American Journal of Plant Sciences
Vol.4 No.12C(2013), Article ID:41198,9 pages DOI:10.4236/ajps.2013.412A3011
Expression Analysis of Two Genes Coding for Trehalose-6-Phosphate Synthase (TPS), in Sugarcane (Saccharum spp.) under Water Stress
![]()
1Institute of Genetics and Biochemistry, Federal University of Uberlandia, Uberlandia, Brazil; 2Center of Education, Communication and Arts, Western Paraná State University, Cascavel, Brazil; 3Biotechnology Unit, UNAERP, Ribeirão Preto, Brazil.
Email: szingaretti@unaerp.br
Copyright © 2013 Nilson Nicolau Junior et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Received October 29th, 2013; revised December 3rd, 2013; accepted December 13th, 2013
Keywords: Differential Expression; EST; Trehalose; Water Stress
ABSTRACT
The accumulation of trehalose (α-D-glucopyranosyl-[1,1]-α-D-glucopyranoside), a sugar with osmoprotectant properties, is very common in microorganisms, invertebrates and in resurrection plants. However, in the majority of higher plants, it is found in trace amounts. Trehalose is synthesized from the UDP-glucose and glucose-6-phosphate in a twostep process with two enzymes, trehalose-6-phosphate synthase or TPS (EC 2.4.1.15 and EC 2.4.1.36) and trehalose-6- phosphate phosphatase or TPP (EC 3.1.3.12). The trehalose-6-phosphate synthase and its product of the trehalose-6- phosphate (T6P) are probable signaling molecules in the carbohydrate metabolism, contributing to enhancing the plants tolerance to water stress. Water scarcity is one of the most important factors that influence productivity in sugarcane (Saccharum spp.) and it activates a cascade of metabolic events and necessary morphologic changes for the survival of the plant under stress. Here we show the in silico expression study of TPS in different libraries from the SUCEST project. Our results showed that the TPS genes are present in all tissues and that they are divided into two subfamilies (class I and II). It is shown that STPS1 belongs to the class I, therefore, it does not have an active phosphatase (TPP) domain, whereas, the STPS2 has an active TPP domain (class II) determined by the presence of phosphatase boxes. Expression analyses based on the semi-quantitative method of the reverse transcription polymerase chain reaction (RT-PCR) show that the STPS1 gene is up-regulated in the tolerant cultivar under stress and down-regulated in susceptible plants. The STPS2 gene does not show considerable variations in the expression levels under the same treatments. The discovery of active genes such as STPS1 and STPS2 in plants under water stress, contributes for the concerning about the cascade of responses in plants under water deficit and points out to target genes for plant breeding.
1. Introduction
The non-reducing disacchacaride, Trehalose (α-D-glucopyranosyl-[1,1]-α-D-glucopyranoside) usually found in bacteria, fungi and invertebrates, is synthesized from the UDP-glucose and glucose-6-phosphate in a two-step process with two main enzymes—trehalose-6-phosphate synthase (TPS) and trehalose-6-phosphate phosphatase (TPP) [1,2]. The biosynthesis starts with the formation of trehalose-6-phosphate (T6P) from UDP-glucose and glucose-6-phosphate by the action of the TPS enzyme, subsequently the TPP enzyme acts on the trehalose-6-phosphate resulting in trehalose. This sugar is known as a protective molecule in response to several stress conditions in a high number of microorganisms. In bacteria as E. coli, the accumulation of trehalose is extremely important for the viability maintenance in low temperatures [3]. In yeast, evidences accumulated in the last years indicate that the intracellular level of trehalose can determine the stress response and the yeast viability under extreme ambient conditions [4,5]. For a long time, the presence of trehalose in angiosperms was questioned except for the discovery of this sugar in leaves of resurrection plants such as Myrothamus flabellifolius [6,7]. However, during the last decade, genes from the trehalose biosynthesis were identified in angiosperms such as Arabidopsis thaliana [8-10], rice, potato [11] and cotton [12]. Leyman et al. [9] found 11 similar TPS genes presumably functional in the genome of Arabidopsis. These genes were divided into two subfamilies (class I and class II) by the presence/absence of the phosphatase boxes, conserved domains found in the genes that code the TPP enzyme [13]. In Arabidopsis of the T6P, an intermediary compound from the trehalose biosynthesis pathway is indispensable for the carbohydrate use during the plant and flower development [14-18]. Besides, there is a strong evidence that the TPS enzyme also acts as a signaling molecule [15]. Trehalose is an efficient osmoprotectant in many organisms, however, according to Kosmas et al. [12], the enhanced stress resistance in transgenic plants that super-expressed TPS genes, is not assigned to the accumulation of this carbohydrate, since, its concentrations in the plant are unlikely to be sufficient for its osmoprotectant effects. Avonce et al. [19] verified that the super-expression of the AtTPS1 gene (class I) in Arabidopsis conferred drought tolerance without causing morphologic changes in the plant. These plants showed a glucose and ABA insensitive phenotypes, by the action of trehalose-6-phosphate or AtTPS1 that are probably involved in the signal transduction of genes related to glucose and ABA biosynthesis. The transformation of tobacco plants, with the same gene of Arabidopsis, conferred enhanced tolerance to many stress types, such as, drought, dissecation, low temperatures and salinity [20]. According to Chary et al. [21], the AtTPS6 gene (class II) from Arabidopsis, is important in the control of cellular morphogenesis because it fully complements mutants with defects in the cell shape (csp-1). Sugarcane plants (Saccharum officinarum L.), encoding a mushroom TPS gene (Grifola frondosa) showed enhanced water stress tolerance and high productivity in drought conditions [22]. Nowadays sugarcane can be considered as one of the most important crops in the world; Brazil produced more then 620 million tons in 2010/2011 harvest (http://www.portalunica.com.br). One of the factors that influence the productivity of the sugarcane crops is the rain regime. Long drought periods decreased the productivity, thus the water stress appearing as one of the most important factors in the sugarcane crop management. In plants, the water deficit generates a response complex initiated by the stress perception, which leads to a cascade of molecular events culminating in many levels of physiological, metabolical and of developmental responses [23,24]. The Brazilian sugarcane cultivars are hybrid, the result of an interspecific crossing between Saccharum officinarum, S. barbieri, S. sinense and the wild species S. spontaneum and S. robustum. Due to its multiple origins, the sugarcane shows one of the most complex genomes of the plant kingdom, with a diploid number of chromosomes that ranges from 100 to 130 [25]. Facing the difficulty to work with the sugarcane genome, the generation of a database with 237.954 ESTs (Expressed Sequence Tag) during SUCEST project [26], made an increment possible in the molecular studies involving specific sugarcane genes. The study of these sequences generated a great amount of data that had being used to understand the metabolic pathways and a set of important biological processes in the sugarcane breeding.
The aim of this work was to analyze the sequence from two TPS genes (STPS1 and STPS2), which were differentially expressed in previous macroarray studies of two sugarcane cultivars, SP90-1638 and SP83-2847, under water stress. The sugarcane ESTs (STPS1 and STPS2) were sequenced and their functions were predicted by an in silico analysis of the motifs. In order to understand the role of STPS1 and STPS2 genes in the answer to water scarcity, a semi-quantitative RT-PCR expression analysis was performed by using leaf samples from tolerant and susceptible cultivars under water stress in the periods of 24, 72 and 120 h.
2. Materials and Methods
2.1. Plant Material and Growth Conditions
Two sugarcane cultivars (Saccharum spp.), SP90-1638 and SP83-2847, respectively classified as susceptible and tolerant to water stress, were provided by the Center of Sugarcane Technology (CTC), Brazil. The experiment was established in a greenhouse with controlled temperature and humidity (day/night regime). The seedlings were cultivated in 10 liters plastic pots containing agricultural soil. After 56 days of growth in greenhouse, it was initiated the water suppression. Control plants were watered regularly. Leaf samples were collected for RNA extraction at 24, 72 and 120 h after the irrigation suppression that corresponding to the T1, T2 and T3 treatments respectively. Collected samples were kept in a freezer at −80˚C.
2.2. cDNA Sequencing and Bioinformatics
STPS1 and STPS2 EST clones were acquired from Brazilian Clone Collection Center (Brazil). The real size of clones was determined using PCR amplification using the T7 and Sp6 universal primers and two specific primers (STPS1-A/STPS1-B and STPS2-A/STPS2-B) presented in Table 1. In order to obtain the complete clone sequences, from the sequence fragments generated in the sequencing, it was used the CAP program, that uses a dynamic algorithm to compute the maximum alignment by overlapping among fragments [27]. The clones obtained nucleotide sequences were compared with the NCBI database through the BLASTX search [28]. The conceptual translation and the search for conserved do-
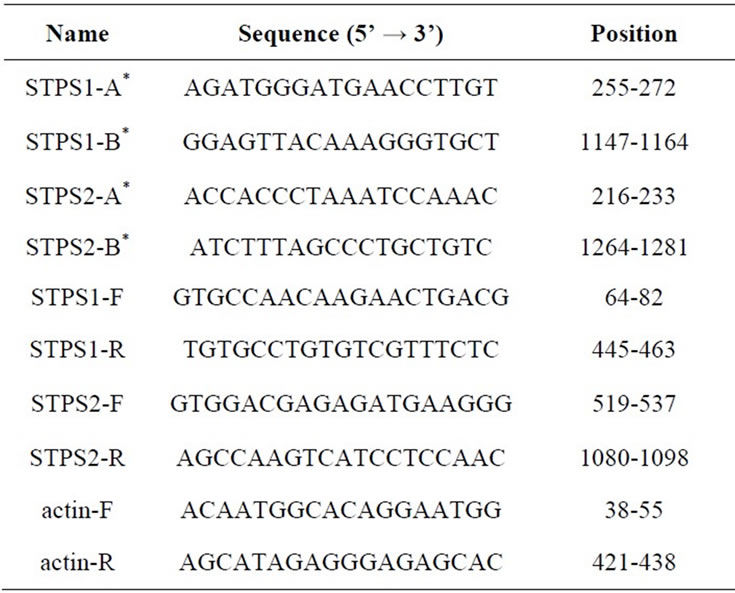
Table 1. Primers used in the sequencing (*) and RT-PCR for the STPS1 e STPS2 genes.
mains was performed with the ORF-Finder tool [29] and CD Search [30]. Multiple alignment analyses was performed with CLUSTALW program [31] using amino acid sequences from STPS1, STPS2 and public available TPS sequences from Arabidopsis thaliana (TPS 1 to 11), gb accession number: AEE36123.1, AEE29528.1, AEE29529.1, AEE85356.1, AEE83948.1, AEE34738.1, AEE27983.1, AEE35042.1, AEE30444.1, AEE33662.1 and AEC06795.1 respectively. The dendrograms were constructed with the Neighbour Joinig method [32].
2.3. In Silico Expression
An in silico expression study using ESTs related to the TPS was performed in the SUCEST database (http://sucest.lad.ic.unicamp.br). The nucleotide sequences from the selected ESTs were used in the search for similarity with amino acid sequences (BLASTX) from the NCBI genbank. The ESTs, with the most representative similarities (e-values < e-30) with the TPS genes, were selected for the in silico expression analyses that was performed using the frequency of the TPS transcript per library.
2.4. Expression Analysis
The expression analysis were performed with Total RNA extracted from sugarcane leaf samples of two cultivars, SP90-1638 (susceptible) and SP83-2847 (tolerant), using the Trizol reagent (Invitrogen). Total RNA (1 µg) treated with DNAse I was reverse transcribed using 200 U of Superscript III RT (Invitrogen) and Oligo dT in the presence of RNAseOUT (Invitrogen). The RT-PCR experiments were performed with gene-specific primers (STPS1-F/STPS1-R and STPS2-F/STPS2-R) as described in Table 1. The sugarcane actin gene was used as internal control. In order to quantify the transcripts, 26 cycles for the TPSs genes and 27 cycles for actin gene were performed using the following PCR conditions: 95˚C for 2 min, 27 or 26 cycles at 95˚C for 45 s, 55˚C for 45 s and 72˚C for 45 s and an elongation step at 72˚C for 5 min. The band intensities in the agarose gel were measured and the expression level was calculated by dividing the intensity from the bands of studied genes by the respective of actin bands intensity.
3. Results
3.1. Sequence Identification and Amino Acid Deduction
The sequenced fragments, overlapped by the CAP program, generated two consensus sequences, one with 1876 bp, STPS1 (gb: EU761244.1) and another with 1822 bp, STPS2 (gb: EU761245.1), representing the cDNA sequences from the studied clones. The sequence similarity between the cDNAs studied and the genes of TPS was confirmed by the BLASTX search. The complete sequences of the TPSs, found in the NCBI genbank, show an average size of 3000 nucleotides, thus, the genes studied in this work are characterized as TPS fragments. The STPS1 gene translated sequence (557 amino acid residues) shows high significant similarities (positives) with the TPS (class I) proteins from Zea mays (maize), Setaria italica (foxtail millet), Brachypodium distachyon, Oryza sativa (rice) and Arabidopsis thaliana with the respective percentages of similarity: 99%, 96%, 93%, 93%, 82%, 80% (Table 2). Besides the positives, the identity percentage between the sequences showed significant high values that together with the low e-values allow the confidence in the results of the BLASTX for the studied sequences (Tables 2 and 3).
3.2. Conserved Domains and Dendrograms
Using the CD Search tool from NCBI, it was possible to identify highly conserved regions known as TPS and TPP domains in the STPS1 and STPS2 incomplete amino acid sequences Figure 1. The analyses allowed observing that both protein sequences show the TPS and TPP domains. The TPP domain, ubiquitous in plants, is the key for the two subfamilies classification proposed by Leyman et al. [9] for the TPS proteins. Our studies indicate that TPP domain from STPS1 is similar to the TPS class I proteins, and it does not contain the region known as phosphatase boxes (LDYD|GD|T|LM| e GDDRSD). Different from STPS1 the STPS2 protein contains the phosphatase boxes usually found in the class II proteins. Two dendrograms were designed performing the Neighbor Joining method, using the amino acid sequences from 11 TPS genes identified in Arabidopsis, and the STPS1
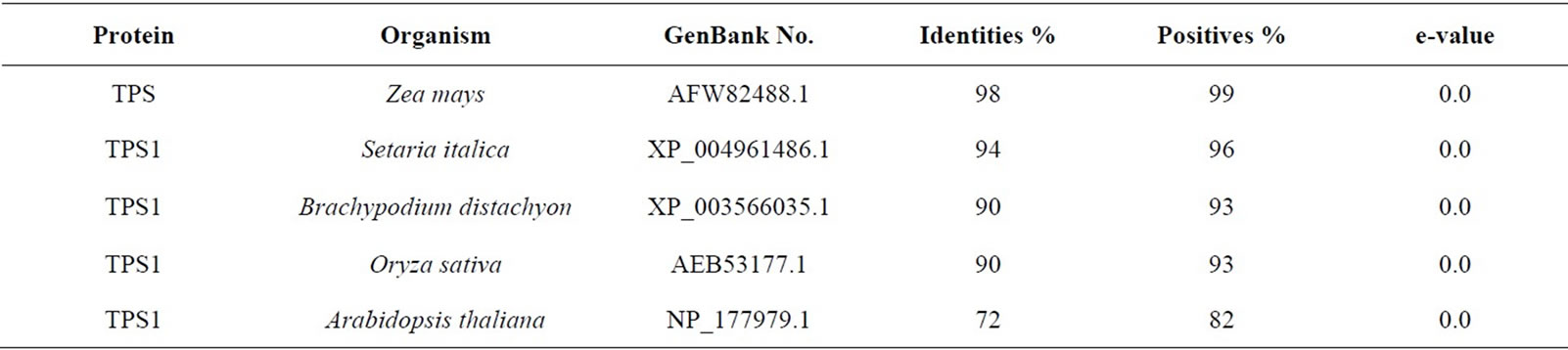
Table 2. Similarity of the sugarcane TPS1 to other TPSs using BlastX tool.
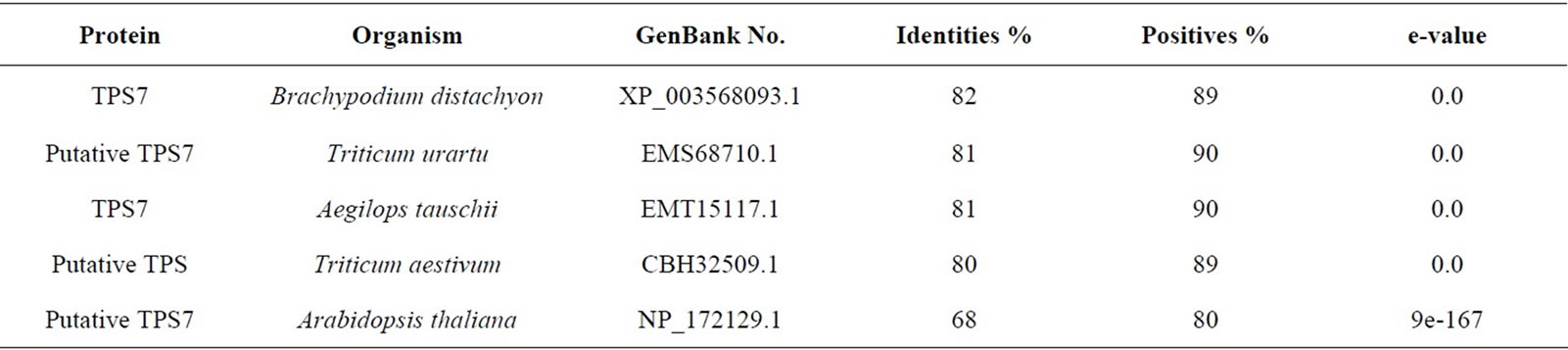
Table 3. Similarity of the sugarcane TPS2 to other TPSs.
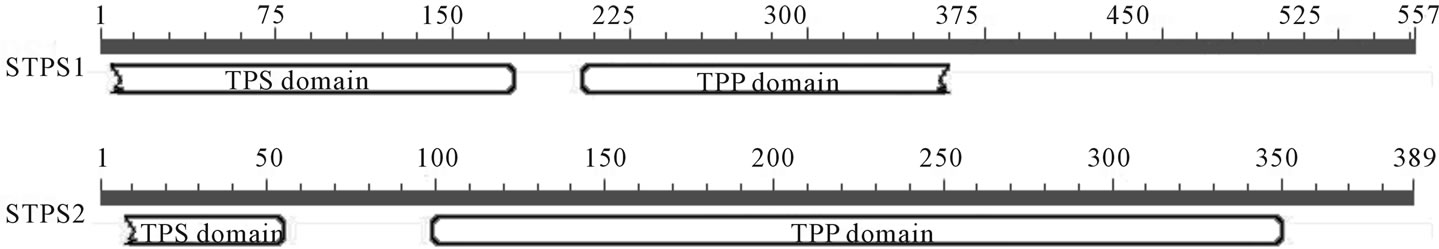
Figure 1. Results from the conserved domain search (CD-Search) tool for the STPS1 and STPS2 incomplete amino acid sequences.
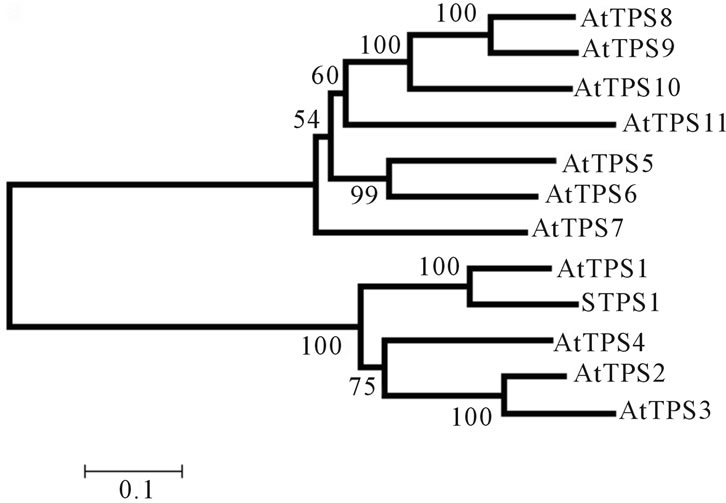
and 2 genes from Saccharum spp., Figures 2(a) and (b), respectively. The first dendrogram, Figure 2(a), shows that STPS1 sequence forms a distinct group with the class I proteins from Arabidopsis (AtTPS1, 2, 3 and 4), whereas the class II representatives are more distant from the STPS1 protein. In the second dendrogram, Figure 2(b), the STPS2 protein forms a group with the class II proteins (AtTPS5, 6, 7, 8, 9, 10 and 11). This result corroborated with the presence of phosphatase boxes detected in the TPP domains of the STPS2 amino acid sequence.
3.3. Expression of TPS Genes
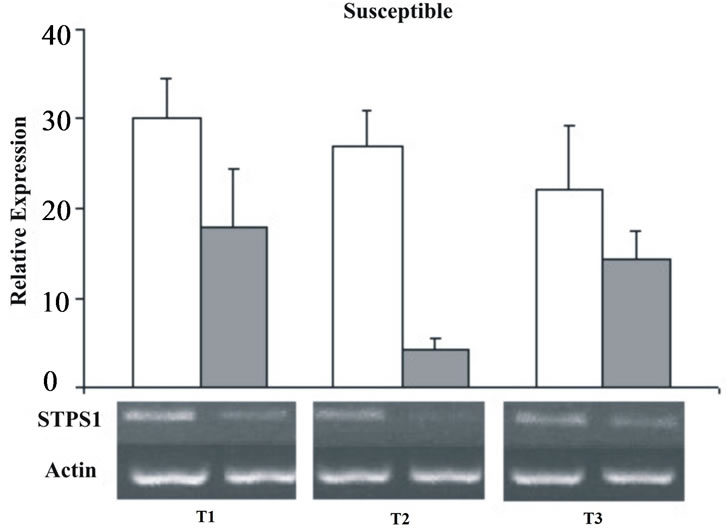
From the 38 EST clusters related to TPS in SUCEST database, 14 cluster were selected by their high similarity with TPSs genes from other organisms (e-values < e-30 in the BLASTX) and by the non-alignment among them (CLUSTALW). These 14 clusters represent 110 distinct TPS transcripts isolated from libraries of the SUCEST project, originated from different tissues and treatments (Figure 3). The in silico TPS expression showed similar frequencies (from 2.47 × 10−4 to 5.14 × 10−4) for the majority of the studied libraries, except for the CL, AD, RZ, and LV libraries that showed the highest frequencies (29.00 × 10−4, 6.80 × 10−4, 6.76 × 10−4, 6.58 × 10−4 respectively). In order to understand the STPS1 and STPS2 expression in the early stages of sugarcane water stress, it was performed a semi-quantitative RT-PCR. The amplification was performed with specific primers for the two genes and the actin gene as an internal control (Table 1) that was used for the data standardization. Although deriving from ESTs found in other tissues (AD and FL) the STPS1 and STPS2 genes showed expression in the leaves of both, control and stressed plants of the sugarcane cultivars studied. Except for the T2 (72 h), the STPS1 gene seems to have an expression level near or superior to the control in the tolerant plant under water stress. In the susceptible plant, the STPS1 gene showed to be down-regulated under water stress for the three studied periods (Figure 4). The expression of STPS2 gene does not showed significant differences between the control and stressed plants for the both cultivars studied,
 (a)
(a)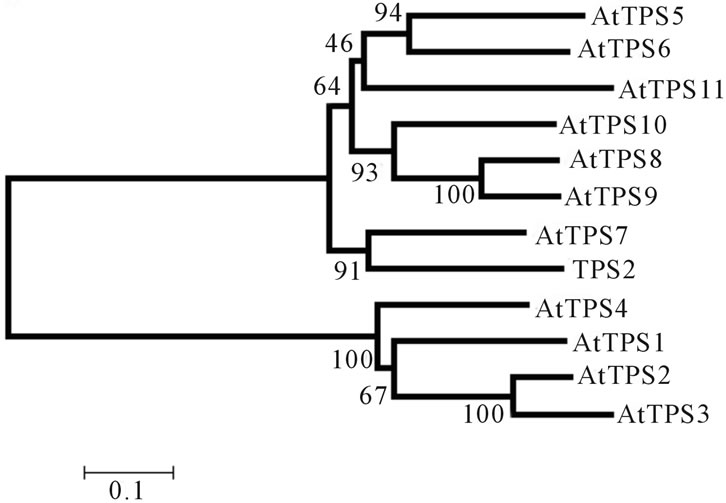 (b)
(b)
Figure 2. Dendrograms based on the neighbor joining method (bootstrap × 1000) showing the distance among the incomplete STPS1 (a) and STPS2 (b) sugarcane amino acid sequences and the TPS proteins from A. thaliana.
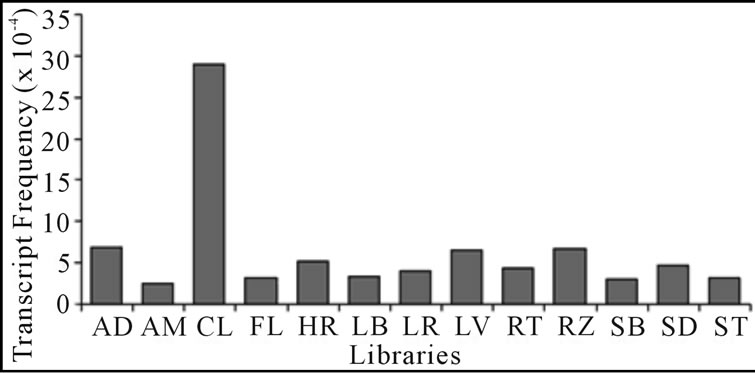
Figure 3. In silico expression of sugarcane TPS genes from different libraries of the SUCEST project. The libraries are: AD (mixture of tissues from root to shoot zone, stem and apical meristem of plantlets cultivated in vitro and infected with Gluconacetobacter diazotroficans); AM (apical meristem of young plants); CL (pool of calli treated for 12 h at 4˚C and 37˚C in the dark or light); FL (flowers harvested at different developmental stages); HR (mixture of tissues from root to shoot zone, stem and apical meristem of plantlets cultivated in vitro and infected with Herbaspirilum diazotroficans); LB (lateral buds from mature plants); LR (leaf roll from immature plants); FL (etiolated leaves from plantlets grown in vitro); RT (0.3 cm-length roots from mature plants and root apex); RZ (root to shoot zone of young plants); SB (stalk bark from mature sugarcane plants); SD (developing seeds); ST (first and fourth internodes of immature plants).
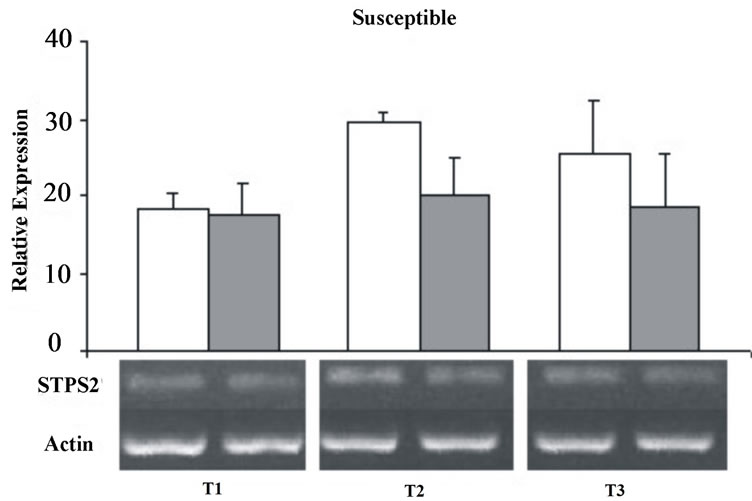
except for the T2 were it was down-regulated under stress condition (Figure 5).
4. Discussion
4.1. TPS in Silico Expression
The 14 TPS clusters identified in sugarcane may represent isoforms for the same gene as already identified in other organisms as Arabidopsis. This probable high number of isoforms of TPS in sugarcane may be due to the polyploidy and the hybrid condition associated to the Saccharum spp. As well as in sugarcane, transcripts of TPS genes in several plant tissues like in Arabidopsis [8,12], rice [33] and maize [34] were found. The in silico expression indicates a constitutive expression of TPS genes in several tissues and treatment approaches in the hibrid sugarcane plants.
According to Avonce et al. [35], the genes AtTPS1, AtTPS6, AtTPS7 AtTPS8 and AtTPS11 showed constitutive expression in plants of Arabidopsis. A transcripts high frequency was found in calli libraries under temperature stress (CL). Although some research results show that for the majority of higher plants, the level of trehalose produced is not enough for its osmoprotectant effect, studies carried out with rice [36,37] and tobacco [20] have shown that the inclusion of TPS genes enhanced the stress tolerance to low temperatures.
4.2. Genes of the TPS of Saccharum spp.
The nucleotide sequence of the two ESTs clones showed 1876 bp for STPS1 and 1887 bp for STPS2. These genes show the complete C-terminal region including the phosphatase domain (TPP) and a fragment of the synthase domain (TPS). These two domains, different from prokaryotes, are fused in the TPS of eukaryotes. In higher plants, the TPS gene was firstly described by Leyman et al. [9] in Arabidopsis where the TPSs were classified in two subfamilies that correspond to the enzymes that have or not have the phosphatase boxes in their respective TPP domains. The TPP domain is the key for the classification of the TPS genes in plants. The STPS1 gene fragment detected initially in flower libraries (FL) showed the phosphatase domain without the phosphatase boxes, and high similarity with AtTPS1 from Arabidopsis. According to van Dijken et al. [17], in mutant plants of Arabidopsis, higher levels of relative AtTPS1 expression were detected in metabolically or-
 (a)
(a)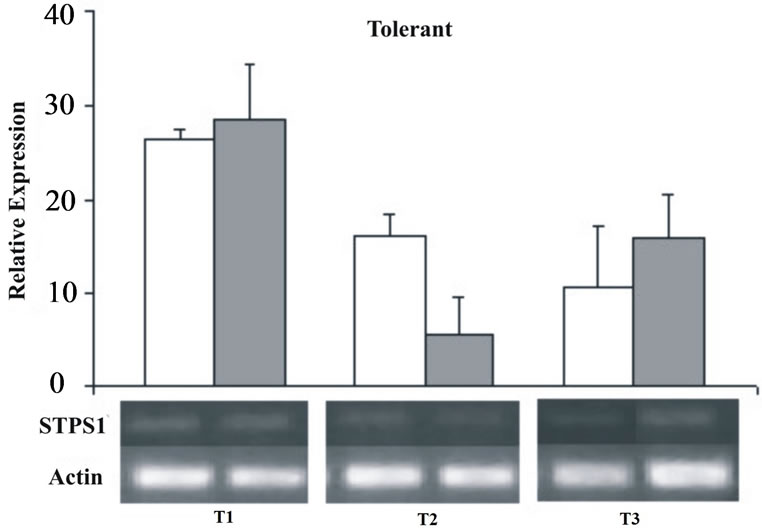 (b)
(b)
Figure 4. Relative expression of the STPS1 gene in the control (white bars) and stressed sugarcane plants (grey bars). It was performed three repetitions for each treatment in both cultivars, susceptible and tolerant. T1 = 24 h; T2 = 72 h; T3 = 120 h.
gans as flower buds and siliques. The AtTPS1 gene, is capable to restore growth and the trehalose synthesis in mutant yeasts, and is admittedly essential for the development of the plant and flowering; besides, TPS generates as product of the T6P that is indispensable for the growth and use of the carbohydrate Arabidopsis [8,15,17]. The STPS2 gene shows similarity with the AtTPS7 from Arabidospis, gene that corresponds to the second TPS class. In spite of its constitutive expression, the AtTPS7 gene was unable to complement the tps1 and tps2 mutants of S. cereviseae, because of its inactive TPS domain [10]. However, evidences indicate that genes form a second class control the morphology of the cell playing an important role in the plant development [21].
4.3. Expression of the TPSs in Saccharum spp.
The data acquired in the semi-quantitative RT-PCR analyses confirmed that two genes of TPS are constitu-
 (a)
(a)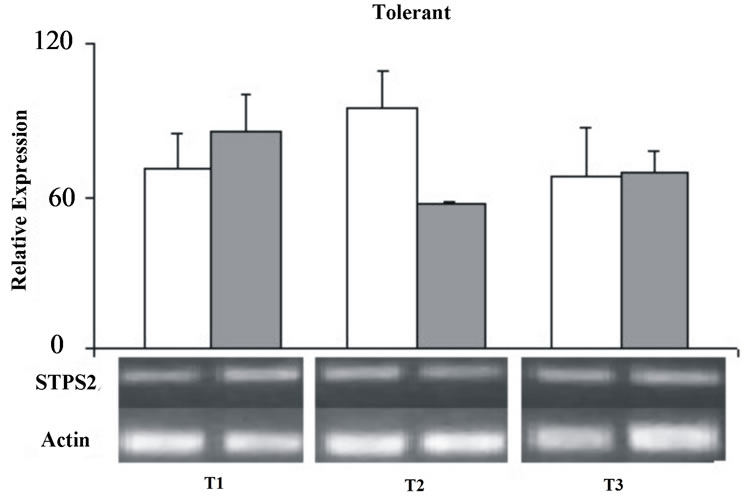 (b)
(b)
Figure 5. Relative expression of the STPS2 gene in the control (white bars) and stressed sugarcane plants (grey bars). It was performed three repetitions for each treatment in both cultivars, susceptible and tolerant. T1 = 24 h; T2 = 72 h; T3 = 120 h.
tively expressed in young leaves of sugarcane in normal and drought conditions (Figures 4 and 5).
The STPS1 gene was differentially expressed in the two cultivars of sugarcane when under water stress (Figure 4). The up-regulation of STPS1 in tolerant plants as well as its down-regulation in susceptible plants at the beginning of water stress corroborates with the waterstress tolerant status of SP83-2847 cultivar. The enhanced expression of the TPS1 gene and the accumulation of T6P act on the growth metabolism and sugar regulation in Arabidopsis plants [15,17]. It may be related to the stress tolerance observed in plants as well as the results of our studies. Facing the difficulty of the classic breeding with polyploidy, as sugarcane cultivars, the enhanced expression of STPS1 in water-stress tolerant plants and its similarity to the AtTPS1 reveal the potential use of this gene as a molecular marker in plant breeding programs and as a target for transgenic plants. Transgenic sugarcane plants (Saccharum officinarum L.), carrying the TPS gene from a mushroom (Grifolia frondosa), showed a significant increase in the drought tolerance [38]. According to Almeida et al. [20], tobacco plants modified to express the AtTPS1 gene, showed resistance to several types of abiotic stress including water stress. Moreover, the super-expression of the AtTPS1 in Arabidopsis increased the T6P production, and displayed the formation of plants with ABA and sugar insensitive phenotypes and tolerance to drought stress [19]. According to Schluepmann et al. [15,16], the T6P plays a role in the regulation of the carbohydrate status and is linked to the expression of genes related to nutritional, biotic and abiotic stresses. Studies with tobacco modified by trehalose biosynthesis genes from E. coli, showed that increased levels of T6P in leaves have an important role in the regulation of the photosynthesis [39]. Moreover, Lunn et al. [40] showed that T6P, besides acting as a regulator of the sugar status in plants, also mediates changes induced by sucrose in the starch synthesis rate. Except for the T2, the STPS2 gene did not show significant differential expression between the control and stressed plants for both sugarcane cultivars. Although the class II TPSs from Arabidopsis is classified as unknown functions, there are some evidences showing that they play a role in the control of the cellular morphology and the plant development [21]. Besides it is also known that the class II genes mediate response signals that activate the gene SnRK1 [41], a kinase protein that plays a role key in the global control of the carbon metabolism in the plant [42]. To our knowledge, this work is the first expression report from the two TPS subfamilies in sugarcane cultivars. The expressions of the TPSs, STPS1 and STPS2, in normal and water stress conditions, together with the presence of many other TPSs detected in all sugarcane tissues are indicators of the enzyme importance and reveal the necessity of further studies using specific genetic tools to enhance the quality of the sugarcane cultivars.
5. Acknowledgements
We thank the Copersucar (Center of Sugarcane Technology, Piracicaba-Brazil), for the sugarcane seedlings, the São Paulo State Foundation for Research Support (FAPESP) that with the National Council for Scientific and Technological development (CNPq) granted financial support to this work.
REFERENCES
- E. Cabib and F. L. Leloir, “The Biosynthesis of Trehalose-6-Phosphate,” Journal of Biological Chemistry, Vol. 31, 1958, pp. 259-275.
- J. M. Thevelein, “Regulation of Trehalose Metabolism and Its Relevance to Cell Growth and Function,” In: R. Brambl and G. A. Marzluf, Eds., The Mycota, Springer, Berlin-Verlag, 1996, pp. 395-420.
- O. Kandror, A. DeLeon and A. L. Goldberg, “Trehalose Synthesis Is Induced upon Exposure of Escherichia coli to Cold and Is Essential for Viability at Low Temperatures,” Proceedings of the National Academy of Sciences, Vol. 99, No. 15, 2002, pp. 9727-9732. http://dx.doi.org/10.1073/pnas.142314099
- Van Laere, “Trehalose, Reserve and/or Stress Metabolite?” FEMS MicrobiologyReviews, Vol. 63, No. 3, 1989, pp. 201-210.
- A. Wiemken, “Trehalose in Yeast, Stress Protectant rather Than Reserve Carbohydrate,” Antonie van Leeuwenhoek, Vol. 58, No. 3, 1990, pp. 209-217. http://dx.doi.org/10.1007/BF00548935
- G. Bianchi, A. Gamba, R. Limiroli, N. Pozzi, R. Elster, F. Salamini and D. Bartels, “The Unusual Sugar Composition in Leaves of the Resurrection Plant Myrothamnus flabellifolia,” Physiologia Plantarum, Vol. 87, No. 2, 1993, pp. 223-226. http://dx.doi.org/10.1111/j.1399-3054.1993.tb00146.x
- P. M. Drennan, M. T. Smith, D. Goldsworthy and J. van Staden, “The Occurrence of Trehalose in the Leaves of the Desiccation Tolerant Angiosperm Myrothamnus flabellifolius Welw,” Journal of Plant Physiology, Vol. 142, No. 4, 1993, pp. 493-496. http://dx.doi.org/10.1016/S0176-1617(11)81257-5
- M. A. Blazquez, E. Santos, C. L. Flores, J. M. MartinezZapater, J. Salinas and C. Gancedo, “Isolation and Characterization of the Arabidopsis TPS1 Gene, Encoding Trehalose-6-Phosphate Synthase,” Plant Journal, Vol. 13, No. 5, 1998, pp. 685-689. http://dx.doi.org/10.1046/j.1365-313X.1998.00063.x
- A. Leyman, P. Van Dijck and J. M. Thevelein, “An Unexpected Plethora of Trehalose Biosynthesis Genes in Arabidopsis Thaliana,” Trends in Plant Science, Vol. 6, 2001, pp. 510-513. http://dx.doi.org/10.1016/S1360-1385(01)02125-2
- G. Vogel, O. Fiehn, L. J. R. D. Bressel, T. Boller, A. Wiemken, R. A. Aeschbacher and A. Wingler, “Trehalose Metabolism in Arabidopsis: Occurrence of Trehalose and Molecular Cloning and Characterization of Trehalose-6- Phosphate Synthase Homologues,” Journal of Experimental Botany, Vol. 52, No. 362, 2001, pp. 1817-1826. http://dx.doi.org/10.1093/jexbot/52.362.1817
- O. Goddijn and S. Smeekens, “Sensing Trehalose Biosynthesis in Plants,” Plant Journal, Vol. 14, No. 2, 1998, pp. 143-146. http://dx.doi.org/10.1046/j.1365-313X.1998.00140.x
- S. A. Kosmas, A. Argyrokastritis, M. G. Loukas, E. Eliopoulos, S. Tsakas and P. J. Kaltsikes, “Isolation and Characterization of Drought-Related Trehalose 6-Phosphate-Synthase Gene from Cultivated Cotton (Gossypiumhirsutum L.),” Planta, Vol. 223, No. 2, 2006, pp. 329-339. http://dx.doi.org/10.1007/s00425-005-0071-5
- C. Thaller, S. Schippa and G. M. Rossolini, “Conserved Sequence Motifs among Bacterial, Eukaryotic and Archaeal Phosphatases That Define a New Phosphohydrolase Superfamily,” Protein Science, Vol. 7, No. 7, 1998, pp. 1651-1656. http://dx.doi.org/10.1002/pro.5560070722
- P. J. Eastmond, A. J. H. van Dijken, M. Spielman, A. Kerr, A. F. Tissier, H. G. Dickinson, J. D. G. Jones, S. C. Smeekens and I. A. Graham, “Trehalose-6-Phosphate Synthase 1, Which Catalyses the First Step in Trehalose Synthesis, Is Essential for Arabidopsis Embryo Maturation,” Plant Journal, Vol. 29, No. 2, 2002, pp. 225-235. http://dx.doi.org/10.1046/j.1365-313x.2002.01220.x
- H. Schluepmann, T. Pellny, A. van Dijken, S. Smeekens and M. Paul, “Trehalose 6-Phosphate Is Indispensable for Carbohydrate Utilization and Growth in Arabidopsis thaliana,” Proceedings of the National Academy of Sciences, Vol. 100, No. 11, 2003, pp. 6849-6854. http://dx.doi.org/10.1073/pnas.1132018100
- H. Schluepmann, A. van Dijken, M. Aghdasi, B. Wobbes, M. Paul and S. Smeekens, “Trehalose Mediated Growth Inhibition of Arabidopsis Seedlings Is Due to Trehalose- 6-Phosphate Accumulation,” Plant Physiology, Vol. 135, No. 2, 2004, pp. 879-890. http://dx.doi.org/10.1104/pp.104.039503
- J. van Dijken, H. Schluepmann and S. C. Smeekens, “Arabidopsis Trehalose-6-Phosphate Synthase 1 Is Essential for Normal Vegetative Growth and Transition to Flowering,” Plant Physiology, Vol. 135, No. 2, 2004, pp. 969-977. http://dx.doi.org/10.1104/pp.104.039743
- B. Zang, H. Li, W. Li, X. W. Deng and X. Wang, “Analysis of Trehalose-6-Phosphate Synthase (TPS) Complexes in Rice,” Plant Molecular Biology, Vol. 76, No. 6, 2011, pp. 507-522. http://dx.doi.org/10.1007/s11103-011-9781-1
- N. Avonce, B. Leyman, J. O. Gallardo-Mascorro, P. Van Dijck, J. M. Thevelein and G. Iturriaga, “The Arabidopsis Trehalose-6-P Synthase AtTPS1 Gene Is a Regulator of Glucose, Abscisic Acid, and Stress Signaling,” Plant Physiology, Vol. 136, No. 3, 2004, pp. 3649-3659. http://dx.doi.org/10.1104/pp.104.052084
- M. Almeida, E. Villalobos, S. S. Araújo, B. Leyman, P. van Dijk, L. Alfaro-Cardoso, P. S. Fevereiro, J. Torné and D. M. Santos, “Transformation of Tobacco with an Arabidopsis thaliana Gene Involved in Trehalose Biosynthesis Increases Tolerance to Several Abiotic Stresses,” Euphytica, Vol. 146, 2005, pp. 165-176. http://dx.doi.org/10.1007/s10681-005-7080-0
- S. N. Chary, G. R. Hicks, Y. G. Choi, D. Carter and N. V. Raikhel, “Trehalose-6-Phosphate Synthase/Phosphatase Regulates Cell Shape and Plant Architecture in Arabidopsis,” Plant Physiology, Vol. 146, No. 1, 2008, pp. 97-107. http://dx.doi.org/10.1104/pp.107.107441
- S. Zhang, B. Yang, C. Feng, R. Chen, J. Luo, W. Cai and F. Liu, “Expression of the Grifola frondosa Trehalose Synthase Gene and Improvement of Drought-Tolerance in Sugarcane (Saccharum officinarum L.),” Journal of Integrative Plant Biology, Vol. 48, No. 4. 2006, pp. 453-459. http://dx.doi.org/10.1111/j.1744-7909.2006.00246.x
- E. A. Bray, “Molecular Responses to Water Deficit,” Plant Physiology, Vol. 103, No. 4, 1993, pp. 1035-1040.
- G. C. Dedemo, F. A. Rodrigues, P. G. Roberto, C. B. Neto, S. C. França and S. M. Zingaretti, “Osmoprotection in Sugarcane under Water Deficit Conditions,” Plant Stress, Vol. 7, No. 1, 2013, pp. 1-7.
- A. D'Hont, L. Grivet, P. Feldmann, S. Rao, N. Berding and J. C. Glaszmann, “Characterisation of the Double Genome Structure of Modern Sugarcane Cultivars (Saccharum spp.) by Molecular Cytogenetics,” Molecular Genetics and Genomics, Vol. 250, No. 4, 1996, pp. 405- 413.
- L. Vettore, F. R. da Silva, E. L. Kemper and P. Arruda, “The Libraries That Made SUCEST,” Genetics and Molecular Biology, Vol. 24, No. 1-2, 2001, pp. 1-4. http://dx.doi.org/10.1590/S1415-47572001000100002
- X. Huang, “A Contig Assembly Program Based on Sensitive Detection of Fragment Overlaps,” Genomics, Vol. 14, 1992, pp. 18-25. http://dx.doi.org/10.1016/S0888-7543(05)80277-0
- S. F. Altschul, W. Gish, W. Miller, E. W. Myers and D. J. Lipman, “Basic Local Alignment Search Tool,” Journal of Molecular Biology, Vol. 215, No. 3, 1990, pp. 403- 410.
- T. Rombel, K. F. Sykes, S. Rayner and A. S. Johnston, “ORF-FINDER: A Vector for High-Throughput Gene Identification,” Gene, Vol. 282, No. 1-2, 2002, pp. 33-41. http://dx.doi.org/10.1016/S0378-1119(01)00819-8
- A. Marchler-Baue and S. H. Bryant, “CD-Search: Protein Domain Annotations on the Fly,” Nucleic Acids Research, Vol. 32, Suppl. 2, 2004, pp. 327-331. http://dx.doi.org/10.1093/nar/gkh454
- M. J. D. Thompson, D. G. Higgins and T. J. Gibson, “CLUSTAL W: Improving the Sensitivity of Progressive Multiple Sequence Alignment through Sequence Weighting, Position-Specific Gap Penalties and Weight Matrix Choice,” Nucleic Acids Research, Vol. 22, No. 22, 1994, pp. 4673-4680. http://dx.doi.org/10.1093/nar/22.22.4673
- N. Saitou and M. Nei, “The Neighbour-Joining Method: A New Method for Reconstructing Phylogenetic Trees,” Molecular Biology and Evolution, Vol. 4, No. 4, 1987, pp. 406-425
- M. H. R. Pramanik and R. Imai, “Functional Identification of a Trehalose 6-Phosphate Phosphatase Gene That Is Involved in Transient Induction of Trehalose Biosynthesis during Chilling Stress in Rice,” Plant Molecular Biology, Vol. 58, 2005, pp. 751-762. http://dx.doi.org/10.1007/s11103-005-7404-4
- W. Jiang, F.-L. Fu, S.-Z. Zhang, L. Wu and W.-C. Li, “Cloning and Characterization of Functional Trehalose- 6-Phosphate Synthase Gene in Maize,” Journal of Plant Biology, Vol. 53, No. 2, 2010, pp. 134-141. http://dx.doi.org/10.1007/s12374-010-9098-7
- N. Avonce, A. Mendoza-Vargas, E. Morett and G. Iturriaga, “Insights on the Evolution of Trehalose Biosynthesis,” BMC Evolutionary Biology Vol. 6, No. 1, 2006, p. 109. http://dx.doi.org/10.1186/1471-2148-6-109
- K. Garg, J. K. Kim, T. G. Owens, A. P. Ranwala, Y. D. Choi, L. V. Kochian and R. J. Wu, “Trehalose Accumulation in Rice Plants Confers High Tolerance Levels to Different Abiotic Stresses,” Proceedings of the National Academy of Sciences USA, Vol. 99, No. 25, 2002, pp. 15898-15903. http://dx.doi.org/10.1073/pnas.252637799
- C. Jang, S. Oh, J. S. Seo, W. B. Choi, S. Y. Song, C. H. Kim, Y. S. Kim, H. S. Seo, Y. D. Choi, B. H. Nahm and J. K. Kim, “Expression of a Bifunctional Fusion of the Escherichia coli Genes for Trehalose-6-Phosphate Synthase and Trehalose-6-Phosphate Phosphatase in Transgenic Rice Plants Increases Trehalose Accumulation and Abiotic Stress Tolerance without Stunting Growth,” Plant Physiology, Vol. 131, No. 2, 2003, pp. 516-524. http://dx.doi.org/10.1104/pp.007237
- Z. Z. Wang, S. Z. Zhang, B. P. Yang and Y. R. Li, “Trehalose Synthase Gene Transfer Mediated by Agrobacterium tumefaciens Enhances Resistance to Osmotic Stress in Sugarcane,” Scientia Agricultura Sinica, Vol. 36, No. 2, 2003, pp. 140-146.
- T. K. Pellny, O. Ghannoum, J. P. Conroy, H. Schluepmann, S. Smeekens, J. Andralojc, K. P. Krause, O. Goddijn and M. J. Paul, “Genetic Modification of Photosynthesis with E. coli Genes for Trehalose Synthesis,” Plant Biotechnology Journal, Vol. 2, No. 1, 2004, pp. 71-82. http://dx.doi.org/10.1111/j.1467-7652.2004.00053.x
- J. E. Lunn, R. Feil, J. H. M. Hendriks, Y. Gibon, R. Morcuende, D. Osuna, W. R. Scheible, O. Carillo, M. R. Hajirezaei and M. Stitt, “Sugar-Induced Increases in Trehalose 6-Phosphate Are Correlated with Redox Activation of ADP Glucose Pyrophosphorylase and Higher Rates of Starch Synthesis in Arabidopsis thaliana,” Biochemical Journal, Vol. 397, No. 1, 2006, pp. 139-148. http://dx.doi.org/10.1042/BJ20060083
- J. E. Harthill, S. E. Meek, N. Morrice, M. W. Peggie, J. Borch, B. H. Wong and C. Mackintosh, “Phosphorylation and 14-3-3 Binding of Arabidopsis Trehalose-Phosphate Synthase 5 in Response to 2-Deoxyglucose,” Plant Journal, Vol. 47, No. 2, 2006, pp. 211-223. http://dx.doi.org/10.1111/j.1365-313X.2006.02780.x
- N. G. Halford, S. Hey, D. Jhurreea, S. Laurie, R. S. McKibbin, M. Paul and Y. Zhang, “Metabolic Signalling and Carbon Partitioning: Role of Snf1-Related (SnRK1) Protein Kinase,” Journal of Experimental Botany, Vol. 54, No. 382, 2003, pp. 467-475. http://dx.doi.org/10.1093/jxb/erg038

