Advances in Bioscience and Biotechnology
Vol.3 No.8(2012), Article ID:25831,7 pages DOI:10.4236/abb.2012.38142
A novel strain of Trichoderma viride shows complete lignocellulolytic activities
![]()
Enzyme Technology Laboratory, Biotechnology Division, Department of Botany, University of Calicut, Malappuram, India
Email: *sailasben@yahoo.co.in
Received 5 September 2012; revised 11 October 2012; accepted 29 November 2012
Keywords: Trichoderma viride; Morphology; Liquid Fermentation; Bluish-Green Pigment
ABSTRACT
In this study we describe a novel dark-green strain of Trichoderma viride exhibiting complete ensemble of cellulase, hemicellulase and ligninase activities on specific plate assays. To assess the cellulase production in detail, basal salt medium (BSM) was fortified with synthetic (carboxymethyl cellulose (CMC), glucose, sucrose, dextrose, lactose or maltose) and natural (flours of banana, banana peel, jack seed, potato or tapioca) carbon as well as nitrogen (yeast extract, beef extract, peptone, NaNO3 or NH4NO3) sources. Temperature and pH optima were 28˚C and 4, respectively for the growth of the fungus in CMC-BSM with 137 U/mL cellulase activity, which was enhanced to 173 U/mL at 1.25% CMC concentration. Flours of potato and banana peel supported comparable yields of cellulase to that of CMC, while sodium nitrate was the preferred nitrogen source. The water soluble bluish-green pigment (a probable siderophore) extracted from the spores showed an absorption maximum at 292 nm. To sum up, the complete lignocellulolytic potential of this fungus offers great industrial significance, coupled with the production of a new pigment.
1. INTRODUCTION
Lignocellulosic biomass comprises cellulose, hemicellulose and lignin, of which lignin is the most recalcitrant to biodegradation and only higher fungi are capable of degrading this polymer via an oxidative process involving an array of oxidases, peroxidases and hydrolytic enzymes [1,2]. Cellulose, hemicellulose and lignin form structures called microfibrils, which are organized into macrofibrils that mediate structural stability in the plant cell wall [2]. The complex lignin polymer forms a cementing envelop around cellulose in lignocellulosic wood; thus the success of the bioconversion of lignocelluloses into fermentable sugars lies in the removal of lignin wrap around the cellulose fibers packed inside. Cellulose is a fibrous, insoluble and crystalline polysaccharide consisting of D-glucose residues linked by β-1, 4-glucosidic bonds; owing to its highly ordered crystalline structure, cellulose is more resistant to hydrolysis than hemicellulose. Hemicellulose macromolecules are often polymers of pentoses (xylose and arabinose), hexoses (mostly mannose) and a number of sugar acids, while cellulose is a homogenous polymer of glucose [3]. Cellulose is the most abundant biopolymer in nature, being degraded to glucose through the synergistical hydrolysis of three classes of cellulases, viz., including endo-β-1,4-glucanase (EC 3.2.1.4), exoglucanase or cellobiohydrolase (EC 3.2.1.91) and β-glucosidase (EC 3.2.1.21). Glucose from the hydrolysis of cellulose can easily be fermented to useful industrial products such as ethanol, lactic acid, single cell protein and other value-added products [4]. There are different methods for the production of lowcost enzymes by the activity of microorganisms.
Some members of all classes of microorganisms, viz., fungi, actinomycetes and bacteria are known to produce cellulase isoforms [5,6]. Among them, mycelial fungi constitute a group unique of microorganisms showing prominent lignocellulolytic activity. Lignocellulolytic fungi are often divided into three groups, viz., white rot, brown rot and soft rot fungi. Trichoderma spp., the ascomycetous green-spored having cosmopolitan distribution are classified under white rot fungi. The surface colony colour is white and scattered greenish patches become visible as the conidia are formed and may form concentric rings. The highly branched conidiophores bear phialides singly or in groups. Trichoderma spp. are ubiquitous colonizers of cellulosic materials and can thus often be found wherever decaying plant material is available as well as in the rhizosphere of plants, where they can also induce systemic resistance against pathogens [6]. At present, the various strains of Trichoderma spp. are recoganized by the researchers as the steadiest and safest fungi for the production of cellulase and hemicellulases. Mostly T. reesei and its variants are widely employed for the commercial production of hemicellulases and cellulases. T. reesei might be a good producer of hemi-cellulolytic and cellulolytic enzymes but is unable to degrade lignin. The ability of fungi to degrade lignocellulosic materials is due to their highly efficient enzymatic system [7]. T. viride have mostly been studied for cellulase production. There exist many studies describing the optimization of process parameters for the production of cellulase using submerged fermentation from T. viride [8]. Species of Trichoderma are shown to have lignolytic, hemicellulolytic and cellulolytic activities, but no organism is reported yet having the embodiment of all these enzyme complexes together. Based upon this background, the specific objectives of this study are to: 1) Confirm the novel isolate as T. viride by morphological characteristics; 2) Check the lignocellulolytic activities of T. viride; 3) Analyze cellulase production potential of T. viride in detail; and 4) Check the nature of pigment produced by T. viride.
2. MATERIALS AND METHODS
2.1. Microorganism
Pure T. viride culture procured from the Kerala Agricultural University, Mannuthi, Kerala (Receipt No.17 dated 15-12-2011) was used for the present study. Stock cultures were maintained on potato dextrose agar (PDA) slants at 4˚C. Analytical grade chemicals from Merck India Ltd. and Himedia were used.
2.2. Morphological Characterization
Morphological characteristics of T. viride were determined by lactophenol cotton blue staining under binocular microscope attached with Image Analyzer (Nikon Eclipse 400). Phase-contrast images of T. viride spores were taken using Leica M80 (Germany).
2.3. Screening for Lignocellulosic Activity
6 day old T. viride on PDA slants was cultured on basal mineral salt medium (BSM) containing (g/L) NaNO3—2, K2HPO4—1, KCl—0.5, MgSO4·7H2O—0.5, proteose peptone—2, agar—20 containing 0.5% CMC (for cellulase activity), xylan (for hemicellulase activity) or lignin (for ligninase activity) was used for preliminary screening of cellulase, hemicellulase or ligninase activities, respectively. The agar plates were incubated at 28˚C. Growth after incubation indicates lignocellulolytic activity of T. viride.
2.4. Screening for Cellulolytic Activity on CMC Medium
Cellulolytic activity of T. viride on CMC medium was determined qualitatively by iodine plate assay. 6 day old T. viride from PDA slant were cultured on BSM-agar medium containing CMC for enrichment and incubated for 4 days, then the plates were flooded with iodine solution (containing 1% iodine crystals and 2% potassium iodide), incubated at 30˚C for 15 min, and excess stain was washed off for visual observation. Cellulolytic activity was indicated by the clear zone around the colonies.
2.5. Screening for Hemicellulase Activity on Xylan Medium
Hemi-cellulolytic activity of T. viride on xylan medium was determined qualitatively by congo-red plate assay. 6 day old T. viride on PDA slant was cultured on BSM agar containing xylan and incubated for 4 days, then the plate were flooded with 10 ml Congo-red (0.1%) solution, incubated for 15 min, then excess stain was drained off, and 10 ml of 1 M NaCl was added for destaining; opaque clear zone formation around the colonies indicates positive result.
2.6. Screening for Ligninase Activity on Lignin Medium
6 day old T. viride from PDA slant was cultured on BSM containing lignin for enrichment. T. viride culture was grown for 5 - 6 days on BSM-lignin medium; growth on lignin medium indicates positive result.
2.7. Pigment Extraction
Pigmented biomass of five days old culture on PDA agar plate was collected in various solvents (distilled water, acetone, hexane, chloroform or methanol) and stirred well on a magnetic stirrer (10 min), followed by centrifugation (9000 × g for 15 min, 4˚C). The supernatant was pooled and its absorption spectrum was determined using UV-visible spectrophotometer (Elico BL 200 double beam biospectrophotometer).
2.8. Cellulase Production Characteristics of T. viride
For making the inoculum uniform; spore suspension in sterile ddH2O was stirred well on a magnetic stirrer aseptically for 5 min, and number of spores in 1 mL of suspension was calculated by counting the spores using Neubauer haemocytometer. Different parameters like effects of incubation period, pH, temperature, CMC concentration, synthetic carbon sources, natural carbon sources and nitrogen sources were optimized.
For initial studies, BSM supplemented with CMC (0.5%) was used. In all studies, 100 µL of spore suspension (~7.2 × 106 spores/mL) was used for 10 mL medium. All flasks were incubated in an environmental shaker (Scigenics Biotech) at 150 rpm, 28˚C for 10 days. Various culture parameters studied for cellulase production were: pH (1 to 8 using different buffers), temperature (25˚C, 28˚C, 31˚C, 33˚C, 35˚C and 37˚C), CMC concentrations (0.75%, 1.0%, 1.25% and 1.5%), various synthetic carbon sources (0.5% glucose, lactose, maltose, dextrose and sucrose), natural carbon sources (0.5% banana flour, banana peel flour, tapioca flour, potato flour and jack seed flour) and nitrogen sources (0.2% peptone, beef extract, yeast extract, sodium nitrate and ammonium nitrate). Growth and yield were monitored up to 7 days. The culture broth was centrifuged (9000 × g for 15 min at 4˚C) and the supernatant was used for cellulase assay.
2.9. Cellulase Assay
Enzyme production was quantitatively measured by DNS (3,5-dinitrosalicylic acid) spectrophotometric assay by the method of Miller [9]. Fermented culture broth was centrifuged at 9000 × g for 10 min at 4˚C and supernatant was used as crude enzyme solution for assays. Reaction mixture contains 0.5 mL of 1% CMC in 0.1 M sodium citrate buffer (pH, 4.8) and 0.5 mL of crude enzyme solution added, then incubated at 50˚C for 30 min. The reaction was stopped by adding 3 mL DNS and incubated in a boiling water bath for 5 min and cooled. The absorbance was measured at 540 nm (Elico double beam bl 200 bio spectrophotometer). One unit (U/mL) of cellulase activity is defined as the amount of protein (cellulase) required to liberate 1 micromole of reducing sugar (D-glucose) from CMC per min under the assay conditions. Cellulase activity was calculated using the formula,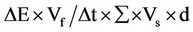 . Where, ΔE = absorbance at 540 nm, Vf = final volume of reaction mixture including DNS, Vs = crude supernatant (ml) containing cellulase used, Δt = incubation time for hydrolysis, ∑ = extinction coefficient of glucose (0.0026), d = diameter of cuvette.
. Where, ΔE = absorbance at 540 nm, Vf = final volume of reaction mixture including DNS, Vs = crude supernatant (ml) containing cellulase used, Δt = incubation time for hydrolysis, ∑ = extinction coefficient of glucose (0.0026), d = diameter of cuvette.
2.10. Statistics
All studies were repeated at least three times to check the reproducibility of the results, and average values were given with standard deviation. Microsoft Excel was used to draw the figures.
3. RESULTS AND DISCUSSION
Results shown here are the average values of at least 3 independent experiments. The fungal culture started to grow as white mycelia on PDA plate, then spread all over the surface of the medium as a dark-green mat. It was stained using lactophenol cotton blue and observed under Image Analyzer. The mycelia, conidiophores bearing warted phialides singly or in clusters and numerous conidiospores were observed (Figure 1A). Usually the phialides were ellipsoidal or flask shaped inflated at the base dark-green coloured spores with smooth outer wall were found to be spherical and biconcave, resembling human erythrocytes (Figures 1B and C). This preparation was observed under phase-contrast microscope. Bissette [10] described that the species in Trichoderma (Ascomycetes, Hypocreales) have narrow and flexuous conidiophores and branches, with branches and phialides uncrowded, frequently paired, and seldom with more than three elements in a whorl. Its type species viride typically produced warted conidia [11], as we described (Figure 1A). Thus, the novel fungal culture used in this study can be confirmed as T. viride.
3.1. Lignocellulolytic Activity
Figure 2 clearly shows that the fungus harbours an en-
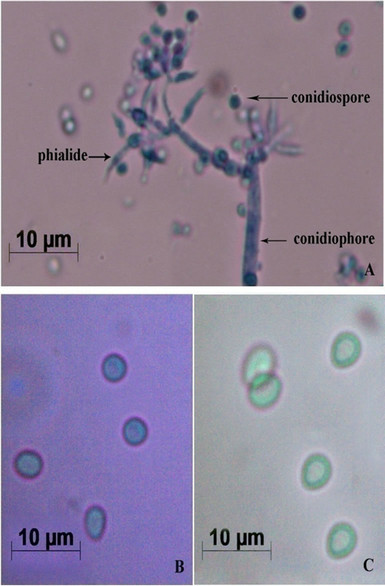
Figure 1. Morphological characterization of T. viride. (A) Mycelia bearing conidiophores and conidiospores after staining with lactophenol cotton blue (Images by Image Analyzer, Scale bar = 10 µM (Nikon Eclipse E400, Towa Optical, Japan, fitted with Nikon digital camera, DXM 1200F, Japan); (B) Conidiospores under Bright Field Microscope (Scale bar = 10 µM, Leica M80, Germany); and (C) Conidiospores under Phase Contrast Microscope (Scale bar = 10 µM, Leica M80, Germany).
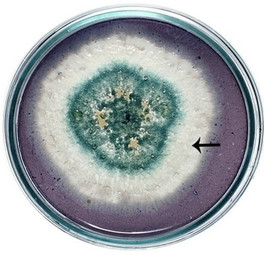
semble of enzymes for the complete degradation of lignocelluloses. Upon iodine plate assay of T. viride on BSM-CMC agar plate showed a clear zone around the mycelial colony, which indicates cellulolytic activity (Figure 2(a)). Xylan degradation around the colonies appeared as yellow and opaque zone against a reddish purple colour for undegraded xylan (Figure 2(b)). On lignin medium, mycelia were grown as bluish-green patches (Figure 2(c)), however, its growth was not as fast as in CMC or xylan medium. These tripartite plate assays clearly show that T. viride harbours an ensemble of lignocellulolytic enzymes for the complete degradation of lignocellulosic materials. The biodegradation of wood constituents is currently understood as a multienzymatic process with the mediation of small molecules [12]. Cellulolytic fungi can easily be screened within two days for the production of cellulolytic enzymes (especially, endoglucanase and exoglucanase) by staining technique or by measuring the amount of reducing sugar (glucose) produced with the dinitrosalicyclic acid reagent method [13]. Thus, we employed DNS method for analyzing the cellulase activity of the culture in liquid media.
3.2. Cellulase Production by T. viride
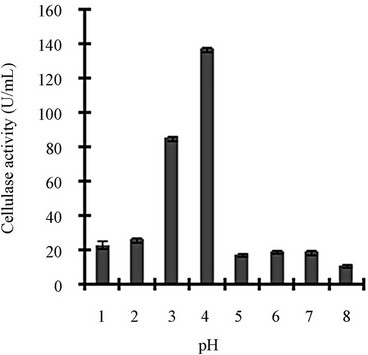
Figure 3 shows a view of the cellulase production profile of the culture in BSM supplemented with various carbon and nitrogen sources. Optimum pH was 4, with maximum yield 137 U/mL (Figure 3(a)). Like pH, temperature also played a crucial role, whose optimum was 28˚C with 128 U/mL at pH 4 (Figure 3(b)). Optimum concentration of CMC was 1.25% in the BSM, at this concentration the cellulase yield was 173 U/mL (Figure 3(c)). However, when CMC was replaced by simple sugars, the yield was reduce tremendously (Figure 3(d)); on the contrary, natural starch supplements (0.5%) showed best improvement in cellulase production (148 U/mL), which was comparable to that of CMC (Figure 4(e)). Simple nitrogen source like sodium nitrate (0.2%) was the preferred nitrogen source (Figure 4(f)).
Optimization of the time course is of prime importance for the biosynthesis of cellulases by fungi [14]. CMC is the preferred carbon source used for the selective enrichment culture of cellulase producing microorganisms. Using CMC as the carbon source, Gashe [15] showed that the CMCase (cellulose) activity of Trichoderma sp. as 167 U/mL. The incubation period was directly related to the production of enzymes and other metabolites up to a certain extent, and we showed that maximum cellulase activity was mostly obtained on the day 6. Aspergillus niger KK2 showed CMCase (129 U/g), β-glucosidase (100 U/g) and β-xylosidase (193 U/g) activities concurrently after 5 to 6 days of solid-state fermentation [16], which is comparable to our results. Most of the filamentous fungi, especially Trichoderma spp. have been found to be able to grow and metabolize within a range of pH, 3 to 5 [17]. At the declining phase of cellulase yield, acidic proteins would be released that would adversely affect the yield and fermentation process [4]. Carbon sources in majority of commercial cellulase fermentations are cellulosic biomass including straw, spent hulls of cereals and pulses, rice or wheat bran, rice or wheat straw, sugarcane bagasse, water hyacinth, paper industry waste and other cellulosic biomass [18]. Other commonly used substrates are wheat bran, orange peel, corn cob residue, rice straw, sugarcane bagasse, etc. [19]. Lactose, cellobiose, sophorose, gentibiose, are the known inducer of cellulase production but only lactose can be used for economically feasible industrial production [19]. Though addition of organic nitrogen sources such as beef extract and peptone resulted in increased growth and cellulase production, they were not effective replacement for inorganic nitrogen sources because of their higher cost [20]. High concentration of nitrogen sources may lead to vitrification (the medium appears to be yellow and glassy) that is usually unsuitable for the micro-organisms.
 (a)
(a) (b)
(b) (c)
(c)
Figure 2. Lignocellulolytic activities of T. viride. (a) Iodine plate assay showing cellulase activity on 4th day of incubation at 28˚C; (b) Congo-red plate assay showing xylanase activity on 4th day of incubation at 28˚C; and (c) Growth of T. viride on lignin medium on 6th day of incubation at 28˚C.
 (a)
(a)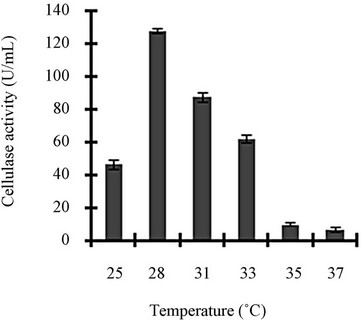 (b)
(b) (c)
(c)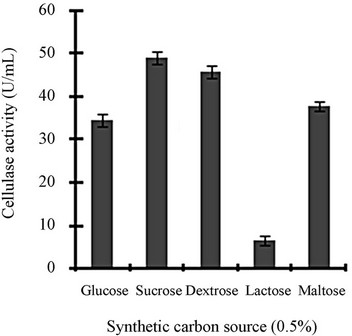 (d)
(d) (e)
(e) (f)
(f)
Figure 3. Optimization of parameters for cellulase production: (a) Effect of pH on cellulase production from T. viride on BSM supplemented with 0.5 % CMC on 6th day of incubation at 150 rpm and 28˚C; (b) Effect of temperature on cellulase production from T. viride on BSM supplemented with 0.5% CMC on 6th day of incubation at 150 rpm, 28˚C and pH 4; (c) Effect of CMC concentration on cellulase production from T. viride on BSM on 6th day of incubation at 150 rpm, 28˚C and pH 4; (d) Effect of synthetic carbon sources on cellulase production from T. viride on BSM on 6th day of incubation at 150 rpm, 28˚C and pH 4; (e) Effect of natural carbon sources (0.5%) on cellulase production from T. viride on BSM on 6th day of incubation at 150 rpm, 28˚C and pH 4, where BF (banana flour), BP (banana peel), JF (jack seed flour), PF (potato flour), TF (tapioca flour); and (f) Effect of nitrogen sources on cellulase production from T. viride on BSM supplemented with 0.5% CMC on 6th day of incubation at 150 rpm, 28˚C and pH 4, where YE (yeast extract) and BE (beef extract).
 (a)
(a)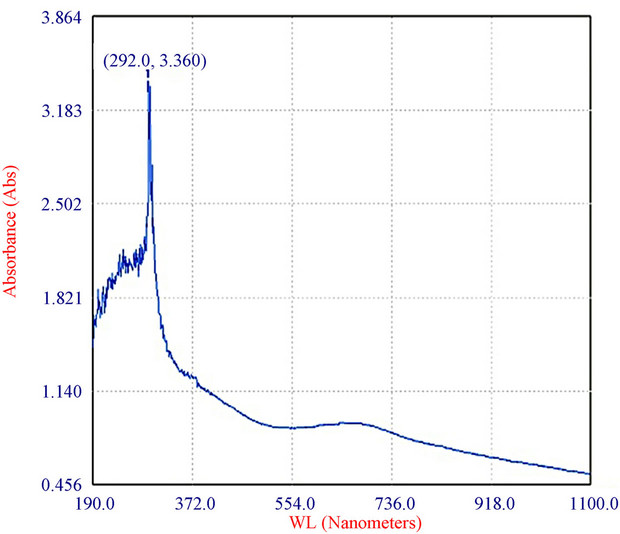 (b)
(b)
Figure 4. Characterization of the bluish-green pigment: (a) Pigment extracted from spores of T. viride in distilled water and centrifuged (9000 × g, 15 min); and (b) Absorption spectrum of pigment of T. viride extracted as above (by ELICO BL 200 Double Beam Bio Spectrophotometer).
3.3. Pigment Production
The fungus used in this study produced a water-soluble greenish pigment (Figure 4(a)). In fact the mycelia were colourless and only the spores were colored. Upon stirring, the pigment bound to the wall of the spores partially diffused in to water, and yet more pigment diffused into it after centrifugation. The absorption maximum of the pigment was at 292 nm (Figure 4(b)). T. viride is shown to produce variously coloured pigments, which range from green [21], yellow [22] to brown [23]. Very recently, Xiaoyi et al., [22] showed optimization of fermentation conditions in liquid state for a T. viride strain to produce yellow pigment, the viridin. From this, the pigment produced by the strain reported herein seems to be a new one, which needs to be elucidated further.
4. CONCLUSION
Characteristics of the fruiting bodies show that the novel strain described in this study is T. viride. Form the screening tests, it is evident that apart from cellulases, it is an efficient producer of ligninases and hemicellulases. From the optimization studies in the liquid medium, T. viride shows good cellulase yield. Moreover, this strains produces an extractable and water soluble bluish-green pigment, probably a siderophore, which needs to be elucidated. Briefly, this study shows that T. viride we used in this study is a potent strain for the complete degradation of lignocelluloses and subsequent fermentation for valuable products like ethanol, preferably by solid-state fermentation.
5. ACKNOWLEDGEMENTS
The authors are thankful to the Kerala State Council for Science, Technology and Environment for a research grant, No. (T) 422/SRS/2009/- CSTE.
REFERENCES
- Martinez, D., Larrondo, L.F., Putnam, N., Gelpke, M.D.S., Huang, K., Chapman, J., Helfenbein, K.G., Ramaiya, P., Detter, J.C., Larimer, F., Coutinho, P.M., Henrissat, B., Berka, R., Cullen, D. and Rokhsar, D. (2004) Genome sequence of the lignocellulose degrading fungus Phanerochaete chrysosporium strain RP78. Nature Biotechnology, 22, 695-700. doi:10.1038/nbt967
- Rubin, E. (2008) Genomics of cellulosic biofuels. Nature, 454, 841-845. doi:10.1038/nature07190
- Howard, R.L. and Abotsi, E. (2003) Issues of bioconversion and enzyme production. African Journal of Biotechnology, 2, 602-619.
- Chandra, M.A., Karala, P.K., Sharma and Sangwan, R.S. (2009) Cellulase production by six Trichoderma spp., fermented on medicinal plant processings. Journal of Industrial Microbiology and Biotechnology, 36, 605-609. doi:10.1007/s10295-009-0544-9
- Glick, B.R. and Pasternak, J.J. (1989) Isolation, characterization and manipulation of cellulose gene. Biotechnology Advances, 7, 361-386. doi:10.1016/0734-9750(89)90180-8
- Kubicek, C.P., Komon-Zelazowska, M. and Druzhinina, I.S. (2008) Fungal genus Hypocrea Trichoderma: From barcodes to biodiversity. Journal of Zhejiang University Science, 9, 753-763. doi:10.1631/jzus.B0860015
- Sánchez, C. (2009) Lignocellulosic residues: Biodegradation and bioconversion by fungi. Biotechnology Advances, 27, 185-194. doi:10.1016/j.biotechadv.2008.11.001
- Gautam, S.P. (2010) Optimization of the medium for the production of cellulase by the Trichoderma viride using submerged fermentation. International Journal of Environmental Science, 1, 656-665.
- Miller, G.L. (1959) Use of dinitrosalicyclic acid reagent for determination of reducing sugar. Analytical Chemistry, 31, 421-428. doi:10.1021/ac60147a030
- Bissett, J. (1991) A revision of the genus Trichoderma. II. Infrageneric classification. Canadian Journal of Botany, 69, 2357-2372. doi:10.1139/b91-297
- Lieckfeldt, E., Samuels, G.J., Nirenberg, H.I. and Petrini, O. (1999) A morphological and molecular perspective of Trichoderma viride: Is it one or two species? Applied and Environmental Microbiology, 65, 2418-2428.
- Leonowicz, A., Matuszewska, A., Luterek, J., Ziegenhagen, D., Wojtaś-Wasilewska, M., Cho, N.S., Hofrichter, M. and Rogalski, J. (1999) Biodegradation of lignin by white rot fungi. Fungal Genetics and Biology, 27, 175- 185. doi:10.1006/fgbi.1999.1150
- Sazci, A., Erenler, K. and Radford, A. (1986) Detection of cellulolytic fungi by using Congo red as an indicator: A comparative study with the dinitrosalicyclic acid reagent method. Journal of Applied Microbiology, 61, 559- 562. doi:10.1111/j.1365-2672.1986.tb01729.x
- Kuhad, R.C., Singh, A. and Eriksson, K.E.L. (1997) Microorganisms and enzymes involved in the degradation of plant fiber cell walls. Advances in Biochemical Engineering/Biotechnology, 57, 45-125. doi:10.1007/BFb0102072
- Gashe, B.A. (1992) Cellulase production and activity by Trichoderma sp. A-001. Journal of Applied Microbiology, 73, 79-82. doi:10.1111/j.1365-2672.1992.tb04973.x
- Kang, S.W., Park, Y.S., Lee, J.S., Hong, S.I. and Kim, S.W. (2004) Production of cellulases and hemicellulases by Aspergillus niger KK2 from lignocellulosic biomass. Bioresource Technology, 91, 153-156. doi:10.1016/S0960-8524(03)00172-X
- Molla, A.H., Fakhru’l-Razi, A. and Alam, M.Z. (2004) Evaluation of solid state bioconversion of domestic wastewater sludge as a promising environmental friendly disposal technique. Water Research, 38, 4143-4152. doi:10.1016/j.watres.2004.08.002
- Belghith, H., Ellouz-Chaabouni, S. and Gargouri, A. (2001) Biostoning of denims by Penicillium occitanis (Pol6) cellulases. Journal of Biotechnology, 89, 257-262. doi:10.1016/S0168-1656(01)00309-1
- Singhania, R.R., Sukumaran, R.K. and Pillai, A. (2006) Solid-state fermentation of lignocellulosic substrates for cellulase production by Trichoderma reesei NRRL 11460. Indian Journal of Biotechnology, 5, 332-336.
- Sun, T., Liu, B.H., Li, Z.H. and Liu, D.M. (1999) Effect of air pressure amplitude on cellulase production by Trichoderma viride SL1 in periodic pressure solid state fermentation. Process Biochemistry, 34, 25-29. doi:10.1016/S0032-9592(98)00060-0
- Betina, V. (1995) Photoinduced conidiation in Trichoderma viride. Folia Microbiologica, 40, 219-224. doi:10.1007/BF02814196
- Xiaoyi, Z., Yingde, C.U., Ning, L.U., Xin, Y.U. and Meimei, Z. (2012) Optimization of fermentation condition in liquid culture of T. viride to produce yellow pigment. Center for Improved Engineering and Science Education, 61, 3205-3212.
- Chitale, A., Jadhav, D.V., Waghmare, S.R., Sahoo, A.K. and Ranveer, R.C. (2012) Production and characterization of brown coloured pigment from Trichoderma viride. Electronic Journal of Environmental Agricultural and Food Chemistry, 11, 529-537.
NOTES
*Corresponding author.

