Advances in Bioscience and Biotechnology
Vol.1 No.4(2010), Article ID:2864,9 pages DOI:10.4236/abb.2010.14046
Genetic variability of Aspergillus terreus from dried grapes using RAPD-PCR
![]()
Vels University, Pallavaram, Chennai, India.
Email: banunkl@yahoo.com
Received 6 May 2010; revised 7 June 2010; accepted 18 June 2010.
Keywords: Aspergillus Terreus, Dried Grapes, Genetic Variability, RAPD-PCR
ABSTRACT
RAPD was used to examine the genetic variability among five isolates of Aspergillus terreus spp. Two random primers were selected for the RAPD assay PG01–5’ CAGGTGTTGC 3’ and PG02–5’ CTGGACAGAC 3’ (Progen Technologies). The characterization of Aspergillus terreus species have been mostly applied on the basis of morphology, phenotype and physiology. DNA Polymorphisms are based on differences in DNA sequences and have advantages over protein polymorphisms. But morphological characterization besides molecular tools will remain a basic and powerful key in the identification of Aspergillus terreus species. The objective of the present study was to isolate the fungal contaminants from dried grapes and compare the genomic profile of the Aspergillus terreus speices isolated from the dried grapes, through RAPD analysis. In the present study with primer PG 01 four different discriminations was there among the A. terreus isolates. There was a homology of genotype between the isolates 1 & 3. And with primer PG 02 four different discriminations were there and there was a homology between 1 & 3. The predominant type was type I in primer I & II. The other isolates belonged to 2, 3 and 4. No similarity was detected for isolates 3, 4 and 5 indicating great genomic diversity of A. terreus.
1. INTRODUCTION
Aspergillus terreus is a common soil saprophyte that has been recovered from desert soil, grasslands and compost heaps. A. terreus produces a variety of secondary metabolites that are economically significant, such as lovastatin, a antihypercholesteroleic drug [1]. Importantly, most members of this species have decreased susceptibility to the antifungal drug amphotericin B (AMB) in vitro and in vivo [2]. Accordingly, the ability to distinguish A. terreus from other species of Aspergillus can be important to the clinician for therapeutic decision making and prognosis.
In addition, A. terreus produces globose, heavy-walled hyaline cells laterally on the hyphae, which are called accessory conidia or are sometimes referred to as aleuroconidia. A. terreus is the only member of the genus Aspergillus that produces such structures [3].
The genus Aspergillus is divided into seven subgenera, which in turn are subdivided into several sections with each section comprised of a few to several closely related species. Using morphological methods as delineators, Raper and Fennell recognized A. terreus as the only known species within the section Terrei, subgenus Nidulantes, along with the rarely encountered varieties of A. terreus var. aureus and A. terreus var. africanus [3]. Both of these varieties of A. terreus differ significantly from A. terreus morphologically in that both grow as bright orange colonies and A. terreus var. africanus sometimes produces sclerotium-like bodies on laboratory media.
Several recent studies have clearly demonstrated that morphological methods are poor markers of species in the genus Aspergillus, and molecular methods may be useful in species identification [4,5]. Phylogenetic analyses of DNA sequences derived from ITS regions and showed the presence of three main clades, which included A. terreus, A. carneus and A. niveus within the section Terrei [1]. However, that study concluded the section Terrei warranted further detailed molecular analysis including more loci and isolates. It has been known for sometime that apart from A. terreus, numerous cryptic species may be present within the section Terrei. Phylogenetic analyses using sequence information from multiple protein coding regions may further clarify the taxonomy of these isolates within the section.
Previous phylogenetic studies in the section Fumigati, genus Aspergillus have revealed the presence of numerous species which were not morphological methods alone. For example, A. lentulus as a novel, cryptic species within the section Fumigati by employing both the maximum likelyhood algorithms, as well as utilizing DNA sequence information from the ITS region, the mitochondrial cytochrome b region, and protein coding region for the β-tubulin gene, the rodlet A gene and the salt responsive gene [4]. Phylogenetic analyses revealed that A. lentulus isolates were present as a monophyletic clade in the majority of the single-locus genealogies, thus, fulfilling the phylogenetic species recognition concept, which requires both genealogical concordance and genealogical nondiscordance in the genealogies evaluated.
Invasive fungal infection due to Aspergillus species have become a major cause of morbidity and mortality among immunocompromised patients. Aspergillus fumigatus is most frequently isolated from clinical specimens, but other important species include A. flavus, A. niger and A. terreus. A. terreus appears to be emerging as a cause of opportunistic infections [5-7].
Aspergillus spp. moulds can cause various diseases, invasive aspergillosis (IA), aspergilloma, allergic bronchopulmonary aspergillosis, rhinocerebral aspergillosis and otomycosis being the most frequent forms. IA is the most life threatening one, which recently possess a threat not only to patients suffering from hematological malignancies [8]. It also occurs in a much broader patient population then the classical immunocompromised hosts and includes mechanically and ventilated intensive care unit patients and patients receiving coreicosteroids for treatment of chronic lung diseases [9].
RAPD-PCR typing has proven useful in many epidemiological investigations [2,10]. The polymerase chain reaction (PCR) has been adapted for fingerprinting microorganisms by using paired primers derived from previously characterized sequences for PCR amplification. This technique has been applied to bacteria, [11], viruses [12] and fungi .
Based on PCR amplifications, a new DNA polymorphism assay, random amplification of polymorphic DNA (RAPD), was developed [13,14]. Random sections of genomic DNA are amplified by using a single arbitrary nucleotide sequence to make polymorphic DNA patterns.
No prior sequence information is required for this method, which has been used to differentiate organisms with close relationships and particularly to differentiate isolates below the species level [15-19]. Furthermore, PCR makes it possible to analyze a small amount of genetic material and handle a large number of specimens in a short time. Furthermore, RAPD was applied to these strains to develop an accurate and simple method for differentiating Aspergillus terreus.
The method has been applied to two fungal plant pathogens, Fusarium solani and Leptosphaeria maculans [15,20]. The application of the random amplification of polymorphic DNA (RAPD) assay to the human pathogen A. fumigatus was described in the year 1993 [16].
Recently, the development of molecular biology has made it possible to study taxonomy by investigating genetic materials directly in addition to observing the expresses behavior of genetic materials. Several molecular characteristics, such as DNA base composition, DNA hybridization, and rRNA relatedness, etc., have been applied in fungal classification [21]. PCR was adapted to fingerprint microorganisms by using paired primers derived from previously characterized sequences for PCR amplification.
Besides, with RAPD, the appearance of new ligation sites only creates new annealing sites without impairing pre-existing ones, which would continue to be amplified. Consequently, RAPD allows an independence of characters, which is a clear advantage over techniques that directly or indirectly make use of restriction enzymes, such as restriction fragment length polymorphism (RFLP). Various phenotypic and genotypic methods have been used successfully in fingerprinting A. fumigatus. However, no comparable data are available for A. terreus. For those reasons, we aimed to compare the genomic profile of different strains of A. terreus isolated from five samples through RANDOM AMPLIFICATION OF POLYMORPHIC DNA (RAPD) analysis.
2 MATERIALS AND METHODS
2.1. Collection of Samples
Five different samples of dry grapes were collected for the isolation of Aspergillus terreus.
2.2. Morphological Studies
2.2.1. Isolation and Identification
Five different samples of dry grapes were collected for the isolation of A. terreus. Grapes were surface sterilized with 0.1% Mercuric chloride and placed on the CDA medium. The plates were then incubated at room temperature for about 3-4 days. Triplicates were maintained for each sample. The identification and nomenclature of Aspergillus terreus was done with standard manual [3].
2.2.2. Subculturing
Among the various colonies identified, only Aspergillus terreus colonies were subcultured onto the Czapek-Dox Agar plates amended with chloramphenicol under sterile conditions, to obtain the pure culture. These plates were then incubated at room temperature (30°C) for about 3-4 days.
2.3. Molecular Genotyping
The Czapek-Dox Broth was prepared and 100 ml of the medium was dispersed equally into ten separate 250ml conical flasks. These were sterilized in an autoclave at 121°C for 15 minutes at 15 lbs pressure. The flasks were inoculated with the isolates of Aspergillus terreus under sterile conditions and were then subjected for incubation at room temperature for about 3-4 days.
The flasks were observed every day for the growth of the fungus. The mycelial mats should be harvested within 36-40 hours of incubation. The harvested mats were then stored at 4℃ under sterile conditions for further use.
2.3.1. DNA Extraction and Isolation [22]
0.2-0.5 g (dry wt) of lyophilized pad was taken in a 50 ml disposable centrifuge tube, by brief shaking the pad was powdered with the help of a spatula or glass rod. 10 ml (for 0.5 g pad) of CTAB extraction buffer was added gently mixed to wet all the powdered pad. It was placed in 65℃ water bath for 30 minutes. It was Cooled and equal volume chloroform: ethanol (24:1) was added and mixed. The tubes were centrifuged in table-top centrifuge for 10 minutes at full speed. The aqueous supernatant was transferred to a new tube. An equal volume of isopropanol was added and centrifuged at low speed for 5 minutes. The supernatant was poured out. The pellet containing DNA was rinsed with 70% ethanol. It was air dried and 1-5 ml of TE containing 20 µg/ml of Rnase A was added. The samples were placed in 65℃bath, or pellets were allowed to resuspend overnight at 4℃.
2.4. RAPD Analysis
The PCR Primer Used in this work was PG01–5’ CAGGTGTTGC 3’ and PG02–5’ CTGGACAGAC 3’ (Progen Technologies). The primer is 10 nucleotides in length, had G-C and A-T contents more or less equal in amounts and did not contain palindromic sequences.
Amplification was carried out with a 50 µL reaction mixture containing the following.
Primer (2 µM/µL) - 8.0 µL
10X Buffer - 5.0 µL
2 MmM dNTP Mix - 5.0 µL
Taq DNA polymerase (U/µL) - 0.5 µL
Template DNA (50 ng) - 2.0 µL
Sterile Distilled Water - 29.5 µL
Total - 50.0 µL
2.4.1. PCR Amplification Condition
The Amplification was carried out with Eppendorf Mastercycler® ep thermal cycler. The program used to carry out the amplification process is an initial denaturation at 94℃ for 5 min, with 34 cycles of denaturation at 94℃ for 40 sec, annealing at 36℃ for 30 sec, extension at 72℃ for 90 sec and a final extension at 72℃ for 10 mins. Both the primers were run under the same conditions.
A blank control with all the PCR reagents except DNA was always included. The resulting banding patterns were indexed by capital letters and even a single band mismatch led to a different letter code.
The amplified products were separated on a 0.8-1% agarose gel in 1X TBE at 75V for 3 h. The gel was stained with ethidium bromide and the amplified product was visualized under a UV transilluminator. Gel products were sized to a relative to a 1 kb ladder standard. A molecular weight marker was run in parallel. RAPD patterns were compared visually. An isolate was defined as different when a band with a density equal to or greater than that that of the 0.8 kb band of the marker occurred and no corresponding band (not even a trace) was detectable in the other isolates. To demonstrate reproducibility of the patterns, all isolates were investigated three times independently, with different subcultures.
2.5. Discrete Character Programme
Tree was constructed by using Phylip 3.69.
3 RESULTS
Altogether, five different fungi were isolated from the dried grapes. Among this Aspergillus terreus is the most prevalent organism (Figure 1 & 2).
3.1. Morphological Characteristics of Aspergillus Terreus
Aspergillus terreus belongs to the genus Aspergillus, sub-genus Nidulantes and it was identified in the laboratory by morphological methods. The A. terreus isolates did not differ in their macroscopic and microscopic morphology on Czapek-Dox agar, A. terreus showed rapid growth with very variable colony appearance ranging from heavily sporulating colonies to fluffy, poorly sporulating phenotypes. Microscopically, conidiophores are typically long, colourless (hyaline) and smooth giving rise to sub-spherical vesicles that are biseriate. Conidia range in size from 1.5-2.4 µm in diameter, are smooth walled, slightly elliptical and striate (Figure 3 & 4).
3.2. Molecular Characterization
3.2.1. Isolation of DNA
Genomic DNA was isolated from Aspergillus terreus obtained from five different dried grapes samples based
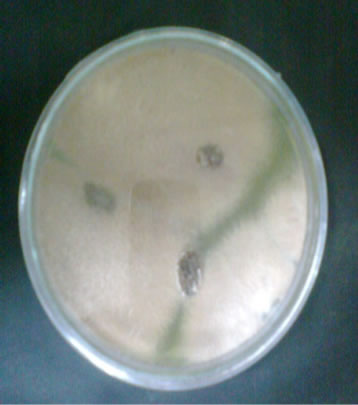


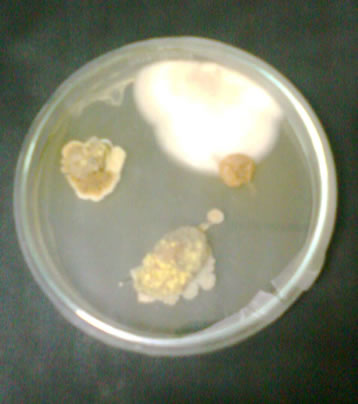
Figure 1. Plates showing fungal colonies on dried grapes.
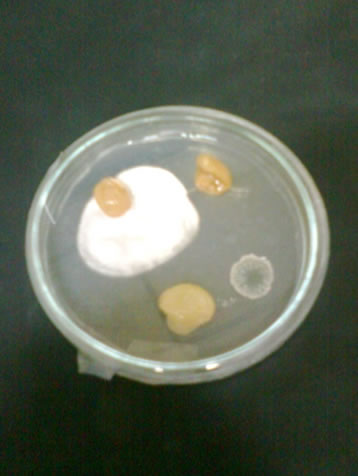


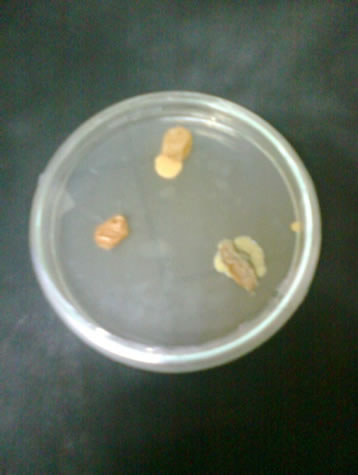
Figure 2. Plates showing fungal colonies on dried grapes.
on the protocol standardized. The isolated genomic DNA was checked on 0.8-1% agarose gel (Figure 5).
3.2.2. RAPD Analysis
3.2.3. Genotypic Typing
The isolated genomic DNA of Aspergillus terreus from dried grapes was amplified using the primers PG01–5’ CAGGTGTTGC 3’ and PG02–5’ CTGGACAGAC 3’ (Progen Technologies) and the amplification was carried out with EPPENDORF Mastercycler ep thermal cycler. The amplified DNA was checked in 0.8-1% agarose gel electrophoresis as shown in Plate VI & VII. The amplified DNA bands of Aspergillus terreus isolates were observed and compared to estimate the genetic diversity. Though there were many bands in the gel, only the brighter bands were selected for the analysis of genetic diversity.
The isolates 1 & 2 produced the bands at 1500 kb, 1750 kb and 2000 kb. The isolates 3 & 4 produced the bands at 1250 kb and 1500 kb. The isolates 4 & 5 produced the bands at 1200 kb. Only the isolate 4 produced a single band at 2750 kb. The isolate 1, 2, 3 and 4 produced the bands at 1500 kb (Table 1). The isolates 1, 3 and 5 produced the bands ay 1200 kb. The isolates 4 and 5 produced bands at 1500 kb. Only the isolate 2 produced a single band at 1250 kb. The isolate 5 produced a single band at 1750 kb and 2000 kb (Table 2). Two different primers were used for typing in RAPDPCR. Both the primers discriminated four RAPD types within the five isolates of Aspergillus terreus. The predominant type was type I in primer I and primer II. The other isolates belonged to II, III and IV (Table 3).
3.2.4. Phylogenetic Tree
Phylogenetic tree based on random amplified polymorphic DNA Profiles (RAPD) showing the genetic relationships among the four isolates of Aspergillus terreus isolated from fresh grapes were generated with the primers PG01 and PG02.
4. DISCUSSION
The present study shows A. terreus to be common in the environmental as a contaminant of various food items and to be common as a cause of Aspergillosis in Immunocompromised patients.
The molecular genotyping of Aspergillus species has proven useful in many epidemiologic situations. One of the most widely used genotyping techniques is RAPDPCR, a relatively technically simple and rapid procedure. Although RAPD-PCR has been criticized for a lack of
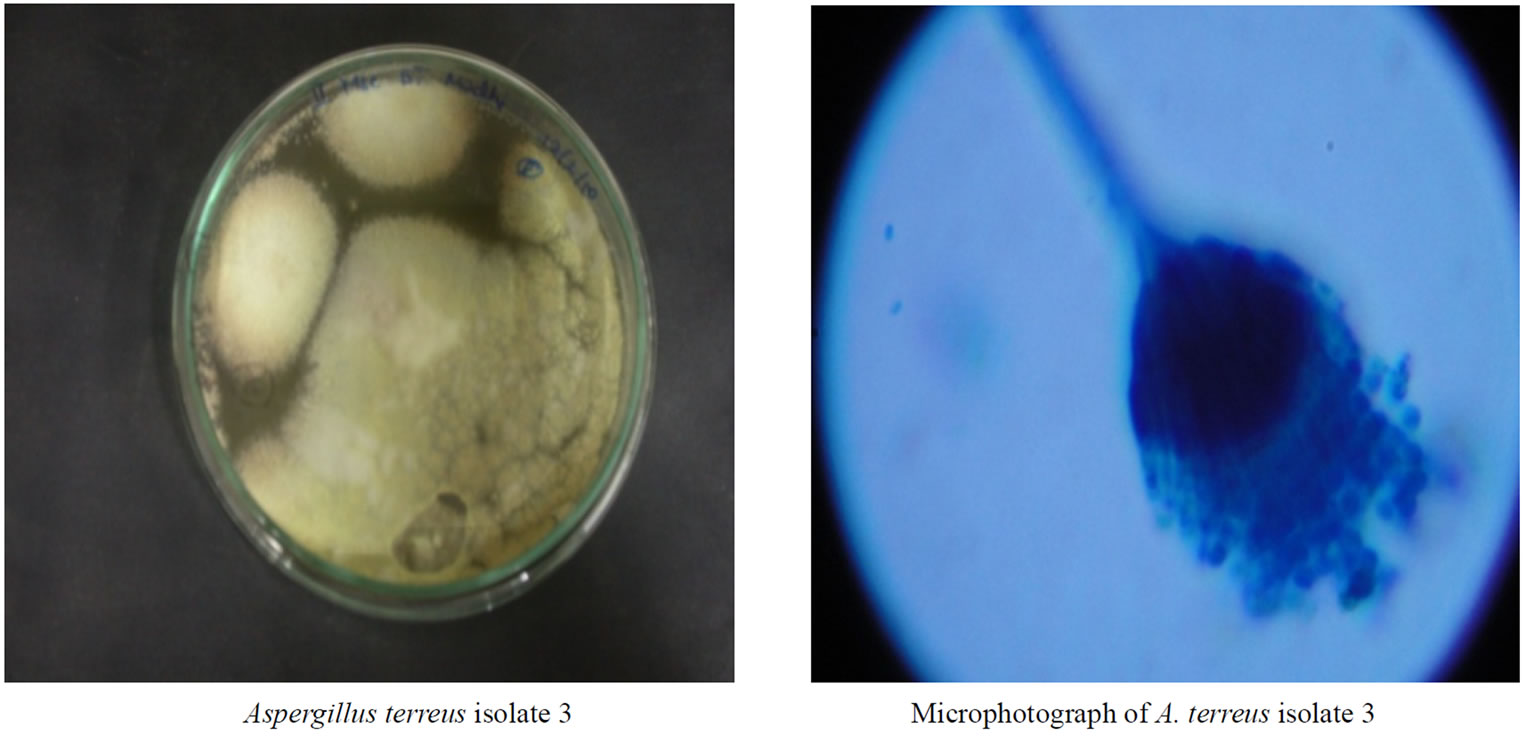


Figure 3. Plates with Aspergillus terreus isolates & its microphotographs.


Figure 4. Plates with Aspergillus terreus isolates & its microphotographs.
reproducibility, this method has been used with the success for A. terreus isolates.
Five primers to test 14 isolates and found primers R108 and CII to be highly discriminatory [23]. Symoens and colleagues tested 43 isolates of environmental and clinical origins with the primers NS3 and NS7 [24]. Among epidemiologically unrelated species, the primers were highly discriminatory. In addition, they found RAPD-PCR to be useful tool in demonstrating the clonal origin of contamination of the environment in a hematology unit.
In the present study there are five different fungi were isolated (Figure 1, 2 and 3). Among these Aspergillus terreus was given special emphasis for its genetic diversity. Five different isolates of A. terreus were isolated and they were subjected to morphological characterization. No difference was found in the macroscopic and microscopic morphology of the five different A. terreus isolates. Similar results were obtained by Rath [10]. The first report of different methods of strain differentiation for A. terreus was given by them. All isolates were produced aleuriospores, which are not only found in A. terreus, but also in A. flavipens, A. niveus and A. carneus [3] were found in the isolates under investigation. The RAPD analysis we performed was intended to assess the genetic diversity within the A. terreus species. RAPD fingerprints allow intraspecific genetic diversity studies but are not very good tools for such analysis at the interspecific level.
In the present study with primer PG 01 four different discriminations was there among the A. terreus isolates (Table 1; Figure 6). There is a homology of genotype between the isolates 1 & 3. And with primer PG 02 four different discriminations was there and there is a homology

Lane 1-1 kb ladder; Lane 2-Aspergillus terreus isolate 1; Lane 3-Aspergillus terreus isolate 2; Lane 4-Aspergillus terreus isolate 3; Lane 5-Aspergillus terreus isolate 4; Lane 6-Aspergillus terreus isolate 5; Lane 7-1 kb ladder.
Figure 5. Genomic DNA of Aspergillus terreus isolates.

Table 1. RAPD Profile of Aspergillus terreus isolates from different samples with Primer PG01.

Table 2. RAPD Profile of Aspergillus terreus isolates from different samples with Primer PG02.

Table 3. RAPD patterns of five A. terreus produced by Primers PG01 & PG02.
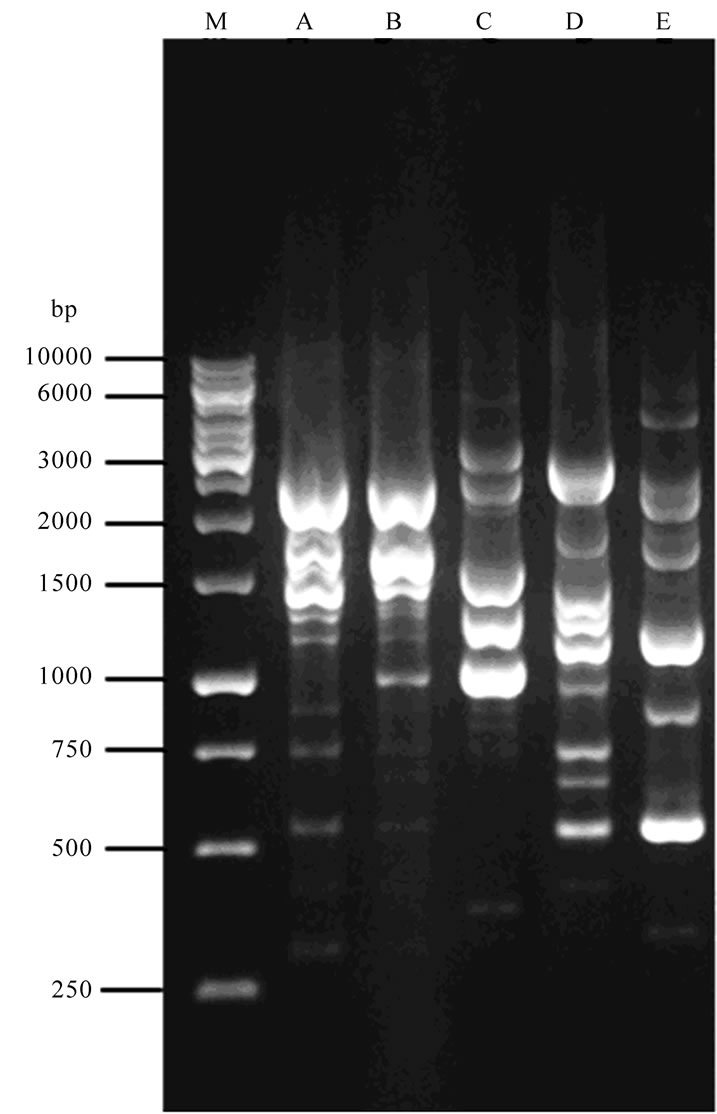
Lane 1-10000 bp ladder; Lane 2-Aspergillus terreus isolate 1; Lane 3-Aspergillus terreus isolate 2; Lane 4-Aspergillus terreus isolate 3; Lane 5-Aspergillus terreus isolate 4; Lane 6-Aspergillus terreus isolate 5.
Figure 6. RAPD-PCR of Aspergillus terreus isolates with primer PG01.
between 1 & 3 (Table 2; Plate 7). The predominant type was type I in primer I & II. The other isolates belonged to 2, 3 and 4. No similarity was detected for isolates 3, 4 and 5 indicating great genomic diversity of A. terreus (Table 3). In fact, RAPD technology is somewhat outdated. It usually generates high false positive rate and the detected bands need further sequencing to generate sufficient diversity information.

Lane 1-10000 bp ladder; Lane 2-Aspergillus terreus isolate 1; Lane 3-Aspergillus terreus isolate 2; Lane 4-Aspergillus terreus isolate 3; Lane 5-Aspergillus terreus isolate 4; Lane 6- Aspergillus terreus isolate 5.
Figure 7. RAPD-PCR of aspergillus terreus isolates with primer PG02.
RAPD-PCR typing of A. terreus has proven useful in many epidemiological investigations. Varga [1] applied RAPD-PCR and showed a high degree of variability among A. terreus isolates, even among those indistinguishable based on their internal transcribed spacer sequences. Florl [25] reported no strain similarity in M. D. Anderson Cancer Centre in Houston, TX, and at the University Hospital of Innsbruck, Austria was detected, indiating great genetic diversity of A. terreus.
Rath [10] reported nine different genotypes of A. terreus by RAPD analysis. Previously, this technique was used successfully in fingerprinting other Aspergillus [16, 10] showing a high genetic diversity in A. fumigatus and A. flavus. Similar results were also found for A. terreus in the present study (Figure 1) Therefore, RAPD seems to demonstrate diversity within a species.
Florl [25] reported the genotypic diversity of 12 A. terreus. Among the 12 strains ten were from one geo-

Figure 8. Phylogenetic tree of Aspergillus terreus isolates.
graphic area, were investigated by phenotypic and genotypic method. In that RAPD analysis was the most discriminatory technique compared to RFLP and Southern Blot Hybridization.
5. CONCLUSIONS
In the present study it was concluded that in contrast to the phenotypic methods, the analysis of RAPD patterns is also a useful tool for strain differentiation of A. terreus.
6. ACKNOWLEDGEMENTS
We thank the Department of Biotechnology, Vels University and our Chairman, Dr. Ishari K. Ganesh for providing us the opportunities for doing this work.
REFERENCES
- Varga, J., Toth, B., Kocscube, S., Rigo, K., Teren, J. and Kozakiewicz, Z. (2005) Evolutionary relationships among Aspergillus terreus isolates and their relatives. Antonie van Leewenhoek, 88(2), 141-150.
- Baddley, J.W., Pappas, P., Smith, A.C. and Moser, S. (2003) Epidemiology of Aspergillus terreus at a university Hospital. Journal of clinical Microbiology, 41(12), 5525-5529.
- Raper, B.K. and Fennel, D.I. (1965) The genus Aspergillus. Williams and Wilkins, Baltimore.
- Balajee, S.A., Gribskov, J.L., Hanley, E., Nickle, D. and Marr K.A. (2005) Aspergillus lentulus sp. nov. a new sibling species of A. fumigatus. Eukaryotic Cell, 4(3), 625-632.
- Hong, S.B., Shin, H.D. and Nong, J. (2008) New taxa of Neosatorya and Aspergillus in Aspergillus section Fumigatic. Antonie Van Leeuwen hoek, 93(1-2), 87-98.
- Hara, K.S., Ryu, J.H., Lie, J.T. and Roberts, G.D. (1989). Disseminated Aspergillus terreus infection in immunocompound hosts. Mayo Clinic Proceedings, 64, 770-775.
- Kransinki, K., Hotzman, R.S., Hanna, B., Greco, M.A., Graft, M. and Bhopal, M. (1985) Nosocomial fungal infection during hospital renovation. Infectious Control, 6(7), 278-282.
- Widerhold, N.P., Lewis R.E. and Kontoyiamnis D.D. (2003) Invasive aspergillosis in patients with hemotologic malignancies. Pharmacotheraphy, 23(12), 1592- 1610.
- Bitle, J., Marcfhetti, O. and Calandra, T. (2005) Changing face of health—case associated fungal infections. Current Opinion in Infectious Diseases, 18(4), 314-319.
- Rath, P.M., Kamphoff, S. and Ansorg, R. (1999) Value of different methods for the characterization of Aspergillus terreus strains. Journal of Medical Microbiology, 48(2), 161-166.
- Watson, M.W., Lambden and Clarke, I. 1991. Genetic diversity and amplification of the chlamydial protein gene. Journal of Clinical Microbiology, 29(6), 1188- 1193.
- Aubin, J.T., Collandre, D., Candotti, D., Ingrand, C., Rouzioux and Muraux J.M. (1991) Several groups among human Herpes virus, six strains can be distinguished by Southern blotting and PCR. Journal of Clinical Microbiology, 29(2), 367-372.
- Welsh, J. and Mc Clelland, M. (1990) Fingerprinting genomes using PCR with arbitrary primers. Nucleic Acids Research, 18(24), 7213-7218.
- Williams, J.G.K., Kubelik, A.R., Livak, K.J., Rafalski, J.A. and Tingey, S.V. (1990) DNA polymorphism amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Research, 18(22), 6531-653.
- Crowhurst, R.N., Hawthorne, B.T., Rikkerink, E.H.A. and Templeton, M.D. (1991) Differentiation of Fusarium solani F.sp. Cucurbitae races 1 and 2 by Random amplification of polymorphic DNA. Currents in Genetics, 20, 391-396.
- Loudon, K.W., Burnie, J.P., Coke, A.P. and Matthews, R.C. (1993) Application of polymerase chain reaction to fingerprinting Aspergillus fumigatus by random amplification of polymorphic DNA. Journal of clinical Microbiology, 31, 1117-1121.
- Rafalski, J.A and Tingey, S.V. (1993). Genetic diagnostics in plant breeding: RAPDs, microsattelites and machines. Trends in Genetics, 9(8), 275-280.
- Williams, J.K., Reiter, R.S., Young, R.M. and Scolnik (1993) Genetic mapping of mutations using phenotypic pools and mapped RAPD markers. Nucleic Acid Research, 21(11), 2697-2702.
- Yoon, C. and Glacose, D.A. (1993) Association of random amplified polymorphic DNA markers with stromatal type in Hypoxylon trancatum. Mycologia, 85, 369-380.
- Goodwin, P.H and Annis, S.L. (1991) Rapid identification of genetic variation and Pathotype of Leptosphaeria masculans by random amplification of polymorphic DNA assay. Applied Environmental Microbiology, 57(9), 2482-2486.
- Kurtzman, C.P. (1985) Molecular taxonomy of the fungi, In: Bennett, I.W. and Lasuse, L.L. Eds., Gene Manipulation in Fungi, Academic press, Orlando, 35-63.
- Saghai-Maroof, M.A., Soliman, K.M., Jorgensen, R.A. and Allard R.W. (1984) Fungal DNA isolation.
- Birch, M., Anderson M.J. and Denning D.W. (1995) Molecular typing of Aspergillus species by random amplification of polymorphic DNA. European Journal of Clinical Microbiology, 18(11), 838-841.
- Symones, F., Boucher, J.P., Heinemann, S. and Norland, N. (2000) Molecular typing of Aspergillus terreus isolates by Random amplification of polymorphic DNA. Journal of Hospital Infection, 44(4), 273-280.
- Florl, L.C., Grif, K. and Kontoyiannis, D. (2007) Molecular typing of Aspergillus terreus isolates collected in Houston, Texas and Innsbruck, Austria: Evidence of great genetic diversity. Journal of Clinical Microbiology, 45(8), 2686-2690.

