Advances in Bioscience and Biotechnology
Vol.2 No.4(2011), Article ID:6870,5 pages DOI:10.4236/abb.2011.24042
The early stage of polyphosphate accumulation in Saccharomyces cerevisiae: comparative study by extraction and DAPI staining
![]()
1Skryabin Institute of Biochemistry and Physiology of Microorganisms, Russian Academy of Sciences, Moscow, Russia;
2Institute of Cell Biophysics, Russian Academy of Sciences, Moscow, Russia.
Email: alla@ibpm.pushchino.ru
Received 16 June 2011; revised 13 July 2011; accepted 20 July 2011.
Keywords: Saccharomyces Cerevisiae; Inorganic Polyphosphate; Localization; Cytosol; Mitochondria; Vacuole; Exopolyphosphatase; Glucose; Ethanol; DAPI
ABSTRACT
Inorganic polyphosphate (PolyP), a bioactive polymer with multiple functions, plays a key role in biomineralizaion and phosphorus homeostasis in yeasts. After phosphate starvation, the cells of Saccharomyces cerevisiae restored their pool of PolyP during the first 30 min of incubation in the media containing phosphate and carbon sources. The cells of parent strain accumulated PolyP both in glucose and ethanol-containing media. In the medium with glucose, the strain with inactivated PPX1 and PPN1 genes (encoding two major yeast polyphosphatases) accumulated 2-fold more PolyP than the parent strain. The PolyP in the mutant cells had a greater average chain length compared to the parent strain. The strain with inactivated exopolyphosphatase genes РРХ1 and PPN1 was incapable of PolyP synthesis in the medium with ethanol. The in vivo staining of cells with DAPI show that in the cells of parent strain PolyP appeared first in cytoplasm and mitochondria under cultivation in glucose-containing medium and in cytoplasm and vacuoles in the medium with ethanol. In the ∆PPX1∆PPN1 double mutant PolyP accumulated in cytoplasm and vacuoles under cultivation in glucose-contained medium.
1. INTRODUCTION
Inorganic polyphosphates (PolyP) are the linear polymers containing a few to several hundred orthophosphate (Pi) residues, which are linked by energy-rich phosphoanhydride bonds and perform in living cells a set of functions including participation in stress response, membrane transport, and gene expression regulation [1-5]. PolyP plays the key role in the processes of biomineralization and phosphorus homeostasis [6,7]. The cells of Saccharomyces cerevisiae under certain growth conditions are able to accumulate PolyP in great amounts strongly depending on the growth stage, culture conditions, and a carbon source [8,9].
One of convenient method of PolyP detection in situ is fluorescence microscopy using a fluorochrome 4’,6’- diamino-2-phenylindole 2HCl (DAPI), which is common for DNA detection at concentration of 0.1 to 1 µg/mL. At a high concentration (3 - 50 µg/mL), it stains also PolyP granules and lipid inclusions [10-12]. The emission maximum of DAPI is 456 nm. DNA induces a strong increase in the fluorescence intensity depending on the concentration. PolyP shows a shift of the fluorescence maximum to 525 nm. Pure DAPI-DNA complexes fluorescence is blue-white, while pure DAPI-PolyP complexes fluorescence is yellow [1,12]. DAPI-lipid fluorescence is white-yellow, week and fades in a few seconds [12]. In living cells mixed fluorescence spectra may be observed [13,14]. The green fluorescence of DAPI appears if PolyP are co-localyzed with DNA, or if a mixed spectrum of DAPI and DAPI-PolyP appears.
DAPI staining was used for identification of the vacuolar PolyP [10,13,14] and cell surface PolyP [11] in yeast. However, these studies were carried out with stationary growth stage cells without a comparison of DAPI staining results and chemical extraction of PolyP.
The goal of this research was to compare the PolyP accumulation and DAPI fluorescence at the early stage of biosynthesis of PolyP in S. cerevisiae cells. Besides, the mutants with inactivated genes encoding two yeast exopolyphosphatases, PPX1 and PPN1, are available [15,16]. These enzymes splitting PolyP to phosphate (Pi) differ in their properties and localization in cells [17,18]. So, it is possible to observe the accumulation of PolyP under suppression of its degradation. The effect of carbon source and inactivation of the PPN1 and PPX1 genes on PolyP accumulation and DAPI staining at early stage of biosynthesis of these biopolymers in S. cerevisiae cells is also taken into account in this study.
2. MATERIALS AND METHODS
2.1. Research Objects and Culture Conditions
Saccharomyces cerevisiae strains CRY (parent strain) and the strain CNX with inactivated PPX1 and PPN1 genes CNX (∆PPX1∆PPN1) were kindly provided by Prof. N. Rao and Prof. A. Kornberg (Stanford University) [15,16]. The yeast was cultivated in the YPD medium supplemented with 20 mM potassium phosphate (+P medium) or the YPD phosphate-deficient medium containing 0.02 mM of Pi (−P medium). The (−P) medium was prepared according to [19]. For studying the early stage of PolyP accumulation, the cells grown in the (−P) medium to the stationary growth stage were re-inoculated into the (+P) medium with 2% glucose or 1% ethanol as a carbon source. The initial cell concentration in these experiments was 5 g·L–1 of wet biomass. The cells were cultivates for 30 min and harvested by centrifugation at 3000 g for 10 min, twice washed with distilled water.
2.2. Extraction and Assay of Phosphorus Compounds
Five PolyP fractions were obtained accordingly [8]: PolyP1 was extracted with 0.5 N HClO4; PolyP2 was extracted with NaClO4 (2 g) and HClO4 (0.5 mL) per 1 g of wet biomass; PolyP3 was extracted with weak solution of NaOH, pH 9 - 10; and PolyP4 was extracted with 0.05 N NaOH, pH 12. All fractions were extracted at 0˚C. PolyP5 was assayed by the appearance of Pi during hydrolysis of residual biomass in 0.5 N HClO4 at 90˚C for 20 min [8]. After each extraction procedure, the residual biomass was separated by centrifugation and the supernatants were analyzed. PolyP and Pi were assayed as described previously [8,20].
2.3. Electrophoresis
PolyP was precipitated from extracts with saturated Ba(NO3)2 solution, pH 8.2 [8], and subjected to electrophoresis in 30% polyacrylamide gel with 7 M urea. The gels were stained with 0.05% toluidine blue, 25% methanol and 1% glycerol, and then washed with distilled water.
2.4. Fluorescence Microscopy
The cell suspension in PBS, pH 7.4, was incubated in the presence of 10 µg·mL–1 DAPI (Sigma, USA) at 30˚ for 15 min. Then the cells were centrifuged, re-suspended in PBS with 1% of agarose, placed onto specimen slides, covered with a cover glass, and sealed with nail polish at the periphery of the cover glass to prevent evaporation. The preparations were studied in a Leica TSC SP5 laser scanning microscope (Leica, Germany) with a PL APO 63x/1.40 Oil λbl (lambda blue) objective. Fluorescence was excited by a laser with λ = 405 nm and registered in two spectral ranges: the first channel of 416 - 500 nm and the second channel of 500 - 635 nm. The images were recorded in transmitted light (as a contrasting method) by the DIC method.
3. RESULTS AND DISCUSSION
The amount of PolyP decreased in the cells of parent strain and mutant strain ∆PPX1∆PPN1 without exopolyphosphatases during cultivation in the (−P) medium (Table 1). However, the content of PolyP in ∆PPX1∆PPN1 cells still higher than that in the parent cells. DAPIstained cells of the parent strain under Pi-limitation had insignificant fluorescence (Figure 1(a)). It is in agreement with the low level of PolyP. The cells of the mutant strain showed blue fluorescence of the nuclei and green fluorescence of the cytoplasm and the vacuole (Figure 2(a)). Green fluorescence may be a sum of DAPI selffluorescence (blue) and DAPI-PolyP-fluorescence (yellow) in the case of cytoplasm and vacuoles and a sum of DAPI-DNA and DAPI-PolyP complexes fluorescence in the case of nuclei and mitochondria. The fluorescence of cytoplasm and vacuoles seems to result from the fact the ∆PPX1∆PPN1 cells grown in Pi-deficient medium contains 2-fold more PolyP than the cells of parent strain (Table 1).
After 30 min of cultivation in the (+P) medium (Table 2) the PolyP pools restored to the levels observed at the stationary stage of growth in the (+P) medium (Tables 1 and 2). The cells of parent strain restored the PolyP pool both under use of glucose and ethanol as energy sources. Ethanol did not provide PolyP accumulation in mutant
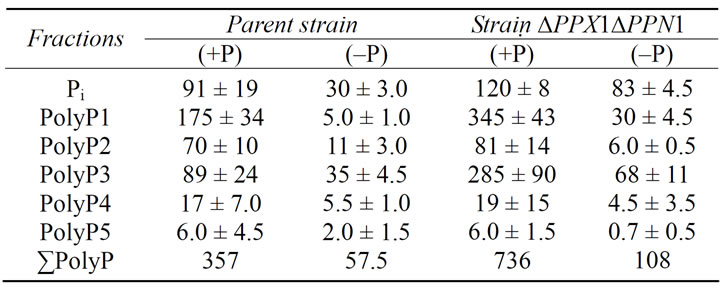
Table 1. The content of Pi and PolyP (µmol/g of dry biomass) in the cells of S. cerevisiae at stationary growth stage in the YPD medium (+Р) (20 mM Рi) and YPD medium (–P) (0.05 mM Рi) with glucose. The average results of four experiments are presented.
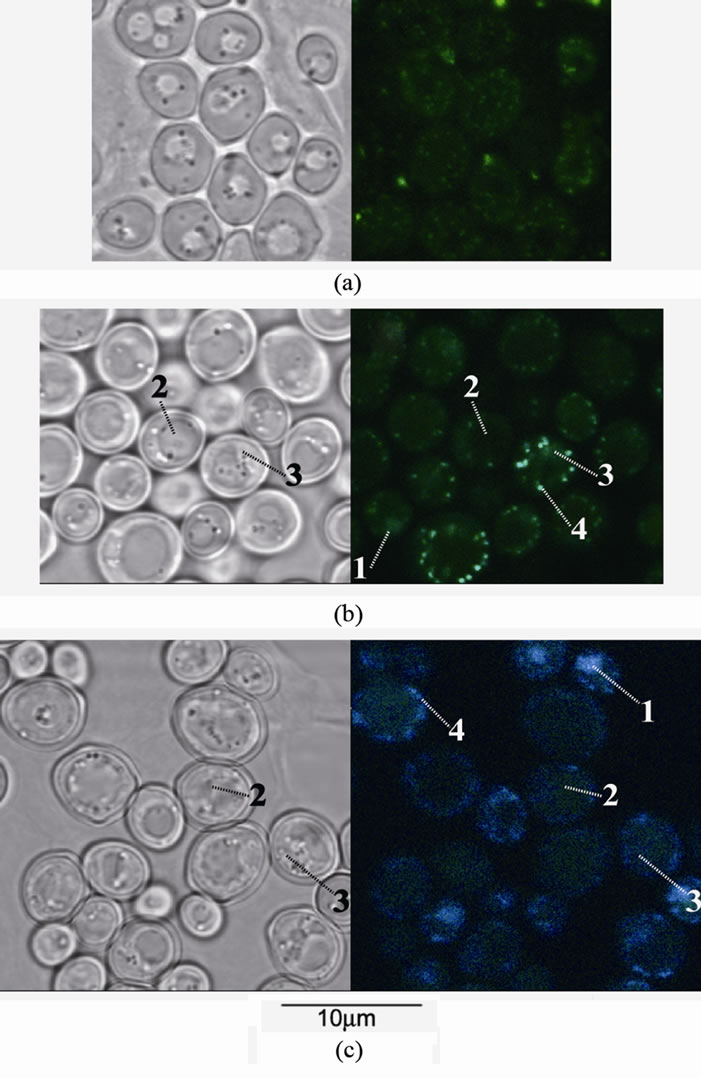
Figure 1. The cells of S. cerevisiae (parent strain) stained by DAPI: left—transmission mode, right—fluorescent mode, (a) the cells grown in (−P) medium with glucose, (b) the cells grown for 30 min in (+P) medium with glucose, (c) the cells grown for 30 min in (+P) medium with ethanol. 1—nucleus, 2—vacuole, 3—cytoplasm, 4—mitochondrial network.
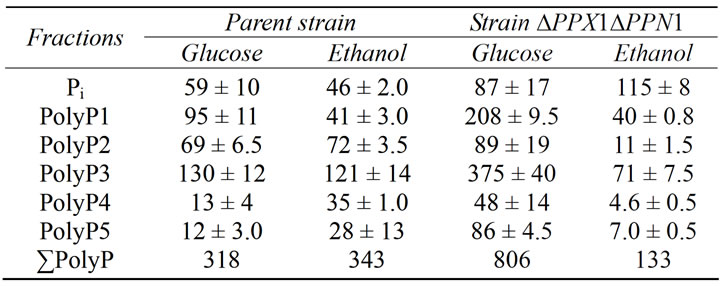
Table 2. The content of Pi and PolyP (µmol/g of dry biomass) in the cells of S. cerevisiae after 30 min cultivation in (+P) medium (20 mM Рi) with glucose or ethanol. The average results of four experiments are presented.
cells (Table 2). In the parent strain, with ethanol used as a carbon source, the content of acid-soluble fractions was lesser than that on glucose, while the content of PolyP3 and PolyP5 was higher. Such effect was observed earlier in the wild type strain [9].
The results of electrophoresis of PolyP accumulated
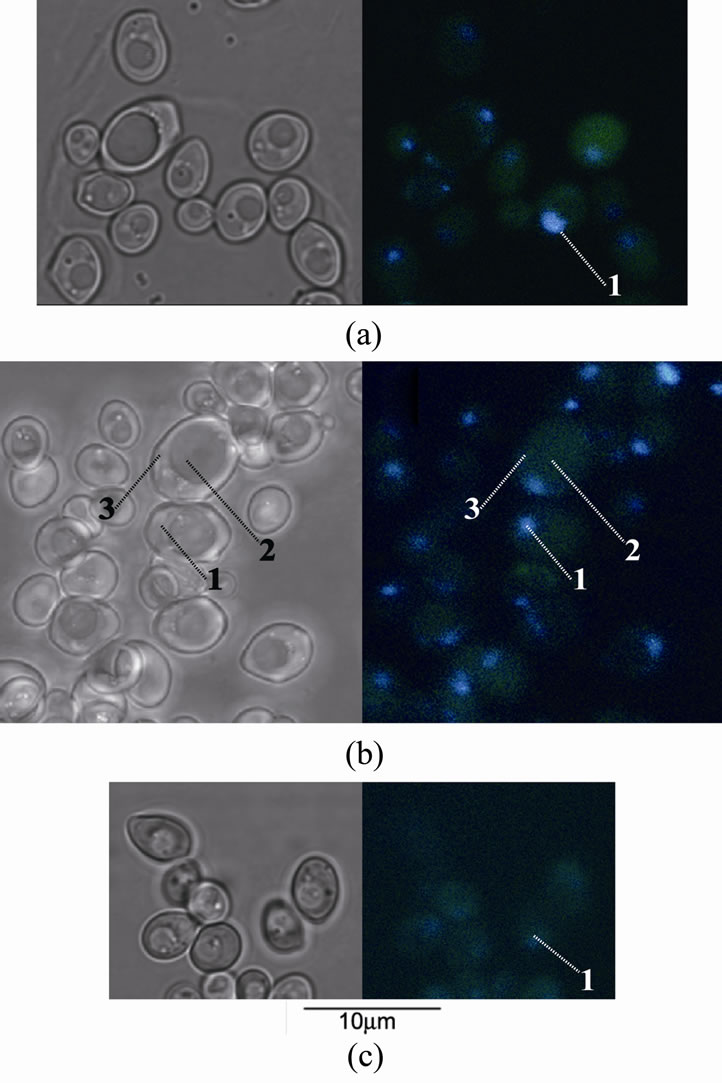

Figure 2. The cells of S. cerevisiae (strain with inactivated genes PPX1 and PPN1) stained by DAPI: left—transmission mode, right—fluorescent mode, (a) the cells grown in (–P) medium with glucose, (b) the cells grown for 30 min in (+P) medium with glucose, (c) the cells grown for 30 min in (+P) medium with ethanol. 1—nucleus, 2—vacuole, 3—cytoplasm.
during 30 min in the (+P) medium with glucose are shown in Figure 3" target="_self"> Figure 3. The average chain length of PolyP in the mutant was greater compared to the parent strain, except for fraction PolyP4. This fraction is believed to be associated with the cell wall, and its metabolism is different from other fractions under study [1].
As whole, the ∆PPX1∆PPN1 cells accumulated more PolyP in the first 30 min than the cells of parent strain in the medium with glucose. It suggested that exopolyphosphatases participate in PolyP metabolism at the first time of its biosynthesis, and biosynthesis and degradation of these polymers occur in parallel.
After 30 min of cultivation in the (+P) medium with glucose, DAPI fluorescence of the parent strain cells considerably increased. The cytoplasm fluoresced in the green spectral region. The nuclei were weakly fluorescent. Fluorescence in the area of vacuoles was not observed either. The small granules fluoresced most brightly
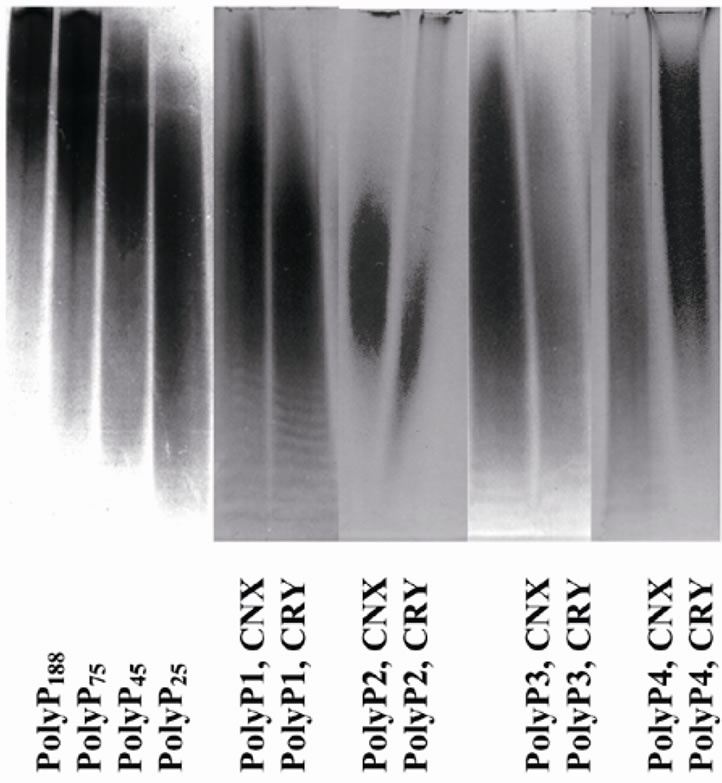
Figure 3. Electrophoregram of PolyP in 20% polyacrylamide gel in the presence of 7 M urea. PolyP were extracted from the cells after 30 min cultivation in (+P) medium with glucose.
(Figure 1(b)). These granules, according to the literature data, are fragments of the mitochondrial network [13]. Previously we have shown that under PolyP accumulation in glucose-containing medium the mitochondrial fraction accumulates PolyP [21]. It seems that the green fluorescence in this case is associated with formation of DAPI-mitochondrial DNA and DAPI-PolyP complexes. The green fluorescence of cytoplasm may be associated with accumulation of PolyP in this compartment.
A different picture was observed with ethanol used as a carbon source (Figure 2(c)). Mitochondria devoid of PolyP [22] under these conditions fluoresced in the blue spectral region typical of DAPI-DNA complexes, along with the nuclei. The cytoplasm and the vacuole fluoresced to approximately the same extent in the green region of the spectrum (Figure 2(c)).
In the mutant strain, under PolyP accumulation in the glucose-containing medium, the green fluorescence of the cytoplasm and vacuoles increased and single areas of yellow fluorescence appeared (Figure 2(b)). The green granules (parts of the mitochondrial network) were not observed. Probably, it was due to defectiveness of mitochondria in this strain [23]. It should also be noted that mitochondria of the mutant strain contain relatively little PolyP at the early growth stage; PolyP in these organelles of mutant strain is accumulated later [23]. With ethanol, cell fluorescence was not different in intensity or character from that of the cells grown under phosphate deficiency (Figures 2(a) and (c)). It is in agreement with the chemical extraction data showing that mutant cells cannot accumulate PolyP in the medium with ethanol as a carbon source. It would be interesting to note that blue fluorescence of the nuclei typical of DAPI-DNA complexes in the mutant strain is much more pronounced than in the parent strain. The nature of this phenomenon cannot be explained as yet; probably, the nuclear envelope permeability for the dye is different for different strains.
Earlier, we demonstrated that cytoplasm, mitochondria, vacuoles and nuclei of S. cerevisiae cells possesses PolyP by subcellular fractionation [24]. However the methods for obtaining of pure fractions of the organelles need long time. The in vivo DAPI staining is a simple method for vital observing of PolyP. The data obtained suggest, that PolyP localization at the initial stage of its synthesis in S. cerevisiae cells depends on the carbon source. PolyP appears first of all in the cytoplasm and mitochondria under cultivation in glucose-containing medium and in the cytoplasm and vacuoles in the medium with ethanol. In the mutant with inactivated genes encoding two exopolyphosphatases (PPX1 and PPN1) PolyP accumulates in the cytoplasm and vacuoles. The role of mitochondria in PolyP synthesis needs further investigation. Comparison of chemical extraction data (Table 2) and DAPI staining (Figures 1 and 2) shows that staining character actually depends on PolyP content. However this dependence is not simple. Nevertheless, DAPI staining is useful for visualization of PolyP localization in living cells. The method of confocal microscopy in vivo should be accompanied with chemical extraction of PolyP. The results show that at the early stage of PolyP biosynthesis after Pi starvation the polymer is immediately localized in different cell compartments, first of all in the cytoplasm. The vacuolar localization of PolyP is more typical of depleted (including Pi-depleted) media [10,25], while formation of DAPI-staining PolyPcontaining granules in S. cerevisiae is observed at the stationary growth stage [14]. Commercial PolyP with average chain length of 25, 45, 75 phosphate residues from Sigma (USA) and with average chain length of 188 phosphate residues from Monsanto (USA) were used as markers.
The comparison of PolyP content and chain length at the early stage of accumulation after Pi deprivation suggest participation of exopolyphosphatases in PolyP metabolism at the first stage of its accumulation. The content and chain lengths of the PolyP1 and PolyP3 fractions increased to the maximum under inactivation of the PPN1 and PPX1 genes. The synthesis of PolyP is probably accompanied by degradation of polymers by exopolyphosphatases. In the double mutant without exopolyphosphatases, this degradation did not occur and the PolyP content and chain length generally increased.
4. ACKNOWLEDGEMENTS
We thank E. Makeeva for help in preparation of the manuscript.
REFERENCES
- Kulaev, I.S., Vagabov, V.M. and Kulakovskaya, T.V. (2004) The biochemistry of inorganic polyphosphates. John Wiley & Sons Ltd., Chichester. doi:10.1002/0470858192
- Rao, N.N., Gomez-Garcia, M.R. and Kornberg, A. (2009) Inorganic polyphosphate: Essential for growth and survival. Annual Review of Biochemistry, 78, 605-647. doi:10.1146/annurev.biochem.77.083007.093039
- Docampo, R., Ulrich, P. and Moreno, S.N. (2010) Evolution of acidocalcisomes and their role in polyphosphate storage and osmoregulation in eukaryotic microbes. Philosophical Transactions of the Royal Society B: Biological Sciences, 365, 775-784. doi:10.1098/rstb.2009.0179
- Chaves, F.P., Lunsdorf, H.L. and Jerez, C.A. (2004) Growth of polychlorinated-biphenyl-degrading bacteria in the presence of biphenyl and chlorobiphenyls generates oxydative stress and massive accumulation of inorganic polyphosphate. Applied and Environmental Microbiology, 70, 3064-3072. doi:10.1128/AEM.70.5.3064-3072.2004
- Campos, E., Façanha, A., Moraes, J. et al. (2007) A mitochondrial exopolyphosphatase activity modulated by phosphate demand in Rhipicephalus (Boophilus) microplus embryo. Insect Biochemistry and Molecular Biology, 37, 1103-1107. doi:10.1016/j.ibmb.2007.06.003
- Gebremariam, S.Y., Beutel, M.W., Christian, D. and Hess, T.F. (2011) Research advances and challenges in the microbiology of enhanced biological phosphorus removal-a critical review. Water Environment Research, 83, 195- 219. doi:10.2175/106143010X12780288628534
- Omelon, S.J. and Grynpas, M.D. (2008) Relationships between polyphosphate chemistry, biochemistry and apatite biomineralization. Chemical Reviews, 108, 4694- 4715. doi:10.1021/cr0782527
- Vagabov, V.M., Trilisenko, L.V. and Kulaev, I.S. (2000) Dependence of inorganic polyphosphate chain length on the orthophosphate content in the culture medium of the yeast Saccharomyces cerevisiae. Biochemistry, 65, 349- 355.
- Vagabov, V.M., Trilisenko, L.V., Kulakovskaya, T.V., et al. (2008) Еffect of carbon source on polyphosphate accumulation in Saccharomyces cerevisiae. FEMS Yeast Research, 8, 877-882. doi:10.1111/j.1567-1364.2008.00420.x
- Allan, R.A. and Miller, J.J. (1980) Influence of S-adenosylmethionine on DAPI-induced fluorescence of polyphosphate in the yeast vacuole. Canadian Journal of Microbiology, 26, 912-920. doi:10.1139/m80-158
- Tijssen, J.P.F., Beekes, H.W. and Van Steveninck, J. (1982) Localization of polyphosphate in Saccharomyces fragilis, as revealed by 4’6’-diamidino-2-phenylindole fluorescence. Biochimica et Biophysica Acta, 721, 394- 398. doi:10.1016/0167-4889(82)90094-5
- Serafim, L.S., Lemos, O.C., Levantesi, C., et al. (2002) Methods for detection and visualization of intracellular polymers stored by polyphosphate-accumulating microorganisms. Journal of Microbiological Methods, 51, 1- 18. doi:10.1016/S0167-7012(02)00056-8
- Puchkov, E.O. (2006) The viability assessment of ethanol-producing yeast by computer-aided fluorescence microscopy. Mikrobiologiia, 75, 193-200.
- Puchkov, E.O. (2010) Brownian motion of polyphosphate complexes in yeast vacuoles:characterization by fluorescence microscopy with image analysis. Yeast, 27, 309-315.
- Wurst, H., Shiba, T. and Kornberg, A. (1995) The gene for a major exopolyphosphatase of Saccharomyces cerevisiae. Journal of Bacteriology, 177, 898-906.
- Sethuraman, A., Rao, N.N. and Kornberg, A. (2001) The endopolyphosphatase gene: Essential in Saccharomyces cerevisiae. Proceedings of the National Academy of Sciences, 98, 8542-8547. doi:10.1073/pnas.151269398
- Lichko, L.P., Andreeva, N.A., Kulakovskaya, T.V., et al. (2003) Exopolyphosphatases of the yeast Saccharomyces cerevisiae. FEMS Yeast Research, 3, 233-238. doi:10.1016/S1567-1356(02)00205-2
- Andreeva, N.A., Kulakovskaya, T.V. and Kulaev, I.S. (2006) High molecular mass exopolyphosphatase from the cytosol of the yeast Saccharomyces cerevisiae is encoded by the PPN1 gene. Biochemistry, 71, 975-977.
- Rubin, G.M. (1973) The nucleotide sequence of Saccharomyces cerevisiae 5.8 S ribosomal ribonucleic acid. Journal of Biological Chemistry, 11, 3860-3875.
- Kulakovskaya, T.V., Andreeva, N.A., Karpov, A.V., et al. (1999) Hydrolysis of tripolyphosphate by purified exopolyphosphatase of Saccharomyces cerevisiae cytosol: Kinetic model. Biochemistry, 64, 990-993.
- Pestov, N.A., Kulakovskaya, T.V. and Kulaev, I.S. (2004) Inorganic polyphosphate in mitochondria of Saccharomyces cerevisiae at phosphate limitation and phosphate excess. FEMS Yeast Research, 4, 643-648. doi:10.1016/j.femsyr.2003.12.008
- Andreeva, N.A., Kulakovskaya, T.V., Kulakovskaya, E.V., et al. (2008) Polyphosphates and exopolyphosphatases in cytosol and mitochondria of Saccharomyces cerevisiae during growth on glucose or ethanol under phosphate surplus. Biochemistry, 73, 65-69.
- Pestov, N.A., Kulakovskaya, T.V. and Kulaev I.S. (2005) Effects of inactivation of the PPN1 gene on exopolyphosphatases, inorganic polyphosphates and function of mitochondria in the yeast Saccharomyces cerevisiae. FEMS Yeast Research, 5, 823-828. doi:10.1016/j.femsyr.2005.03.002
- Lichko, L., Kulakovskaya, T., Pestov, N. and Kulaev, I. (2006) Inorganic polyphosphates and exopolyphosphatases in cell compartments of the yeast Saccharomyces cerevisiae under inactivation of PPX1 and PPN1 genes. Bioscience Reports, 26, 45-54. doi:10.1007/s10540-006-9003-2
- Trilisenko, L.V., Vagabov, V.M. and Kulaev, I.S. (2002) The content and chain length of polyphosphate from vacuoles of Saccharomyces cerevisiae VKM Y-1173. Biochemistry, 67, 592-596.
NOTES
* The work was supported by the Russian Foundation for Basic Research (grant 11-04-01498)

