American Journal of Plant Sciences
Vol.5 No.3(2014), Article ID:42677,8 pages DOI:10.4236/ajps.2014.53049
Response of Arabidopsis Clones to Toxic Compounds Released by Various Rhizoctonia Species
![]()
Plant Protection Institute of the Centre for Agricultural Research of the Hungarian Academy of Sciences, Budapest, Hungary.
Email: *gyoros@aim.com
Copyright © 2014 Gyula Oros et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. In accordance of the Creative Commons Attribution License all Copyrights © 2014 are reserved for SCIRP and the owner of the intellectual property Gyula Oros et al. All Copyright © 2014 are guarded by law and by SCIRP as a guardian.
Received December 9th, 2013; revised January 14th, 2014; accepted January 26th, 2014
KEYWORDS
Arabidopsis; Rhizoctonia; Toxin; Athelia; Ceratobasidium; Ceratorhiza; Thanatephorus; Waitea
ABSTRACT
Response of 3 Arabidopsis clones to 41 strains of eight Rhizoctonia species was studied in model experiments. The seed germination was decelerated in most of the cases, although the inhibitory effect varied within large limits. The pre-emergence damping off and root neck rot leading to damping off were the most frequent symptoms of disease syndrome caused by toxic metabolites. The clone transformed with cDNA clone overexpressing gstf4 gene exhibited significantly improved tolerance as compared to parental one, meanwhile the sensitivity of Dmannose pyrophosphorylase/mannose-1-pyrophosphatase deficient clone dramatically increased. Strains of R. solani of AG-2, AG-4 and AG-7 and Athelia rolfsii produced the most toxic metabolites, however, no strict relationships were revealed between taxonomic position of Rhizoctonia strains and toxicity of their metabolites.
1. Introduction
Damping off caused by various soil-borne pathogenic microbes is a well-known phenomenon in plant cultivation. The traditional control measures (seed dressing, soil fumigation) cannot be applied in large scale due to high risk of irreversible environmental pollution and possible toxic residuals in food and forage. Moreover, the soilborne fungi (mainly Fusarium spp, Pythium, Rhizoctonia) can attack several hundred cultivated plant species during all vegetation period that makes impossible the long lasting protection of plants with selectively acting synthetic compounds or antibiotic preparations. Recently, none of the marketed fungicides is translocated basipetally, which counteracts their use after development of foliage against root parasiting fungi. Although, Rhizoctonia species are not among the top ten pathogens [1], economic importance of their control is increasing from severe to catastrophic yield losses reported from main wheat cultivating areas [2-4]. Changes in agricultural practices of cereals with special regard to minimum or no tillage cultivation [5,6] led to increased importance of Rhizoctonia infections [7,8]. The most plausible method is the breeding of tolerant varieties.
The Rhizoctonia species are abundant in soils as mutualistic members of microbial consortium associated with plants. Their relationship may change from symbiosis (as in various orchids) to parasitism. The stunted growth can be most frequently observed, which might be related to the effect of Rhizoctonia toxins although few evident experimental proof has been demonstrated yet [9]. Presence of parasitic hyphae can also be commonly seen even in symptomless tissues, and in normal conditions does not cause visible disease symptoms either in phyllosphere or in roots. However, when environmental stress factors overwhelm the homeostatic regulation of plant, the disease syndrome may develop at various degrees. Both biotic and abiotic stress manifests as an oxidative stress [10], thus by scavenging free radicals an active antioxidative defense system comprising enzymatic and non-enzymatic antioxidants reduces the level of oxidative stress in plant cells and counteracts evolution of disease. Regulation of substrate pools such as glutathione or ascorbic acid is the key element of cell protection [11], and the capacity of the biochemical path sustaining the oxido-reductive potential of cells has crucial importance in elimination of harmful free radicals [12] and confers in resistance to pathogens [13,14]. The high level of ascorbic acid reduces the disease severity [15], while the deficiency increases the sensitivity to environmental stress factors [12]. Glutathione S-transferases play an important role in adaptation of various organisms to adverse factors [16-18], and in detoxification of electrophilic compounds either exogenic or of natural origin [19,20]. The formation of glutathione adducts catalyzed by GST is usually the first step of deterioration [21]. The level of GST increased in okra [22] and ricinus [23] seedlings damaged by soil-borne Rhizoctonia, suggesting activation of glutathione conjugation system in defense mechanisms counteracting invasion and eliminating grave consequences of the infection. The deteriorative capacity of plant and associated microbes is related to the tolerance of host plant to the pathogen. Thus, the selection of tolerant plants to these pathogenic fungi is important for quality food production as well.
There is an urgent need in sustainable management strategy to combat damages induced by soil habiting Rhizoctonia complex. No major resistance genes to this pathogen have been identified so far in spite of increasing efforts in studies of physiology and genetics of Rhizoctonia/host interaction [8]. Currently, the genetic engineering is widely used in breeding technologies providing genetically modified (GM) plants with a very high resistance against abiotic and biotic factors. Some plants have been objected for increasing tolerance to unfavorable environmental conditions [24] and remediative capacity [25] by gene engineering that can be a determinative factor for producing healthy food [26].
The genomes and physiology of Arabidopsis thaliana are broadly studied, and numerous mutans or transgenic clones are described in literature. The Arabidopsis model system greatly help us to understand specific genetic and biochemical processes. The aim of this work was to get information on the effect of toxic metabolites of Rhizoctonia species complex on seed germination of Arabidopsis.
The scoring of disease incidence and severity meet difficulties in the case of Rhizoctonias as some symptoms of disease syndrome can be induced by toxic metabolites of Rhizoctonia alone [9]. In agar cultures, the above two effects can be separated during first days of germination, however, later the damage caused by parasiting hyphae secondary infections of germlings caused by various microbes of spermosphere can mask the visual effect of Rhizoctonia toxins even in agar cultures. To avoid the altering effects, the observations were limited up to five days, as the aim of this work was the evaluation of sensitivity of Arabidopsis to toxic metabolites of Rhizoctonia released to the medium.
2. Materials and Methods
Response of three A. thaliana clones to metabolites of 41 Rhizoctonia strains five genera (Athelia, Ceratobasidium, Ceratorhiza, Thanatephorus and Waitea) was compared in model experiment applying vertical agar diffusion technique. The seeds were surface sterilized for 2 min in 70% alcohol followed by 5 min in commercial bleach (0.5% hypochlorite) then washed five times in sterile water and diluted in sterile water slightly solidified with 0.1% agar before stratification for 48 hours at 4˚C in the dark.
2.1. Plant Material
Seeds of Arabidopsis thaliana wild type (Col-5; #195), the vitamin C deficient mutant vtc1-1 [12,27] and a transgenic clone overexpressing glutathione S-transferase [28] (gst f; #193) were used for all exepriments.
Col-5 [wild type] and vtc1-1 seeds were obtained from Arabidopsis stock Center (www.arabidopsis.info).
A-193—For gstf, ecotype: Col-5 was transformed with cDNA clone overexpressing gstf4 gene (Zea mays, EMBL: U12679/X79515; Uniprot: P46420) driven by cauliflower mosaic virus 35S promoter. cDNA were introduced by using the floral dip transformation method [29] using the hygromycin phosphotransferase (hpt) gene as selectable marker. The pCAMBIA1301 binary vector was used for transformation.
vtc1 ecotype is deficient in the function of GDP-mannose pyrophosphorylase/mannose-1-pyrophosphatase. This enzyme provides GDP-mannose, which is used for cell wall carbohydrate biosynthesis, protein glycosylation and for ascorbate (vitamin C) biosynthesis. Total acorbic acid content in vtc1 clone is about 40% compared to wild type. Earlier work revealed that vtc1 mutant has increased resistance against Pseudomonas syringae or Peronospora parasitica [30] but more sensitive to Alternaria brassicicola [15].
2.2. Test Fungi
Rhizoctonia strains were originated of different locations and various hosts:
Rhizoctonia solani Kühn strains of CBS collection: B-415 (AG-1, Pinus sylvestris L., Canada, CBS 522.96), B-432 (AG-2, Daucus carota L., Netherlands, CBS 326.84 ), B-446 (AG-3, Solanum tuberosum L., Spain, CBS 117248), B-417 (AG-4, Citrus sp., Argentina, CBS 341.35), B-430 (AG-4, Phaseolus sp., England, CBS 340.51), B-418 (AG-5, Zea mays L., Netherlands, CBS 339.84), B-419 (AG-6, Erigeron canadensis (L.) Cronquist, CBS 137.82, USA), B-420 (AG-7, soil, Japan, CBS 214.84), B-421 (AG-8, Triticum aestivum L., Australia, CBS 101782), B-422 (AG-9, S. tuberosum, USA, CBS 970.96), B-423 (AG-10, T. aestivum, USA, CBS 971.96), B-424 (AG-11, Lupinus angustifolius L., Australia, CBS 974.96), B-434 (AG-E, Malus sp., Netherlands, CBS 340.84).
R. solani strains isolated in Hungary: B-151 (S. tuberosum cv Desirée), B-245 (Allium cepa L., China, Henan), B-246 (S. tuberosum cv Gül Baba), B-399 (Sesamum indicum L.), B-403 and B-404 (S. tuberosum cv Ella), B-409 (Hibiscus rosa-chinensis L., Lybia, Tripoli), B-410 (S. tuberosum cv Kisvárdai rózsa), B-411 (S. tuberosum cv Desirée), B-412 (S. tuberosum cv Cleopatra), B-413 (Malus sp. L.), B-433 (Festuca arundinacea Schreb.), B-444 (Viola × wittrockiana Gams.), B-446 (S. tuberosum cv Százszorszép), B-521 (Impatiens balsamina L.), B-522 (Oxalis triangularis A.St.-Hil., Oxalidales, Oxalidaceae), B-573 (T. aestivum)., B-548 (Phragmipedium schlimii (Lind. & Reich.f.) Rolfe, Orchidaceae), B-553 (Phalenopsis Orchidaceae), B-557 (Dendrobium × Phalenopsis hybrid, Orchidaceae), B-560 (Doritis pulcherissima Lindley, Orchidaceae).
R. fragariae S. Husain & W. E. McKeen 1963 (teleomorph: Ceratorhiza fragariae (S. S. Husain & W. E. McKeen) R. T. Moore 1987), B-438 (Fragaria × ananassa Duchense, Canada, CBS 335.62).
R. cerealis E. P. Hoeven 1977 (teleomorph: Ceratobasidium cereale D. I. Murray & Burpee 1984 Basidiomycota, Cantharellales), B-447 (T. aestivum, Germany, CBS 559.77).
R. stahlii Burgeff 1936 (teleomorph: Mycelium radicis Platantherae chloranthae (Custer) Rchb.), Thanatephorus Donk sp., Basidiomycota, Cantharellales, Germany), B-441 (P. chlorantha (Custer) Rchb.), Asparagales, Orchidaceae, Germany, CBS 119.92).
R. ramicola W. A. Weber & D. A. Roberts (teleomorph: Ceratorhiza ramicola (W. A. Weber & D. A. Roberts) R. T. Moore 1987, Basidiomycota, Cantharellales), B-427 (Pittosporum tobira (Thunb.) W. T. Aiton, Asparagales, Orchidaceae, Florida, USA, CBS 400.51).
R. carotae Rader 1948 (teleomorph: Athelia arachnoidea (Berk.) Jülich 1972, Basidiomycota, Atheliales), B-440 (D. carota, USA, CBS 464.48).
Athelia rolfsii (Curzi) C. C. Tu & Kimbr. 1978 (Basiiomycota, Atheliales), B-442 (S. tuberosum L., Italy, CBS 464.48).
R. zeae Voorhees 1938 (teleomorph: Waitea circinata Warcup and P. H. B. Talbot 1962), B-405 (F. arundinacea, Hungary).
The strains were maintained on potato dextrose agar (Merck, Darmstadt, Germany) amended with 2 g soya peptone L44, 0.5 g Trypton T L43 (Oxoid, Basingstoke, sUK) and 1 g yeast extract L21 (Oxoid).
2.3. Assay for Toxin Sensitivity
The culture medium consisted of agar No. 1. (Oxoid) (11 gl−1), soya peptone (5 gl−1), yeast extract (3 gl−1), starch (5 gl−1), glucose (5 gl−1) glycerol (2 gl−1), Na-bglycerophosphate × 6H2O (0.5 gl−1), Tween 80 (0.25 gl−1), KH2PO4 (0.5 gl−1), Na2HPO4 × 7H2O (0.5 gl−1), KCl (0.25 gl−1), MgSO4 (0.15 gl−1), CaCl2 (0.15 gl−1), FeSO4 × 5H2O (0.025 gl−1), CuSO4 × 5H2O (5 mgl−1), MnSO4 × H2O (5 mgl−1), Ni(NO3)2 × 6H2O (1 mgl−1) and CoCl2 × 6H2O (1 mgl−1) and vitamins pyridoxine×HCl (1 mgl−1), thiamine × HCl (10 mgl−1), riboflavine (1 mgl−1) and nicotinamide (20 mgl−1). The agar plates (5 ml medium in 50 mm diameter) were centrally inoculated, and incubated at ambiental conditions for 21 days. The thickness of medium reduced up to 1 mm, and the mycelium completely covered the surface. These cultures were covered with 5 ml agar solution (10 gl−1), and after 8 hours stratified seeds were dropped over the surface (25 - 30 seeds in 200 microl). The plates were incubated in ambiental conditions, and the germination was followed up to 10 days. Number of germinated seeds was counted in each morning, moreover, the germlings were studied under dissecting microscope to check their response to metabolites of Rizoctonia strains released into the medium. The half time of germination and ratio of survivors were calculated.
2.4. Data Analysis
Fisher’s test was applied to evaluate significance of differences between variant at p = 0.05 level. The results of counting of germinated/dormant and healthy/diseased individuals were calculated as percentages. The half time of germination (hours requested for germination of the 50% of seeds) was determined by linear regression analysis, where the percent values were transformed into probits. Similarity of Arabidopsis clones and relationships between parameters of assessment were investigated by Canonic Correlation Analysis (CCA). Statistical functions of Microsoft Office Excel 2003 (Microsoft, Redmond, USA) and Statistica5 program (StatSoft 5.0., Tulsa, USA) were used for analysis of data. The graphical presentation of result of data analysis was edited uniformly in MS Office Power Point 2003.
3. Results
3.1. Germination of Seeds
The number of dropped seeds varied between 25 ± 6 and they disseminated in a spot with diameter between 10 - 14 mm on agar plate. On control plates 96% - 100% of them have germinated indicating the good quality of seed material. Seeds of each clone germinated synchronously within 48 hours. The half time of germination approached by linear regression varied within −12 and 18 hours, where the 48 hours of stratification should be taken into consideration for negative value. Predominant majority of Rhizoctonia strains did not break through the agar layer in first five days of incubation, thus most of seeds in each variant could germinate and develop cotyledons until meeting with hyphae.
The hyphae of strains started to colonize the surface after six days of incubation, but their attack cannot be accurately evaluated due to airborne contaminations and bacteria arising of spermosphere. The first visual symptom of toxicose was the formation of brown spot on root-neck of the germlings even before full opening of cotyledons that was followed with damping off within 24 hours. The killed seeds became dark brown. Seedlings that survived the effect of toxic metabolites were subsequently destroyed by fungal attack rapidly in the case of most aggressive strains (B-151, B-245, B-420, B-427 and B-557), while in the case of other strains the variation in number of survivors within series was too high for correct assessment of pathogenicity, thus the data analysis was limited to 5 days of incubation. In some cases germination started after 24 - 72 hours lag phase (Table 1).
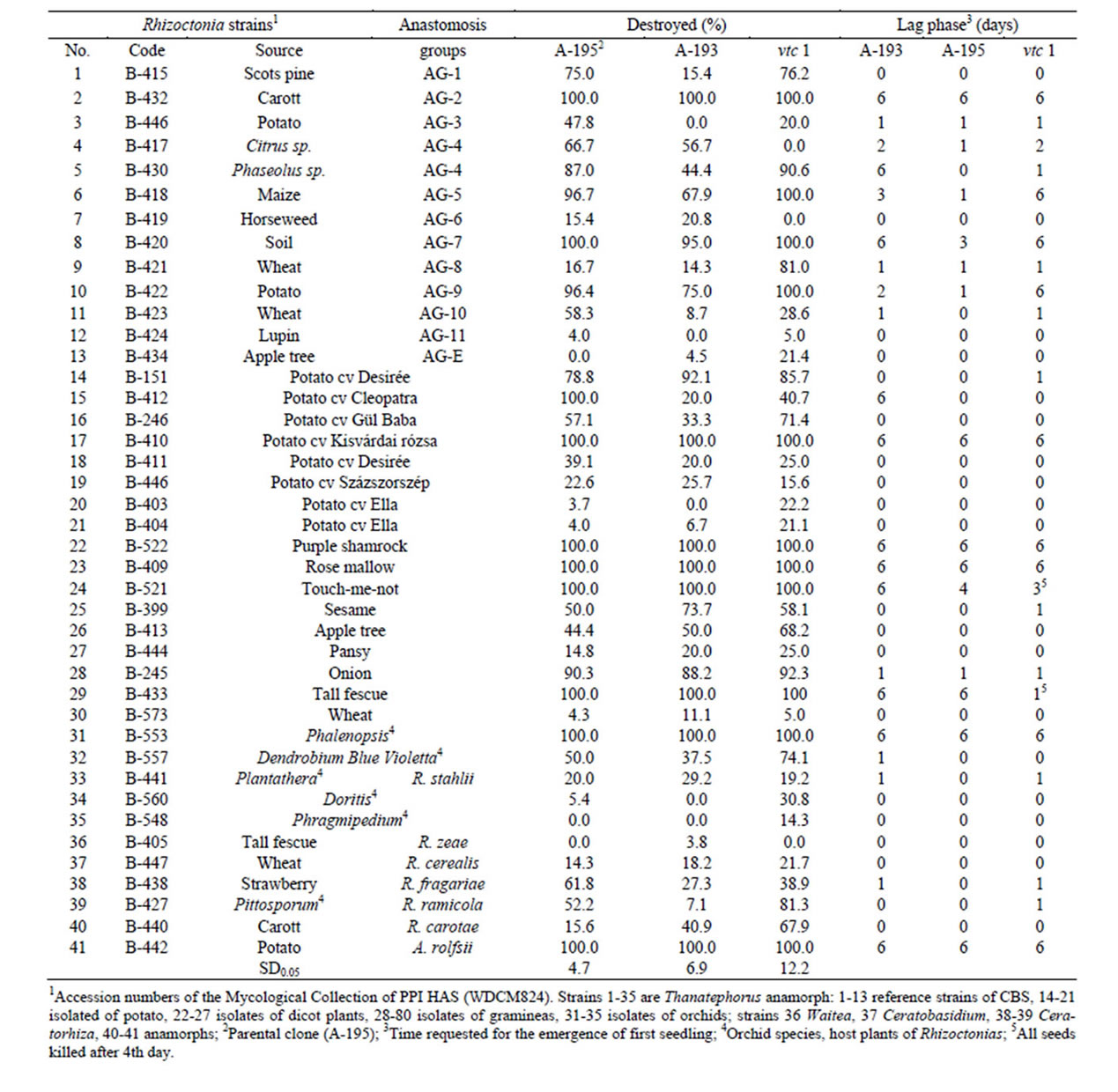
Table 1. Toxic effect of Rhizoctonia metabolites to germinating Arabidopsis seeds.
3.2. Toxic Effect of Metabolites
Response of germinating seeds was evaluated in two aspect. First the lethal effect, i.e., the ratio of non germinated seeds was calculated of recorded data at 5th days of incubation for each strain (Table 1). The reaction to metabolites of Rhizoctonias varied within large limit and strain dependents manner. The toxicity of AG-3 (B-446 and B-433) and AG-4 (B-417 and B-430) strain pairs proved to be significantly different. Similarly, great differences were revealed between strains isolated of the same host, the toxicity of potato originated ones varied between 4% and 100%. The genetic manipulation altered significantly the response Arabidospsis clones to Rhizoctonia metabolites (Table 2). The clone A-193 with improved capacity of glutathione conjugation system (GCS) tolerated at higher level the toxins than both parental (A-195) and vitamin C deficient mutant (vtc1), while the latter slightly got behind of A-195.
The other aspect of evaluation of toxic effects was the delay of germination. Toxins of ten strains, except A. rolfsii (B-442) all of them Thanatephorus anamorphs, killed the seeds before seed coat rupture. The introduction of gstf4 gene (A-193) counteracted to this effect in three cases (Table 2), that means the overexpression of GST increased the vitality of Arabidopsis. In some cases the germination started after 24 - 72 hours of lag phase (Table 1). In this case, the overexpression of GST unlocked the inhibitory effect as well. The lag phase of germination poorly correlated with lethal effect (Canonic R2 = 0.389, χ2 = 13.75, p = 0.13), that means the retardation of emergence is not strictly connected to response of seedlings to toxic effect in subsequent stages of germination (Table 3). The half time of germination (Table 4) varied within large limits (13 - 150 hr), nevertheless, this parameter could be calculated surprisingly high precision (the regression could be fitted in p < 0.05 in all cases) that underlines again the good quality of seed material as well as indicates the reproducibility of toxin production following the method applied in our experiments.
The presence gstf4 gene (A-193) accelerated the germination at about 20 hr as related to parental and vitamin C deficient clones, although, this protective effect varied in Rhizoctonia strain dependent manner.
The CCA revealed that the effect on half time of germination correlated slightly better to lethal effect on seedlings than to lag phase (Table 3). Plotting the strains as canonic scores (plots are not shown) no grouping was revealed in the case of HT vs LE and HT vs LP. However, two groups was formed on plot LE vs LP (Table 4). The major one (Group A, 22 strains) did not show any correlation, but in the other one (Group B, 19 strains) the correlation between lag phase and lethal effect was significant (R2 = 0.75). The difference between this two
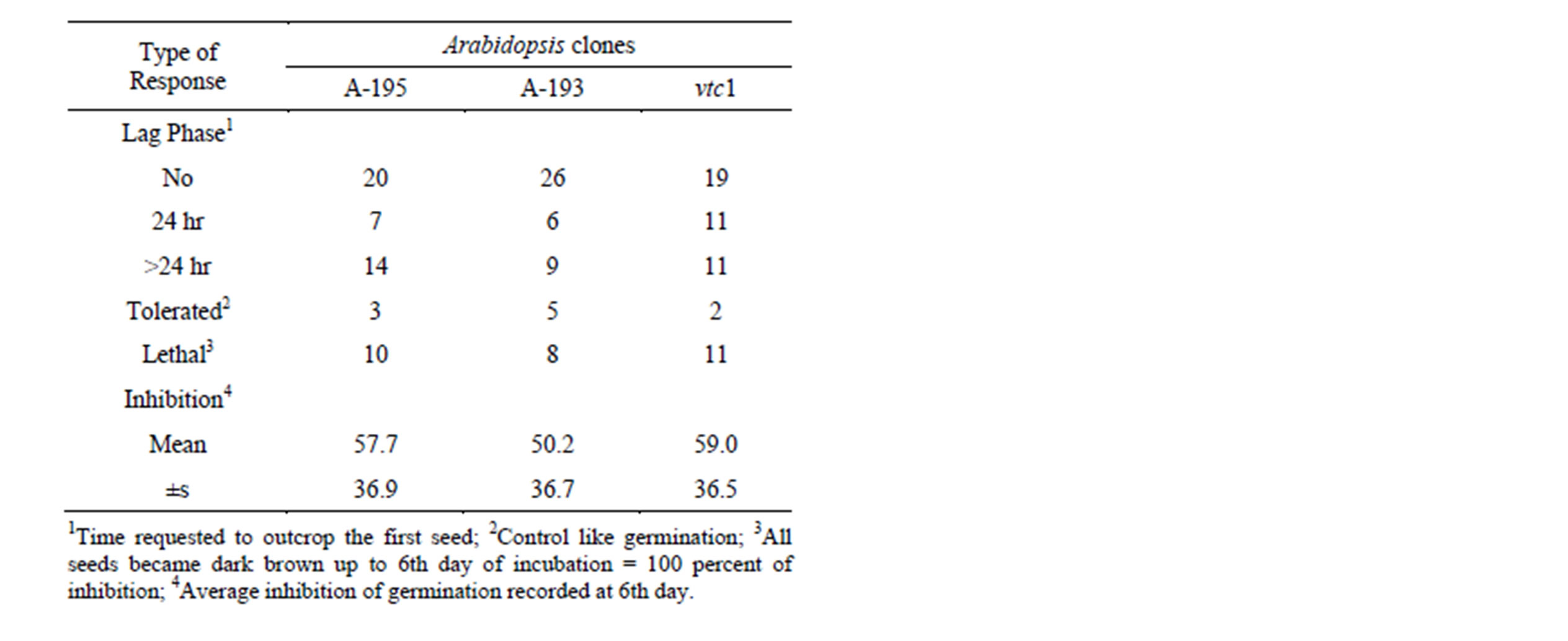
Table 2. Response of germinating seeds of Arabidopsis to toxic metabolites of Rhizoctonia strains.

Table 3. Similarity of response as evaluated by various parameters of germination.
groups is in the selective effect of their toxins to Arabidopsis clones. The parental clone (A-195) exhibited slightly higher sensitivity to B group than to A (57% and 50%, resp.), while the genetically altered A-193 and vtc1 clones responded exhibited higher sensitivity to A group (48% and 59%, resp.) than to B group (41% and 54%, resp.).
No significant relationships is between selective toxicity of taxonomic position, source and origin of strains could be revealed.
The seedlings of A-193 clone differed at higher extent than vtc1 one of parental A-195 in tolerance to Rhizoctonia toxins, but the response of germinating seeds of vtc1 clone altered at higher extent of parental A-195 that those of A-193 (Table 5). The Person’s coefficient between responses of A-193 and vtc1 as evaluated by half time of germination was lower than in scoring of lethal effect (0.553 < 0.873) that indicates changes in sensitivity spectrum of clones during the seed germination.
4. Discussion
Rhizoctonia species are well known soil borne pathogens, which are habiting mainly in rhizosphere, however, they may survive as saprobionts in the upper layer of the soil
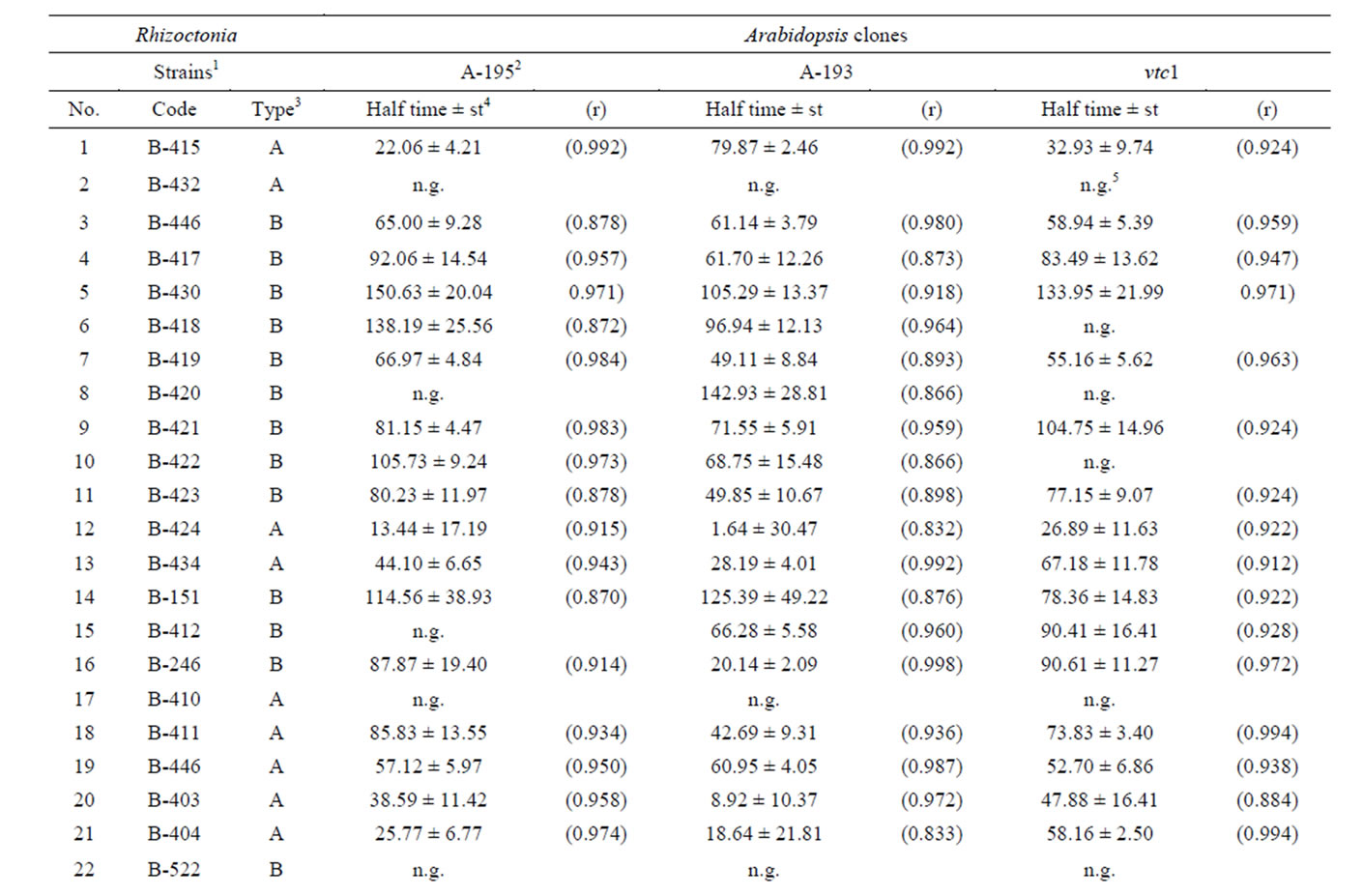

Table 4. Influence of Rhizoctonia metabolites on germination of Arabidopsis seeds.

Table 5. Similarity of response of Arabidopsis clones to Rhizoctonia toxins.
The sub-matrix up to diagonal (A) shows similarity (Pearson’s coeffients) of clones as evaluated on the base of incidence of diseased individuals (Table 1), while the sub-matrix down the diagonal (B) on the base of delay of germination (Table 3).
forming a mycelial web, thus the undisturbed soil enhance the risk of the infection of young roots. The disease syndrome may evolve rapidly to a fatal consequence in formerly symptomless host (damping off and wilting). Our data support the assumption of crucial role of mycotoxins in evolution of disease syndrome [9,22]. Like to series of host plants [8,10,31] the strain dependent response of A. thaliana clones to Rhizoctonia was demonstrated in our experiments as well. The strains examined highly diverged in toxic properties. The poor correlation between reaction of Arabidopsis seedlings in subsequent steps indicates the qualitative differences in composition of toxic substances released into the medium. Further research requested for both analysis of this complex and elucidation of toxic properties of each component.
The fact of increased tolerance of transgenic clone bearing overexpressed alien GST and increased sensitivity of ascorbic acid deficient clone evidently support the importance of elimination of free radicals in response of plants to pathogen attack.
5. Conclusions
Rhizoctonia strains of various taxonomic position release medium metabolites toxic to Arabidopsis thaliana. Neither taxonomic position nor origin of strains was related to inhibitory effect of their toxins.
The genetic manipulation of A. thaliana significantly altered the response of germinating seeds to Rhizoctonia toxins. Contrary to ascorbic acid deficiency the overexpression of alien glutathione-S-transferase significantly improved the tolerance of seeds.
The improvement of tolerance of transgenic plant to wild range of soil-borne Rhizoctonia strains was demonstrated here, illustrating the usefulness of A. thaliana as a model for research of biochemical background of resistance to soil-borne pathogens. In our opinion, this approach can significantly accelerate the progress in breeding of wheat tolerant to brown patch disease
Acknowledgements
The research work was supported by The Hungarian Scientific Research Fund (Grant K-67688).
REFERENCES
- R. Dean, J. A. L. Van Kan, Y. A. Pretorius, K. E. Hammond-Kosack, A. Di Pietro, P. D. Spanu, J. J. Rudd, M. Dickman, R. Kahmann, J. Ellis and G. D. Foster, “The Top 10 Fungal Pathogens in Molecular Plant Pathology,” Molecular Plant Pathology, Vol. 13, No. 4, 2012, pp. 414-430. http://dx.doi.org/10.1111/j.1364-3703.2011.00783.x
- M. G. Cromey, R. C. Butler, H. J. Boddington and A. R. Moorhead, “Effects of Sharp Eyespot on Yield of Wheat (Triticum aestivum) in New Zealand,” New Zealand Journal of Crop and Horticultural Science, Vol. 30, No. 1, 2002, pp. 9-17. http://dx.doi.org/10.1080/01140671.2002.9514194
- G. J. Kövics and N. Lőrincz, “Causal Agents of StemBase Diseases of Winter Wheat in Eastern Hungary,” Analele Universitátii din Oradea. Tom. VII. Scientific Communication Session: Partea I-a. Fascicula Agricultură- Horticultura, Vol. 7, 2001, pp. 37-44.
- M. Anees, V. Edel-Hermann and C. Steinberg, “Build up of Patches Caused by Rhizoctonia solani,” Soil Biology & Biochemistry, Vol. 42, No. 10, 1998, pp. 1661-1672.
- K. L. Schroeder and T. C. Paulitz, “Effect of Inoculum Density and Soil Tillage on the Development and Severity of Rhizoctonia Root Rot,” Phytopathology, Vol. 98, No. 3, 2008, pp. 304-314.
- R. Mrabet, “Effects of Residue Management and Cropping Systems on Wheat Yield Stability in a Semiarid Mediterranean Clay Soil,” American Journal of Plant Sciences, Vol. 2, No. 2, 2011, pp. 202-216.
- W. W. Bockus and J. P. Shroyer, “The Impact of Reduced Tillage on Soil-borne Plant Pathogens,” Annual Review of Phytopathology, Vol. 36, No. 1, 1998, pp. 485-500. http://dx.doi.org/10.1146/annurev.phyto.36.1.485
- G. Oros, Z. Naár and D. Magyar, “Susceptibility of Wheat Varieties to Soil-Borne Rhizoctonia Infection,” American Journal of Plant Sciences, Vol. 4, No. 11, 2013, pp. 2240- 2258. http://dx.doi.org/10.4236/ajps.2013.411277
- F. E. Bartz, N. J. Glassbrook, D. A. Danehower and M. A. Cubeta, “Modulation of the Phenylacetic Acid Metabolic Complex by Quinic Acid Alters the Disease-Causing Activity of Rhizoctonia solani on Tomato,” Phytochemistry, Vol. 89, 2013, pp. 47-52. http://dx.doi.org/10.1016/j.phytochem.2012.09.018
- A. Rao, S. D. Ahmad, S. M. Sabir, S. I. Awan, A. H. Shah, S. R. Abbas, S. Shafique, F. Khan and A. Chaudhary, “Potential Antioxidant Activities Improve Salt Tolerance in Ten Varieties of Wheat (Triticum aestivum L.),” American Journal of Plant Sciences, Vol. 4, No. 6A, 2013, pp. 69-76. http://dx.doi.org/10.4236/ajps.2013.46A010
- C. H. Foyer and G. Noctor, “Ascorbate and Glutathione: The Hearth of the Redox Hub,” Plant Physiology, Vol. 155, No. 1, 2011, pp. 2-18.
- P. L. Conklin, E. H. Williams and R. L. Last, “Environmental Stress Sensitivity of an Ascorbic Acid-Deficient Arabidopsis mutant,” Proceedings of the National Academy of Sciences of the United States of America, Vol. 93, No. 18, 1996, pp. 9970-9974. http://dx.doi.org/10.1073/pnas.93.18.9970
- L. Colville and N. Smirnoff, “Antioxidant Status, Peroxidase Activity, and PR Protein Transcript Levels in Ascorbate-Deficient Arabidopsis thaliana vtc Mutants,” Journal of Experimental Botany, Vol. 59, No. 14, 2008, pp. 1-12. http://dx.doi.org/10.1093/jxb/ern229
- J. Zhou, A. Sun and D. Xing, “Modulation of Cellular Redox Status by Thiamine-Activated NADPH Oxidase Confers Arabidopsis Resistance to Sclerotinia scleroticorum,” Journal of Experimental Botany, Vol. 64, No. 11, 2013, pp. 3261-3272. http://dx.doi.org/10.1093/jxb/ert166
- C. J. Botanga, G. Bethke, Z. Chen, D. R. Gallie, O. Fiehn and J. Glazebrook, “Metabolite Profiling of Arabidopsis Inoculated with Alternaria brassicicola Reveals That Ascorbate Reduces Diseasse Severity,” Molecular PlantMicrobe Interactions, Vol. 25, No. 12, 2012, pp. 1628- 1638. http://dx.doi.org/10.1094/MPMI-07-12-0179-R
- R. Edwards and D. P. Dixon, “Plant Glutathione Transferases,” In: H. Sies and L. Packer, Eds., Gluthione Transferases and Gamma-Glutamyl Transpeptidases Methods in Enzymology, Elsevier B.V., Amsterdam, 2005, Vol. 401, pp. 169-186. http://dx.doi.org/10.1016/S0076-6879(05)01011-6
- S. Srivastava, S. Mishra, S. Dwivedi and R. D. Tripathi, “Role of Thiol Metabolism in Arsenic Detoxificationin Hydrilla verticillata (L.f.) Royle,” Water Air and Soil Pollution, Vol. 212, No. 1-4, 2010, pp. 155-165. http://dx.doi.org/10.1007/s11270-010-0329-9
- M. Basantani and A. Srivastava, “Plant Glutathione Transferases—A Decade Falls Short,” Canadian Journal of Botany, Vol. 85, No. 5, 2007, pp. 443-456. http://dx.doi.org/10.1139/B07-033
- R. Bhat, R. V. Rai and A. A. Karim, “Mycotoxins in Food and Feed: Present Status and Future Concerns,” Comprehensive Reviews in Food Science and Food Safety, Vol. 9, No. 1, 2010, pp. 57-81. http://dx.doi.org/10.1111/j.1541-4337.2009.00094.x
- S. Barnes, J. Prasain, T. D’Alessandro, A. Arabshahi, N. Botting, M. A. Lila, G. Jackson, E. M. Janle and C. M. Weaver, “The Metabolism and Analysis of Isoflavones and Other Dietary Polyphenols in Foods and Biological Systems,” Food and Function, Vol. 2, No. 5, 2011, pp. 235-244. http://dx.doi.org/10.1039/c1fo10025d
- J. Coleman, M. Blake-Kalff and E. Davies, “Detoxification of Xenobiotics by Plants: Chemical Modification and Vacuolar Compartmentation,” Trends Plant Science, Vol. 2, No. 4, 1997, pp. 144-151. http://dx.doi.org/10.1016/S1360-1385(97)01019-4
- A. Bittsanszky, R. V. Rai and G. Oros, “Response of Glutathion Conjugation System to Soil Borne Rhizoctonia Infection of Okra,” Acta Phytopathologicaet Entomolog ica Hungarica, Vol. 47, No. 2, 2012, pp. 191-202.
- A. Bittsánszky, S. A. Deepak and G. Oros, “Selective Response of Ricinus communis Seedlings to Soil Borne Rhizoctonia Infection,” Ecocycles, Vol. 1, No. 1, 2014, 6 p, in Press.
- V. P. Roxas, J. Wang, S. Lodhi and R. D. Allen, “Engineering Stress Tolerance in Transgenic Plants,” Acta Physiologica Plantarum, Vol. 19, No. 4, 1997, pp. 591- 594. http://dx.doi.org/10.1007/s11738-997-0058-x
- P. C. Abhilash, S. Jamil and N. Singh, “Transgenic Plants for Enhanced Biodegradation and Phytoremediation of Organic Xenobiotics RID C-2740-2009,” Biotechnology Advances, Vol. 27, No. 4, 2009, pp. 474-488. http://dx.doi.org/10.1016/j.biotechadv.2009.04.002
- H. Sandermann, “Higher Plant Metabolism of Xenobiotics: The ’Green Liver’ Concept,” Pharmacogenetics, Vol. 4, No. 5, 1994, pp. 225-241. http://dx.doi.org/10.1097/00008571-199410000-00001
- P. L. Conklin, S. A. Saracco, S. R. Norris and R. L. Last, “Identification of Ascorbic Acid-Deficient Arabidopsis thaliana Mutants,” Genetics, Vol. 154, No. 2, 2000, pp. 847-856.
- A. Bittsanszky, G. Gyulai and T. Kőmíves, “Arabidopsis thaliana Overexpressing Zea mays Glutathione S-Transferase: Modelling an Effective Phytoremediation,” Journal of Biotechnology, Vol. 150, 2010, pp. S486-S486. http://dx.doi.org/10.1016/j.jbiotec.2010.09.743
- S. J. Clough and A. F. Bent, “Florap Dip: A Simplified Method for Agrobacterium-Mediated Transformation of Arabidopsis thaliana,” The Plant Journal, Vol. 16, No. 6, 1998, pp. 765-743.
- C. Barth, W. Moeder, D. F. Klessig and P. L. Conklin, “The Timing of Senescence and Response to Pathogens Is Altered in the Ascorbate-Deficient Arabidopsis Mutant Vitamin c-1,” Plant Physiology, Vol. 134, No. 4, 2004, pp. 1784-1792. http://dx.doi.org/10.1104/pp.103.032185
- M. Khandaker, A. Khair and K. A. Bhuiyan, “Disease Reaction of Different Crops against Virulent Potato Isolates of Rhizoctonia solani Kuhn,” Bangladesh Journal of Botany, Vol. 37, No. 1, 2008, pp. 75-80. http://dx.doi.org/10.3329/bjb.v37i1.1567
NOTES
*Corresponding author.

