITS region of rDNA. They could able to clearly distinguish Withania somnifera from Withania coagulans and suggested the multidisciplinary approaches should be tried in order to resolve the ambiguities persist at taxonomic as well as genomic levels in this important medicinal herb. Recently, single primer amplification reaction (SPAR) methods have proved their utility to provide novel insight in genetic variation at interand intra specific levels in various plant taxa including desert trees [35,36]. However, most appropriate and authentic approach to determine the nature of polyploidy in case if Withania somnifera would be the most convincing genomic in-situ hybridization (GISH) technique as demonstrated by Raina and Mukai [37] in Aracis hypogaea to reveal the amphidiploids nature.
REFERENCES
- B. A. Mir, S. Loul, A. Kumar, M. K. Kaul, A. S. Soodan and S. N. Raina, “Intraspecific Variation in the Internal Transcribed Spacer (ITS) Regions of rDNA in Withania somnifera (Linn.) Dunal,” Indian Journal of Biotechnolog, Vol. 9, 2010, pp. 325-328.
- A. Kumar, B. A. Mir, D. Sehgal, T. H. Dar, S. Koul, M. K. Kaul, S. N. Raina and G. N. Qazi, “Utility of a Multidisciplinary Approach for Genome Diagnostics of Cultivated and Wild Germplasm Resources of Medicinal Withania somnifera, and the Status of New Species, W. ashwagandha, in the Cultivated Taxon,” Plant Systematics and Evolution, Vol. 291, No. 3-4, 2011, pp. 141-151. doi:10.1007/s00606-010-0372-4
- M. M. Bhandari, “Flora of the Indian Desert,” Scientific Publishers, Jodhpur, 1990.
- A. Chevallier, “The Encyclopedia of Medicinal Plants,” Dorling Kindersley, London, 1996.
- G. S. Chatha and S. S. Bir, “Cytological Evolution in Woody Taxa of Gamopetalae and Monochlamydae from South India,” Plant Sciences, Vol. 9, 1998, pp. 199-256.
- V. Singh and R. P. Pandey, “Ethnobotany of Rajasthan,” Scientific Publishers, Jodhpur, 1998.
- W. Emboden, “Narcotic Plants,” Studio Vista, London, 1979.
- C. K. Atal, “Schwarting AE. Ashwagandha––An Ancient Indian Drug,” Economic Botany, Vol. 15, No. 3, 1961, pp. 256-263. doi:10.1007/BF02862166
- C. K. Atal and A. E. Schwarting, “Schwarting. Intraspecific Variability in Withania somnifera. I. A Preliminary Survey,” Llyodia, Vol. 25, 1962, pp. 78-87
- N. S. Dhalla, K. C. Gupta, N. S. Sastry and C. L. Malhotra, “Comparative Studies on Withania somnifera Dunnl. and Withania ashwagandha Kaul,” Indian Journal of Pharmacology, Vol. 23, 1961, pp. 126-127.
- K. N. Kaul, “The Origin, Distribution and Cultivation of Ashwagandha the so Called Withania somnifera of Indian Literature,” Symposium on the Utilization of Indian Medicinal Plants, Council of Scientific & Industrial Research, New Delhi, 1957, pp. 7-8.
- S. Negi, A. Singh and M. Lakshmikumaran, “Genetic Variation and Relationship among and within Withania species as Revealed by AFLP Markers,” Genome, Vol. 43, No. 6, 2000, pp. 975-980.
- H. Y. MohanRam and I. Kamini, “Embryology and Fruit Development in Withania somnifera Dunal.,” Phytomorphology, Vol. 14, No. 3, 1964, pp. 574-587.
- S. R. Baquar, S. Akhtar and A. Hussain, “Meiotic Chromosome Numbers in Some Vascular Plants of Indus Delta I,” Botaniska Notiser, Vol. 118, 1965, pp. 289-298.
- P. N. Bhaduri, “Chromosome Numbers of Some Solanaceous Plants of Bengal,” Journal of the Indian Botanical Society, Vol. 12, 1933, pp. 56-64.
- J. Miege, “Troisieme Liste de Numbers Chromosomiques d’Especes d’Afrique Occidentale,” Ann Fac Sci Univ Dakar., Vol. 5, 1960, pp. 75-85.
- S. S. Bir and M. Sidhu, “Cytological Observations on Weed Flora of Patiala District, Punjab,” In: S. S. Bir, Eds., Recent Researches in Plant Sciences, Kalyani Publishers, Ludhiana, 1980, pp. 261-271.
- D. Rawat, S. K. Sharma, A. Mahmoudi and S. R. Rao, “Cytogenetic Rationale for Probable Amphidiploids Origin of Dipcadi erythraeum Webb. & Berth—A Rare and Endemic Plant of Indian Thar Desert,” Caryologia, Vol. 64, No. 1, 2011, pp. 75-83.
- V. D. Turkov and G. A. Shelepina, “Karyotype Characteristics of Cultivated Potato (Solanum tuberosum L.),” Dokl Vses Akad Skh IM VI Lenina, Vol. 11, 1974, pp. 18-19.
- R. M. Devarmumath, S. C. Hiremath, S. R. Rao, A. Kumar and S. Bewal, “Genome Analysis of Finger Millet E. coracana by Interspecific Hybridization among Diploid Wild Species of Eleusine (Poaceae),” Cytologia, Vol. 70, 2005, pp. 427-434. doi:10.1508/cytologia.70.427
- R. M. Devarmumath, S. C. Hiremath, S. R. Rao and A. Kumar, “Genome Interrelationship in the Genus Eleusine (Poaceae) as Revealed through Heteroploid Crosses,” Caryologia, Vol. 58, No. 4, 2005, pp. 300-307.
- P. Scartezzini, F. Antognoni, L. Conte, A. Maxia, A. Troia and F. Poli, “Genetic and Phytochemical Difference between Some Indian and Italian Plants of Withania somnifera (L.) Dunal,” Natural Product Research: Formerly Natural Product Letters, Vol. 21, No. 10, 2007, pp. 923- 932. doi:10.1080/14786410701500169
- R. C. Verma, “Cytogenetics of Crotalaria and Phlox,” Ph.D. Thesis, University of Jodhpur, Jodhpur, 1981.
- A. Kumar and S. R. Rao, “Chromosome Studies in Capparis deciduas,” Nucleus, Vol. 46, No. 1-2, 2003, pp. 95-98.
- A. Kumar and S. R. Rao, “Cytological Investigations in Some Tree Species of Rajasthan II. Male Meiosis Studies in the Genus Anogeissus (DC.),” Caryologia, Vol. 55, No. 1, 2002, pp. 63-72.
- V. Kumar and B. Subramaniam, “Chromosome Atlas of Flowering Plants of the Indian Subcontinent,” Botanical Survey of India, Ministry of Environment & Forests, Government of India, 1987.
- Y. S. Bedi, B. S. Gill and S. S. Bir, “Cytological Evolution in Woody Taxa of Gamopetalae and Monochlamydae,” Journal of Cytology and Genetics, Vol. 20, 1985, pp. 162-203.
- W. Gottschalk, “Die Chromosomenstruktur der Solanceen unter Berucksichtigung Phylogenetischer Fragestellungen,” Chromosoma, Vol. 6, 1954, pp. 539-626. doi:10.1007/BF01259953
- G. R. Rao and A. Kumar, “Chromosome Pairing in Species Hybrids of Solanum nigrum Complex,” Current Science, Vol. 49, No. 22, 1980, pp. 868-870.
- K. N. Subramanyam, H. Kishore and P. Misra, “Hybridization of Haploids of Potato in the Plains of India,” Current Science, Vol. 41, 1972, p. 580.
- E. W. M. Van Breukelen, M. S. Ramanna and J. G. Hermsen, “Monohaploids (n=x=12) from Autotetraploid Solanum tuberosum (2n=4x=48) through Two Successive Cycles of Female Parthenogenesis,” Euphytica, Vol. 24, No. 3, 1975, pp. 567-574. doi:10.1007/BF00132892
- M. Wagenvoort and W. Lange, “The Production of Aneudihaploids in Solanum tuberosum L. Group tuberosum (The Common Potato),” Euphytica, Vol. 24, No. 3, 1975, pp. 731-741. doi:10.1007/BF00132912
- M. Sanjapa and N. Sathyananda, “In IOPB Chromosome Number Reports LXIV,” Taxon, Vol. 28, No. 4, 1979, pp. 391-408.
- J. Venkateswarlu and M. K. Rao, “Breeding Systems, Cross Ability Relationships and Isolating Mechanisms in the Solanum nigrum Complex,” Cytologia, Vol. 37, 1972, pp. 317-326. doi:10.1508/cytologia.37.317
- S. K. Sharma, S. Kumaria, P. Tandon and S. R. Rao, “Single Primer Amplification Reaction (SPAR) Reveals Interand Intra-Specific Natural Genetic Variation in Five Species of Cymbidium (Orchidaceae)” Gene, Vol. 483, No. 1-2, 2011, pp. 54-62. doi:10.1016/j.gene.2011.05.013
- S. K. Sharma, D. Rawat, S. Kumar, A. Kumar, S. Kumaria and S. R. Rao, “Single Primer Amplification Reaction (SPAR) Reveals Intra-Specific Natural Variation in Prosopis cineraria (L.) Druce.,” Trees-Structure and Function, Vol. 24, No. 5, 2010, pp. 855-864. doi:10.1007/s00468-010-0455-4
- S. N. Raina and Y. Mukai, “Detection of a Variable Number of 18S-5.8S-26S and 5S Ribosomal DNA Loci by Fluorescent in Situ Hybridization in Diploid and Tetraploid Arachis Species,” Genome, Vol. 42, No. 1, 1999, pp. 52-59.
NOTES
*Corresponding author.

