Advances in Microbiology
Vol. 2 No. 2 (2012) , Article ID: 19693 , 20 pages DOI:10.4236/aim.2012.22023
Over-Production of P60 Family Proteins, Glycolytic and Stress Response Proteins Characterizes the Autolytic Profile of Listeria monocytogenes
1Centro de Biomedicina Molecular e Estrutural (IBB), Faculdade de Ciências e Tecnologia, Universidade do Algarve, Faro, Portugal
2BioFig, Faculdade de Ciências e Tecnologia, Universidade do Algarve, Faro, Portugal
3Department of Infection, Immunity and Inflammation, University of Leicester, Leicester, UK
Email: *mfaleiro@ualg.pt
Received April 11, 2012; revised April 27, 2012; accepted May 10, 2012
Keywords: Listeria monocytogenes; autolysis proteome; stress proteins
ABSTRACT
Listeria monocytogenes is a foodborne pathogen capable of surviving under challenging conditions both outside and inside the host. During the transition from exponential to stationary phase it experiences a series of environmental changes that require an appropriate response to maintain cell viability. In this study the autolytic behaviour of a L. monocytogenes strain was investigated by two-dimensional electrophoresis. The study was done at the permissive autolysis temperature, 30˚C and at 20˚C, an autolysis non-permissive temperature. An autolytic strain proteome was also compared to a non-autolytic strain at the permissive autolysis temperature. The autolytic strain proteome at 30˚C in comparison to 20˚C evidenced increased synthesis of the P60 autolysin, glycolytic enzymes and proteins related with environmental stress responses. The over-production of P45 autolysin, was observed when the autolytic strain proteome was compared with the non-autolytic strain. The proteomes at the non-permissive temperature and the proteome of the non-autolytic strain were characterized by a diminished synthesis of several stress related proteins. The lack of autolysis seems to be associated to the over-production of proteins linked to fatty acid and amino acid synthesis, transcription regulation and cell morphogenesis as evidenced by the proteome at the non-permissive temperature and the non-autolytic strain. Autolysis proteome evidenced the over-production of P60 autolysins, glycolysis and stress proteins whereas the proteome obtained in conditions of absence of autolysis reveal a completely different group of proteins. Possible targets to activate listerial autolysis were identified.
1. Introduction
Autolysis in Listeria monocytogenes, a Gram-positive bacterium associated with the foodborne disease listeriosis, has been known for a long time [1,2]. The phenomenon occurs as a result of the activity of peptidoglycan hydrolases called autolysins [3]. These peptidoglycan hydrolases can participate in a significant number of important biological processes, namely in cell wall turnover, cell separation, cell division and their contribution in pathogenicity, either for Gram positive (e.g. Streptococcus pneumoniae, Staphylococcus aureus) and Gram negative bacteria (e.g. Helicobacter pylori) [4-6], also has been demonstrated. In L. monocytogenes, six autolysins have been identified P60, P45, Ami, Mur, Auto and the Lmo0327 [7-13] and at least three of these autolysins, P60, Ami and Auto are involved in survival and virulence of L. monocytogenes [8,14,15].
The peptidoglycan hydrolases constitute a distinctive family of enzymes whose principal function is to exactly cut the peptidoglycan in order that new murein strands can be inserted. Their activity must be firmly controlled to avoid the strand breakage resulting in cell lysis. The autolysis process is widespread amongst bacteria and other microorganisms, such as yeast and fungi (e.g. Saccharomyces cerevisae and Aspergillus nidulans) [16,17]. In Gram positive bacteria, teichoic acids are considered the principal regulators of the peptidoglycan hydrolases [18,19]. The binding of murein hydrolases to teichoic acids is mediated by repeat elements known as cholinebinding domains [20] and the connection of murein hydrolases to the cell wall occurs through the modification of teichoic acids. The addition of D-alanine ester linkages to teichoic acids modifies the surface charge and protects the cell from autolysis [21]. Besides the requirement for choline-binding domains and D-alanylation of teichoic acids, other studies have demonstrated that pyruvylation of unspecified carbohydrates (presumably teichoic acids) also contribute to the control of the murein hydrolases; namely a Bacillus anthracis mutant defective in csaAB genes, that are responsible for pyruvylation, displayed deficient cell division and autolysis [22]. Autolysis can be induced by the action of different factors, namely the dissipation of proton motive force (PMF) across the cell membrane, increase of cell wall pH, temperature mineral salts, ethanol and EDTA [23-25].
In our laboratory we found the process of autolysis in L. monocytogenes is more complex than may have been thought previously, with different strains of L. monocytogenes having very great differences in the pattern of their autolysis. For example, some L. monocytogenes isolates promptly and consistently undergo autolysis when grown in minimal medium, whereas other strains do not. Clearly there are differences in the control of autolysis in these strains but to date it is unknown how L. monocytogenes autolysins are regulated.
The objective of this study was to compare the intracellular proteome and the extracellular proteome of two L. monocytogenes strains that differ in the propensity to autolysis, one that had a propensity to autolyse and one which did not. Differences in the proteomes of these two strains provided important clues about this cellular process that can result in cell death and can be further explored to trigger listerial death in contaminated foods.
2. Material and Methods
2.1. Bacterial Strains and Growth Conditions
Two strains of L. monocytogenes were used in this study, C897 (a cheese isolate of 1/2a serotype prone to autolysis) and EGD (a clinical isolate of 1/2a serotype and a nonautolytic strain). For routine growth of listeria, tryptic soy agar medium (Merck, Darmstadt, Germany) was used at 30˚C. To study the protein synthesis under autolysis, the defined medium of Trivett and Meyer [26] (TM) was used. This medium has been used to investigate physiological responses in L. monocytogenes [1,27,28]. These cultures were incubated at two temperatures, at 30˚C at which the autolysis process is observed in C897 and at 20˚C at which autolysis is absent. The culture at both temperatures was done with shaking (120 rpm). All listeria cultures were inoculated to obtain an initial optical density at 600 nm (OD600nm) of 0.02 - 0.05, using 16 h overnight cultures in TM as inocula.
2.2. Induction of Lysis by Penicillin G
Penicillin induction of lysis was done according to Fontana et al. [29]. Overnight listeria cultures were diluted in fresh medium to obtain an initial optical density at 600 nm (OD600nm) of 0.02 - 0.05 and left to grow at 37˚C, with shaking, until they reached OD600nm of 0.15 - 0.2. Penicillin G (100 U/ml) was added to the cultures and the OD600nm was measured over 6 h and after 24 h. The viability was determined at the time of penicillin addition and at the end of 24 h using the drop method [30].
2.3. Protein Extracts and Protein Determination
Listeria cultures were grown until the end of exponential phase (OD600nm = 0.5 - 0.6) and the cells were collected by centrifugation (3000 ×g, 10 min at 4˚C). The protein extraction from the supernatant was done according to Trost et al. [31]. Briefly, the listerial cultures were centrifuged to eliminate the bacterial cells and the supernatant was filtered using 0.22 mm filters. Protein precipitation was initiated by the addition of PMSF (0.2 mM) and sodium-deoxycholate (0.2 mg/ml) to the samples and incubation on ice for 30 min. Afterwards, TCA (7%, w/v) was added and the samples were incubated overnight at 4˚C. The samples were centrifuged at 4020 × g for 15 min at 4˚C. The supernatant was removed and the protein pellet was washed twice with cold acetone. The pellet was air-dried and the protein was dissolved in solubilisation buffer (7 M urea, 2 M thiourea, 4% (w/v) CHAPS, 40 mM DTT, 0.8% (v/v) pharmalyte pH 4 - 7 (GE Healthcare). The samples were maintained at −80˚C until analyzed.
The protein extraction from the bacterial cells was done according to Folio et al. [32]. The collected cells were immediately washed after sampling with washing buffer (100 mM Tris-HCl pH 7.0, 100 mM EDTA and 0.1 ml 100× protease inhibitor mix (GE Healthcare, Madrid, Spain). Cell samples were resuspended in lysis buffer (25 mM Tris-HCl pH 7.0, 25 mM EDTA, 1% (v/v) DTT and 0.25 ml of 100× protease inhibitor mix) and were lysed by sonication on ice for 15 min. Contaminating nucleic acids were eliminated by the addition of 1 µl (1 unit) DNase RQ1 (Promega) and 5 µl RNase A (10 mg/ml, Promega). The samples were centrifuged at 3000 × g for 10 min at 4˚C and acetone (5× the volume of the supernatant) was added to each sample. The proteins were precipitated at −20˚C for 1 h and collected by centrifugation (18,000 × g for 30 min at 4˚C). Samples were air-dried and proteins were solubilised by the addition of 400 μl solubilisation buffer (7 M urea, 2 M thioureia, 4% (w/v) CHAPS, 40 mM DTT, 0.8% (v/v) Pharmalyte [GE Healthcare]). The total protein concentration was determined by using a Bio-Rad Protein Assay kit, according to the manufacturer’s instructions. The protein samples were kept at −80˚C until use.
2.4. Two-Dimensional Gel Electrophoresis (2-DE) and Image Analysis
The protein profile of the cell extracts and the supernatant of the L. monocytogenes cultures were analyzed by 2-DE. Approximately 300 - 400 mg of protein was used. The protein samples were separated in the first dimension using an 18 cm pH 3 - 10 or pH 4 - 7 Immobiline Dry Strip (GE Healthcare). Rehydration and isoelectric focusing was done using an IPGphor Isoelectric Focusing System (GE Healthcare). The Immobiline strips were re-hydrated for 11 h at 20˚C, 30 V and 50 µA/strip. Proteins were focused at 20˚C, 50 µA/strip, 100 V for 1 h, 500 V for 1 h and then a gradient at 8000 V with a final voltage from 8000 V until 60,000 Vhr. After isoelectric focusing, the strips were maintained at −80˚C or were immediately placed in an equilibration buffer (6 M urea, 75 mM Tris-HCl pH 8.8, 2% (w/v) SDS, 29.3% (v/v) glycerol, 0.002% (w/v) bromophenol blue and 10 mg/ml DTT (GE Healthcare) for 20 min. After this first step, the strips were transferred to fresh equilibration buffer supplemented with 25mg/ml iodoacetamide (GE Healthcare). Protein separation in the second dimension was performed in 12.5% (w/v) SDS-polyacrylamide gels in an Ettan Dalt six apparatus (GE Healthcare). The gels were run in triplicate to confirm the reproducibility of the protein patterns. Protein spots were visualised by Coomassie Blue R-250 staining. The determination of the protein profiles was done using an Image Scanner II (GE Healthcare), in combination with computational image analysis done by using Image Master 2D Platinum software, version 6 (GE Healthcare). The statistical analysis was performed using the Student’s t-test (confidence level 0.05 and 0.01). Mean normalized spot volume, standard deviation (SD) and coefficient of variance (CV) were determined for each spot.
2.5. Protein Identification
The spots of interest were manually excised from the stained 2-DE gels and analyzed by MALDI-TOF or LCMS/MS by the Protein and Nucleic Acid Chemistry Laboratory at the University of Leicester, UK or Aberdeen Proteome Facility (extracellular spots) and proteins were identified using MASCOT software.
3. Results and Discussion
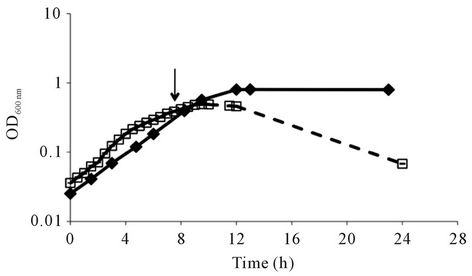
Listeria monocytogenes as many other bacterial pathogens meets a variety of challenging conditions either outside or inside the host, including insufficiency in nutrients (carbon, nitrogen, iron, and other nutrients), pH variations, oxidative stress and strong adverse conditions directed by the host innate and adaptive immune responses. The successful establishment of infection largely depends on the bacterial ability to modify and adjust their physiological state and virulence phenotype in agreement with the encountered challenging conditions. Bacterial cells exhibit growth-phase dependent physiological events and during the entry into the stationary phase an autolysis process may be trigger. The stationary growth phase is defined as the time that bacterial growth rate starts to decline and the development of multiple stress responses on promotion of survival can be activated. The comprehension of the cellular events at this time point of the growth phase is vital to fully understand the mechanisms of listerial response to environmental changes. The autolytic behaviour of L. monocytogenes C897 was observed when grown in the chemically defined TM medium at 30˚C, whereas autolysis was not seen with EGD in TM at 30˚C (Figure 1(a)). Consistent with this difference, C897 was more susceptible to lysis by penicillin (Figure 1(b)).
In contrast to the events at 30˚C, at lower growth temperature (20˚C) the autolysis process was not observed in C897 (Figure 1(c)). The generation time achieved by the autolytic strain and non-autolytic strain when grown at 30˚C was 2.33 ± 0.02 h and 2.08 ± 0.06 h, respectively and were not significantly different (P < 0.05).
To understand the events leading up to autolysis, two proteomic analysis were done: 1) the autolytic extracellular proteome (C897 grown at 30˚C) was compared to the non-autolytic extracellular proteome (C897 grown at 20˚C), and to the extracellular proteome of the nonautolytic strain EGD grown at 30˚C, 2) the autolytic intracellular proteome (C897 grown at 30˚C) was compared to the non-autolytic intracellular proteome (C897 grown at 20˚C) and to the non-autolytic intracellular proteome of EGD grown at 30˚C.
3.1. The Intracellular Proteome of the Autolytic Strain at Autolysis Condition
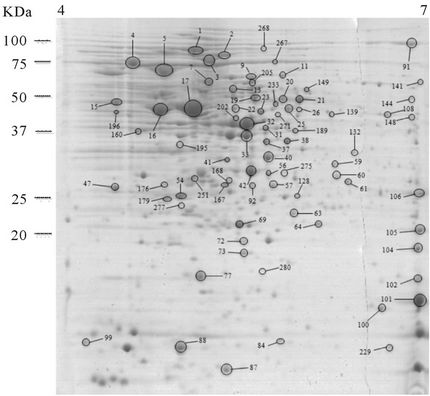
It was expected that the analysis of the intracellular proteome would reveal the cellular events that may be deregulated to allow the development of autolysis and also the cells’ efforts to control this potentially lethal cellular event. To accomplish this the intracellular proteome of the autolytic strain C897 at an autolysis condition (30˚C) was compared to the intracellular proteome at a nonautolysis condition (20˚C) and to the intracellular proteome of the non-autolytic strain EGD (30˚C). Data are summarized in Table 1. A representative gel of each experiment is shown in Figure 2(a). Twenty proteins were significantly more expressed (P < 0.05) by the autolytic strain, C897, at 30˚C in comparison to the intracellular proteome at the non-autolysis condition (20˚C). The identified proteins are distributed in ten functional categories, with some of these categories including at least three
 (a)
(a)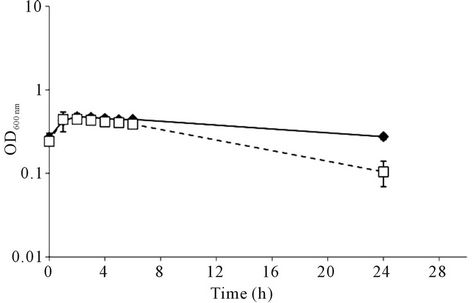 (b)
(b)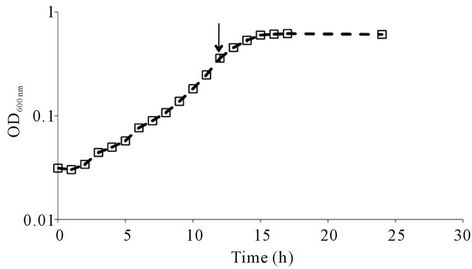 (c)
(c)
Figure 1. Autolysis behaviour of Listeria monocytogenes C897. (a) growth of L. monocytogenes C897 (¨) and EGD (t) in TM medium at 30˚C; (b) induction of lysis by penicillin G in L. monocytogenes C897 (¨) and EGD (t) cultures in BHI at 37˚C, (c) growth of L. monocytogenes C897 in TM medium at 20˚C. Data are the mean of three to four independent experiments. The standard error bars are within the area of the symbols in (a) and (c). The arrow indicates the time point at which samples were taken for 2-DE analysis.
proteins, which may indicate a higher importance of these cellular events in the autolysis process or its control. These categories are glycolysis, metabolism of amino acids and related molecules and the adaptation to stress conditions. A noteworthy aspect is the increased expression of six proteins of glycolysis, namely pyruvate kinase, phosphoglycerate kinase, enolase, glyceraldehyde 3-phosphate dehydrogenase (Gap protein), L-lactate dehydrogenase 1 (LDH-1) and Lmo 1634 (similar to alcohol dehydrogenase). The overproduction of six proteins involved in glucose metabolism by the autolytic strain indicates a strong need for energy.
Two proteins involved in the use of glucose were also over-produced by the C897 strain. These proteins, the phosphocarrier protein Hpr (spot 88) and the catabolite control protein CcpA (spot 31), are involved in the carbon catabolite repression (CCR) process (Table 1). The involvement of CcpA in stress responses of L. monocytogenes has been observed when listerial cells were exposed to salt stress [33]. Moreover the simultaneous overproduction of CcpA and the chaperones DnaK and GroEL (Table 1) was already reported for Lactobacillus plantarum [34] and L. monocytogenes [35] so is possible that CcpA may operate as a positive regulator of the two chaperones. Regarding the proteins involved in metabolism of amino acids and related molecules it is noteworthy that there was an increased synthesis of cysteine synthase (CysK) and D-alanine aminotransferase (DaaA). It is known that CysK is over-produced in B. subtilis under cold and oxidative stresses [36,37]. D-alanine is a key element in the synthesis of murein and one of the principal proteins involved in its synthesis is the Dalanine aminotransferase, which catalyzes the conversion of D-glutamic acid to α-ketoglutaric acid and D-alanine. The deletion of the genes that encode D-alanine aminotransferase (dat or daaA) and alanine racemase (dal) produces a phenotype that is entirely dependent on exogenous alanine and its absence induces lysis of L. monocytogenes cells in log phase of growth [38]. The over-expression of these two proteins under autolysis condition is indicative of the fatal murein break due to autolysis in C897 cells.
3.2. In Autolysis Condition the Autolytic Strain Shows a Stress Adaptation Response
The intracellular proteome of the C897 strain at 30˚C, in comparison to the intracellular proteome at the nonautolysis condition (20˚C), revealed the over-production of five proteins associated with adaptation to stress conditions (Table 1) and one of these were also over-produced by C897 at 30˚C, in comparison to EGD at 30˚C. Three proteins, in particular, that fit into the functional category of adaptation to atypical conditions were identified; namely the general stress protein Ctc, peroxide resistance protein Dpr and the Lmo1601 protein (stress protein-like protein). Two other protein spots related with stress response, the DnaK protein and the 60 KDa chaperonine, GroEL also were identified (Table 1).
The over-expression of Ctc in L. monocytogenes exposed to stress conditions, namely cold, salt and osmotic
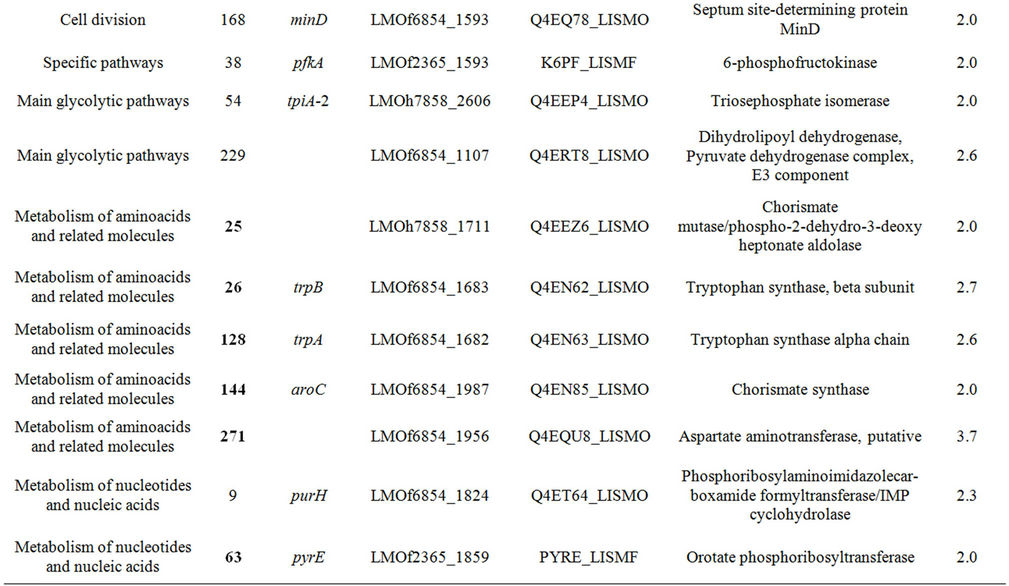
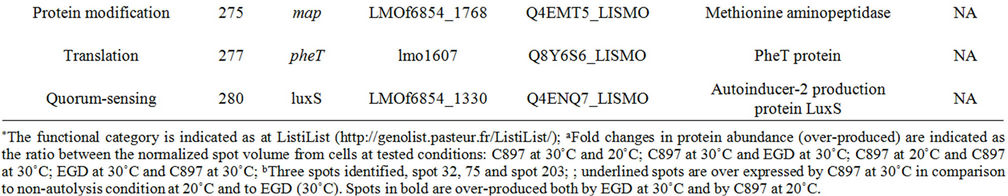
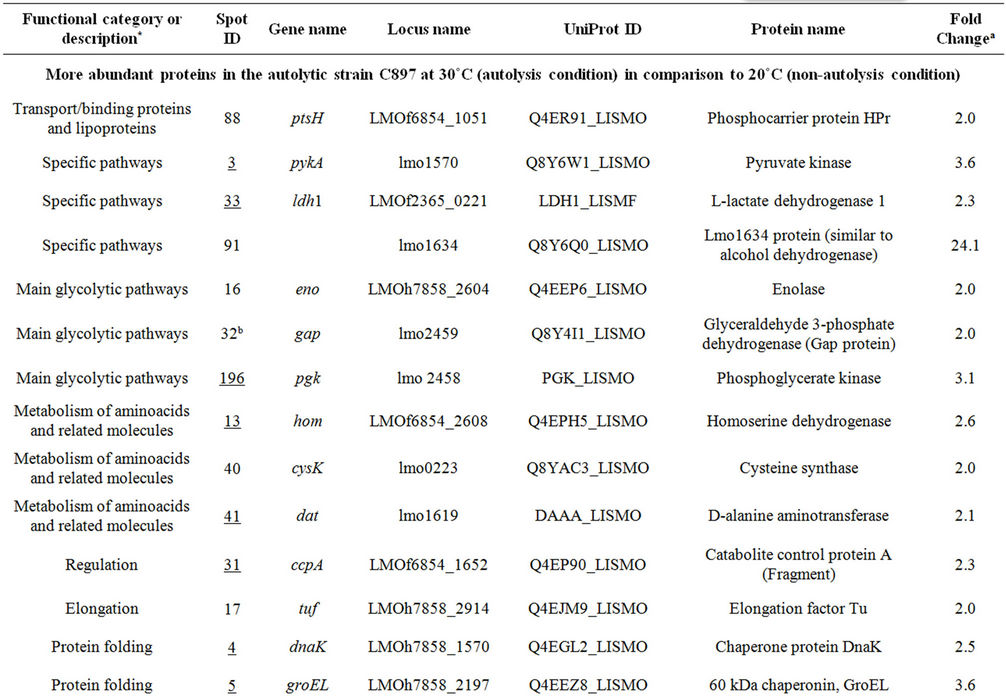
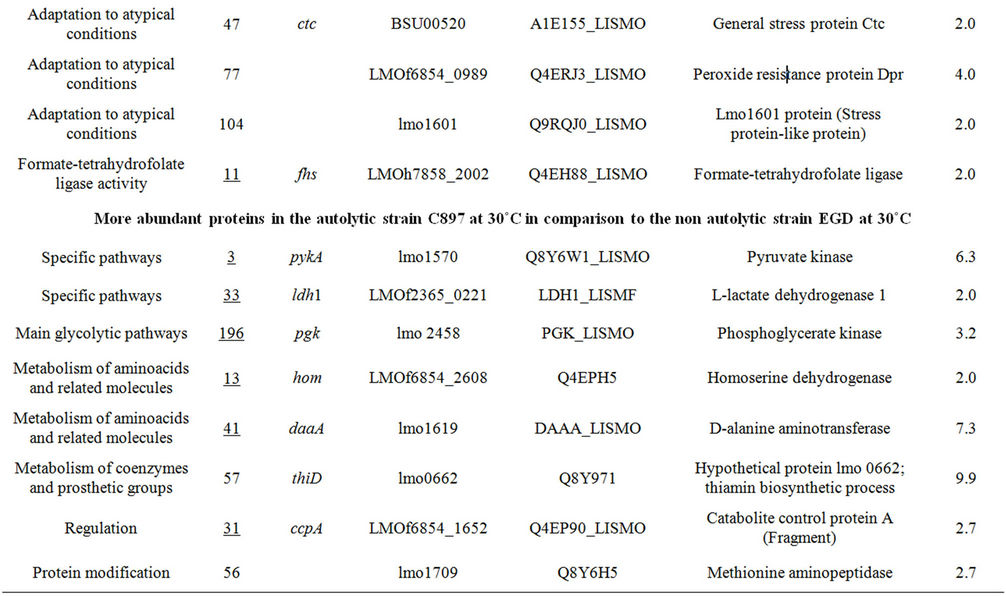

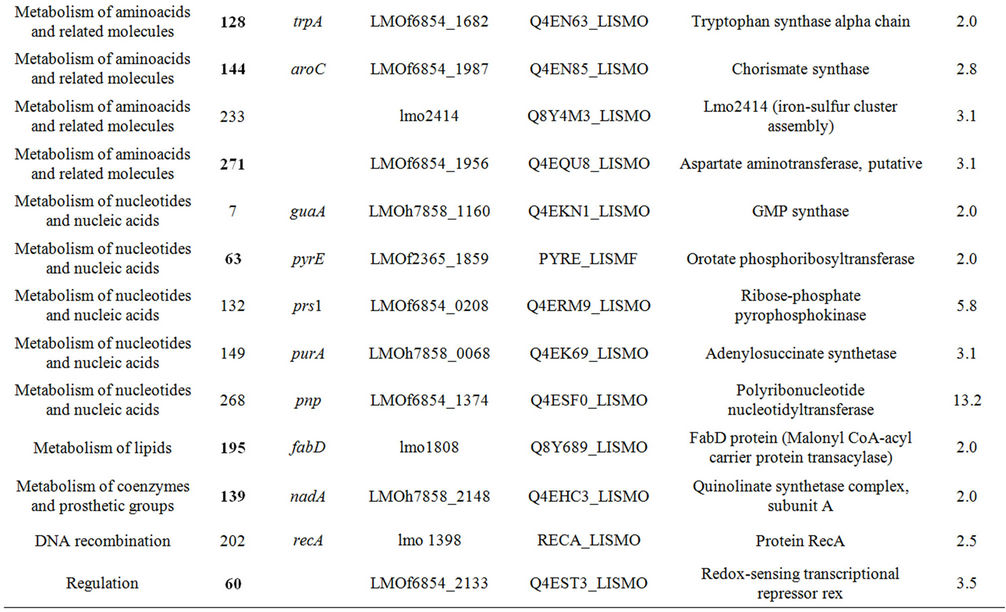
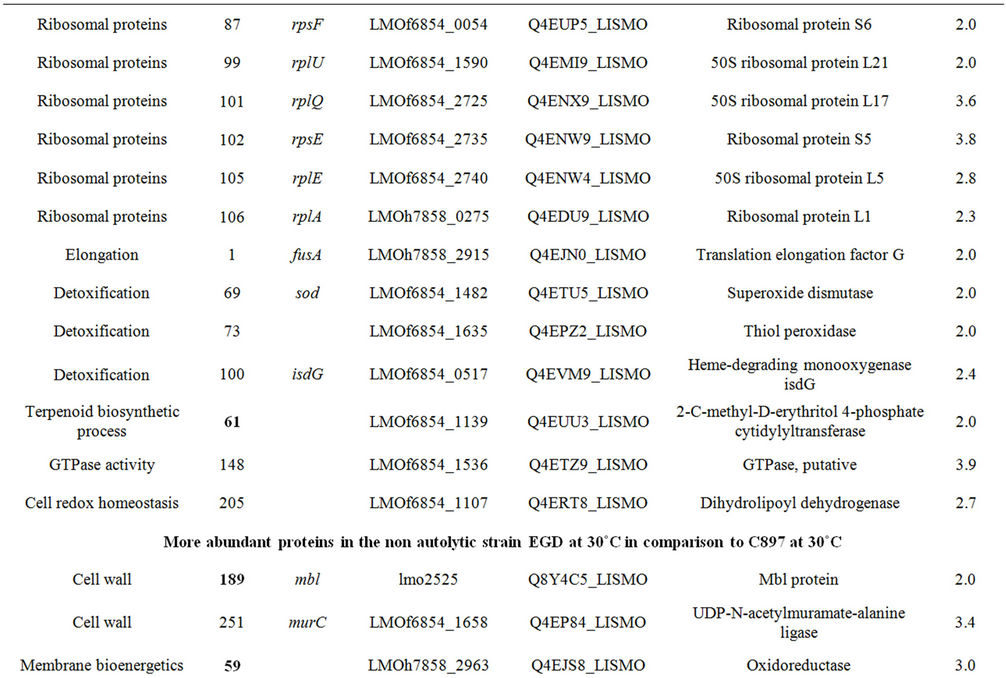

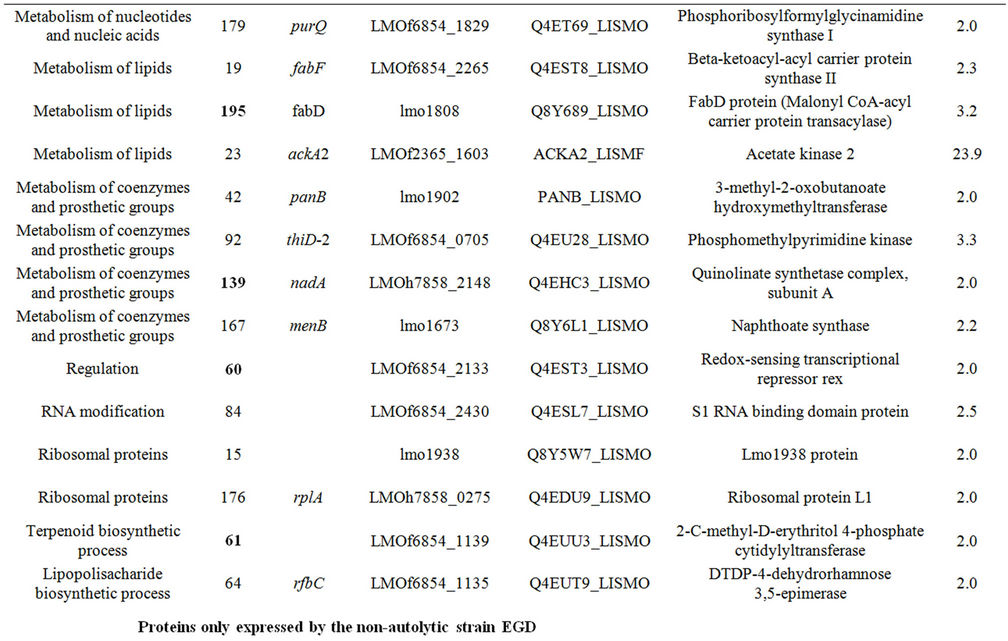

Table 1. Differentially expressed intracellular proteins. Differential protein abundance is indicated as fold change (minimum 2-fold, P < 0.05).
shock, already has been reported [33,35,39]. Likewise, in another Gram positive bacterium, B. subtilis, the overexpression of Ctc was observed when cells were under osmotic, heat and oxidative stresses and during glucose limitation [40]. Dpr that responds to peroxide stress has been associated with response to lactate exposition in Lactobacillus plantarum [41] and Lactococcus lactis [42] and was reported to protect Streptococcus pyogenes against several stresses [43]. It seems that the Lmo1601 protein (stress protein-like protein) may be involved in the maintenance of the redox balance of the cell because this protein has 59% similarity to the general stress protein YtxH from B. subtilis. Another over-produced protein worthy of note due to its involvement in stress response is the elongation factor Tu (EF-Tu). In Escherichia coli the EF-Tu seems to have a role additional to its translation elongation function, in that it may act as a chaperone-like protein protecting cells from stress [44]. It also has been reported to protect L. monocytogenes cells from salt and cold stress [33,35]. None of these proteins have previously been related to autolysis, except DnaK.
The heat shock proteins DnaK and GroEL were overproduced by C897 at 30˚C in comparison to 20˚C. The expression of these two proteins is elevated when bacterial cells are exposed to several environmental stress conditions and they play a crucial role in L. monocytogenes in vivo survival [45,46]. Besides their recognized value for cell protection under stress, an important role in protein folding has been attributed to GroEL even when bacterial cells grow at optimal temperature [47]. The association of induction of heat shock proteins with autolysis has been reported in S. aureus and E. coli [48,49]. Recently a dnaK mutant of S. aureus was described that showed a reduced autolysis rate in comparison to the wild type strain suggesting a possible role of DnaK on autolysins activity [50]. Considering the results obtained by Singh et al. [50] it is tempting to hypothesise that DnaK alone, or in combination with GroEL activate the activity of P60 and/or P45.
The observation of the over-production of a significant number of stress response proteins by the C897 strain strongly suggests that it is stress that triggers autolysis.
3.3. The Intracellular Proteome of the Autolytic Strain at Non-Autolysis Condition
Thirty-night proteins over-expressed at the non-autolysis condition (20˚C) were identified. Data are summarized in the Table 1, Supplementary Table A and a representative gel is shown in Figures 2(b). These proteins can be distributed among eighteen functional categories. The functional categories that have the highest number of spots are the metabolism of amino acids (8 proteins), metabolism of nucleotides and nucleic acids (5 proteins) and ribosomal proteins (6 proteins).
Two oxidative stress proteins were significantly expressed: superoxide dismutase and thiol peroxidase. A component of the SOS response, the activator RecA also was more expressed. The over-production of proteins involved in cell detoxification in non-autolysis and autolysis conditions indicates the higher sensitivity of the autolytic strain to oxidative stress.
At the non-autolysis condition, the intracellular overexpression of the autolysin P60 was observed. This autolysin was at lower levels in the extracellular proteome at 20˚C (see section 3.7) and thus we can hypothesize that intracellular/extracellular proteome ratio is an important factor in autolysis inhibition.
A significant set of over-produced proteins was also over-produced by the non-autolytic EGD and will be discussed in section 3.5. Intracellular protein expressed in both strains in the absence of autolysis.
3.4. The Intracellular Proteome of the Non-Autolytic Strain
Comparison of the intracellular proteomes of the nonautolytic EGD grown in conditions that induce autolysis of C897 can provide information on key elements of the autolysis inhibition process. Twenty-nine proteins that were over-produced in EGD were identified. Data are summarized in Table 1 and a representative gel is shown in Figure 2(c). The functional categories with the highest number of spots were the metabolism of amino acids (5 proteins), followed by the metabolism of coenzymes and prosthetic groups and the metabolism of nucleotides and nucleic acids (4 proteins) metabolism of lipids and main glycolytic pathways (3 proteins). The five proteins for the metabolism of amino acids and related molecules were common to proteins over-produced by the autolytic
 (a)
(a)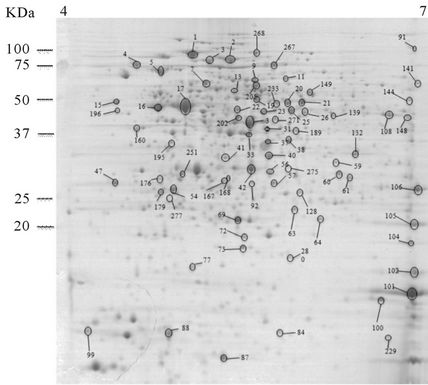 (b)
(b)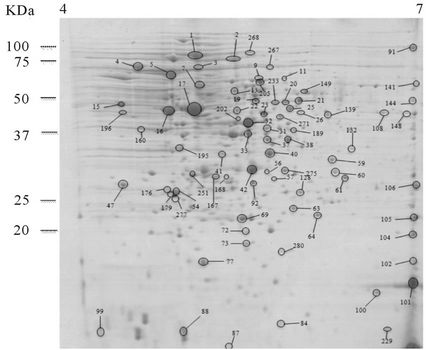 (c)
(c)
Figure 2. 2-DE maps of the intracellular proteome of the autolytic strain C897 grown at 30˚C (autolysis permissive temperature) (a) at 20˚C (non-autolysis permissive temperature); (b) and the non-autolytic strain EGD grown at 30˚C; (c) The identified spots are indicated.
strain at 20˚C in comparison to 30˚C (see following section). Two proteins related with the cell wall and one protein related with cell division, Mbl, UDP-N-acetylmuramate alanine ligase and the septum determining protein MinD, respectively were identified. UDP-N-acetylmuramate-alanine ligase (MurC) is one of the fundamental cytoplasmic peptidoglycan biosynthetic enzymes. This enzyme catalyzes the ATP-dependent ligation of L-alanine (Ala) and UDP-N-acetylmuramic acid (UNAM) to form UDP-N-acetylmuramyl-L-alanine (UNAM-Ala). The gene minD belongs to the minicell genetic locus constituted by other two genes minC and minE [51]. The investigation of the expression of different combinations of the min genes revealed that MinC is a division inhibitor and MinD induces MinC activity [51,52]. The possible role of Mbl will be discussed below, as is a common over produced spot by EGD and the autolytic strain at 20˚C.
The membrane lipid homeostasis and the capacity of bacterial cells to modify the lipid composition according to different environments may dictate bacterial survival. Consistent with the hypothesis that lipid composition influences autolysis, two proteins that are key elements of the fatty acid synthesis (b-ketoacyl-acyl carrier protein synthase II (FabF) and the FabD protein, Malonyl CoAacyl carrier protein transacylase) were over-expressed in the non-autolytic strain and in the autolytic strain at 20˚C.
In contrast to over-production of several glycolytic enzymes in C897 at 30˚C, only two glycolytic proteins were higher in EGD at 30˚C (6-phosphofructokinase and triosephosphate isomerase). This is consistent with the previous suggestion that the requirement of glycolysis is an important part of the autolysis phenomenon.
The LuxS (S-ribosylhomocysteinase) protein was only found in the intracellular proteome of EGD (Table 1, Supplementary Table A, Figure 2). Autolysis is linked to quorum sensing in S. pneumoniae [53,54] which is regulated by the ComDE, which is generally considered to be a component of a quorum sensing mechanism [53]. The participation of quorum-sensing components in L. monocytogenes autolysis regulation has not been reported.
3.5. Intracellular Protein Expressed in Both Strain in the Absence of Autolysis
Analysis of data collected in the absence of autolysis (C897 at 20˚C and EGD at 30˚C) revealed a set of common proteins (Table 1, Supplementary Table A). From the 12 identified proteins five are involved in the metabolism of amino acids and two in the metabolism of coenzymes and prosthetic groups. The significance of increases in the abundance of proteins associated with the metabolism of the amino acids tyrosine, phenylalanine and tryptophan (chorismate mutase, chorismate synthase, tryptophan synthase, beta and alpha subunits) relies in the role of these aromatic amino acids in the maintenance of the structure and function of membrane proteins [55, 56]. An important clue to the inhibition of autolysis is the component of the cell wall, the Mbl protein (similar to MreB-like protein). Mbl is an isoform of MreB that, together with other proteins, provide a rod shape to bacterial cells. B. subtilis has three MreB isoforms (MreB, Mbl and MreBH) which seems to be responsible for the positioning of peptidoglycan synthases, a peptidoglycan hydrolase (LytE) and other membrane-associated cell morphogenesis proteins, such as MreC and MreD [57]. From these three MreB isoforms, Mre-BH seems to be responsible for the control of the autolytic activity by directing the localization of the cell wall hydrolase LytE [58]. Search of the Listeria genome indicates that only two MreB isoforms are present, the MreB and Mbl. So far the roles of MreB and its Mbl isoform in L. monocytogenes have not been clarified. From our data it is possible to propose that Mbl plays an important role in autolysis inhibition. The absence of the over-production of the glycolysis enzymes L-lactate dehydrogenase 1 (LDH-1), Lmo 1634 (similar to alcohol dehydrogenase) and glyceraldehyde 3-phosphate dehydrogenase (Gap protein) in the non-autolysis condition may be explained by the over-production of the redox-sensing transcriptional repressor rex (spot 60) which senses the NADH/NAD+ ratio in the cell and which has been shown in Streptomyces coelicolor and Bacillus subtilis to indirectly modulate the metabolism by regulation of genes encoding proteins of the respiratory chain [59,60], whereas in Staph. aureus it regulates the transcription process during the switch from aerobic to anaerobic growth [61]. In Staph. aureus the proteins Adh1, Ldh1 and GapA1 are Rex-regulated, but only Adh1 and Ldh1 have a Rex binding site [61]. The genes regulated by Rex and its role in the aerobic and anaerobic growth of L. monocytogenes remains to be clarified.
In both strains at absence of autolysis the 2-C-methylD-erythritol 4-phosphate cytidyltransferase which, is involved in the terpenoid (or isoprenoid) biosynthetic process, was over-produced (Table 1, Supplementary Table A). Isoprenoids include a large number of compounds (more than 30,000) participating in a significant number of physiological processes either in eukaryotes and prokaryotes. In bacteria they have important roles in electron transport chains, with ubiquinone and menaquinone standing out in these roles. Bactoprenols function as carbohydrate carriers in the biosynthesis of peptidoglycan [62, 63].
3.6. The Extracellular Proteome of the Autolytic Strain at 30˚C Evidence Over-Production of P60 Family Proteins
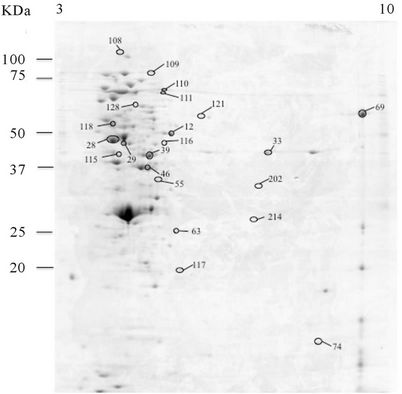
Three spots were identified as being significantly (P < 0.05) more abundant in the culture supernatant of the autolytic strain C897 when grown at 30˚C, compared to growth at 20˚C. One of them is the autolytic enzyme P60 (invasion associated protein). Data are summarized in the Table 2 and Supplementary Table A and a representative gel is shown in Figure 3(a). Five proteins were identified in the extracellular proteome of the autolytic strain C897 grown at 30˚C compared to the extracellular proteome of the non-autolytic strain EGD grown at 30˚C. Among them is the P45 autolysin, an amino acid ABC transporter and Lmo1333 which is similar to the B. subtilis YqzC protein (Table 2, Supplementary Table A). P60 was equally abundant in the extracellular proteome of C897 and EGD at 30˚C.
The differences in the autolytic behaviour of C897 at 30˚C in comparison to the lack of autolysis at 20˚C could be linked to the higher abundance of P60 in the extracellular proteome of C897 at 30˚C whereas the higher abundance of P45 in the extracellular proteome of C897 in comparison to EGD could be linked to the different autolytic behaviour of these two strains.
P60 is classified as an endo-N-acetylmuramidase [15] and is implicated in cell separation because P60 depletion results on the formation of long chains of cells [14]. The P45 protein displays 55% similarity and 38% identity to P60 and also exhibits peptidoglycan hydrolase activity [13]. To date, and in contrast to reported phenotype alterations due to P60 depletion, the exact function of P45 has not been described. As these two autolysins were detected in higher quantities in the extracellular proteome of the autolytic strain at 30˚C, in comparison to the growth conditions in absence of autolysis (P60 linked to temperature and P45 linked to strain) become evident their impact on the autolysis process.
3.7. The Extracellular Proteome at Non-Autolysis Conditions (Temperature and Strain)
The analysis of the extracellular proteome at non-autolysis condition was done by collecting the supernatant of EGD at 30˚C and C897 at 20˚C at the end of exponential phase (an approximate OD600nm of 0.5 - 0.6, Figures 1(a) and (b)). Seven extracellular proteins of the autolytic strain were identified as being expressed only at the nonautolytic condition (20˚C), (Table 2, Supplementary Table A and Figure 3(b)) and only two proteins belong to the same functional category (metabolism of nucleotides and nucleic acids). Among the eight proteins, a cell surface
 (a)
(a)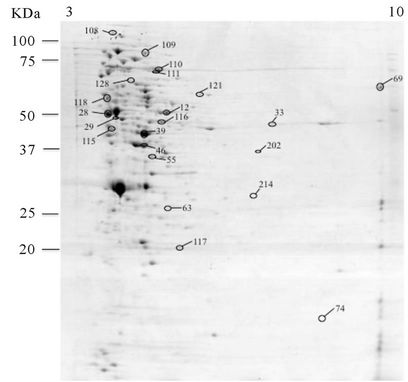 (b)
(b)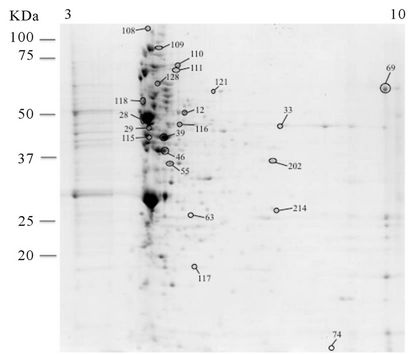 (c)
(c)
Figure 3. 2-DE maps of the extracellular proteome of the autolytic strain C897 grown at 30˚C (autolysis permissive temperature) (a) at 20˚C (non-autolysis permissive temperature); (b) and the non-autolytic strain EGD grown at 30˚C; (c) The identified spots are indicated.
protein (Lmo2185, SvpA, NEAr transporter) and a protein related with bioenergetics, an ATP synthase, were identified. The production of SvpA protein increases significantly when cells experience iron deprivation and studies on cellular fractions showed that in iron-rich media SvpA is entirely secreted into the culture supernatant [64]. The comparison of the extracellular proteome of C897 and EGD at 30˚C showed five over-expressed proteins of the extracellular proteome of the non-autolytic strain, EGD (Table 2, Supplementary Table A and Figure 3(c)). One of these spots belongs to the transport/ binding lipoproteins and is an EII Mant PTS permease
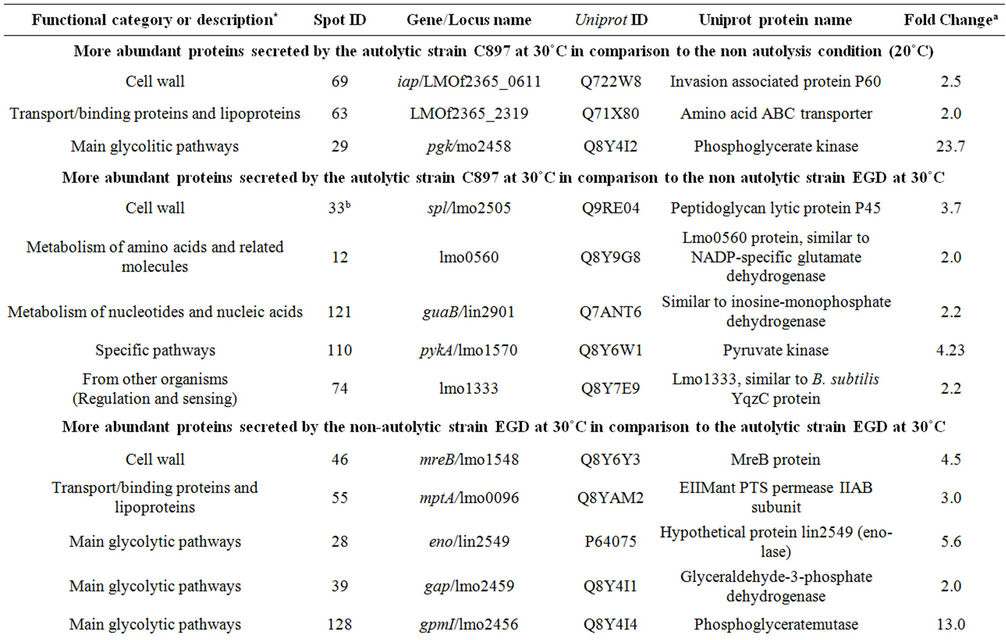
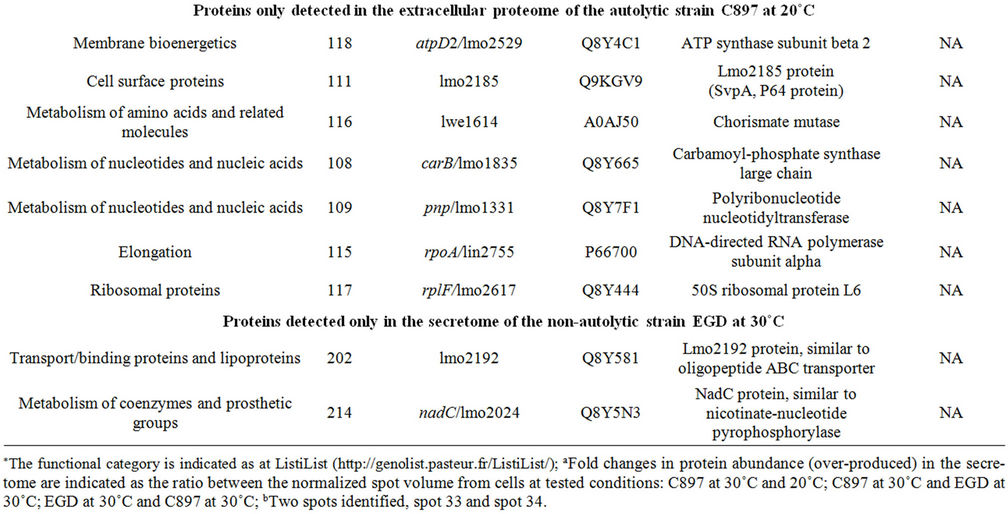
Table 2. Proteins identified in the extracellular proteome of C897 and EGD strain.
IIAB subunit. An increase expression of this mannosespecific PTS enzyme IIAB also was detected in the intracellular proteome of L. monocytogenes LO28 when the bacterial cells were exposed to salt stress and this increase may be related to higher energy requirements [33].
4. Conclusions
Our results show that at the end of exponential phase the autolytic strain increases the production of a series of proteins with a recognized role in combating the action of environmental stressors. This does not occur with the non-autolytic EGD and suggest that C897 is more sensitive to stresses that eventually can trigger the autolysis process. At the autolysis permissive temperature (30˚C) the secretion of two autolysins (P60 and P45) was greater than at the non-autolysis permissive temperature (20˚C) and by the non-autolytic strain EGD. On other hand increased amounts of the P60 autolysin was observed in the intracellular proteome at non-permissive temperature. These findings suggest that at lower temperature the export of autolysins is inhibited and this may be due to alterations in the cell wall as evidenced by the over-production of proteins involved in cell wall synthesis. Differences in over-produced proteins of the glycolytic pathway, fatty acid and amino acids synthesis, transcription regulation, cell wall synthesis and morphogenesis are elements that may assure the less susceptible profile of EGD to autolysis. A significant number of proteins related to fatty acid and amino acid synthesis, transcription regulation and cell morphogenesis were commonly over-produced by the non-autolytic strain EGD and by the autolytic strain at autolysis non-permissive temperature, indicating a set of important cellular events important to the lack of autolysis.
From the collected data the cell shape Mbl protein and the transcriptional repressor Rex can constitute possible cell targets to trigger autolysis.
5. Acknowledgements
This work was supported by Fundação para a Ciência e a Tecnologia (PTDC/AGRI-ALI/2006 and IBB/CBME, LA, FEDER/POCI). ML Faleiro is grateful for the grant SFRH/BSAB/859/2008.
REFERENCES
- E. Jones, G. Shama, D. Jones, I. S. Roberts and P. A. Andrew, “Physiological and Biochemical Studies on Phychrotolerance in Listeria monocytogenes,” Journal of Applied Microbiology, Vol. 83. No. 1, 1997, pp. 31-35. doi:10.1046/j.1365-2672.1997.d01-391.x
- L. Pine, G. B. Malcom, J. B. Brooks and M. I Danesshvar, “Physiological Studies on the Growth and Utilization of Sugars by Listeria Species,” Canadian Journal of Microbiology, Vol. 35, No. 2, 1989, pp. 245-254. doi:10.1139/m89-037
- J. V. Holtje, “From Growth to Autolysis: The Murein Hydrolases in Escherichia coli,” Archives of Microbiology, Vol. 164, No. 4, 1995, pp. 243-254. doi:10.1007/BF02529958
- A. M. Berry, R. A. Lock, D. Hansman and J. C. Paton, “Contribution of Autolysin to Virulence of Streptococcus pneumoniae,” Infection and Immunity, Vol. 57, No. 8, 1989, pp. 2324-2330.
- N. Mani, L. M. Baddour, D. Q. Offutt, U. Vijaranakul, M. J. Nadakavukaren and R. K. Jayaswal, “Autolysis-Defective Mutant of Staphylococcus aureus: Pathological Considerations, Genetic Mapping and Electron Microscopic Studies,” Infection and Immunity, Vol. 62, No. 4, 1994, pp. 1406-1409.
- E. A. Marcus and D. R. Scott, “Cell Lysis Is Responsible for the Appearance of Extracellular Urease in Helicobacter pylori,” Helicobacter, Vol. 6, No. 2, 2001, pp. 93-99. doi:10.1046/j.1523-5378.2001.00014.x
- A. Bubert, M. Kuhn, W. Goebel and S. Kohler, “Structural and Functional Properties of the p60 Proteins from Different Listeria Species,” Journal of Bacteriology, Vol. 174, No. 24, 1992, pp. 8166-8171.
- D. Cabanes, O. Dussurget, P. Dehoux and P. Cossart, “Auto, a Surface Associated Autolysin of Listeria monocytogenes Required for Entry into Eukaryotic cells and Virulence,” Molecular Microbiology, Vol. 51, No. 6, 2004, pp. 1601-1614. doi:10.1111/j.1365-2958.2003.03945
- S. A Carroll, T. Hain, U. Technow, A. Darji, P. Phashalidis, S. W. Joseph and T. Chakraborty, “Identification and Characterization of a Murein Hydrolase, MurA, of Listeria monocytogenes, a Muramidase Needed for Cell Separation,” Journal of Bacteriology, Vol. 185, No. 23, 2003, pp. 6801-6808. doi:10.1128/JB.185.23.6801-6808.2003
- A. M McLaughlan and S. J. Foster, “Molecular Characterization of an Autolytic Amidase of Listeria monocytogenes EGD,” Microbiology, Vol. 144, No. Pt 5, 1998, pp. 1359-1367.
- E. Milohanic, R. Jonquieres and P. B. Coossart, “The Autolysin Ami Contributes to the Adhesion of Listeria monocytogenes to Eukaryotic Cells via Its Cell wall Anchor,” Molecular Microbiology, Vol. 39, No. 5, 2001, pp. 1212- 1224. doi:10.1111/j.1365-2958.2001.02208.x
- M. Popowska and Z. Markiewicz,“Characterization of Listeria monocytogenes Protein Lmo0327 with Murein Hydrolase Activity,” Archives of Microbiology, Vol. 186, No. 1, 2006, pp. 69-86. doi:10.1007/s2003-006-0122-8
- K. Schubert, A. M. Bichlmaier, E. Mager, K. Wolff, G. Ruhland and F. Fiedler, “P45, an Extracellular 45 kDa Protein of Listeria monocytogenes with Similarity to Protein p60 and Exhibiting Peptidoglycan Lytic Activity,” Archives of Microbiology, Vol. 173, No. 1, 2000, pp. 21-28. doi:10.1007/s20030050003
- S. Pilgrim, A. Kolb-Mäurer, I. Gentshev, W. Goebel and M. Kuhn, “Deletion of the Gene Encoding p60 in Listeria monocytogenes Leads to Abnormal Cell Division and Loss of Actin-Based Motility,” Infection and Immunity, Vol. 71, No. 6, 2003, pp. 3473-3484. doi:10.1128/IAI.71.6.3473-3484.2003
- M. D. Wuenscher, S. Köhler, A. Bubert, U. Gerike and W. Goebel, “The iap Gene of Listeria monocytogenes Is Essential for Cell Viability, and Its Gene Product, p60, Has Bacteriolytic Activity,” Journal of Bacteriology, Vol. 175, No. 11, 1993, pp. 3491-350.
- R. Gonzalez, A. J. Martinez-Rodriguez and A. V. Carrascosa, “Yeast Autolytic Mutants Potentially Useful for Sparking Wine Production,” International Journal of Food Microbiology, Vol. 84, No. 1, 2003, pp. 21-26. doi:1016/S0168-1605(02)00389-6
- T. Emri, Z. Molnár, M. Szilágyi and I. Pócsi, “Regulation of Autolysis in Aspergillus nidulans,” Applied Biochemistry and Biotechnology, Vol. 151, No. 2-3, 2008, pp. 211- 220. doi:10.1007/s12010-008-8174-7
- F. C. Neuhaus and J. Baddiley, “A Continuum of Anionic Charge: Structures and Functions of D-Alanyl-teichoic Acids in Gram-Positive Bacteria,” Microbiology and Molecular Biology, Vol. 67, No. 4, 2003, pp. 686-723. doi:10.1128.MMBR.67.4.686-723.2003
- I. Fedtke, D. Mader, T. Kohler, H. Moll, G. Nicholson, R. Biswas, K. Henseler, F. Götz, U. Zähringer and A. Peschel, “A Staphylococcus aureus ypfP Mutant with Strongly Reduced Lipoteichoic Acid (LTA) Content: LTA Governs Bacterial Surface Properties and Autolysin Activity,” Molecular Microbiology, Vol. 65, No. 4, 2007, pp. 1078- 1091. doi:10.1111/j.1365-2958.2007.05854.x
- J. R.Scott and T. C. Barnett, “Surface Proteins of GramPositive Bacteria and How They Get There,” Annual Reviews of Microbiology, Vol. 60, 2006, pp. 397-423. doi:10.1146/annurev.micro.60.080805.142256
- J. Wecke, K Madela and W. Fischer, “The Absence of DAlanine from Lipoteichoic Acid and Wall Teichoic Acid Alters Surface Charge, Enhances Autolysis and Increases Susceptibility to Methicillin in Bacillus subtilis,” Microbiology, Vol. 143, No. 9, 1997, pp. 2953-2960. doi:10.1099/00221287-143-9-2953
- S. Mesnage, T. Fontaine, M. Mignot, M. Delepierre, M. Mock and A. Fouet, “Bacterial SLH Domain Proteins Are Non-Covalently Anchored to the Cell Surface via a Conserved Mechanism Involving Wall Polysaccharide Pyruvylation,” The EMBO Journal, Vol. 19, No. 17, 2000, pp. 4473-4484. doi:10.1093/emboj/19.17.4473
- L. O. Ingram, “Mechanism of lysis of Escherichia coli by Ethanol and other Chaotropic Agents,” Journal of Bacteriology, Vol. 146, No. 1, 1981, pp. 331-336.
- S. R. Watt and A. J. Clarke, “Role of Autolysins in the EDTA-Induced Lysis of Pseudomonas aeruginosa,” FEMS Microbiology Letters, Vol. 124, No. 1, 1994, pp. 113-120. doi:10.1111/j.1574-6968.1994.tb07270.x
- T. Ochiai, “Salt-Sensitive Growth of Staphylococcus aureus: Stimulation of Salt-Induced Autolysis by Multiple Environmental Factors,” Microbiology and Immunology, Vol. 43, No. 7, 1999, pp. 705-709.
- T. L. Trivett and E. A Meyer, “Citrate Cycle and Related Metabolism of Listeria monocytogenes,” Journal of Bacteriology, Vol. 107, No. 3, 1971, pp. 770-779.
- A. Adrião, M. Vieira, I. Fernandes, M. Barbosa, M. Sol, R. P. Tenreiro, L. Chambel, B. Barata, I. Zilhão, G. Shama, S. Perni, S. J. Jordan, P. W. Andrew and M. L. Faleiro, “Marked Intra-Strain Variation in Response of Listeria monocytogenes Dairy Isolates to Acid or Salt Stress and the Effect of Acid or Salt Adaptation on Adherence to Abiotic Surfaces,” International Journal of Food Microbiology, Vol. 123, 2008, pp. 142-150. doi:10.1016/j.ijfoodmicro.2007.12.016
- M. L.Faleiro, P. W. Andrew and D. Power, “Stress Response of Listeria monocytogenes Isolated from Cheese and other Foods,” International Journal of Food Microbiology, Vol. 84, No. 2, 2003, pp. 207-216. doi:10.1016/S0168-1605(02)00422-1
- R. Fontana, M. Boaretti, A. Grossato, E. A. Tonin, M. M. Lléo and G. Satta, “Paradoxical Response of Enterococcus faecalis to the Bactericidal Activity of Penicillin is Associated with Reduced Activity of one Autolysin,” Antimicrobial Agents and Chemotherapy, Vol. 34, No. 2, 1990, pp. 314-320.
- C.-Y. Chen, G. W. Nace and P. L. Irwin, “A 6 × 6 Drop Plate Method for Simultaneous Colony Counting and MPN Enumeration of Campylobacter jejuni, Listeria monocytogenes and Escherichia coli,” Journal of Microbiological Methods, Vol. 55, No. 2, 2003, pp. 475-479. doi:10.1016/S0167-7012(03)00194-5
- M. Trost, D. Wehmhöner, U. Kärst, G. Dieterich, J. Wehland and L. Jänsch, “Comparative Proteome Analysis of Secretory Proteins from Pathogenic and Non-Pathogenic Listeria Species,” Proteomics, Vol. 5, No. 6, 2005, pp. 1544-1557. doi:10.1002/pmic.200401024
- P. Folio, P. Chavant, I. Chafsey, A. Belkorchia, C. Chambon and M. Hébraud, “Two-Dimensional Electrophoresis Database of Listeria monocytogenes EGDe Proteome and Proteomic Analysis of Mild-Log and Stationary Growth Phase Cells,” Proteomics, Vol. 4, 2004, pp. 3187-3201. doi:10.1002/pmic.200300841
- O. Duché, F. Tremoulet, P. Glaser and J. Labadie, “Salt Stress Proteins Induced in Listeria monocytogenes,” Applied and Environmental Microbiology, Vol. 68, No. 4, 2002, pp. 1491-1498. doi:10.1128/AEM.68.4.1491-1498.2002
- C. Castaldo, R. A. Siciliano, L. Muscariello, R. Marasco and M. Sacco, “CcpA Affects Expression of the groESL and dnaK operons in Lactobacillus plantarum,” Microbial Cell Factories, Vol. 5, 2006, p. 35. doi:10.1186/1475-2859-5-35
- G. Cacace, M. F. Mazzeo, A. Sorrentino, V. Spada, A. Malorni and R. A Siciliano, “Proteomics for the Elucidation of Cold Adaptation Mechanisms in Listeria monocytogenes,” Journal of Proteomics, Vol. 73, No. 10, 2010, pp. 2021-2030. doi:10.1016/j.prot.2010.06.011
- H. Antelmann, J. Bernhardt, R. Schmid, H. Mach, U. Völker and M. Hecker, “First Steps from a Two-Dimensional Protein Index towards a Response-Regulation Map for Bacillus subtilis,” Electrophoresis, Vol. 18, No. 8, 1997, pp. 1451-1463. doi:10.1002/elps.1150180820
- P. Graumann, K. Schröder, R. Schmid and M. A Marahiel, “Cold Shock Stress-Induced Proteins in Bacillus subtilis,” Journal of Bacteriology, Vol. 178, No. 15, 1996, pp. 4611- 4619.
- R. J. Thompson, H. G. A. Bouwer, D. A. Portnoy and F. R. Frankel, “Pathogenicity and Immunogenicity of a Listeria monocytogenes Strain that Requires D-Alanine for Growth,” Infection and Immunity, Vol. 66, No. 8, 1998, pp. 3552-3561.
- R. Gardan, O. Duché, S. Leroy-Sétrin, European Listeria Genome Consortium and J. Labadie, “Role of ctc from Listeria monocytogenes in Osmotolerance,” Applied and Environmental Microbiology, Vol. 69, No. 1, 2003, pp. 154-161. doi:10.1128/AEM.69.1.154-161.2003
- U. Völker, S. Engelmann, B. Maul, S. Riethdorf, A. Völker, R. Schmid, H. Mach and M. Hecker, “Analysis of the Induction of General Stress Proteins of Bacillus subtilis,” Microbiology, Vol. 140, No. Pt 4, 1994, pp. 741-52. doi:10.1099/00221287-140-4-741
- B. Pieterse, J. Rob, R. J. Leer, F. H. J. Schuren and M. J. van der Werf, “Unravelling the Multiple Effects of Lactic acid stress on Lactobacillus plantarum by Transcription Profiling,” Microbiology, Vol. 151, No. Pt12, 2005, pp. 3881-3894. doi:10.1099/mic.0.28304-0
- A. Hartke, S. Bouché, J-C. Giard, A. Benachour, P. Boutibonnes and Y. Auffray, “The Lactic Acid Stress Response of Lactococcus lactis subsp. lactis,” Current Microbiology, Vol. 33, No. 3, 1996, pp. 194-199. doi:10.1007/s002849900099
- C.-C. Tsou , C. Chiang-Ni, Y.-S. Lin, W.-J. Chuang, M. T. Lin, C. C. Liu and J. J. Wu, “An Iron-Binding Protein, Dpr, Decreases Hydrogen Peroxide Stress and Protects Streptococcus pyogenes Against Multiple Stresses,” Infection and Immunity, Vol. 76, No. 9, 2008, pp. 4038- 4045. doi:10.1128/IAI.00477-08
- T. D. Caldas, A. E. Yaagoubi and G. Richarme, “Chaperone Properties of Bacterial Elongation Factor EF-Tu,” The Journal of Biological Chemistry, Vol. 273, No. 19, 1998, pp. 11478-11482. doi:10.1074/jbc.273.19.11478
- T. Hanawa, M. Fukuda, H. Kawakami, H. Hirano, S. Kamiya and T. Yamamoto, “The Listeria monocytogenes DnaK Chaperone Is Required for Stress Tolerance and Efficient Phagocytosis with Macrophages,” Cell Stress Chaperones, Vol. 4, No. 2, 1999, pp. 118-128.
- C. G. M.Gahan, J. O’Mahony and C. Hill, “Characterization of the groESL Operon in Listeria monocytogenes: Utilization of Two Reporter Systems (gfp and hly) for Evaluating in Vivo Expression,” Infection and Immunity, Vol. 69, No. 6, 2001, pp. 3924-3932. doi:10.1128/IAI.69.6.3924-3932.2001
- O. Fayet, T. Ziegelhoffer and C. Georgopoulos, “The groES and groEL Heat Shock Gene Products of Escherichia coli Are Essential for Bacterial Growth at All Temperatures,” Journal of Bacteriology, Vol. 171, No. 3, 1989, pp. 1379-1385.
- M. W. Qoronfleh, J. E. Gustafson and B. J. Wilkinson, “Conditions that Induce Staphylococcus aureus Heat Shock Proteins also Inhibit Autolysis,” FEMS Microbiology Letters, Vol. 66, No. 1, 1998, pp. 103-107. doi:10.1111/j.1574-6968.1998.tb13189.x
- J. K. Powell and K. Y. Young, “Lysis of Escherichia coli by Beta-Lactams which bind Penicillin-Binding Proteins 1a and 1b: Inhibition by Heat Shock Proteins,” Journal of Bacteriology, Vol. 173, No. 13, 1991, pp. 4021-4026.
- V. K. Singh, S. Utaida, L. S. Jackson, R. K. Jayaswal, B. J. Wilkinson and N. R. Chamberlain, “Role of dnaK locus in Tolerance of Multiple Stresses in Staphylococcus aureus,” Microbiology, Vol. 153, No. 9, 2007, pp. 3162- 3173. doi:10.1099/mic.0.2007/009506-0
- P. A. J. de Boer, R. E. Crossley and L. I. Rothfield, “Central Role for the Escherichia coli minC Gene Product in Two Different Cell Division-Inhibition Systems,” Proceedings of the National Academy of Sciences of USA, Vol. 87, No. 3, 1990, pp. 1129-1133. doi:10.1073/pnas.87.3.1129
- P. A. J. de Boer, R. E. Crossley and L. I. Rothfield, “A Division Inhibitor and a Topological Specificity Factor Coded for by the Minicell Locus Determine Proper Placement of the Division Septum in E. coli,” Cell, Vol. 56, No. 4, 1989, pp. 641-649. doi:10.1016/0092-8674(89)90586-2
- J. P. Claverys, M. Prudhomme and B. Martin, “Induction of Competence Regulons as General Stress Responses in Gram-positive Bacteria,” Annual Reviews in Microbiology, Vol. 60, 2006, pp. 451-475. doi:10.1146/annurev.micro.60.080805.142139
- G. E. Piñas, P. R. Cortes, A. G. A. Orio and J. Echenique, “Acidic Stress Induces Autolysis by a CSP-Independent ComE Pathway in Streptococcus pneumoniae,” Microbiology, Vol. 154, No. Pt 5, 2008, pp. 1300-1308. doi:10.1099/mic.0.2007/015925-0
- R. R. Draheim, A. F. Bormans, R. Z. Lai and M. D. Manson, “Tryptophan Residues Flanking the Second Transmembrane helix (TM2) Set the Signalling State of the Tar Chemoreceptor,” Biochemistry, Vol. 44, No. 4, 2005, pp. 1268-1277. doi:10.1021/bi048969d
- A. S. Miller and J. J. Falke, “Side Chains at the Membrane-Water Interface Modulate the Signalling State of a Transmembrane Receptor,” Biochemistry, Vol. 43, No. 7, 2004, pp. 1763-1770. doi:10.1021/bi0360206
- R. Carballido-López and A. Formstone, “Shape Determination in Bacillus subtilis,” Current Opinion in Microbiology, Vol. 10, 2007, pp. 611-616. doi:10.1016/j.mib.2007.09.008
- R. Carballido-López and A. Formstone, Y. Li, S. D. Herlich, P. Noirot and J. Errington, “Actin Homolog MreBH Governs Cell Morphogenesis by Localization of the Cell Wall Hydrolase LytE,” Developmental Cell, Vol. 11, No. 3, 2006, pp. 399-409. doi:10.1016/j.devcel.2006.07.017
- D. Brekasis and M. Paget, “A Novel Sensor of NADH/ NAD Redox Poise in Streptomyces coelicolor A3(2),” The EMBO Journal, Vol. 22, No. 18, 2003, pp. 4856-4865. doi:10.1093/emboj/cdg453
- [61] M. Schau, Y. Chen and F. M. Hulett, “Bacillus subtilis YdiH Is a Direct Negative Regulator of the cydABCD Operon,” Journal of Bacteriology, Vol. 186, No. 14, 2004, pp. 4585-4595. doi:10.1128/JB.186.14.4585-4595.2004
- [62] M. Pagels, S. Fuchs, J. Pané-Farré, C. Kohler, L. Menschner, M. Hecker, P. J. McNamarra, M. C. Bauer, C. von Wachenfeldt, M. Liebeke, M. Lalk, G. Sander, C. von Eiff, R. A. Proctor and S. Engelmann, “Redox Sensing by a Rex-family Repressor Is Involved in the Regulation of Anaerobic Gene Expression in Staphylococcus aureus,” Molecular Microbiology, Vol. 76, No. 5, 2010, pp. 1142- 1161. doi:10.1111/j.1365-2958.2010.07105.x
- [63] Y. Boucher and W. F. Doolittle, “The Role of Lateral Gene Transfer in the Evolution of Isoprenoid Biosynthesis Pathways,” Molecular Microbiology, Vol. 37, No. 4, 2000, pp. 703-716. doi:10.1046/j.1365-2958.2000.02004.x
- [64] R. S. Putra, A. Disch, J. M. Bravo and M. Rohmer, “Distribution of Mevalonate and Glyceraldehyde 3-Phosphate/ Pyruvate Routes for Isoprenoid Biosynthesis in some Gram-Negative Bacteria and Mycobacteria,” FEMS Microbiology Letters, Vol. 164, No. 1, 1998, pp. 169-175. doi:10.1111/j.1574-6968.1998.tb13082.x
- [65] S. M. C. Newton, P. E. Klebba, C. Raynaud, Y. Shao, X. Jiang, I. Dubail, C. Archer, C. Frehel and A. Charbit, “The svpA-srtB Locus of Listeria monocytogenes: Furmediated Iron Regulation and Effect on Virulence,” Molecular Microbiology, Vol. 55, No. 3, 2005, pp. 927-940. doi:10.1111/j.1365-2958.2004.04436.x
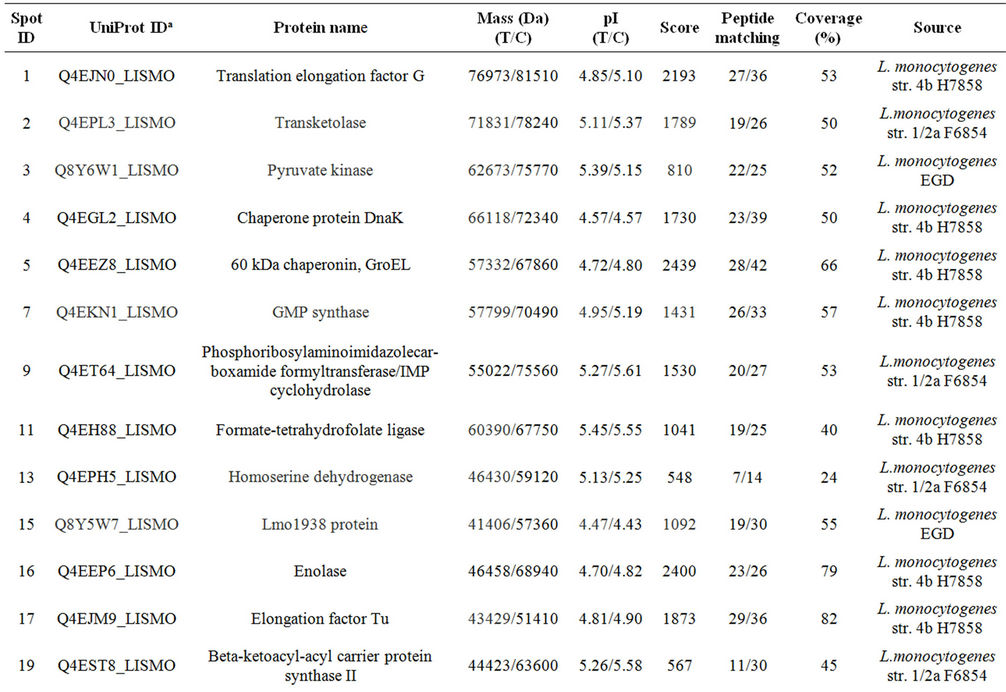
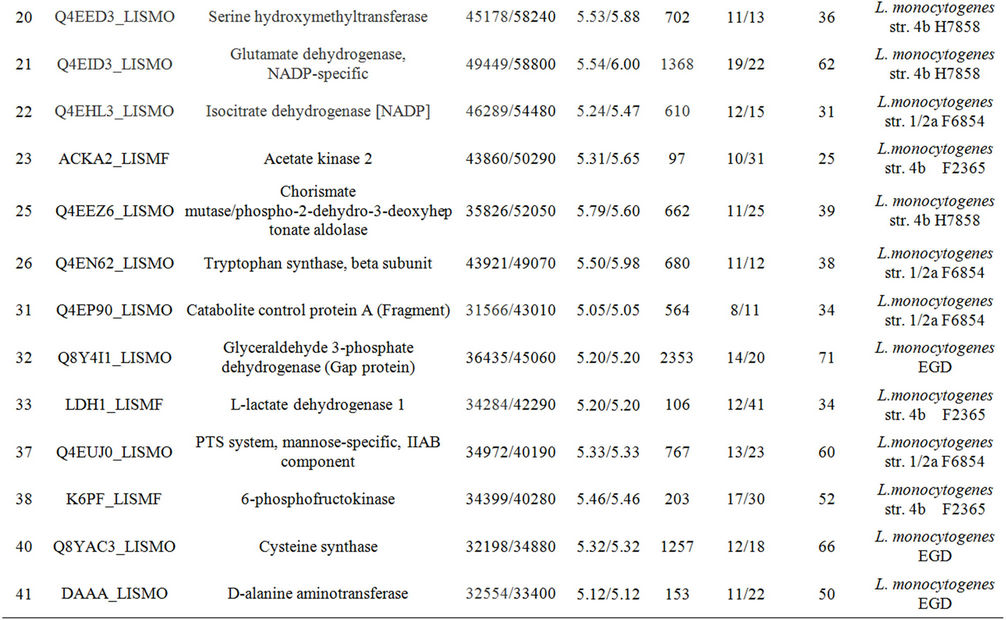
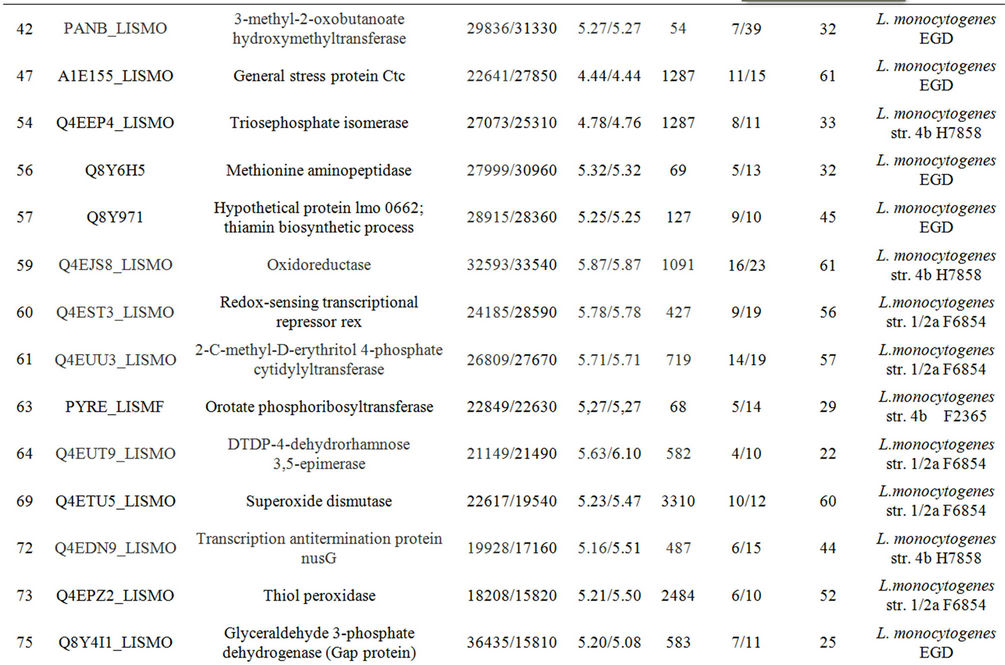
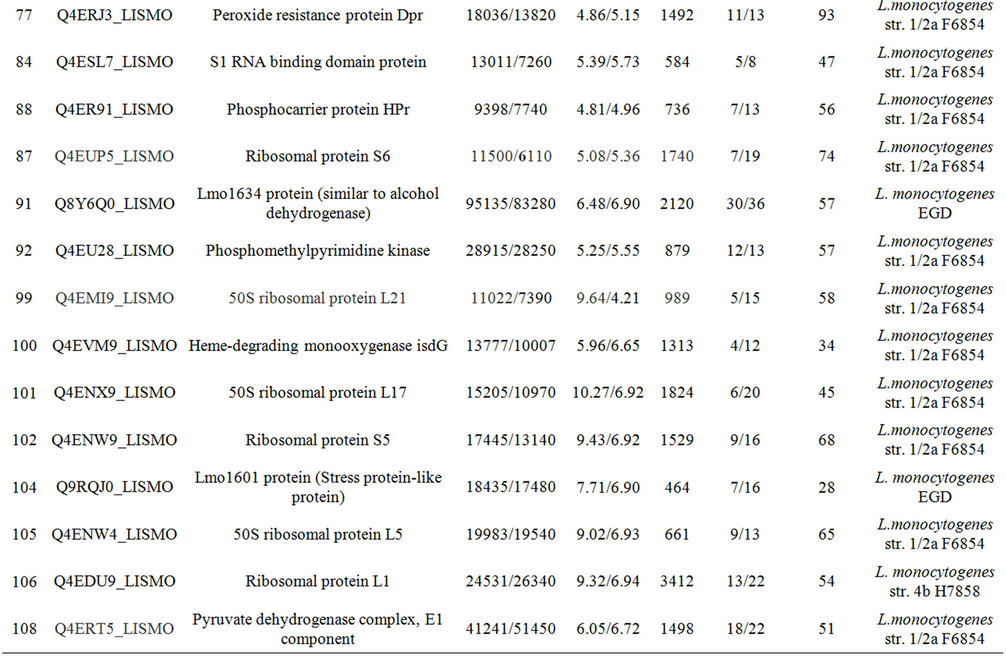
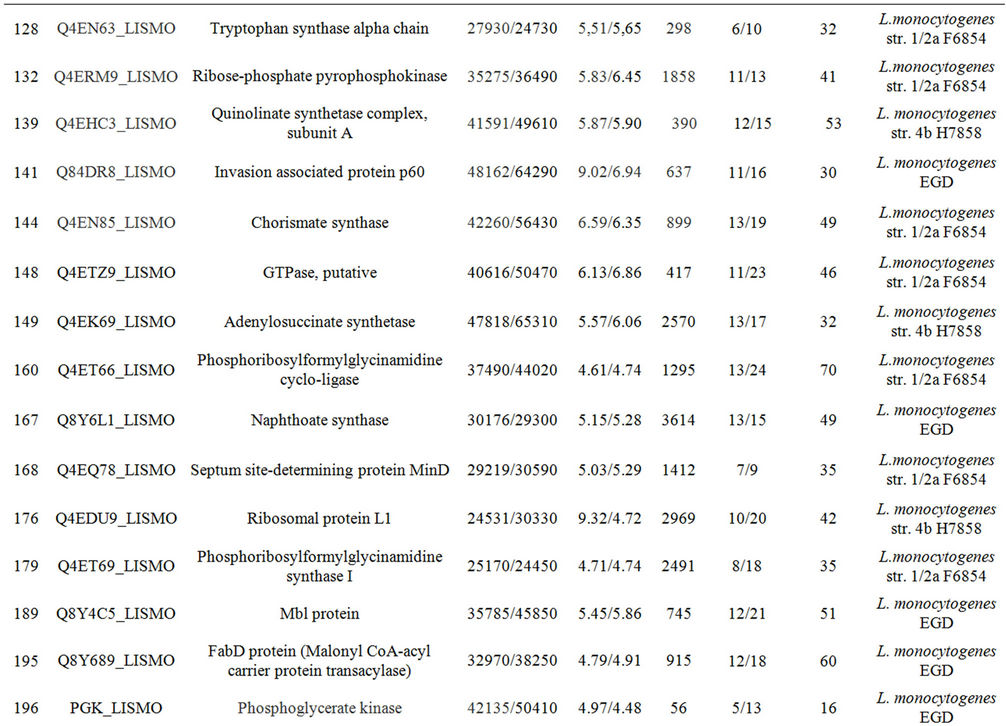
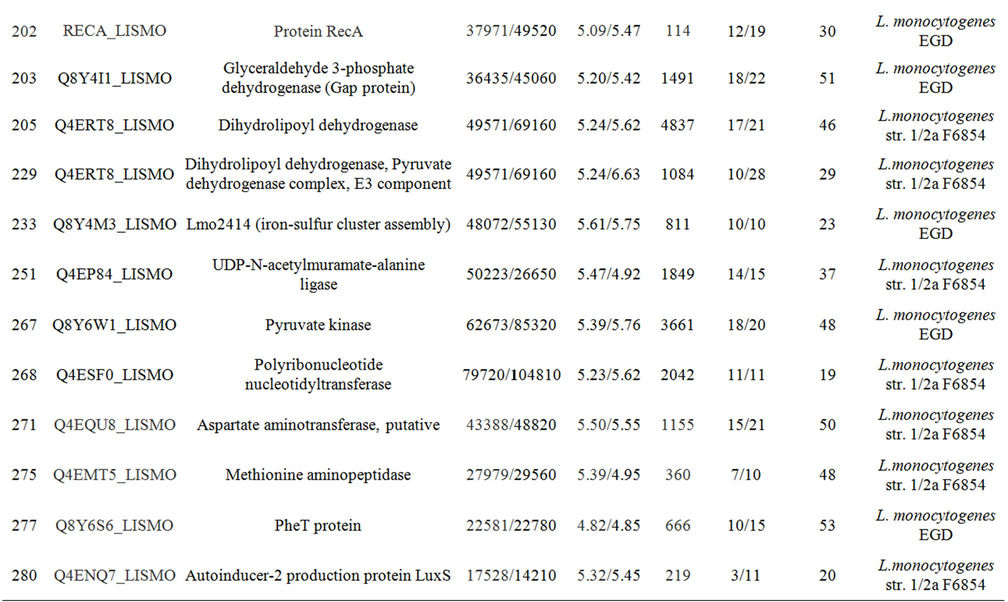
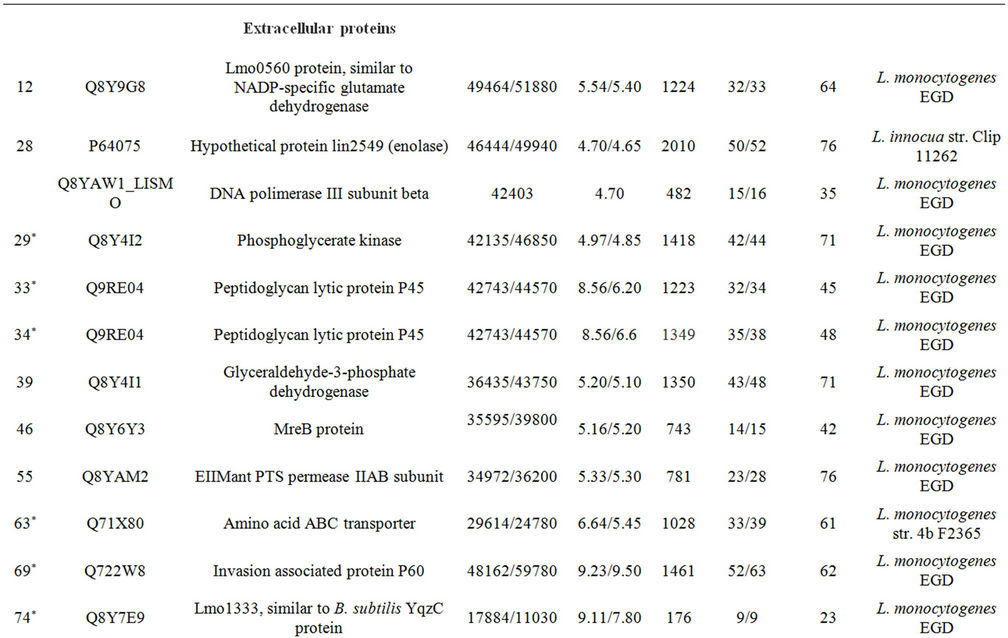
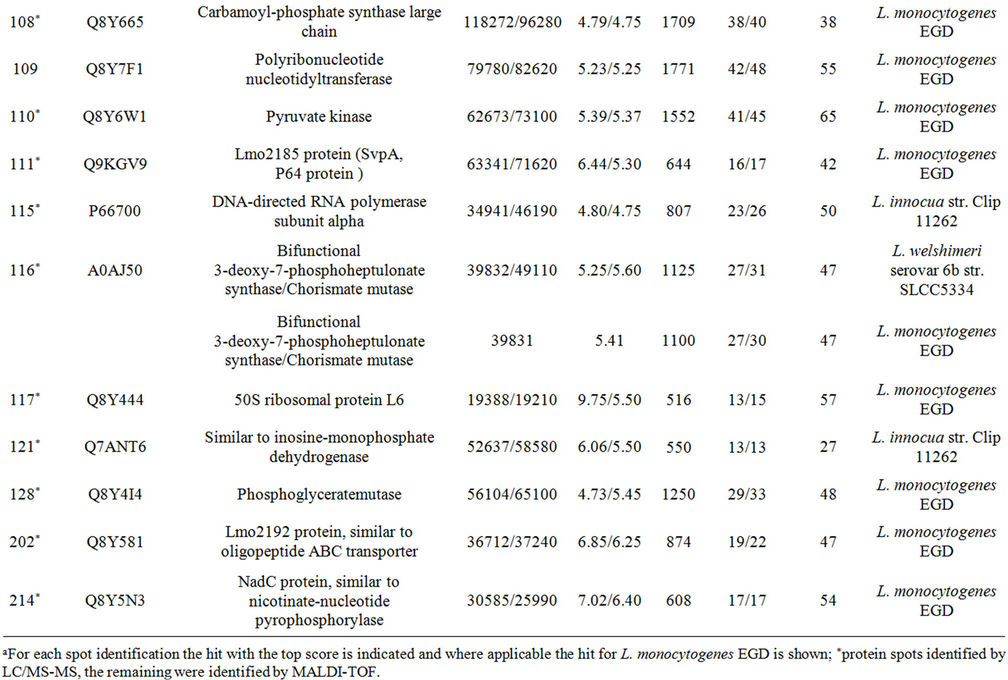
Supplementary Table A. Identification of proteins differentially expressed under autolysis and non-autolysis condition.
NOTES
*Corresponding author.

