Food and Nutrition Sciences
Vol.4 No.9A(2013), Article ID:36478,7 pages DOI:10.4236/fns.2013.49A1018
Evaluation of the Anti-Inflammatory Activities of 5,8,11-cis-Eicosatrienoic Acid
![]()
1Department of Biotechnology, Ming Chuan University, Taoyuan, Chinese Taipei; 2Department of Marine Biotechnology and Resources, National Sun Yat-sen University, Kaohsiung, Chinese Taipei; 3Department of Life Science, Fu-Jen Catholic University, Chinese Taipei.
Email: #joannawu@mail.mcu.edu.tw
Copyright © 2013 Lun-Chung Hsu et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Received May 16th, 2013; revised June 16th, 2013; accepted June 26th, 2013
Keywords: 5,8,11-cis-Eicosatrienoic Acid; Anti-Inflammatory Activities; COX-2; iNOS; PUFAs
ABSTRACT
The main eicosanoids inflammatory mediators, prostaglandins and leukotrienes, are both generated from arachidonic acid (AA; 20:4 n-6). AA is a member of polyunsaturated fatty acids (PUFAs). Numerous studies have demonstrated that various contents of PUFAs can modulate the inflammatory responses. However, fewer studies have examined n-9- PUFAs and their effects on the inflammatory responses. In the present study, the role of 5,8,11-cis-eicosatrienoic acid (ETrA; 20:3 n-9, also called Mead acid) in the inflammatory responses has been investigated. The anti-inflammatory activities of ETrA were examined using an in vitro macrophage system and the inhibitory effect was confirmed by western blot analysis for iNOS and COX-2 expressions. The interactions between ETrA and COX-2 protein were simulated to produce a computer modeling protein-ligand complexes and the results suggest a possible mechanism for the effects of ETrA. In this study, we described a significant inhibition of the inflammatory activities initiated by ETrA. Since ETrA is a substance presented in the tissues of young animals, we therefore anticipate that ETrA can be utilized as a natural therapeutic supplement to inhibit inflammatory activities.
1. Introduction
Inflammation is a defensive reaction in response to harmful stimuli such as infectious, toxic or traumatic injuries. It is also associated with an extensive range of diseases, such as asthma, rheumatoid arthritis, cardiovascular disease and cancer [1,2]. This complex biological response is induced by the release of cytokines and inflammatory mediators. Eicosanoids are key mediators of inflammation. The main eicosanoids inflammatory mediators, prostaglandins and leukotrienes, are both generated from arachidonic acid (AA; 20:4 n-6) via the cyclooxygenase and 5-lipoxygenase pathways respectively [3,4].
AA is a member of polyunsaturated fatty acids (PUFAs). PUFAs play important roles in maintaining membrane protein function and membrane fluidity. PUFAs are also involved in cell signalling, gene expression and a variety of cellular functions [5]. Numerous studies have demonstrated that various contents of PUFAs can modulate the inflammatory responses. For example, inflammatory cells typically contain a high proportion of the n-6 PUFA and low proportions of the other 20-carbon PUFAs [6,7]. In addition, n-6 PUFAs have been found to regulate activities of inflammatory cells, the production of cytokines and the various balances within the immune system [8,9]. Similarly, studies suggested that dietary supplementation of n-3-PUFAs have beneficial impact in suppression of inflammation, especially eicosapentaenoic acid (EPA; 20:5 n-3) and docosahexaenoic acid (DHA; 22:6 n-3) [10,11]. EPA and DHA are abundant in marine fish oils and they can reduce AA content in cell membrane phospholipids to moderate the generation of eicosanoids [12,13]. Several n-3-PUFAs derived anti-inflammatory lipid mediators were identified including resolvin E1 [14] generated from EPA and protectin D1 [15] generated from DHA. Although the mechanism that n-3-PUFAs affect inflammation is still not fully defined in molecular details, the anti-inflammatory actions, activities and potential therapeutic effects of n-3 PUFAs were studied extensively [11].
On the contrary, fewer studies have examined n-9- PUFAs and their effects on the inflammatory responses. Among n-9-PUFAs, 5,8,11-cis-eicosatrienoic acid (ETrA; 20:3 n-9, also called Mead acid) has been suggested to play a role in the inflammatory responses. It can be synthesized endogeneously at extremely low level but the amount is raised during essential fatty acid (EFA) deficiency [16]. Interestingly, high levels of ETrA were discovered in the tissues of young chickens, fetal calves and newborn pigs but the amount of ETrA is reduced and replaced by n-6-PUFA progressively [17]. It has been shown that dietary supplement with ETrA can inhibit inflammatory mediator Leukotriene B4 synthesis in rats [18,19]. Additionally, an n-9 PUFA-rich diet containing ETrA suppressed platelet-activating factor (PAF) generation [20]. However, studies suggested that the anti-inflammatory effects of n-9 PUFA are different from those of dietary n-3 PUFA, that n-9 PUFA-rich diet significantly attenuated galactosamine/lipopolysaccharide induced liver injury more effectively than the n-3 PUFArich diet in the experimentally induced inflammation in mice [21] and the effects of dietary ETrA on AA content in the phospholipid subclasses and AA release were quite different from those of dietary DHA [20].
In this study, the anti-inflammatory activities of ETrA were examined using an in vitro macrophage system and the inhibitory effect was confirmed. To elucidate the possible mechanisms of the anti-inflammatory activities of ETrA, we therefore applied computational ligand docking and modeling approaches to simulate the protein-ligand complexes and demonstrate the possible interaction between ETrA and COX-2 protein.
2. Materials and Methods
2.1. Cell Culture
Murine RAW 264.7 macrophages (ATCC, No TIB-71) were cultured in Dulbecco’s modified Eagle’s medium (DMEM) containing 10% heat-inactivated FBS, 2 mM glutamine, 1 mM pyruvate, 4.5 g/Liter glucose, 50 U/mL penicillin and 50 μg/mL streptomycin at 37˚C in a humidified 5% CO2: 95% air incubator under standard conditions as described above before LPS or ETrA treatment [22].
2.2. In Vitro Lipopolysaccharide (LPS)— Stimulated Macrophage System
The anti-inflammatory activity method used was performed as described [22]. Briefly, inflammation in macrophages was induced by 16 h incubation in a medium containing Escherichia coli LPS (0.01 μg/mL, Sigma). To measure the anti-inflammatory activity, ETrA
(1, 5, 10 or 25 μM) was added to the cells 5 min before LPS challenge. The photographs of the cells were taken using a microscope digital camera system (400× magnification, Leica DMI3000 SPOT CCD) and cells were then washed with ice-cold phosphate-buffered saline (PBS), lysed in ice-cold lysis buffer (50 mM Tris, pH 7.5, 150 mM NaCl, 1% Triton X-100, 100 μg/mL phenylmethylsulfonyl fluoride, 1 μg/ml aprotinin), and then centrifuged at 20,000 g for 30 min at 4˚C. The supernatant was decanted from the pellet and retained for western blot analysis.
2.3. Western Blot Analysis for iNOS and COX-2
Western blotting was performed according to the method described in Wen et al., 2005 [23]. An equal volume of sample buffer (2% sodium dodecyl sulfate (SDS), 10% glycerol, 0.1% bromophenol blue, 2% 2-mercaptoethanol, and 50 mM Tris-HCl, pH 7.2) was added to the sample, which was then loaded onto a tricine SDS-polyacrylamide gel and electrophoresed at 150 V for 90 min. The proteins were transferred to a polyvinylidene difluoride membrane (PVDF; Immobilon-P, Millipore, 0.45 μM pore size) at 125 mA overnight at 4˚C in transfer buffer (50 mM Tris-HCl, 380 mM glycine, 1% SDS, and 20% methanol). The membrane was blocked for 50 min at room temperature with 5% non-fat dry milk in Trisbuffered saline (TTBS; 0.1% Tween 20, 20 mM TrisHCl, 137 mM NaCl, pH 7.4), and then incubated for 180 min at room temperature with antibodies against iNOS (1:1000 dilution; BD Pharmingen, San Diego, CA, USA; polyclonal antibody), COX-2 (1:1000 dilution; Cayman Chemical, USA; polyclonal antibody) proteins. The blots were then visualized in ECL solution (NEN. LifeScience, Boston, MA, USA) for 30 s and finally exposed to X-ray film (Koda X-OMAT LS; Eastman Kodak Company, Rochester, NY, USA). The membranes were re-probed with a monoclonal mouse anti-β-actin antibody (1:2500, Sigma) as the loading control. After X-film scanning, the integrated optical density of the bands was estimated (Image Pro plus 4.5 software; Media Cybernetics, Silver Spring, USA), and normalized to the background values. Relative variations between the bands of the drug-treatment samples and the LPS samples were calculated using the same image.
2.4. Software
Patchdock was used to perform ligand docking and modeling to simulate the protein-ligand complexes [24]. The structural coordinate of ETrA and the produced model was represented and viewed using Pymol Molecular Viewer (http://www.pymol.org).
2.5. Miscellaneous
Dulbecco’s modified Eagle’s medium (DMEM), fetal bovine serum (FBS), sodium pyruvate, L-glutamine, antibiotic-antimycotic solution, and trypsin-EDTA were purchased from Invitrogen Co. (Grand Island, NY, USA). Lipopolysaccharide (LPS; Escherichia coli) were purchased from Sigma Co., Ltd. (St Louis, MO, USA). All utilized electrophoretic reagents and the protein standard markers were purchased from GE Healthcare BioSciences (Piscataway, NJ, USA).
3. Results
3.1. The Effect of ETrA on LPS-Induced Macrophage
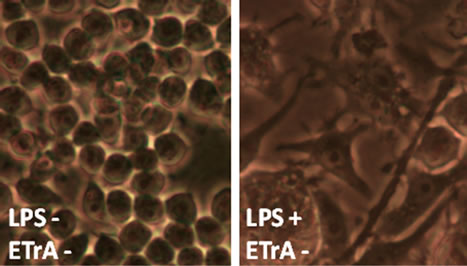
LPS is an endotoxin, which induces septic shock syndrome and stimulates the production of inflammatory mediators such as NO and prostaglandins. It has been shown that LPS-treated RAW264.7 cells increased in cell size and acquired distinct dendritic morphology [25]. To evaluate the effect of ETrA on the inflammation of cells, various concentrations of ETrA were added to the culture media to perform LPS challenge. Representative micrographs were shown in Figure 1. Macrophage cells were viable and indistinguishable from the normal cultural morphology (Figure 1(a)) when 10 or 25 μM of ETrA was applied to the LPS-induced macrophage (Figure 1(c) and (d)), while morphological changes were observed from the LPS treated macrophage cells without ETrA (Figure 1(b)). These results indicate that ETrA can help macrophages against LPS challenge.
 (a)
(a)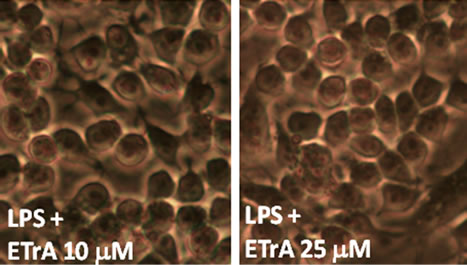 (b)
(b)
Figure 1. The effect of ETrA on LPS-induced macrophage. Representative micrographs (400× magnification) of Murine RAW 264.7 macrophages cultured by 16 hours incubation in the medium (a) without LPS; (b) containing 0.01 μg/mL LPS. To examine the effect of ETrA, (c) 10 μM of ETrA; (d) 25 μM of ETrA was added to the cell cultures before LPS challenge.
3.2. The Effect of ETrA on LPS-Induced iNOS and COX-2 Protein Expression in Macrophage
Inducible nitric oxide synthase (iNOS) and cyclooxgenase-2 (COX-2) are important enzymes that mediate inflammatory processes. Therefore, inflammation is positively correlated to the expression of the iNOS and COX-2 proteins. Compounds that are able to block or reduce LPS-induced expression of iNOS or COX-2 proteins might be beneficial in the treatment of inflammatory responses.
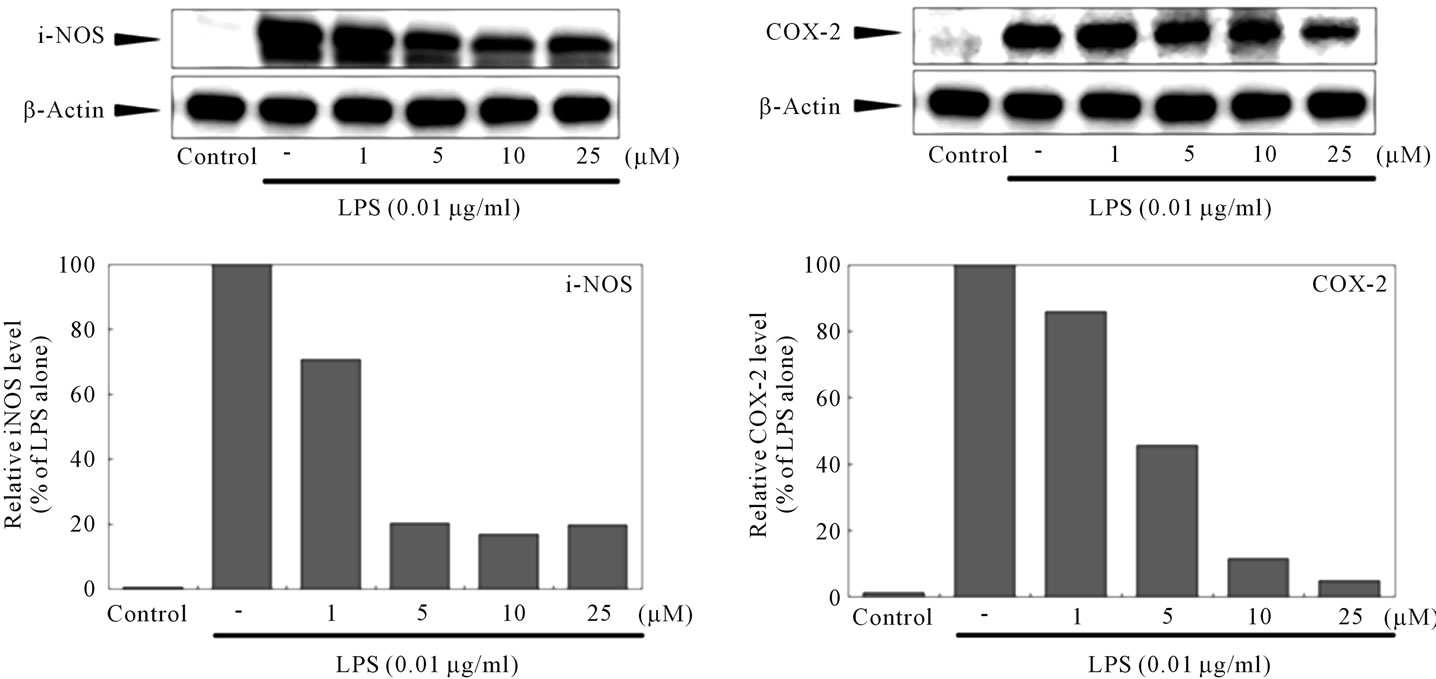
To examine the anti-inflammatory effect of ETrA, the expressions of pro-inflammatory iNOS and COX-2 in the LPS-stimulated RAW 264.7 macrophage cells were evaluated by western blot analysis (Figure 2). The dose responses for inhibition of the LPS-induced 130-kDa iNOS protein expression by ETrA are shown in Figure 2(a). At 1, 5, 10 and 25 μM doses of ETrA, the levels of iNOS protein expression were significantly reduced to 71%, 20%, 16% and 19% of the control level (i.e. the absence of ETrA) respectively. The dose responses for inhibition of the LPS-induced 71-kDa COX-2 protein expression by ETrA are shown in Figure 2(b). At 1, 5, 10 and 25 μM doses of ETrA, the levels of COX-2 protein expression were also reduced significantly to 86%, 46%, 11% and 5% of the control level respectively. From the above observations, both iNOS and COX-2 were significantly inhibited by ETrA.
3.3. Molecular Docking of ETrA to COX-2
ETrA and arachidonic acids resemble structure similarity. Both are carboxylic acid with a 20-carbon chain and a number of cis double bounds. It is speculated that ETrA can be regarded as an analog of arachidonic acid (AA). Since AA is the substrate of COX enzymes and COX-2 inhibitor is an important target of new drugs, we therefore simulated the interaction of ETrA and Cox-2 in order to investigate if ETrA antagonize AA metabolism.
PatchDock is a geometry-based molecular docking algorithm. The PatchDock method performs structure prediction of protein-protein and protein-small molecule complexes. We used the PatchDock web server to compute the scores of docked complexes. The Protein Data Bank (PDB) [26] was used for data mining of 3D coordinate of COX-2 enzyme (PDB code, 1CVU: COX-2) and the structural coordinate file of ETrA was created using the molecular graphics and modeling package, Pymol.
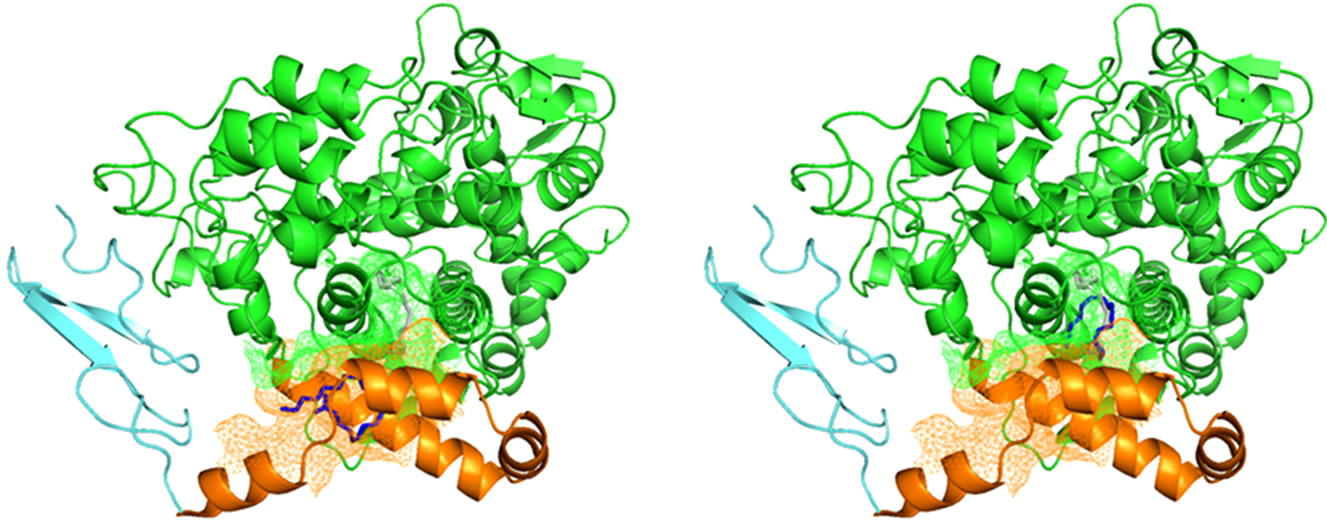
The crystallization of COX proteins revealed the structural basis at the molecular level [27,28]. COX-2 is a membrane-anchored protein that exists as dimmers and the substrate AA gains access to the active site via a hydrophobic channel (Figure 3). The initial PatchDock docking output of the full length COX-2 enzyme and ETrA showed that ETrA was docked at the entrance of the hydrophobic channel heading to the active site of COX-2 (Figure 3(a)). Considering the highly hydrophobility of the membrane binding domain of COX-2 protein may interfere the docking result of the fatty acid, we truncated the membrane binding domain of COX-2 using the molecular editing features of the Pymol and the resulting coordinate file was used for the sequential docking attempt. The automatic docking output of the COX-2 enzyme and ETrA are showed in Figure 3(b). In either cares, ETrA is located in the hydrophobic channel and thus block the catalytic domain and almost all interactions were of the hydrophobic type. The results indicated
 (a) (b)
(a) (b)
Figure 2. The effect of ETrA on pro-inflammatory iNOS and COX-2 protein expression in LPS-induced macrophage. (a) Western blots for iNOS and β-actin proteins from RAW 264.7 cells and the relative density of immunoblot; (b) Western blots for COX-2 and β-actin proteins from RAW 264.7 cells and the relative density of immunoblot. The LPS-stimulated group was taken to be 100%. Band intensities were quantified by densitometry and are indicated as a percentage change relative to that of the LPS stimulated group.
 (a) (b)
(a) (b)
Figure 3. Model for the binding of ETrA to COX-2. (a) The PatchDock docking output based on the deposited X-ray crystallographic coordinates (PDB code, 1CVU: COX-2); (b) docking output based on a membrane binding domain truncated COX-2 coordinates. The hydrophobic channel is shown in orange mesh and the active site is shown in green mesh. The blue molecular in the orange or green mesh is ETrA. The gray molecular in the active site indicates the position of AA in the co-crystal structure.
that ETrA has an excellent complementarity with COX-2. Therefore, we expect that ETrA can antagonize AA metabolism and that ETrA may act as a natural inhibitor of the functions of cyclooxygenase.
4. Discussion
Inhibitors of COX enzymes are important target of nonsteroidal anti-inflammatory drugs. Because of the side effect of the traditional COX inhibitors, the newer COX-2 selective inhibitors that do not affect COX-1 but selectively block COX-2 represent a new class of drugs provides the benefits of reducing inflammation without irritating the stomach. However, selective inhibitors of COX-2 may conduct a small risk of myocardial infarction [29]. Therefore, a natural therapeutic supplement to inhibit inflammatory activities is of importance.
In this study, the anti-inflammatory effect of ETrA was demonstrated in an LPS-stimulated macrophage in vitro assay and confirmed that ETrA significantly inhibits two important pro-inflammatory proteins, iNOS and COX-2. Although the molecular mechanisms involved in this inhibition are not understood in great detail, the ability of ETrA to inhibit iNOS expression is approximately equal to that of COX-2.
We have used PatchDock simulations to investigate the interaction of COX-2 enzyme and the fatty acid, ETrA. These results were compared with reference simulations of arachidonate to explore the effect of enzyme on substrate conformation and positioning in the active site. The cyclooxygenases are functional homodimers, each subunit contains an active site consists of a hydrophobic channel that begins at the opening of the membrane binding domain and protrudes into the body of the catalytic domain of the enzyme [30].
The program PatchDock produced a reasonable docking result as ETrA is oriented within the hydrophobic channel. Comparing this result to that of AA bounds to COX-2 [31], the molecular conformations of the two fatty acids are different from each other. In any case, ETrA is oriated in the hydrophobic channel and thus block the catalytic domain (Figure 3). This structurebased pharmacophore interprets that ETrA can antagonize AA metabolism.
Although no direct connection of the antagonized COX-2 activity and the reduced COX-2 expression level was established. These are many cases of COX-2 inhibitor results in the decreased expression of pro-inflammatory proteins, such as the well-known drug Celecoxib [32] and the chamomine extract [33], presumably due to the reduced PGE2 biosynthesis and effect the overall balance of cell signal transduction. Recent studies have suggested that different impacts of saturated and unsaturated free fatty acids on COX-2 expression and complex intracellular signals including p38, JNK and MAPKs as well as the PPAR family were involved [34,35]. Recent study on the monounsaturated oleic acid (OA; 18:1n-9), a major component of olive oil, also demonstrated an inhibitory effect on the expression of iNOS and COX-2, while the saturated stearic acid (OA; 18:0) did not inhibit expression of iNOS and COX-2 in LPS-stimulated BV2 microglia. This implies that double bonds in unsaturated fatty acids may play a critical role in the anti-inflammatory effects [36].
In summary, our results demonstrate the anti-inflammatory activities of ETrA and suggest that ETrA has the potential to be an effective therapeutic compound. Since ETrA is a normal constituent of fetal tissues [16] and it can be synthesized endogeneously during essential fatty acid deficiency [19], it can be expected to be benign or at least well-tolerated by human bodies. In addition, it has been mentioned that, comparing to other potential antiinflammatory PUFAs, ETrA has greater chemical stability because it is substantially less unsaturated than EPA or DHA [18] and this could be an important advantage of pharmaceutical application. Taken together, ETrA warrants further investigation as a potential novel anti-inflammatory agent for the treatment of inflammation associated diseases.
5. Acknowledgements
This work was supported by the grant from IsRed Pharmatheutical and Biotechnology Research Corporation (Pingtung, Taiwan) and Ming Chuan University (Grant No. S10142).
REFERENCES
- C. Nathan, “Points of Control in Inflammation,” Nature, Vol. 420, No. 6917, 2002, pp. 846-852. doi:10.1038/nature01320
- G. Srikrishna and H. H. Freeze, “Endogenous Damage-Associated Molecular Pattern Molecules at the Crossroads of Inflammation and Cancer,” Neoplasia, Vol. 11, No. 7, 2009, pp. 615-628.
- G. Folco, and R. C. Murphy, “Eicosanoid Transcellular Biosynthesis: From Cell-Cell Interactions to in Vivo Tissue Responses,” Pharmacological Reviews, Vol. 58, No. 3, 2006, pp. 375-388. doi:10.1124/pr.58.3.8
- J. Claria and M. Romano, “Pharmacological Intervention of Cyclooxygenase-2 and 5-Lipoxygenase Pathways. Impact on Inflammation and Cancer,” Current Pharmaceutical Design, Vol. 11, No. 26, 2005, pp. 3431-3447. doi:10.2174/138161205774370753
- P. C. Calder and G. C. Burdge, “Bioactive Lipids,” In: A. Nicolaou and G. Kokotos, Eds., Fatty Acids, The Oily Press, Bridgewater, 2004, pp. 1-36. doi:10.1533/9780857097934.1
- P. C. Calder, “Polyunsaturated Fatty Acids, Inflammatory Processes and Inflammatory Bowel Diseases,” Molecular Nutrition & Food Research, Vol. 52, No. 8, 2008, pp. 885-897. doi:10.1002/mnfr.200700289
- P. C. Calder, “Dietary Modification of Inflammation with Lipids,” Proceedings of the Nutrition Society, Vol. 61, No. 3, 2002, pp. 345-358. doi:10.1079/PNS2002166
- M. L. Salem, “Systemic Treatment with n-6 Polyunsaturated Fatty Acids Attenuates EL4 Thymoma Growth and Metastasis through Enhancing Specific and Non-Specific Anti-Tumor Cytolytic Activities and Production of TH1 Cytokines,” International Immunopharmacology, Vol. 5, No. 6, 2005, pp. 947-960. doi:10.1016/j.intimp.2004.12.013
- J. Thanasak, V. P. Rutten, J. T. Schonewille, A. Hoek, A. C. Beynen, J. P. Noordhuizen and K. E. Muller, “Effect of a Dietary n-6 Polyunsaturated Fatty Acid Supplement on Distinct Immune Functions of Goats,” Journal of Veterinary Medicine Series A, Vol. 5, No. 1, 2004, pp. 1-9. doi:10.1111/j.1439-0442.2004.00595.x
- P. C. Calder, “n-3 Polyunsaturated Fatty Acids, Inflammation, and Inflammatory Diseases,” The American Journal of Clinical Nutrition, Vol. 83, No. 6, 2006, pp. 1505S-1519S.
- J. W. Fetterman Jr. and M. M. Zdanowicz, “Therapeutic Potential of n-3 Polyunsaturated Fatty Acids in Disease,” American Journal of Health System Pharmacy, Vol. 66, No. 13, 2009, pp. 1169-1179. doi:10.2146/ajhp080411
- C. L. Shen, J. Peterson, O. L. Tatum and D. M. Dunn, “Effect of Long-Chain n-3 Polyunsaturated Fatty Acid on Inflammation Mediators during Osteoblastogenesis,” Journal of Medicinal Food, Vol. 11, No. 1, 2008, pp. 105-110. doi:10.1089/jmf.2007.540
- D. S. Kelley, P. C. Taylor, G. J. Nelson, P. C. Schmidt, A. Ferretti, K. L. Erickson, R. Yu, R. K. Chandra and B. E. Mackey, “Docosahexaenoic Acid Ingestion Inhibits Natural Killer Cell Activity and Production of Inflammatory Mediators in Young Healthy Men,” Lipids, Vol. 34, No. 4, 1999, pp. 317-324. doi:10.1007/s11745-999-0369-5
- H. Seki, Y. Tani and M. Arita, “Omega-3 PUFA Derived Anti-Inflammatory Lipid Mediator Resolvin E1,” Prostaglandins & Other Lipid Mediators, Vol. 89, No. 4, 2009, pp. 126-130. doi:10.1016/j.prostaglandins.2009.03.002
- N. G. Bazan, “Neuroprotectin D1-Mediated Anti-Inflammatory and Survival Signaling in Stroke, Retinal Degenerations, and Alzheimer’s Disease,” Journal of Lipid Reseach, Vol. 50, Suppl. 9, 2009, pp. S400-S405. doi:10.1194/jlr.R800068-JLR200
- H. D. Adkisson 4th, F. S. Risener Jr., P. P. Zarrinkar, M. D. Walla, W. W. Christie and R. E. Wuthier, “Unique Fatty Acid Composition of Normal Cartilage: Discovery of High Levels of n-9 Eicosatrienoic Acid and Low Levels of n-6 Polyunsaturated Fatty Acids,” FASEB Journal, Vol. 5, No. 3, 1991, pp. 344-353.
- T. Hamazaki, N. Suzuki, R. Widyowati, T. Miyahara, S. Kadota, H. Ochiai and K. Hamazaki, “The Depressive Effects of 5,8,11-Eicosatrienoic Acid (20:3n-9) on Osteoblasts,” Lipids, Vol. 44, No. 2, 2009, pp. 97-102. doi:10.1007/s11745-008-3252-8
- M. J. James, R. A. Gibson, M. A. Neumann and L. G. Cleland, “Effect of Dietary Supplementation with n-9 Eicosatrienoic Acid on Leukotriene B4 Synthesis in Rats: A Novel Approach to Inhibition of Eicosanoid Synthesis,” The Journal of Experimental Medicine, Vol. 178, No. 6, 1993, pp. 2261-2265. doi:10.1084/jem.178.6.2261
- L. G. Cleland, R. A. Gibson, M. A. Neumann, T. Hamazaki, K. Akimoto and M. J. James, “Dietary (n-9) Eicosatrienoic Acid from a Cultured Fungus Inhibits Leukotriene B4 Synthesis in Rats and the Effect Is Modified by Dietary Linoleic Acid,” The Journal of Nutrition, Vol. 126, No. 6, 1996, pp. 1534-1540.
- S. Watanabe, M. Doshi, K. Akimoto, Y. Kiso and T. Hamazaki, “Suppression of Platelet-Activating Factor Generation and Modulation of Arachidonate Metabolism by Dietary Enrichment with (n-9) Eicosatrienoic Acid or Docosahexaenoic Acid in Mouse Peritoneal Cells,” Prostaglandins & Other Lipid Mediators, Vol. 66, No. 2, 2001, pp. 109-120. doi:10.1016/S0090-6980(01)00152-6
- M. Doshi, S. Watanabe, T. Niimoto, H. Kawashima, Y. Ishikura, Y. Kiso and T. Hamazaki, “Effect of Dietary Enrichment with n-3 Polyunsaturated Fatty Acids (PUFA) or n-9 PUFA on Arachidonate Metabolism in Vivo and Experimentally Induced Inflammation in Mice,” Biological & Pharmaceutical Bulletin, Vol. 27, No. 3, 2004, pp. 319-323. doi:10.1248/bpb.27.319
- Y. H. Jean, W. F. Chen, C. Y. Duh, S. Y. Huang, C. H. Hsu, C. S. Lin, C. S. Sung, I. M. Chen and Z. H. Wen, “Inducible Nitric Oxide Synthase and Cyclooxygenase-2 Participate in Anti-Inflammatory and Analgesic Effects of the Natural Marine Compound Lemnalol from Formosan Soft Coral Lemnalia cervicorni,” European Journal Pharmacology, Vol. 578, No. 3, 2008, pp. 323-331. doi:10.1016/j.ejphar.2007.08.048
- Z. H. Wen, G. J. Wu, Y. C. Chang, J. J. Wang and C. S. Wong, “Dexamethasone Modulates the Development of Morphine Tolerance and Expression of Glutamate Transporters in Rats,” Neuroscience, Vol. 133, No. 3, 2005, pp. 807-817. doi:10.1016/j.neuroscience.2005.03.015
- D. Schneidman-Duhovny, Y. Inbar, R. Nussinov and H. J. Wolfson, “Patch Dock and Symm Dock: Servers for Rigid and Symmetric Docking,” Nucleic Acids Research, Vol. 33, Suppl. 2, 2005, pp. W363-W367. doi:10.1093/nar/gki481
- R. K. Saxena, V. Vallyathan and D. M. Lewis, “Evidence for Lipopolysaccharideinduced Differentiation of RAW264.7 Murine Macrophage Cell Line into Dendritic like Cells,” Journal of Bioscience, Vol. 28, No. 1, 2003, pp. 129-134. doi:10.1007/BF02970143
- H. M. Berman, J. Westbrook, Z. Feng, G. Gilliland, T. N. Bhat, H. Weissig, I. N. Shindyalov and P. E. Bourne, “The Protein Data Bank,” Nucleic Acids Research, Vol. 28, No. 1, 2000, pp. 235-242. doi:10.1093/nar/28.1.235
- P. J. Loll, D. Picot and R. M. Garavito, “The Structural Basis of Aspirin Activity Inferred from the Crystal Structure of Inactivated Prostaglandin H2 Synthase,” Nature Structural Biology, Vol. 2, No. 8, 1995, pp. 637-643. doi:10.1038/nsb0895-637
- R. G. Kurumbail, A. M. Stevens, J. K. Gierse, J. J. McDonald, R. A. Stegeman, J. Y. Pak, D. Gildehaus, J. M. Miyashiro, T. D. Penning, K. Seibert, P. C. Isakson and W. C. Stallings, “Structural Basis for Selective Inhibition of Cyclooxygenase-2 by Anti-Inflammatory Agents,” Nature, Vol. 384, No. 6610, 1996, pp. 644-648. doi:10.1038/384644a0
- A. Risser, D. Donovan, J. Heintzman and T. Page, “NSAID Prescribing Precautions,” American Family Physician, Vol. 80, No. 12, 2009, pp. 1371-1378.
- M. G. Malkowski, S. L. Ginell, W. L. Smith and R. M. Garavito, “The Productive Conformation of Arachidonic Acid Bound to Prostaglandin Synthase,” Science, Vol. 289, No. 5486, 2000, pp. 1933-1937. doi:10.1126/science.289.5486.1933
- J. R. Kiefer, J. L. Pawlitz, K. T. Moreland, R. A. Stegeman, W. F. Hood, J. K. Gierse, A. M. Stevens, D. C. Goodwin, S. W. Rowlinson, L. J. Marnett, W. C. Stallings and R. G. Kurumbail, “Structural Insights into the Stereochemistry of the Cyclooxygenase Reaction,” Nature, Vol. 405, No. 6782, 2000, pp. 97-101. doi:10.1038/35011103
- G. D. Basu, L. B. Pathangey, T. L. Tinder, S. J. Gendler and P. Mukherjee, “Mechanisms Underlying the Growth Inhibitory Effects of the Cyclo-Oxygenase-2 Inhibitor Celecoxib in Human Breast Cancer Cells,” Breast Cancer Research, Vol. 7, No. 4, 2005, pp. R422-435. doi:10.1186/bcr1019
- J. K. Srivastava, M. Pandey and S. Gupta, “Chamomile, a Novel and Selective COX-2 Inhibitor with Anti-Inflammatory Activity,” Life Sciences, Vol. 85, No. 19, 2009, pp. 663-669. doi:10.1016/j.lfs.2009.09.007
- A. Kadotani, Y. Tsuchiya, H. Hatakeyama, H. Katagiri and M. Kanzaki, “Different Impacts of Saturated and Unsaturated Free Fatty Acids on COX-2 Expression in C(2)C(12) Myotubes,” American Journal of Physiology Endocrinology and Metabolism, Vol. 297, No. 6, 2009, pp. E1291-E1303. doi:10.1152/ajpendo.00293.2009
- C. D. Funk, “Prostaglandins and Leukotrienes: Advances in Eicosanoid Biology,” Science, Vol. 294, No. 5548, 2001, pp. 1871-1875. doi:10.1126/science.294.5548.1871
- Y. T. Oh, J. Y. Lee, J. Lee, H. Kim, K. S. Yoon, W. Choe and I. Kang, “Oleic Acid Reduces LipopolysaccharideInduced Expression of iNOS and COX-2 in BV2 Murine Microglial Cells: Possible Involvement of Reactive Oxygen Species, p38 MAPK, and IKK/NF-Kappa B Signaling Pathways,” Neuroscience Letters, Vol. 464, No. 2, 2009, pp. 93-97. doi:10.1016/j.neulet.2009.08.040.
NOTES
*These authors contributed equally to this work.
#Corresponding author.

