American Journal of Plant Sciences
Vol.5 No.18(2014), Article
ID:49145,7
pages
DOI:10.4236/ajps.2014.518290
Genetic Diversity of Five Different Lily (Lilium L.) Species in Lithuania Revealed by ISSR Markers
Judita Žukauskienė1, Algimantas Paulauskas1, Judita Varkulevičienė2*, Rita Maršelienė2, Vytautė Gliaudelytė1
1Faculty of Natural Sciences, Vytautas Magnus University, Kaunas, Lithuania
2Kaunas Botanical Garden, Vytautas Magnus University, Kaunas, Lithuania
Email: *j.varkuleviciene@bs.vdu.lt
Copyright © 2014 by authors and Scientific Research Publishing Inc.
This work is licensed under the Creative Commons Attribution International License (CC BY).
http://creativecommons.org/licenses/by/4.0/



Received 8 June 2014; revised 15 July 2014; accepted 8 August 2014
ABSTRACT
To study the genetic diversity and structure of lily (Lilium L.), we collected 35 samples from Vytautas Magnus University Kaunas Botanical Garden, and analyzed their mutual Simple Sequence Repeat (ISSR) molecular markers. For genetic analysis of lily we chose the lily 6 markers. ISSR data revealed a relatively high level of genetic diversity at the different levels of the group, with 95% of polymorphic loci, effective number of alleles of 1.21, the average expected heterozygosis of 0.15 and Shannon’s information index of 0.26. ANOVA analysis and UPGMA-dendrogram suggested a hierarchical structure between species.
Keywords:ISSR, Lily Species, Genetic Variability, Lithuania

1. Introduction
Species of lilies were used as ornamental plant for thousands of years. In the genera there are about 110 of wild species. Nevertheless the first hybrids derived from the 19th century [1] . Classifications of lily (Lilium L.) species were set before several centenaries. DNA technology has helped to identify and distinguish the lily species and also detected their genetic diversity [2] [3] . In Lithuania, just naturally grows wild lily (Lilium martagon), other species are introduced [4] .
Identification of hybrids using DNA analysis was started at 20th century, nine decades. Natures of hybrids were confirmed analyzing DNA of parental forms and their offspring using RAPD method [5] .
Plant population genetics study systematic biology and conservation, is a big interest in the genetic diversity in natural populations [6] [7] . Reflecting on the evolutionary history of the species, genetic diversity in natural populations is very useful for those who study plant population genetics, systematic biology and conservation [6] [7] . Plant species across the wider geographical region can be divided into different ecotypes; this division could be influenced by ecological factors, which are more important for the geographical variation of plants [8] . Therefore, successful management, preservation and reestablishment of sustainable populations of rare, threatened, or endangered species depend on a complete understanding of the species, including levels and structure of genetic variation [9] [10] .
In Lithuania, quite a lot various lily (Lilium L.) species are introduced. However, not all species of our weather conditions are suitable. That’s why we have to do morphologic and genetic researches to learn out plants resources. It is also important to compare the results with the global genetic results and the plant of origin. In Lithuania there weren’t any lily genetic researches done.
Molecular markers are recognized as a good method for the identification of plant species [11] [12] . Accordingly, these markers could be helpful in the breeding aimed at new cultivar development through early detection and screening of plants that have so far escaped cultivation [13] . Also to improve plants taxonomy or in fine cultivars number in collections [14] [15] , genetic diversity of plants was detected using various methods. Such methods as restriction fragment length polymorphism (RFLP), analysis of chloroplast DNA (cpDNA) [3] random amplified polymorphic DNA (RAPD), simple sequence repeat (SSR) and intersimple sequence repeat (ISSR) have enabled researchers to investigate genetic diversity among various plant species across natural populations [16] -[20] . The efficiency of these methods has already been demonstrated in phylogenetic and population genetic studies [3] . Local populations of traditional cultivars provide a valuable resource for plant breeding as well as for the preservation of genetic diversity [21] . The exploration, evaluation, and conservation in situ and ex situ of genetic diversity in natural populations are imperative to guarantee sustainable development [22] . Random amplified polymorphic DNA (RAPD) method has often been used for estimating genetic variation within and among plant populations, cultivars and species, and also was used for determining the hybridism of interspecific hybrids in the genus Lilium [2] [3] . Recently, the usefulness of intersimple sequence repeat (ISSR) markers that amplify the genomic sequence between two simple sequence repeats (SSR or microsatellites) using anchored simple sequence repeat (ASSR) primers has also been reported [23] .
To find out recourses of lily in Lithuanian, it is necessary to evaluate the variability using of genetic methods. It is also very important to evaluate and compare the genetic variation between species and wild populations of wild samples. There are some problems with naturally grown wild lily populations and introduced Lilium species, varieties and hybrids. There could be several origin possibilities; they could be sowed out from introduced lilies, also there could be hybrids.
After molecular DNA analysis to choose lily we can more efficiently and faster to make the biggest part of selection work processes. Also there would be a possibility during plant regeneration process to control proportions of variability and stability.
2. Materials and Methods
2.1. Sampling
Thirty five samples of lily individuals of 3 different species (Lilium martangon L. (18 samples), Lilium pardalinum Kellogg (4 samples), Lilium henryi Baker (4 samples)), three samples of varieties Lilium henryi var. citrinum and 6 samples of hybrid—Lilium hybridum hort. (originator Petras Balčikonis) were chosen from Kaunas Botanical Garden of Vytautas Magnus University natural fields (meadows) and collection. Fresh young leaf material—was sampled according to blossom time. DNA extraction was done using Thermo scientific extraction kit.
2.2. DNA Analysis Using ISSR Method
Six primers were used for ISSR analysis (Applied bio systems) (Table 1). DNA amplification reaction was performed in 25 μl reaction mix made of: 12.5 μl 2 × PCR Master Mix (0.625 U/μl Tag DNA polymerase; 2 mM MgCl2; 0.2 mM each dNTP) (Thermo scientific, Lithuania), 2 μl primer (concentration 10 pmol/μl), 6.5 μl twice distilled water and 4 μl (25 ng/ μl) DNA. PCR reaction was performed in a thermo cycler (Master cycler®
Table 1. Attributes of the primers used for generating Lilium ISSR amplification and number of bands per primer.
Eppendorf, Germany) under the following conditions: initial denaturation for 5 min at 94˚C, 45 cycles of denaturation for 30 s at 94˚C, primers annealing for 30 s at 50˚C - 55˚C (depends on primer), extension for 60 s at 72˚C followed by a final extension for 10 min at 72˚C (Table 1).
Of 6 ISSR primer pairs [24] approved for Lilium, 4 primer pairs were chosen PCR reactions involving five 3’-ASSR primers [25] 2 primer pairs were chose (Table 1). For analyses of 35 different lilies samples using 6 ISSR primers were used.
Polymorphisms were scored as presence (1) or absence (0) after resolution on 2% agarose gels prepared in 1XTBE buffer. A standard molecular marker—Gene RulerTM DNA Ladder Plus (MBI Ferments, Lithuania), was used to determine molecular weights of the amplified bands. Gels were examined visually under UV light using ethidium bromide. The gels were photographed and documented with Hero lab transliuminator and Win32 (Hero lab, Germany) system.
2.3. Data Analysis
Calculation of the observed number of alleles, Nei’s [26] gene diversity (H), Shannon’s Information Index (I) and generation of a Nei’s genetic distance based. Relationships among different lily species were evaluated using a dendrogram based on Nei and Li (1979) genetic distances. It was generated by the UPGMA (unweight pair group method) cluster analysis method. Dendrogram were carried out with POPGENE V 1.31 software. A principal coordinate analysis (PCoA) was performed with the Gen AlEx 6 program [27] .
3. Results
The ISSR profile showed a high polymorphism in analyzed lily species. The primers yielded a total of 80 highly reproducible ISSR bands ranging from 125 to 1000 bp in size (Figure 1). From the summary in Table 2, 95% (76/80) of the bands were polymorphic, indicating a high level of genetic diversity among all analyzed groups (3 species, 1 varietet and 1 hybrid) and while a moderate polymorphism within groups was found in the range of 21.69% - 84.34%. Highest polymorphism was detected in natural Lilium martagon group (84%). In other research groups polymorphism was considerably lower. From the average expected heterozygosis (He), a parameter regarded as an index of gene diversity [28] was estimated to be 0.21 ± 0.005 at the species level, ranging from 0.009 to 0.14 (Table 2).
The number of alleles per locus (Na), effective number of alleles per locus (Ne), and the Shannon information index (I) at the group level were 1.95 ± 0.22, 1.21 ± 0.23, and 0.26 ± 0.17, respectively. As another measure of gene diversity, Shannon’s information index (I) was much higher in wild lily (0.244 ± 0.18) (Table 2) lowest in Lilium henry f. citrinum 0.13 ± 0.27. Lilium exhibited considerable genetic differentiation among species (GST = 0.211), representing 11% genetic variability among analyzed groups and 89% within groups. The fixation index (FST = 0.107) among analyzed groups showed a certain degree of genetic subdivision among the 3 lily species 1varietet and 1 hybrid. In the model with two hierarchical levels, both variance components among and within groups were highly significant (P < 0.001; 107 permutations). Furthermore pair-wise Nm at the species level was 0.9. Nm predicts the relative importance of gene flow and genetic drift based on the observed pattern of genetic differentiation in a population.
To further determine phylogenetic relationships among the 3 different species (Lilium martangon L., Lilium pardalinum Kellogg, Lilium henryi Baker), 1 varieties Lilium henryi var. citrinum and 1 hybrid—Lilium hybri-
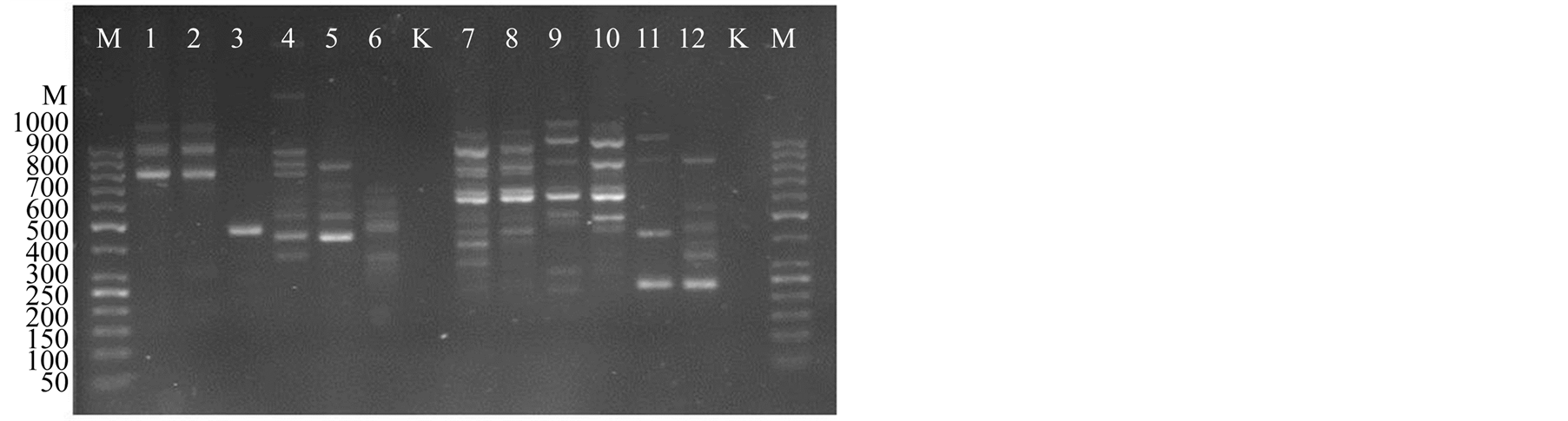
Figure 1. Spectrum of UBC 15 ISSR markers with different lily samples (M—marker, 1 - 12—Lilium samples, K—control).
Table 2. Genetic variability in the Lilium species studied in this work.
dum hort. (originator P. Balčikonis), UPGMA clustering was carried out using Nei’s unbiased genetic distance matrix. The dendrogram (Figure 1) clearly showed a differentiation in among hybrid from Lithuania (P. Balčikonis) and species, because the 3 species (Lilium martagon, Lilium pardalinum and Lilium henryi) and Lilium henryi var. citrinum were clustered into separate group, while other researched groups were clustered in another group. These findings suggest that the maternal origins of the individuals in bigger group are closely related. Therefore, UPGMA analysis is useful to show genetic structure in different groups (Figure 2).
4. Discussion
This study offered the first detailed analysis of the genetic diversity and population structure for 3 different lily species 1 varieties and 1 hybrid grown in Lithuania based on ISSR molecular markers that have been proven to be valuable for the determination of genetic diversity. The ISSR-PCR identified relatively high genetic diversity (PPB, He and I), considerable genetic differentiation among species (GST) and the gene flow (Nm) at the species level. Compared with other Lilium species, polymorphism of analyzed groups is higher than that of 13 Lilium species and of 16 interspecific hybrids (16%) [24] , and that of the Asiatic hybrid lily-Montreux and Connecticut King (52.4%) [23] , and also that of the Lilium tsingtauence (85%) [24] and higher than in the Lilium maculatum var. bukosanense (88.2%) [3] .
The effective gene flow per generation (Nm) of all analyzed groups is higher than one successful migrant per generation (0.5), possibly to overcome the diversifying effect of random drift [24] . Lilium is a long-lived perennial grassy species, which should enable the species to maintain a high level of genetic diversity. Furthermore, its high genetic diversity could come from seedling establishment in the early period. Our observations suggest that events that potentially reduce genetic diversity and increase genetic differentiation among populations such as genetic drift or inbreeding might have not occurred yet with the restricted distribution. Based on Nei’s gene diversity and AMOVA analyses for 3 lily species, 1 varieties and 1 hybrid genetic differentiation among species (GST) and fixation index (FST) also signify a genetic structure. The non-random temporal or spatial distribution of genetic variations of alleles and genotypes is often referred to as the genetic structure of a population [29] .
The level and distribution of genetic variability among and within groups can be shaped with a great number of factors including pollen and seed dispersal, successional stages, geographic distribution range, adult density,
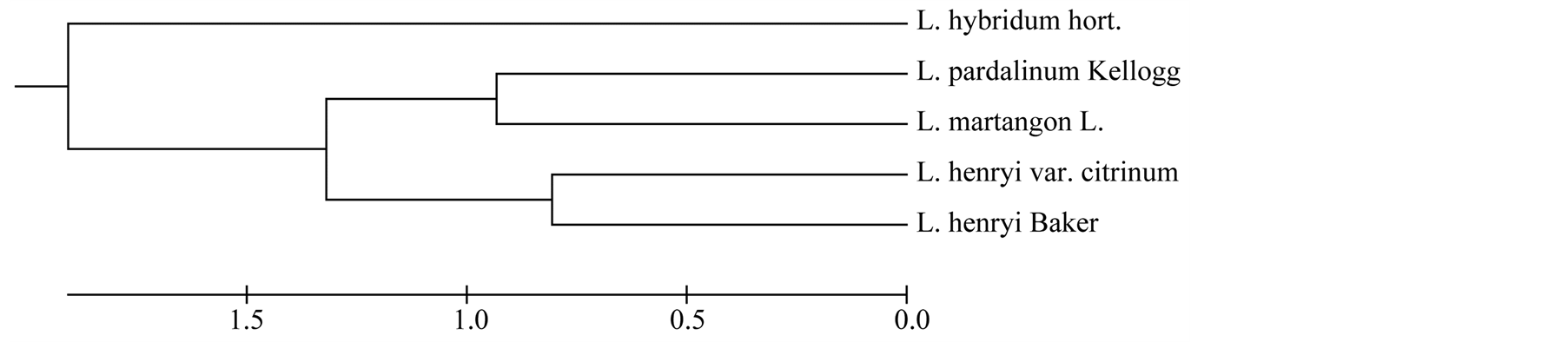
Figure 2. UPGMA (unweight pair-group method with arithmetic average) cluster based on genetic distances.
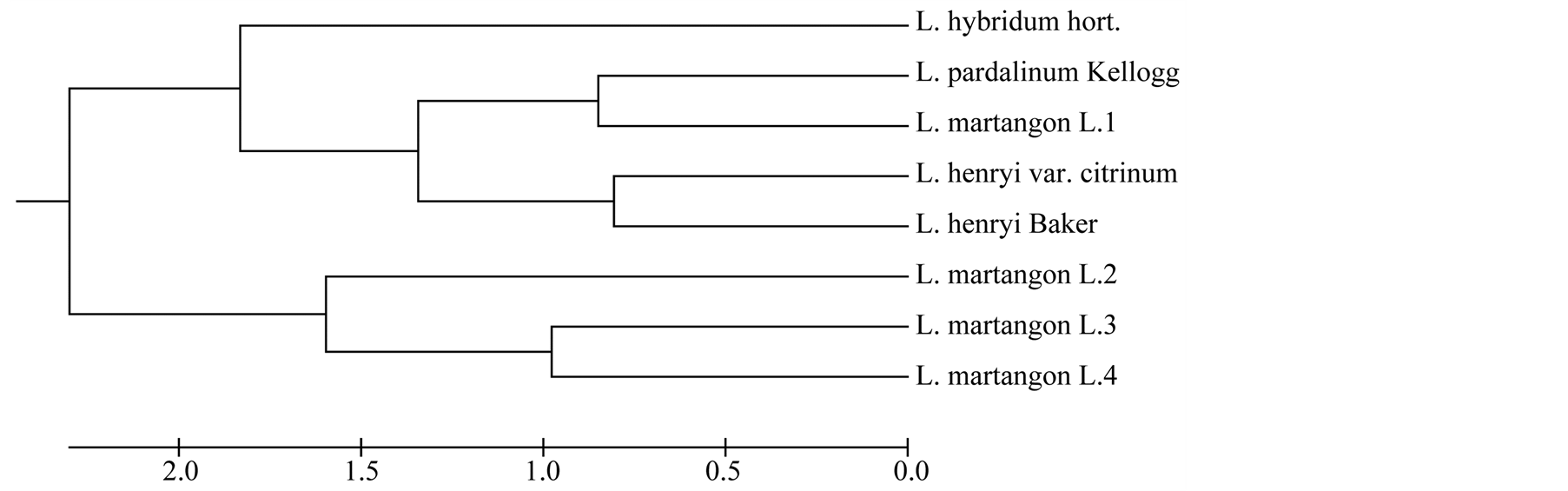
Figure 3. UPGMA (unweight pair-group method with arithmetic average) cluster based on analyzed sample genetic distances.
mating system, colonization history, and natural selection [24] .
We hope that the genetic diversity and structure of analyzed five different lily groups reported here offer important information for this endangered species (Figure 3).
Checking theory about Lilium martagon L. origin in Lithuania populations we could think about three different origin possibilities. First is connected to 1 group of wild lily. This group has genetic similarities with species Lilium pardalinum Kellogg. The secon Lilium martagon L. group is in different UPGMA group from species and hybrids. This could reveal about purity of this group.
5. Conclusion
Talking about Kaunas wild lily populations, we could accept one Lilium martagon L. group and think about mixing their origin.
References
- Jefferson-Brown, M.J. and Howland, H. (1995) The Gardener’s Guide to Growing Lilies. Timber Press, Portland.
- Wen, C.S. and Hsiao, J.Y. (2001) Altitudinal Genetic Differentiation and Diversity of Taiwan Lily (Lilium longiflorum var. formosanum; liliaceae) Using RAPD Markers and Morphological Characters. International Journal of Plant Science, 162, 287-295. http://dx.doi.org/10.1086/319588
- Arzate-Fernández, A.M., Miwa, M., Shimada, T., Yonekura, T. and Ogawa. K. (2005) Genetic Diversity of Miyamasukashi-Yuri (Lilium maculatum Thunb. var. bukosanense), an Endemic and Endangered Species at Mount Buko, Saitama, Japan. Plant Species Biology, 20, 57-65. http://dx.doi.org/10.1111/j.1442-1984.2005.00124.x
- Varkuleviciene, J. and Marseliene, R. (2010) Specific Lilies Use in Green Plantation. Journal name, 1, 208-213. http://www.krastotvarka.vhost.lt/documents/2010_35.pdf
- Beata, P. (2007) RAPD-Molecular Marker in Detection of Hybrids la Hybrids. Scientific Works of the Lithuanian Institute of Horticulture and Lithuanian University of Agriculture, 26, 246-250
- Guo, W., Wang, R., Zhou, S., Zhang, S. and Zhang, Z. (2003) Genetic Diversity and Clonal Structure of Phragmites austral is in the Yellow River Delta of China. Biochemistry Systematic and Ecology, 31, 1093-1109. http://dx.doi.org/10.1016/S0305-1978(03)00032-2
- Stevens, L., Salomon, B. and Sun, G.L. (2007) Microsatellite Variability and Heterozygote Excess in Elymus trachycaulus Populations from British Columbia in Canada. Biochemical Systematics and Ecology, 35, 725-736. http://dx.doi.org/10.1016/j.bse.2007.05.017
- Yahiaoui, S., Igartua, E., Moralejo, M., Ramsay, L., Molina-Cano, J.L., Ciudad, F.J., Lasa, J.M., Gracia, M.P. and Casas, A.M. (2008) Patterns of Genetic and Eco Geographical Diversity in Spanish Barleys. Theoretical and Applied Genetics, 116, 271-282. http://dx.doi.org/10.1007/s00122-007-0665-3
- Luan, S.S., Chiang, T.Y. and Gong, X. (2006) High Genetic Diversity vs. Low Genetic Differentiation in Nouelia insignis (Asteraceae), a Narrowly Distributed and Endemic Species in China, Revealed by ISSR Fingerprinting. Annals of Botany, 98, 583-589. http://dx.doi.org/10.1093/aob/mcl129
- Balode, A. (2013) Diversity of the Martagon Lily (Lilium martagon L.) in Latvia. In: Scripta Horti Botanici Universitatis Vytauti Magni, VDU, Kaunas, 17, 25-35.
- Pounders, C.T., Rinehart, T. and Sakhanokho, H. (2007) Evaluation of Interspecific Hybrids between Lagerstroemia indica and Lagerstroemia speciosa. HortScience, 42, 1317-1322.
- Wadl, P.A., Skinner, J.A., Dunlap, J.R., Rees, S.M., Rinehart, T.A., Pantalone, V.R. and Trigiano, R.N. (2009) Honeybee-Mediated Controlled Pollination in Cornus florida and C. kousa Intra and Interspecific Crosses. HortScience, 44, 1527-1533.
- Dean, D., Wang, X., Klingeman, W.E. and Ownley, B.H. (2011) Screening and Characterization of 11 Novel Microsatellite Markers from Viburnum dilatatum. HortScience, 46, 1456-1459.
- Gismondi, A., Fanali, F., Martínez Labarga, J.M., Grilli Caiola, M. and Canini, A. (2013) Crocus sativus L. Genomics and Different DNA Barcode Applications. Plant Systematics and Evolution, 299, 1859-1863. http://dx.doi.org/10.1007/s00606-013-0841-7
- Gismondi, A., Impei, S., Di Marco, G., Crespan, M., Leonardi, D. and Canini, A. (2014) Detection of New Genetic Profiles and Allelic Variants in Improperly Classified Grapevine Accessions. Genome, 57, 111-118. http://dx.doi.org/10.1139/gen-2013-0218
- Archak, S., Gaikwad, A.B., Gautam, D., Rao, E.V.V.B., Swamy, K.R.M. and Karihaloo, J.L. (2003) DNA Fingerprinting of Indian Cashew (Anacardium occidentale L.) Varieties Using RAPD and ISSR Techniques. Euphytica, 130, 397-404. http://dx.doi.org/10.1023/A:1023074617348
- Deshpande, A.U., Apte, G.S., Bahulikar, R.A., Lagu, M.D., Kulkarni, B.G., Suresh, H.S., Singh, N.P., Rao, M.K.V., Gupta, V.S., Pant, A. and Ranjekar, P.K. (2001) Genetic Diversity across Natural Populations of Three Montane Plant Species from the Western Ghats, India Revealed by Intersimple Sequence Repeats. Molecular Ecology, 10, 2397-2408. http://dx.doi.org/10.1046/j.0962-1083.2001.01379.x
- Knox, E.B. and Palmer, J.D. (1999) The Chloroplast Genome Arrangement of Lobelia thuliniana (Lobeliaceae): Expansion of the Inverted Repeat in an Ancestor of the Companulales. Plant Systematics and Evolution, 214, 49-64. http://dx.doi.org/10.1007/BF00985731
- Gabrielsen, T.M. and Brochmann, C. (1998) Sex after All: High Levels of Diversity Detected in the Arctic Clonal Plant Saxifraga cernua Using RAPD Markers. Molecular Ecology, 7, 1701-1708. http://dx.doi.org/10.1046/j.1365-294x.1998.00503.x
- Wolfe, A.D., Xiang, Q. and Kephart, S.R. (1998) Assessing Hybridization in Natural Populations of Penstemon (Scrophulariaceae) Using Hypervariable Intersimple Sequence Repeat (ISSR) Bands. Molecular Ecology, 7, 1107-1125. http://dx.doi.org/10.1046/j.1365-294x.1998.00425.x
- Nevo, E. (1998) Genetic Diversity in Wild Cereals: Regional and Local Studies and Their Bearing on Conservation ex Situ and in Situ. Genetic Resources and Crop Evolution, 45, 355-370. http://dx.doi.org/10.1023/A:1008689304103
- Kölliker, R., Herrmannm, D., Boller, B. and Widmer, F. (2003) Swiss Mattenklee Landraces, a Distinct and Diverse Genetic Resource of Red Clover (Trifolium pratense L.). Theoretical and Applied Genetics, 107, 306-315. http://dx.doi.org/10.1007/s00122-003-1248-6
- Yamagishi, M., Abe, H., Nakano, M. and Nakatsuka, A. (2002) PCR-Based Molecular Markers in Asiatic Hybrid Lily. Scientia Horticulturae, 96, 225-234. http://dx.doi.org/10.1016/S0304-4238(02)00095-X
- Guo, W., Jeong, J., Kim, Z., Renqing, W., Kim, E. and Kim, S. (2011) Genetic Diversity of Lilium tsingtauense in China and Korea Reveals by ISSR Markers and Morphological Characters. Biochemical Systematics and Ecology, 39, 352-360. http://dx.doi.org/10.1016/j.bse.2011.05.002
- Abe, H., Nakano, M., Nakatsuka, A., Nakayama, M., Koshioka, M. and Yamagishi, M. (2002) Genetic Analysis of Floral Anthocyanin Pigmentation Traits in Asiatic Hybrid Lily Using Molecular Linkage Maps. Theoretical and Applied Genetics, 105, 1175-1182. http://dx.doi.org/10.1007/s00122-002-1053-7
- Nei, M. (1973) Analysis of Gene Diversity in Subdivided Populations. Proceedings of the National Academy of Sciences of the United States of America, 70, 3321-3323. http://dx.doi.org/10.1073/pnas.70.12.3321
- Peakall, R. and Smouse, P.E. (2006) Genalex 6: Genetic Analysis in Excel. Population Genetic Software for Teaching and Research. Molecular Ecology Notes, 6, 288-295. http://dx.doi.org/10.1111/j.1471-8286.2005.01155.x
- Nei, M. and Li, W.H. (1979) Mathematical Model for Studying Genetic Variation in Terms of Restriction Endonucleases. Proceedings of the National Academy of Sciences of the United States of America, 76, 5269-5273. http://dx.doi.org/10.1073/pnas.76.10.5269
- Escudero, A., Iriondo, J.M. and Torres, M.E. (2003) Spatial Analysis of Genetic Diversity as a Tool for Plant Conservation. Biological Conservation, 113, 351-365. http://dx.doi.org/10.1016/S0006-3207(03)00122-8
NOTES

*Corresponding author.


