Advances in Microbiology
Vol.3 No.1(2013), Article ID:29203,7 pages DOI:10.4236/aim.2013.31011
Phylogenetic Identification of Pathogenic Fungi from Apple in Batu City, Malang, Indonesia
Biology Department, Postgraduate Program, Brawijaya University, Malang, Indonesia
Email: 116090117011005@mail.ub.ac.id, calistus@ub.ac.id, laras@ub.ac.id
Received December 7, 2012; revised January 8, 2013; accepted February 10, 2013
Keywords: Apple; DNA; Fungi; Pathogen; Phylogenetic
ABSTRACT
Pathogenic fungi cause dying off of apple tree and giving negative impact to farmers. The objective of this study was to identify tree pathogenic fungi isolates from apple tree in the Batu City and to know phylogenetic relationship of the isolates based on the DNA sequences. The method in this study included subculture of three pathogenic fungi isolates (M1, M4, and MB1) using V8 Juice Agar medium for one to three days, continued with DNA isolation, amplification using ITS5/ITS4 primer, purification, sequencing, and sequence homology analysis of DNA amplicons of each isolates with reference isolates. The results showed similarity value of DNA sequence of the three isolates M1, M4, and MB1 of more than 99%. The three isolates M1, M4, and MB1 have a similarity value of DNA sequence with species of Pythium splendens of 99.84%, 99.67% and 99.83%, respectively. On the other hand similarity value between those isolates with Phytophthora was less than 76%. It was concluded that the three isolates are Py. splendens.
1. Introduction
Pathogenic fungi are one of the causes of herbaceous plants and woody plants diseases. It causes rotting of the roots, stems base, death of plants, which in turn give negative impact to farmers [1]. One example of the pathogenic fungi that attack plant is Phytophthora [2]. More than 66% of root diseases and more than 90% of stem base rot diseases of woody plants caused by it [3]. Several species of Phytophthora also cause natural ecosystems damage and change the composition of flora [2]. In addition, the development of agro-industry in Australia is affected by this pathogen, especially in the agrofruit crops and ornamental plants [1]. Approximately more than $250 million per year was spent by farmers to overcome it [1].
In many countries, some types of Phytophthora have been identified in apple crop, like P. cactorum, P. drechsleri, P. cryptogea, P. citricola, P. cambivora, and P. arecae [3]. There are also other spesies of pathogenic fungi which were often associated with Phytophthora, such as Pythium, for example Pythium ultimum causes collar rot diseases like Phytophthora cactorum [4]. Pythium and Phytophthora are Pythiaceae. It causes the death of the plant, has a broad host range, and causes much harm to farmer [5]. This fungi causes of root disease in plants, the death of the seeds and small plants, as well as significantly decrease in the ability of roots to obtain nutrients from the soil because structural damage [6]. Several species of Pythium cause disease on apple crops, such as Py. splendens in California and Florida [6], Py. rostratifingens, Py. undulatum, Py. sterilum in Tunisia [7], Py. intermedium, Py. irregulare, Py. Heterothallicum, Py. Sylvaticum, and Py. MM1 (aff. Macrosporum) at Washington [8].
Nugrahani [9] obtained four pathogenic fungi isolates from apple crops. Those isolates were suspected to be members of Phytophthora [9]. However, the phenotypes of those isolates were not able to identify the species name. Therefore, it is necessary to identify the phylogenetic relationship with Phytophtora species to know the species name of those isolates based on DNA sequences as well as knowing phylogenetic relationship of three isolates with reference isolates (members of the Phytophthora and other group, like members of the Pythium).
The development process of analysis and study of phytopathogenic fungi increased along with the increase of members of Phytophthora found in recent years [10]. The development of molecular techniques provides great opportunities for the development of a sensitive detection device, specific, effective and efficient method [11]. Polymerase Chain Reaction (PCR) is a methods and devices used in the detection of molecular approaches, which serves in vitro DNA amplification. The product of PCR can be sequenced then used to differentiate organisms at the level of genus, species and even the specific genotype [10].
The existence of conservative areas, such as genes encoding ribosomal RNA (rRNA) 18 S, 5.8 S, and 28 S simplify the process of phylogenetic identification [12]. There were areas known as the internal transcribed spacer (ITS) among the genes encoding structural protein [13]. ITS region mutates and a significant variation between species can be determined from the area [13]. ITS4 and ITS5 in this study were expected to amplify the ITS1 and ITS2 (universal DNA barcode marker for fungi) [14], 5.8 S rDNA complete sequence, 18 S rDNA and 28S rDNA partial sequence [15]. It was used for identify the specific name of the species of the isolates analyzed.
Development of those methods is very useful to understand the phylogenetic relationship among isolates [11]. Furthermore, it also supports to determine appropriate methods for controlling those pathogens. Infection among groups of different taxonomic fungi has variation and association with the tissue of hosts infected and also nutrients needed by pathogens for its growing. So, closely related pathogens will have similar handling methods. The advantages of this research are the existence of a proper diagnosis of pathogenic fungi, development of biological control methods of pathogens using microbial antagonists (like Trichoderma sp.), reduction of pesticides application on farms, and increasing agricultural and plantation production.
2. Materials and Methods
2.1. The Source of Pathogenic Fungi Isolate
Pathogenic fungi isolates used in this study were obtained from Nugrahani research [9]. They were isolated from soil samples under canopy of apple plants (from five different sides) which showed symptoms of diseases caused by the pathogenic fungi. Pathogenic fungi were isolated by modified baiting method [9].
2.2. DNA Isolation of Pathogenic Fungi
Three isolates of pathogenic fungi (M1, M4, and MB1) was sub cultured in V8 juice medium. Each isolates were sampled and inoculated in 25 ml of medium Jungle Juice (V8 Juice instead of medium order) then incubated in a shaker incubator at 28˚C with speed of 120 rpm for three days. DNA genome was isolated from the mass of fungi mycelium after incubated for three days using a modified method [16].
2.3. Amplification of DNA Sequences Pathogenic Fungi
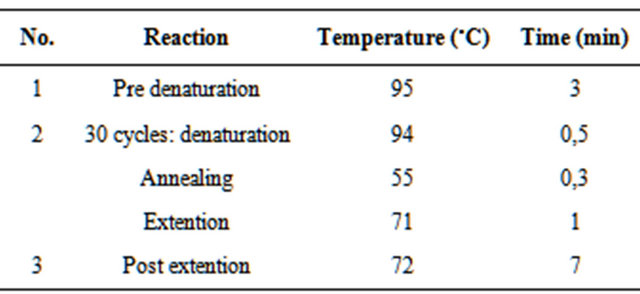
ITS region was amplified using ITS5 (5’-GGAAGTAAAAGTCGTAACAAGG-3’) and ITS4 (5’-TCCTCCGCTTATTGATATGC-3’) [12]. Composition of the compounds used for the amplification of DNA sequences (Table 1) [17] with modifications to the DNA sequence amplification reaction condition are shown in Table 2. The primers ITS5 and ITS4 will produced a single band of approximately 900 bp [18]. The PCR result were visualized by electrophoresis and documented by UV transluminator [16].
2.4. DNA Sequencing Pathogenic Fungi
Purification and sequencing of DNA amplicons performed at Macrogen, South Korea.
2.5. Phylogenetic Analysis
Data sequences of all fungi isolates, several isolates of the reference sequence (Phytophthora) and reference isolates as outgroup (Pythium and Peronospora) from the GeneBank (http://www.ncbi.nlm.nih.gov) were presented in FASTA format. Alignment of DNA sequences was done using ClustalW program. Phylogeny tree was constructed using Phylogeny in MEGA 5.03 program with 1000 bootstrap, inversion using Maximum Likelihood methods and evolutionary distance were analyzed according to Tamura-Nei model [16].
3. Result
Amplification of DNA sequences by primers ITS5 and ITS4 produced amplicons of 900 bp (Figure 1). These results have similar with the results of other research.
Table 1. Composition of the compounds used for the amplification of DNA sequences.
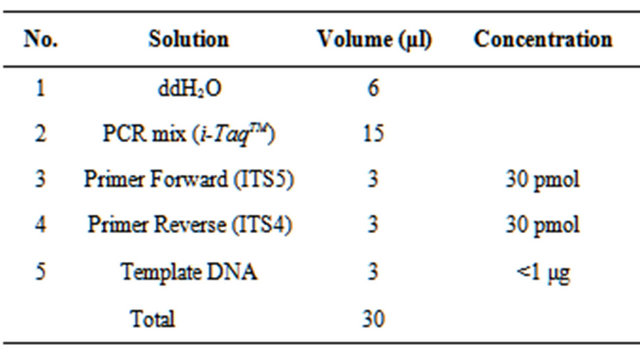
Table 2. The DNA sequence amplification reaction condition.
Based on the analysis, it is known that the three isolates belong to the same species (Figure 2). The results of DNA sequences show isolates M1 had similarity value of 99.73% with isolates MB1 (difference in evolutionary distance of 0.0027) and 99.46% with M4 isolates (the differences in the evolutionary distance of 0.0054). M4
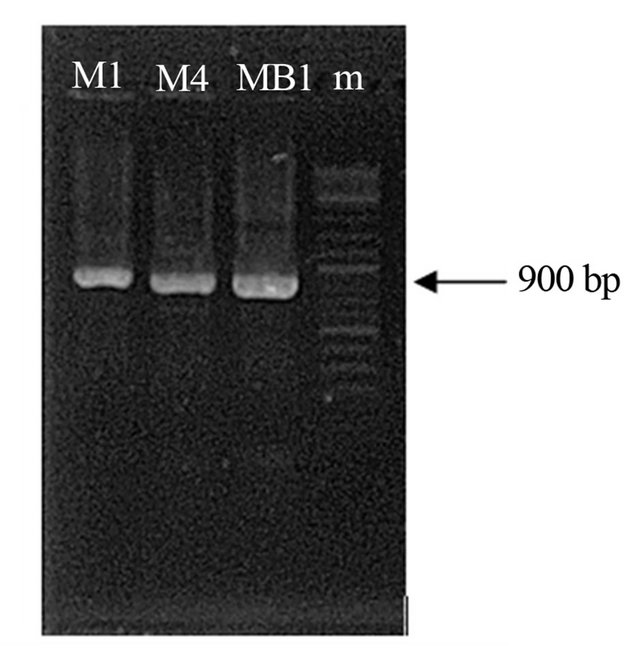
Figure 1. DNA amplicons with primer ITS4/ITS5. M1, M4 and MB1 is a pathogenic fungi isolates, while m is a marker (GeneRuler™ DNA Ladder Mix 100).
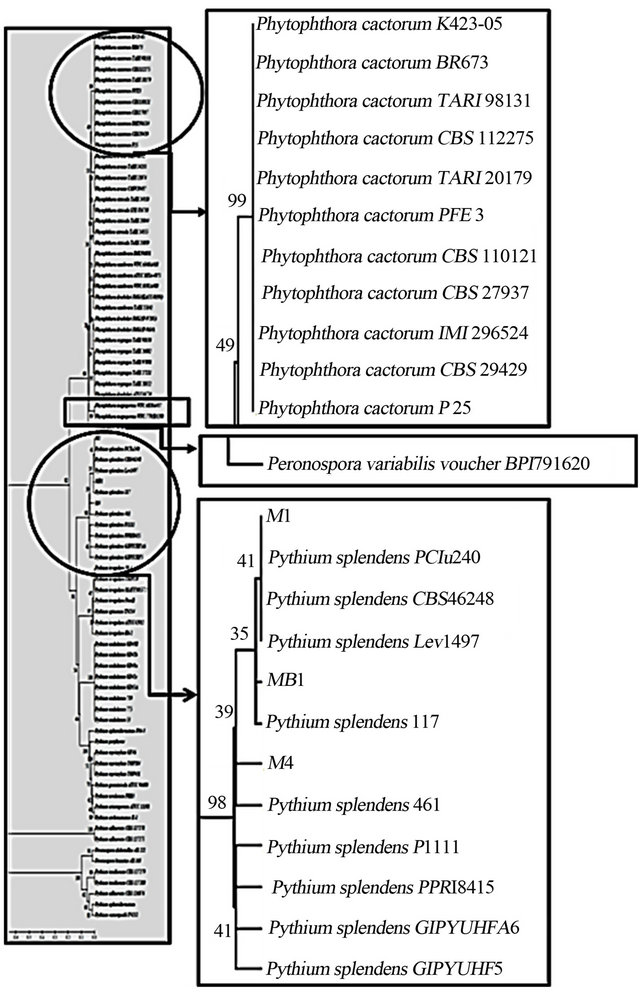
Figure 2. Phylogeny tree of pathogenic fungi and isolates members of the Phytophthora, Pythium, and Peronospora based on maximum likelihood algorithm with Tamura-Nei method of analysis.
isolates had a similarity value of 99.19% with MB1 isolates (differences in the evolutionary distance of 0.0081). Phylogeny tree showed that the three isolates were belong to genera Pythium and species of Pythium splendens with similarity values of more than 99%, while the similarity value of those isolates with Phytophthora were less than 76% (Figure 2).
This study also reviewed several morphological characters of pathogenic fungi isolates (M1, M4 and MB1) which showed the characters of Phytophthora and Pythium which was used by Nugrahani [9]. Dendogram which was built based on morphological data showed that those isolates had similarity value of 88% with Phytophthora and Pythium (Figure 3).
4. Discussion
At present, amplification of DNA sequence by primers ITS5 and ITS4 similar with the results of Cooke et al. [18] which is always produce a single band around 900 bp. It is consisted of small sub unit (SSU) 18S partial sequence, the ITS1, ITS2, 5.8S rDNA complete sequence, and large subunit (LSU) 28S rDNA partial sequence [15].
The DNA sequence data was utilized to construct phylogenetic tree using Maximum Likelihood method [19]. This method uses a calculation to find a tree that has the best calculation of the variation in data sequence. It based on the amount of sequence change or mutation that provides variations in sequences [20]. It was used to determine the relationship among the sequences which more diverse [20]. Maximum Likelihood methods considered to be most suitable to analyze the data because the rapid reproduction of pathogenic fungi which made possible the mutation and rapid changes as well so it can easily be used to distinguish the three isolates [20].
The results of DNA sequences show isolates M1 had similarity value of 99.73% with isolates MB1 and 99.46% with M4 isolates. M4 isolates had a similarity value of 99.19% with MB1 isolates. Phylogeny tree showed that the three isolates were belong to genera Pythium and species of Pythium splendens with similarity values of more than 99%, while the similarity value of those isolates with Phytophthora were less than 76%
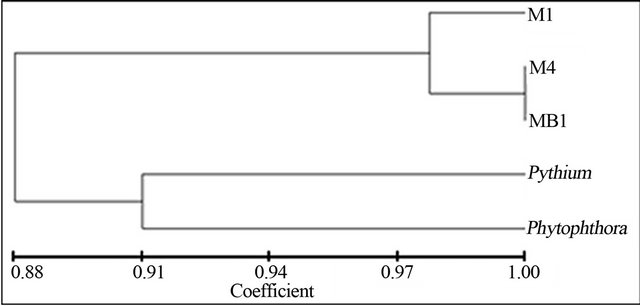
Figure 3. Dendogram of phenotype similarity of pathogenic fungi, Phytophthora and Pythium.
(Figure 2). Similarity value of 100% indicates that the test isolates were single strain and if the similarity value of 99% indicates similar species, whereas the similarity value of 89% - 99% is included into similar genus [19, 21]. DNA sequence differences in one species only ranged between 0.2% - 1% [19,21].
The analysis showed that three isolates are Pythium splendens and it was distinguished strain specific, because the similarity value of three isolates only 99%. Species were included in one strain if they have a similarity value of 100% [19]. Determination of a strain isolates required more than one pair of primer to obtain the valid results, although the ITS region is one of DNA barcode marker for fungi. However, ITS regions, SSU and LSU in the barcode gap analysis of inter-specific and intra-specific variation [14], were not show good results despite the high success rate of amplification. Barcode gap analysis showed that the ability of ITS to indicate an average gap in barcode at inter-spesific variation of 0.372664 and intraspecific variation of 0.025013. Therefore, primers that may be used for further identification of the strain or the intra-specific level were needed.
The morphological analysis shows there are error result. Base on the species concept [22], if the interspecific similarity value of isolates phenotype more than 50%, the isolates were grouped into one Genus, and if the similarity value more than 80% the isolates were grouped into one Species. However the morphological character identified in this research were not enough for distinguishing those isolates. So, the similarity value of 88% can not be used to determine the species.
Morphological character which using in identification affect the result of phenetic classification. Morphological character identified [9] such as round, single, papillate sporangium with size variation; unbranched, smoothwalled, and 2.5 - 5 µm width sporangiophore; and round or oval intercalary clamidiosphore with diameter of 15 µm (Figure 4) can be used for identifying the species (Phytophtora or Phytium). On the other hand some characters such as the upright grow of mycelium, reproductive structures, the thickness of sporangium wall, pigmentation of sporangium, the existance of papillae, the evidence of septate hyphae, and cytoplasm character can not be used for identification. According to other researcher [23], morphological character specific for Genus Phytophthora including nonseptate hyphae, sexual reproduction structure with biflagellate zoospore, posses sexual organs in the form of antheridium and oogonium, the sporangiosphore can not be distinguished from its mycelium, and posses ovoid sporangium in the shape of lemon and had papillae. The evidence of papillate sporangium is the specific character of Phytophthora sp. which distinguished it from Pythium sp. Only several member of primitive Phytophthora that was not found to have papillate sporangium (Figures 5 and 6).
Those observation lead to the conclusion that the isolates identified more likely to be the member of Genus Pythium based on the morphological character such as rounded sporangium and the absent of papillate sporangium (Figure 4). Taxonomy determination using morphological character for species identification need a long time, intensity of researchers and have high error probability [25]. However the physiological and bio chemical
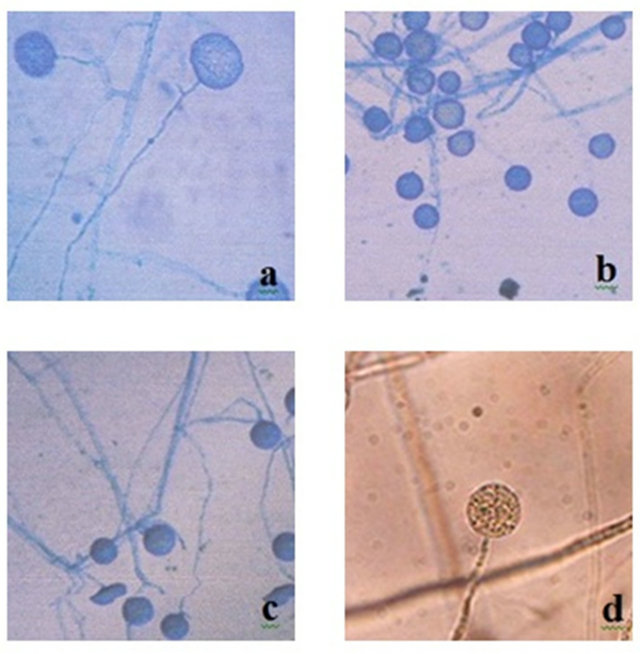
Figure 4. Morphological of nonpapillate sporangium; (a) M1, (b) M4, (c) MB1 [9], (d) Pythium splendens (magnification 400×) [24].
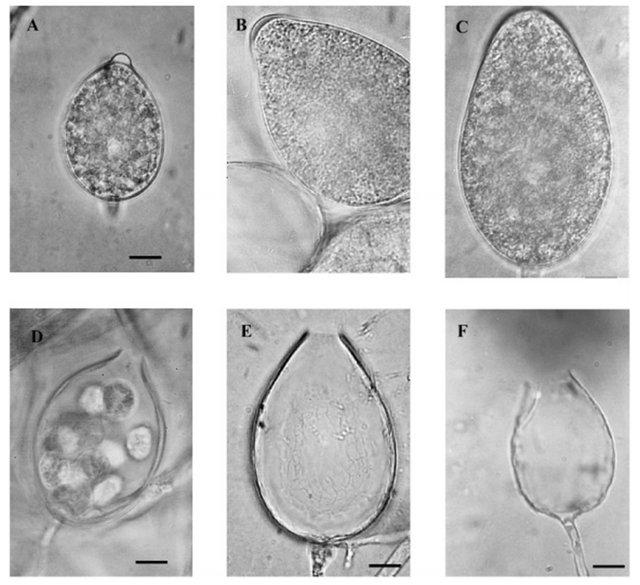
Figure 5. Morphological sporangium of Phytophthora [26]; (A) Papillate sporangium of P. cactorum, (B) Semipapillate sporangium of P. citricola, (C) Nonpapillate sporangium of P. pistaciae, (D) Narrow exit pore of P.cactorum, (E) Medium exit pore of P.citricola, (F) Wide exit pore of P. lateralis.
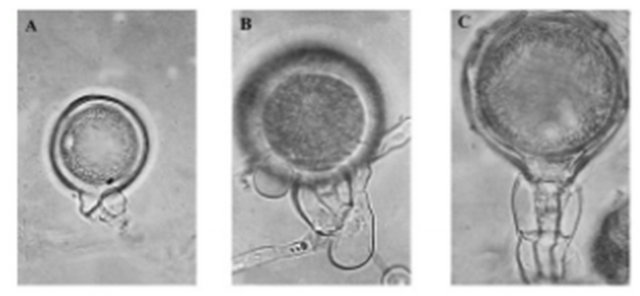
Figure 6. Morphological antheridium of Phytophthora [26] (A) Paragynous antheridium of P. citricola. (B) Amphigynous one-cell antheridium of P. inundata. (C) Amphigynous two-cell antheridium of P. Cambivora.
test are needed to be done in future studies, so it could be further validation for the conclusion.
There are some key character for traditional classification of Phytophthora [27] such as morphological of sporangium (papillate, semipapillate, and nonpapillate) combination character of antheridium type (amphigynous and paragynous) (Figures 5 and 6), and reproduction system (homothallic and heterothallic). Error in characterization of those isolates [23] could be the result of failure in identification of some character of Phytophthora and Pythium. Therefore, in this research phylogenetic relationship based on molecular identification was used to determine the true name of the pathogenic fungi.
Phytophthora produces asexual spores at suitable environmental conditions. Asexual spores are called sporangia that are formed in sporangium. Sporangium has variation of the shape like ovoid, obovoid, ellipsoid, limoniform (like lemon) and pyriform (like pear). Sporangia would germinate and form germ tubes in contact with plants [5]. Sexual spores are called zoospores that are produced through the fusion of antheridium and oogonium. Zoospores can be spread by splashing water flow at ground level. These spores have flagella that can help the movement to approach the host [28]. On the other hand Pythium produces spherical bodies called conidia (asexual reproduction or vegetative spores) which are easily separated and serves to help its spread. Especially, Pythium splendens is not like other Pythium species that have a structure or swimming motile zoospores, but when germinate to form germ tubes that can be used to penetrate or infected the host tissue. In addition, the thick-walled sexual spores (oospore) are only formed when the sexual reproduction happen. Oospore allows the fungi to survive in unfavorable conditions [29].
Plants attacked by Phytophthora characterized by the rot in the stem bark. In addition, the infected plant part would be browned rot. The infection can extend to the roots and to the stems base start from external part, causing the death of plant. Sometimes, it was not widespread, but the skin on the affected plants will rot, become dry and detached. It leads to cancer cell development [30]. Phytophthora attack annual plants and seedling of perennial plants. Old crops which are attacked by Phytophthora would show symptoms such as leaf number decreased, leaf yellowing and death of twigs or branches. It also showed the death of small roots and the presence of necrotic brown lesions form on the old roots. The root systems of young or old plants are destroyed, followed by the death of the plant [3]. The typical symptoms caused by Pythium attacks can be seen in the morning by the present of spider threads—like hypha with drops of dew hanging. It often attacks young plants near the soil which causes leaf spot which eventually will grow to be a wide spot [30]. It commonly causes seed rot as well as preand post-emergence damping-off of plants. Pythium causes lobed stems 1 - 15 cm and also results in root-lobed that significantly reduce root length. In addition, the effect of it causes the bark totally broken and has hollow stem pith. The young stems of woody plants will collapse. Eventually plants become blacked rotten and the roots of infected plants will brown and watery. It causes a seed rot, damping-off, and seedling root rot, caused a watery and soft decay of older plants in wet soils. Pythium attack can also be found in warm temperatures and subtropical regions, and it has a wide host range and is not only founded on the plant parasite [30].
Handlings of pathogenic fungi have generally been carried out by spraying a systemic fungicide. Systemic fungicide worked its way into the vascular system of plant that inhibited infection into plant tissue. However, the use of systemic fungicide continuously leads to possibility of development of new strains of pathogenic fungi. In addition, it can also cause environmental pollution and mortality of non-target organisms in the surrounding environment. Therefore, this result can support the development of biological control methods using specific microbial antagonists and proper management to overcome the attacks of pathogens and also to understand the sensitivity of those pathogenic fungi caused by additional of organic compound and environmental stress.
Molecular biology gives impact more and more heavily on mycology [11]. Species identification based on phenotypic identification is often time-consuming, need high passion of the researchers and have high error probability [25], and it is also has subjective nature of phenotypic characteristics, which are readily influenced by culture conditions [11]. Conversely, molecular phylogenetic identification involving gene sequencing is more objective, uninfluenced by growth conditions, and capable of discriminating between fungi that fail to produce distinctive morphological features [11].
5. Conclusion
Conclusions from this study were three isolates of pathogenic fungi (M1, M4 and MB1) are similar species with DNA sequence similarity value of more than 99%. These three isolates are closely related and members of Pythium splendens with DNA sequence similarity value of more than 99%.
REFERENCES
- A. Dreth, G. Wagels, B. Smith, B. Sendall, C. O’Dwyer, G. Irvine and J. A. G. Irwin, “Development of A DNABased Method for Detection and Identification of Phytophthora Species,” Australasian Plant Pathology, Vol. 35, No. 2, 2004, pp. 147-159. doi:10.1071/AP06018
- L. Schena and D. E. L. Cooke, “Assessing the Potential of Regions of the Nuclear and Mitochondrial Genome to Develop a “Molecular Tool Box” for the Detection and Characterization of Phytophthora Species,” Journal of Microbiological Methods, Vol. 67, No. 1, 2006, pp. 70-85. doi:10.1016/j.mimet.2006.03.003
- Balitjestro, “Phytophthora sp. Patogen Penyebab Penyakit pada Tanaman Jeruk dan Buah Subtropika,” 2011. http://balitjestro.litbang.deptan.go.id
- A. Belenin, Z. Borecki and D. F. Millikan, “Identification of Pythium ultimum in the Collar Rot Complex of Apple,” Phytophatology, No. 66, 1976, pp. 127-129. doi:10.1094/Phyto-66-127
- D. C. Erwin and O. K. Riberio, “Phytophthora Diseases Worldwide,” APS Press, St. Paul, 1996.
- R. C. Ploetz, “Tropical Fruit Crops and the Diseases That Affect Their Production,” UNESCO-Encyclopedia of Life Support Systems (EOLSS), Florida, 2005.
- M. Souli, N. Boughalleb, P. Abad-Campos, L. A. Álvare, A. Pérez-Sierra, J. Armengol, J. Garcia-Jimenez and M. S. Romdhani, “Diversity of the Pythium Community Infecting Crown and Roots Apple in Tunisia,” Research in Plant Biology, Vol. 1, No. 4, 2011, pp. 16-22.
- M. Mazzola, P. K. Andrews, J. P. Reganold and C. A. Levesque, “Frequency, Virulence, and Metalaxyl Sensitivity of Pythium spp. Isolated from Apple Roots Under Conventional and Organic Production Systems,” Plant Disease, No. 86, 2002, pp. 669-675. doi:10.1094/PDIS.2002.86.6.669
- M. Nugrahani, “Eksplorasi Kapang Antagonis Terhadap Phytophthora sp. Patogen pada Tanaman Apel,” Scholar Thesis, Brawijaya University, Malang, 2011.
- T. W. Darmono, I. Jamil, and D. A. Santosa, “Pengembangan Penanda Molekular untuk Deteksi Phytophthora palmivora pada Tanaman Kakao,” Menara Perkebunan, Vol. 74, No. 2, 2006, pp. 87-96.
- A. M. Borman, C. J. Linton, S.-J. Miles and E. M. Johnson, “Molecular Identification of Pathogenic Fungi,” Journal of Antimicrobial Chemotherapy, Vol. 61, No. S1, 2008, pp. i7-i12.
- T. J. White, T. Bruns, S. Lee, and J.Taylor, “Amplification and Direct Sequencing of Fungal Ribosomal RNA Genes for Phylogenetics,” In: M. A. Innis, D. H. Gelfand, J. J. Sninsky and T. J. White, Eds., PCR Protocols: A Guide to Methods and Applications, Academic Press, San Diego, 1990, pp. 315-322.
- A. A. Appiah, J. Flood, S. A. Archer and P. D. Bridge, “Molecular Analysis of the Major Phytophthora Species on Cocoa,” Plant Pathology, Vol. 53, No. 2, 2004, pp. 209- 219.
- C. L. Schoch, K. A. Selfert, S. Huhndorf, V. Robert, J. L. Spouge and C. A. Levesque, “Nuclear Ribosomal Internal Transcribed Spacer (ITS) Region as a Universal DNA Barcode Marker for Fungi,” Proceedings of the National Academy of Sciences of the USA, Vol. 109, No. 16, 2011, pp. 1-6. doi:10.1073/pnas.1117018109
- Y. Matsuda, T. Sameshima, N. Moriura, K. Inoue, T. Nonomura, K. Kakutani, H. Nishimura, S. Kusakari, S. Takamatsu and H. Toyoda, “Identification of Individual Powdery Mildew Fungi Infecting Leaves and Direct Detection of Gene Expression by Single Conidium Polymerase Chain Reaction,” The American Phytopathological Society, Vol. 95, No. 10, 2005, pp. 1137-1143.
- J. J. Doyle and J. L. Doyle, “Isolation of Plant DNA from Fresh Tissue,” Focus, No. 12, 1990, pp. 13-15.
- Intron Biotechnology, “2x PCR Master Mix Solution (iTaqTM),” 2011. http://eng.intronbio.com
- D. E. L. Cooke, A. Drenth, J. M. Duncan, G. Wagels and C. M. Brasier, “A Molecular Phylogeny of Phytophthora and Related Oomycetes,” Fungal Genetics and Biology, Vol. 30, No. 1, 2000, pp. 17-32. doi:10.1006/fgbi.2000.1202
- Suharjono, A. P. W. Marhendra, A. Triwiratno, S. Wuryantini and L. O. Rahayu, “Sistematik Filogenetik Isolat Kapang Indigenous Indonesia Sebagai Entomopatogen Kutu Sisik (Lepidoshapes beckii Newman) Hama Tanaman Jeruk,” Biota, Vol. 15, No. 2, 2010, pp. 231-236.
- N. L. P. I. Dharyamanti, “Filogenetika Molekuler: Metode Taksonomi Organisme Berdasarkan Sejarah Evolusi,” Balai Besar Penelitian Veteriner, Bogor, 2011.
- B. D. Shenoy, R. Jeewon and K. D. Hyde, “Impact of DNA Sequence-Data on the Taxonomy of Anamorphic Fungi,” Fungi Diversity, No. 26, 2007, pp. 1-54.
- C. R Vanderpool, “Bacterial Chemistry and Structure,” 2000. http://zoey.med.howard.edu
- S. Purwantisari and R. B. Hastuti, “Isolasi dan Identifikasi Jamur Indigenous Rhizosfer Tanaman Kentang dari Lahan Pertanian Kentang Organik di Desa Pakis, Magelang,” BIOMA, Vol. 11, No. 2, 2009, pp. 45-53.
- A. E. Dorrance, “Characterization of Pythium spp. from Three Ohio Fields for Pathogenicity on Corn and Soybean and Metalaxyl Sensitivity,” Plant Management Network, St. Paul, 2004.
- C. C. Siricord, “Detection of Phytophthora Spesies by MALDI-TOF Mass Spectrometry,” Murdoch University, Perth, 2005.
- M. Gallegly and C. Hong, “Identifying Species by Morphology and DNA Fingerprints,” APS Press, St. Paul, 2008.
- N. O. Villa, K. Kageyama, T. Asano and H. Suga, “Phylogenetic Relationships of Pythium and Phytophthora Species Based on ITS rDNA, Cytochrome Oxidase П and β-Tubulin Gene Sequences,” Mycologia, Vol. 98, No. 3, 2006, pp. 410-422. doi:10.3852/mycologia.98.3.410
- C. J. Alexopoulos, C. W. Mims and M. Blackwell, “Introductory Mycology,” 4th Edition, John Wiley & Sons Inc., New York, 1996.
- J. Y. Uchida, “Pythium Root Rot of Spathiphyllum,” Hawaii Institute of Tropical Agriculture and Human Resources, Hawaii, 1990.
- H. Semangun, “Pengantar Penyakit Tumbuhan,” Gadjah Mada University Press, Yogyakarta, 1996.

