Natural Science
Vol. 5 No. 8 (2013) , Article ID: 35653 , 8 pages DOI:10.4236/ns.2013.58115
Hereditary constitution analysis of Shaolingyuan ancient human in Xi’an, northwestern China*
![]()
1Department of Oral Biology, Clinic of Oral Rare Diseases and Genetic Diseases, School of Stomatology, The Fourth Military Medical University, Xi’an, China; #Corresponding Author: xhduan@fmmu.edu.cn
2Department of Orthodontics, School of Stomatology, The Fourth Military Medical University, Xi’an, China; #Corresponding Author: shaojl@fmmu.edu.cn
3Shaanxi Provincial Institute of Archaeology, Xi’an, China
Copyright © 2013 Lin Li et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Received 6 February 2013; revised 6 March 2013; accepted 13 March 2013
Keywords: HVR-I; Chinese; Ancient; 3000 Years Ago
ABSTRACT
In order to identify the kinship of Shaolingyuan ancient human excavated from Shaolingyuan archaeological site with high level of certainty, and infer racial origins more clearly and reliably, this paper analyzed the hereditary constitution of this population. We used the “Reverse root canal technique” to extract ancient DNA from 28 teeth in 28 skeletal remains (3057 - 2784 BP) of Shaolingyuan archaeological site, obtained the sequences of mtDNA Hypervariable region I (HVR-I) by PCR amplifications; then used MEGA 5.5 software to construct phylogenetic trees and compared the sequences among the sequences of inter-races, intra-races. The phylogenetic tree showed that there were two major clusters, Cluster 1 with 16 individuals, and Cluster 2 with 5 individuals. Either the genetic gap or the geographic position of the individuals was small. The frequency of SNP site 16223 T > C was 71.4%, significantly higher than other sites. The comparisons of different population demonstrated that there is no significant difference among them. All of them shared the same haplogroup L1’2’3’4’5’6, close to African. Finally, we confirm that there is a very close genetic relationship between some individuals in this cemetery. We regarded Shaolingyuan Western Zhou cemetery as a family cemetery, and these people belong to East Asia lineage.
1. INTRODUCTION
Ancient DNA research provides the innovative ways to archaeologists and anthropologists to study the history [1-3]. The ancient mitochondrial DNA (mtDNA) has been used for the research of human evolution, population affinities, and genetic relationship identification. MtDNA has its own polymorphic property, distinctive features of maternal inheritance, high copy number, high mutation rate, and low recombination. Furthermore, its hypervariable region-I (HVR-I) has been an ideal genetic marker for individual identification and kinship determination [4,5]. Previous phylogenetic analyses of the sequence of the mtDNA HVR-I were used to study the ethnic migration and evolution, genetic relationship among different species [6].
The Shaolingyuan archaeological site is located in the southeast of Chang’an area in Xi’an, northwestern China. Shaolingyuan tombs and the materials accompanied in tombs reflected the historical period of 3057 - 2784 BP in northern China. There is no report about the hereditary constitution of the individuals in Shaolingyuan area [7]. Here we designed a method to extract ancient DNA from teeth, and detect their mtDNA HVR-I by sequencing, phylogenetic analyzing [8-11] and comparing it with the populations of different races, and the same race from different areas in China. We aimed to find the genetic relationship in Shaolongyuan population, and the racial origin of Shaolingyuan tombs.
2. MATERIAL AND METHODS
2.1. Sample Collection and Preparation
Investigation was carried out on 28 human remains, which were excavated from Shaolingyuan archaeological site in Xi’an (northern China), and stored in the Shaanxi provincial institute of archaeology. This population lived among the years of 3057 - 2784 BP, and the institute determined their gender and life period by their skeleton characteristics [7]. We selected 28 teeth from 28 different individuals. The pulp cavities of the chosen teeth were not perforated. Several precautions were taken to prevent contamination during this study [12]. The processing of the experimental samples was carried out in a sterile laminar flow cabinet. First removed the superficial dirt of the teeth using the blade and brush, soaked the samples in 1 mol/L HCL 5 - 10 min, then washed teeth with distilled water, anhydrous ethanol successively, and air dry naturally. Each sample was exposed to UV light for 15 min. Disposable gloves and masks were used in every step [13,14].
2.2. DNA Extraction, PCR Amplification and Sequencing
The orthograde entrance technique combined with Chelex-100 method to extract ancient DNA [13]. The Sterile Root canal reamers and files were used to entering the teeth cavity from the apical foramen reversely in accordance with the order from small numbers to large, rasped root canal wall lightly. 5 - 10 mg dentin scrap and pulp tissue can be acquired Each sample was mixed with 200 µl 5% Chelex-100 and 2 µl proteinase K, then shook for 12 h, 150 r/min, 56˚C, then, shook for 15 s. After water bath at 100˚C 8 min, concussion 15 s; The samples were centrifuged under 12000 r/min for 5 min and kept the supernatant. The quantification was performed using Nanodrop ND-2000 spectrophotometer. Amplification was performed using AMFlSTR SGM PlusTM Primer set on 9600 Gene Amp PCR system according to manufacturers recommendations. Extraction controls without powdered sample were processed in parallel to test for contamination during the extraction process. Amplification was performed in another UV sterile room.
Hypervariable region I (HVR-I) of the mtDNA control region was amplified using the primer pairs (L16035 5’-GAAGCAGATTTGGGTACCAC-3’, H16398 5’-CAAGGGACCCCTATCTGAGG-3’), PCR conditions were as follows: pre-denaturation at 94˚C for 5 mins, followed by 38 cycles at 94˚C for 1 min, 55 s at 55˚C, and 72˚C for 4 mins, and final extension at 72˚C for 10 mins. PCR amplifications were carried out in 50 ml of reaction mixture. HVR-I amplification products were visualized on a 2% agarose gel, then, purified these products and sequenced both directions, were analyzed on an ABI Prism 3730 (PE Applied Biosystems) automated DNA sequencer in the Sequencing Service of the Sunny Biological Technology co., LTD (Shanghai, China). In this experiment, DNA extraction, quantitation, amplification and data analysis were performed in separated laboratory areas. The separate hoods used for DNA extraction and PCR preparation were irradiated with UV rays before and after the experiment. Disposable sterile tubes, sterile reagents and solutions were used. Negative controls were included in each amplification.
2.3. Phylogenetic Analysis
The phylogenetic trees were constructed by the MEGA 5.5 software tool using the Maximum Parsimony (MP) method, bootstrap values of 50 and higher are indicated showing on the nodes. Reliability of nodes was assessed with non-parametric bootstraps [15]. One thousand bootstraps were generated [16,17] since MP method reconstructs correct phylogenetic trees with a high probability in the relatively closely related samples [18]. In this analysis, genetic distances were estimated on the basis of sequence data, the bootstrapped data sets were used to generate genetic distances by DNADIST 15. These sets of distances were then used to generate consensus trees by CONSENSE, in which the percentage of support for the branches represents the bootstrap value for the nodes of the trees [19].
2.4. Sequence Comparing
In order to identify the mutation position, we used mitotool software to compare the mtDNA HVR-I sequences of 28 individuals with rCRS searching from PUBMED and mitotool software [20] to investigate the relationship between Shaolingyuan ancient people and other populations.
The intrarace contrast crowds are Han Chinese from Shandong, Hubei, Yunnan, Inner Mongolia, Guangzhou, and Liaoning; Daur and Ewenki from Inner Mongolia, Miao from Hunan province. We also compared Shaolingyuan ancient population to people from other countries, including Japan, Russia, and Morocco. All sequences of the compared people are searched in PUBMED (AY 255134-AY255180) [16].
3. RESULTS
By the phylogenetic analysis of mtDNA HVR-I, we found two major clusters in the MP-tree. Cluster 1 contained 16 individuals, and M79, M207 consisted a subcluster of cluster 1. 5 individuals belong to Cluster 2. M258 and M64 formed two branches independently. Every member in the same cluster came from an identical ancestor. In contrast, M293, M310, and M464 were scatted relatively.
The MP tree (Figure 1) suggests that the genetic gap of some individuals in this cemetery is relatively small, and indicates a close relationship among the samples plainly, especially for the members in Cluster 1, and Cluster 2. Either the sequence difference among the samples—M354, M297, M307, M340, M467, M451, M85,
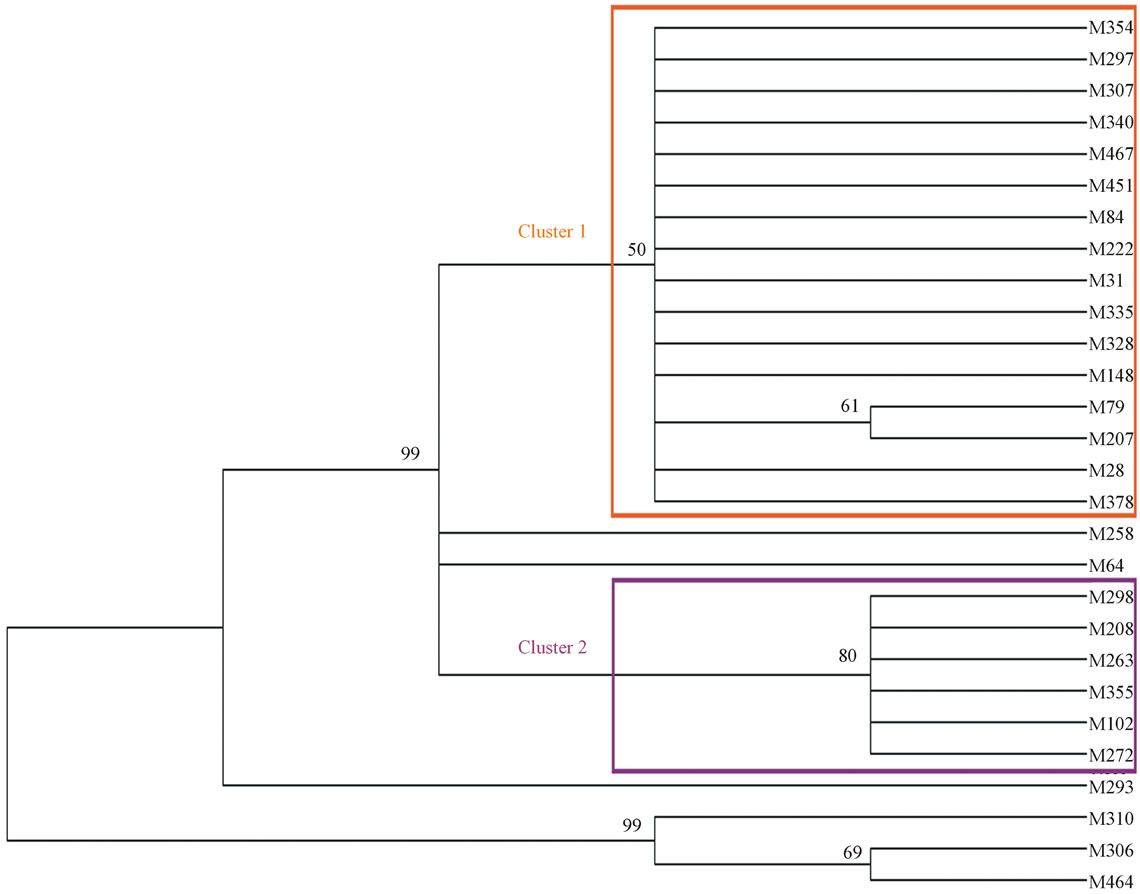
Figure 1. Phylogenetic tree of Shaolingyuan ancient population. Trees were generated by a bootstrap test with 1000 replications based on Maximum Parsimony (MP) method. Bootstrap values of 50 and higher are labeled on the nodes.
M222, M31, M335, M328, M148, M79, M207, M28, M378, or the samples—M298, M208, M263, M355, M102, M272 are small. These findings indicated that there was a very close genetic relationship in some individuals of this cemetery.
The cemetery is divided into five areas (Block1 - Block5) according to the geographical positions. As shown in Figure 2, with regard to Cluster 1, 9 of 16 individuals were gathered in Block 5; 3 in Block 2; others scattered. For Cluster 2, 3 of 6 individuals gathered in Block 3, others randomly distributed to the rest blocks. Therefore, individuals of the same family are buried intensively, especially for Cluster 1. Geographically,M467 was in the center of the Cluster 1, and the distance between M467 and M84 (the farthest individual to the central M467) is just about 83 meters.
The result of mtDNA HVR-I sequence comparison are shown in Tables 1 and 2. The mutation points of Shaolingyuan ancient population are shown in Table 1. The ratio of SNP site 16223 T > C are 71.4%, significantly higher than other sites. As Table 2 shows all of them shared the same haplogroup, but have different mutation points. As shown in Figure 3, this kind of haplogroup belongs to L*.
4. DISCUSSION
It is known that human mtDNA is a closed double-stranded circular molecule constructed by 16569-nt. MtDNA contains a 1121-nt non-coding region and a coding region. The non-coding region referred to as the control region or D-loop region, and its evolution rate is the highest among mtDNA genome. It is also the most polymorphic region within mtDNA genome. There are two hypervariable regions in mtDNA, HVR-I and HVRII. These two regions have been treated as an ideal forensic marker for individual identification, and kinship determination [5,20,21].
The researches on mtDNA variation and phylogenetic level have been advanced in the past few years [15,19]. Some researchers have studied on the attribution and evolutionary relationship of the crowd [17,22]; others have estimated divergence time of the crowd [23,24]. In contrast, the analysis of genetic relationship between individuals in this method is relatively rare. Moreover, according to the archaeological research of Shaolingyuan

Figure 2. The Burial maps of Shaolingyuan in Xi’an (3057 - 2784 BP). Orange and purple color represent two different family clusters respectively Green represents all the left individuals with no close relationship. Cluster 1: 9 of 16 individuals were gathered in block 5; 3 in block 2; others scattered. Cluster 2: 3 of 6 individuals gathered in block 3, others distributed to the rest blocks.
site, the time span of these tombs covered the whole period of Western Zhou dynasty, which was inferred to be a family cemetery of the civilian. There were some weapons accompanied with several individuals [7]. It is quite important to identify the genetic relationship among those individuals with weapons, thus we might infer whether the tomb was a family cemetery or a military residence. Here we found a very close genetic relationship in some individuals of this cemetery. With regard to the kinship between the scattered tombs, it is hard to determine the further relationship and family construct. But we may believe that Shaolingyuan Western Zhou cemetery is a family cemetery.
The comparison of mtDNA HVR-I sequence was performed between Shaolingyuan ancient individuals to different references, such as other Chinese ethnic people,
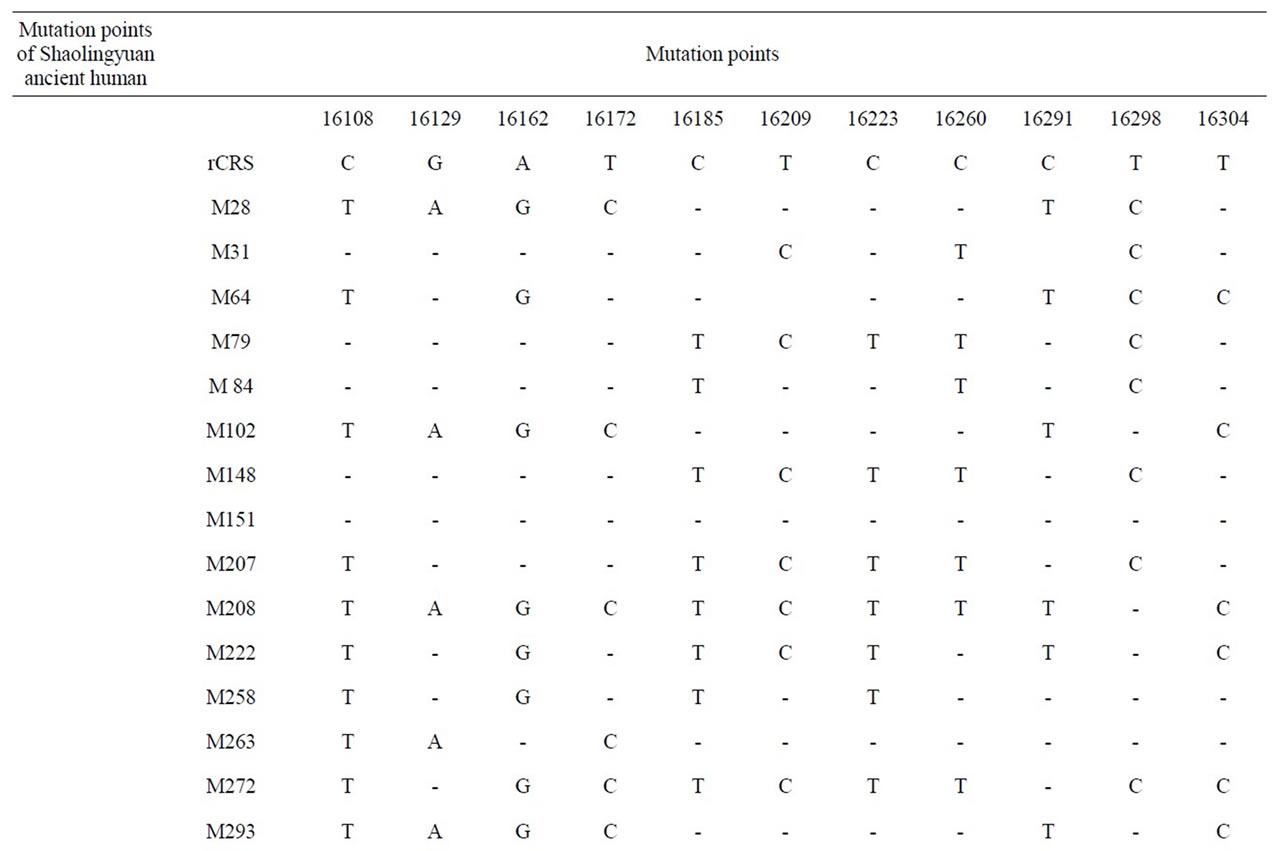
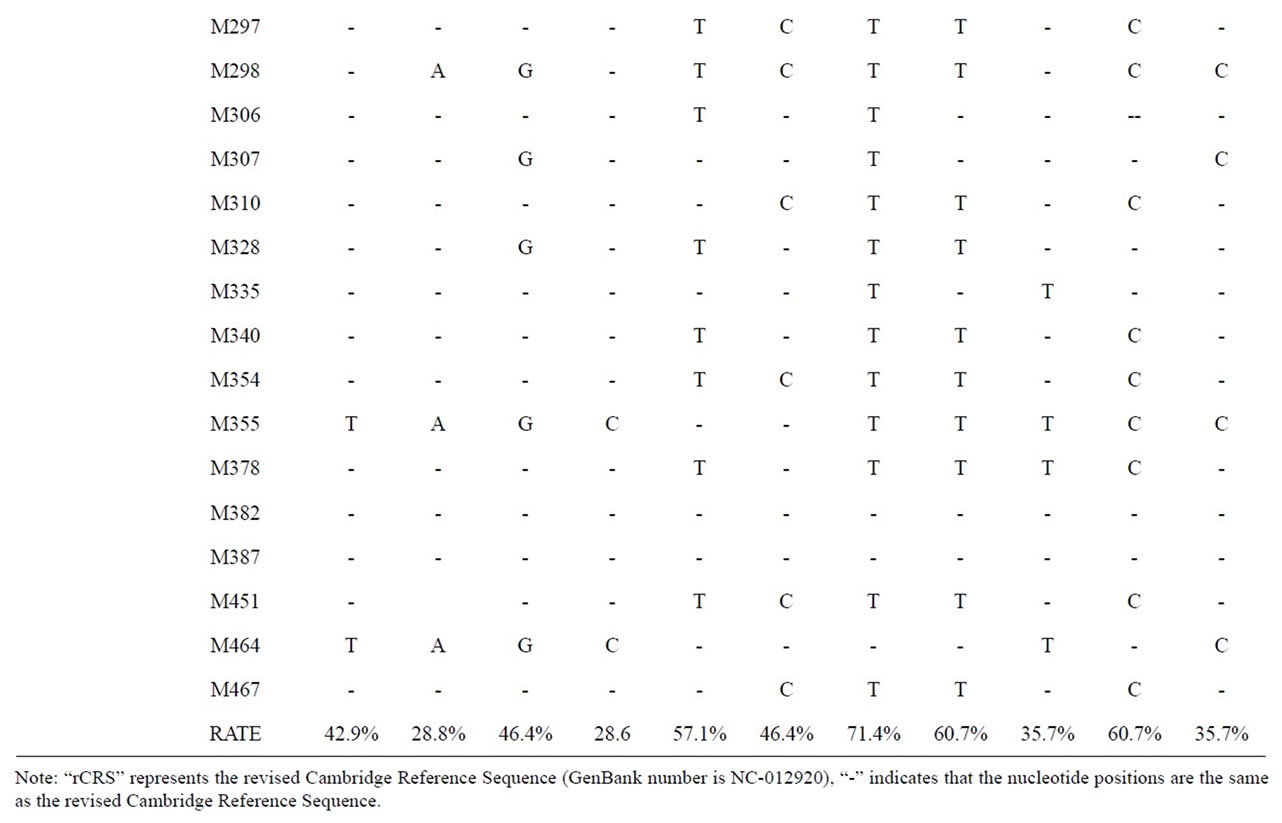
Table 1. Mutation points of Shaolingyuan ancient human.
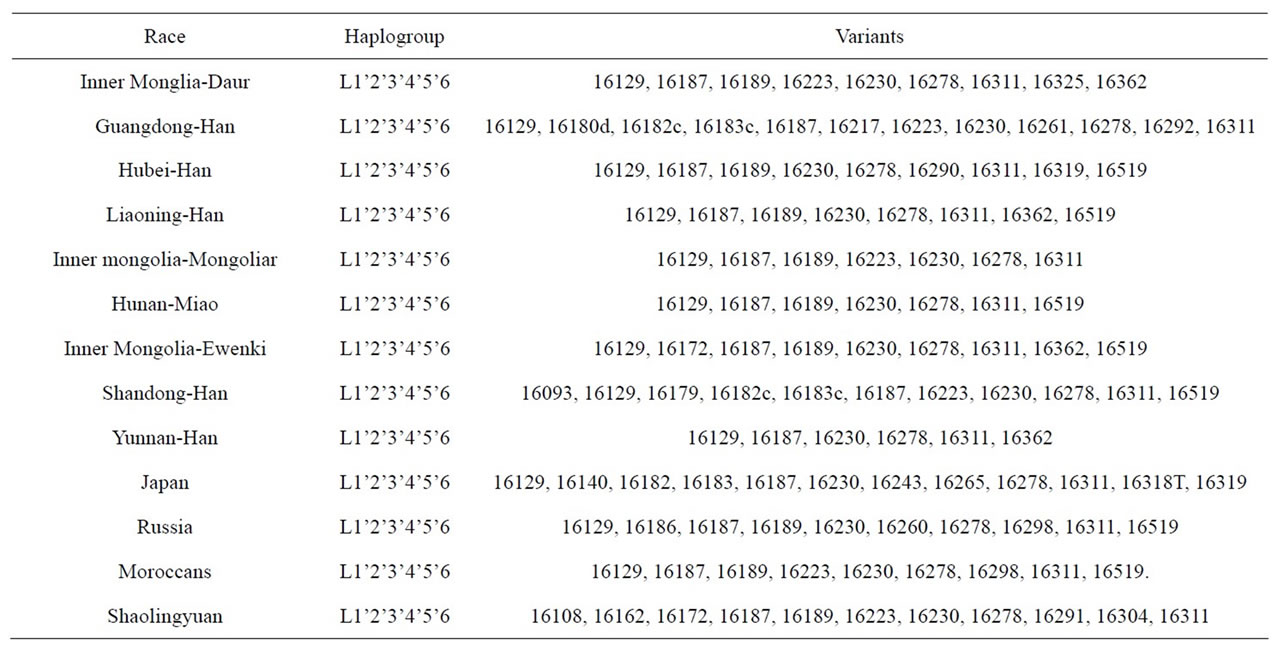

Table 2. Comparisons of the mtDNA HVR-I region sequence.
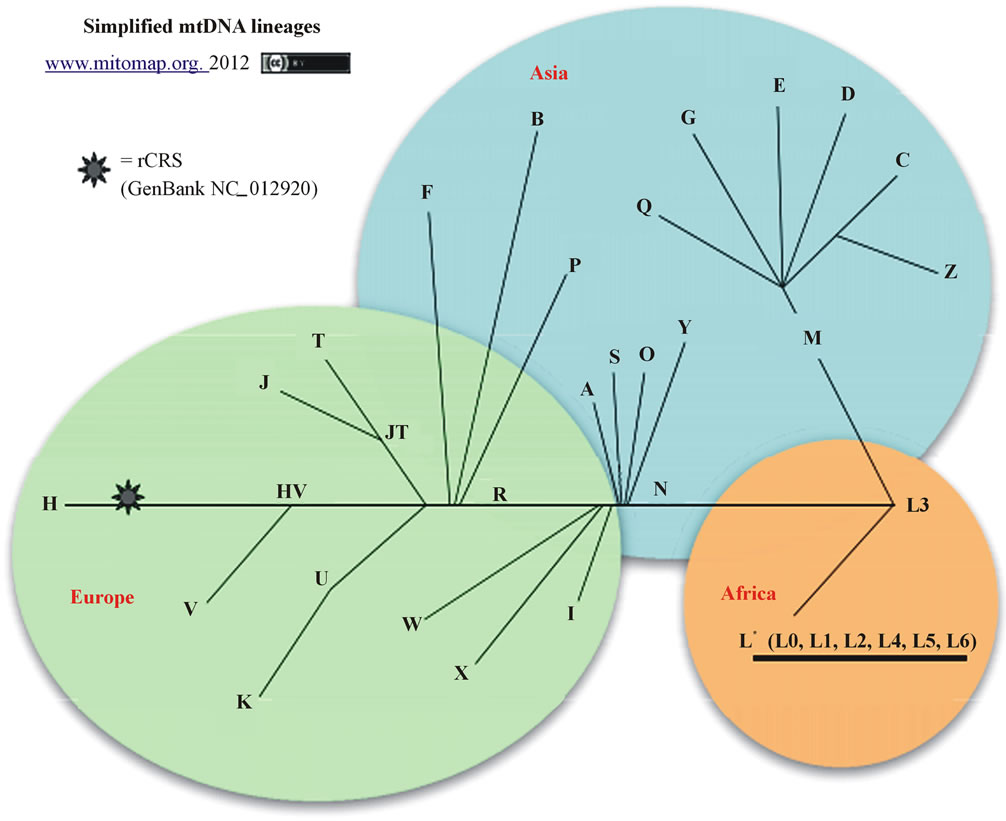
Figure 3. Simplified mtDNA lineages from the web of Mitomap. The African belongs to L haplotype, the underlined part is the experimental samples haplotype belonged to.
and non-Chinese individuals, including Japanese, Swiss, Russian, Moroccans. However, there was no significant sequence difference among those populations. All these crowds are belonged to the haplogroup L, according to Figure 3, which also exists in Africa, the L3 branch. According to the previous studies, the frequencies of position 16223 (C to T) mutation are higher in the population of East Asia (Mongolian 65%, Xinjiang 48.5%, Japan 41.7%), lower in African, and rare in European [25]. While in present study, the mutation frequency of position 16223 (C to T) in Shaolingyuan ancient people is about 71.4%.
Previous physical anthropology studies have shown that the ancient residents of Shaanxi region have been part of the East Asian lineage as early as in the Yangshao period [26]. Historical records and Anthropological studies have shown that, about 60,000 years ago, modern humans oriented from African moved into East Asia, in the following tens of thousands of years, gradually moved northward throughout mainland China [27]. After a long period of ignorance, the earliest Chinese civilization began to sprout in the middle and upper reaches of the Yellow River about 8500 years ago. At this point, the predecessor of the Huaxia developed.
The Shaolingyuan site could reflect WesternZhou dynasty (3057 - 2784 BP). Western Zhou dynasty marked a prosperous period of the ancient society in China. During this period, various ethnic groups and tribes of the territory began to integration, and Huaxia had further formation and development, becoming modern Han predecessor. In summary, combined the analysis of mtDNA HVRI with the historical research, we may infer that Shaolingyuan ancient population belongs to East Asia lineage.
5. CONCLUSION
Here phylogenetic analysis offers us a sure verdict that there are very close genetic relationships in some of the individuals from Shaolingyuan tombs and the results of the sequence comparison give support to these ancient groups which belong to East Asia lineage, from the perspective of mtDNA. Further study should be performed on sequencing the complete mtDNA genome [28-31].
6. ACKNOWLEDGEMENTS
Thanks to the Shaanxi Provincial Institute of Archaeology staffs for their helpful support in sample collection. Thanks to Dr. Xue-juan Li, and Xiao-yang Wang of University of Shaanxi Normal, China, for their advice and helps in writing this paper. Thanks Prof. Jianghua Lai in Xi’an Jiaotong University gave us comments on our manuscript. This work was supported in part by grants of National Scientific Foundation of China (81070819, 31070835; 81271116).
![]()
![]()
REFERENCES
- Kirsanow, K. and Burger, J. (2012) Ancient human DNA. Annals of Anatomy, 194, 121-132. doi:10.1016/j.aanat.2011.11.002[, #7]
- Rasmussen, M., Ying-rui, L., Lindgreen, S., Pedersen, J.S., Albrechtsen, A., Moltke, I., Metspalu, M., Metspalu, E., Kivisild, T., Gupta, R., Bertalan, M., Nielsen, K., Gilbert, M.T.P., Yong, W., Raghavan, M., Campos, P.F., Kamp, H.M., Wilson, A.S., Gledhill, A., Tridico, S., Bunce, M., Lorenzen, E.D., Binladen, J., Xiao-sen, G., Jing, Z., Xiuqing, Z., Hao, Z., Zhuo, L., Minfeng, C., Orlando, L., Kristiansen, K., Bak, M., Tommerup, N., Bendixen, C., Pierre, T.L., Grønnow, B., Meldgaard, M., Andreasen, C., Fedorova, S.A., Osipova, L.P., Thomas, F., Higham, G., Ramsey, C.B., Hansen, T.O., Nielsen, F.C., Crawford, M.H., Brunak, S., Sicheritz-Ponten, T., Villems, R., Nielsen, R., Krogh, A., Jun, W. and Willerslev, E. (2010) Ancient human genome sequence of an extinct Palaeo-Eskimo. Nature, 463, 757-762. doi:10.1038/nature08835
- Rohland, N. and Hofreiter, M. (2007) Ancient DNA extraction from bones and teeth. Nature Protocols, 2, 1756- 1762. doi:10.1038/nprot.2007.247
- Umetsua, K. and Yuasa, I., 2005Recent progress in mitochondrial DNA analysis. Legal Medicine, 7, 259-262. doi:10.1016/j.legalmed.2005.01.005
- Anderson, S., Bankier, A.T., Barrell, B.G., de Bruijn, M.H., Coulson, A.R., Drouin, J., Eperon, I.C., Nierlich, D.P., Roe, B.A., Sanger, F., Schreier, P.H., Smith, A.J., Staden, R. and Young, I.G. (1981) Sequence and organization of the human mitochondrial genome. Nature, 290, 457-465. doi:10.1038/290457a0
- Brown, T.A. and Brown, K.A. (1993) Ancient DNA: using molecular biology to explore the past. Bioessays, 16, 719-726. doi:10.1002/bies.950161006
- Shaanxi Provincial Institute of Archaeology, The archaeology report of the Shaolingyuan tombs. Science Press, 2009.
- Adachi, N., Umetsub, K., Takigawa, W. and Sakaue, K. (2004) Phylogenic analysis of the human ancient mitochondrial DNA. Journal of Archaeological Science, 31, 1339-1348. doi:10.1016/j.jas.2004.02.011
- Oota, H., Saitou, N., Matsushita, T. and Ueda, S. (1995) A genetic study of 2000-year-old human remains from Japan using mitochondrial DNA sequences. American Journal of Physical Anthropology, 98, 133-145. doi:10.1002/ajpa.1330980204
- Oota, H., Saitou, N., Matsushita, T. and Ueda, S. (1999) Molecular genetic analysis of remains of a 2000-year-old human population in China and its relevance for the origin of the modern Japanese population. The American Journal of Human Genetics, 64, 250-258. doi:10.1086/302197
- Metress, J.F. and Conway, T. (1975) Standardized system for recording dental caries in prehistoric skeletons. Journal of Dental Research, 54, 908. doi:10.1177/00220345750540043901
- Dongya, Y. and Kathy, W. (2005) Contamination controls when preparing archaeological remains for ancient DNA analysis. Journal of Archaeological Science, 32, 331-336. doi:10.1016/j.jas.2004.09.008
- Alakoc, Y.D. and Aka, P.S. (2009) “Orthograde entrance technique” to recover DNA from ancient teeth preserving the physical structure. Forensic Science International, 188, 96-98. doi:10.1016/j.forsciint.2009.03.020
- Cobb, J.C. (2002) Ancient DNA recovered by a nondestructive method. Ancient Biomolecules, 4, 169-172. doi:10.1080/1358612021000028461
- Archie, J.W. and Felsenstein, J. (1993) The number of evolutionary steps on random and minimum length trees for random evolutionary data. Theoretical Population Biology, 43, 52-79. doi:10.1006/tpbi.1993.1003
- Kong, Q.P., Yao, Y.G., Sun, C., Bandelt, H.J., Zhu, C.L. and Zhang, Y.P. (2003) Phylogeny of East Asian mitochondrial DNA lineages inferred from complete sequences. The American Journal of Human Genetics, 73, 671-676. doi:10.1086/377718
- Barros, M.C., Sampaio, I. and Schneider, H. (2003) Phylogenetic analysis of 16S mitochondrial DNA data in sloths and anteaters. Genetics and Molecular Biology, 26, 5-11. doi:10.1590/S1415-47572003000100002
- Saitou, N. and Imanishi, T. (1989) Relative efficiencies of the Fitch Margoliash, maximum-parsimony, maximumlikelihood, minimum-evolution, and neighbor-joining methods of phylogenetic tree construction in obtaining the correct tree. Molecular Biology and Evolution, 6, 514- 525.
- Starikovskaya, Y.B., Sukernik, R.I., Theodore, Schurr, G., Kogelnik, A.M. and Wallace, D.C. (1998) MtDNA diversity in Chukchi and Siberian Eskimos: implications for the genetic history of ancient beringia and the peopling of the New World. The American Journal of Human Genetics, 63, 1473-1491. doi:10.1086/302087
- Fan, L. and Yao, Y.G. (2011) Mitotool: A web server for the analysis and retrieval of human mitochondrial DNA sequence variations. Mitochondrion, 11, 351-356. doi:10.1016/j.mito.2010.09.013
- Wallace, D.C. (2007) Why do we still have a maternally inherited mitochondrial DNA? Insights from evolutionary medicine. Annual Review of Biochemistry, 76, 781-821. doi:10.1146/annurev.biochem.76.081205.150955
- Smith, A.B. and Peterson, K.J. (2002) Dating the time of origin of major clades: Molecular clocks and the fossil record. Annual Review of Earth and Planetary Sciences, 30, 65-88. doi:10.1146/annurev.earth.30.091201.140057
- Soares, P., Rito, T., Trejaut, J., Mormina, M., Hill, C., Hundal, E.T., Braid, M., Clarke, D.J., Loo, J.H., Thomson, N., Denham, T., Donohue, M., Macaulay, V., Lin, M., Oppenheimer, S. and Richards, M.B. (2011) Ancient voyaging and Polynesian origins. The American Journal of Human Genetics, 88, 239-247. doi:10.1016/j.ajhg.2011.01.009
- Feng, J., Zhang, J.Y. and Liu, M. (2011) Association of mtDNA haplogroup F with healthy longevity in the female Chuang population, China. Experimental Gerontology, 9, 1-7.
- Zheng, X.Y. (1994) Ethnic ingredient research of the ancient residents northwestern China. China Academic Journal Electronic Publishing House, 2008, 85-89.
- Cui, Y.Q., Duan, R.H., Ji, C.N., Zhu, H., Li, W., Min, M.Y. and Zhou, H. (2002) Analys is of mitochondrial DNA from the ancient ruins of Jiao-he, China. Chemical Journal of Chinese Universities, 23, 1510-1514.
- Zhu, H. (2006) Ancient race of Northwest China. Archive of Cult Relics, 5, 60-65.
- Ballinger, S.W., Schurr, T.G., Torroni, A., Gan, Y.Y., Hodge, J.H., Hassan, K., Chen, K.H. and Wallace, D.C. (1992) Southeast Asian mitochondrial DNA analysis reveals genetic continuity of ancient mongoloid migrations. Genetics, 130, 139-152.
- Simon, Y.W., Ho, M., Thomas, P. and Gilbert (2010) Ancient mitogenomics. Mitochondrion, 10, 1-11.
- Izagirre, N., Dela, C. and Ru, A. (1999) An mtDNA analysis in ancient Basque populations: Implications for haplogroup V as a marker for a major Paleolithic expansion from Southwestern Europe. The American Journal of Human Genetics, 65, 199-207. doi:10.1086/302442
- Ricaut, F.X., Thomas, T., Mormina, M., Cox, M.P., Bellatti, M., Foley, R.A. and Marta, M.L. (2010) Ancient Solomon islands mtDNA: Assessing holocene settlement and the impact of European contact. Journal of Archaeological Science, 37, 1161-1170. doi:10.1016/j.jas.2009.12.014
NOTES
*Funding: National Scientific Foundation of China (81070819, 31070835; 81271116).

