Paper Menu >>
Journal Menu >>
 J. Biomedical Science and Engineering, 2010, 3, 415-421 JBiSE doi:10.4236/jbise.2010.34057 Published Online April 2010 (http://www.SciRP.org/journal/jbise/). Published Online April 2010 in SciRes. http://www.scirp.org/journal/jbise Insilico structural analysis of parasporin 2 protein sequences of non-toxic bacillus thuringiensis Ayyasamy Mahalakshmi, Rajaiah Shenbagarathai PG Research Department of Zoology and Biotechnology, Lady Doak College, Madurai-2, Tamilnadu, India. Email: shenbagarathai@rediffmail.com Received 17 September 2009; revised 5 December 2009; accepted 7 December 2009. ABSTRACT The unusual and remarkable property of parasporin 2 of non-insecticidal Bacillus thuringiensis is specifi- cally recognizing and selectively targeting human leukemic cell lines. The 37-kDa inactive nascent protein is proteolytically cleaved to the 30-kDa ac- tive form that loses both the N-terminal and the C-terminal segments. Accumulated cytological and biochemical observations on parasporin-2 imply that the protein is a pore-forming toxin. To confirm the hypothesis, insilico analysis was performed us- ing homology modeling. The resulting model of parasporin 2 protein is unusually elongated and mainly comprises long β-strands aligned with its long axis. It is similar to aerolysin-type β-pore-forming toxins, which strongly reinforce the pore-forming hypothesis. The molecule can be divided into three domains. Domain 1, comprising a small β-sheet sandwiched by short α-helices, is probably the tar- get-binding module. Two other domains are both β-sandwiches and thought to be involved in oli- gomerization and pore formation. Domain 2 has a putative channel-forming β-hairpin characteristic of aerolysin-type toxins. The surface of the protein has an extensive track of exposed side chains of serine and threonine residues. The track might orient the molecule on the cell membrane when domain 1 binds to the target until oligomerization and pore forma- tion are initiated. The β-hairpin has such a tight structure that it seems unlikely to reform as postu- lated in a recent model of pore formation developed for aerolysin-type toxins. Parasporin 2 (Accession no: BAD35170) protein sequence analysis indicated two different domains namely, aerolysin toxin and clos- tridium toxin domain based on different database searches (CDD and Pfam). It showed a close similar- ity with the available PDB template (PDB id: 2ZTB) of parasporin which has cytocidal activity against MOLT-4, HL60 and Jurkat cell lines. Based on the PSI Blast analysis, 3D structures of the domains were predicted by using Swiss model server. Accu- racy of the prediction of 3D structure of different domains of parasporin protein was further vali- dated by Ramachandran plot and PROCHECK (G-value). The structure is dominated by β-strands (67%, S1-12), most of which are remarkably ex- tensive, running all or most of the longer axis of the molecule. This study helped to elucidate the 3D structure of parasporin 2 (Acc. No. BAD35170) which might enable to probe further its specific mechanism of action. Though the similarity is ob- served in the domain architecture, there is variation in the regions of the domains even among the same group of parasporin 2. Docking of this model struc- ture and experimental structure with specific recep- tors of the cancer cells will facilitate to explore mechanism of parasporin 2 action and also provide information about its evolutionary relationship with toxic Cry proteins. Keywords: Parasporin; Homology Model; Non-Toxic; Bacillus thuringiensis; Cell Lines 1. BACKGROUND Since the incidence of new cancer patients is increasing annually due to altered food habits and life styles, efforts are being made worldwide to identify new molecular markers and therapeutic agents for the purpose of diagno- sis and treatment of the same. The existing chemothera- peutics not only affect tumor cells but also normal cells. Hence, search for compounds which can specifically target the cancer cells will overcome the existing problem [1]. At present, four genealogically different parasporins are identified as parasporin-1 to parasporin-4 that has the ability to specifically act against cancer cells [2]. This area of research is still under exploration, since a very few of the literature is available related to parasporin structure and mechanism of action. Parasporin 2 is known to inter- act with GP-I protein and the cell death induced by parasporin-2 is non-apoptotic, although the apoptotic process occurs when the cell damage proceeded slowly. 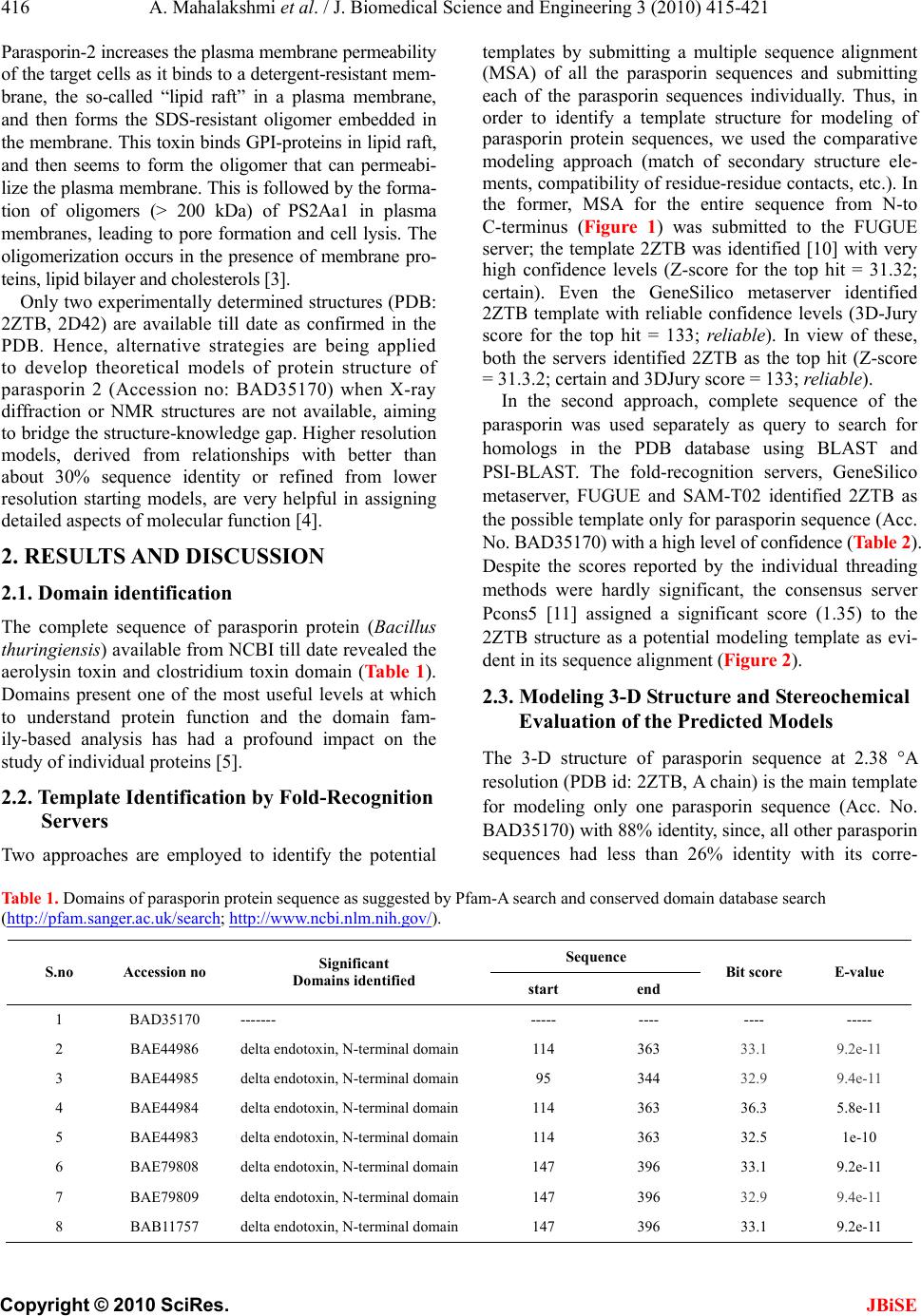 A. Mahalakshmi et al. / J. Biomedical Science and Engineering 3 (2010) 415-421 Copyright © 2010 SciRes. JBiSE 416 Parasporin-2 increases the plasma membrane permeability of the target cells as it binds to a detergent-resistan t mem- brane, the so-called “lipid raft” in a plasma membrane, and then forms the SDS-resistant oligomer embedded in the membrane. This toxin binds GPI-proteins in lipid raft, and then seems to form the oligomer that can permeabi- lize the plas ma me mbrane. Th is is follo wed by the forma- tion of oligomers (> 200 kDa) of PS2Aa1 in plasma membranes, leading to pore formation and cell lysis. The oligomerization occurs in the presence of membrane pro- teins, lipid bilaye r and cholesterols [3]. Only two experimentally determined structures (PDB: 2ZTB, 2D42) are available till date as confirmed in the PDB. Hence, alternative strategies are being applied to develop theoretical models of protein structure of parasporin 2 (Accession no: BAD35170) when X-ray diffraction or NMR structures are not available, aiming to bridge the structure-knowledge gap. Higher resolution models, derived from relationships with better than about 30% sequence identity or refined from lower resolution starting models, are very helpful in assigning detailed aspects of molecular function [4]. 2. RESULTS AND DISCUSSION 2.1. Domain identification The complete sequence of parasporin protein (Bacillus thuringiensis) available from NCBI till date revealed the aerolysin toxin and clostridium toxin domain (Ta bl e 1). Domains present one of the most useful levels at which to understand protein function and the domain fam- ily-based analysis has had a profound impact on the study of individual proteins [5]. 2.2. Template Identification by Fold-Recognition Servers Two approaches are employed to identify the potential templates by submitting a multiple sequence alignment (MSA) of all the parasporin sequences and submitting each of the parasporin sequences individually. Thus, in order to identify a template structure for modeling of parasporin protein sequences, we used the comparative modeling approach (match of secondary structure ele- ments, compatibility of residue-residue contacts, etc.). In the former, MSA for the entire sequence from N-to C-terminus (Figure 1) was submitted to the FUGUE server; the template 2ZTB was identified [10] with very high confidence levels (Z-score for the top hit = 31.32; certain). Even the GeneSilico metaserver identified 2ZTB template with reliable confidence levels (3D-Jury score for the top hit = 133; reliable). In view of these, both the servers identified 2ZTB as the top hit (Z-score = 31.3.2; certain and 3DJury score = 133; reliable). In the second approach, complete sequence of the parasporin was used separately as query to search for homologs in the PDB database using BLAST and PSI-BLAST. The fold-recognition servers, GeneSilico metaserver, FUGUE and SAM-T02 identified 2ZTB as the possible template only for parasporin sequence (Acc. No. BAD35170) with a high level of confidence (Table 2). Despite the scores reported by the individual threading methods were hardly significant, the consensus server Pcons5 [11] assigned a significant score (1.35) to the 2ZTB structure as a potential modeling template as evi- dent in its sequence alignment (Figure 2). 2.3. Modeling 3-D Structure and Stereochemical Evaluation of the Predicted Models The 3-D structure of parasporin sequence at 2.38 A resolution (PDB id: 2ZTB, A chain) is the main template for modeling only one parasporin sequence (Acc. No. BAD35170) with 88% identity, since, all other p ar as por in sequences had less than 26% identity with its corre- Table 1. Domains of parasporin protein sequence as suggested by Pfam-A search and conserved domain database search (http://pfam.sanger.ac.uk/search; http://www.ncbi.nlm.nih.gov/). Sequence S.no Accession no Significant Domains identified start end Bit score E-value 1 BAD35170 ------- ----- ---- ---- ----- 2 BAE44986 delta endotoxin, N-terminal domain 114 363 33.1 9.2e-11 3 BAE44985 delta endotoxin, N-terminal domain 95 344 32.9 9.4e-11 4 BAE44984 delta endotoxin, N-terminal doma i n 114 363 36.3 5.8e-11 5 BAE44983 delta endotoxin, N-terminal domai n 114 363 32.5 1e-10 6 BAE79808 delta endotoxin, N-terminal domain 147 396 33.1 9.2e-11 7 BAE79809 delta endotoxin, N-terminal domain 147 396 32.9 9.4e-11 8 BAB11757 delta endotoxin, N-termi n a l d omain 147 396 33.1 9.2e-11  A. Mahalakshmi et al. / J. Biomedical Science and Engineering 3 (2010) 415-421 Copyright © 2010 SciRes. JBiSE 417 Figure 1. Multiple sequence alignment of parasporin protein residues *-conserved residues. Ta b le 2 . Summary of the best template sequence profile that was generated at the end of 3rd iteration of PSI Blast analysis using the parasporin protein sequence of B.thuringiensis as query. Threshold PSI-BLAST E-value = 0.001. S.no Accession no Query region Template sequence id Region of template seq aligned % identityGap (%)E-valueAnnotation of the template 1 BAD35170 28-278 2ZTB 2-252 88 0 2e-114Parasporin 2 crystal structure of Bacillus thuringiensis 2 BAE44986 119-383 1DLC 18-251 26 15 1e-12 Crystal Structure Of Insecticidal Delta-endotoxin. 3 BAE44985 101-364 1DLC 19-251 26 15 Crystal Structure Of Insecticidal Delta-endotoxin. 4 BAE44984 109-383 1DLC 5-251 26 16 6e-13 Crystal Structure Of Insecticidal Delta-endotoxin. 5 BAE44983 109-383 1DLC 5-251 26 16 1e-12 Crystal Structure Of Insecticidal Delta-Endotoxin 6 BAE79808 153-416 1DLC 19-251 25 15 2e-12 Crystal Structure Of Insecticidal Delta-Endotoxin 7 BAE79809 153-416 1DLC 19-251 26 15 2e-12 Crystal Structure Of Insecticidal Delta-Endotoxin 8 BAB11757 153-416 1DLC 19-251 25 15 2e-12 Crystal Structure Of Insecticidal Delta-Endotoxin  A. Mahalakshmi et al. / J. Biomedical Science and Engineering 3 (2010) 415-421 Copyright © 2010 SciRes. JBiSE 418 Figure 2. Sequence alignment of the parasporin protein (Acc. No. BAD35170) with its tem- plate (PDB id: 2ZTB-A chain). sponding template (Figure 3). The final averaged and optimized model passed all the tests implemented in the stereochemistry-evaluating WHATCHECK suite [12-15] and in the VERIFY3D program, which uses contact po- tentials to assess whether the modeled amino acid resi- dues occur in the environment typical for globular pro- teins with h ydrophobic core an d solvent-e xposed surface (Eisenberg et al., 1997). Moreover, the reasonable ener- gies are rarely observed for misfolded structures. Thus, the scores reported for our model by WHATCHECK (Z-score-4.1) and VERIFY3D (average score 0.3, no regions scored lower than 0) suggest that both its three-dimensional fold and the conformation of individ- ual residues are reasonable. The selected model, the value of the objective func- tion, reported as current energy is in the same range as that if the template is aligned with its own sequence. On an average, 99.1% of the residues are found in the al- lowed region of Ramachandran map, PROCHECK con- siders the model to be very good if it has 90% of the residues in the most favored region. The inter-atomic distances are within acceptable range. Verify3D score is greater than zero for the entire model (1 to 274 residues). The models were also evaluated using Colorado3D server, which facilitates the change of amino acid win- dow size when calculating the overall score. Two win- dow sizes, 5 and 21, were used to calculate the average Verify3D and ProsaII score per residue for the model [12,13]. The scores calculated using these two window sizes were found to be very similar. The template and target models were rendered with the residues color-coded based on ProsaII and Verify3D scores. With ProsaII score-based coloring, most of the residues are green and yellow (i.e., average score) in both the target and tem- plate proteins. Z-score of a model is a measure of com- patibility between its sequence and structure, the model Z-score should be comparable to that of the template. With Verify3D score-based coloring, even the template proteins has residues in red color (i.e., bad score) al- though the number of such residues are more in the tar- gets. 2.4. Description of the Model The structure is dominated by β-strands (67%, S1–12), most of which are remarkably extensive, running all or most of the longer axis of the molecule (Figure 3). The longest strand (S8), comprising 21 residues, runs the whole length of the molecule; three others (S11, S12, S14) span two-thirds of the molecule’s length. The molecule can be divided into three domains: domain 1 (Val29-His85, Gly145-Ser173), domain 2 (Ile36-Pro53, Ala81-Asp138, Glu160-Ser175, Gly211-Gln236), and do- main 3 (Val54-Asn80, Phe139-Leu159, Thr237-Ala250). Domain 1 is composed of a short N-terminal β-strand Figure 3. Homology model of the parasporin protein sequence (Acc.No. BAD35170).  A. Mahalakshmi et al. / J. Biomedical Science and Engineering 3 (2010) 415-421 Copyright © 2010 SciRes. JBiSE 419 (S1: 30-34), a α/β structure (H1 and S2: 46-66), a β-hairpin (S9 and S10: 152-168), and a α-helix (H2: 79 -83). α-Helices are found only in this domain. The two α-helices, H1 and H2, are close together and short, oc- cupying only 8.9% of the molecule. Domains 2 and 3 are both β-sandwiches: the former is made of a two-stranded β-hairpin (S6 and S7) and a curled anti-parallel five stranded β-sheet including S3, S9, S12, S5 and S8; and the latter is of antiparallel three-stranded and two- stranded β-sheets (S4, S9 and S12; S5 and S8). These results (Figure 3, Table 3) are consistent with earlier reports [14]. This model is expected to yield insight into the molecular function and mechanism of parasporin action [15]. 3. CONCLUSIONS The initial step in cytocidal action of PS2Aa1 is the spe- cific binding of this cytotoxin to a putative receptor lo- cated in the lipid rafts, followed by its oligomerization and pore formation in plasma membrane.Secondary structures observed in the model (Accession No: BAD 35170) are organized virtually in the same way as the experimentally determined parasporin-2 protein (PDb id: 2ZTB). Even though 2ZTB and 2D42 belong to the parasporin type 2 protein sequence, BAD35170 bears greater similarity to 2ZTB in its domain organization. 4. METHODS 4.1. Databases The amino acid sequences of the experimentally chara- cterized parasporin sequences (Ta ble 4) were retrieved from the protein sequence database at NCBI http:// www.ncbi.nlm.nih.gov. The 3-D structures of proteins Table 3. Structural superposition report of the model generated for BAD35170 with its corresponding template, 2ZTB. Table 4. List of parasporin sequences retrieved from NCBI database. S.no Accession no Type of protein 1 BAD35170 Parasporin 2 Ab cytotoxic Crystal protein (Bacillus thuringiensis) 2 BAE44986 Putative uncharacterized protein 1 (Bacillus thuringiensis) 3 BAE44985 Putative uncharacterized protein 2 (Bacillus thuringiensis) 4 BAE44984 Putative uncharacterized protein 3 (Bacillus thuringiensis) 5 BAE44983 Putative uncharacterized protein 4 (Bacillus thuringiensis) 6 BAE79808 81-k da leukemia protein (Bacillus thuringiensis) 7 BAE79809 Cry 31-like 82-k da protein (Bacil- lus thuringiensis) 8 BAB11757 81-k da leukemia toxin (Bacillus thuringiensis) were obtained from the protein data bank [16]. The fold classification of proteins is from the SCOP data- base [17]. 4.2. Servers Protein sequence databases we r e searched using PSI-B LAST [18] servers at NCBI, PHYRE (successo r of 3 D-PSSM) [19], SAM-T02 [20] and GeneSilico Metaserver [21] were used for fold-recognition. Multiple sequence alignments were obtained using the CLUSTALW server [22]. Verify3D [13] and Colorado3D [23] were used to evaluate the models. All the servers were used with default values for the various parameters, except where mentioned otherwise. 4.3. Software and Hardware SwissPDBviewer [24] was used for visualization and/or rendering. The stereochemical quality of the generated model was assessed using PROCHECK [10]. Default values were used for all the parameters, unless specified otherwise. 4.4. Te m plate-Target Sequence Al ignmen t The parasporin sequences were submitted for structure prediction using comparative modeling technique. The preliminary models were obtained using unrefined pairwise alignments reported by PSI-BLAST [25]. Energy minimization was carried out using GROMOS96 [26] until all inconsistencies in geometry were rectified and all the short contacts were relieved. The stereochemical and energetic properties of modeling intermediates and of the final model were evaluated using WHATCHECK [27] and VERIFY3D [28]. Semi-automated and manual manipulations with protein structures and sequence–structure alignments were conducted using SWISS-PDB VIEWER [24]. All the servers provide alignment of the submitted  A. Mahalakshmi et al. / J. Biomedical Science and Engineering 3 (2010) 415-421 Copyright © 2010 SciRes. JBiSE 420 parasporin sequence (target) with the sequence of the potential hits (templates). 4.5. Validation of Predicted 3-D Structures The stereochemical properties of predicted 3-D structures were assessed by PROCHEC K and t he residue envi ronm ents by Verify3D and Colorado3D. Regions that are found by these servers as poorly modeled were improved by manual adjustment of alignments and re-modeling. 5. ACKNOWLEDGEMENTS The authors acknowledge N.Lavanya Roselin for acquisition of data. Authors express their heartfelt thanks for DBT-BIF in providing the necessary infrastructure facility in the collection, analysis, and inter- pretation of data; in the writing of the manuscript; and in the decision to submit the manuscript for publication. BLAST server: http://www.ncbi.nlm.nih.gov/BLAST/ Colorado3D: http://asia.genesilico.pl/colorado3d/ FUGUE: http://www-cryst.bioc.cam.ac.uk/fugue/ GeneSilico Metaserver: http://genesilico.pl/meta PDB: http://www.rcsb.org PHYRE: http://www.sbg.bio.ic.ac.uk/~phyre PROCHECK: http://www.biochem.ucl.ac.uk/~roman / p r o c h ec k / p r o c he c k . html SCOP database: http://scop.mrc-lmb.cam.ac.uk/scop/ SwissPDBviewer: http://ca.expasy.org/spdbv/ Verify_3D: http://nihserver.mbi.ucla.edu/Verify_3D/ REFERENCES [1] Mizuki, E., Ohba, M., Akao, T., Yamashita, S., Saitoh, H. and Park, Y.S. (1999) Unique activity associated with non-insecticidal Bacillus thuringiensis parasporal inclu- sions: In vitro cell-killing action on human cancer cells. Journal of Applied Microbiology, 86, 477-86. [2] Katayama, H., Yokota, H., Akao, T., Nakamura, O., Ohba, M., Mekada, E. and Mizuki, E. (2005) Pa raspo rin-1, a nove l cytotoxic protein to human cells from non-insecticidal parasporal inclusions of Bacillus thuringiensis. Biochem- istry, 137, 17-25. [3] Abe, Y., Shimada, H. and Kitada, S. (2008) Raft-targeting and oligomerization of parasporin-2, a Bacillus thuringiensis crystal protein with Anti-Tumour activity. Biochemistry, 143(2), 269-275. [4] Murray, D. and Honig, B. (2002) Electrostatic control of the membrane targeting of C2 domains. Molecular Cell, 9, 145-154. [5] Copley, R.R., Doerks, T., Letunic, I. and Bork, P. (2002) Protein domain analysis in the era of complete genomes. FEBS Letters, 20, 129-134. [6] Godzik, A. (2003) Fold recognition methods. Methods of Biochemical Analysis, 44, 525-546. [7] Kurowski, M.A., Bujnicki, J. M. (2003) GeneSilic o protein structure prediction meta-server. Nucleic Acids Research, 31, 3305-3307. [8] Akiba, T., Abe, Y., Kitada, S., Kusaka, Y., Ito, A., Ichi- matsu, T., Katayama, H., Akao, T., Higuchi, K., Mizuk i, E . , Ohba, M., Kanai, R. and Harata, K. (2009) Crystal struc- ture of the parasporin-2 of Bacillus thuringiensis toxin that recognizes cancer cells. Journal of Molecular Biol- ogy, 386, 121-133. [9] Lundstrom, J., Rychlewski, L., B ujnicki, J.M. an d Elofsson, A. (2001) Pcons: A neural-network-based consensus predic- tor that improves fold recognition. Protein Science, 10, 2354-2362. [10] Laskowski, R.A., MacArthur, M.W., Moss, D.S. and Thornton, J.M. (1993) PROCHECK: A program to check the stereochemical quality of protein structures. J Appl Cryst, 26, 283-291. [11] Hooft, R.W.W., Vriend, G., Sander, C. and Abola, E.E. (1996) Errors in protein structures. Nature, 381, 272-272. [12] Sippl, J. (1993) Recognition of error s in three dimensional structures of proteins. Proteins, 17, 355-362. [13] Lüthy, R., Bowie, J.U. and Eisenberg, D. (1992) Assessment of protein models with three-dimensional profiles. Nature, 5, 83-5. [14] Brenner, S.E. (2001) A tour of structural genomics. Nature Reviews Genetics, 2, 801-809. [15] Berman, H.M., Westbrook, J., Feng, Z., Gilliland, G., Bhat, T.N., Weissig, H., Shindyalov, I.N. and Bourne, P.E. (2000) The protein data bank. Nucleic Acids Research, 28, 235-242. [16] Andreeva, A., Howorth, D., Brenner, S.E., Hubbard, T.J., Chothia, C. and Murzin A.G. (2004) SCOP database in 2004: Refinements integrate structure and sequence fam- ily data. Nucleic Acids Research, 32, 226-229. [17] Altschul, S.F., Madden, T.L., Schaffer, A.A., Zhang, J., Zhang, Z., Miller, W. and Lipman, D.J. (1997) Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Research, 25, 3389-3402. [18] Shi, J., Blundell, T.L. and Mizuguchi, K. (2001) FUGUE: sequence-structure homology recognition using environment- specific substitution tables and structure-dependent gap penalties. Journal of Molecular Biology, 29, 243-57. [19] Kelley, L.A., MacCallum, R.M., Sternberg, M.J.E. (2000) Enhanced genome annotation using structural profiles in the program 3D-PSSM. Journal of Molecular Biology, 299, 499-520. [20] Karplus, K., Karchin, R., Draper, J., Casper, J., Mandel- Gutfreund, Y., Diekhans, M. and Hughey, R. (2003) Combining local-structure, fold-recognition, and new fold methods for protein structure prediction. Proteins, 53, 491-496. [21] Kurowski, M.A. and Bujnicki, J.M. (2003) GeneSilico protein structure prediction meta-server. Nucleic Acids Research, 31, 3305-3307. [22] Thompson, J.D., Higgins, D.G. and Gibson, T.J. (1994) CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Research, 22, 4673-4680. [23] Sasin, M. and Bujnicki, J.M. (2004) COLORADO3D, a web server for the visual analysis of protein structures. Nucleic Acids Research, 32, 586-589. [24] Guex, N. and Peitsch, M.C. (1997) SWISS-MODEL and the Swiss-PdbViewer: An environment for comparative protein modeling. Electrophoresis, 18, 2714-2723. [25] Altschul, S.F, Madden, T.L., Schäffer, A.A., Zhang, J., Zhang, Z., Miller, W. and Lipman, D.J. (1997) Gapped blast and PSI-blast: A new generation of protein database search programs. Nucleic Acids Research, 25(17), 3389-3402.  A. Mahalakshmi et al. / J. Biomedical Science and Engineering 3 (2010) 415-421 Copyright © 2010 SciRes. JBiSE 421 [26] van Gunsteren, W.F., Billeter, S.R., Eising, A.A., Hünen- berger, P.H., Krüger, P., Mark, A.E., Scott, W.R.P. and Tironi, I.G. (1996) Biomolecular simulation. The GRO- MOS96 Manual and User Guide, Zürich, Groningen. [27] Hooft, R.W.W., Vriend, G., Sander, C. and Abola, E.E. (1996) Errors in protein structures. Nature, 381, 272-272. [28] Eisenberg, D., Lüthy, R. and Bowie, J.U. (1997) VERIFY3D: Assessment of protein models with three-dimensional profiles. Methods Enzymol, 277, 396-404. |

