Computational Chemistry
Vol.05 No.03(2017), Article ID:77542,10 pages
10.4236/cc.2017.53009
ONIOM Method Characterization of Hydrogen Bonding Sites of Mycolactone A/B, a Buruli Ulcer Toxin
Kadjo François Kassi, Mamadou Guy-Richard Koné, Sopi Thomas Affi, Nahossé Ziao*
Laboratoire de Thermodynamique et de Physico-Chimie du Milieu, UFR SFA, Université Nangui Abrogoua, Abidjan, Côte-d’Ivoire

Copyright © 2017 by authors and Scientific Research Publishing Inc.
This work is licensed under the Creative Commons Attribution International License (CC BY 4.0).
http://creativecommons.org/licenses/by/4.0/



Received: May 23, 2017; Accepted: July 8, 2017; Published: July 11, 2017
ABSTRACT
Mycolactone molecules are responsible of Buruli ulcer disease. In this work, we are interested in the geometric, energetic and spectroscopic characterization of the hydrogen bonding interactions in mycolactone A/B, using quantum chemical method, especially ONIOM(HF/6-311+G(d,p):AM1) and ONIOM (B3LYP/6-311+G(d,p):AM1) levels. ONIOM two layers method has been used because mycolactones compounds are very large, taking into account diffuse and polarization functions are important whenever the matter is intermole- cular interactions. Geometric, energetic and spectroscopic parameters of hydrogen bonding reaction on each of the nine oxygen heteroatoms of mycolactone A/B have revealed that the O5sp2 heteroatom is far away the hydrogen bonding site. The identification of such a site constitutes a tool for working out a methodology for the annihilation of the destruction effects of mycolactones.
Keywords:
Hydrogen Bonding, Mycobacterium ulcerans, Mycolactone, ONIOM, Quantum Chemistry

1. Introduction
Buruli ulcer is a disease caused by Mycobacterium ulcerans, a microorganism belonging to the family of bacteria responsible for tuberculosis and leprosy [1] . Longtime neglected, this disease that prevails in tropical and subtropical humid countries, has increased in West Africa since 1980 [2] . This situation led the World Health Organization (WHO) to classify this disease as emerging and to recognize it as a public health and development problem [3] . Mycobacterium ulcerans secretes a toxin called mycolactone, responsible for extremely deep tissue damage, because of its cytotoxic and immunosuppressive properties. Nowadays, six (06) different natural molecular structures of mycolactones named A/B, C, D, E, F and G, have been isolated [4] . The mycolactone is constituted of a lactone ring linked to two lateral chains. Especially, the form A/B is the subject of this study (Figure 1).
Despite the progress in medical management, the therapeutic arsenal against Buruli ulcer remains limited [5] . Indeed, antibiotic therapy and restorative surgery remain the reference treatment, with high cost and numerous relapses (16% to 28%), in case of serious infection [6] . The mode of the toxin’s action remains unknown. The relationship between mycolactone and the proteins responsible for the appearance of Buruli ulcer are related to the conformation of the molecules and their intermolecular interactions. Hydrogen bond is one of the most important inter-molecular interactions involved in supramolecular chemistry, protein-ligand interactions [7] [8] and especially crystal engineering [9] [10] . Polyfunctional molecules, generally, comprise several heteroatoms which are capable to receive Hydrogen bonds. This work, part of Buruli ulcer control program, focuses on mycolactones A/B. It aims to determine, by quantum chemical methods, some physicochemical properties of these mycolactone molecules, in particular, geometric and energetic parameters of the hydrogen bonds established on the heteroatoms, in order to determine hydrogen bonding site. Final aim is to propose an experimental methodology of the annihilation of the destructive effects of mycolactone A/B.
2. Experimentation Section
2.1. Computational Details
Mycolactone A/B possesses nine (09) heteroatoms, all those are sp2 or sp3 hybridized oxygen atoms, shown in red color at a 3D molecular structure of mycolactone A/B (Figure 2). Heteroatoms are numbered from 1 to 9 and these numbers will also correspond respectively to the names of the different hydrogen bond complexes.
ONIOM method, developed by Morokuma et al. [11] , is used because of the
Figure 1. 2D structure of mycolactone A/B.
Figure 2. 3D Molecular structure of mycolactone A/B (visualized with Gauss View 5.0 software).
high number of atoms in mycolactone A/B. It consists of cutting the studied system into several layers, each of the layers being treated at a different calculation level. It therefore allows to describe precisely the part of the system which has particular interest for the study, called the internal layer, and to describe in a less precise manner the rest of the system, called the outer layer or the environment. ONIOM method permits to obtain the energy of the real system at a high level of computation called high level, 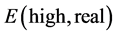 , by means of extrapolation according Equation (1), where
, by means of extrapolation according Equation (1), where  is the energy of the real system at the low calculation level,
is the energy of the real system at the low calculation level, 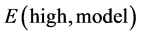 the energy of the model system at the higher calculation level, and
the energy of the model system at the higher calculation level, and 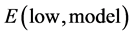 the energy of the model system at the low calculation level. An example of description of the ONIOM two layers in mycolactone A/B is shown in Figure 3.
the energy of the model system at the low calculation level. An example of description of the ONIOM two layers in mycolactone A/B is shown in Figure 3.
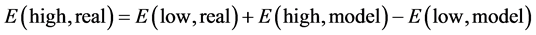 (1)
(1)
All calculations were performed, using Gaussian 03 software [12] at the ONIOM (HF/6-311+G(d,p):AM1) and ONIOM(B3LYP/6-311+G(d,p):AM1). The presence of diffuse and polarization functions in the basis sets is important in order to take into account the lone pairs of the heteroatoms, as well as intermolecular interactions.
2.2. Geometry Optimization
Nine hydrogen bond complexes were constructed on each of the oxygen heteroatoms, a water molecule being the probe, as Hydrogen Bonding Donor. Such hydrogen bond can be characterized by geometric parameters (Figure 4).
Before optimization, for all complexes, the angle of the linearity α has been set at 180˚ and the angle of the directionality β, at 109.5˚ for sp3 hybridized oxygen and 120˚ for sp2 hybridized oxygen. According to Gillespie’s V.S.E.P.R (Valence Shell Electron Pair Repulsion) theory (Figure 5), the distance d between an oxygen atom of mycolactone and a hydrogen atom of the probe is set at 2 Å. These values correspond respectively to the angles and the minimum approach distance of the hydrogen bond [14] .
Figure 3. Description of a model for cutting a complex of mycolactone A/B according ONIOM method (Figure from gaussview software).
Figure 4. Geometric parameters α, β and d describing hydrogen bond [13] .
Figure 5. Definition of linearity and directionality angles describing hydrogen bond.
2.3. Energetic Parameters
Hydrogen bonding between a donor molecule 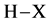 and an acceptor molecule
and an acceptor molecule 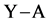 occurs according reaction 2. The Hydrogen bond complex
occurs according reaction 2. The Hydrogen bond complex 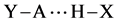 is the product. The variation in electronic energy, at 0 K, is given by Equation (3):
is the product. The variation in electronic energy, at 0 K, is given by Equation (3):
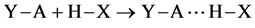 (2)
(2)
 (3)
(3)
The internal energy, at 298.15 K, corresponds to the sum of the electronic, rotational, translational and vibrational contributions, so that it’ variation can be written according Equation (4):
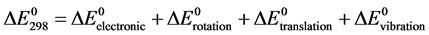 (4)
(4)
Geometry optimization of both reactants and products, gives access to all contributions (including nuclear repulsion energies). In ideal gas approximation, rotational and translational contributions are given according Equation (5):
 (5)
(5)
 includes ZPVE (Zero Point Vibrational Energy) energy, i.e. lowest vibrational level energy, due to 3N-6 normal vibrational modes (3N-5 for the linear molecules), each with frequency
includes ZPVE (Zero Point Vibrational Energy) energy, i.e. lowest vibrational level energy, due to 3N-6 normal vibrational modes (3N-5 for the linear molecules), each with frequency


As a result, internal energy variation at 298.15 K is given by Equation (7):

Enthalpy and free enthalpy variations, At 298.15 K, enthalpy and free enthalpy are respectively given by Equations (8) and (9), and entropy variation, by Equation (11):


where

2.4. Spectroscopic Parameters
Spectroscopic descriptors can serve as Hydrogen bond scale. The X-H bond linking the donor atom X and the hydrogen atom H increases or decreases according Hydrogen bond’s strength. Therefore the stretch vibration wave-num- ber can be measured. When the donor is a water molecule, the displacement 


At ONIOM(HF/6-311+G(d,p):AM1) and ONIOM(B3LYP/6-311+G(d,p):AM1) levels, frequencies of vibrator 
3. Results and Discussion
3.1. Geometric Parameters
All geometry optimizations succeeded at the two levels of computation. Examples of initial guess geometry and then optimized geometries of two hydrogen bond complexes are shown in Figure 6. Geometric parameters are given in Table 1.
Figure 6. Examples of non-optimized and optimized geometries of Hydrogen bond complexes computed at ONIOM(B3LYP/6-311+G(d,p):AM1) level.
Table 1. Geometric parameters of Hydrogen bond complexes of mycolactone A/B.
At level ONIOM(HF/6-311+G(d,p):AM1), value of the linearity angles α on the O5sp2 heteroatom equals 169.78˚ and is the closest angle to the ideal value of 180˚, the directionality angle β equals 120.10˚ and is the closest to the ideal angle of 120˚, the hydrogen bond length d equals 2.00 Å and is the smallest (Table 1). Indeed, as far as the lengths of the H bonds (distance d) are concerned, the practice is to consider a contact as a real H bond if the distance d is less than the sum of the Van der Waals radius, taking 1.52 Å [15] , and 1.2 Å [16] , respectively for a contact with the oxygen and hydrogen atoms; meaning that d ≤ 2.72 Å. It is also known that the shorter the H bond length is, the stronger it is. So, according geometric parameters, O5sp2 heteroatom is the major hydrogen bonding site. At level ONIOM(B3LYP/6-311+G(d,p):AM1), the closest angle α, 171.40˚, rather concerns the O8sp3 heteroatom, the closest angle β, 108.40˚, is found on O2sp3 (the ideal β angle for sp3 oxygen equals 109.5˚), and the shortest length d, 1.86 Å, is found for O7sp3. Values of geometric parameters computed at ONIOM (B3LYP/6-311+G(d,p):AM1) level don’t allow an undoubtedly conclusion about the major hydrogen bonding site.
3.2. Energetic Parameters
All values of enthalpy 

Spontaneity is much greater with the O5sp2 heteroatom, since the corresponding values are the lowest, i.e. −19.53 kJ/mol at both ONIOM (HF/6-311+ G(d,p):AM1) and ONIOM(B3LYP/6-311+G(d,p):AM1) levels. Therefore, the O5sp2 heteroatom gives the most stable complexes. Values of 
Table 2. Energetic parameters of Hydrogen bond complexes of mycolactone A/B. (Entropy, in J/mol・K).
Table 3. Frequency displacements Δν(O-H) (cm−1) of Hydrogen bond complexes of mycolactone A/B.
positive values of free enthalpies, meaning that there is no possibility of spontaneous reaction for these different sites. It’s noticeable that all the latter heteroatoms are sp3 hybridized. It seems that sp3 hybridized oxygen cannot be hydrogen bond major site.
3.3. Spectroscopic Parameters
All frequency shifts are positive (Table 3), corresponding to a decrease in the bond O-H vibration frequency, under hydrogen bonding process. The higher the shift Δν(O-H) is, the stronger the Hydrogen bong will be. Highest values correspond to O5sp2 heteroatom, i.e. 159.68 cm−1 and 233.20 cm−1 respectively at ONIOM(HF/6-311+G(d,p):AM1) and ONIOM(B3LYP/6-311+G(d,p):AM1) levels. Spectroscopic parameters designate then the O5sp2 oxygen atom as the major Hydrogen bonding site.
4. Conclusion
ONIOM two layers method has been successfully used to identify mycolactone A/B hydrogen bonding major sites. This compound possesses up to nine oxygen heteroatoms, all sp2 or sp3 hybridized. Geometric, energetic and spectroscopic parameters have been computed at both ONIOM(HF/6-311+G(d,p):AM1) and ONIOM(B3LYP/6-311+G(d,p):AM1) levels. Results show that sp2 hybridized oxygen is the major hydrogen bonding site and then sp3 hybridized oxygen is unlikely subject to hydrogen bonding. Two sp2 oxygen atoms, O3sp2 and O5sp2, involved in mesomerism process are the very major sites. Our analysis has permit to undoubtedly designate the major site as O5sp2 atom. Annihilating such a site would render the action of Mycobacterium ulcerans ineffective, assuming that all intermolecular interactions will occur on the same site as Hydrogen bonding.
Cite this paper
Kassi, K.F., Koné, M.G.-R., Affi, S.T. and Ziao, N. (2017) ONIOM Method Characterization of Hydrogen Bonding Sites of Mycolactone A/B, a Buruli Ulcer Toxin. Computational Chemistry, 5, 103-112. https://doi.org/10.4236/cc.2017.53009
References
- 1. WHO (2008) Buruli Ulcer: Progress Report, 2004-2008, Weekly Epidemiological Record, 83, 145-154. http://www.who.int/wer
- 2. Young, V.R. (1970) The role of skeletal and cardiac muscle in the regulation of protein metabolism. In: Munro, H.N., Ed., Mammalian Protein Metabolism, Vol. 4, Academic Press, New York, 587-674.
https://doi.org/10.1016/b978-0-12-510604-7.50018-9 - 3. Marston, B.J., Diallo M.O. and Horsburgh Jr., C.R. (1995) Emergence of Buruli Ulcer Disease in the Daloa Region of Cote d’Ivoire. American Society of Tropical Medicine and Hygiene, 52, 219-224.
https://doi.org/10.4269/ajtmh.1995.52.219 - 4. Torrado, E., Adusumilli, S., Fraga, A.G., Small, P.L. and Castro, A.G. (2007) Mycolactone-Mediated Inhibition of Tumor Necrosis Factor Production by Macrophages Infected with Mycobacterium ulcerans Has Implications for the Control of Infection. Infection and Immunity, 75, 3979-3988.
https://doi.org/10.1128/IAI.00290-07 - 5. Stanford, J.L., Revill, W.D., Gunthorpe, W.J. and Grange, J.M. (1974) The Production and Preliminary Investigation of Burulin, a New Skin Test Reagent for Mycobacterium ulcerans. Journal of Hygiene (London), 74, 7-16.
https://doi.org/10.1017/S0022172400046659 - 6. George, K., Chatterjee, M., Gunawardana, D., Welty, G., Hayman, D., Lee, J. and Small, P.R. (1999) Mycolactone: A Polyketide Toxin from Mycobacterium ulcerans Required for Virulence. Science AAAS, 283, 854-857.
https://doi.org/10.1126/science.283.5403.854 - 7. Rablen, P.R., Lockman, J.W. and Jorgensen, W.L. (1998) Ab Initio Study of Hydrogen-Bonded Complexes of Small Organic Molecules with Water. The Journal of Physical Chemistry A, 102, 3782-3797.
https://doi.org/10.1021/jp980708o - 8. Jeffrey, G.A. and Saenger, W. (1991) Hydrogen Bonding in Biological Structures. Springer-Verlag, Berlin.
https://doi.org/10.1007/978-3-642-85135-3 - 9. Desiraju, G.R. (1997) Designer Crystals: Intermolecular Interactions, Network Structures and Supramolecular Synthons. Chemical Communications, 1475-1482.
https://doi.org/10.1039/a607149j - 10. Aakeroy, C.B. (1997) An Infinite Hydrogen-Bonded Sheet in Guanidinium Trifluoromethanesulfonate. Actacryst, 53, 569-586.
- 11. Berthelot, M., Laurence, C., Safar, M. and Besseau, F. (1998) Hydrogen-Bond Basicity pKHB Scale of Six-Membered Aromatic N-Heterocycles. Journal of the Chemical Society, Perkin Transactions, 2, 283-290.
https://doi.org/10.1039/a706696a - 12. Gaussian 03, Revision C.01, Frisch, M.J., Trucks, G.W., Schlegel, H.B., Scuseria, G.E., Robb, M.A., Cheeseman, J.R., Montgomery, Jr., J.A., Vreven, T., Kudin, K.N., Burant, J.C., Mil-lam, J.M., Iyengar, S.S., Tomasi, J., Barone, V., Mennucci, B., Cossi, M., Scalmani, G., Rega, N., Petersson, G.A., Nakatsuji, H., Hada, M., Ehara, M., Toyota, K., Fukuda, R., Hasegawa, J., Ishida, M., Nakajima, T., Honda, Y., Kitao, O., Nakai, H., Klene, M., Li, X., Knox, J.E., Hratchian, H.P., Cross, J.B., Adamo, C., Jaramillo, J., Gomperts, R., Stratmann, R.E., Yazyev, O., Austin, A.J., Cammi, R., Pomelli, C., Ochterski, J.W., Ayala, P.Y., Morokuma, K., Voth, G.A., Salvador, P., Dannenberg, J.J., Zakrzewski, V.G., Dapprich, S., Daniels, A.D., Strain, M.C., Farkas, O., Malick, D.K., Rabuck, A.D., Raghavachari, K., Foresman, J.B., Ortiz, J.V., Cui, Q., Baboul, A.G., Clifford, S., Cioslowski, J., Stefanov, B.B., Liu, G., Liashenko, A., Piskorz, P., Komaromi, I., Martin, R.L., Fox, D.J., Keith, T., Al-Laham, M.A., Peng, C.Y., Nanayakkara, A., Challacombe, M., Gill, P.M.W., Johnson, B., Chen, W., Wong, M.W., Gonzalez, C. and Pople, J.A. (2004) Gaussian, Inc., Wallingford.
- 13. Desiraju, G.R. and Steiner, T. (1999) The Weak Hydrogen Bond in Structural Chemistry and Biology. The Weak Hydrogen Bond in Structural Chemistry and Biology, Oxford University Press, Oxford.
- 14. Jeffrey, G.A. and Saenger, W. (1991) Hydrogen Bonding in Biological Structures. Springer, Berlin.
https://doi.org/10.1007/978-3-642-85135-3 - 15. Jorly, J. and Eluvathingal, D.J. (2007) Red-, Blue-, or No-Shift in Hydrogen Bonds: A Unified Explanation. Journal of the American Chemical Society, 129, 4620-4632.
https://doi.org/10.1021/ja067545z - 16. Bondi, A. (1964) Van der Waals Volumes and Radii. Journal of Physical Chemistry, 68, 441-451.
https://doi.org/10.1021/j100785a001








