Advances in Breast Cancer Research
Vol.2 No.4(2013), Article ID:36728,5 pages DOI:10.4236/abcr.2013.24019
Polymorphisms at GSTM1, GSTP1, GSTT1 Detoxification Genes Loci and Risk of Breast Cancer in Kazakhstan Population
Institute of Molecular Biology & Biochemistry, Almaty, Kazakhstan
Email: mbbtimur@mail.ru
Copyright © 2013 T. S. Balmukhanov et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Received May 20, 2013; revised June 21, 2013; accepted June 29, 2013
Keywords: Breast Cancer; Gene Polymorphism; GSTM1, GSTP1, GSTT1 Genes; Kazakhstan
ABSTRACT
Associations of null polymorphism (copy number variation) of detoxification genes GSTM1, GSTT1 and GSTP1 (at rs2495636, 105 Ile/Val) with the breast cancer (BC) were assessed in two main ethnic groups of the Republic of Kazakhstan (Kazakhs and Russians). Total number of patients was 181, and of controls 397. Statistically significant difference was observed between BC patients and healthy individuals in alleles frequency (χ2 = 4.89, р = 0.007) of GSTP1 gene at rs2495636 (105 Ile/Val) among the Kazakhs ethnic group. Difference in genotypes distribution (χ2 =5.26, р = 0.076) at this site is approximating to be statistically significant. In the Russian group, no differences were found in genotypes and alleles atrs 2495636 of GSTP1 gene between cases and controls. There was no significant difference between null polymorphism (copy number variation) of GSTM1 and GSTT1 genes among cases and controls in both ethnic groups.
1. Introduction
Breast cancer (BC) is one of the most common cancers among women worldwide. Recent studies and modern GWAS describe that the associations between BC and polymorphisms in genes are involved in xenobiotic detoxification.
GSTM1, GSTP1, and GSTT1 genes belong to the glutathione-S-transferases superfamily of genes, whose products are Phase II metabolizing enzymes, which act in coordination with Phase I metabolizing enzymes in the carcinogen metabolism. The Phase I enzymes usually activate the carcinogens to reactive intermediates and the GSTs are active in the detoxification of a wide variety of these potentially toxic and carcinogenic electrophiles by conjugating them to Glutathione [1]. The variations in metabolic activities in each phase or in the coordination of these two phases regulate the clearance of DNA toxic metabolites and might be partially responsible for individual host susceptibility to cancer.
About 50% of both Caucasians and Japanese lack the GSTM1 gene due to inherited homozygous deletion of both alleles [2]. The individuals with GSTM1 null polymorphism lack the ability of detoxifying specific substrate epoxide intermediates [3]. The GSTM1 null genotype is also positively associated with high DNA adduct levels, suggesting its role in carcinogenesis [4].
Several investigators have speculated that the early loss of GSTP1 function leads to increased vulnerability to oxidant and heterocyclic amine carcinogens, implicated in prostate carcinogenesis [5]. Hence, heritable differences in GSTP1 function may also be associated with prostate cancer development. An A to G polymorphism at nucleotide 313 of GSTP1 results in an amino acid substitution (Ile105Val) at the substratebinding site of GSTP1 [6]. The substitution of the less bulkier and more hydrophobic valine results in substrate-dependent alterations of GSTP1 catalytic activity [6,7]. Positive associations have been reported between the GSTP1 I105V polymorphism and risk of oral and breast cancers [8,9].
The interest to the associations of the polymorphic variations of GSTPs genes with BC is growing owing to the fact that its variations’ important role in the sensitivity to the cancer chemotherapy and tumors progression [10]. The results of investigations describing the associations of rs2495636 site of GSTP1 gene with BC in different world populations are controversial. The studies performed in Turkey, Thailand and China did not reveal this association [11-13]. The data demonstrating positive association of GSTP1 gene polymorphism that was taken alone with BC and in combination with some other polymerphisms of candidate-genes were also published [14].
The GSTT1 gene is also one of the members of GST’s enzymes family. It has lower glutathione binding activity, along with increased catalytic efficiency. It is shown that null genotype of GSTT1 contributes to colorectal cancer risk in Asian populations [15]. The data describing the associations of GSTT1 gene polymorphisms with pseudoexfoliative glaucoma in Pakistani population [16], breast cancer susceptibility [17] and the results of pooled analysis [18] were published earlier.
Republic of Kazakhstan is a multinational state located in the middle of Central Asia with population of about 17 million and its main ethnical groups are Kazakhs (Asians) and Russians (Caucasians). Specific ethnic differences in the distribution of the genotypes and in the frequencies of alleles, which can be used as markers of BC among representatives of different races and nations, were described earlier in numerous publications.
The present study was undertaken to determine the distribution of genotypes and alleles frequency of GSTP1 (rs2495636), GSTM1 null polymorphisms and GSTT1 (Ile105Val) polymorphism among BC cases and controls so as to understand whether these polymorphisms are associated with the risk of BC in Kazakhstan population.
2. Materials and Methods
Venous blood samples (5 ml) were collected in EDTA vials from 181 women diagnosed with BC from Almaty (Kazakh Research Institute of Oncology and Radiology). Blood samples from 397 healthy women donors (Almaty) without clinical symptoms and with no family history of cancers were used as a controls. The average age of the BC patients was 50.3 ± 11.6 years (Kazakhs), 55.7 ± 11.7 (Russians) and of the donors 50.07 ± 8.47 (Kazakhs), 54.8 ± 5.9 (Russians) years. Deoxyribonucleoside triphosphates, restriction endonuclease BstMAI, Taq DNA polymerase, BSA were purchased by “SibEnzime” (Russia).
Methods. DNA was isolated from the blood using DNA extraction kits “Axygen” (USA). The sequences of the oligonucleotide primers were obtained from literature sources. Individual SNPs were detected by restriction fragment length polymorphism (RFLP) analysis after PCR amplification.
PCR reactions were carried out using gradient Mastercycle “Eppendorf”. Amplification was performed in a buffer containing 67 mM TrisHCl, pH 8.8, 16.6 mM (NH4)2SO4, 2 mM MgCl2, 0.01% Tween 20, 0.15 mg/ml BSA, 2 pM of each of the primers, 0.25 mM each of the four dNTPs, 1 unit/μl of Taq DNA polymerase. The oligonucleotide sequences of the primers for the amplification of GSTM1 gene were F 5’-GAACTCCCTGAAAAGCTAAAGC-3’ and R 5’-GTTGGGCTCAAAACGGTGG-3’; for GSTT1 gene—F 5’-ttccttactggtcctcacatctc-3’ and R: 5’-TCACCGGATCATGGCCAGCA-3’ according to KhanM.I. [8]. The sequences of primers used for β globin gene amplification used as a positive control of amplification were F: 5’-CAACTTCATCCACGTTCACC-3’ and R: 5’-GAAGAGCCAAGGACAGGTAC-3’. GSTP1 gene (rs2495636) was amplified using primers F: 5’-ACCCCAGGGCTCTATGGGAA-3’; R: 5’-TGAGGGCACAAGAAGCCCCT-3’ and digested with BstMAI restrictase [9].
Electrophoresis was carried out in 8% polyacrylamide gel at 40 mA and 150 V for 2 - 3 h.
Statistical analysis. The Pearson χ2 criterion (p < 0.05), odds ratios (OR), 95% confidence intervals (CI) tests were applied to the data analysis. The distribution of genotypes for both investigated groups was in accordance with the Hardy-Weinberg equilibrium (HWE).
3. Results and Discussion
Null polymorphism is a result of lengthy deletion resulting in the formation of truncated form of the protein with decreased enzymatic activity. In current investigation null polymorphism of the GSTP1 and GSTM1 genes was tested by means of multiplex PCR. Electrophoretically it is possible to detect the two variants of amplified products: 0/0 (homozygous deletion)—absence of the bands, and either +/0 or +/+ (heterozygous deletion and homozygous with functional genes without deletions) were presented by bands of 459 bp for GSTT1 gene and of 209 bp for GSTM1 gene (Figure 1). Beta-globine gene was used as a positive control (268 bp). Such approach permits combining the genotypes of the samples according either presence or absence of even sole copy of tested genes.
No differences were found in distribution of GSTP1 and GSTM1 genes null polymorphism between patients and controls in both the Kazakh and in Russian groups. Also, no interethnic differences were shown between Kazakhs and Russians in the genotypes distribution. The results are presented in Table 1.
To evaluate the interaction between the two genotypes, we examined the combined effect of the GSTT1 and GSTM1 genotypes. Reference group of samples having active variants of the genes was used as a base line. The analysis revealed no significant difference among cases and controls in distribution of the genotype combination neither in Kazakh nor in Russian groups.
Thereby the associations of null polymorphic variants of GSTT1 and GSTM1 genes with BC were not found in the two tested ethnic groups of Kazakhstan.
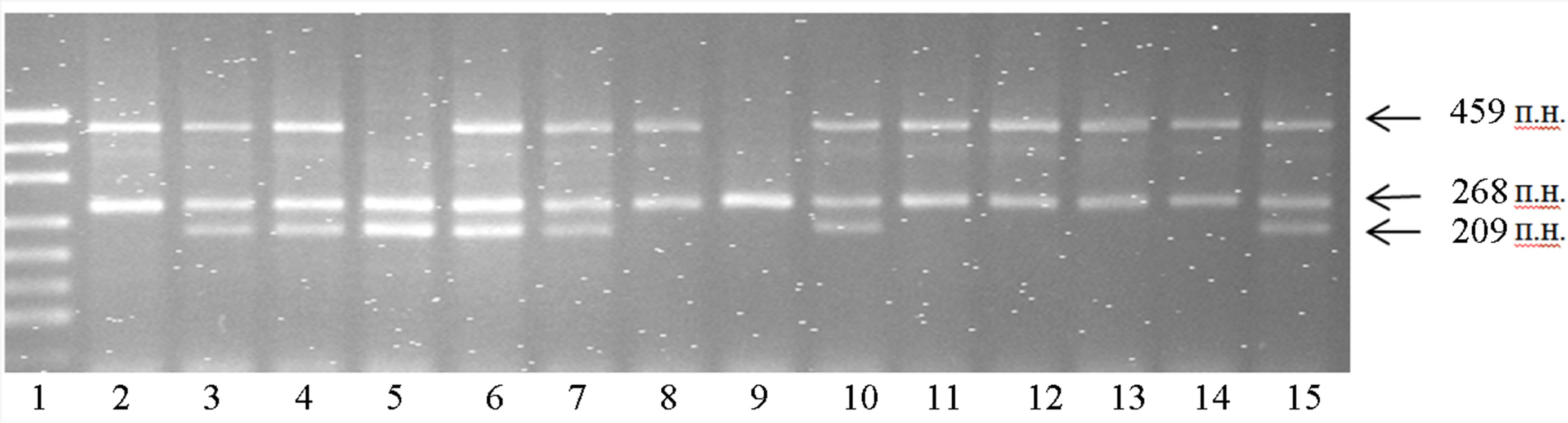
Figure 1. PAAG-analysis of GSTT1 and GSTM1 genes amplified products. Lane 1—molecular mass marker pUC19/Kzo9 I; lanes 2, 8, 11, 12, 13, 14—genotype Т1М0; lanes 3, 4, 6, 7, 10, 15—genotype Т1М1; lane 9—genotype Т0М0.
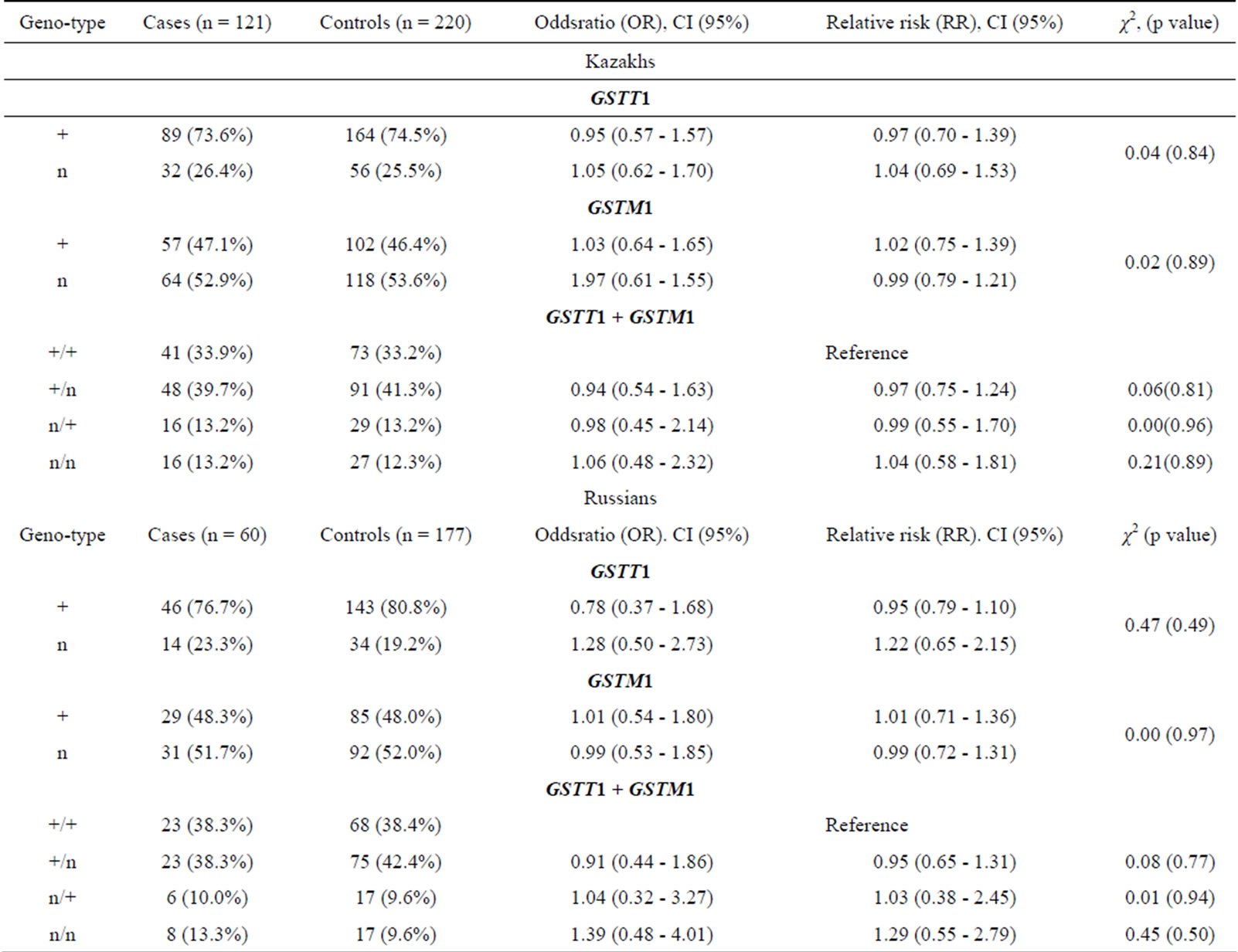
Table 1. Null polymorphism of GSTT1 and GSTM1 genes in Kazakh and Russian ethnic groups.
The search of association of BC with 105 Ile/Val polymorphism of GSTP1 gene at site rs2495636 was performed by restriction fragment length polymorphism (RFLP) analysis.
The PCR product of 176 bp was subjected to restriction digestion by BstMAI at 37˚ for 4 hour and DNA band was resolved by electrophoresis. The genotypes were determined based on the band pattern.
The Val allele contains BstMAI specific sequence whereas the Ile allele lack such sequence; thus electrophoretically Ile/Ile genotype is identified as an undigested band of 176 bp, the Val/Val genotype—as bands of 91 bp and 85 bp and Ile/Val genotype, is determined by three fragments of 176 bp, 91 bp and 85 bp (Figure 2). The results are presented in the Table 2.
As it can be seen from the results presented in Table 2, no significant differences were found between cases and control in allele frequencies and genotypes distribution of the tested sites of GSTT1 and GSTM1 genes in the Russian group. Interethnic differences in alleles and genotypes
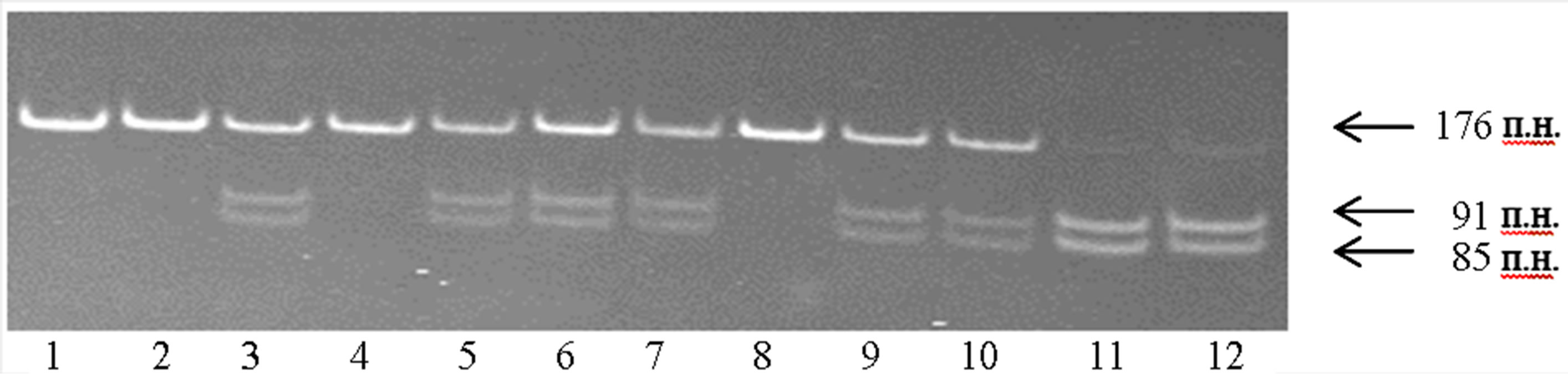
Figure 2. 8% PAAG-analysis of the RFLP-PCR products of GSTР1 gene (rs2495636). Lanes 1, 2, 4, 8—genotype A/A (Ile/Ile); lanes 3, 5, 6, 7, 9, 10 A/G (Ile/Val); lanes11, 12—genotype G/G (Val/Val).
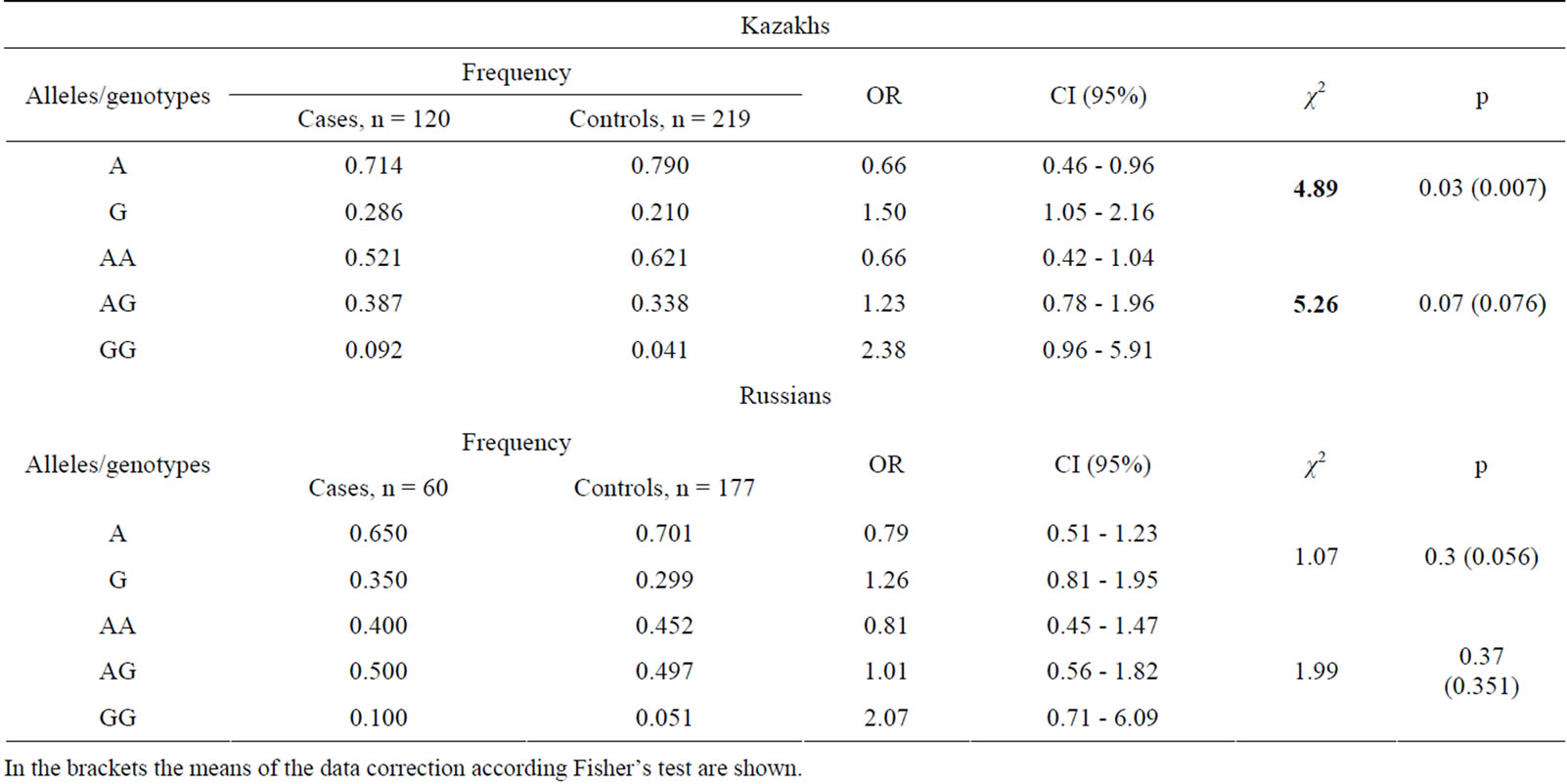
Table 2. Genotypes distribution and allele frequency atrs 2495636 of GSTP1 gene in Kazakh and Russian ethic groups.
In the brackets the means of the data correction according Fisher’s test are shown.
distribution between the Kazakhs and Russians also were not revealed. In both ethnic groups, the genotypes’ distribution was in accordance with HWE.
In the Kazakh group the statistically significant differences in allele frequency and genotypes distribution between cases and controls were demonstrated. The value of odds ratio (OR) for the allele G is equal to 1.50 (95% CI = 1.05 - 2.16) and OR for genotype GG—2.38 (95% CI = 0.96 - 5.91), which can be indicative for regarding of this polymorphism as a risk factor of BC in Kazakh ethnic group of Kazakhstan population.
REFERENCES
- R. C. Strange, J. T. Lear and A. A. Fryer, “Glutathione S-Transferase Polymorphisms: Influence on Susceptibility to Cancer,” Chemico-Biological Interactions, Vol. 111-112, 1998, pp. 351-364. http://dx.doi.org/10.1016/S0009-2797(97)00172-5
- T. R. Rebbeck, “Molecular Epidemiology of the Human Glutathione S-Transferase Genotypes GSTM1 and GSTT1 in Cancer Susceptibility,” Cancer Epidemiology, Biomarkers & Prevention, Vol. 6, No. 9, 1997, pp. 733-743.
- V. Nazar-Stewart, A. G. Motulsky and D. L. Eaton, “The Glutathione S-Transferase mu Polymorphism as a Marker for Susceptibility to Lung Carcinoma,” Cancer Research, Vol. 53, No. S10, 1993, pp. 2313-2318.
- J. K. Wiencke, K. T. Kelsey, R. A. Lamela and W. A. Toscano Jr., “Human Glutathione S-Transferase Deficiency as a Marker of Susceptibility to Epoxide-Induced Cytogenetic Damage,” Cancer Research, Vol. 50, No. 5, 1990, pp. 1585-1590.
- W. G. Nelson, A. M. De Marzo and T. L. DeWeese, “The Molecular Pathogenesis of Prostate Cancer: Implications for Prostate Cancer Prevention,” Urology, Vol. 57, No. S4, 2001, pp. 39-45. http://dx.doi.org/10.1016/S0090-4295(00)00939-0
- F. Ali-Osman, O. Akande and G. Antoun, “Molecular Cloning, Characterization, and Expression in Escherichia coli of Full-Length cDNAs of Three Human Glutathione S-Transferase Pi Gene Variants. Evidence for Differential Catalytic Activity of the Encoded Proteins,” The Journal of Biological Chemistry, Vol. 272, No. 15, 1997, pp. 10004-10012. http://dx.doi.org/10.1074/jbc.272.15.10004
- K. Sundberg, A. S. Johansson and G. Stenberg, “Differences in the Catalytic Efficiencies of Allelic Variants of Glutathione Transferase P1-1 towards Carcinogenic Diol Epoxides of Polycyclic Aromatic Hydrocarbons,” Carcinogenesis, Vol. 19, No. 3, 1998, pp. 433-436. http://dx.doi.org/10.1093/carcin/19.3.433
- S. K. Park, D. S. Yim, K. S. Yoon, I. M. Choi, J. Y. Choi, K. Y. Yoo, D. Y. Noh, K. J. Choe, S. H. Ahn, A. Hirvonen and D. Kang, “Combined Effect of GSTM1, GSTT1, and COMT Genotypes in Individual Breast Cancer Risk,” Breast Cancer Research and Treatment, Vol. 88, No. 1, 2004, pp. 55-62. http://dx.doi.org/10.1007/s10549-004-0745-x
- K. Mitrunen, N. Jourenkova, V. Kataja, M. Eskelinen, V. M. Kosma, S. Benhamou, H. Vainio, M. Uusitupa and A. Hirvonen, “Glutathione S-Transferase M1, M3, P1, and T1 Genetic Polymorphisms and Susceptibility to Breast Cancer,” Cancer Epidemiology, Biomarkers & Prevention, Vol. 10, No. 3, 2001, pp. 229-236.
- A. L. Oliveira, F. F. Rodrigues, R. E. Santos, T. Aoki, M. N. Rocha, C. A. Longui and M. B. Melo, “GSTT1, GSTM1, and GSTP1 Polymorphisms and Chemotherapy Response in Locally Advanced Breast Cancer,” Genetics and Molecular Research, Vol. 9, No. 2, 2010, pp. 1045- 1053. http://dx.doi.org/10.4238/vol9-2gmr726
- T. Pongtheerat, M. Treetrisool and W. Purisa, “Glutathione S-Transferase Polymorphisms in Breast Cancers of Thai Patients,” Asian Pacific Journal of Cancer Prevention, Vol. 10, No. 1, 2009, pp. 127-132.
- B. Kiran, M. Karkucak, H. Ozan, T. Yakut, K. Ozerkan, S. Sag and M. J. Ture, “GST (GSTM1, GSTT1, and GSTP1) Polymorphisms in the Genetic Susceptibility of Turkish Patients to Cervical Cancer,” Journal of Gynecologic Oncology, Vol. 21, No. 3, 2010, pp. 169-173. http://dx.doi.org/10.3802/jgo.2010.21.3.169
- L. C. Sakoda, C. R. Blackston, K. Xue, J. A. Doherty, R. M. Ray, M. G. Lin, H. Stalsberg, D. L. Gao, Z. Feng, D. B. Thomas and C. Chen, “Glutathione S-Transferase M1 and P1 Polymorphisms and Risk of Breast Cancer and Fibrocystic Breast Conditions in Chinese Women,” Breast Cancer Research and Treatment, Vol. 109, No. 1, 2008, pp. 143-155. http://dx.doi.org/10.1007/s10549-007-9633-5
- B. O. Van Emburgh, J. J. Hu, E. A. Levine, L. J. Mosley, N. D. Perrier, R. I. Freimanis, G. O. Allen, P. Rubin, G. B. Sherrill, C. S. Shaw, L. A. Carey, L. R. Sawyer and M. S. Miller, “Polymorphisms in CYP1B1, GSTM1, GSTT1 and GSTP1, and Susceptibility to Breast Cancer,” Oncology Report, Vol. 19, No. 5, 2008, pp. 1311-1321.
- D. Xu, S. Yan, J. Yin and P. Zhang, “Null Genotype of GSTT1 Contributes to Colorectal Cancer Risk in Asian Populations: Evidence from a Meta-Analysis,” Asian Pacific Journal of Cancer Prevention, Vol. 12, No. 9, 2011, pp. 2279-2284.
- M. I. Khan, S. Micheal, F. Akhtar, W. Ahmed, B. Ijaz, A. Ahmed and R. Qamar, “The Association of Glutathione S-Transferase GSTT1 and GSTM1 Gene Polymorphism with Pseudoexfoliative Glaucoma in a Pakistani Population,” Molecular Vision, Vol. 26, No. 16, 2010, pp. 2146-2152.
- A. C. Ramalhinho, J. A. Fonseca-Moutinho and L. A. Breitenfeld Granadeiro, “Positive Association of Polymorphisms in Estrogen Biosynthesis Gene, CYP19A1, and Metabolism, GST, in Breast Cancer Susceptibility,” DNA and Cell Biology, Vol. 31, No. 6, 2012, pp. 1100- 1106. http://dx.doi.org/10.1089/dna.2011.1538
- F. D. Vogl, E. Taioli, C. Maugard, W. Zheng, L. F. Pinto, C. Ambrosone, F. F. Parl, V. Nedelcheva-Kristensen, T. R. Rebbeck, P. Brennan and P. Boffetta, “Glutathione S-Transferases M1, T1, and P1 and Breast Cancer: A Pooled Analysis,” Cancer Epidemiology, Biomarkers & Prevention, Vol. 13, No. 9, 2004, pp. 1473-1479.

