Journal of Cancer Therapy
Vol.4 No.1(2013), Article ID:27909,43 pages DOI:10.4236/jct.2013.41031
Tumorigenic Responses of Cancer-Associated Stromal Fibroblasts after Ablative Radiotherapy: A Transcriptome-Profiling Study
![]()
1Institute of Clinical Medicine, Tromsø, Norway; 2Microarray Resource Centre Tromsø (MRCT), Institute of Clinical Medicine, University of Tromsø, Tromsø, Norway; 3Department of Oncology, University Hospital of North-Norway, Tromsø, Norway.
Email: *ruth.h.paulssen@uit.no
Received November 7th, 2012; revised December 11th, 2012; accepted December 19th, 2012
Keywords: Stereotactic Ablative Radiotherapy (SART); Gene Expression; Cancer-Associated Fibroblasts (CAFs)
ABSTRACT
Cancer-associated fibroblasts (CAFs) are key elements in the progression of cancer and thereby represent important targets for cancer therapies. Increased attention has been given to ablative radiotherapy in the clinics. Therefore, in this study we have aimed at identifying the transcriptional responses occurring in primary CAFs exposed to high-dose irradiation. Established primary CAFs obtained from non-small-cell lung cancer (NSCLC) patient material were irradiated with a single dose of 18 Gy and total RNA was isolated 24 hrs after treatment. Radiation-induced transcriptional alterations were investigated by gene expression analysis using genome-wide microarrays. Obtained results were verified by qRT-PCR of relevant genes. Confirmation of gene expression outcomes was achieved by diverse functional and expression assays including DNA damage response, measurements of reactive oxygen species (ROS) by flow cytometry and senescence-associated β-galactosidase. Irradiation resulted in differential expression of 680 genes of which 557 were upand 127 down-regulated. Of those, 153 genes were differentially expressed with a fold-change greater than 1.0 and an adjusted p-value less than 0.05 across different comparisons (non-irradiated vs. irradiated). Expression patterns revealed profound changes in biological functions and processes involved in DNA repair, apoptosis, p53 pathway, autophagy, senescence, ROS production and immune response. CAFs display proand anti-tumorigenic effects after having received a single high-dose radiation. The measured effects will have an impact on the tumor microenvironment in respect to tumor growth and metastasis.
1. Introduction
Hypofractionated radiotherapy is emerging as a new treatment option for solid neoplasms of different kinds, including but not limited to medically inoperable stage I non-smal-cell lung cancers (NSCLC) [1,2]. Advances in radiotherapy (RT)-technology thus now permit accurate delivery of high-dose (or ablative) RT to target earlystage small tumors with acceptable toxicity to the surrounding normal tissue [3]. Clinical outcomes indicate that ablative RT is associated with improved local control and overall survival compared with conventional RT regimens [2,4]. Despite the encouraging clinical outcomes presented in recent years, little attention has been paid to the biological reactions associated to ablative doses of radiation to tumors.
Tumor progression is a multi-step process orchestrated by many different cell associated to the malign tissue, such as cancer-associated fibroblasts (CAFs), pericytes, lymphocytes, endothelial cells, extracelluar matrix and myeloid cells [5-7]. CAFs are frequently found in the reactive stroma of cancers, and their presence in large number is associated with poor prognosis. When acting on cancer cells, CAFs promote tumor growth and invasion, and enhance angiogenesis by secreting factors that activate endothelial cells and pericytes. Through the secretion of cytokines, CAFs and immune cells can exert both, tumor-suppressing and tumor-promoting effects [8].
Exposure of normal fibroblasts to ionizing radiation has been a common model to study cellular responses to genotoxic stress [9,10]. Aiming for a better understanding of the collateral effects induced by ionizing radiation on normal tissue, some studies have tried to decode the overall changes in gene expression on normal fibroblasts by applying microarray gene technology [9-13]. After functional categorization of differentially expressed genes, some common traits on altered pathways can be denoted from those studies including DNA damage responses, regulation of cell cycle and proliferation, programmed cell death, p53 target genes, signaling pathways, ROS scavenging and ECM remodeling. Despite the available knowledge on normal tissue fibroblast responses to RT, few reports have focused on the effects of RT on intratumoral reactive stromal fibroblast (CAFs). In a recent study from our laboratory we show that CAF survive to relative high single radiation doses, however the cellular phenotype becomes profoundly altered, characterized by the induction of permanent DNA damage responses and the acquisition of a senescent phenoltype and growth arrest [14]. Of note, in that study we show that the expression of some matrix metalloproteinases becomes altered along with overexpression of cell surface integrins.
This work is intended to complement our initial study on radiation-induced responses by tumor-associated fibroblast. Following the rational of using single high radiation doses to reproduce the effects provoked by sterotactic ablative radiation therapy regimens, in this study we perform transcriptional profiling to rule out in which degree CAFs become pro-tumorigenic and/or anti-tumorigenic after irradiation. To our knowledge this is the first time that such a study has been conducted with freshly prepared CAFs.
2. Materials and Methods
2.1. Human Material, Cell Isolation and CAFs Cultures
Human CAFs were harvested from freshly resected nonsmall lung cell carcinoma (NSCLC) tumor tissues. Tumors from eight patients with an average age of 58 years (range 44 - 71) were included in this study. The Regional Ethical Committee (REK-Nord) approved the study, and all patients provided written informed consent. Fibroblasts from tumors were isolated and characterized following standard procedures. Briefly, tumor resections were collected and cut into 1 - 1.5 mm3 pieces. Enzymatic digestion of tissues was carried out for 1.5 hrs in 10 mL DMEM/HAM’S F-12 containing bacterial collagenase (Cat.#C-9407, Sigma-Aldrich, St. Louis, MO, USA) at a final concentration of 0.8 mg/mL. Digested tissue was spun down to eliminate collagenase, and resuspended in fresh growth medium (DMEM/HAM’S F-12 supplemented with 10% FBS). Pure fibroblast cultures were obtained by selective cell detachment from the primary culture mix used 2 mM PBS-EDTA solution, and by further cell propagation in the presence of 10% FBS. Cells were used for experiments after the second passage (2 - 3 weeks). The resulting cultures were characterized for purity and cell identity by flow cytometry using a FITC-conjugated anti-human α-SMA (smooth muscle α-actin) antibody (Abcam; Cat.#ab8211) (data not shown), and by immuno-fluorescent staining with anti-FAP (fibroblast activation protein) antibody (Abcam; Cat.#ab53066) on formalin-fixed CAF cultures.
2.2. Irradiation of Cells
Radiation protocols were established after initial doseescalating pilot trials. Hence, adherent CAFs cultured in flasks were irradiated at RT with high energy photons produced by a Varian clinical linear-accelerator, delivered as single doses of 2, 6, 12 and 18 Gy. Standard parameters for dose delivery was depth 30 mm, beam quality 15 MV, dose-rate 6 Gy/min and field size 20 × 20 cm. Radiation-doses were confirmed to be correct within an acceptable ± 4% by Thermo-Luminescent Dosimeters (TLDs) [15]. Cell survival/death after radiation was assessed by light microscopy during the initial three weeks, by automated cell index analyses (xCelligence, Roche) and by MTT cell proliferation and viability assays [16] at different time points [14]. The extent of radiation-induced apoptosis and necrosis in CAFs was determined by staining for annexin V and propidium iodide respectively (data not shown).
2.3. DNA Damage Response (DDR) Foci Staining
CAFs were cultured in 2-well chamber slides (Nunc, Thermo Fisher Scientific, NY, USA), fixed with 4% PFA in PBS for 10 min at RT and permeabilized with 0.2% Triton in PBS for 8 min. Slides were then exposed to blocking buffer containing 2% HSA in PBS, for 30 min at RT. Primary antibody (Rabbit anti human 53BP1; Cat. #ab36823, Abcam, Cambridge, UK) was diluted in blocking buffer and incubated with CAFs for 45 min at RT. After washing with PBS, cells were incubated with secondary antibody (anti rabbit-Alexa546, Cat.#A11010, Molecular Probes/Invitrogen, Leiden, The Netherlands) in blocking buffer, 30 min at RT. A second wash was followed by preparation of slides in DAPI-FluoromountG (Cat.#0100-20, Southern Biotech, Birmingham, AL, USA). Specimens were examined in a fluorescence microscope (Zeiss Axiophot, Germany) equipped with a Nikon DS-5MC digital camera, and images were processed with Adobe® Photoshop Software (CS5).
2.4. Senescence Associated β-Galactosidase Assays
CAFs were seeded at a density of 20,000 cells per well in uncoated 6-well plates and left for attachment for 24 hrs. Adherent cells were irradiated with a single fraction of 18 Gy. Five days post-irradiation, cultures were washed twice in PBS, and fixed for 5 min at RT with freshly prepared paraformaldehyde (2%). β-galactosidase (5- bromo-4chloro-3-indolyl-B-D-galactopyranoside) staining was achieved following instructions from the manufacturer with the “Senescence Cells Histochemical Staining Kit” (Cat.no CS0030, Sigma-Aldrich, St. Louise, MO, USA). The number of β-galactosidase active and sensecent cells was determined by counting blue cells under a Nikon Eclipse TS100 model light microscope, and randomly selected fields were photographed at 1000× magnification, using an Idea SPOT digital camera.
2.5. ROS Measurements
Intracellular ROS was determined using the CMH2DCFDA probe (Molecular Probes, Invitrogen, Carlsbad, CA, USA) and flow cytometry. When oxidized, this probe can be detected by fluorescence with excitation at 485 nm and emission at 515 nm. Irradiated CAFs were seeded in 6 well cell culture plates (1 × 105) with 2 ml of complete growth medium and incubated overnight at 37˚C, 5% CO2. The next day, the cells were washed 2 times with Hanks Balanced Salt Solution with CaCl2 and MgCl2 (HBSS) (GIBCO, Invitrogen, Carlsbad, CA) and incubated for 30 min with 2.5 μM CM-H2DCFDA in HBSS. Subsequently the cells were trypsinized, resuspended in 0.5 ml HBSS and immediately analysed by flow cytometry (FACSAria, BD Biosciences). Ten thousand events were collected and analyzed by BD FACSDiva Version 5.0.2. The numbers presented are the median values of the fluorescent intensity.
2.6. RNA Preparation and Quality/Quantity Control
Disruption and homogenization of cells were performed in lysis buffer using the MagNa Lyser Instrument (Roche Applied Science, Germany) and according to the manufacturer’s protocol. Subsequently, total RNA was isolated with the MagNa Pure Compact Instrument and the MagNa Pure Compact RNA Isolation Kit (Roche Applied Science, Germany) as previously described [17]. RNA was quantified by measuring absorbance at 260 nm, and RNA purity was determined by the ratios OD260 nm/280 nm and OD230/280 nm using the NanoDrop instrument (NanoDrop® ND-1000, Wilmington, USA). The RNA integrity was determined by electrophoresis using the BioRad Experion Bioanalyzer (Data not shown). All the RNA preparations were verified for possible genomic DNA contamination by a minus-RT-PCR conducted directly on RNA samples, with human genomic DNA as a positive control and amplification of the human housekeeping gene cyclophilin A. Genomic DNA was not detected in RNA preparations (Data not shown).
2.7. Probe Generation and Array Hybridization
Prior to probe generation 1 mg total RNA was amplified with the Ambion®MessageAmpTM II aRNA Amplification Kit (Applied Biosystems Inc., USA). RNA samples were transcribed to cDNA using the Invitrogen superscript cDNA synthesis kit (InvitrogenTM, USA). cDNA and Cy3-labelled with the NimbleGen One-Color DNA Labeling Kit and according to the manufacturer’s protocol (Roche NimbleGen, Germany). Labelled probes were hybridized to a NimbleGen oligo arrays (12 × 135K) with the Roche NimbleGen hybridization system for 16 hours at 42˚C. The microarray slides were washed according to the manufacturer’s protocol and scanned with an Axon GenePix 4000B scanner (Molecular Devices Inc., USA). Features were extracted by using the NimbleScan v2.5 software.
2.8. Data Analysis and Statistical Analysis
Normalization was carried out using quantile normalization [18]. For the purpose of finding differentially expressed genes, we applied an empirical Bayes analysis using the LIMMA package [19], and significance was determined at the 0.05 level corrected for false discovery rate (FDR) using the Benjamini-Hochberg method [20]. Principal component analysis (PCA) was carried out on the data in order to visualize the data structure and look for potential outlier samples [21]. The differentially expressed genes were annotated by Protein Analysis THrough Evolutionary Relationships (PANTHER; http://www.panther.org/). Gene SetEnrichment Analysis (GSEA) was performed using the R statistical package (http://www.broad.mit.edu/gsea/). The microarray data were prepared according to minimum information about a microarray experiment (MIAME) recommendations and deposited in the Gene Expression Omnibus (GEO) database: http://www.ncbi.nlm.nih.gov/geo/. The GEO accession number for the series is GSE37318.
2.9. Validation of Gene Expression by Quantitative Real-Time Polymerase Chain Reaction
Total RNA was prepared representative for four irradiated and non-treated CAF samples each. These were reverse transcribed using Transcriptor First Strand cDNA synthesis Kit (Roche Applied Science, Germany) as described in the manufacturer’s protocol. TaqMan real-time PCR amplification was performed with an ABI HT7900 Instrument (Applied Biosystems) using primers and probes designed with the Universal Probe Library (Roche Applied Science, Germany) and Taqman® Gene Expression Assays (Applied Biosystems). The following primers and probes and inventory assays have been used: MMP11-fw, 5’-GGTGCCCTCTGAGATCGAC-3’, MMP 11-rev, 5’-TTCACAGGGTCAAACTTCCAG-3’, MMP- 11 probe #4; ATG16L2-fw, 5’-GGCCACAATGACCAGAAGAT-3’, AGT16L2-rev, 5’-GGATGACCTGGGTGCAGT-3’, AGT16L2 probe #56; COL7A1-fw, 5’-GCTGGTGCTGCCTTTCTC-3’, COL7A1-rev, 5’-TCCAGGCCGAACTCTGTC-3’, COL7A1 probe #71; IL12A-fw, 5’-CACTCCCAAAACCTGCTGAG-3’, IL12A-rev, 5’-TCTCTTCAGAAGGTGCAAGGGTA-3’, IL12A probe #50; GDF15-fw, 5’-CGGATACTCACGCCAGAAGT-3’, GDF15-rev, 5’-AGAGATACGCAGGTGCAGGT, GDF15 pobe # 28; 5’PPIA-fw, 5’-TGCTGGACCCAACACAAAT-3’, PPIA-rev, 5’-CACATGCTTGCCATCCAA-3’, PPIA (cyclophilin) probe #48, CYP3A7, Hs00426361_m1; TP53I3, Hs00936520_m1; MDM2, Hs1066930:m1; FDXR, Hs01031618_g1. Samples of each experiment were run in triplicate and averaged for final quantification. The fold-changes were calculated as described previously by analysis of relative gene expression data using real-time quantitative PCR and the 2(-∆∆(T)) method [22].
3. Results
3.1. Survival Rates and CAFs Responses to Ablative Radiation Doses
All the data generated in this study come from cell cultures directly prepared from lung tumor specimens obtained from 8 different donors after surgical resection (Figure 1). In a previous study from our group we describe in detail the isolation and characterization of the tumor fibroblast cultures that have been used also for this study [14].
Of note, flow cytometry analyses using the fibroblast specific marker α-SMA reveal a purity above 99.5%, which gives full credibility to the data generated with this material. The intrinsic radioresistance of CAFs in dose escalating trial experiments have also been evaluated and described by us previously [14]. A single dose of 18 Gy happens to be sub-lethal for CAFs, however we have observed that single doses above 12 Gy induce enduring DNA damage responses (nuclear foci of 53BP1) and the cells enter into a permanent cell growth arrest or sensecence. Hence, based on our previous experience we have chosen the use of a single dose of 18 Gy as the basis
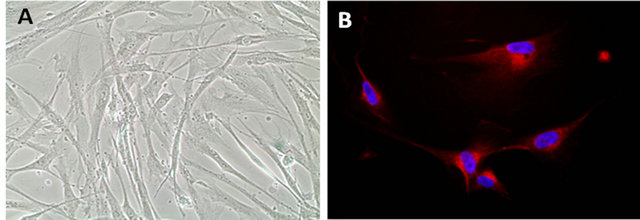
Figure 1. A 200× magnification phase contrast micrograph of three weeks old CAFs in monolayer culture is shown in (A). CAFs are characterized by immunostaining with antiFAP (fibroblast activating protein), a specific marker of reactive fibroblasts (400× magnification) and as shown in (B).
protocol for conducting this study. In previous studies done by others on normal fibroblasts, 24 hours post-irradiation seems to be the time point where the largest number of genes is altered [10]. We have therefore set up 24 hours as the single and most relevant time point to analyse changes in gene expression.
3.2. Radiation-Induced Gene Expression
Large scale molecular responses in CAFs obtained from NSLC patient material to ablative doses of radiation were studied by measuring genome-wide transcript-level changes using human genome survey microarrays as described in Materials and Methods. After pre-processing and normalization, a total of 680 differentially expressed were found in the group of irradiated CAF samples, of which 127 genes were downand 553 were up-regulated transcripts with p < 0.05 (Supplemental list 1). Of all differentially expressed genes, 601 could be annotated by GO terms for biological processes in Kegg (http://www.genome.jp/kegg/) and are summarized in Supplemental list 2. Principal component analysis (PCA) revealed that the major variability in the data set is caused by the difference between irradiated and non-irradiated CAFs (Data not shown). The number and distribution of 153 differentially expressed genes with fold change greater than 1 and an adjusted p-value less than 0.05 across different comparisons is shown in Figure 2.
3.3. Radiation-Induced Pathways after Functional Categorization
Candidate genes which relate to senescence have been extracted from the GenAge database (http://genomics.senescence.info/genes/human.html)
(Table 1). In addition, the expression levels of some differentially expressed genes found in this study have been verified by qRT-PCR (Table 2).
Differentially expressed genes were annotated with PANTHER (http://www.pantherdb.org/) to different biological processes: cell cycle regulation and DNA repair
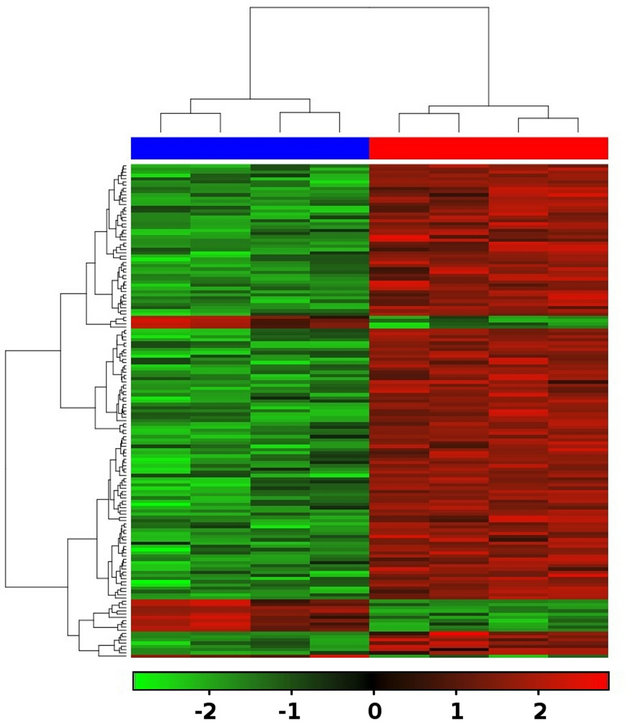
Figure 2. Heatmap of 153 differentially expressed genes in irradiated CAFs with fold change greater than 1.0 and an adjusted p-value less than 0.05 are shown. CAF samples are shown as red bar (top) and non-irradiated CAF samples as blue bar (top). Legend (bottom) shows the fold change [log2 (radiated) - log2 (normal)] color coded values in the heatmap.
(Table 3), oxidative stress (Table 4), apoptosis (Table 5), autophagy (Table 6), and cell communication/cell adhesion (Table 7).
Increased expression levels of interleukin 12 (IL-12), autophagy related protein 16-2 (ATG16L2), growth differentiation factor 15 (GDF15), cytochrome P450, family 3, subfamily A, polypeptide 7 (CYP3A7), NADH adenodoxin osireductase (FDXR), transformed 3t3 cell double minute 2 (MDM2), collagen type 7 alpha 1 (COL7A1) and stromelysin 3 (MMP11), and tumor protein p53 inducible protein 3 (TP53IP3) were confirmed by qRTPCR (Table 2). Figure 3 depicts differential expression of 37 selected genes that are discussed in more detail below.
3.4. Validation of Expression Data by Functional Assays
Gene expression analysis indicated that some p53-dependent pathways and the cell cycle regulation machinery were altered after IR. One end point of such disturbances is the development of stress-induced cellular senescence. In our previous study we have performed quantitative and qualitative measurements of senescence on CAFs after exposure to either high single dose or fractionated
Table 1. Differentially expressed genes related to ageing/senescence in irradiated CAFs.
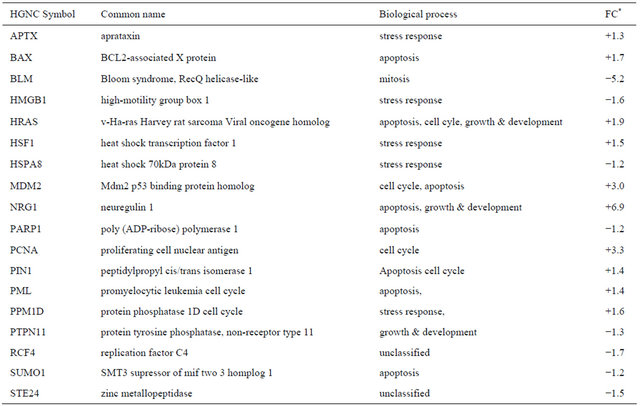

*Values are shown as fold-change representing eight independent microarray experiments.
Table 2. Verification of selected expressed genes in irradiated CAFs by qPCR.

*Values are shown as fold-change (FC) representing eight independent microarray experiments and #RT-PCR validations in triplicate ± S.D. of each representative RNA sample.
Table 3. Differentially expressed genes (p < 0.05) related to cell cycle regulation and DNA repair in irradiated CAFs.


*Values are shown as fold-change (FC) representing eight independent microarray experiments.
Table 4. Differentially expressed genes (p < 0.05) related to oxidative stress in irradiated CAFs.
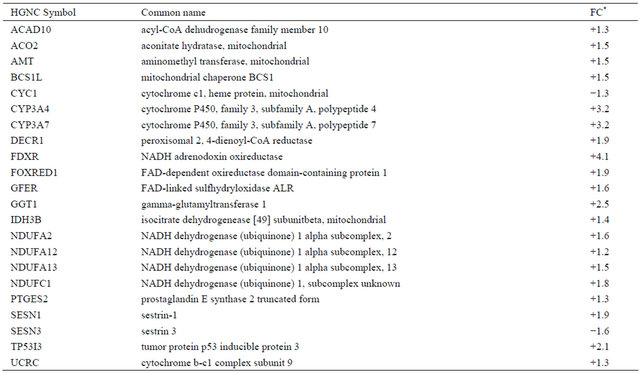

*Values are shown as fold-change (FC) representing eight independent microarray experiments.
Table 5. Differentially expressed genes (p < 0.05) related to apoptosis in irradiated CAFs.
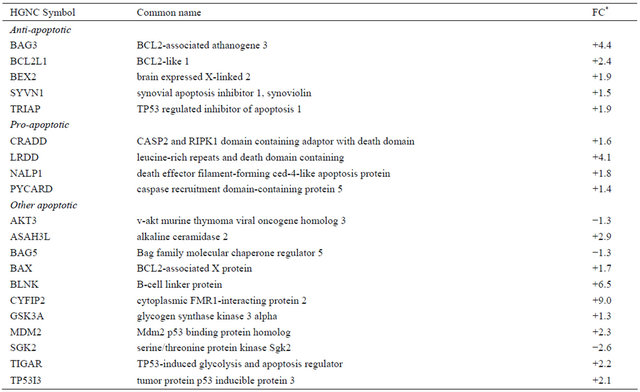

*Values are shown as fold-change (FC) representing eight independent microarray experiments.
Table 6. Differentially expressed genes (p < 0.05) related to autophagy in irradiated CAFs.
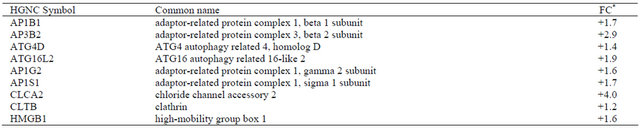

*Values are shown as fold-change (FC) representing eight independent microarray experiments.
Table 7. Differentially expressed genes (p < 0.05) related to cell communication/cell adhesion and other biological processes in irradiated CAFs.
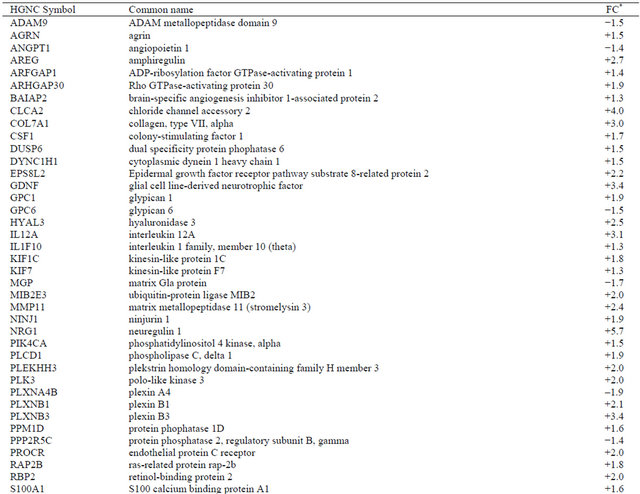

*Values are shown as fold-change (FC) representing eight independent microarray experiments.
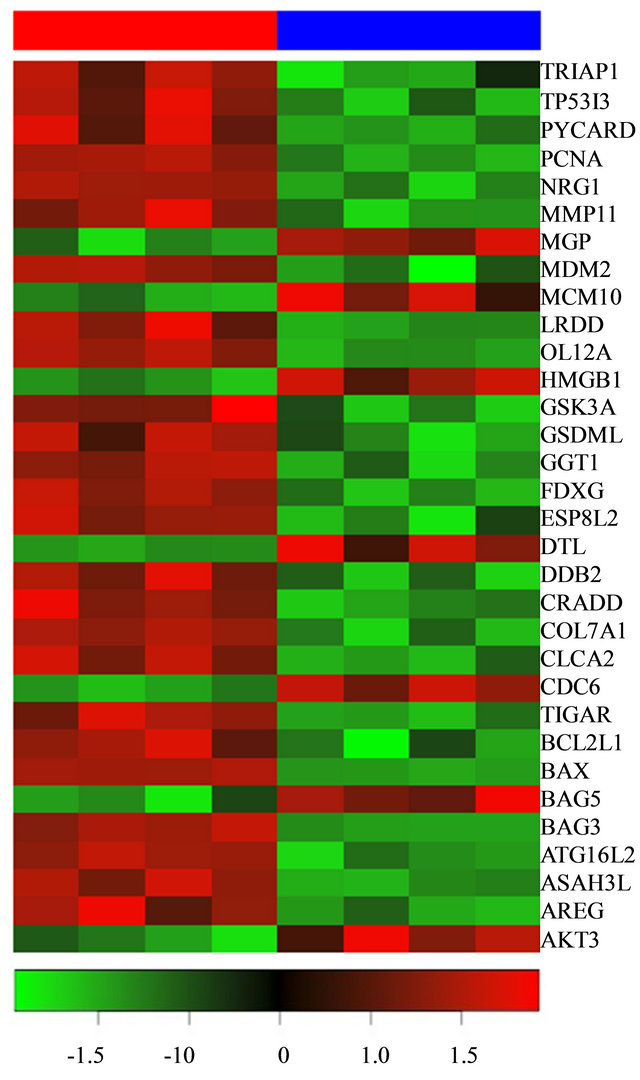
Figure 3. Heatmap of 37 selected genes in irradiated CAF samples are shown as red bar (top) and non-irradiated CAF samples as blue bar (top). Legend (bottom) shows the fold change [log2 (radiated) - log2 (normal)] color coded values in the heatmap.
irradiation [14]. Our data show a potent and permanent induction of cell senescence after a single insult of 18 Gy and Figure 4 illustrates the acquisition of the senescent phenotype by CAFs after exposure to 18 Gy.
One of the most drastically altered pathways after irradiation corresponded to DNA damage responses and chromatin rearrangements (Table 3). Previously we have conducted quantitative dose-dependent measurements of DNA damage and repair induced by IR on CAFs. In that study we show that IR provoke substantial DNA damage even at low radiation doses, however we saw that arising nuclear foci were able to resolve at radiation doses lower than 12 Gy. After a single dose with 18 Gy a strong and permanent DNA damage response on cells was established. An illustration of these observations is presented in Figure 5.
Intracellular levels of reactive oxygen species (ROS) were determined as described in detail in the Materials & Methods. ROS production by CAFs was significantly increased in irradiated CAFs (Figure 6). Elevated ROS production has been found to be predominantly a consequence of altered mitochondrial gene expression (Table 4). Increased expression of cytochrome P450, family 3, subfamily A, polypeptide 7 (CYP3A4) and NADH adrenodoxin oxireductase (FDXR) was confirmed by qPCR (Table 2).
Genes associated with regulation of autophagy and apoptosis were also transformed after IR (Table 5 and Table 6). However, expression of apotopic genes like MDM2 p53 binding protein homolog (MDM2) and tumor protein p53 inducible protein 3 (TP53I3), and expression of ATG16 autophagy related 16-like 2 (ATG16L2) gene was confirmed by qPCR (Table 2).
In pilot experiments, apoptosis and autophagy were measured on CAFs at different points after IR by staining for annexin V on living cells and by using Cyto-ID Autophagy Detection Kit (ENZO life Sciences) respecttively (data not shown). These assays failed to show induction of neither apoptosis nor autophagy in CAFs for up to two weeks after exposure to 18 Gy, indicating that the counter balance between the proand anti-signaling mechanisms prevented the ultimate induction of such pathways.
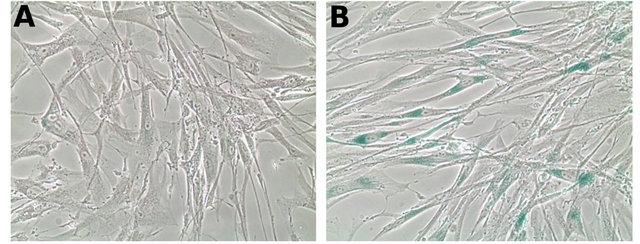
Figure 4. Induction of cellular senescence was determined by β-galactosidase staining assay. Adherent CAFs were irradiated (1 × 18 Gy) and 5 days later examined for the expression of intra-cytoplasmic β-galactosidase. (A) control conditions (0 Gy); (B) irradiated cultures (18 Gy).
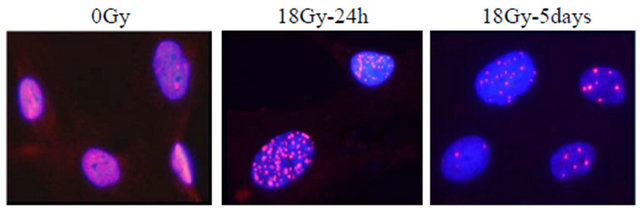
Figure 5. Persistent DDR (DNA Damage Response). Cells were immunostained for 53BP1 (red, Alexa546) and nuclei were stained with DAPI (Blue). Homogenous nuclear staining of 53BP1 was observed in non-irradiated cells, whereas numerous nuclear foci were observed 24 hours post-radiation. After five days most cells still demonstrated multiple nuclear foci, with a reduced number of foci when compared to earlier time points.
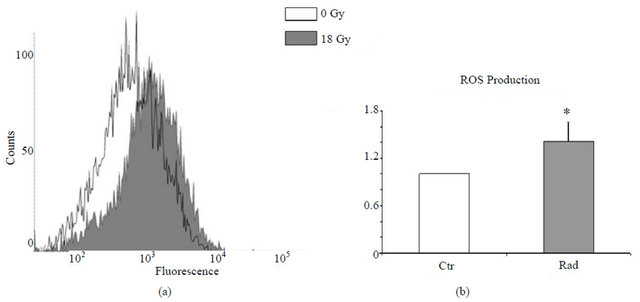
Figure 6. The intracellular ROS production was counted as total H2O2 accumulated in cells 24 hours after IR. H2O2 levels were determined in CAFs from 4 different donors by incubating cells with 2.5 mM CM-H2DCFDA for 30 minutes followed by flow cytometric analyses. Ten thousand events were collected for each measurement and the mean values of the relative fluorescence were registered. Panels in (a) illustrate ROS staining in CAFs from one representative donor. Continuous lines correspond to fluorescence patterns in non-irradiated cells; filled peaks correspond to fluorescence intensity in irradiated cells. In (b), mean fluorescent intensities resulting from FACS analysis from four randomly selected donors is presented. Fluorescence intensity from non-irradiated cultures was averaged and normalized (open bars) whereas the irradiated counterparts are presented with X-fold increase in surface labeling relative to control cells (filled bars). The asterix (*) indicates significant differences (p < 0.05) in mean values when compared to the levels in untreated cells.
In addition, the increased expression of genes involved in cell adhesion, collagen type VII alpha (COL7A1) and stromelysin 3 (MMP11), and the cytokine interleukin 12A (IL2A) has been confirmed accordingly (Table 2).
4. Discussion
The aim of this study was to examine the impact of high-dose ionizing radiation on activated cancer-associated fibroblasts (CAFs) and to predict the consequences of the induced changes on post-radiation tumorigenesis. The overall study showed that approximately 80 % of all differentially expressed genes were up-regulated and 20 % were down-regulated after treatment (Supplemental list 1). In line with previous reports performed on normal tissue fibroblasts, our data show that a large extent of differentially expressed genes belongs to cellular mechanisms related to cell stress, cell cycle, apoptosis, DNA repair, and other pro-survival pathways. It is expected that ionizing radiation induces severe genotoxic stresses in cells, followed by alterations on the expression of genes involved in biological processes like DNA repair, cell proliferation, cell cycle, apoptosis and p53 pathway as has been also shown in a previous study with primary human skin fibroblasts [9]. Cell proliferation and cell cycle checkpoints are some of the pathways profoundly affected by ablative doses of ionizing radiation (AIR) on CAFs. Hence, a list of genes involved in cell cycle regulation such as cell division cycle homologs CDC6 and CDC34, proliferating cell nuclear antigen (PCNA) and different cylins are all up-regulated, whereas denticless protein homolog (DTL) and minichromosome maintenance component 10 (MCM10) are down-regulated (Table 3). Secondly, (RPA2), Mdm2 p53 binding protein homolog (MDM2), and damage-specific DNA binding protein 2 (DDB2), were both up-regulated. In line with our observations, an increased expression of cylins and PCNA has been also observed in normal fibroblasts after lower dose ionizing radiation [11]. In addition, many histones showed increased expression (Table 3). They do not only play an important role in transcription regulation, but are also important team players in DNA repair mechanisms by being responsible for sustaining chromosomal stability [23]. Putting it together, these data suggest that CAFs intend to counteract mitotic catastrophe after irradiation.
In normal fibroblast radiation exposure, as well as administration of genotoxic agents, are able to induce growth arrest, a phenomenon described as cellular senescence [24]. Senescence is often accompanied by stimuli like oxidative DNA damage, and a compromised replicative cell cycle [25]. Although cellular senescence can be ascribed as a self-regulated tumor suppressor mechanism, thereby protecting the organisms from developing cancer, it has been acknowledged that senescent stromal fibroblasts are able to expedite epithelial tumorigenesis [26]. Ionizing radiation of CAFs revealed differential expression of a considerable number of genes that are related to senescence and ageing. Candidate genes listed have been shown to be involved in sensecence at different extents (Table 1).
In our hands, high-dose ionizing radiation results in an increase of reactive oxygen species (ROS) formation in CAFs. The provoked enhanced ROS production is reflected by alterations in mainly mitochondrial-generated ROS rather than activation of the NADPH-oxidase (NOX) system (Figure 3 and Table 4). Of note, the disturbances in the redox homeostasis of irradiated CAFs have the potential to lead to increased tumor growth [27]. Previous studies have shown that normal fibroblasts express multiple ROS scavenger genes when irradiated with lower doses [13]. Here, enhanced expression was only observed for one ROS scavenger, namely gamma glutamyl transferase 1 (GGT1).
The expression of genes involved in the regulation of apoptosis was also modified after AIR, including both proand anti-apoptotic signaling pathways (Table 5). Thus, the transcription of genes involved in caspase-mediated apoptosis (caspase recruitment domain-containing protein (PYCARD) and caspase and RIP adapter with death domain (CRADD) was enhanced, which correlates with observations made in irradiated normal tissue fibroblast [11]. However, in our in vitro assays we could not observe apoptotic cells during the first two weeks post irradiation. Taken these observations together, we may conclude that the anti-apoptotic signals had stronger influence on the final outcomes on cells.
Of importance, a number of genes regulated by or connected to the tumor suppressor gene p53 were shown to be upor down regulated. The observed over-expression of MDM2 can result in excessive inactivation of tumor protein p53 in CAFs, thereby diminishing its tumor suppressor function and therefore affecting tumorigenesis [28] or may act in a p53-independent manner as an oncogene [29]. It is hereby noted that nine different MDM2 transcripts showed increased expression levels (see also Supplemental list 1). Their functional and not yet defined biological implications on the tumor microenvironment still needs to be elucidated [30]. Increased expression levels for MDM2 variants have been reported to be related to radio-sensitivity in the lung [31], and implies the possibility that CAF to some extent become radio-resistant after treatment [32]. On the other hand, up-regulation of the TP53-induced glycolysis and apoptosis regulator (TIGAR) may protect irradiated CAFs against reactive oxigen species and apoptosis induced by TP53/p53. It has been shown that fibroblasts are also able to induce an up-regulation of TIGAR expression in cancer cells, thereby protecting cancer cells against apoptosis and autophagy, as recently described in an autophagic tumor stroma model of cancer [33].
Autophagy, which can be ascribed as both cell survival and cell death mechanism has been shown to be activated in tumor cells after ionizing radiation [34]. Here, enhanced expression of a large number of other genes related to autophagy has been observed (Table 6). A role for autophagy in CAFs has been recently shown to be linked to promotion of tumor cell survival [33]. However, the establishment of an autophagic response in CAFs after AIR still needs to be confirmed, and whether such stress-induced changes occurring in CAFs could have an impact on tumor cells also remains to be determined.
Pro-angiogenic signals released by CAFs contribute importantly to the overall sustainability of tumors [35,36]. However, the observed decreased expression of the proangiogenic factor angiopoietin 1 ANGPT1, a factor that mediates reciprocal interactions between the endothelium and surrounding matrix [37], might result in suppressed tumor growth. Several anti-angiogenic molecules showed increased expression in CAFs after highdose ionizing radiation which might have a direct impact on tumor growth. The matrix-derived angiogenesis inhibitor thrombospondin type I, domain containing 1 (THSD 1) [38], which is able to confer signals like thrombospondin 1 [39-41], and interleukin 12A (IL-12A) [42], both may be able to reduce tumor growth rate. In contrast, several semaphorin receptors which are able to confer both, proand anti-angiogenic signals are differentially expressed in irradiated CAFs (Table 7). The observed differential expression of plexins PLXNB1, PLXNB3 and PLXA4B imply a role for CAFs on endothelial cell migration [43]. Their role in normal lung and in invasive growth and cell migration in lung cancer has been discussed [44,46]. Interestingly, the activation of PLXNB1 and PLXNB3 in irradiated CAFs might inhibit integrin-based adhesion and cell migration on the ECM as has been reported earlier for normal fibroblasts [47].
Together with immune cells, fibroblasts are able to support the survival of the tumor [48,49]. Although many genes involved in signaling pathways for growth factors (i.e. EGF/EGFR) have been identified in CAFs [48-51], a differential expression of these particular growth factors or receptors has not been observed by us. Interestingly, two members of the epidermal growth factor (EGF) family downstream of the EGF receptor pathway have been found with increased expression in irradiated CAFs: the autocrine growth factor and mitogen for fibroblasts amphiregulin (AREG), and epidermal growth factor recaptor pathway substrate 8-related protein 2 (EPS8L2). Increased expression of AREG might have an effect on tumor growth since it has been shown that AREG can inhibit growth and apoptosis in certain aggressive adenoma cell lines in culture [52]. However, increased expression of EPS8 has been linked to progression of squamous carcinogenesis and might confer pro-tumorigenic effects on the surrounding tumor tissue [53]. It is therefore tempting to speculate that alternative signaling mechanisms may deregulate EGFR signaling during carcinogenesis, such as over-expression or activation of EGF/EGFR pathway intermediates.
CAFs express extracellular matrix (ECM) proteins and matrix regulatory agents, thereby providing a favourable microenvironment for cancer cells which is followed by a stimulation of proliferation of tumors and metastasis [54-56]. The increased expression of collagen type VII, alpha COL7A1 might sustain the stability of the cellular organization within the tumor microenvironment by contributing to epithelial basement membrane organization and adherence by interacting with ECM proteins. On the other hand, the matrix metalloproteinase 11 (stromelysin 3) (MMP11) is the solely significant differentially exessed matrix metalloproteinase after irradiation. The enhanced expression of MMP11 in CAFs could promote cancer progression by remodeling the ECM and confers anti-apoptotic and anti-necrotic effects on tumor cells [57]. Numerous studies have linked increased MMP11 expression in different human cancers with poor disease prognosis [58,59], and have been recently suggested to represent a new prognostic indicator for gastric cancer [60]. In addition, the ECM protein Gla protein (MGP) showed decreased expression which can be associated with poor prognosis of lung cancer as has been recently reported for colorectal adearcinoma [61]. A potential role for MGP as a prognostic marker for breast cancer has also been implied [62]. Increased expression levels of multiple splice variants for neuregulin-1 (NRG1) in CAFs (see also Supplemental list 1) might have an impact on neighboring epithelial cells by inducing their growth and differentiation [63,64]. Elevated expression of gasdermin B (GSDMB) imply a regulatory role for irradiated CAFs on the adjacent epithelial cells, as augnted expression of GSDML has been correlated to cancer progression in the gastric epithelium [65].
In conclusion, CAFs are able to survive ablative radiation doses, but such insult provoke prominent cellular damages and consequently numerous genes implicated in DNA repair, cellular stress and cell survival pathways are activated. Our analyses show that CAFs display both proand anti-tumorigenic effects after having received a single high-dose of ionizing radiation. These effects might have an impact on the tumor microenvironment in respect to tumor growth and metastasis. The complexity of the observed differential gene expression patterns and implicative biological effects are interesting future objecttives for further elucidations.
5. Acknowledgements
Funding was provided by the North Norway Regional Health Authority (Helse Nord) and Aakre Foundation. Microarray experiments were conducted at the Microarray Resource Centre Tromsø (MRCT), which is supported by the National Program for Research and Functional Genomics in Norway (FUGE), University of Tromsø and University Hospital North Norway. The authors have no potential conflicts of interest to disclose.
REFERENCES
- J. H. Heinzerling, B. Kavanagh and R. D. Timmerman, “Stereotactic Ablative Radiation Therapy for Primary Lung Tumors,” Cancer Journal, Vol. 17, No. 1, 2011, pp. 28-32. doi:10.1097%2FPPO.0b013e31820a7f80
- D. Palma and S. Senan, “Stereotactic Radiation Therapy: Changing Treatment Paradigms for Stage I Non Small Cell Lung Cancer,” Current Opinion in Oncology, Vol. 23, No. 2, 2011, pp. 133-139. doi:10.1097%2FCCO.0b013e328341ee11
- B. D. Kavanagh, M. Miften and R. A. Rabinovitch, “Advances in Treatment Techniques: Stereotactic Body Radiation Therapy and the Spread of Hypofractionation,” Cancer Journal, Vol. 17, No. 3, 2011, pp. 177-181. doi:10.1097%2FPPO.0b013e31821f7dbd
- T. B. Lanni Jr., I. S. Grills, L. L. Kestin and J. M. Robertson, “Stereotactic Radiotherapy Reduces Treatment Cost while Improving Overall Survival and Local Control over Standard Fractionated Radiation Therapy for Medically Inoperable Non-Small-Cell Lung Cancer,” American Journal of Clinical Oncology, Vol. 34, No. 5, 2011, pp. 494-498. doi:10.1097%2FCOC.0b013e3181ec63ae
- M. P. Lisanti, U. E. Martinez-Outschoorn, B. Chiavarina, S. Pavlides, D. Whitaker-Menezes, A. Tsirigos, A. Witkiewicz, Z. Lin, R. Balliet, A. Howell and F. Sotgia, “Understanding the ‘Lethal’ Drivers of Tumor-Stroma Co-Evolution: Emerging Role(s) for Hypoxia, Oxidative Stress and Autophagy/Mitophagy in the Tumor MicroEnvironment,” Cancer Biology and Therapy, Vol. 10, No. 6, 2010, pp. 537-542. doi:10.4161%2Fcbt.10.6.13370
- K. Pietras and A. Ostman, “Hallmarks of Cancer: Interactions with the Tumor Stroma,” Experimental Cell Research, Vol. 316, No. 8, 2010, pp. 1324-1331. doi:10.1016%2Fj.yexcr.2010.02.045
- M. Allen and J. L. Jones, “Jekyll and Hyde: The Role of the Microenvironment on the Progression of Cancer,” The Journal of Pathology, Vol. 223, No. 2, 2011, pp. 162-176.
- K. Räsänen and A. Vaheri, “Activation of Fibroblasts in Cancer Stroma,” Experimental Cell Research, Vol. 316, No.17, 2010, pp. 2713-2722. doi:10.1016%2Fj.yexcr.2010.04.032
- E. Kis, T. Szatmari, M. Keszei, R. Farkas, O. Esik, K. Lumniczky, A. Falus and G. Safrany, “Microarray Analysis of Radiation Response Genes in Primary Human Fibroblasts,” International Journal of Radiation Oncology, Biology, Physics, Vol. 66, No. 5, 2006, pp. 1506-1514. doi:10.1016%2Fj.ijrobp.2006.08.004
- S. Tachiiri, T. Katagiri, T. Tsunoda, N. Oya, M. Hiraoka and Y. Nakamura, “Analysis of Gene-Expression Profiles after Gamma Irradiation of Normal Human Fibroblasts,” International Journal of Radiation Oncology, Biology, Physics, Vol. 64, No. 1, 2006, pp. 272-279. doi:org/10.1016%2Fj.ijrobp.2005.08.030
- O. K. Rødningen, J. Øvergaard, J. Alsner, T. Hastie and A. L. Børresen-Dale, “Microarray Analysis of the Transcriptional Response to Single or Multiple Doses of Ionizing Radiation in Human Subcutaneous Fibroblasts,” Radiotherapy and Oncology, Vol. 77, No. 3, 2005, pp. 231-240. doi:10.1186%2Fbcr1151
- H. Landmark, S. A. Nahas, J. Aaroe, R. Gatti, A. L. Borresen-Dale and O. K. Rodningen, “Transcriptional Response to Ionizing Radiation in Human Radiation Sensitive Cell Lines,” Radiotherapy and Oncology, Vol. 83, No. 3, 2007, pp. 256-260.
- O. K. Rødningen, A. L. Børresen-Dale, J. Alsner, T. Hastie and J. øvergaard, “Radiation-Induced Gene Expression in Human Subcutaneous Fibroblasts is Predictive of Radiation-Induced Fibrosis,” Radiotherapy and Oncology, Vol. 86, No. 3, 2008, pp. 314-320. doi:10.1016%2Fj.radonc.2007.09.013
- T. Hellevik, I. Pettersen, V. Berg, J. O. Winberg, B. T. Moe, K. Bartnes, R. H. Paulssen, L. T. Busund, R. Bremnes, A. Chalmers and I. Martinez-Zubiaurre, “CancerAssociated Fibroblasts from Human NSCLC Survive Ablative Doses of Radiation but Their Invasive Capacity is Reduced,” Radiation Oncology, Vol. 7, No. 1, 2012, pp. 59-71. doi:10.1186%2F1748-717X-7-59
- S. Stathakis, J. S. Li, K. Paskalev, J. Yang, L. Wang and C. M. Ma, “Ultra-Thin TLDs for Skin Dose Determination in High Energy Photon Beams,” Physics in Medicine and Biology, Vol. 51, No. 14, 2006, pp. 3549-3567. doi:10.1088%2F0031-9155%2F51%2F14%2F018
- T. Mosmann, “Rapid Colorimetric Assay for Cellular Growth and Survival: Application to Proliferation and Cytotoxicity Assays,” Journal of Immunological Methods, Vol. 65, No. 1-2, 1983, pp. 55-63. doi:10.1016%2F0022-1759%2883%2990303-4
- R. H. Paulssen, L. Olsen and T. C. Sogn, “MagNa Pure Compact RNA Isolation Kit: Isolation of High-Quality Total RNA from a Broad Range of Sample Material,” Biochemica, Vol. 2, 2006, pp. 14-16.
- B. M. Bolstad, R. A. Irizarry, M. Astrand and T. P. Speed, “A Comparison of Normalization Methods for High Density Oligonucleotide Array Data Based on Variance and Bias,” Bioinformatics, Vol. 19, No. 2, 2003, pp. 185-193. doi:10.1093%2Fbioinformatics%2F19.2.185
- G. K. Smyth, “Linear Models and Empirical Bayes Methods for Assessing Differential Expression in Microarray Experiments,” Statistical Applications in Genetics and Molecular Biology, Vol. 3, No. 1, 2004, pp. 1544-6115. doi:10.2202%2F1544-6115.1027
- Y. Benjamini and D. Yekutieli, “False Discovery RateAdjusted Multiple Confidence Intervals for Selected Parameters,” Journal of the American Statistical Association, Vol. 100, No. 469, 2005, pp. 71-81. doi:10.1198%2F016214504000001907
- L. Gidskehaug, H. Stodkilde-Jorgensen, M. Martens and H. Martens, “Bridge-PLS Regression: Two-Block Bilinear Regression without Deflation,” Journal of Chemometrics, Vol. 18, No. 3-4, 2004, pp. 208-215. doi:10.1002%2Fcem.862
- K. J. Livak and T. D. Schmittgen, “Analysis of Relative Gene Expression Data using Real-Time Quantitative PCR and The 2(-Delta Delta C(T)) Method,” Methods, Vol. 25, No. 4, 2001, pp. 402-408.
- G. Li and D. Reinberg, “Chromatin Higher-Order Structures and Gene Regulation,” Current Opinion in Genetics & Development, Vol. 21, No. 2, 2011, pp. 175-186. doi:10.1016%2Fj.gde.2011.01.022
- A. Krtolica, S. Parrinello, S. Lockett, P. Y. Desprez and J. Campisi, “Senescent Fibroblasts Promote Epithelial Cell Growth and Tumorigenesis: A Link Between Cancer and Aging,” Proceedings of the National Academy of Sciences of the United States of America, Vol. 98, No. 21, 2001, pp. 12072-12077. doi:10.1073%2Fpnas.211053698
- J. L. Wang and P. C. Wang, “The Effect of Aging on the DNA Damage and Repair Capacity in 2BS Cells Undergoing Oxidative Stress,” Molecular Biology Reports, Vol. 39, No. 1, 2012, pp. 233-241. doi:10.1007%2Fs11033-011-0731-4
- C. Bavik, I. Coleman, J. P. Dean, B. Knudsen, S. Plymate and P. S. Nelson, “The Gene Expression Program of Prostate Fibroblast Senescence Modulates Neoplastic Epithelial Cell Proliferation through Paracrine Mechanisms,” Cancer Research, Vol. 66, No. 2, 2006, pp. 794-802. doi:10.1158%2F0008-5472.CAN-05-1716
- T. B. Kryston, A. B. Georgiev, P. Pissis and A. G. Georgakilas, “Role of Oxidative Stress and DNA Damage in Human Carcinogenesis,” Mutation Research, Vol. 711, No. 1-2, 2011, pp. 193-201. doi:10.1016%2Fj.mrfmmm.2010.12.016
- P. L. Miliani de Marval and Y. Zhang, “The RP-Mdm2- p53 Pathway and Tumorigenesis,” Oncotarget, Vol. 2, No. 3, 2011, pp. 234-238.
- J. J. Manfredi, “The Mdm2-p53 Relationship Evolves: Mdm2 Swings Both Ways as an Oncogene and a Tumor Suppressor,” Genes & Development, Vol. 24, No. 15, 2010, pp. 1580-1589. doi:10.1101%2Fgad.1941710
- L. C. Harris, “MDM2 Splice Variants and Their Therapeutic Implications,” Current Cancer Drug Targets, Vol. 5, No. 1, 2005, pp. 21-26. doi:10.2174%2F1568009053332654
- K. Ogawa, S. Murayama and M. Mori, “Predicting the Tumor Response to Radiotherapy Using Microarray Analysis,” Oncology Reports, Vol. 18, No. 5, 2007, pp. 1243- 1248.
- W. F. Guo, R. X. Lin, J. Huang, Z. Zhou, J. Yang, G. Z. Guo and S. Q. Wang, “Identification of Differentially Expressed Genes Contributing to Radioresistance in Lung Cancer Cells Using Microarray Analysis,” Radiation Research, Vol. 164, No. 1, 2005, pp. 27-35. doi:10.1667%2FRR3401
- U. E. Martinez-Outschoorn, S. Pavlides, A. Howell, R. G. Pestell, H. B. Tanowitz, F. Sotgia and M. P. Lisanti, “Stromal-Epithelial Metabolic Coupling in Cancer: Integrating Autophagy and Metabolism in the Tumor Microenvironment,” The International Journal of Biochemistry & Cell Biology, Vol. 43, No. 7, 2011, pp. 1045-1051. doi:10.1016%2Fj.biocel.2011.01.023
- H. Chaachouay, P. Ohneseit, M. Toulany, R. Kehlbach, G. Multhoff and H. P. Rodemann, “Autophagy Contributes to Resistance of Tumor Cells to Ionizing Radiation,” Radiotherapy and Oncology, Vol. 99, No. 3, 2011, pp. 287- 292. doi:10.1016%2Fj.radonc.2011.06.002
- M. Ao, O. E. Franco, D. Park, D. Raman, K. Williams and S. W. Hayward, “Cross-Talk between ParacrineActing Cytokine and Chemokine Pathways Promotes Malignancy in Benign Human Prostatic Epithelium,” Cancer Research, Vol. 67, No. 9, 2007, pp. 4244-4253. doi:10.1158%2F0008-5472.CAN-06-3946
- R. F. Hwang, T. Moore, T. Arumugam, V. Ramachandran, K. D. Amos, A. Rivera, B. Ji, D. B. Evans and C. D. Logsdon, “Cancer-Associated Stromal Fibroblasts Promote Pancreatic Tumor Progression,” Cancer Research, Vol. 68, No. 3, 2008, pp. 918-926. doi:10.1158%2F0008-5472.CAN-07-5714
- A. D. Blann, K. S. Ramcharan, P. S. Stonelake, D. Luesley and G. Y. Lip, “The Angiome: A New Concept in Cancer Biology,” Journal of Clinical Pathology, Vol. 64, No. 7, 2011, pp. 637-643. doi:10.1136%2Fjcp.2011.088948
- J. M. Ko, P. L. Chan, W. L. Yau, H. K. Chan, K. C. Chan, Z. Y. Yu, F. M. Kwong, L. D. Miller, E. T. Liu, L. C. Yang, P. H. Lo, E. J. Stanbridge, J. C. Tang, G. Srivastava, S. W. Tsao, S. Law and M. L. Lung, “Monochromosome Transfer and Microarray Analysis Identify a Critical Tumor-Suppressive Region Mapping to Chromosome 13q14 and THSD1 in Esophageal Carcinoma,” Molecular Cancer Research, Vol. 6, No. 4, 2008, pp. 592-603. doi:10.1158%2F1541-7786.MCR-07-0154
- L. C. Armstrong and P. Bornstein, “Thrombospondins 1 and 2 Function as Inhibitors of Angiogenesis,” Matrix Biology, Vol. 22, No. 1, 2003, pp. 63-71. doi:10.1016%2FS0945-053X%2803%2900005-2
- P. Bornstein, “Thrombospondins Function as Regulators of Angiogenesis,” Journal of Cell Communication and Signaling, Vol. 3, No. 3-4, 2009, pp. 189-200. doi:10.1007%2Fs12079-009-0060-8
- F. de Fraipont, A. C. Nicholson, J. J. Feige and E. G. Van Meir, “Thrombospondins and Tumor Angiogenesis,” Trends in Molecular Medicine, Vol. 7, No. 9, 2001, pp. 401-407. doi:10.1016%2FS1471-4914%2801%2902102-5
- F. Cavallo, E. Quaglino, L. Cifaldi, E. Di Carlo, A. Andre, P. Bernabei, P. Musiani, G. Forni and R. A. Calogero, “Interleukin 12-Activated Lymphocytes Influence Tumor Genetic Programs,” Cancer Research, Vol. 61, No. 8, 2001, pp. 3518-3523.
- A. Sakurai, C. Doci and J. S. Gutkind, “Semaphorin Signaling in Angiogenesis, Lymphangiogenesis and Cancer,” Cell Research, Vol. 22, No. 1, 2012, pp. 23-32. doi:10.1038%2Fcr.2012.21
- T. Ito, M. Kagoshima, Y. Sasaki, C. Li, N. Udaka, T. Kitsukawa, H. Fujisawa, M. Taniguchi, T. Yagi, H. Kitamura and Y. Goshima, “Repulsive Axon Guidance Molecule Sema3A Inhibits Branching Morphogenesis of Fetal Mouse Lung,” Mechanisms of Development, Vol. 97, No. 1-2, 2000, pp. 35-45. doi:10.1016%2FS0925-4773%2800%2900401-9
- M. Kagoshima and T. Ito, “Diverse Gene Expression and Function of Semaphorins in Developing Lung: Positive and Negative Regulatory Roles of Semaphorins in Lung Branching Morphogenesis,” Genes to Cells: Devoted to Molecular and Cellular Mechanisms, Vol. 6, No. 6, 2001, pp. 559-571. doi:10.1046%2Fj.1365-2443.2001.00441.x
- V. A. Potiron, J. Roche and H. A. Drabkin, “Semaphorins and Their Receptors in Lung Cancer,” Cancer Letters, Vol. 273, No. 1, 2009, pp. 1-14. doi:10.1016%2Fj.canlet.2008.05.032
- D. Barberis, S. Artigiani, A. Casazza, S. Corso, S. Giordano, C. A. Love, E. Y. Jones, P. M. Comoglio and L. Tamagnone, “Plexin Signaling Hampers Integrin-Based Adhesion, Leading to Rho-Kinase Independent Cell Rounding, and Inhibiting Lamellipodia Extension and Cell Motility,” FASEB Journal, Vol. 18, No. 3, 2004, pp. 592-594.
- N. A. Bhowmick, E. G. Neilson and H. L. Moses, “Stromal Fibroblasts in Cancer Initiation and Progression,” Nature, Vol. 432, No. 7015, 2004, pp. 332-337. doi:10.1038%2Fnature03096
- M. Akdis, S. Burgler, R. Crameri, T. Eiwegger, H. Fujita, E. Gomez, S. Klunker, N. Meyer, L. O’Mahony, O. Palomares, C. Rhyner, N. Ouaked, A. Schaffartzik, W. Van De Veen, S. Zeller, M. Zimmermann and C. A. Akdis, “Interleukins, from 1 to 37, and Interferon-Gamma: Receptors, Functions, and Roles in Diseases,” The Journal of Allergy and Clinical Immunology, Vol. 127, No. 3, 2011, pp. 701-721. doi:10.1016%2Fj.jaci.2010.11.050
- R. Kalluri and M. Zeisberg, “Fibroblasts in Cancer,” Nature Reviews. Cancer, Vol. 6, No. 5, 2006, pp. 392-401. doi:10.1038%2Fnrc1877
- H. Nakagawa, S. Liyanarachchi, R. V. Davuluri, H. Auer, E. W. Martin Jr., A. de la Chapelle and W. L. Frankel, “Role of Cancer-Associated Stromal Fibroblasts in Metastatic Colon Cancer to the Liver and Their Expression Profiles,” Oncogene, Vol. 23, No. 44, 2004, pp. 7366- 7377. doi:10.1038%2Fsj.onc.1208013
- A. Hurbin, L. Dubrez, J. L. Coll and M. C. Favrot, “Inhibition of Apoptosis by Amphiregulin via an Insulin-Like Growth Factor-1 Receptor-Dependent Pathway in NonSmall Cell Lung Cancer Cell Lines,” The Journal of Biological Chemistry, Vol. 277, No. 51, 2002, pp. 49127- 49133. doi:10.1074%2Fjbc.M207584200
- H. Wang, V. Patel, H. Miyazaki, J. S. Gutkind and W. A. Yeudall, “Role for EPS8 in Squamous Carcinogenesis,” Carcinogenesis, Vol. 30, No. 1, 2009, pp. 165-174. doi:10.1016%2Fj.oos.2009.06.212
- J. A. Joyce and J. W. Pollard, “Microenvironmental Regulation of Metastasis,” Nature Reviews. Cancer, Vol. 9, No. 4, 2009, pp. 239-252.
- M. Liu, J. Xu and H. Deng, “Tangled Fibroblasts in Tumor-Stroma Interactions,” International Journal of Cancer, Vol. 129, No. 8, 2011, pp. 1795-1805. doi:10.1002%2Fijc.26116
- L. E. Littlepage, M. D. Sternlicht, N. Rougier, J. Phillips, E. Gallo, Y. Yu, K. Williams, A. Brenot, J. I. Gordon and Z. Werb, “Matrix Metalloproteinases Contribute Distinct Roles in Neuroendocrine Prostate Carcinogenesis, Metastasis, and Angiogenesis Progression,” Cancer Research, Vol. 70, No. 6, 2010, pp. 2224-2234. doi:10.1158%2F0008-5472.CAN-09-3515
- P. Basset, C. Wolf and P. Chambon, “Expression of the Stromelysin-3 Gene in Fibroblastic Cells of Invasive Carcinomas of the Breast and Other Human Tissues: A Review,” Breast Cancer Research and Treatment, Vol. 24, No. 3, 1993, pp. 185-193. doi:10.1007%2FBF01833259
- D. Peruzzi, F. Mori, A. Conforti, D. Lazzaro, E. De Rinaldis, G. Ciliberto, N. La Monica and L. Aurisicchio, “MMP11: A Novel Target Antigen for Cancer Immunoerapy,” Clinical Cancer Research , Vol. 15, No. 12, 2009, pp. 4104-4113. doi:10.1158%2F1078-0432.CCR-08-3226
- C. W. Cheng, J. C. Yu, H. W. Wang, C. S. Huang, J. C. Shieh, Y. P. Fu, C. W. Chang, P. E. Wu and C. Y. Shen, “The Clinical Implications of MMP-11 and CK-20 Expression in Human Breast Cancer,” Clinica Chimica Acta, Vol. 411, No. 3-4, 2010, pp. 234-241. doi:10.1016%2Fj.cca.2009.11.009
- [61] Z. S. Zhao, Y. Q. Chu, Z. Y. Ye, Y. Y. Wang and H. Q. Tao, “Overexpression of Matrix Metalloproteinase 11 in Human Gastric Carcinoma and Its Clinicopathologic Significance,” Human Pathology, Vol. 41, No. 5, 2010, pp. 686-696. doi:10.1016%2Fj.humpath.2009.10.010
- [62] C. Fan, D. Sheu, H. Fan, K. Hsu, C. Allen Chang and E. Chan, “Down-Regulation of Matrix Gla Protein Messenger RNA in Human Colorectal Adenocarcinomas,” Cancer Letters, Vol. 165, No. 1, 2001, pp. 63-69. doi:10.1016%2FS0304-3835%2801%2900416-5
- [63] K. Yoshimura, K. Takeuchi, K. Nagasaki, S. Ogishima, H. Tanaka, T. Iwase, F. Akiyama, Y. Kuroda and Y. Miki, “Prognostic Value of Matrix Gla Protein in Breast Cancer,” Molecular Medicine Reports, Vol. 2, No. 4, 2009, pp. 549-553. doi:10.3892%2Fmmr_00000135
- [64] J. C. Montero, R. Rodriguez-Barrueco, A. Ocana, E. DiazRodriguez, A. Esparis-Ogando and A. Pandiella, “Neuregulins and Cancer,” Clinical Cancer Research, Vol. 14, No. 11, 2008, pp. 3237-3241. doi:10.1158%2F1078-0432.CCR-07-5133
- [65] N. V. Hayes and W. J. Gullick, “The Neuregulin Family of Genes and Their Multiple Splice Variants in Breast Cancer,” Journal of Mammary Gland Biology and Neoplasia, Vol. 13, No. 2, 2008, pp. 205-214. doi:10.1007%2Fs10911-008-9078-4
- [66] H. Komiyama, A. Aoki, S. Tanaka, H. Maekawa, Y. Kato, R. Wada, T. Maekawa, M. Tamura and T. Shiroishi, “Alu-Derived Cis-Element Regulates TumorigenesisDependent Gastric Expression of GASDERMIN B (GSDMB),” Genes & Genetic Systems, Vol. 85, No. 1, 2010, pp. 75-83. doi:10.1266%2Fggs.85.75
Supplement
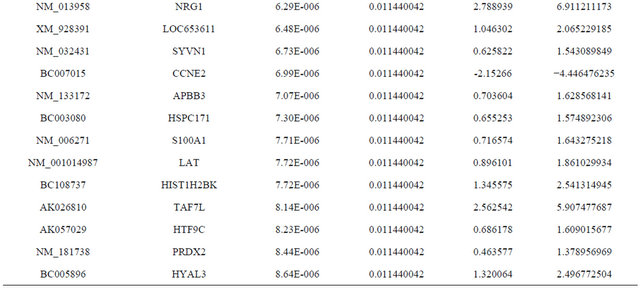
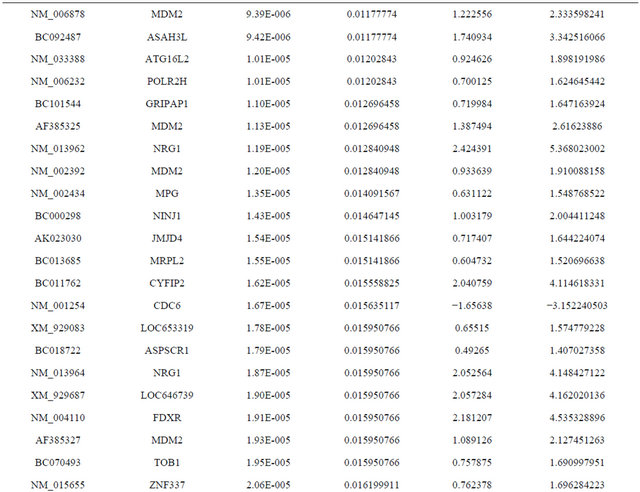
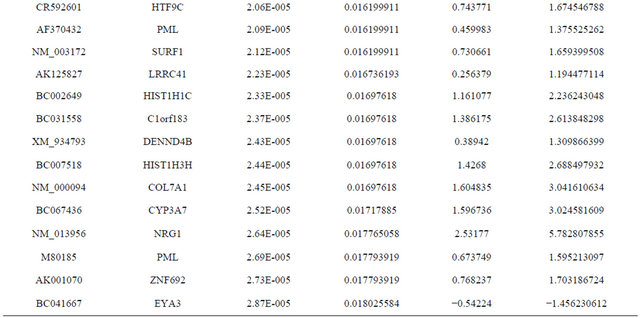
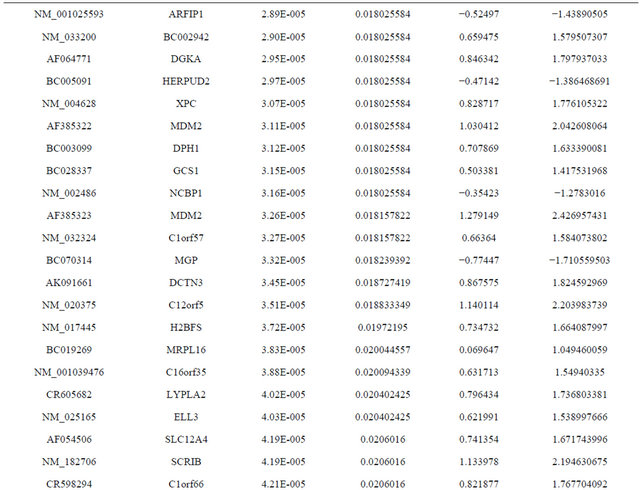
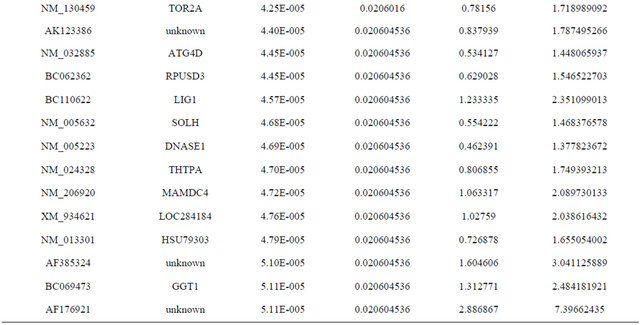
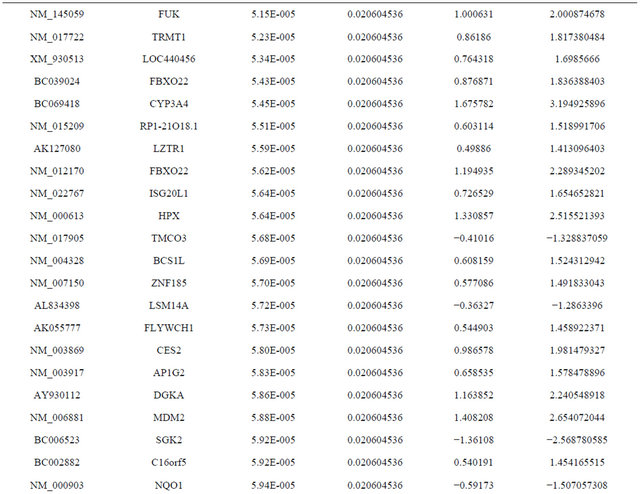
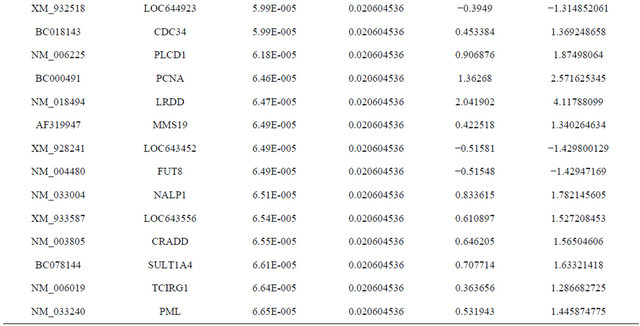
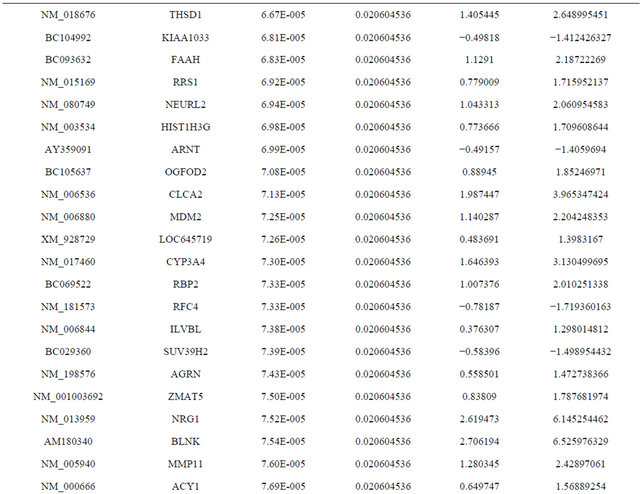
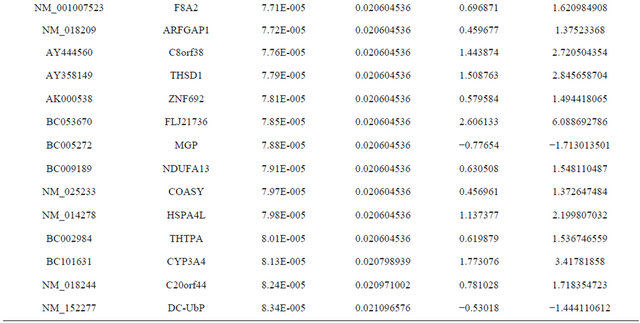
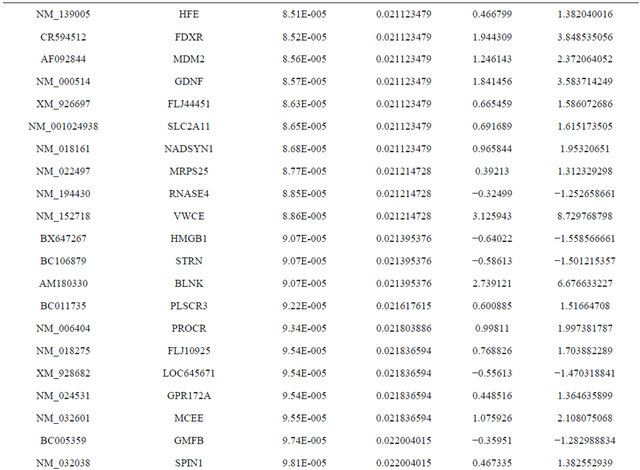
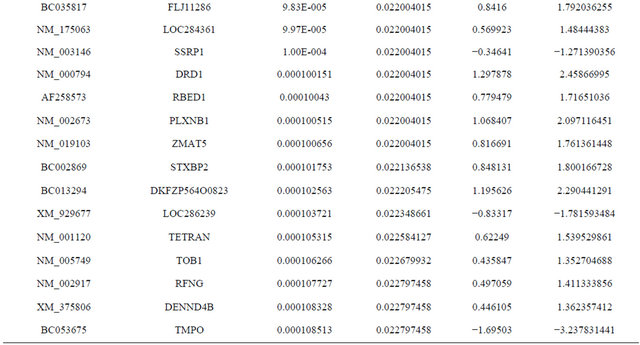
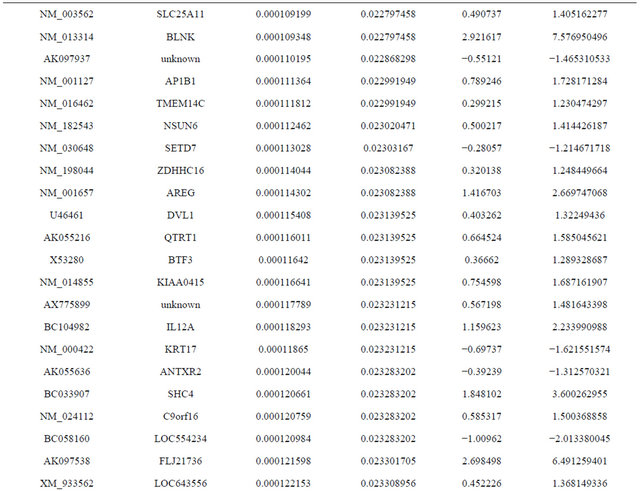
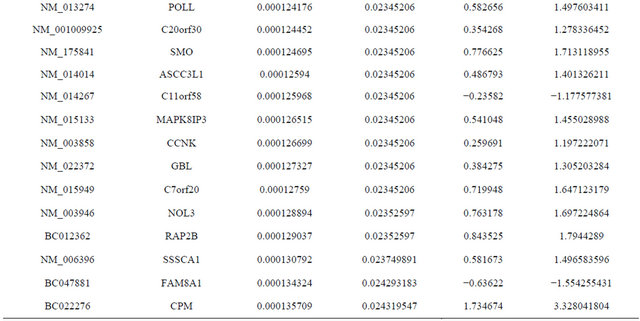
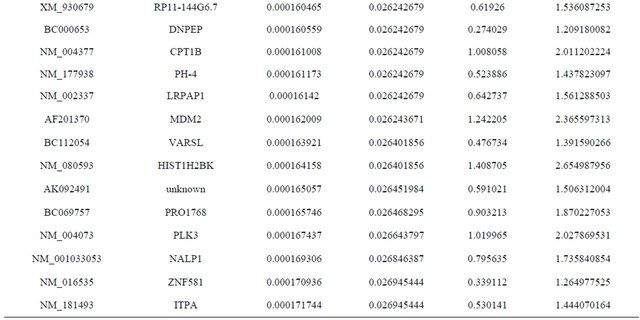
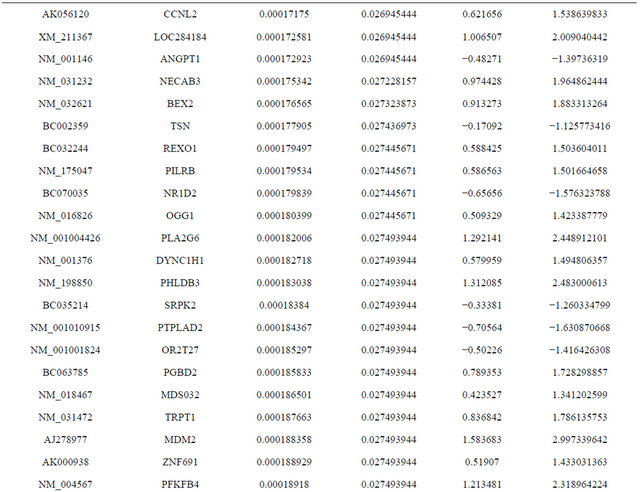
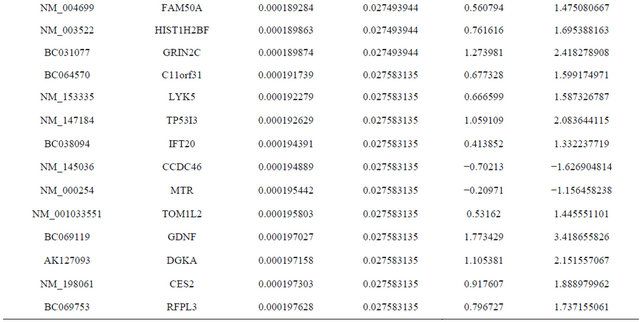
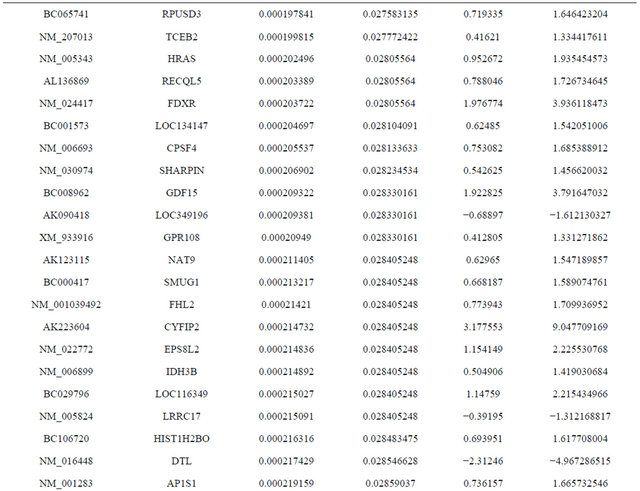
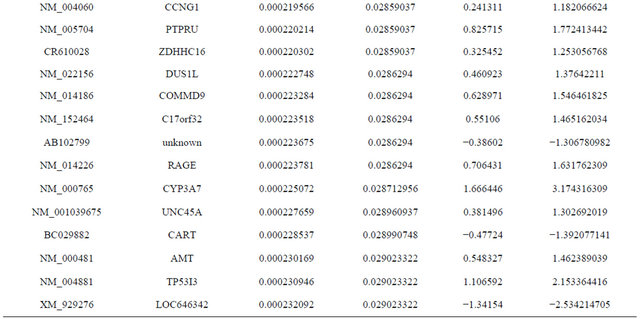
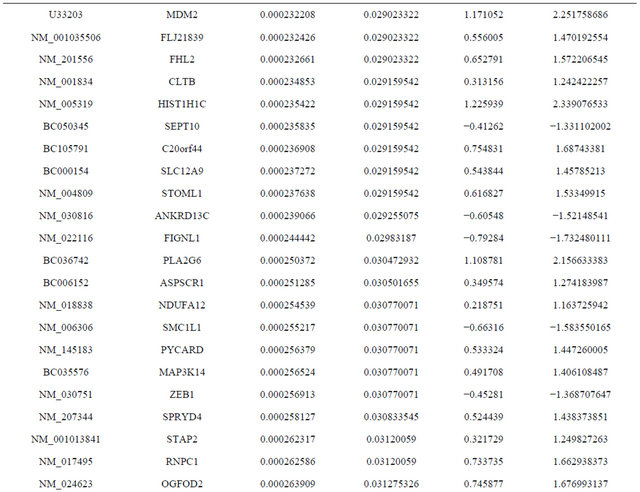
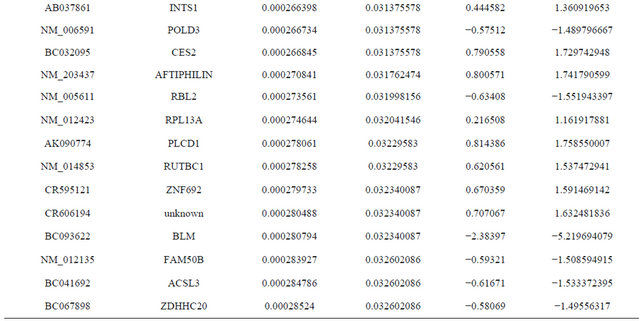
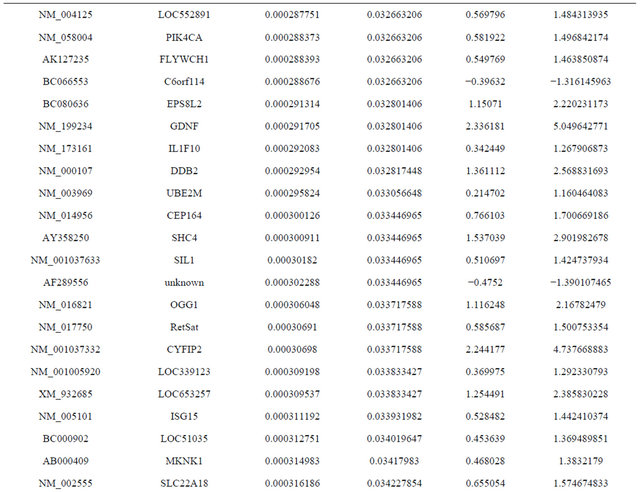
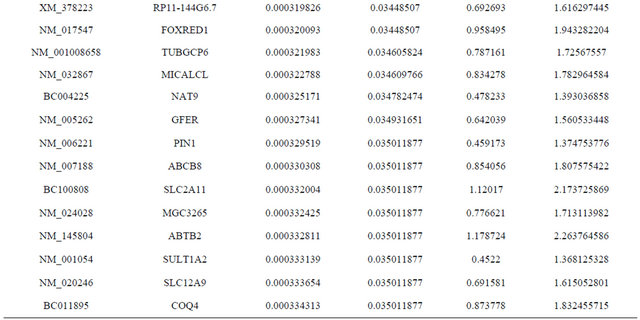
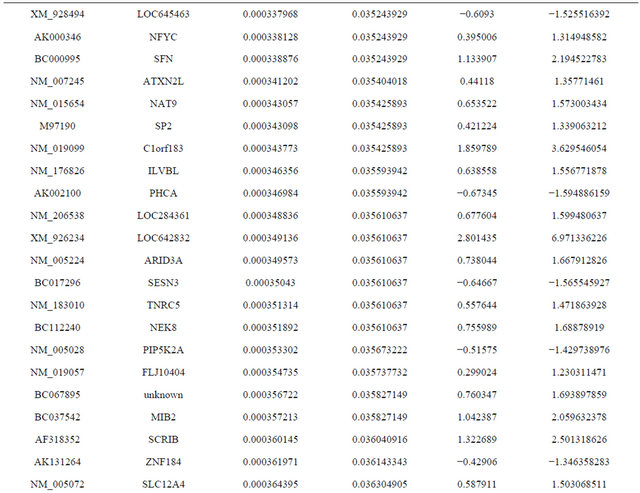
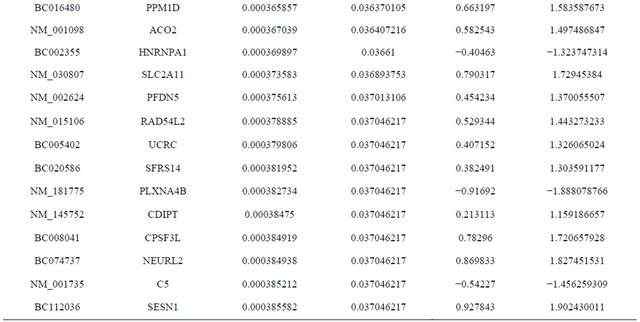
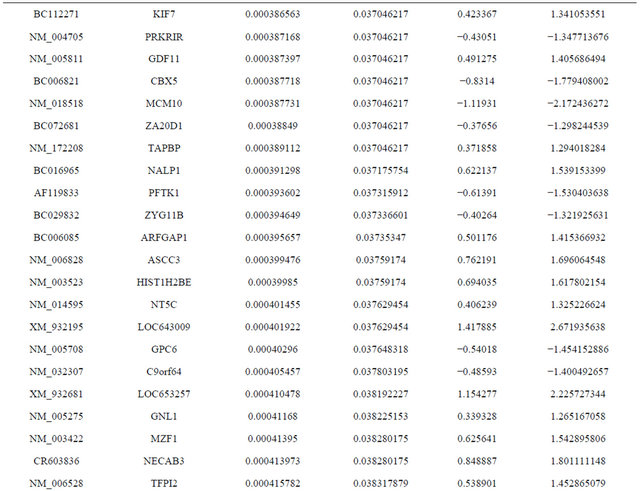
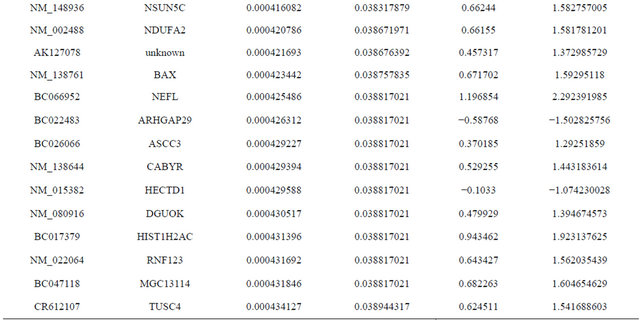
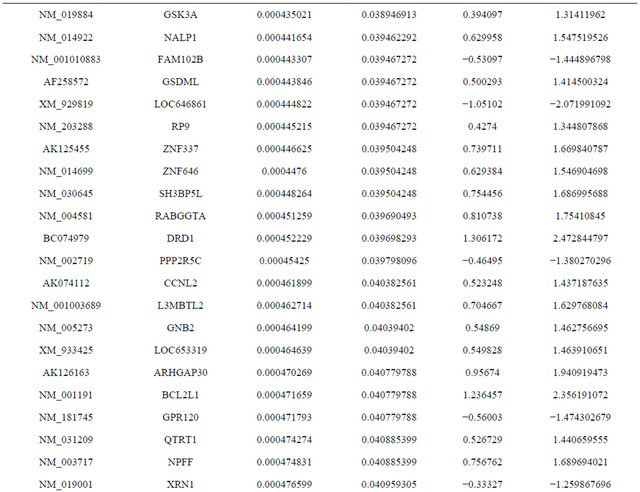
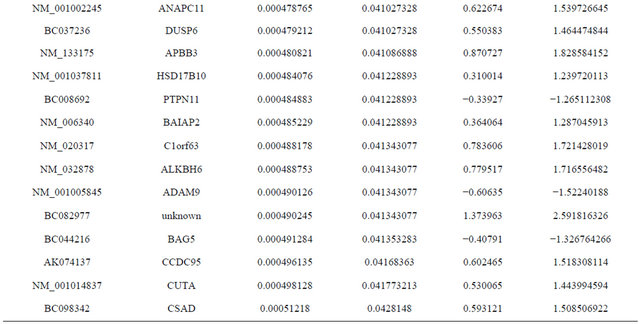
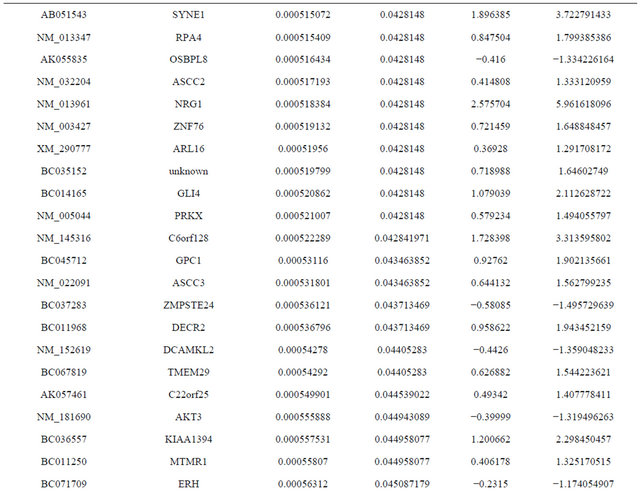
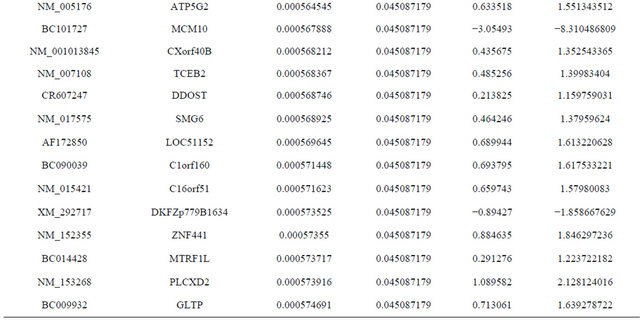
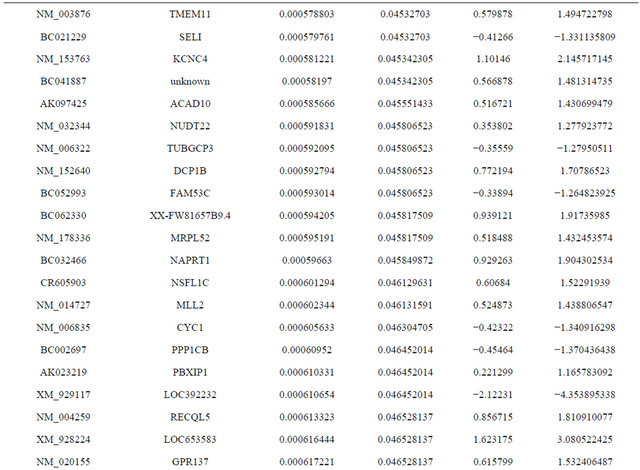
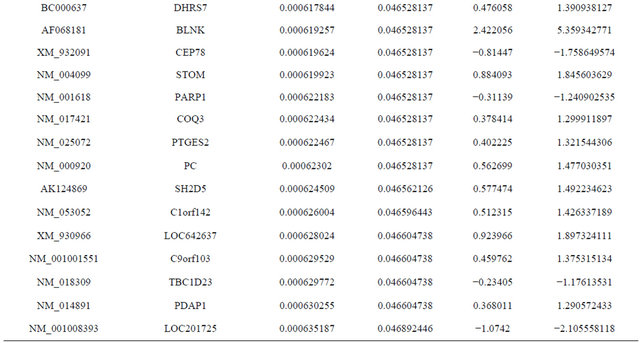
Supplemental list 1. Differentially expressed genes in irradiated cancer-associated fibroblasts.
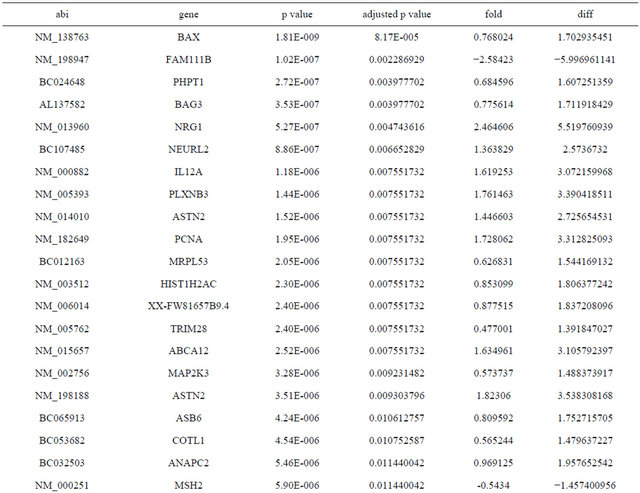
Continued
Continued
Continued
Continued
Continued
Continued
Continued

Continued
Continued
Continued
Continued
Continued
Continued
Continued
Continued
Continued
Continued
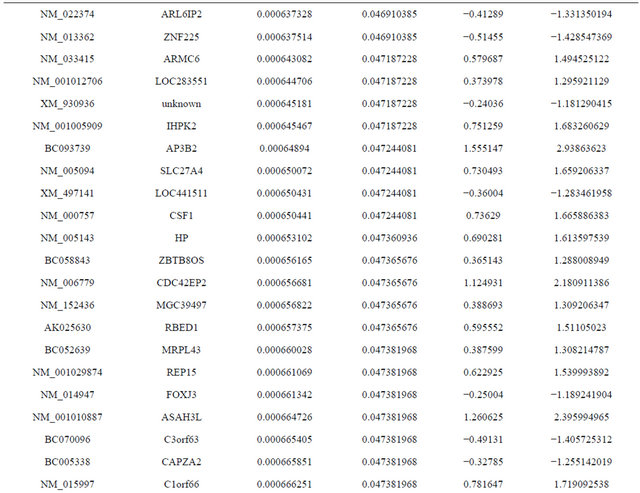
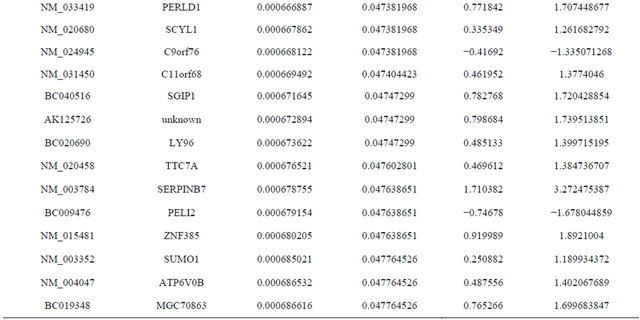
Continued
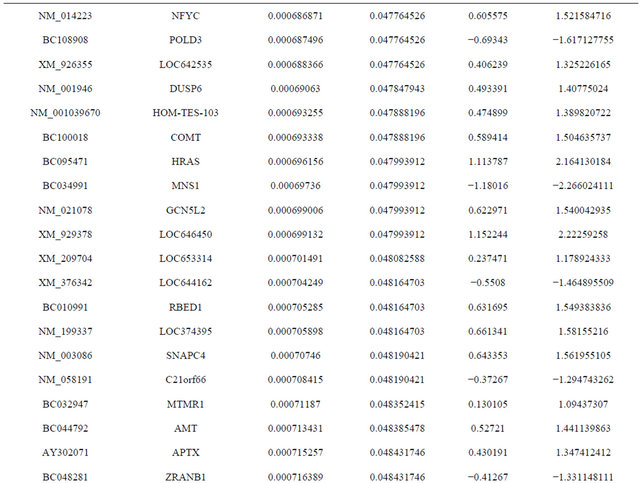
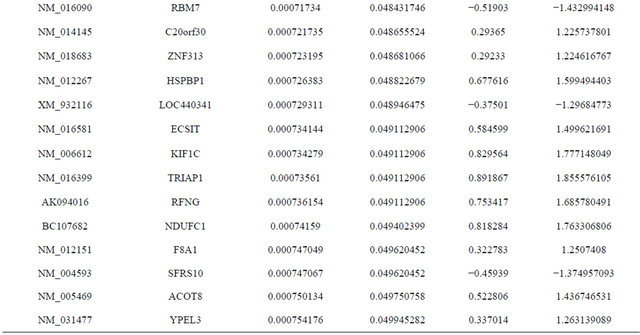
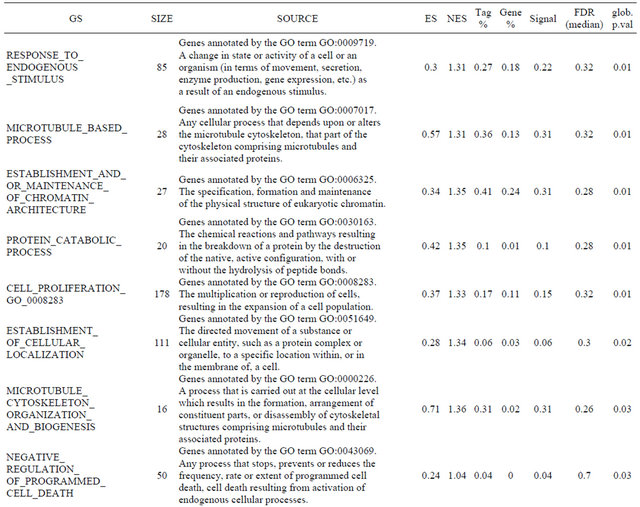
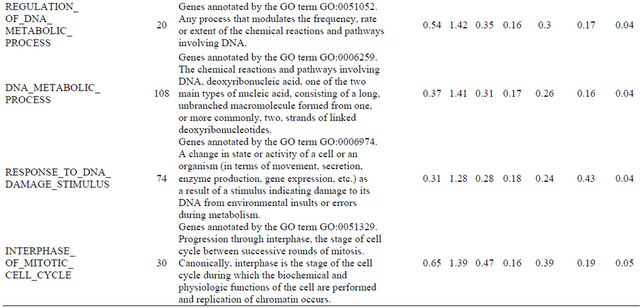
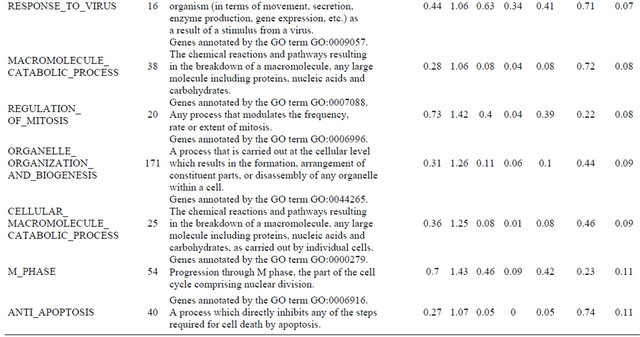
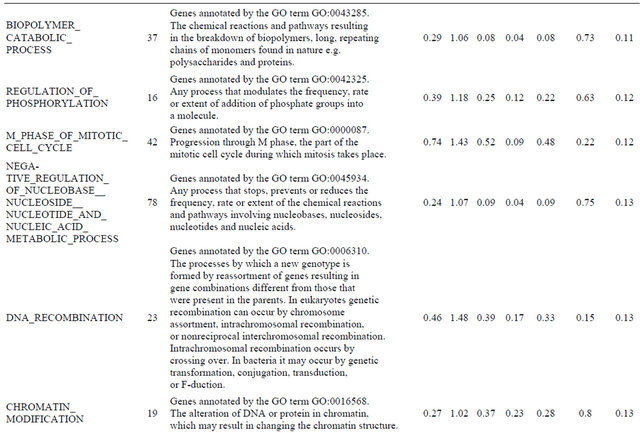
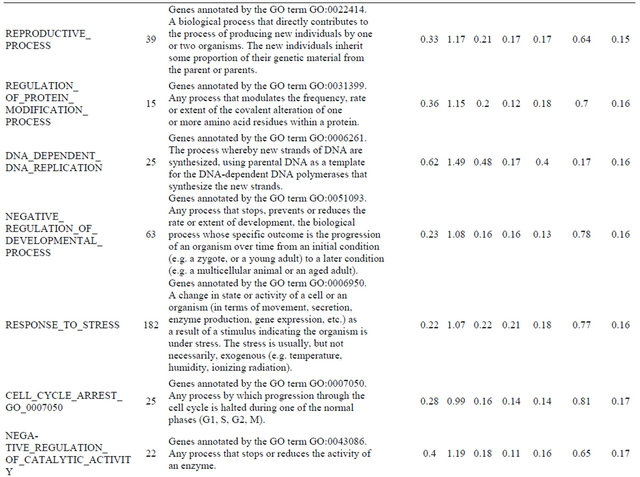
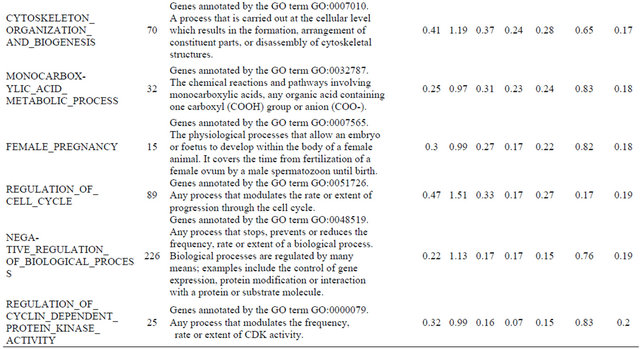
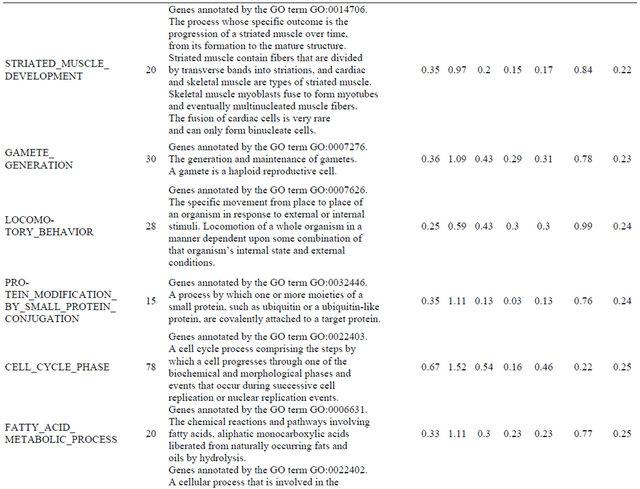
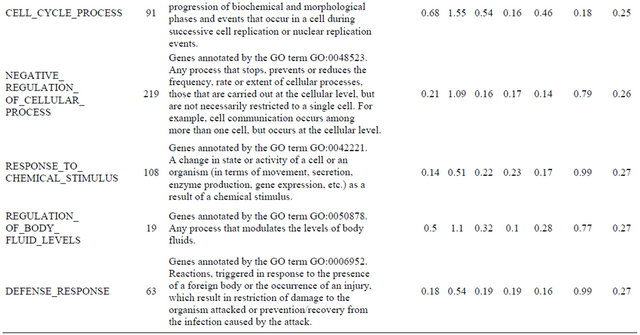
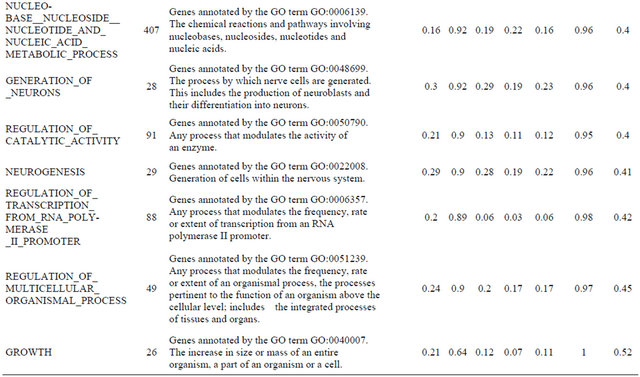
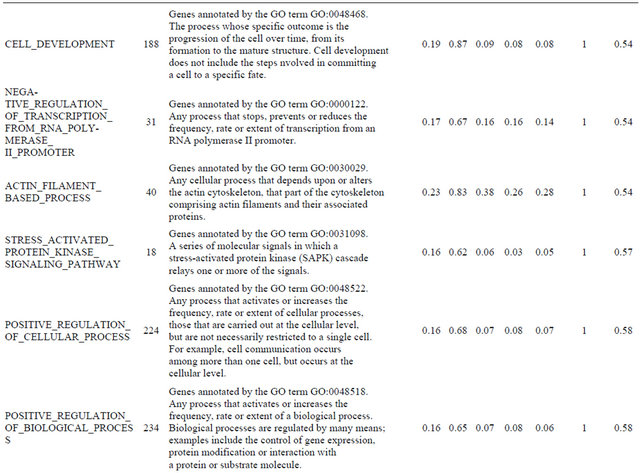
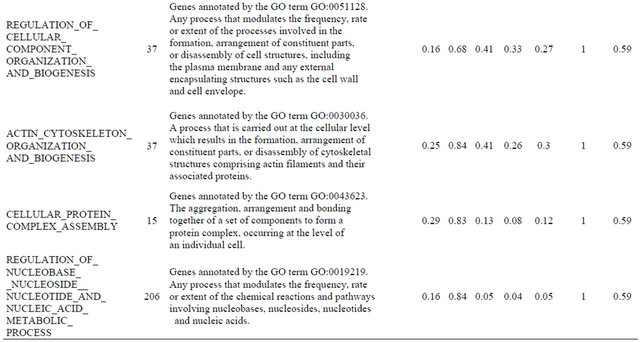
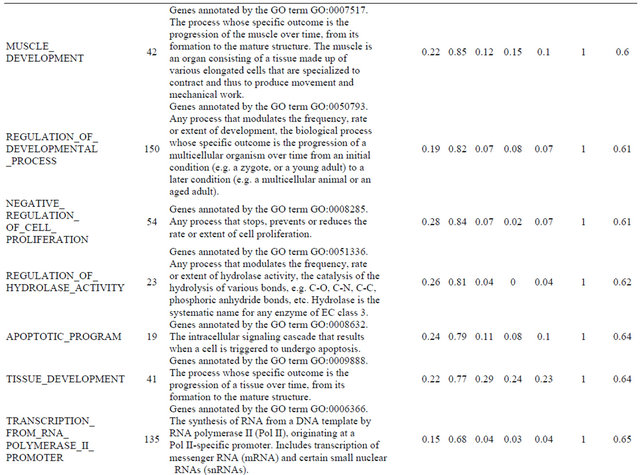
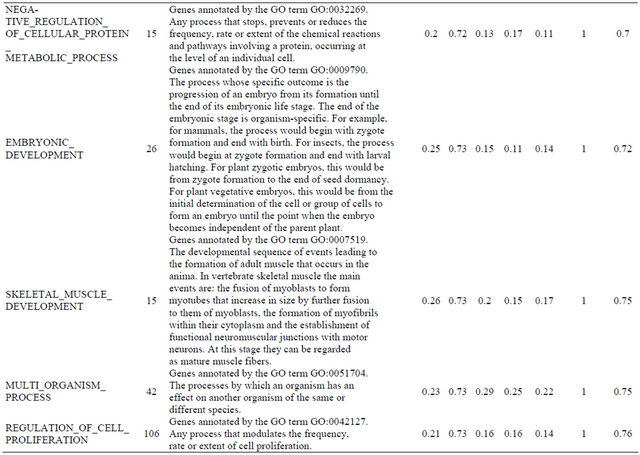
Supplemental list 2. Annotation of differentailly expressed genes in cancer-associated fibroblasts.
Continued

Continued

Continued
Continued
Continued

Continued
Continued

Continued
NOTES
*Corresponding author.

