Food and Nutrition Sciences
Vol.5 No.1(2014), Article ID:42013,7 pages DOI:10.4236/fns.2014.51012
NMR-Metabolomic Study on Monocultivar and Blend Salento EVOOs including Some from Secular Olive Trees
Di.S.Te.B.A., Prov.le Lecce-Monteroni, University of Salento, Lecce, Italy.
Email: laura.delcoco@unisalento.it, sandra.depascali@unisalento.it, *fp.fanizzi@unisalento.it
Received November 29th, 2013; revised December 29th, 2013; accepted January 6th, 2014
ABSTRACT
The aim of the present work has been to characterize, by NMR-based metabolic profiling, extravirgin olive oils (EVOOs) from a subarea (Salento) of Apulia, leader EVOO producer among the Italian regions. According to the European Union (EU) definition, Protected Designation of Origin (PDO) products are mostly closely linked to the concept of terroir due to the place of origin, climate and local know-how. Moreover, the authenticity and traceability of several products such as olive oils with specific geographical origin require to be preserved by analytical methods. In this regard, about a hundred EVOO samples (monovarietal and blend samples, cultivars Ogliarola Salentina and Cellina di Nardò, basis of “Terra d’Otranto” PDO, campaign 2012-2013) were therefore analyzed by 1H NMR spectroscopy and multivariate statistical analysis. Both unsupervised (PCA) and supervised (OPLS-DA) statistical analyses allowed differentiation of monocultivar oils and blends characterization. Other features such as the age of the trees (young, <100 years, and secular olive trees, >100 years) could also be investigated. Cellina samples showed a higher content of aldehydic and phenolic compounds, while Ogliarola samples were characterized by NMR signals in the range of δH 6.5 - 5.6, which could be ascribed to higher carotenoids content. Higher polyphenols and polyunsaturated fatty acid content were also found in young over secular tree EVOOs.
Keywords:Cellina Di Nardò; Ogliarola Salentina; EVOOs; 1H NMR Spectroscopy; Multivariate Statistical Analysis, MVA; Principal Component Analysis, PCA; Orthogonal Partial Least Squares Discriminant Analysis (OPLS-DA)
1. Introduction
Olea Europaea L. (family, Oleaceae), commonly known as “olive”, is among the oldest known cultivated trees in the world and in particular the most abundant in the Mediterranean basin. The health beneficial effects of olive fruit and oil, in particular of extravirgin olive oils (EVOOs), are well known and documented [1]. Olive oil chemical composition has already been investigated by means of several techniques, such as gas chromatography (GC-MS) and high throughput analytical methods as well as nuclear magnetic resonance (NMR) and/or mass spectrometry (MS). These last are the typical approaches used in metabolomic research to evaluate the quality of food and beverages such as oil and wine [2-]">4]. In particular, 1H NMR spectroscopy, coupled with chemometrics studies, allows the identification of EVOOs related to specific production areas and/or olive cultivars [5,]">6]. The absolute concentrations and relative proportions of olive oil’s minor components are characteristic of each oil, and may be used for production area and/or potential adulterations identification purposes. The fine composition of olive oil and therefore its sensory characteristics are influenced by several factors such as climate and soil conditions, agricultural practices, as well as the nature of the cultivar used for its production. Recognition of the influence of these factors has led researchers to study the oil obtained from the same cultivar over the course of several years and in different geographical areas [7,8,]. In this study, EVOOs from Salento, a small geographical area that stretches between the Adriatic and Jonian seas of Apulia region in Italy, were studied by a NMR-based metabolomic approach. Ogliarola di Lecce (also known as Salentina) and Cellina di Nardò are the most popular olive cultivars in the Jonian-Salentina area and they are the basis of “Terra d’Otranto” Protected Designation of Origin (PDO) production (alone or in combination at least for 60% [9]). The production area named “Terra d’Otranto” includes the entire territory of the province of Lecce and part of the provinces of Brindisi and Taranto. The PDO EVOO “Terra d’Otranto” is characterized by green to yellow color, with a high content of antioxidant and aromatic substances that is able to play a very important role from both a nutritional and organoleptic point of view [10]. In this regard, 93 monovarietal and blend EVOOs from typical Salento (Apulia) cultivars (campaign 2012-2013, Cellina and Ogliarola, essentially) were analyzed by 1H NMR spectroscopy and multivariate statistical analysis within the project PIF Filiera Olivicola 100% Pugliese Jonico-Salentina [11].
2. Materials and Methods
2.1. Sample Collection
93 authentic EVOO samples were collected during the harvesting period 2012-2013 from different microareas of Lecce province (Le, Italy): 26 monocultivar Cellina di Nardò; 32 monocultivar Ogliarola Leccese; 35 blend Cellina/Ogliarola samples (Figure 1). The different olive oil samples were labeled as declared by farmers and also classified on the basis of the age of trees. In particular, 29 samples were classified as from “young olive trees” (<100 years) and 42 from “secular olive trees” production (secular, >100 years). Moreover, 4 EVOO samples were ascribed to “very young olive trees” production (<30 years). Samples were analyzed by 1H NMR and multivariate statistical analysis (MVA).
2.2. Chemicals
All chemical reagents for analysis were of analytical grade. CDCl3 (99.8 atom %D) and tetramethylsilane, TMS (0.03 v/v %) were purchased from Armar Chemicals (Switzwerland).
2.3. 1H NMR Spectroscopy
For NMR sample preparation ~140 mg of olive oil was dissolved in deuterated chloroform (CDCl3 with TMS as internal standard) adjusting the mass ratio of olive oil:CDCl3 to 13.5%:86.5%. 600 µL of the prepared mixture was transferred into a 5 mm NMR tube. NMR spectra were recorded on a Bruker Avance III spectrometer (Bruker, Karlsruhe, Germany), operating at 400.13 MHz for 1H observation and a temperature of 300.0 K, equipped with a BBO 5 mm direct detection probe incorporating a z axis gradient coil. NMR spectra were acquired using Topspin 2.1 (Bruker). Automated tuning and matching, locking and shimming using the standard Bruker routines ATMA, LOCK, and TopShim were used to optimize the NMR conditions. Experiments were run in automation mode after loading individual samples on a Bruker Automatic Sample Changer, (BACS-60), interfaced with the software IconNMR (Bruker). Two different 1H NMR experiments were performed for each sample: a standard one-dimensional 1H ZG NMR experiment and a one-dimensional 1H NOESYGPPS NMR pulse sequence with suppression of the strong lipid signals (20 frequencies), in order to enhance signals of minor components present in EVOOs (Bruker). Spectra were obtained by the following conditions: zg pulse program (for 1H ZG NMR) 64 K time domain, spectral width 20.5555 ppm (8223.685 Hz), p1 12.63 μs, pl1 −1.00 db, 16 repetitions; noesygpps1d.comp2 pulse program (for 1H NOESYGPPS NMR) 32 K time domain, spectral width 20.5555 ppm (8223.685 Hz), p1 12.63 μs, pl1 −1.00 db, 32 repetitions.
2.4. NMR Data Reduction and Preprocessing
NMR data were processed using Topspin 2.1 (Bruker) and visually inspected using Amix 3.9.13 (Bruker, Bios pin). 1H NMR spectra were obtained by the Fourier Transformation (FT) of the FID (Free Induction Decay), applying an exponential multiplication with a line-broadening factor of 0.3 Hz. The resulting 1H NMR spectra were manually phased and baseline corrected using the Bruker Topspin software. Chemical shifts were reported with respect to the TMS signal set at 0 ppm. 1H NMR spectra were segmented in rectangular buckets of fixed 0.04 ppm width and integrated, using the Bruker Amix software. Bucketing of 1H ZG NMR spectra (BUCKET-1) and 1H NOESYGPPS NMR spectra (BUCKET-2) were obtained within the range 10.0 - 0.5 ppm (BUCKET-1) and 10.0 - 5.6 ppm (BUCKET-2), respectively. In both cases, the spectral region between 7.60 and 6.90 ppm was discarded because of the peak due to residual protic chloroform signal at 7.24 ppm. The remaining buckets were then normalized to total area to minimize small differences due to total olive oil concentration and/or acquisition conditions among samples. A third data set named BUCKET-3 was generated combining BUCKET-1 and BUCKET-2 in one matrix (1 line per olive oil sample).
2.5. Statistical Analysis
The potential to correlate origin of authentic olive oil samples with NMR data was studied using a combination of established multivariate statistical tools, such as unsu-

Figure 1. Expansion of the extremely south east subarea (Salento) of Apulia region in Italy. In squares, the number of samples collected from each district of Salento peninsula is specified.
pervised (PCA) and supervised (PLS-DA, OPLS-DA) statistical techniques. Multivariate statistical analysis and graphics were obtained using Simca-P version 13.0.2 (Umetrics, Sweden) and different procedures were used: Principal Component Analysis (PCA), Partial Least Squares Discriminant Analysis (PLS-DA) and Orthogonal Partial Least Squares Discriminant Analysis (OPLSDA) [12]. PCA, an unsupervised pattern recognition method, was performed to examine the intrinsic variation in the data set. The Orthogonal Partial Least Squares Discriminant technique (OPLS-DA) is the most recently used for the discrimination of samples with different characteristics (such as cultivars and/or geographical origin) as shown in several recent studies of metabolomics [13,14]. OPLS-DA is a modification of the usual PLSDA method which filters out variation that is not directly related to the response. So, the further improvements made by the OPLS-DA resides in the ability to separate the portion of the variance useful for predictive purposes from the not predictive variance (which is made orthogonal). Furthermore, OPLS-DA focuses the predictive information in one component, facilitating the interpretation of spectral data. The variables used for chemometric analyses were the buckets, which represent the entire NMR spectrum, and describe all the molecules present in oils (both triglycerides and unsaponifiable fractions). The robustness and predictive ability of the OPLS-DA models for discrimination purposes were tested by cross-validation [15,16]. For this reason specific parameters indicative of the goodness of the performances of statistic models were evaluated. The R2 (cum) and Q2 (cum) are the two parameters considered for description of the soundness of the models. The former (R2) explains the total variations in the data whereas the latter (Q2) is a cross validation parameter, which indicates the predictability of the model.
3. Results and Discussion
As reported in details in the experimental section, three different bucket datasets were generated from NMR spectra: BUCKET-1 was obtained within the range 10.0 - 0.5 ppm, BUCKET-2 was obtained within the range 10.0 - 5.6 ppm and BUCKET-3 was the combination of the two previous bucket tables (taking into account only the range 5.0 - 0.5 ppm originating from BUCKET-1 and the whole BUCKET-2). For every bucket table built, multivariate statistical analyses (unsupervised, PCA, and supervised methods, PLS-DA and OPLS-DA) were applied. PCA allowed to obtain a general overview of the natural data grouping. The original datasets were rearranged in a new multivariate coordinate space where the dimensions are ordered by decreasing explained variance in the data. The principal components were displayed as a set of scores (t), which highlight clustering or outliers, and a set of loadings (p), which highlight the influence of input variables on t. In all the models studied, PCA did not show significant trends or clustering with the exception of t2/t4 PCA score plot (Figure 2) for BUCKET-3. It should be noted that this combined bucket table gave also the best results, reported and discussed below, for further statistical analyses. PCA on the monocultivar Cellina and Ogliarola samples with 7 components gave R2 = 0.755 and Q2 = 0.525, a weak model but useful for visualization of data. Looking at the score plot a certain degree of
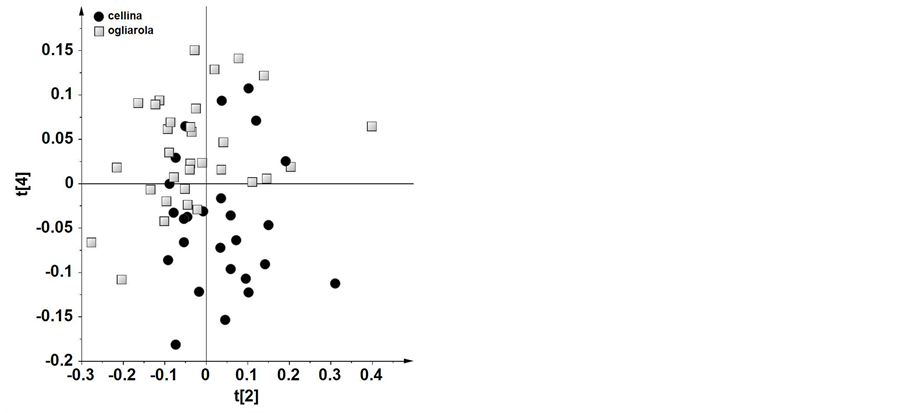
Figure 2. PCA (t2/t4) scoreplot for monovarietal Cellina and Ogliarola samples (7 components gives R2 = 0.755 and Q2 = 0.525).
overlap was observed between the two cultivars. Nevertheless, by examining the loadings of the original bucket variables, Cellina samples were characterized by variables with negative loadings on t4. In particular, signals at 9.64 and 6.64 ppm were attributed to aldehydic and phenolic compounds, respectively. On the contrary, Ogliarola samples were characterized by positive loadings on t4 of signals in the range of δH 6.5 - 5.6, which could be ascribed to carotenoids. In order to improve the separation among oils based on maximizing covariance between the measured data (X) and the response variable (Y), OPLS-DA models were also studied. By this method the identity of each group of samples is specified such that maximum variance of the groups can be attained in the hyperspace. OPLS-DA applied to the same two most representative cultivars of Salento area (Cellina di Nardò and Ogliarola Leccese) gave a good model (1 predictive and 2 orthogonal) with R2 = 0.661 and Q2 = 0.448. The predictive variation, t1, corresponds to 9.01% of all variation in the data and the uncorrelated variation, to1 (orthogonal variation), corresponds to 2.22%. The score plot showed a clear separation of the two groups (Figure 3). Analyzing the loadings, Cellina samples showed higher levels of molecules having signals at 9.64 and 9.24 ppm, attributed to aldehydic protons and at 6.64 ppm, attributed to phenolic protons and lower levels of molecules having signals in the range δH 6.0 - 5.6, that could be ascribed to carotenoids. This can be also consistently observed by simple comparison of two 1H NMR spectra representative of the two cultivars (Figure 4). Interestingly the amount of phenolic compounds is an important factor when evaluating the quality of virgin olive oil because they are involved in resistance to oxidation and give a sharp bitter taste to the oil. The research conducted
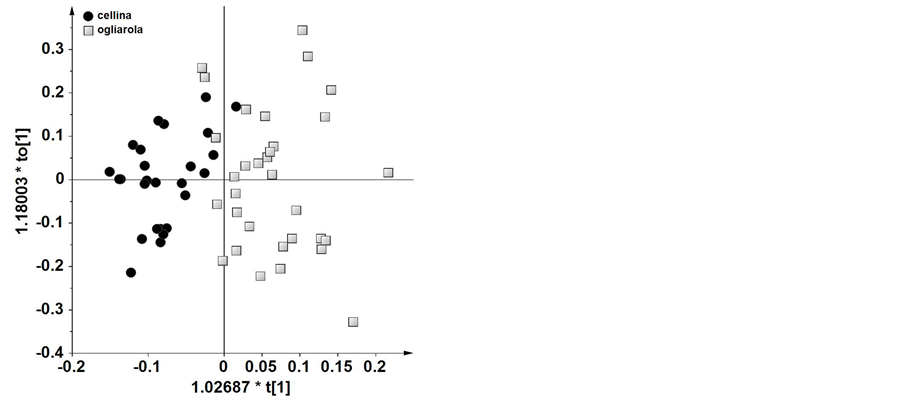
Figure 3. OPLS-DA scoreplot for monovarietal Cellina and Ogliarola samples (1 predictive and 2 orthogonal, R2 = 0.661 and Q2 = 0.448). The predictive variation, t1, corresponds to 9.01% of all variation in the data and the uncorrelated variation, to1 (orthogonal variation), corresponds to 2.22%.

Figure 4. Comparison among NMR spectra of Cellina and Ogliarola Leccese EVOO samples.
on olive oil chemical composition highlights that the polyphenols are remarkably variable according to the variety, the agronomic conditions, the state of ripeness, and the technology of conservation [17].
In general, concentrations of some molecules of the unsaponifiable fraction of EVOOs (such as minor components) and fatty acids resulted significantly different for the two cultivars considered. In addition, OPLS-DA was performed on Cellina di Nardò (26 samples) and Ogliarola Leccese (32 samples) using the statistical models for classification purposes of blend samples (35 Cellina/Ogliarola samples). Interestingly, both the PCA and OPLS-DA models had a good descriptive ability. The performance classification of OPLS-DA for blend EVOOs (Cellina/Ogliarola samples) is shown in the score plot tPS[1] vs. toPS [1] (Figure 5). This is essentially a good representation of the blend samples according to what declared by farmers (70% Cellina and 30% Ogliarola). Therefore, this model could also offer a method for blend classification by investigating the degree of overlap for blend samples with the monovarietal ones, pro-
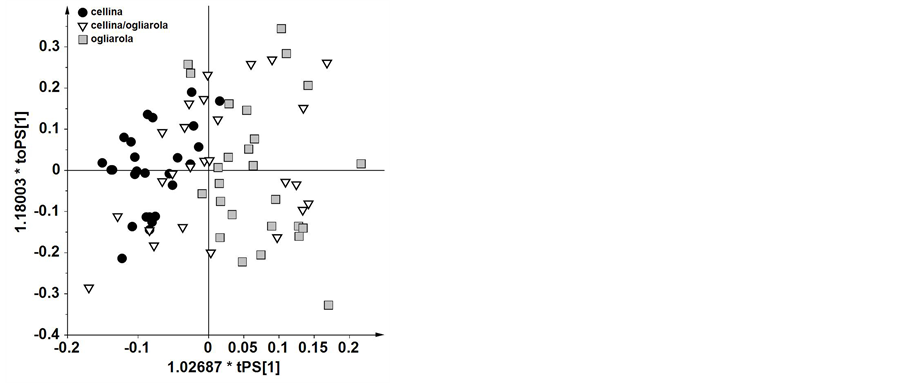
Figure 5. OPLS-DA CV-scoreplot for blend Cellina/Ogliarola samples (1 predictive and 2 orthogonal, R2 = 0.661 and Q2 = 0.448).
viding that they all were obtained in the same relatively small geographical area such as for Salento EVOOs.
In any case, both unsupervised and supervised methods are required for this kind of study, in particular PCA to look for trends among samples and possible outliers, while OPLS-DA to simply interpretation of data in the case of known class information [18].
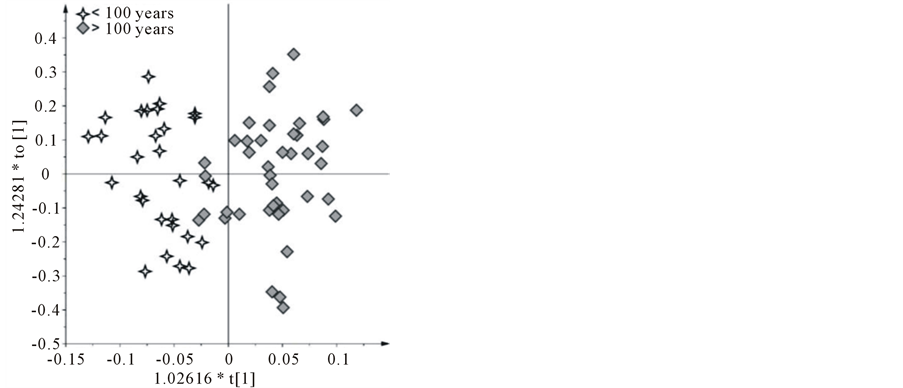
It is well known that Apulia region is the most important area for olive oil production in Italy, accounting for almost 40% of the total country production [19] and about 10% of the genetic olive tree patrimony within the region consists of the secular and monumental olive trees. Furthermore Apulia region approved a law aimed at protecting and enhancing of secular trees and claimed the introduction of the special mention in labeling: “Extra Virgin Olive Oil from the Apulia secular olive trees” (art. 7 L. R. n. 14, 4 June 2007) [20]. Therefore, an attempt for a deeper level of investigation on Salento EVOOs was performed using supervised statistical analysis (OPLS-DA) discriminating samples according to the age of the trees from which the EVOOs were extracted. OPLS-DA gave a good model (1 predictive and 5 orthogonal) with R2 = 0.669 and Q2 = −0.17 (Figure 6(a)). R2 is very high, while negative Q2 indicates that the model is not predictive, but remains still a very good descriptive model. Also in this case, a certain degree of variation in metabolite content was observed from the S-line plot analysis (Figure 6(b)). The S-line plot was used to visualize NMR signals that influence the separation of the groups. For EVOOs from young trees (<100 years) higher polyphenols and aldehydes content was found (signals at 9.52, 9.22 ppm for aldehydes, 6.76 and 6.56 for phenolic compounds), as well as for polyunsaturated fatty acid content (bis-allylic and allylic protons of both linolenic and linoleic acids, signals at 2.78 and 2.06 ppm, respectively).
 (a)
(a)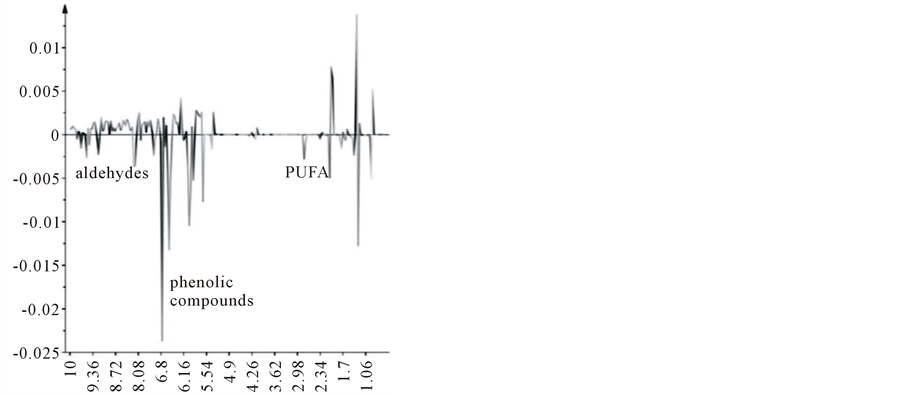 (b)
(b)
Figure 6. (a) OPLS-DA scoreplot (1 predictive and 5 orthogonal, R2 = 0.669 and Q2 = −0.17) based on the age of trees (young, <100 years and secular, >100 years); (b) S-line plot for the model between young and secular EVOOs. The relevance to the model is indicated by the signal amplitude.
4. Conclusion
This study provides an initial evaluation of how natural variability in the olive oil might affect blends originating from specific cultivars. It is worth noting that Ogliarola di Lecce (also known as Salentina) and Cellina di Nardò, which are the basis of “Terra d’Otranto” PDO EVOOs (alone or in combination at least for 60% [9]), are the most popular olive cultivars of the Jonian-Salentina area of Apulia region in Italy. Other varieties can also be present in a proportion not exceeding 40%. Multivariate statistical analyses (unsupervised, PCA, and supervised methods, OPLS-DA) were applied on 1H NMR data from monovarietal and blend EVOOs. In addition, PLS-DA and OPLS-DA were performed on Cellina di Nardò (26 samples) and Ogliarola Leccese (32 samples) using the statistical models for prediction purposes of blend samples (35 Cellina/Ogliarola samples) and finally for discrimination of EVOOs according to intrinsic features such as the age of the trees (young, <100 years and secular, >100 years). Higher polyphenols and polyunsaturated fatty acid content were found in young over secular olive tree productions. This study also suggests a methodological approach for verifying the composition of a blend of olive oil. This could be useful either in compliance with EU regulation no. 644/98 on the registration of geographical indications and designations of origin of agricultural products and foodstuffs [10] or on the mandatory labeling reporting the geographical origin of olive oils related to EU Regulation 182/2009 [21]. Interestingly this latter rule still lacks of official reference method to validate the region and therefore the country of origin for EVOOs. The statistical models obtained showed a very useful tool for both single cultivar and blend EVOOs characterization. Implementation of certification and authentication methods for the olive oil production of Salento area may give an “identity card” of an excellent quality product, reflecting the colors and tastes of Salento.
Acknowledgements
This work was supported by Apulia region project grant (PIF mis. 124 Filiera Olivicola 100% Pugliese JonicoSalentina).
We thank Agricola Nuova Generazione soc. coop. Agricola and Dr Carmelo Buttazzo for providing and organizing EVOO samples.
REFERENCES
- R. W. Owen, A. Giacosa, W. E. Hull, R. Haubner, G. Wurtele, B. Spiegelhalder and H. Bartsch, “Olive-Oil Consumption and Health: The Possible Role of Antioxidants,” Lancet Oncology, Vol. 1, No. 2, 2000, pp. 107- 112. http://dx.doi.org/10.1016/S1470-2045(00)00015-2
- P. Papadia, L. Del Coco, I. Muzzalupo, M. Rizzi, E. Perri, G. Cesari, V. Simeone, D. Mondelli, F. P. Schena and F. P. Fanizzi, “Multivariate Analysis of 1H NMR Spectra of Genetically Characterized Extra Virgin Olive Oils and Growth Soil Correlations,” JAOCS, Vol. 88, No. 10, 2011, pp. 1463-1475. http://dx.doi.org/10.1007/s11746-011-1812-1
- R. Godelmann, F. Fang, E. Humpfer, B. Schütz, M. Bansbach, H. Schäfer and M. Spraul, “Targeted and Nontargeted Wine Analysis by 1H NMR Spectroscopy Combined with Multivariate Statistical Analysis. Differentiation of Important Parameters: Grape Variety, Geographical Origin, Year of Vintage,” Journal of Agricultural and Food Chemistry, Vol. 61, No. 23, 2013, pp. 5610-5619. http://dx.doi.org/10.1021/jf400800d
- F. Longobardi, A. Ventrella, C. Napoli, E. Humpfer, B. Schütz, H. Schäfer, M. G. Kontominas and A. Sacco, “Classification of Olive Oils According to Geographical Origin by Using 1H NMR Fingerprinting Combined with Multivariate Analysis,” Food Chemistry, Vol. 130, No. 1, 2012, pp. 177-183. http://dx.doi.org/10.1016/j.foodchem.2011.06.045
- G. Oms-Oliu, I. Odriozola-Serrano and O. Martín-Belloso, “Metabolomics for Assessing Safety and Quality of PlantDerived Food,” Food Research International, Vol. 54, No. 1, 2013, pp. 1172-1183. http://dx.doi.org/10.1016/j.foodres.2013.04.005
- C. Aghemo, A. Albertino, R. Gobetto, C. Lussiana, A. De Maria and D. Isocrono, “Piedmont Olive Oils: Compositional Characterization and Discrimination from Oils from Other Regions,” European Journal of Lipid Science and Technology, Vol. 114, No. 12, 2012, pp. 1409-1416. http://dx.doi.org/10.1002/ejlt.201100244
- S. Ben Temime, W. Taanalli, B. Baccouri, L. Abaza, D. Daoud and M. Zarrouk, “Changes in Olive Oil Quality of Chétoui Variety According to Origin of Plantation,” Journal of Food Lipids, Vol. 13, No. 1, 2006, pp. 88-99. http://dx.doi.org/10.1111/j.1745-4522.2006.00036.x
- M. Paz Aguilera, G. Beltran, D. Ortega, A. Fernandez, A. Jiménez and M. Uceda, “Characterization of Virgin Olive Oil of Italian Olive Cultivars ‘Frantoio’ and ‘Leccino’, Grown in Andalusia,” Food Chemistry, Vol. 89, No. 3, 2005, pp. 387-391.
- D.M. 6 August 1998-GURI n. 193, Registration in the “Register of Protected Designations of Origin and Protected Geographical Indications” within the Meaning of Reg. CE 1065/97, 2014. http://www.le.camcom.gov.it/P42A0C100S82/D-O-P--Terra-d-Otranto.htm.
- http://eur-lex.europa.eu/LexUriServ/LexUriServ.do?uri=OJ:C:2013:237:0036:0039:EN:PDF
- http://www.regione.puglia.it/web/files/agricoltura/avviso_progetti_filiera.pdf
- J. C. Lindon, J. K. Nicholson and E. Holmes, “The Handbook of Metabonomics and Metabolomics,” Elsevier, Amsterdam, 2007.
- L. Del Coco, F. P. Schena and F. P. Fanizzi, “1H Nuclear Magnetic Resonance Study of Olive Oils Commercially Available as Italian Products in the United States of America,” Nutrients, Vol. 4, No. 5, 2012, pp. 343-355. http://dx.doi.org/10.3390/nu4050343
- L. Mannina, F. Marini, M. Gobbino, A. P. Sobolev and D. Capitani, “NMR and Chemometrics in Tracing European Olive Oils: The Case Study of Ligurian Samples,” Talanta, Vol. 80, No. 5, 2010, pp. 2141-2148. http://dx.doi.org/10.1016/j.talanta.2009.11.021
- H. Eastment and W. Krzanowski, “Cross-Validatory Choice of the Number of Components from a Principal Component Analysis,” Technometrics, Vol. 24, No. 1, 1982, pp. 73-77. http://dx.doi.org/10.1080/00401706.1982.10487712
- S. De Jong, “Multivariate Calibration, H. Martens and T. Naes, Wiley, New York, 1989. ISBN 0 471 90979 3. Price: £75.00, US$138.00. No. of Pages: 504,” Journal of Chemometrics, Vol. 4, No. 6, 1990, p. 441. http://dx.doi.org/10.1002/cem.1180040607
- A. Sacco, M. A. Brescia, V. Liuzzi, F. Reniero, G. Guillou, S. Ghelli and P. van der Meer, “Characterization of Italian Olive Oils Based on Analytical and Nuclear Magnetic Resonance Determinations,” Journal of the American Oil Chemists’ Society, Vol. 77, No. 6, 2000, pp. 619- 625. http://dx.doi.org/10.1007/s11746-000-0100-y
- B. Worley and R. Powers, “Multivariate Analysis in Metabolomics,” Current Metabolomics, Vol. 1, No. 1, 2013, pp. 92-107. http://dx.doi.org/10.2174/2213235X11301010092
- ISMEA, “Oli e Olive Mensa—Studi di Settore, Previsioni di Produzione Olio di Oliva,” 2013. http://www.ismea.it/flex/cm/pages/ServeBLOB.php/L/IT/IDPagina/8118
- 2004. http://ambiente.regione.puglia.it/index.php?option=com_content&view=article&id=818:censimento-ulivi-monumentali&catid=295:servizio-ecologia-ulivi-monumentali&Itemid=389
- The Commission of the European Communities, “Commission Regulation (EC) No 182/2009 of 6 March 2009 Amending Regulation (EC) No 1019/2002 on Marketing Standard for Olive Oil,” Off. J. Eur. Union 2009, L63, 6-8 (accessed November 2013).
NOTES
*Corresponding author.

