Agricultural Sciences
Vol.3 No.3(2012), Article ID:19047,8 pages DOI:10.4236/as.2012.33049
The disease reactions of heirloom bell pepper “California Wonder” to Phytophthora capsici
![]()
1Horticulture Department, University of Georgia Tifton Campus, Tifton, USA; *Corresponding Author: pconner@uga.edu
2Horticulture Department, University of Georgia, Athens, USA
3Plant Pathology Department, University of Georgia, Athens, USA
Received 3 January 2012; revised 21 February 2012; accepted 16 March 2012
Keywords: Pepper; Phytophthora Blight; Root Rot; Stem Blight; Foliar Blight; SSR
ABSTRACT
Greenhouse studies were performed to determine the reactions of 10 “California Wonder” (Capsicum annuum) accessions to the three forms of Phytophthora blight (root rot, stem blight and foliar blight) caused by Phytophthora capsici. Differences in root rot, stem blight and foliar blight severities among accessions were significant. The accessions consistently differentiated into two groups across the three disease syndromes. Simple sequence repeat (SSR) markers showed variability both within and between accessions of California Wonder. The variability in the responses to the three forms of Phytophthora blight does not warrant its usefulness as a standard susceptible control in studies involving the Capsicum-P. capsici pathosystem.
1. INTRODUCTION
“California Wonder” is an heirloom cultivar and one of the oldest bell pepper (Capsicum annuum L.) cultivars. It is an open-pollinated inbred line and is still available commercially and widely used by home and market growers. It has also been used as a standard cultivar [1] in various studies, including those on Phytophthora blight of pepper caused by Phytophthora capsici [2,3]. However, genetic variability among accessions of “California Wonder” has been demonstrated through random amplified polymorphic DNA (RAPD) analysis [1].
P. capsici causes multiple disease syndromes in pepper such as root rot, foliar blight and stem blight. All are considered different disease syndromes because each requires a single and different gene for the expression of its resistance [4]. In 2006, a study to screen the United States Department of Agriculture’s (USDA) C. annuum germplasm collection for resistance to root rot, stem blight and foliar blight was initiated at the University of Georgia’s Department of Horticulture, Tifton, GA and “California Wonder” was one of the cultivars considered as a susceptible control. In a preliminary test to determine the utility of “California Wonder” as the susceptible control, different accessions of “California Wonder” were found to vary in their reaction to the pathogen, with some demonstrating moderately high levels of resistance. “California Wonder” plants, apparently free of root rot were also observed in a field plot naturally infested with P. capsici [5]. To confirm these observations, a study was conducted to determine the reaction of “California Wonder” accessions to root rot, foliar blight, and stem blight, and to investigate the genetic diversity within and among these accessions through the use of SSR molecular markers. Information on the variability of the responses of this cultivar will determine its usefulness in future Capsicum diseases studies and in the development of resistant cultivars.
2. MATERIALS AND METHODS
2.1. Plant Material
“California Wonder” seeds were obtained from 10 different sources (Table 1) and seeds from the different sources were considered different accessions. Seeds from Ferry-Morse Seeds with different lot numbers were obtained from two different stores and were treated as different accessions. “Aristotle”, a crown rot tolerant [6] and widely-grown cultivar in the bell pepper growing areas of southern Georgia and “Camelot”, a susceptible cultivar were included as reference commercial cultivars. “Criollo de Morelos 334” (provided by Dr. P. W. Bosland, New Mexico State University) was included as the resistant control.
For root inoculation tests, seeds from each accession were sown in individual plastic cells of a 12-cell multipot bedding plant container (Compack D812, Hummert

Table 1. “California Wonder” accessions and resistant and susceptible controls used in greenhouse or field tests to determine their reactions against Phytophthora capsici.
International, St. Louis, MO). Each cell was 3.9 cm × 2.7 cm × 5.5 cm and contained Redi Earth plug and seedling mix (Sun Gro, Bellevue, WA). A replicate of each accession consisted of six cells with one seedling each. Each multipot container was then placed in 52.3 cm × 25.9 cm × 6.1 cm F1020 plastic trays with drainage holes (Hummert International, St. Louis, MO).
Two separate sets of test plants were prepared for foliar and stem inoculations. For each set, three seeds were sown in 8.9 cm square Kord green pots (Kord Products, Toronto, Canada). Each pot contained the same growing mix as mentioned in the above. After 14 days, each pot was thinned out to one seedling. The seedlings were then transferred to 18-pocket trays (Kord Products, Toronto, Canada). Each pocket tray contained six accessions with three pots per accession.
The test plants were maintained in the greenhouse, watered twice daily and fertilized twice a week with water-soluble Miracle-Gro all purpose plant food (24N-6P- 16 K) (Scotts Miracle-Gro Products, Inc., Marysville, OH) at the rate indicated in the label. The greenhouse air temperature had a diurnal range of 13˚C to 30˚C.
2.2. Inoculum Preparation and Inoculation Methods
Three virulent isolates each from the A1 and A2 mating types of P. capsici that were collected from different growers’ fields in Tift County, GA were used in the greenhouse inoculation tests. A mixture of zoospores from these isolates was used to inoculate the test plants. The zoospores were produced aseptically by transferring 10 agar plugs from the advancing portion of 5-day-old cultures (25˚C, under dark condition) of P. capsici in 5% (v/v) clarified V8 juice agar [7] to each 100 × 15 mm Petri dish and 10 ml of clarified V8 juice were added thereafter. After 24 h of incubation at 25˚C under dark condition, the V8 juice in each plate was replaced with 10 ml sterile mineral salt solution (MSS) [7] and incubated at 20˚C, 30 cm under two fluorescent lights (cool white, 20 W, 35 µmol∙m–2∙s–1) for 24 h. The MSS from each plate was then replaced with the same volume of fresh MSS and allowed to incubate for three more days.
Zoospores from each isolate were harvested separately. Zoospores were harvested by discarding the MSS, after which each plate was washed with 10 ml of sterile distilled water. After the second washing, 10 ml of sterile distilled water was added to each plate and placed in a refrigerator (1.3˚C) for 45 min. The plates were warmed on top of a laboratory bench and monitored for zoospore release. The zoospore suspension from each Petri dish was transferred to a 100-ml graduated cylinder and left undisturbed for 5 min. The upper 10 ml of the zoospore suspension was pipetted out and transferred to a 15-ml conical centrifuge. The tube was then inverted gently 2 - 3 times to distribute the zoospores in the suspension. One ml of the suspension was transferred to a 2-ml microcentrifuge tube with flat cap and vortexed for 90 sec to encyst the zoospores. The zoospore concentration was determined by using a hemacytometer and adjusted to required inoculum densities.
2.3. Root Inoculation
Two hours prior to inoculation, the trays containing 14-day-old pepper seedlings were placed in water-filled F1020 trays without drainage holes to saturate the roots. Five ml zoospore suspension (2000 zoospores per ml) was then delivered to each cell [8] by using an automatic dispenser (Finpipette, Vantaa, Finland). The saturated condition was maintained for another 48 hrs and disease evaluation was performed 14 days after inoculation. The plants were evaluated based on a 10-point scale [8]: 0 = no response, vigorous, healthy; 3 = brown roots, slight stunting, very small lesions on stems; 5 = brown roots, small lesions on stems, lower leaves wilted, stunted plants; 7 = brown roots, large lesions on stems, girdling, whole plant wilted, and stunted; 9 = death. Even numbers corresponded to intermediate responses.
The root inoculation tests were performed in the greenhouse in a randomized complete block design with five replications. Each replicate consisted of six seedlings and the test was repeated once.
2.4. Foliar Inoculation
Foliar inoculations were performed in 6-wk-old seedlings and arranged in a randomized complete block design with five replications under greenhouse conditions. Each replicate consisted of three seedlings and the test was repeated once. A volume of 100 µl zoospore suspension (5000 zoospores per ml) was placed on the upper surface of a partially expanded leaf [9]. The inoculated seedlings were placed inside a humidity chamber made of 0.1 mm plastic sheets that were also used to cover the mesh opening of the greenhouse benches. A home-use humidifier provided a relative humidity of 100% at night. Foliar blight assessment was performed 14 days after inoculation by using a 0 - 5 foliar blight severity scale [10]: 0 = no visible symptoms, 1 = small circular or irregular spots on upper leaves, 2 = leaf-enlarged symptoms with brownish lesions beginning to appear on stems and <25% of the plant wilted, 3 = leaves defoliated with lesions on leaves covering half of a leaf and 25% - 50% of the plant wilted moderately, 4 = leaves defoliated or dried, with rapidly expanding stem lesions and 50% - 70% of the plant wilted severely, and 5 = plant dead.
2.5. Stem Inoculation
Greenhouse stem inoculations were performed in a randomized complete block design with five replications. Each replicate consisted of three seedlings and the test was repeated once. At eight weeks after planting, stems were tied with sterile absorbent cotton yarn (3 mm in diameter) [4] at two different places 2 - 3 cm apart. One hour before inoculation, the yarns were saturated with sterile distilled water and 45 µl of zoospore suspension (50,000 zoospores per ml) was placed on the upper yarn. Stem blight assessment was performed 14 days after inoculation by using a 0 - 5 stem blight severity scale [10]: 0 = no visible symptoms, 1 = brownish lesion at the inoculation point, 2 = stem lesion extending 1 - 3 cm from inoculation point, 3 = stem lesion progression up to half of the plant height, 4 = stem lesion progressing toward the shoot apex, and 5 = plant dead.
2.6. Field Test
The difference in the reaction of “California Wonder” from Burpee Seed Co. and Rupp Seeds was verified in a field test during fall of 2008 by superimposing it in an ongoing pepper varietal test in a field plot that was naturally infested with P. capsici. “Aristotle”, “Camelot”, and “Criollo de Morelos 334” were included as reference cultivars. The field test was conducted at the Black Shank Farm of the University of Georgia’s Coastal Plain Experiment Station, Tifton, GA. The soil type was fuquay loamy sand (88% sand, 8% silt, 4% clay; pH 5.5 - 6.0; 2% organic matter; loamy, siliceous thermic Arenic Plinthic Paleudults). Raised soil beds measuring 4.6 m long, 0.8 m wide and 0.3 m high were formed by using a commercial tractor-drawn bedder (Kennco’s Blue Bedder Model BL4000BK, Kennco, Mfg., Ruskin, FL). Drip irrigation tapes (Aqua-Traxxs Premium drip tape, Toro Ag Irrigation Business, El Cajon, CA) were installed at a depth of 2.5 cm at the same time with white low-density polyethylene (LDPE) plastic mulch (50 mm thickness). The pepper accessions or cultivars were arranged in a randomized complete block design with five replications and each replicate consisted of a total of 32 plants for each accession. Seven-week-old seedlings were transplanted on double rows with the distance between rows and between plants within a row at 30 cm. All treatments were fertigated once a week (130 kg/ha of nitrogen, 48 kg/ha of P2O5, 27 kg/ha K2O) by injecting the liquid fertilizer into the drip irrigation system. Insects and weeds were controlled according to the practices recommended by the University of Georgia Cooperative Extension Services. All treatments were evaluated at maturity for root rot severity by estimating the percentage of root rot system with root discoloration or browning; and for plant survival which is the percentage of plants that survived the disease.
2.7. Molecular Marker Analysis
Seeds were planted is seedling trays at the University of Georgia South Milledge Greenhouses in Athens, GA. Three leaf disks were collected from 3 or 4 plants of each cultivar/accession using the lid of a microfuge tube as a punch. Samples were immediately put in liquid nitrogen and stored at –80˚C until processing. Samples were ground using 5 mm stainless steel beads (Qiagen, Inc, Valencia, CA) and the TissueLyser system (Qiagen). After grinding, DNA was extracted using GenElute Plant Genome DNA miniprep kit (Sigma-Aldrich Inc., St. Louis, Mo.). DNA concentration was measured on a NanoDrop (Model ND-8000, Thermo Fisher Scientific Inc., Waltham, MA) and diluted if necessary.
Twenty-two previously published SSR primer pairs [11-13] were tested as described previously [14] and 9 were found to be polymorphic within or between California Wonder seed lots (Table 2).
PowerMarker Version 3.25 [15] was used to calculate frequency based distances [16] and draw a tree based on the unweighted pair group method with arithmetic mean (UPGMA) [17] that was visualized in Tree View [18]. Powermarker was also used to calculate heterozygosity [19] and polymorphism information content (PIC) [20], while genotype diversity was calculated as described
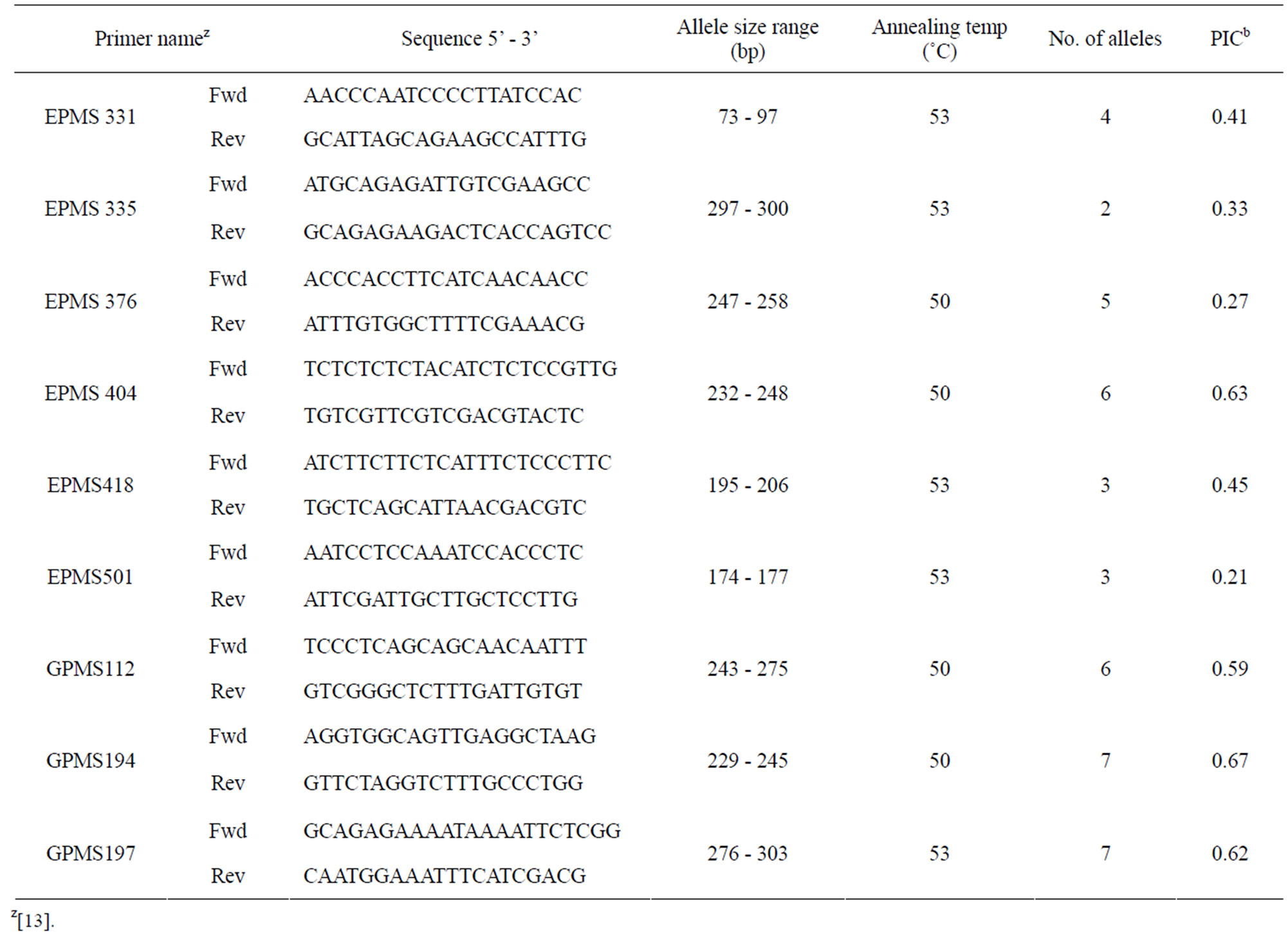
Table 2. The nine SSR primer pairs that showed polymorphic allele sizes within or between “California Wonder” seed sources.
by Tommasini et al. [21].
2.8. Data Analysis
Root rot, foliar blight, and stem blight severity data from the five replications and two experiments were combined. All are ordinal data and were therefore analyzed by one-way layout nonparametric tests [22] using SAS 9.1 software (SAS Institute, Cary, NC). For each disease syndrome, the rank assigned to each observation was determined by PROC RANK. The ranks were then sorted (PROC SORT) by accession and replication, after which, one-way analysis of the marginal effects, computation of the lsmeans for each accession, and determination of the ANOVA-type statistics were performed by using PROC MIXED. Lastly, the ranks were used to estimate the corresponding relative treatment effects and their confidence intervals (95%) for each accession by using LD_CI macro [23]. The relative treatment effects whose values are always between 0 and 1, are directly related to the values of the observations. Hence, a smaller relative treatment effects for an accession indicate smaller values for either root rot severity, foliar blight severity or stem blight severity. The severity values between two accessions are significantly different from each other if their 95% confidence intervals for the relative treatment effects do not overlap. The range and median of the severity rating for each accession were also determined.
Data on root discoloration (%) and plant survival (%) were analyzed by analysis of variance (ANOVA). Mean separation was performed by the least significant difference test (LSD, P = 0.05) after a significant F-test in SAS ver. 9.1 (SAS Institute, Cary, NC).
3. RESULTS AND DISCUSSION
Range, median, mean rank, relative treatments effects and the confidence intervals for the relative treatment effects of each accession are presented separately for root rot, stem blight, and foliar blight severities. Based on their confidence intervals for relative treatment effects, the 10 accessions could be differentiated into two groups (Table 3). The first group consisted of accessions whose root reactions encompassed the 10-point severity rating scale for root rot and their median root rot severity
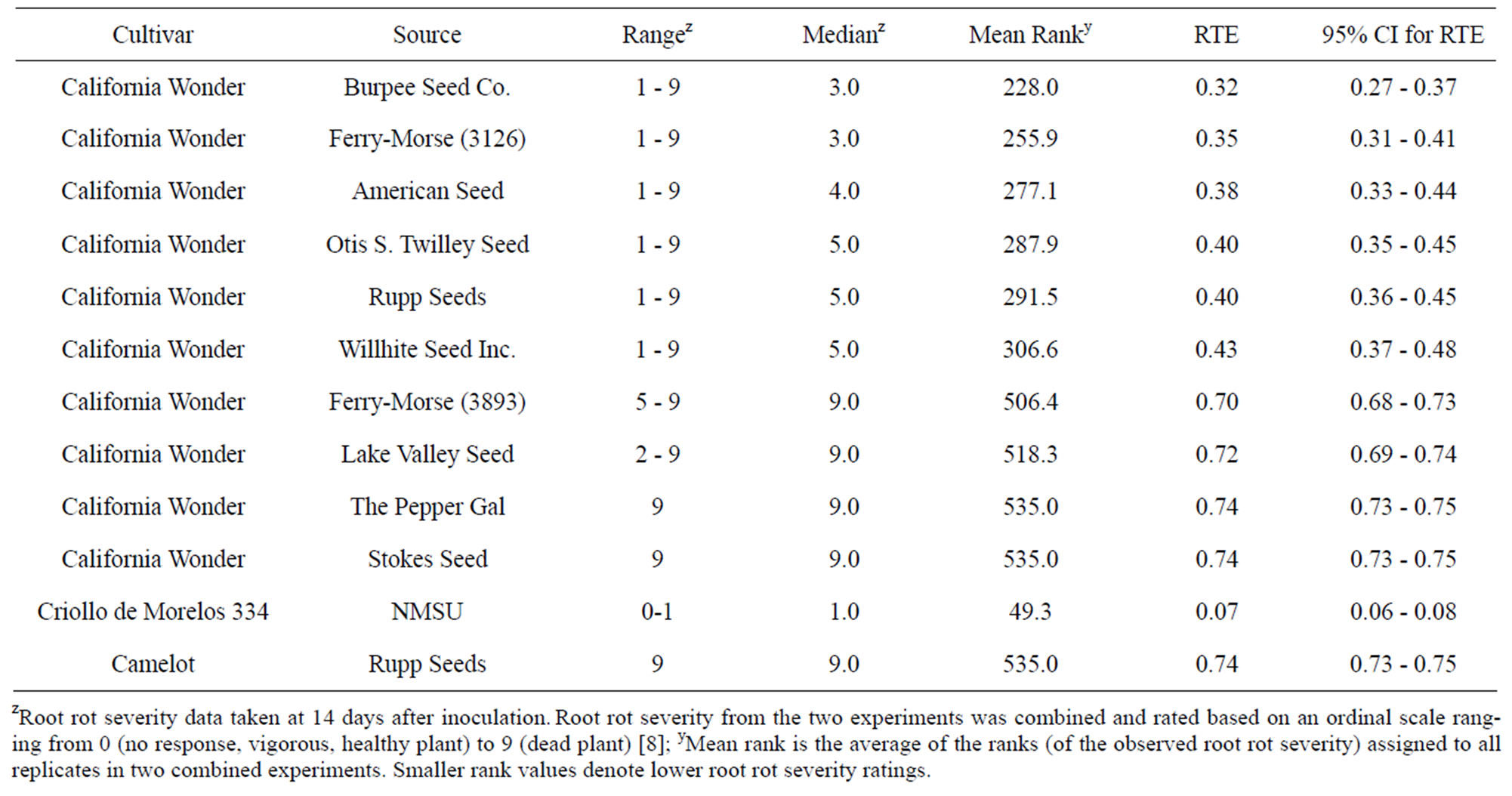
Table 3. Range, median, mean rank, relative treatment effects, and confidence intervals (CI) for relative treatment effects (RTE) of root rot severity on “California Wonder” accessions artificially inoculated with Phytophthora capsici.
ranged from 3 - 5. The second group consisted of four accessions with each having a median root rot severity of 9 and two of these accessions (The Pepper Gal and Stokes Seeds) exhibited consistent root rot severity ratings of 9. Differences among accessions within a group were not significant, but differences between accessions coming from the two different groups were significant. Root rot from the latter group was as severe as that observed on “Camelot” while “Criollo de Morelos 334” had less root rot than all “California Wonder” accessions.
Stem blight severity differentiated the same two groups of accessions with differing levels of susceptibility (Table 4). The first group of six accessions had median stem blight severities ranging from 1 - 2 and the second group with a median stem blight severity of 5. Differences in stem blight severity among accessions within a group were not significant, but differences between accessions coming from the two different groups were significant. The stem blight severity in all “California Wonder” accessions was significantly higher than the stem blight severity in “Criollo de Morelos 334”. On the other hand, the stem blight severities in the accessions from the more susceptible second group were not significantly different from that of “Camelot”.
Lastly, the 10 “California Wonder” accessions differentiated into two groups based on foliar blight severity (Table 5). The first group exhibited a median of 0 with observed foliar blight severity ratings of 0 - 5 (except Ferry-Morse 3126) and the second group with medians of 5 and observed foliar blight severity ratings of 3 - 5. Accessions from Stokes Seed and The Pepper Gal had a consistent foliar blight severity rating of 5. Differences among accessions within a group were not significant, but differences between accessions coming from the two different groups were significant. The only “California Wonder” accession whose foliar blight severity was not significantly more than “Criollo de Morelos 334” was the accession from Rupp Seeds. The foliar blight severities of the accessions from the second group were not significantly different from that of “Camelot”.
Field tests of two “California Wonder” accessions from the more resistant group were undertaken to compare to the standard resistant control of “Criollo de Morelos 334” and the popular cultivar “Aristotle” which demonstrates some tolerance to the disease. The differences in root rot, stem blight, and foliar blight severities between accessions from Rupp Seeds and Burpee Seed Co. were not significant in greenhouse tests, but field tests demonstrated significant differences in root discoloration and plant survival (Table 6). Root discoloration and plant survival of the Burpee Seed Co. accession were not significantly different from those of “Aristotle”, and both accessions were significantly more resistant to the P. capsisci than the susceptible control “Camelot”.
SSR markers were used to investigate the heterozygosity and diversity of the “California Wonder” accessions. Nine of the 22 primers used for amplification showed polymorphisms in allele size within or between California Wonder seed sources. The PIC of these primers ranged from 0.21 to 0.67 (Table 2). As expected the
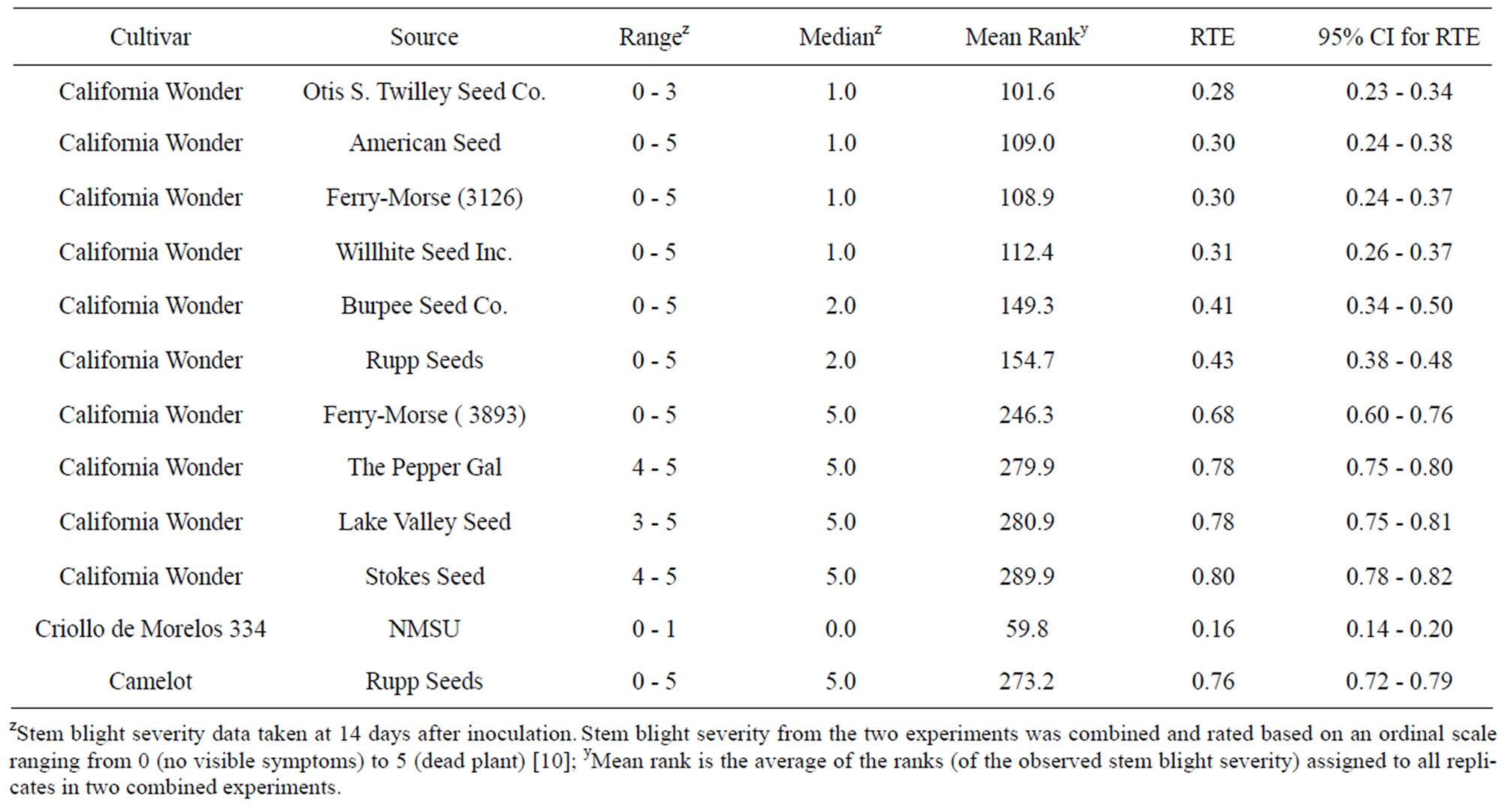
Table 4. Range, median, mean rank, relative treatment effects, and confidence intervals (CI) for relative treatment effects (RTE) of stem blight severity on “California Wonder” accessions artificially inoculated with Phytophthora capsici.
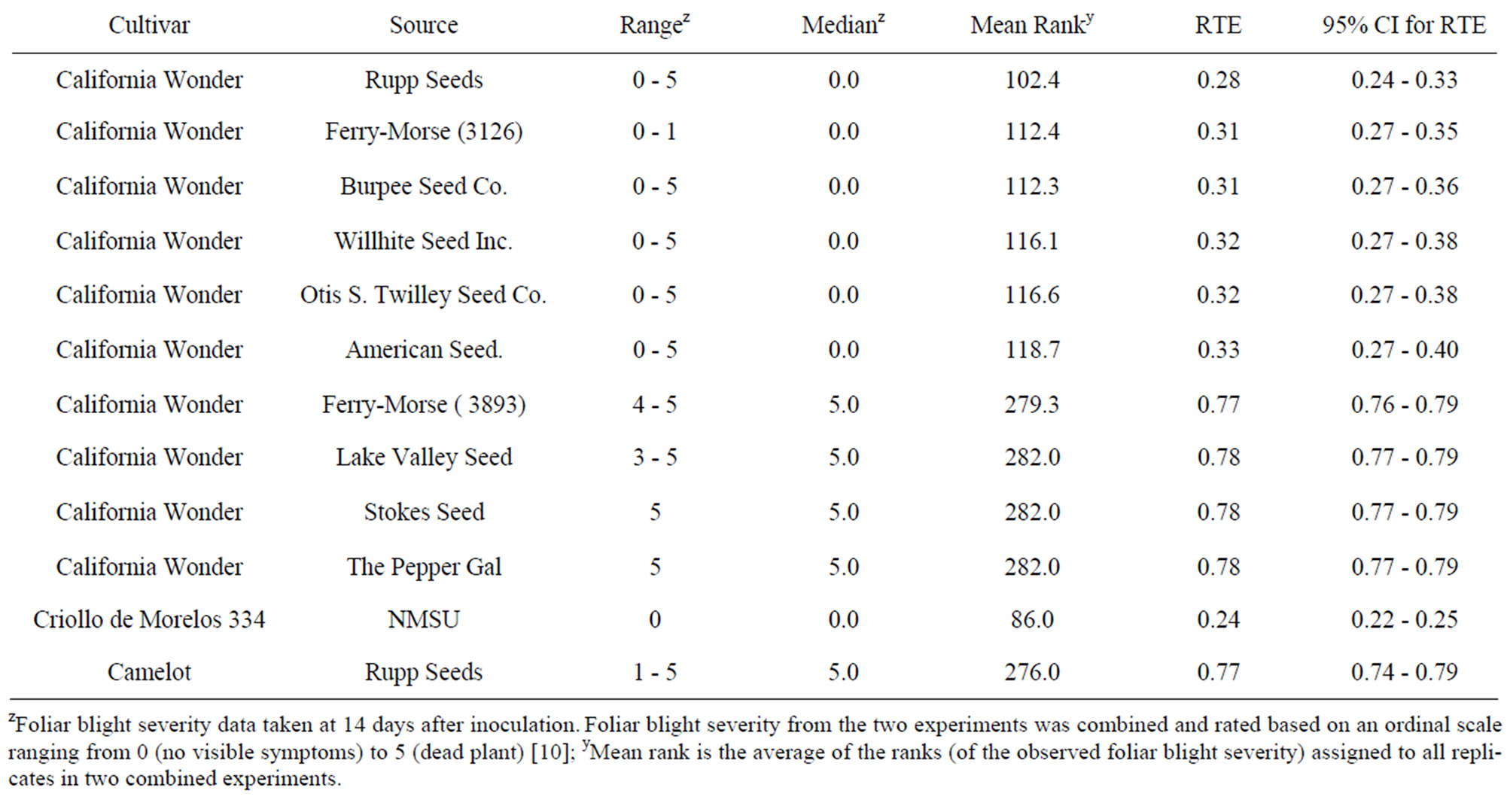
Table 5. Range, median, mean rank, relative treatment effects, and confidence intervals (CI) for relative treatment effects (RTE) of foliar blight severity on “California Wonder” accessions artificially inoculated with Phytophthora capsici.
hybrid cultivars Aristotle and Camelot showed high heterozygosity and low genotype diversity. Genotype diversity is a measure of uniformity of individuals within a cultivar/accession, and ranges from 0 to 1 with zero indicating no variation within the cultivar/accession [21]. “California Wonder” accessions demonstrated lower heterozygosity (Table 7), as would be expected from an open pollinated cultivar of a self pollinating species. However, three sources had relatively high heterozygosity, indicating that outcrossing does occur. The high
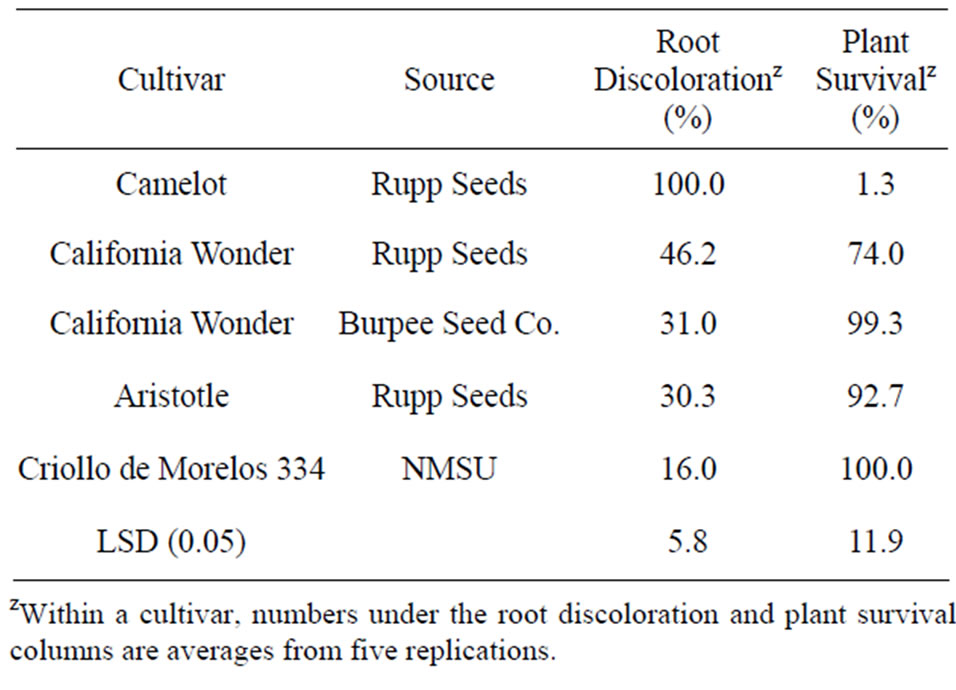
Table 6. Root discoloration and plant survival of “California Wonder” accessions and other selected pepper cultivars planted in a field naturally infested with Phytophthora capsici.
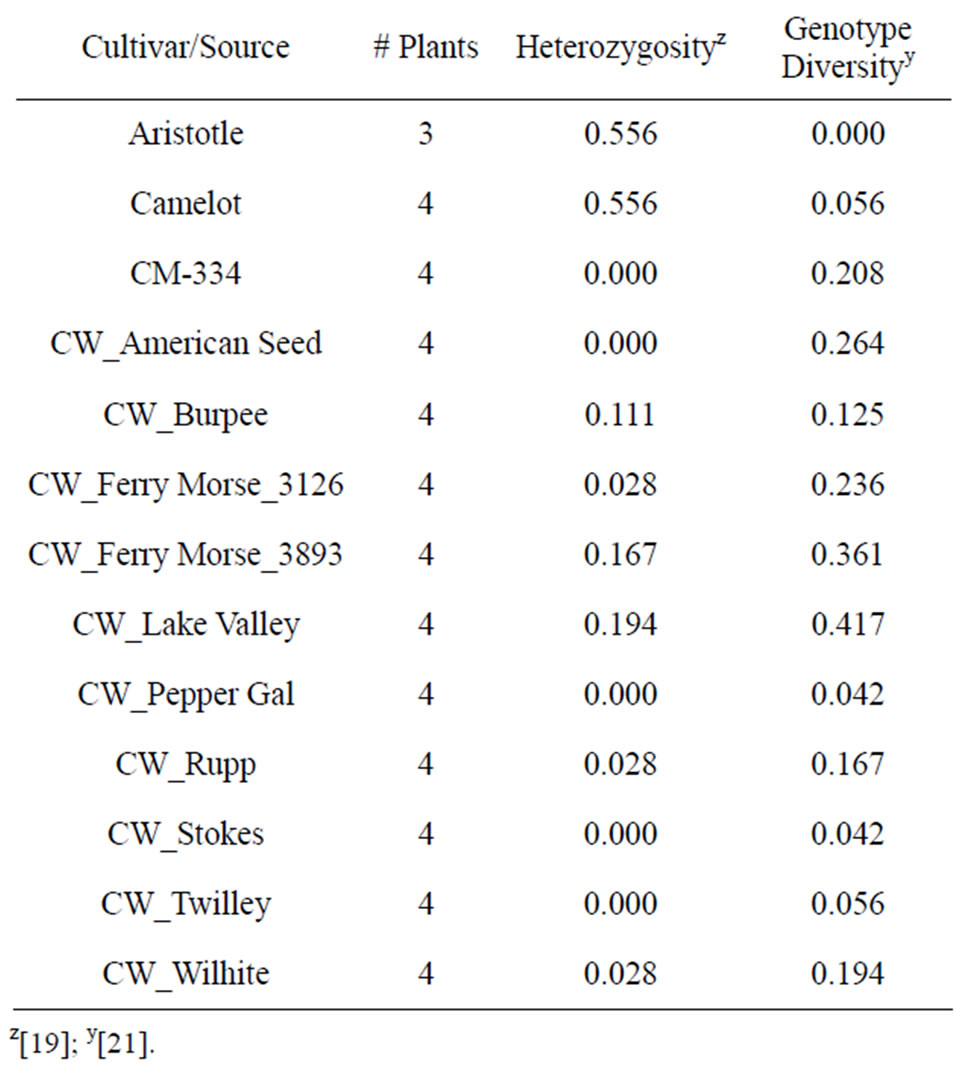
Table 7. Heterozygosity and genotype diversity calculated from SSR analysis of the cultivars and seed sources used in this study.
genotype diversity of many accessions indicates signifycant variation within the seed sources of supposedly pure lines.
A phenogram of SSR data indicate that the “California Wonder” accessions fall into two main clades (Figure 1). All accessions could be differentiated except for Pepper Gal and Stokes. These two seed sources also showed no heterozygosity and low genotype diversity, and may represent a single source as Stokes is a large supplier of seeds. The two clades nearly match the two groupings demonstrated by the resistance tests. The single exception is the “Californing Wonder” Rupp accession which fell into the more resistant grouping in the resistance tests, but was grouped with the more susceptible accessions by the SSR data.
The variability in “California Wonder” could be attributed to a number of sources [1]: 1) since “California Wonder” is an old introduction, there is no strict control over its seed production; 2) a small sample of an originnally genetically diversified population becomes the source of a new population, but contains only a small sample of the genetic variability found in the original Population; 3) modern sources of “California Wonder” may not be related to an original source at all due to seed mixing, mislabeling, or misrepresentation. For example, some packets of seeds of “California Wonder” labeled as “CalWonder” were observed. “CalWonder” could be easily confused with “Calwonder”, which is a bell pepper cultivar different from “California Wonder” [24] and such confusion could lead to mislabeling. 4) cross-pollination with other Capsicum species (2% - 92% occurrence) could be another source of the variability [25]. The presence of heterozygosity as indicated by SSR profiles in many of the seed sources indicate that precautions should be taken to prevent cross pollination in seed production fields.
The 10 accessions studied consistently differentiated into two groups and the accessions which compose each group were consistent across the three disease syndromes. Furthermore, the data indicated an overall variability among “California Wonder” accessions from different sources in terms of their responses to the three disease syndromes. This variability in the observations could be attributed to the high level of genetic variability in cultivar “California Wonder” as previously reported [1]. It was also indicated that the genetic variability in cultivar “California Wonder” existed for the most part, among seed sources rather than within seed sources [1], while our study reported here provided evidence that variability in disease severity was observed within seed sources.
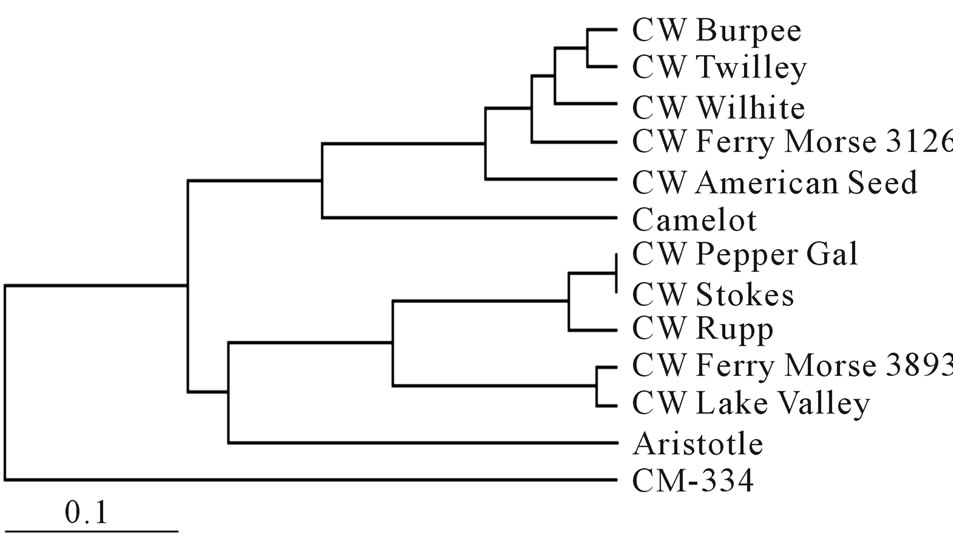
Figure 1. Phenogram based on frequency based distances [16] and the unweighted pair group method with arithmetic mean (UPGMA) of the cultivars and seed sources used in this study.
With this information, a mass screening for resistance to P. capsici would be rendered unrealizable if “California Wonder” is used as the standard susceptible control. Specifically, it could provide an unreliable benchmark on the prevailing disease pressure during an experiment as well as disease pressure from one experiment to another.
REFERENCES
- Votava, E.J. and Bosland, P.W. (2002) A cultivar by any other name: Genetic variability in heirloom bell pepper “California Wonder”. HortScience, 37, 1100-1102.
- Barksdale, T., Papavisas, G. and Johnston, S. (1984) Resistance to foliar blight and crown rot of pepper caused by Phytophthora capsici. Plant Disease, 68, 506-509.
- Saini, S.S. and Sharma, P.P. (1978) Inheritance of resistance to fruit rot (Phytophthora capsici Leon.) and induction of resistance in bell pepper (Capsicum annuum L.). Euphytica, 27, 721-723. doi:10.1007/BF00023707
- Sy, O., Bosland, P.W. and Steiner, R. (2005) Inheritance of Phytophthora stem blight resistance as compared to Phytophthora root rot and Phytophthora foliar blight resistance in Capsicum annuum L. Journal of the American Society of Horticultural Science, 130, 75-78.
- Smith, P.G. and Minges, P.W. (1951) Resistance of strains of California Wonder pepper to Phytophthora root rot. Plant Disease Reporter, 35, 509.
- Kline, W.L., Wyenandt, C.A., Ward, D.L., Sudal, J.F. and Maxwell, N.L. (2011) Evaluation of six nitrogen fertility programs on marketable yield and development of skin separation in bell pepper fruit. HortTechnology, 21, 323- 328.
- Kuhajek, J.M., Jeffers, S.N., Slattery, M. and Wedge, D.E. (2003) A rapid microbioassay for discovery of novel fungicides for Phytophthora spp. Phytopathology, 93, 46-53. doi:10.1094/PHYTO.2003.93.1.46
- Bosland, P.W. and Lindsey, D.L. (1991) A seedling screen for Phytophthora root rot of pepper, Capsicum annuum. Plant Disease, 75, 1048-1050. doi:10.1094/PD-75-1048
- Alcantara, T.P. and Bosland, P.W. (1994) An inexpensive disease screening technique for foliar blight of chile pepper seedlings. HortScience, 29, 1182-1183.
- Kim, Y.J., Hwang, B.K. and Park, K.W. (1989) Expression of age-related resistance in pepper plants infected with Phytophthora capsici. Plant Disease, 73, 745-747. doi:10.1094/PD-73-0745
- Lee, J.M., Nahm, S.H., Kim, Y.M. and Kim, B.D. (2004) Characterization and molecular genetic mapping of microsatellite loci in pepper. Theoretical and Applied Genetics, 108, 619-627. doi:10.1007/s00122-003-1467-x
- Minamiyama, Y., Tsuro, M., Kubo, T. and Hirai, M. (2007) QTL analysis for resistance to Phytophthora capsici in pepper using a high density SSR-based map. Breeding Science, 57, 129-134. doi:10.1270/jsbbs.57.129
- Nagy, I., Stágel, A., Sasvári, Z., Röder, M. and Ganal, M. (2007) Development, characterization, and transferability to other Solanaceae of microsatellite markers in pepper (Capsicum annuum L.). Genome, 50, 668-688. doi:10.1139/G07-047
- McGregor, C., Waters, V., Nambeesan, S., MacLean, D., Candole, B.L. and Conner P. (2011) Genotypic and phenotypic variation among pepper accessions resistant to Phytophthora capsici. HortScience, 46, 1235-1240.
- Liu, K. and Muse, S.V. (2005) PowerMarker: An integrated analysis environment for genetic marker analysis. Bioinformatics, 21, 2128-2129. doi:10.1093/bioinformatics/bti282
- Nei, M., Tajima, F. and Tateno, Y. (1983) Accuracy of estimated phylogenetic trees from molecular data. Journal of Molecular Evolution, 19, 153-170. doi:10.1007/BF02300753
- Sneath, P.H.A. and Sokal, R.R. (1973) Numerical taxonomy. W. H. Freeman, San Francisco.
- Page, R.D.M. (1996) Tree view: An application to display phylogenetic trees on personal computers. Computer Applications in the Biosciences: CABIOS, 12, 357-358.
- Weir, B.S. (1996) Genetic data analysis II. Sinauer Associates, Inc., Sunderland.
- Botstein, D., White, R.L., Skolnick, M. and Davis, R.W. (1980) Construction of a genetic linkage map in man using restriction fragment length polymorphisms. American Journal of Human Genetics, 32, 314-331.
- Tommasini, L., Batley, J., Arnold, G.M., Cooke, R.J., Donini, P., Lee, D., Law, J.R., Lowe, C., Moule, C., Trick, M. and Edwards, K.J. (2003) The development of multiplex simple sequence repeat (SSR) markers to complement distinctness, uniformity and stability testing of rape (Brassica napus L.) varieties. Theoretical and Applied Genetics, 106, 1091-1101.
- Shah, D.A. and Madden, L.V. (2004) Nonparametric analysis of ordinal data in designed factorial experiments. Phytopathology, 94, 33-43. doi:10.1094/PHYTO.2004.94.1.33
- Brunner, E.S., Domhof, S. and Langer, F. (2002) Nonparametric analysis of longitudinal data in factorial experiments. John Wiley & Sons, Inc., New York.
- Whealy, K. (2004) Garden seed inventory: Inventory of seed catalogs listing of all non-hybrid vegetable seeds available in the United States. 6th Edition, Seed Savers, Inc., Iowa.
- Pickersgill, B. (1997) Genetic resources and breeding of Capsicum spp. Euphytica, 96, 129-133. doi:10.1023/A:1002913228101

