Advances in Bioscience and Biotechnology
Vol.5 No.3(2014), Article ID:42709,11 pages DOI:10.4236/abb.2014.53030
Genomic analysis of a novel strain of Bacillus nealsonii, isolated from Surti buffalo rumen
![]()
1Department of Animal Biotechnology, College of Veterinary Science & Animal Husbandry, Anand Agricultural University, Anand, India
2Department of Microbiology, Gujarat Vidyapith, Sadra, India
Email: cgjoshi@rediffmail.com
Copyright © 2014 Neelam M. Nathani et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. In accordance of the Creative Commons Attribution License all Copyrights © 2014 are reserved for SCIRP and the owner of the intellectual property Neelam M. Nathani et al. All Copyright © 2014 are guarded by law and by SCIRP as a guardian.
Received 9 November 2013; revised 16 January 2014; accepted 28 January 2014
KEYWORDS: Bacillus nealsonii AAU1; Anaerobic; Phenotypic Characterization; Genomic Analysis
ABSTRACT
Aim: Whole genome sequencing and functional annotation of Bacillus nealsonii strain AAU1, an amylolytic anaerobic spore forming isolate from ruminal contents of buffalo. Methods and Results: Morphologically, the strain was observed as slender Grampositive rods, occurring in pairs. Optimal growth was observed at 40˚C (range: 30˚C to 45˚C) and pH 6.5 (range: 5.5 to 7.5) when cultivated in Hungate’s medium supplemented with starch. The microorganism showed extracellular constitutive amylolytic activity, proving to be capable of utilizing glucose, maltose, mannose, trehalose, dextrin and starch under anaerobic conditions. Sequence analysis revealed a GC content of 35.1 mol%. Comparison of housekeeping gene sequences for RNA polymerase subunit B (rpoB) and gyrase A (gyrA) identified sequence similarity within the Bacillus genus, confirmed by 16S rRNA gene sequence similarity which identified Bacillus nealsonii DSM 15077 as the closest publically available relative. Chemotaxonomic analysis provided conflicting results with straight-chain saturated C16: 0/C16:0 aldehyde, C16:0 DMA, C14:0 and monounsaturated 16:1w7c and 16:1w9c the major fatty acids in contrast to those reported for B. nealsonii DSM15077. Further characterization using AN-Biolog and physiological parameters provided genotypic and phenotypic support for taxonomic classification of isolate AAU1 with published Bacillus species including B. licheniformis, B. subtilis, B. circulans and B. nealsonii. Conclusion: Based on the data presented, isolate is likely to represent a new strain/subspecies, for which the identifier B. nealsonii AAU1 is proposed. Significance and Impact of Study: The strict anaerobic conditions prevailing in the bovine rumen from where AAU1 was isolated may have resulted in genetic polymorphism influencing its metabolic characteristics. Bacillus nealsonii AAU1; Anaerobic; Phenotypic Characterization; Genomic Analysis
1. INTRODUCTION
The rumen harbors a large and diverse range of microorganisms categorized into Bacteria, Archaea (methanogens) and Eucarya (protozoa and fungi) [1]. Obligatory anaerobes are dominated and supplemented by facultative anaerobes including Streptococcus, Staphylococcus, Bacillus and Lactobacillus species [2]. The complex microbial ecosystem of the rumen functions as an efficient biological fermentor and provides nutrients essential for the growth and productivity of the ruminant host in the form of volatile fatty acids and microbial protein.
The Bacillus genus, introduced by Cohn in the year 1872, comprises more than 200 species and is considered to be among the largest bacterial genera with new additions identified every year. The Bacilli are rod shaped gram positive bacteria, and characterized by spore forming ability and aerobic or facultative anaerobic metabolism [3]. Single spores are formed per cell in response to environmental stress, such as heat, cold, radiation or desiccation; features which support their existence in extreme habitats include desert sands, hot springs and Arctic soils. Sequencing bacterial genomes provide insight into the genetic basis of phenotypic plasticity and their ability to tolerate environmental stresses [4]. Bacillus nealsonii species have previously been isolated from potentially harsh environments including a spacecraft assembly, spores of which were observed to be resistant to ultraviolet (UV) light, γ-radiation, hydrogen peroxide and desiccation [5]. Hence, study of these bacteria can improve understanding of bacterial responses to environmental stress and inform upon their role in complex microbial communities, such as those found within the bovine rumen.
The Surti buffalo (Bubalus bubalis) is a buffalo breed which is popular across Gujarat, India, commonly providing milk and draught power. Buffalo are notable for their feed conversion ability from rough grazing, promoting characterization of their ruminal microbiota. We describe here the isolation and characterization by sequencing and biochemical analysis of a putatively new B. nealsonii strain or isolate with relevance to the future understanding and improvement of ruminant health and nutrition.
2. MATERIALS AND METHOD
2.1. Sampling and Enrichment
Rumen fluid samples were collected aseptically from a Surti Buffalo at the Veterinary College, Anand Agricultural University (AAU), Anand, India. The ruminal fluid samples were enriched in Hungate medium containing starch in roll tubes, maintained in strict anaerobic conditions by purging N2:CO2 (80:20) and sealed with butyl rubber and aluminum seals.
2.2. Media and Culture Conditions
Anaerobic culture techniques and incubations were performed as described elsewhere by Hungate and colleagues [6]. Hungate medium [7], used for enrichment, serial dilution and subsequent cultivation of bacteria from ruminal fluid, contained (per 1000 ml) K2HPO4 0.5 g, KH2PO4 0.2 g, (NH4)2SO4 0.5 g, NaCl 1.0 g, MgSO4 0.02 g, CaCl2 0.05 g, NaHCO3 5.0 g, cysteine hydrochloride 0.5 g, starch 5.0 g, clarified rumen fluid 1% and agar 3% at pH 7.2. The medium was pre gassed with N2:H2 (80:20); sterilized at 10 psi followed by post gassing with N2:CO2 (80:20) and sealed with butyl rubber and aluminium seals.
2.3. Isolation
Serial dilutions were prepared in Hungate medium from enriched samples and 0.1 mL from each dilution was streaked on Hungate Agar medium containing starch in anaerobic bottles. The medium was post gassed with N2:CO2 (80:20). All plates were incubated at 40˚C ± 2˚C, after which colony morphology was recorded. Subsequently, three sequential transfers were sub-cultured from well isolated colonies after dilution to purify the culture.
2.4. Characterization and Substrate Utilization Profile
Carbon utilization by the putative Bacillus isolate was investigated biochemically using the AN-Biolog® microplate assay to determine substrate fermenting potential as per the manufacturer’s instructions. Subsequently, the OD595 was measured using a microtiter plate reader after 72 h incubation. Growth, colony characteristics and morphology of the isolate were monitored according to methods prescribed in Bergey’s Manual.
2.5. FAME Analysis
Fatty acid methyl ester (FAME) analysis was carried out by gas-liquid chromatography (Sherlock Microbial Identification System [MIS]; MIDI, Inc.). Chromatographic data defining the isolate were analyzed using the Sherlock software version 6.0B (S/N; 160277) MIDI, with the SMOORE6 method.
2.6. Genome Sequencing and Assembly
The whole genome sequence of the putative Bacillus isolate was determined by 454 GS-FLX (Roche) and Ion Torrent PGM platform sequencing as per the manufacturer’s instructions. The results were generated by the GS run browser and the sequencing reads were assembled using the GS De Novo Assembler V.2.6 providing consensus contigs.
2.7. Bioinformatic Analysis: 16S rRNA and Housekeeping Genes
Comparison of similarity to published bacterial sequences was confirmed using homology by uploading the assembled contigs into the RDP-Ribosomal Database Project Classifier. For 16S rRNA homology studies, local BLAST of the assembled contigs was performed against a 16S rRNA gene sequence database of 7545 sequences downloaded from NCBI. Comparison of genes encoding the housekeeping proteins rpoB (RNA polymerase open promoter) and gyrase A was undertaken using publically available sequences downloaded from NCBI. Sequences coding for 16S rRNA and gyrA of the genus Bacillus were aligned with that of Bacillus nealsonii AAU1, using ClustalW. Subsequently, an evolutionary distance matrix was generated from these nucleotide sequences in the dataset using Maximum Composite Likelihood method. Phylogenetic analysis was performed using the Neighbor Joining method by MEGA (Molecular Evolutionary Genetics analysis) version 4.0 [8].
2.8. Gene Prediction and Annotation
Whole genome gene prediction and annotation was performed by uploading the assembled contigs onto the RAST-Rapid Annotation using Subsystem Technology server V. 4.0 (http://rast.nmpdr.org/rast.cgi) [9]. Bacterial features identified were studied for metabolic and other functional potentials.
3. RESULT
3.1. Morphology, Physiological Characteristics and Substrate Utilization
Morphologically the organism was found to be gram positive and motile, appearing as long slender rods, singly or in chains with a terminal spore. Colonies were 3 - 4 mm in diameter, irregular, flat, opaque and creamy or white after 4 - 5 days of incubation at 40˚C in Hungate medium containing starch. The isolate grew in 0.5% - 7% NaCl, at pH 5 - 8, but did not grow at pH below 5, and temperature below 15˚C or above 60˚C. The isolate showed acid and gas production in the presence of glucose and mannose in the medium. Results of the AN-Biolog plate revealed the substrate utilization profile of the isolate using 95 different substrates as the sole source of carbon.
Out of the total 95 substrates present in the AN-Biolog plate, 38 were metabolized. Twelve substrate types were not metabolized (aromatic chemicals, carboxylic acids, dicarboxylic acid and ether) while none of the amino sugars and sugar alcohols provided among a panel of ten carbohydrate types were utilized. The B. nealsonii AAU1 strain was found to utilize high numbers of hexoses and amino acids. The substrates utilized included 0/3 amino sugars, 1/2 pentoses, 1/1 ketose, 1/2 deoxysugars, 0/6 sugar alcohols, 3/6 monosaccharides, 2/6 disaccharides, 4/11 oligosaccharides, 3/5 glycosides, 1/1 uronic acid, 10/20 amino acids, 0/2 aromatic chemicals, 0/1 carboxylic acid, 6/15 organic acids, 0/3 dicarboxylic acids, 1/2 esters, 0/1 antibiotic, 1/1 ribonucleotide, 2/3 nucleotides, 0/1 ether and 1/2 lipids (Table 1).
3.2. FAME Analysis
Bacillus nealsonii strain AAU1 was found to contain straight-chain and terminally branched saturated and mono-unsaturated fatty acids with a composition of 63.2%, 4.25% and 29.3%, respectively (Table 2).
Among the fatty acids measured, hexadecanoic acids (16:0/16:0 aldehyde, 16:0 DMA), tetradecanoic acid (14:0), mono-unsaturated 16:1w7c and 16:1w9c dominated. Bacillus nealsonii strain AAU1 contained higher amounts of straight-chain saturated fatty acid (i.e. 63.2%) than the reference. B. nealsonii FO-92T, B. licheniformis ATCC 14580, B. circulans ATCC 4513 and B. subtilis IAM 1026, contained 73.2, 89.1, 86.7, and 80.1% terminally branched saturated fatty acids, respectively.
3.3. Genome Sequencing and Assembly
Whole genome sequencing of the putative Bacillus iso-


Table 1. Substrate utilization profile of the culture as determined by AN-Biolog* Plate.
late yielded a total of 870,068 reads. Assembly using the GS browser resulted in 4.9 Mb sequence in 446 contigs, defined by a GC content of 35.1% (Table 3). Approx-
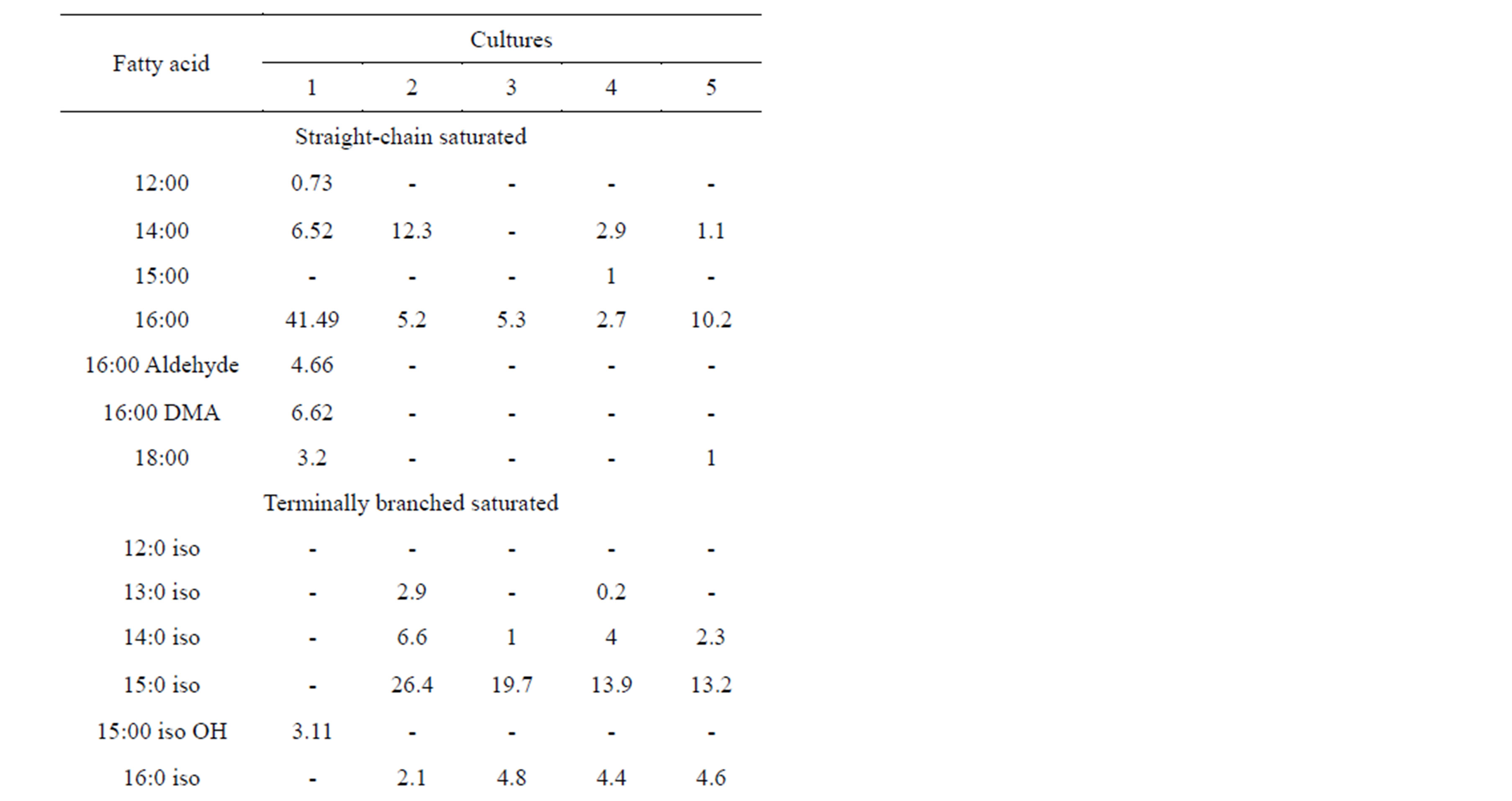
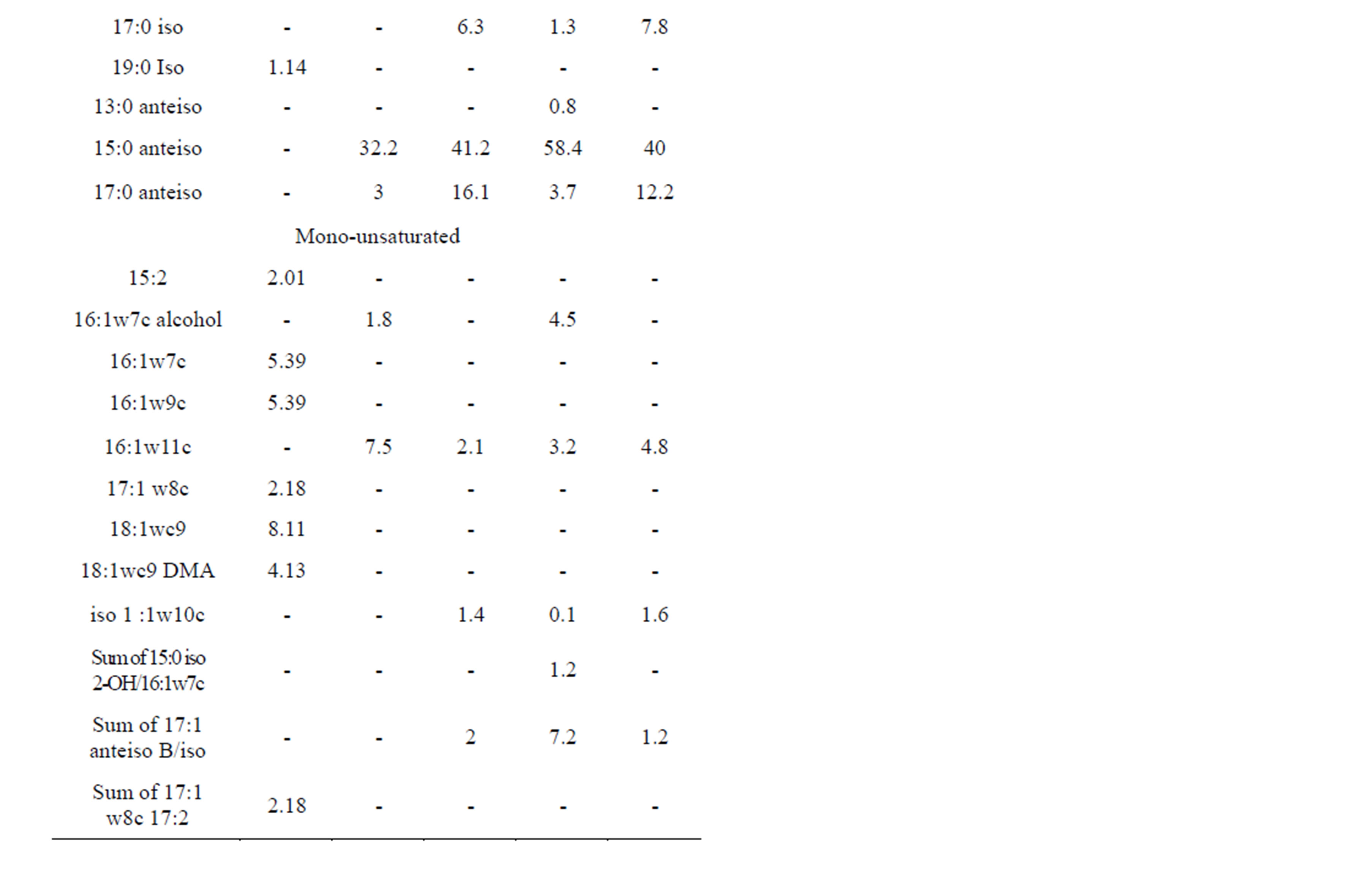
Table 2. Comparison of FAME with neighbors. 1-Our Culture, 2-B. nealsonii FO-92T, 3-B. licheniformis ATCC 14580, 4-B. circulans ATCC 4513, 5-B. subtilis IAM 1026.
imately 99% of the input bases were successfully aligned during assembly (General Genome Features described in Table 4). Details of number of reads and bases used for the genome assembly are mentioned in Table 5.
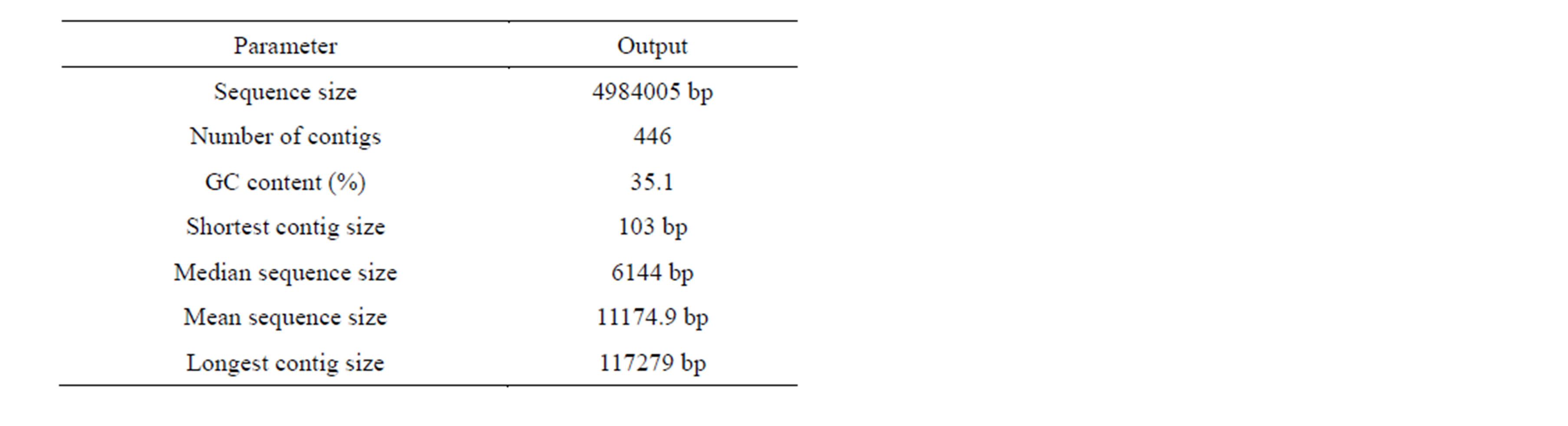

Table 3. Contig statistics of the Bacillus strain as determined from RAST server.
3.4. 16S rRNA and Housekeeping Genes
Comparative 16S rRNA gene sequence analysis using Local BLAST showed the highest level of similarity with the 16S rRNA gene sequence of the anaerobic Bacillus nealsonii DSM 15077 strain (98.50%). RDP results confirmed the taxonomy (Classified as Domain: Bacteria; phylum: Firmicutes; class: Bacilli; order: Bacillales; family: Bacillaceae and genus: Bacillus). Phylogenetic analysis showed the strain to be closely related to Bacillus nealsonii strain B29S based on 16S gene sequences (Figure 1).
While the coding sequences for housekeeping gene gyrase A showed the strain to be more closely related to Bacillus species like the pumilus, halodurans and subtilis (Figure 2). Comparison revealed approximately 78% identity between the strain AAU1 and the reference Bacillus species for the housekeeping genes targeted.
3.5. Gene Prediction and Annotation
Metabolic reconstruction subsystems were assembled to create a metabolic reaction network for the putative Bacillus isolate. Bacillus nealsonii AAU1 comprised of 5202 predicted coding sequences (CDS), out of which about 2034 (39.1%) were functional in specific subsystems. Subsystem analysis revealed the presence of 2994 genes involved in coding for functions related to various subsystems (Figure 3).
Comparison of metabolic characteristics of B. nealsonii with nearest species describe its taxonomic placement and adaptation properties (Table 6).
3.6. Cell Wall Components
Teichoic acids are unique to gram positive bacteria. The peptidoglycan composition predicted by the annotation analysis was similar to that described previously for other Bacillus species. such as the neutrophilic B. subtilis, including many acidic components along with genes coding for peptidoglycan components. The glycosyl transferase cluster (seven genes tuaA-tuaH related to teichuronic acid biosynthesis) were observed to be pre-

Table 4. General genome features and subsystem distribution.

Table 5. Genome assembly input/output details.

Figure 1. Phylogenetic tree based on 16S rRNA gene sequences of genus Bacillus. The bootstrap consensus tree inferred from 500 replicates is taken to represent the evolutionary history of the taxa analyzed. Branches corresponding to partitions reproduced in less than 50% bootstrap replicates are collapsed.

Figure 2. Phylogenetic tree based on gyrase A gene sequences of genus Bacillus. The bootstrap consensus tree inferred from 500 replicates is taken to represent the evolutionary history of the taxa analyzed. Branches corresponding to partitions reproduced in less than 50% bootstrap replicates are collapsed.
sent in the organism. Teichuronic acid is a copolymer comprising of alternate glucuronic acid and glutamate/ N-acetylmannosaminuronic acid. Genes involved in peptidoglycan synthesis such as mraY, murC-murG, ftsL, ddlA, cwlA and glnA were also observed to be present in the B. nealsonii AAU1 genome. Nine sequences encoding the enzymes related to diaminopimelate biosynthesis were also found to be present in the genome as well as genes coding for teichoic and lipoteichoic acid biosynthesis (details shown in full in Table 7).
3.7. ATP Binding Class of Proteins (ABC Transporters)
A large number of genes coding for members of the ABC transporter superfamily were observed in the genome, grouped by binding preference (sugars, proteins and other molecules required to be translocated across the cytoplasmic membrane). The genome encoded 27 sequences related to oligopeptide ATP binding proteins and 51 amino acid ATP binding proteins, including 13 permeases. Analysis also revealed the presence of zinc,
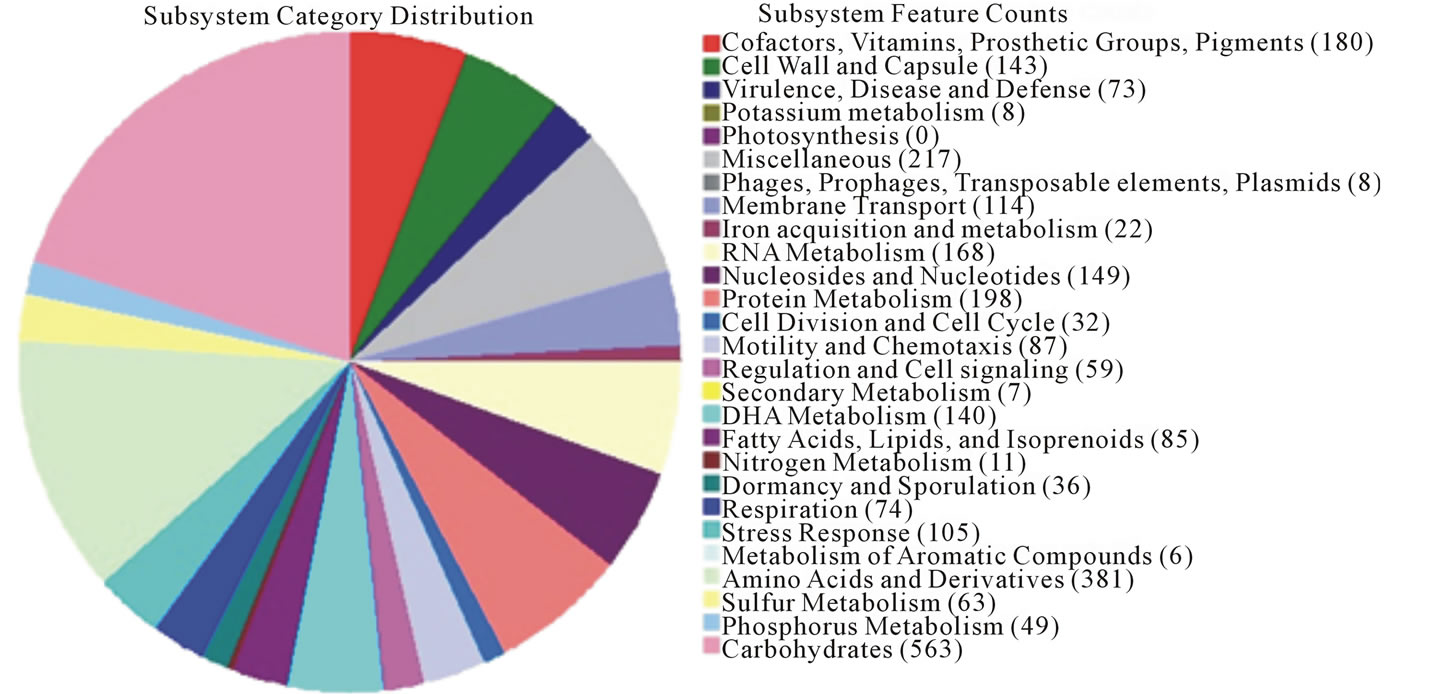
Figure 3. Feature counts of coding sequences falling under specific subsystem categories.

Table 6. Comparison of metabolic characteristics with neighbors. 1-Our Culture, 2-B. nealsonii FO-92T, 3-B. licheniformis ATCC 14580, 4-B. circulans ATCC 4513, 5-B. subtilis IAM 1026.
molybdenum and iron metal permeases, vitamin B12 permease, sulfonate permease, phosphate permease and others associated with translocation and absorption of polyamine molecules like putrescine and spermidine that are important for cell division and play many vital roles. A class of petrobactin was identified and hence the strain is likely to be positive for catechol containing siderophore formation.
3.8. Motility and Chemotaxis, Competence
A total of 77 CDS were present in the subsystem related to motility and chemotaxis. Of these 32 associated with motility and other flagellar genes for motor rotation proteins by proton motive force, basal body formation and flagellar biosynthesis proteins like flh and fli clusters. Others included the methyl accepting chemotaxis proteins and chemotaxis genes like cheA-cheD, cheV and cheR. Genome analysis also revealed the presence of CDS related to bacterial competency. Two CDS coding for competence transcription factors, along with four genes required for regulation of competence, were observed in the B. nealsonii strain AAU1 genome including sinR, active in many other Bacillus and Streptococcus strains [10]. Ten sequences coding for the late competence proteins comEA, comEB, comEC, comER, comGA-GD were also detected. Additionally, a cinA gene encoding the competence-inducible (cin) operon was detected.
3.9. Polyamine (Putrescine, Spermidine, Spermine) Transport and Biosynthesis
Nine of the putative ABC transporters were related to transport proteins integral to putrescine and spermidine utilization. These coded for the potA, potB, potC and potD protein cluster. The mreR family, which regulates the transport of polyamines and genes for spermidine synthase, adenosyl methionine decarboxylase and agma-
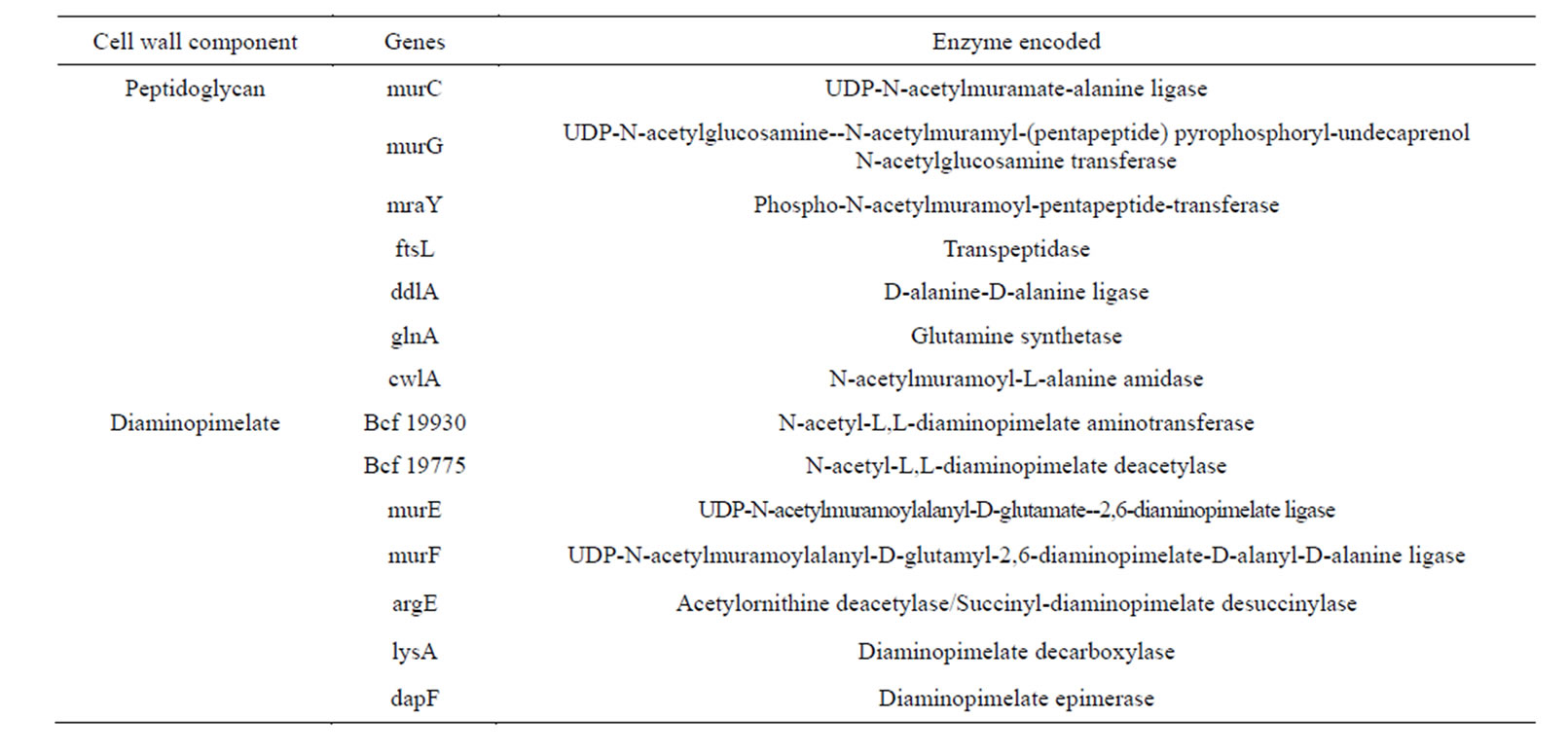
Table 7. Genes coding for enzymes related to peptidoglycan and Diaminopimelic acid component of the cell wall.
tinase, was also detected representing part of the polyamine synthesis pathway.
3.10. Sporulation
Gram positive bacteria produce intracellular structures called endospores by undergoing cellular differentiation in a process known as sporulation [11]. A large number of genes were found to be involved in the eight stage process of endospore formation and its regulation. Genome analysis of the B. nealsonii strain revealed the presence of four CDS encoding regulatory kinases responsible for initiation of sporulation. The genome also revealed the presence of 29 genes coding for spore germination protein factors and enzymes, including GerPA, GerPB, GerPC, GerPD, GerPE, GerPF, GerKA, GerKB, GerKC, YpeB and about 14 genes involved in spore coat development. Eleven CDS encoding small acid-soluble proteins (SASPs) with a functional role in protection were detected. In total, 133 CDS were observed in the genome coding for sporulation, germination, spore coat formation, maturation and responsible for coding transcriptional regulatory factors for the process.
4. DISCUSSION
4.1. Housekeeping Genes
Widespread use of gene sequencing for the identification of bacteria from complex microbial communities has increased the number of candidate new bacterial species. Some genes are shared by a vast majority of bacterial species, including the ribosomal genes and housekeeping genes such as rpoB, gyrA and homologous recombination-associated recA, offering possibilities as universal targets for identification and taxonomy [12]. Sequencing of the housekeeping gene rpoB is increasingly utilized as standard to confirm 16S rRNA based phylogenetic trees and identify closely related bacterial species [13]. The taxonomic resolution of this gene is reported to be more than three times greater than that of the 16S rRNA gene for bacterial genera such as Bacillus and Pseudomonas [14,15]. The phylogenetic analysis based on gyrase A gene sequence showed that the strain AAU1 is more closely related to B. pumilus and B. subtilis species, which are also the most closely related relatives whose whole genome sequences are already published as per the RAST analysis. In case the gyrase A gene sequence of reference Bacillus nealsonii strain was available, probably the strain AAU1 would have been observed to be holding a close position to the nealsonii species for the CDS of gyrA gene.
4.2. Cell Wall
The cell wall plays an important role in the viability and shape determination of bacterial organisms. Cell wall integrity and dynamics govern bacterial growth [16]. Peptidoglycan is the major component for most bacterial cell walls [17]. Teichoic acids (TAs) are reported to be fundamental components of the cell wall in many Grampositive bacteria, influencing surface antigenicity, polarity and hydrophobicity [18]. Teichuronic acid (TUA), a long chain polysaccharide composed of disaccharide repeating units, plays an important role in microbe/host interaction. As reported earlier, teichuronic acid synthetase is an enzyme complex comprising a cluster of seven genes (described above). The complex has been found to be present in the cytoplasmic phase of the cell membrane [19]. Acid polymers and other cell wall polysaccharides play important roles including protection of bacteria, signal transduction, concentration of metal ions from the surrounding environment, nutrient assimilation and interaction with the environment and the host defense system. As a result, gaining knowledge of the unique polysaccharide structures responsible for such interactions and ultimately understanding the genes and gene products responsible for cell wall component biosynthesis is of key interest [20].
4.3. ABC Binding Class of Proteins (ABC Transporters)
ABC transporters play a functional role in the translocation of solutes across membranes against ATP hydrolysis. They have been found to be important and the most highly represented class of genes in genomes of many Bacillus species including B. subtilis and B. halodurans [21], providing gram positive bacteria with protection against toxic substances and compensating in part for their single cell membrane. Petrobactin binding proteins and transporters were also observed and may be helpful in iron acquisition, catechol containing siderophore synthesis and growth during iron depleting conditions [22].
4.4. Motility and Chemotaxis; Competence
Bacterial organisms have developed many strategies to cope with environmental fluctuations and stresses. Examples include motility and chemotaxis mechanisms to improve nutrient access in limiting conditions, antimicrobial synthesis to improve competition with other microbes, development of competence and sporulation [23]. Correlation between bacterial flagellar motility proteins and regulation of the Entner-Doudoroff pathway by induction of methyl-accepting chemotaxis protein has been demonstrated in vitro [24]. The presence of flagellar proteins and methyl-accepting chemotaxis proteins in the B. nealsonii AAU1 strain genome confirms the active participation of enzymes like phosphoglycerate mutase, gluconolactonase, 2-dehydro-3-deoxygluconate kinase and other related enzymes in the non-phosphorylated alternative Entner-Doudoroff pathway used by some anaerobic organisms for sugar utilization.
Natural competence is the ability of organisms to take up DNA from their surroundings by spontaneous transformation, or when grown in favorable media. The study of natural competence can provide insight into the genetic basis of transformation [25], promoting focus on the Bacillus species capacity for transformation. For B. nealsonii strain AAU1 transformation can in part be facilitated by the presence of two genomic sinR gene sequences, encoding regulators of post-exponential-phase responses in competence and sporulation. The CoiA gene sequence observed in the genome codes for the synthesis of an ephemeral protein expressed specifically during competence and required for genetic transformation in Streptococcus pneumoniae, but not for DNA uptake. It has previously been reported that this gene is widely conserved among Gram-positive bacteria [26].
4.5. Polyamine (Putrescine, Spermidine, Spermine) Transport and Biosynthesis
Polyamines are an important requirement for growth in microorganisms and are said to be integral to nucleic acid and protein metabolism as they are cationic in nature and affect synthesis by binding negatively charged nucleic acids [27]. Bacillus subtilis strain 168 has been shown to have a single pathway to polyamine biosynthesis with agmatine as an intermediate comprising of arginine decarboxylase, speA and speE-speB operon, synthesizing spermidine synthase and agmatinase [28]. The presence of sequences coding for adenosylmethionine decarboxylase and agmanitase enzymes indicate the strain’s probable potential for polyamine biosynthesis. The ABC transporter genes observed in the genome also comprise the potABCD cluster involved in transport and utilization of spermidine and putrescine like polyamines, further confirming the presence of a polyamine metabolic subsystem in the B. nealsonii strain AAU1.
4.6. Sporulation
Bacterial endospores are complex structures conserved among gram positive bacteria characterized by low GC content including the Bacillus and Clostridium genera [29]. The main stimuli for spore formation include nutrient depletion, leading to endospore biogenesis for protection against extreme conditions like high temperature, UV radiation, dehydration, vacuum and high pressures [30,31]. The vegetative cell that represents the stage 0 of sporulation initiates the process. The DNA filament is formed and the autophosphorylation of kinases (four kinase genes as mentioned in the results) leads to phosphate transfer with the spo0F gene activating further stage 0 genes and initiating a cascade of germinating events, supported by genes involved in stages of cell division, pre-spore formation, coat synthesis, spore maturation and ultimately development of the endospore. The SASPs present in the genome are annotated as actively functional and have previously been reported to be capable of binding DNA and protecting it from damage. They also help maintain the core pH below that of the vegetative cell fluid. The core lytic enzyme coding sequences observed in the genome may be involved in the hydrolysis of the cortex at later germination with YpeB needed for the activity of other core lytic genes. The process of spore formation remains a topic of interest, providing insight into the ubiquitous global distribution of spores [30]. Bacterial endospores have also been studied as candidates for transfer of life between planets due to their extreme resistance and longevity properties [32].
4.7. Substrate Utilization
Many bacteria are capable of utilizing a large variety of organic substances. This capacity was compared for the B. nealsonii AAU1 strain with established Bacillus species (Table 6). AN-Biolog microtiter plates were initially developed to assist classification of bacterial isolates based on ability to oxidize distinct carbon sources. The method is also used to characterize the metabolic potential of microbial communities [33].
4.8. FAME Analysis
FAMEs have long been recognized as useful biochemical markers for bacterial classification and characterization [34]. The types and relative abundances of fatty acids produced within a cell are largely determined by an organism’s genotype and can be used for identification of different species [35]. Different culture conditions can result in high variability within FAME profiles [36]. The FAME profile for the B. nealsonii AAU1 strain was unfortunately ambiguous (Table 2) and could not identify the isolate. Comparison of the FAME analysis with other Bacillus species highlighted distinct fatty acid profiles in our isolate, indicating marked differences in fatty acid abundance. The relative proportions of unsaturated fatty acid markers such as 16:1w7c, which is commonly abundant in BcT FAME profiles, is primarily affected by saturated fatty acid precursor (16:0) concentration and oxygen availability [37].
5. CONCLUSIONS
Genetic and phylogenetic analyses recommend designation of isolate AAU1 as a novel B. nealsonii strain. The strict anaerobic conditions prevailing in the bovine rumen from where AAU1 was isolated may have resulted in genetic polymorphism influencing its biochemical FAME profile. Nonetheless, 16S rRNA and housekeeping gene similarities indicate phylogenetic placement within the Bacillus genus proximal to B. nealsonii, corroborated by AN-Biolog profiling.
“Nucleotide sequence data reported are available in the GenBank database under the accession number ASRU00000000”.
REFERENCES
- Pandya, P.R., et al. (2010) Bacterial diversity in the rumen of Indian Surti buffalo (Bubalus bubalis), assessed by 16S rDNA analysis. Journal of Applied Genetics, 51, 395-402. http://dx.doi.org/10.1007/BF03208869
- Pattnaik, P., et al. (2001) Purification and characterization of a bacteriocin-like compound (Lichenin) produced anaerobically by Bacillus licheniformis isolated from water buffalo. Journal of Applied Microbiology, 91, 636-645. http://dx.doi.org/10.1046/j.1365-2672.2001.01429.x
- Earl, A.M., et al. (2012) Whole-genome sequences of Bacillus subtilis and close relatives. Journal of Bacteriology, 194, 2378-2379. http://dx.doi.org/10.1128/JB.05675-11
- Brown, C.T., et al. (2011) Whole-genome sequencing and phenotypic analysis of Bacillus subtilis mutants following evolution under conditions of relaxed selection for sporulation. Applied and Environmental Microbiology, 77, 6867-6877. http://dx.doi.org/10.1128/JB.05675-11
- Venkateswaran, K., et al. (2003) Bacillus nealsonii sp. nov., isolated from a spacecraft-assembly facility, whose spores are gamma-radiation resistant. International Journal of Systematic and Evolutionary Microbiology, 53, 165-172. http://dx.doi.org/10.1099/ijs.0.02311-0
- Hungate, R.E., Smith, W. and Clarke, R.T. (1966) Suitability of butyl rubber stoppers for closing anaerobic roll culture tubes. Journal of Bacteriology, 91, 908-909.
- Latham, M.J., et al. (1978) Adhesion of Bacteroides succinogenes in pure culture and in the presence of Ruminococcus flavefaciens to cell walls in leaves of perennial ryegrass (Lolium perenne). Applied and Environmental Microbiology, 35, 1166-1173.
- Tamura, K., et al. (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Molecular Biology and Evolution, 24, 1596-1599. http://dx.doi.org/10.1093/molbev/msm092
- Aziz, R.K., et al. (2008) The RAST server: Rapid annotations using subsystems technology. BMC Genomics, 9, 75. http://dx.doi.org/10.1186/1471-2164-9-75
- Mobberley, J., et al. (2010) Lysogeny and sporulation in Bacillus isolates from the Gulf of Mexico. Applied and Environmental Microbiology, 76, 13. http://dx.doi.org/10.1128/AEM.01710-09
- Wolska, K.I., Grudniak, A.M. and Kraczkiewicz-Dowjat, A. (2007) Genetic and physiological regulation of bacterial endospore development. Polish Journal of Microbiology, 56, 11-17.
- Drancourt, M. and Raoult, D. (2005) Sequence-based identification of new bacteria: A proposition for creation of an orphan bacterium repository. Journal of Clinical Microbiology, 43, 4311-4315. http://dx.doi.org/10.1128/JCM.43.9.4311-4315.2005
- Smuts, H.E. and Lastovica, A.J. (2011) Molecular characterization of the 16S rRNA gene of Helicobacter fennelliae isolated from stools and blood cultures from Paediatric patients in South Africa. The Journal of Pathology, 2011, 217376.
- Ait Tayeb, L., et al. (2005) Molecular phylogeny of the genus Pseudomonas based on rpoB sequences and application for the identification of isolates. Research in Microbiology, 156, 763-773. http://dx.doi.org/10.1016/j.resmic.2005.02.009
- Ki, J.S., Zhang, W. and Qian, P.Y. (2009) Discovery of marine Bacillus species by 16S rRNA and rpoB comparisons and their usefulness for species identification. Journal of Microbiological Methods, 77, 48-57. http://dx.doi.org/10.1016/j.mimet.2009.01.003
- Hayhurst, E.J., et al. (2008) Cell wall peptidoglycan architecture in Bacillus subtilis. Proceedings of the National Academy of Sciences of the United States of America, 105, 14603-14608. http://dx.doi.org/10.1073/pnas.0804138105
- Lahooti, M. and Harwood, C.R. (1999) Transcriptional analysis of the Bacillus subtilis teichuronic acid operon. Microbiology, 145, 3409-3417.
- Archibald, A.R., et al. (1989) Cell wall composition and surface properties in Bacillus subtilis: Anomalous effect of incubation temperature on the phage-binding properties of bacteria containing varied amounts of teichoic acid. Journal of General Microbiology, 135, 667-673.
- Deng, L.L., et al. (2010) The cell wall teichuronic acid synthetase (tuas) is an enzyme complex located in the cytoplasmic membrane of Micrococcus luteus. Biochemistry Research International, 2010, 395758.
- De Kimpe, S.J., et al. (1995) The cell wall components peptidoglycan and lipoteichoic acid from Staphylococcus aureus act in synergy to cause shock and multiple organ failure. Proceedings of the National Academy of Sciences of the United States of America, 92, 10359-10363. http://dx.doi.org/10.1073/pnas.0804138105
- Takami, H., et al. (2000) Complete genome sequence of the alkaliphilic bacterium Bacillus halodurans and genomic sequence comparison with Bacillus subtilis. Nucleic Acids Research, 28, 4317-4331. http://dx.doi.org/10.1093/nar/28.21.4317
- Carlson Jr., P.E., et al. (2010) Genetic analysis of petrobactin transport in Bacillus anthracis. Molecular Microbiology, 75, 900-909. http://dx.doi.org/10.1111/j.1365-2958.2009.07025.x
- Hamoen, L.W., Venema, G. and Kuipers, O.P. (2003) Con- trolling competence in Bacillus subtilis: Shared use of regulators. Microbiology, 149, 9-17. http://dx.doi.org/10.1099/mic.0.26003-0
- Pruss, B.M., et al. (2003) FlhD/FlhC is a regulator of anaerobic respiration and the Entner-Doudoroff pathway through induction of the methyl-accepting chemotaxis protein Aer. Journal of Bacteriology, 185, 534-543. http://dx.doi.org/10.1128/JB.185.2.534-543.2003
- Dubnau, D. (1991) Genetic competence in Bacillus subtilis. Microbiological Reviews, 55, 395-424.
- Desai, B.V. and Morrison, D.A. (2007) Transformation in Streptococcus pneumoniae: Formation of eclipse complex in a coiA mutant implicates CoiA in genetic recombination. Molecular Microbiology, 63, 1107-1117. http://dx.doi.org/10.1111/j.1365-2958.2006.05558.x
- Tabor, C.W. and Tabor, H. (1976) 1,4-Diaminobutane (putrescine), spermidine, and spermine. Annual Review of Biochemistry, 45, 285-306. http://dx.doi.org/10.1146/annurev.bi.45.070176.001441
- Sekowska, A., Bertin, P. and Danchin, A. (1998) Characterization of polyamine synthesis pathway in Bacillus subtilis 168. Molecular Microbiology, 29, 851-858. http://dx.doi.org/10.1046/j.1365-2958.1998.00979.x
- Errington, J. (2003) Regulation of endospore formation in Bacillus subtilis. Nature Reviews Microbiology, 1, 117- 126. http://dx.doi.org/10.1038/nrmicro750
- Nicholson, W.L., et al. (2000) Resistance of Bacillus endospores to extreme terrestrial and extraterrestrial environments. Microbiology and Molecular Biology Reviews, 64, 548-572. http://dx.doi.org/10.1128/MMBR.64.3.548-572.2000
- Stragier, P. and Losick, R. (1996) Molecular genetics of sporulation in Bacillus subtilis. Annual Review of Genetics, 30, 297-341. http://dx.doi.org/10.1146/annurev.genet.30.1.297
- Horneck, G., Rettberg, P., Rabbow, E., Strauch, W., Seckmeyer, G., Facius, R., Reitz, G., Strauch, K. and Schott, J.U. (1996) Biological dosimetry of solar radiation for different simulated ozone column thicknesses. Journal of Photochemistry and Photobiology B, 32, 189-196. http://dx.doi.org/10.1016/1011-1344(95)07219-5
- Smalla, K., et al. (1998) Analysis of BIOLOG GN substrate utilization patterns by microbial communities. Applied and Environmental Microbiology, 64, 1220-1225.
- Welch, D.F. (1991) Applications of cellular fatty acid analysis. Clinical Microbiology Reviews, 4, 422-438.
- Vandamme, P., Pot, B., Gillis, M., de Vos, P., Kersters, K. and Swings, J. (1996) Polyphasic taxonomy, a consensus approach to bacterial systematics. Microbiology and Molecular Biology Reviews, 60, 407-438.
- Venkateswaran, K., Moser, D.P., Dollhopf, M.E., Lies, D.P., Saffarini, D.A., MacGregor, B.J., Ringelberg, D.B., White, D.C., Nishijima, M., Sano, H., Burghardt, J., Stackebrandt, E. and Nealson, K.H. (1999) Polyphasic taxonomy of the genus Shewanella and description of Shewanella oneidensis sp. nov. International Journal of Systematic and Evolutionary Microbiology, 49, 705-724. http://dx.doi.org/10.1099/00207713-49-2-705
- Kaneda, T. (1968) Fatty acids in the genus Bacillus. II. Similarity in the fatty acid compositions of Bacillus thuringiensis, Bacillus anthracis, and Bacillus cereus. Journal of Bacteriology, 95, 2210-2216.
NOTES
*Nathani N. M. & Duggirala S. M. equally contribute to the article.

