Health
Vol.4 No.10A(2012), Article ID:24263,4 pages DOI:10.4236/health.2012.430150
First isolation and identification of H1N1 swine influenza viruses in Colombian pig farms
![]()
Línea de Microbiología y Epidemiología, Posgrado en Salud Animal, Facultad de Medicina Veterinaria y de Zootecnia, Universidad Nacional de Colombia, Bogotá, Colombia; *Corresponding Author: gcramirezn@unal.edu.co
Received 17 September 2012; revised 15 October 2012; accepted 27 October 2012
Keywords: Swine Influenza; H1N1; Colombia; First Isolation; Classical; Pandemic
ABSTRACT
The pig industry in Colombia has grown 30% in the last decade achieving high levels of technology and efficiency; in spite of that, respiratory diseases remain a constraint. Since 1970, serological evidence and histological findings suggested the role of swine influenza virus (SIV) as part of the porcine respiratory disease complex; nevertheless, elusive and molecular typing isolates are missing. This study was aimed at isolating SIV from intensive pig farms and to achieve molecular characterization to determine strains circulating in the field. In order to accomplish this goal, 242 samples were taken from nasal swabs, 25 from bronchial washes and 8 from lung tissue. Samples were collected during a period of three years, between 2008 and 2010 and were originated from 78 farms of the three main pig production regions of the country. The samples were transported in BHI broth with 2% antibiotic and antimycotic solution and stored at –70˚C until processed. The swabs were inoculated in 9 - 11 days old embryo chicken eggs and in MDCK (Madin Darby Canine Kidney) cell cultures with the addition of trypsin. The isolates were identified by the HA (hemoagglutination) test and by RT-PCR targeting the HA (hemagglutinin), NA (Neuraminidase) and M (Matrix) genes. Full length sequence of the HA and NA glycoproteins from four selected virus isolates was conducted (Macrogen®, USA). As a result, fifteen SIV isolates from nine farms distributed in the three regions were obtained. Twelve of the isolates are related to the swine origin H1N1 virus that caused the 2009 influenza pandemic. The remaining three viruses were related to classical swine influenza viruses.
1. INTRODUCTION
The Porcine Respiratory Disease Complex (PRDC), a multifactorial condition is a major treat to the pig Industry worldwide [1]. The PRDC is a mixed infection etiologically related to viral and bacterial infections. Most commonly, viral infections such as the porcine reproductive and respiratory syndrome (PRRS), porcine circovirus-2 (PCV-2), Aujesky’s disease virus and the swine influenza virus (SIV) play a major role in this condition. Bacterial infections including Mycoplasma hyopneumoniae, Actinobacillus pleuropneumoniae, Haemophillus spp, Streptococcus suis and Pasteurella multocida may also be involved [1-3].
Several studies have shown the importance of swine influenza virus infection associated with the porcine respiratory disease complex [4,5]. SIV infection causes coughing, sneezing, nasal discharge, fever, lethargy and difficulty breathing. Although mortality is low and recovery could occur within 7 - 10 days post-infection, morbidity can reach up to 100% of the animals [6]. Swine influenza is caused by Influenza A virus, a member of the Orthomyxoviridae family. Influenza A viruses have been isolated from different species including, humans, pigs, horses, dogs, sea mammals and birds [7,8] between others. The virus has a single-stranded RNA segmented genome. Based on antigenic differences in the surface glycoproteins, hemagglutinin (HA) and neuralminidase (NA), influenza A viruses are further classified into 16 HA and 9 NA subtypes [8,9]. Although other subtypes have been identified, the influenza A virus sub-types H1N1, H1N2 and H3N2 are the most prevalent in pig populations worldwide [7,10]
The pig industry in Colombia has grown about 30% during the last decade. With a swine population close to four million heads, located in four geographical regions it is one of the major contributors to the GDP of the country (ICA, 2012). In spite of achieving high levels of technology and efficiency, respiratory diseases remain a major constraint. In this sense the PRC is a major cause of economic loss in the pig industry of Colombia affecting mainly growing and finishing pigs. Considering SIV as one of the pathogens involved in this complex and having serologic evidence suggesting influenza virus activity since 1971 in pigs from Colombia, establishing a 14% reactivity against the H3N2 SIV subtype in Antioquia [11]. Studies conducted in samples collected between 1991-1994, demonstrated 6.5% reactivity to the H1N1 SIV [12]. Later on in samples from 1997-1999, a reactivity of 41.3% to H3N2 virus and 0.8% to H1N1 virus (Antioquia) was determined [13]. Additional studies reported reactivity of 19.85% in sows (multiparous) and 3.13% in replacement animals [14]. Another study conducted in 2003 reported prevalence of 10% for H3N2 and only a 0.4% for H1N1 for the three major producing regions of Colombia [15]. In spite of this, the role of the SIV in the respiratory disease complex has not been established in Colombia and there are no SIV strains isolated from the field until now.
Based on the fact that SIV has been circulating in Colombian pigs for more than thirty years and that the only information available during this time is related to serological activity against the virus, it is evident that there is a need to study and determine the type of influenza virus causing the infection in the field in addition to establishing the population serological status. Therefore, it is required to isolate these viruses in order to characterize them trying to understand swine influenza virus infection under Colombian conditions. This knowledge will provide clues to propose and develop systematic control programs according to local conditions.
The importance of knowing the characteristics of the swine influenza viruses acting upon the field in Colombia goes beyond animal health and the negative economic impact to the swine industry. The consequences of SIV infection have an impact at the local and at the international level due to the implications of influenza virus infections as a public health treat.
The main goal of this research was to conduct a systematic study on swine influenza virus infection, in the major pig producing areas of the country over a period of three years. The focus of the study was to know the dynamic of this viral infection and to establish characteristics of the strains circulating in the field in Colombia.
2. MATERIALS AND METHODS
2.1. Samples
A statistically representative number of 78 herds were selected from farms located in three of the major swine rearing areas in Colombia: Antioquia (North West region), Valle del Cauca (West region) and Cundinamarca (Central region) Figure 1. A total of 43 animals per farm were used to approach serological status against SIV. The range of age covered animals from nursing, growing to
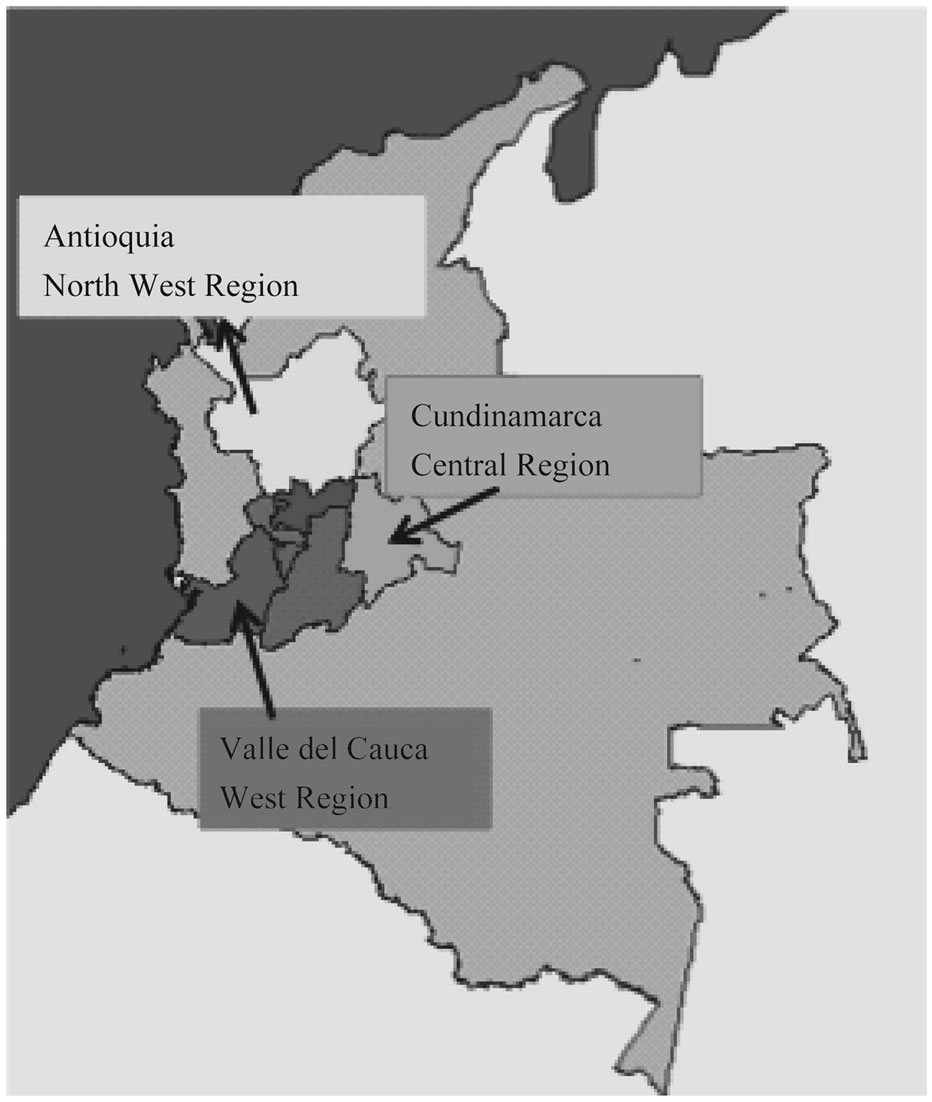
Figure 1. Farms geographical location where H1N1 SIV was isolated in Colombia.
finishing period. It is important to mention that SIV vaccination program had never taken place at the farms in Colombia. In order to attempt virus isolation, samples were collected from animals showing respiratory signs compatible with swine influenza infection.
2.2. Clinical Samples
The present study was conducted over a period of three years from 2008-2010. During this time a total of 275 samples were collected from animals showing clinical respiratory signs of cough, fever and depression. From the 275 samples collected, 242 corresponded to nasal swabs, 25 to lung tissue and the remaining 8 samples were from bronchial aspirates. Samples were collected in BHI media (BD®) supplemented with 2% antibiotic and antimycoticsolution (Sigma®), filtered through 0.22 μm filter (MILLEX® GV) and stored at –70˚C until processing.
2.3. Virus Isolation
The 275 filtered samples were inoculated into MadinDarby Canine Kidney (MDCK) cells and on 9 to 11 dayold SPF chicken embryo eggs (CEE) (Charles Rivers®, USA). Briefly, confluent monolayer of MDBK cells prepared in six well plates were washed twice with PBS and infected with 200 µl of the original sample. Cells were grown in Dulbecco’s Modified Eagle Medium (DMEM) (Gibco®) supplemented with 5% FCS (Gibco®), 1% antibioticand antimycotic solution (Sigma®), 1% L-glutamine (Sigma®), 1% HEPES buffer (Sigma®) and 2 µg/µl TPCK trypsin (Sigma®). The plates were incubated at 37˚C for an hour to allow virus adsorption, followed by an addition of fresh medium and an incubation period at 37˚C for 72 hours. Cells were observed daily to establish cytophatic effect (CPE) characteristic of influenza virus. On the other hand, samples were also inoculated through the allantoic cavity into 9 to 11 days old SPF chicken embryo eggs (CEE) (Charles River®, USA) and incubated at 37˚C for 72 hours.
Following the incubation period, the cell culture supernatant (CS) and the allantoic fluid (AF) were collected and tested by hemagglutination assay (HA) using chicken eritrocytes following standard procedures according to the OIE protocol (Swine influenza, 2008) [16].
2.4. Viral RNA Extraction and RT-PCR
All HA positive samples resulting from cell culture or egg inoculation were tested to determine the subtype of virus isolates. Viral RNA was extracted using the commercial RNeasy® Mini kit (Qiagen®, CA, USA), following the manufacturer’s recommendations. Subtyping of the virus was done by amplification of the HA and NA segments by a duplex RT-PCR assay targeting the HA and NA genes of H3N2 and H1N1 swine influenza viruses [17]. An initial reverse transcription was performed with M-MVL reverse transcriptase (Invitrogen®) using 5 µl of extracted viral RNA as template and the uni12 primer (5’-AGCAAAAGCAGG-3’) [18]. PCR amplification conditions were as recommended by Choi, Y., Goyal, S., Kang, S., Farnham, M., Joo, H., 2002 [17]. Additional RT-PCR targeting the M gene of the influenza A virus was conducted, using primers Mf: 5’ACG GAT GGG AGT GCA GAT AC 3’ and Mr: 5’ GAA GGC CCT CTT TTC AAA CC 3’. The expected size of the amplicon was 153 bp. Amplified products were then separated onto agarose gel and purified using the Qiaquick® gel extraction kit (Qiagen®, CA, USA).
2.5. Sequencing and Phylogenetic Analysis
Both strands of the amplicons were sequenced with the same forward and reverse primers used for the amplification. Full-length HA and NA amplicons were also sequenced using internal primers [18]. Gel purified products were sequenced by Macrogen®, USA, using BigDye® Terminator Cycle Sequencing. DNA sequences were combined and edited using the Lasergene sequencing analysis software package (DNASTAR®, Madison, Wisconsin). Multiple sequence alignments were made using Clustal W to identify related reference influenza genes. Phylogenetic analysis of full-length nucleotide sequence of the HA gene, was achieved from four selected viruses representing isolates from 2008, 2009 and 2010. Three were H1N1 pandemic (pH1N1) influenza viruses and one was H1N1 classical (cH1N1) SIV. Sequence comparisons and phylogenetic relationships through a bootstrap trial of 1000 were determined with the Mega 5.1 program using the Clustal W alignment algorithm and the evolutionary history was inferred using the Neighbor-Joining method [19] for tree construction. Gene sequences of the Colombian strains were compared with swine, avian and human influenza viruses which sequences were retrieved from the NCBI Influenza Virus Resource.
3. RESULTS
A systematic virological surveillance of influenza viruses in pigs from Colombia has led to the isolation of 15 field isolates of swine influenza virus, originated from nine farms located in the three geographical regions analyzed. Virus isolation was demonstrated on MDCK cell culture supernatant and on allantoic fluid through hemoagglutination activity performed using the HA assay. Viruses were grown initially in MDCK cells with TPCK. From the 15 isolates, initially 11 grew well in cell culture, two grew better in 9 to 11 days old chicken-embryo eggs, and the remaining two grew well in both substrates. After two to three passages in tissue culture the samples were inoculated in CEE, resulting in good performance for all of them. All the isolates were from nasal swabs and belonged to the H1N1 subtype. Twelve out of fifteen were pH1N1 viruses and the remaining three were cH1N1 SIVs. The 15 strains isolated in the time period between 2008-2010 belonged to 9 herds distributed in the three different geographical regions evaluated. The first three SIVs isolates were from 2008 and belonged to farms located in Antioquia; four out of eight isolates from 2009, were from Antioquia and the other 4 were from the western region. The last 4 isolates were from 2010 and belonged to farms located in the central region of the country. Information of the strains isolated in this study such as geographical origin, year of isolation and type of virus are reported in Table 1. All the isolates were obtained from nasal swabs and corresponded mainly to growing animals. All the isolated strains were sub-typed by a specific duplex RT-PCR, for HA and NA. Sequence analysis of the HA and NA genes showed that 12 out of 15 isolates corresponded to pH1N1 swine origin influenza virus, and 3 viruses corresponded to cH1N1 swine influenza virus. Interestingly, the three viruses isolated in 2008, corresponded to the cH1N1 swine influenza virus, while the remaining twelve viruses isolated during 2009- 2010, were identified as H1N1 pandemic-like.
The phylogenetic tree in Figure 2 shows clearly that SIVs isolated in Colombia are distributed into two groups: the pandemic and the classic H1N1 viruses. I
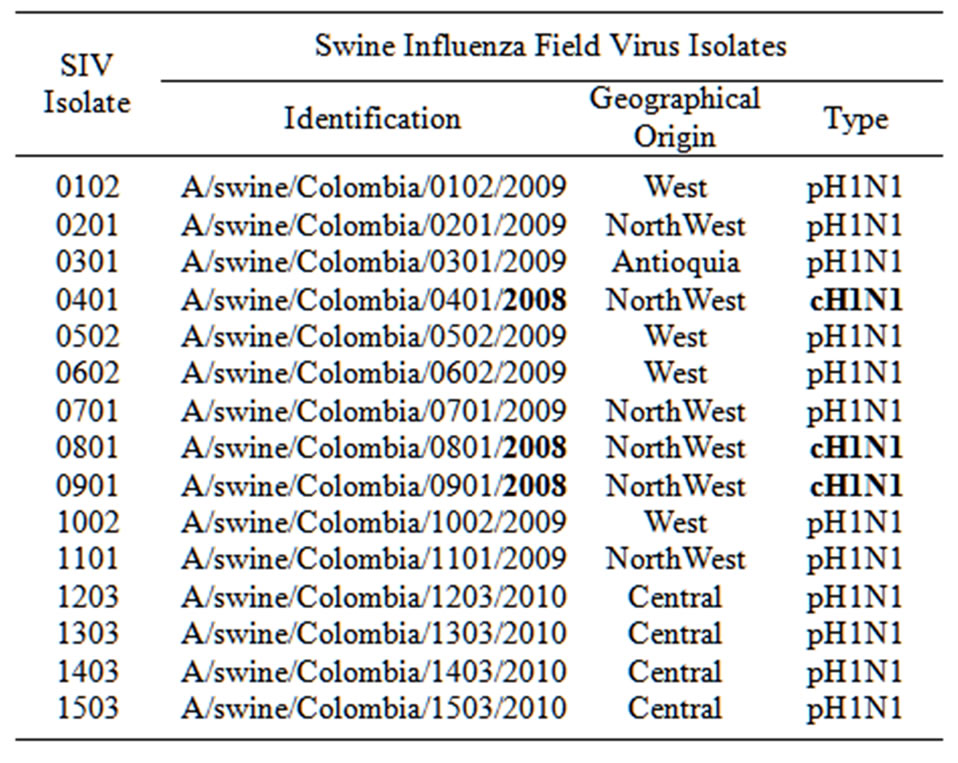
Table 1. Geographical origin and type of swine influenza viruses isolated in colombia during 2008-2010.
the upper part of the tree are pandemic viruses isolated during 2009 and 2010, including the California 04, 2009, originallyisolated from humans and considered the origin of the last influenza pandemic. These viruses show a 98.53% identity with the A/swine/Colombia/0102/2009 (pH1N1), A/swine/Colombia/1101/2009 (pH1N1) and the A/swine/Colombia/1403/2010 (pH1N1) isolated from pigs in Colombia during 2009 and 2010. The identity into these tree pH1N1 Colombian viruses is close to 99%. From Figure 2 is also evident that pH1N1 viruses isolated from Colombia are in the same group or cluster with viruses isolated from the United States, China, Italy, Argentina and Chile, as can be observed in Figure 2(a).
In relation to cH1N1 SIV, tree groups or clusters are shown in Figure 2. The first cluster includes only H1 swineisolates, in the secondcluster are H1 avianvirusesand in the third cluster are avian-like swine H1 viruses. The cH1N1 SIV isolated in Colombia A/swine/Colombia /0401/2008 belongs to the first cluster (Figure 2). This cluster is divided into four subgroupsone includes viruses isolated from the USA between 2008 and 2012. The second subgroupincludes mainly Asian isolates dated from 1957 to 1979 and are related to one from California, 1991. In the third subgroup the majority of the isolates are from the USA over a time frame period between 1957 - 1981. These viruses are related also to a Hong Kong, 1977. Finally, in the fourth of these subgroups, is the Colombian isolate: A/swine/Colombia/0401/2008 H1N1, along with the ancient H1N1 USA virus from 1931 and more recent isolates from Changhua, 2000 and Guandong (China) 2009 (Figure 2(b)); viruses in this subgroup share a 99.47% nucleotide sequence identity between them. Interestingly, the closest homology of the Colombian A/swine/Colombia/0401/2008 H1N1 SIV is with the Changhua (Taiwan) isolate, with only three nucleotides difference.
4. DISCUSION
It is well known that porcine respiratory diseases are one of the major problems to the swine industry, the presence of viral agents such as the PRRS and PCV-2 viruses in Colombia and, the evidence of bacterial infections in swine highlight the importance of a continuous study of the agents involved in this complex with a particular emphasis on influenza virus infection.
Although previous studies had shown serologic reactivity to swine influenza virus in Colombian pigs for more than thirty years, this paper reports the first isolation of several H1N1 SIV strains in commercial Colombian pig farms. There are probably several reasons that could explain this situation. One could be that swine influenza has been under diagnosed due to the similarity of clinical signs whit other pathogens causing respiratory disease in pigs. Another reason might be that some years ago there was limited laboratory facilities or lack of appropriate methods to isolate influenza virus from pigs in the country.
The year 2009 is remarkable in terms of swine influenza virus around the world, not only because of the pandemic it caused but due to the appearance of a novel influenza virus resulting from an unexpected reassortment originated from three different influenza virusesorigin [20]. According to the information previously available, serotypes H1N1 and H3N2 were circulating before 2009 in Colombia, as a result it is not surprising that the isolates from 2008 were of the cH1N1 SIV subtype. But what is important to point out is the evident predominance of pH1N1 swine influenza viruses isolated after that year, between 2009 and 2010. This might suggest that a novel virus with genetic characteristics that are different from the cH1N1 that was circulating previously in the herds, was able to enter the pig population and infect a number of susceptible animals. It seems that the pH1N1 influenza virus was able to surpass and replace the cH1N1 influenza virus that was circulating previously in the herds until 2009. Even though infection with different subtypes of influenza virus results in partial cross protection, the immune response in the animal facing heterologous viruses is subject to immune pressure and as a consequence immune selection [21]. Therefore, we could expect a large number of susceptible animals that were challenged by a new H1N1 influenza virus acting during the influenza pandemic in 2009. Additionally, it is necessary to emphasize that the pig population had never been vaccinated in Colombia, then it could be suggested that probably before this event there was an equilibrium between antibody response and virus challenge on the field, reducing immune pressure and maintaining the virus stable. After the entrance of a novel virus such as the pH1N1 influenza virus into the population is necessary to improve biosecurity measures to
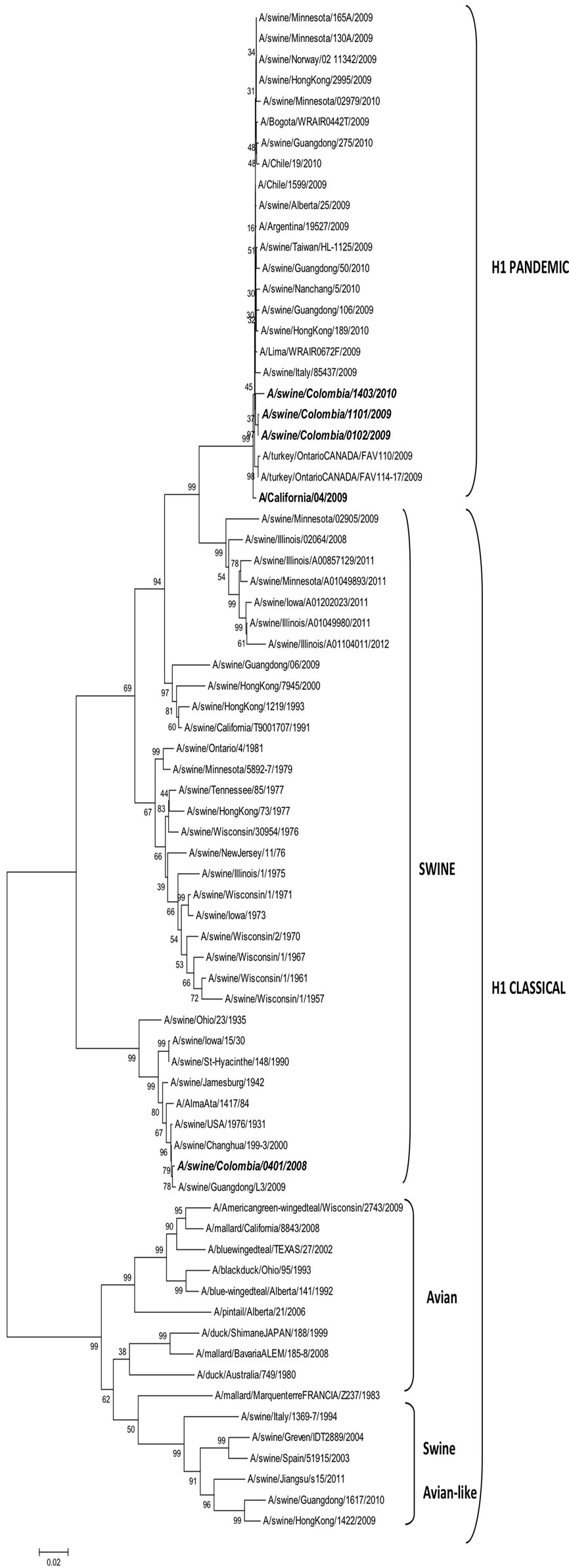
Figure 2. Phylogenetic relations of H1N1 SIVs isolated in Colombia. The evolutionary history was inferred using the Neighbor-Joining method [19]. The optimal tree with the sum of branch length = 1.62215850 is shown. The percentage of replica trees in which the associated taxa clustered together in the bootstrap test (1000 replicates) are shown next to the branches [22]. The tree is drawn to scale, with branch lengths in the same units as those of the evolutionary distances used to infer the phylogenetic tree. The evolutionary distances were computed using the Kimura 2-parameter method [23] and are in the units of the number of base substitutions per site. All positions containing gaps and missing data were eliminated. There were a total of 1675 positions in the final dataset. Evolutionary analyses were conducted in MEGA 5 [3].
 (a)
(a)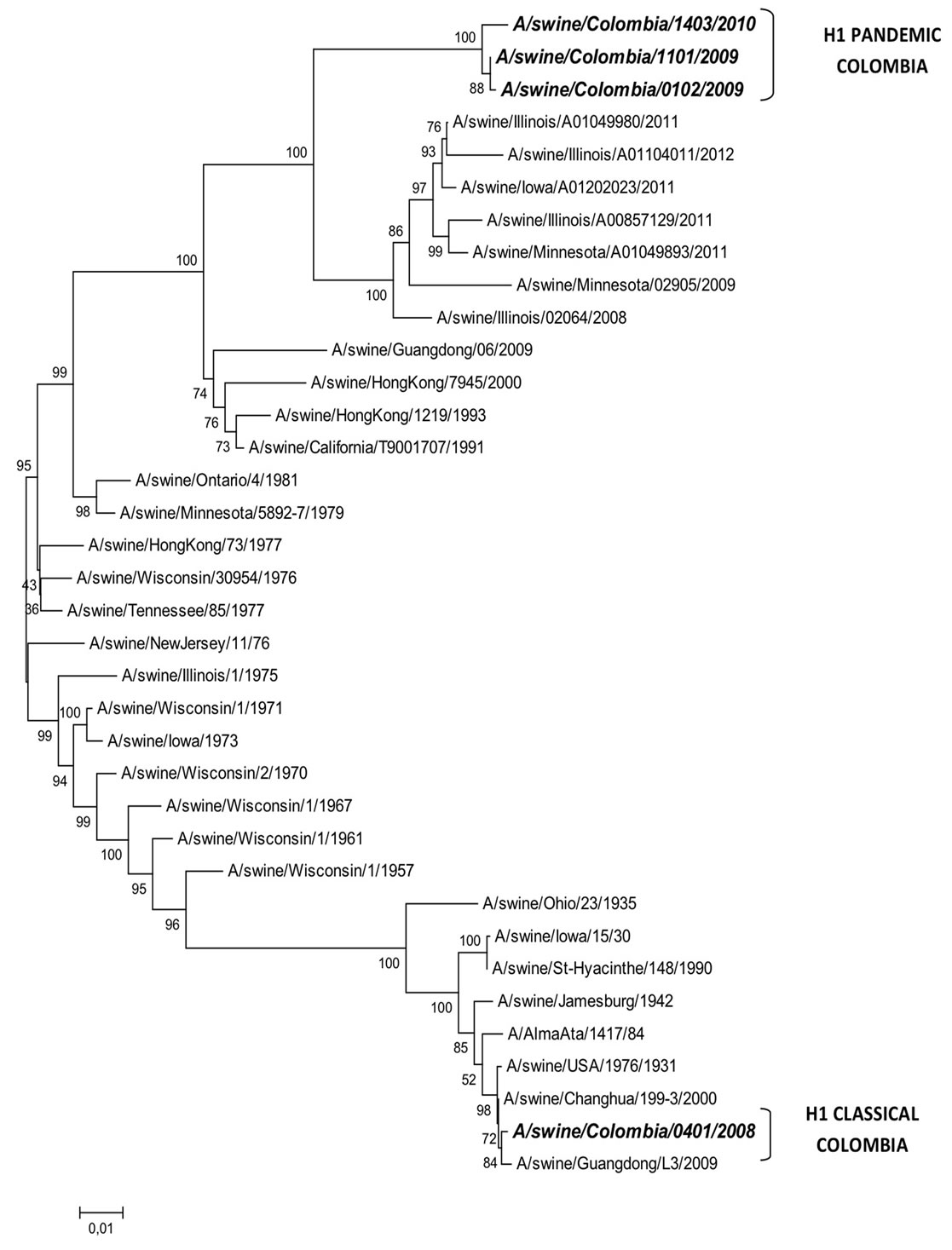 (b)
(b)
Figure 2. (a) Phylogenetic relation of pH1N1 SIVs isolated in Colombia; (b) Phylogenetic relation of cH1N1 SIV isolated in Colombia.
prevent further exposure to the virus.
Isolation of the H1N1 swine origin influenza virus for the first time in the field has remarkable impact. Analysis of full length sequence of the HA gene, the major antigenic determinant of influenza viruses, showed that the pH1N1 viruses isolated in Colombian pigs are phylogenetically related with human viruses from USA, Argentina, Chile, Perú, and with swine viruses originated mainly from USA and Asia. It is well known that in Colombia most of the commercial exchange in terms of live animals and semen is with USA, then it is not surprising to find this close relationship between the SIV isolates. This could also be suggested as the origin of influenza virus entrance to the country several years ago. However, the lack of information regarding the origin of breeding animals in the herds limits our capacity to follow back the origin of the viruses acting on the field.
It is worth noticing that although the difference in the HA sequence is slight, sharing a 99% homology between the HA of pH1N1 viruses isolated in 2009 compared to those isolated in 2010, it suggests that the virus is changing rapidly. In addition, considering that H3N2, and H1N1 pandemic viruses are circulating in the swine population in Colombia and that there is evidence of new viruses such as the H1N2 that emerged in Italy [24] as a result from reassortment between the H3N2 and the pH1N1 influenza virus it is imperative to continue studying swine influenza virus infection in Colombia.
On the other hand, the cH1N1 swine influenza virus isolated from Colombian pigs showed homology with virus from United States, Asia and Europe. Surprisingly, closest homology is with the cH1N1 SIV from Changhua, Taiwan, followed by the North American origin H1N1 viruses. In this case it is difficult to explain the relation to the virus from Taiwan and more research is needed in this topic.
In contrast to the pH1N1 influenza virus, the cH1N1 virus shows a higher genetic variability geographically and temporarily which is not surprising considering the time frame period of nearly 80 years since the first cH1N1 swine influenza virus was reported in 1931. These changes could be explained from the nature of influenza A virus replication itself and the evolution of influenza viruses over time. This shows an adaptation process making this a successful virus capable of survive and persist forever independent of time and space. Examples of the success of this process for the virus are the appearance of the double and triple reassortant SIVs in the 80s and 90s [25,26], the survival of influenza viruses in front of a permanent immunologic pressure in the population, and an increasing range of susceptible host, among others.
In conclusion, it is evident that swine population in Colombia is susceptible to the infection with new influenza viruses that are circulating around the world. In spite of having a natural immune response as a result of previous exposure, because of the absence of a vaccination program, it might not be enough to face future challenges. There is a need to establish a permanent system to monitor influenza virus infection in swine populations in order to know the molecular characteristics of the strains circulating in the field, this will help design strategies to control the disease at a national level. It is also necessary to determine the outcome of influenza virus infections under tropical conditions as well as particular aspects related to the epidemiology of influenza virus in general. Not to mention the importance of the potential range of susceptible hosts available in Colombia, particularly in relation to the large number of avian species that exist in this country and which could play a role in the dissemination of new viruses to different countries. It is essential to understand host-pathogen interactions that have an impact on biological properties of influenza viruses. In summary, due to the strategic geographical position of this country in South America, it acquires importance to know what is happening in the evolution of swine influenza viruses beyond the genetic diversity of the virus not only in swine population but also in avian and human beings in order to prevent new pandemics and/or epidemics of this disease.
5. ACKNOWLEDGEMENTS
We would like to give special thanks to Angela Maria Lora for her contribution on the analysis and construction of phylogenetic trees. This research was sponsored by The Ministry of Agriculture and Rural Development of Colombia, (MADR), the Pig Producers Association of Colombia and the National University of Colombia.
REFERENCES
- Bochev, I. (2007) Porcine respiratory disease complex (PRDC): A review. I. Etiology, epidemiology, clinical forms and pathoanatomical features. Bulgarian Journal of Veterinary Medicine, 10, 131-146.
- Choi Y.K., Goyal, S.M. and Joo, H.S. (2003) Retrospective analysis of etiologic agents associated with respiratory diseases in pigs. Canadian Veterinary Journal, 44, 735-737.
- Tamura, K., Peterson, D., Peterson, N., Stecher, G., Nei, M. and Kumar, S. (2011) MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Molecular Biology and Evolution, in press. doi:10.1093/molbev/msr121
- Hansen, M.S., Pors, S.E., Jensen, H.E., Bille-Hansen, V., Bisgaard, M., Flachts, E.M. and Nielsen, E.M. (2010) An investigation of the pathology and pathogenesis associated with porcine respiratory disease complex in Denmark. Journal of Comparative Pathology, 143, e131. doi:10.1016/j.jcpa.2010.01.012
- Olsen, C.W., Brown, I., Easterday, B.C. and Van Reeth, K. (2006) Swine influenza. In: Straw, B.E., Zimmerman, J.J., D’Allaire, S. and Taylor, D.J., Eds., Diseases of Swine, 9th Edition. Iowa State University Press, Ames, 469-482.
- Van Reeth, K., Nauwynck, H. and Pensaert, M. (1996) Dual infections of feeder pigs with porcine reproductive and respiratory syndrome virus followed by porcine respiratory corona virus or swine influenza virus: A clinical and virological study. Veterinary Microbiology, 48, 325- 335. doi:10.1016/0378-1135(95)00145-X
- Kuntz-Simon, G. and Madec, F. (2009) Genetic and antigenic evolution of swine influenza virus in Europe and evaluation of their zoonotic portential. Zoonoses Public Health, 56, 310-325.
- Webster, R.G., Bean, W.J., Gorman, O.T., Chambers, T.M. and Kawaoka, Y. (1992) Evolutionand ecology of influenza A viruses. Microbiological Reviews, 56, 152-179.
- Fouchier, R.A., Munster, V., Wallensten, A., Bestebroer, T.M., Herfst, S., Smith, D., Rimmelzaann, G.F., Olsen, B. and Osterhaus, A.D. (2005) Characterization of a novel influenza A virus hemagglutinin subtype (H16) obtained from black-head gulls. Journal of Virology, 79, 2814- 2822. doi:10.1128/JVI.79.5.2814-2822.2005
- Ma, W.J., Lager, K.M., Lekcharoensuk, P., Ulery, E.S., Janke, B.H., Solorzano, A., Webby, R.J., Garcia-Sastre, A. and Richt, J.A. (2010) Viral reassortment and transmission after co-infection of pigs with classical H1N1 and triple-reassortant H3N2 swine influenza viruses. Journal of General Virology, 91, 2314-2321. doi:10.1099/vir.0.021402-0
- Hansen, B. (1977) Influenza in swine from Antioquia-Colombia (Influenza en porcinos de Antioquia-Colombia). Bol Oficina Sanit Panam, 82, 35-42.
- Mieles, A. (1995) Evidenciaserológica de influenza en porcinos de variasregiones del país. Trabajo de GradocomoBacterióloga, Universidad Colegio Mayor de Cundinamarca, Bogotá.
- Mogollón, J.D., Rincón, M.A., Preciado, P., Cepeda, M. and Ruiz, S. (2003) Serologic reactivity against swine influenza virus in intensive production herds from Colombia.: ReferenciasparaConsultorios MV (Bogotá). No. 06, pp. 15-20.
- Moscoso, C. and Neira, A. (2001) Prevalence of swine influenza virus antibodies against the H3N2 serotype in intensive procution herds in the country. Trabajo de GradocomoMédicoVeterinario, Universidad Nacional de Colombia, Bogotá.
- Pardo, S. (2001) Detection of positive reactions to swine influenza by the inhibition hemagglutination test. Thesis, Microbiólogo Agrícola y Veterinaria. Pontificia Universidad Javeriana, Bogotá.
- OIE (2005) Swine influenza. http://www.oie.int/eng/normes/mmanual/A_summry.htm
- Choi, Y., Goyal, S., Kang, S., Farnham, M. and Joo, H. (2002) Detection and subtyping of swine influenza H1N1, H1N2 and H3N2 viruses in clinical samples using two multiplex RT-PCR assays. Journal of Virological Methods, 102, 53-59 doi:10.1016/S0166-0934(01)00442-6
- Hoffman, E., Stech, J., Guan, Y., Webster, R.G. and Perez, D.R. (2001) Universal primer set for the full-length amplification of all influenza A viruses. Archives of Virology, 146, 2275-2289. doi:10.1007/s007050170002
- Saitou, N. and Nei, M. (1987) The neighbor-joining method: A new method for reconstructing phylogenetic trees. Molecular Biology and Evolution, 4, 406-425.
- Smith, G.J., Vijaykrishna, D., Bahl, J., Lycett, S.J., Worobey, M., Pybus, O.G., Ma, S.K., Cheung, C.L., Raghwani, J., et al. (2009) Origins and evolutionary genomics of the 2009 swine-origin H1N1 influenza A epidemic. Nature, 459, 1122-1125. doi:10.1038/nature08182
- Murcia, P.R., Hughes, J., Battista, P., Lloyd, L., Baillie, G.J., Ramirez-Gonzalez, R.H., Ormond, D., Oliver, K., Elton, D., Mumford, J.A., Caccamo, M., Kellam, P., Grenfell, B.T., Holmes, E.C. and Wood, J.L.N. (2012). Evolution of an Eurasian avian-like influenza virus in naïve and vaccinated pigs. PLoS Pathogens, 8, e1002730. doi:10.1371/journal.ppat.1002730
- Felsenstein, J. (1985) Confidence limits on phylogenies: An approach using the bootstrap. Evolution, 39, 783-791 doi:10.2307/2408678
- Kimura, M. (1980) A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. Journal of Molecular Evolution, 16, 111-120. doi:10.1007/BF01731581
- Moreno, A., Di Trani, L., Faccini, S., Vaccari, G., Nigrelli, D., Boniotti, M.B., Falcone, E., Boni, A., Chiapponi, Ch., Sozzi, E. and Cordioli, P. (2011) Novel H1N2 swine influenza reassortant strain in pigs derived from the pandemic H1N1/2009 virus. Veterinary Microbiology, 149, 472-477. doi:10.1016/j.vetmic.2010.12.011
- Ma, W., Vincent, A.L., Janke, B.H., Gramer, M.R. and Richt, J.A. (2009) The role of swine in the generation of novel influenza viruses. Zoonoses Public Health, 56, 326- 337. doi:10.1111/j.1863-2378.2008.01217.x
- Vincent, A.L., Ma, W., Lager, K.M., Janke, B.H. and Richt, J.A. (2008) Swine influenza viruses a North American perspective. Advances in Virus Research, 72, 127-154. doi:10.1016/S0065-3527(08)00403-X
ABREVIATIONS
SIV: Swine Influenza Virus;
PRDC: Porcine Respiratory Disease Complex;
PRRS: Porcine Reproductive and Respiratory Syndrome;
GDP: Gross Domestic Product;
BHI: Brain Hearth Infusion Broth;
CEE: Chicken Embryonated Egg

