Advances in Microbiology
Vol.04 No.15(2014), Article ID:51679,14 pages
10.4236/aim.2014.415122
Genome Annotation and Comparative Genomics of ORF Virus
A. K. M. Firoj Mahmud1, K. M. Zillur Rahman2, Shuvra Kanti Dey2, Tahsina Islam2, Ali Azam Talukder2*
1Molecular Biology Department, Umea University, Umea, Sweden
2Deptartment of Microbiology, Jahangirnagar University, Savar, Bangladesh
Email: *aat@juniv.edu, tunubj@yahoo.com
Copyright © 2014 by authors and Scientific Research Publishing Inc.
This work is licensed under the Creative Commons Attribution International License (CC BY).
http://creativecommons.org/licenses/by/4.0/



Received 29 September 2014; revised 7 November 2014; accepted 20 November 2014
ABSTRACT
ORF virus (ORFV), the etiological agent of contagious pustular dermatitis in small ruminants, be- longs to members of the genus Parapoxvirus of the Poxviridae. The genome of the ORFV is dsDNA of 139,962 bp which has about 89% coding region, 63% GC content and codes 130 proteins. There are four unique genes within the genome revealed by homology search of them two posses’ strong regulatory region and transmembrane helices. One of the ORF-039 contains signal peptide indicating the possibilities to be secretory protein coding gene. Comparative genomic analysis reveals significant differences in Bovine Papular Stomatitis Virus (BPSV) strain BV-AR02 and ORFV strain OV-SA00, and these may account for differences in host range. Interspecies sequence variability is observed in all functional classes of genes but is the highest in putative virulence/host range genes. Notably, ORFV contains genes which are homologous of Vaccinia virus. Phylogenetic analysis reveals that although divergent, ORFV virus is distinct from other known mammalian cowpox virus. An improved understanding of Parapoxvirus (PPV) biology will permit the engineering of novel vaccine viruses and expression vectors with enhanced efficacy and greater versatility. The novel vaccine will have a significant role in the economy of a country through the control of disease in an economically important and small ruminant caused by ORFV.
Keywords:
ORF Virus, Contagious Pustular Dermatitis, Comparative Genomics, GLIMMER, ACT, Uniue Genes, Novel Vaccine

1. Introduction
Genome annotation is the analysis of genome sequence of a particular species which includes all the possible analysis of DNA sequence that can be done by computational means. Raw DNA sequence produced by the genome-sequencing projects is taken for the analysis. Analysis and interpretation is carried out which is necessary to extract its biological significance and organize the information into the context of our understanding of biological processes. By genome annotation, an individual gene and its protein (or RNA) product can be predicted by in depth analysis of several gene features, such as ORFs length, % (G + C) content, promoter, polyA tail, homology, etc. The focal point of each such record is the validity of a ORF as a potential functional gene. Moreover, analysis may also include a brief description of the evidence for the assigned or proposed function. Several genome annotation tools are freely available for further analysis of genome sequence. Users can use several individual tools for each task or can use some integrated tools that do many task simultaneously. GLIMMER [1] , GeneMark [2] , ORF finder [3] and FrameD [4] are well-used tools for predicting Open Reading Frame (ORF). BLAST is the most used homology tool for homology prediction but there are many other options are available, such as MPsrch, PSI-BLAST and WU-BLAST [5] . Artemis, Apollo, JBROWSE, etc. are used for Genome browsers for curation and further annotation [6] . There are also many Genome annotation pipelines available, such as PASA and MAKER, in which several analysis tools are integrated [6] . Users are required to select the tools or pipeline according to their needs that serve best for their purpose
Contagious pustular dermatitis; ORF is a common epitheliotropic viral disease of sheep, goats and wild ruminants and is characterized by the formation of papules, nodules, or vesicles caused by the ORF virus [7] . Humans always posses a high risk due to zoonotic characteristics of this disease and often humans can contract this disorder through direct contact with infected animals by the fomites that carry the ORF virus [8] . Purulent-appearing papule is the major symbol causes locally and generally no systemic symptoms is obseved [3] . Normally it infects the finger, hand, arm, face and even the penis [6] - [9] . ORF virus (ORFV) is an oval and enveloped virus containing dsDNA genome within the genus Parapoxvirus, family Poxviridae [6] . The genus also includes pseudocowpox virus (PCPV) and bovine papular stomatitis virus (BPSV) in cattle and parapoxvirus (PPV) of red deer in New Zealand [10] .
Mechanisms involved in ORFV virulence are not well studied [10] . Several putative virulence genes have been identified, such as the vascular endothelial growth factor homologue, an interleukin (IL-10) homologue, a double-stranded RNA-binding protein, a factor inhibiting the cytokines granulocyte-macrophage colony-stimu- lating factor and IL-2 [11] - [16] . The whole genome of ORF virus strain OV-IA82 has been sequenced by the Plum Island Animal Disease Center, USA in 2004 [17] . The genome has high 63% GC content with 89% coding region which codes 130 proteins and length is 139,962 bp [17] . The complete genomic sequence available for OV-IA82 enables to deduce the different complex area including other molecular aspects of pathogenesis of this virus in details [18] . In this study, we tried to identify unique genes in ORFV genome and characterize them by Insilco process.
2. Materials and Methods
There are certain kinds of genome annotation tools available for analysis the whole genome, some of which Artemis and Artemis comparison Tools (ACT) has been used extensively for Open Reading Frame (ORF) visualization, editing, determine GC plot as well as retrieve desired nucleotide length within the genome for further analysis [19] . The GLIMMER has been chosen for predicting the genes or ORF in the genome of ORFV based up on its accuracy and flexibility of changing stop and start codons according to the requirement [20] [21] . Potential protein coding ORFs were identified by the following criteria: ORF size larger than 60 amino acid (aa), presence of potential transcriptional start and stop sites, a high GLIMMER score and homology to other known Parapoxvirus or cellular ORFs [21] . To find the similarity and homology, sequences of the GLIMMER pedicted protein, coding ORF were compared with the protein databases such as SwissProt, Trembl, UniProt, etc. In this study, among the tools which are available to find the similarity compared to protein databases, the Blast module Blastall, which supports all five Blast programs (blastp, blastn, blastx, tblastn and tblastx) has been chosen for finding the unique genes. Promoter, poly A signal and CpG island were analyzed for each unique genes in order to determine their potentiality as genes. For promoter prediction upstream ~350 bases were subjected within Neural Network Promoter Prediction Tool developed by University of Berkly, USA with a cut off value 0.8 [22] . PolyA signals were determined by polyADQ which is a poly (A) signal search engine developed by Cold Spring Harbor Laboratory, USA [22] [23] . The software CpGIE was downloaded via the website: [24] . The following cutoff values were used to determine the CpG island in a given genomic sequence: ≥200 nt, G + C content 50%, and an observed: expected CpG ratio 0.6 [25] [26] . To determine the Trans Membrane (TM) domain and peptide signal of the unique genes TMHMM and SignalP 3.0 software was used respectively [27] . The Artemis and Artemis Comparison Tool (ACT), written by Kim Rutherford, Sanger Institute, UK [19] were used for genome analysis and the pair wise comparisons between OV- IA82, VACV and BPSV genomes. For phylogenetics analysis various B2L genes of ORFV were selected on the basis of host specificity such as human, sheep and goats [28] [29] . In order to determine distant relationship BPSV and psedocowpox B2L gene also selected. All the genes sequences were downloaded from gene bank and stored for phylogenetic tree construction as listed in Table 1. The nucleotide sequences of diverse ORf viruses and others were aligned using the Bioedit program and Mega 5 software [22] . One thousand bootstrap replicates were subjected to nucleotide sequence distance and neighbor-joining methods, and the consensus phylogenetic tree was drawn.
3. Results
In this study we tried to identify unique genes in ORFV on the basis of the availability of complete genome sequence of OV-IA82 by Insilco process in order to determine the novel vaccine strain. Recent advances in bioinformatics enable us to further analysis of the ORFV genome and the results are given below.
3.1. Coding Potential and Functional Analysis
GLIMMER predicted 130 potential open reading frames which supported previous studies. Like other poxviruses, ORFV genomes contain a large central coding region (ORF 12) bounded by two identical inverted terminal repeat (ITR) regions, (ORF 26 and 48). Homology search by blast revealed four ORFs 039, 116, 124 and 125 as unique genes as no similarity has been found with others rather than with ORFV. Only one ORF (039), previously described for ORFV strains NZ2 and NZ7, is completely located within the ITRs of the genome.
3.2. Regulatory Regions of the Unique Genes
Among the four unique genes, all posses promoter above cut off value 0.9 in the nearer upstream region which is
Table 1. List of genes for phylogenetic analysis with origin and gene bank accession number.
an indication of their translation probabilities. On the contrary, ORF 124 and 125 lack polyA signal as well as CpG island. Besides ORF 039 and 116 have strong promoter as well as CpG island 51 and 52 bp long respectively. ORF 039 has polyA signal in their 6 bp downstream region but ORF 116 lacks polyA signal.
3.3. Transmembrane Domains and Signal Peptides of the Unique Genes
All four unique genes were further evaluated to determine the TM helics. ORF 039 contains TM helices in between 2 - 20 amino acid (aa) in their C-terminal region in which inside helices is in 21 - 51 aa. On the other hand, ORF 116 contains TM helices in between 26 - 48 aa, in which 1 - 28 aa are inside the membrane and 49 - 52 aa are outside the membrane (Figure 1).
Besides, only ORF 039 was found having peptide signal at 31 and 32 aa with a probability 0.82. For confirmation both the PAM Matrix (PM) algorithm and Hidden Markov Model (HMM) were used (Figure 2). Other three ORFs lack signal peptides, by which we can assume that they are not secretory proteins.
3.4. Comparative Genomics
Three genomes of Orf virus (ORFV) strain OV-IA82, bovine papular stomatitis virus (BPSV) strain BV-AR02 and vacccinia virus (VACV) strain VACV-A4L were used for comparative genomics in order to find the unique genes (Figure 3).
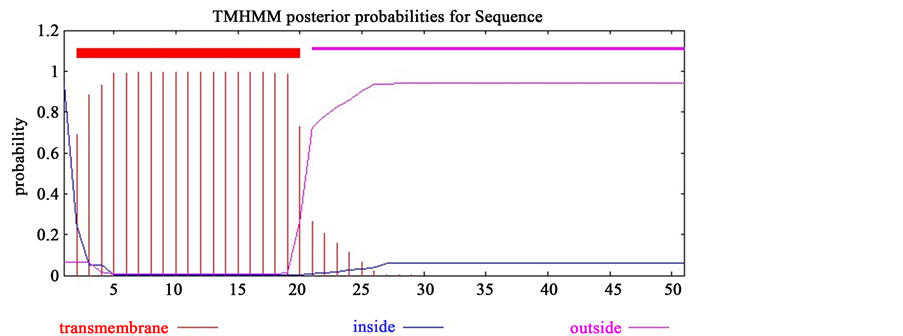
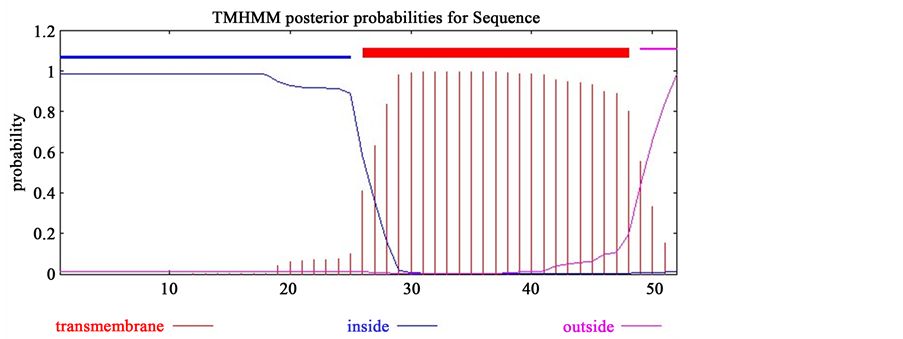
Figure 1. Determination of TM helics. Here, (a) represents the output result of TMHMM for ORF 039 and (b) represents the TMHMM result for ORF 116 in which transmembrane helices are shown in bold red line and inside and outside membrane helices are shown in blue and purple line, respectively.
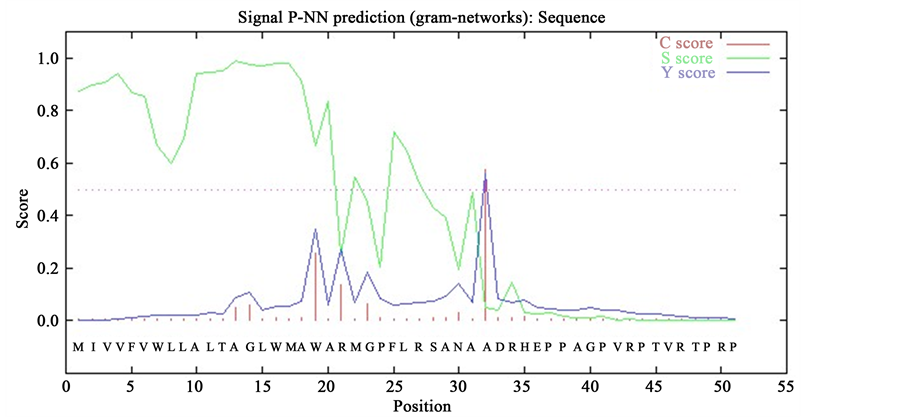
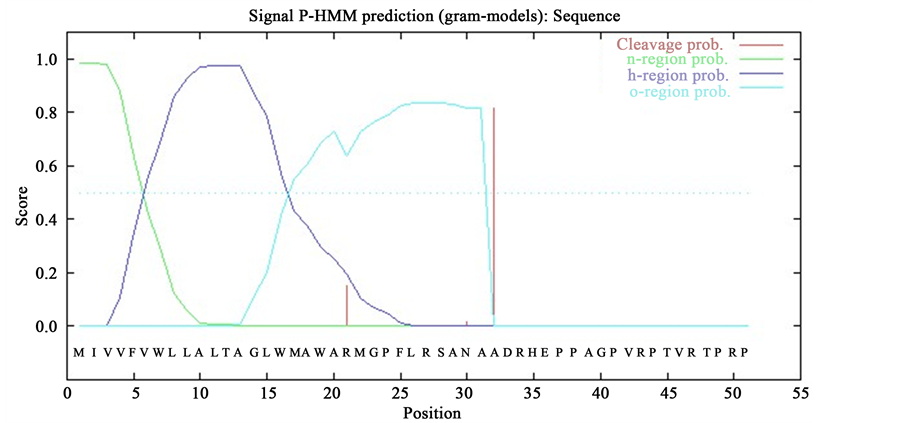
Figure 2. Signal peptide determination of ORF 039 by SignalP 3.0. Here, (a) and (b) represent the SignalP PAM matrix (PM) prediction and Hidden Markov Model (HMM) prediction, respectively.
3.5. Comparison of BPSV with ORFV
At the genomic level, BPSV and ORFV genomes share 67% to 75% nucleotide identity and contain 127 genes with the same relative order and orientation. Among them 15 genes are unique to PPVs. BPSV and ORFV contain 15 and 16 ORFs, respectively, that share no significant homology to known proteins while search for homology blast and are primarily located at the right end of the genome. Fourteen ORFs (001, 005, 012, 013, 024, 073, 113, 115, 116, 119, 120, 121, 124 and 125) was observed in both BPSV and ORFV. Besides, four ORFs (039, 116, 124 and 125) are present only in ORFV with 29% to 64% amino acid identities and one (ORF 133) is unique to BPSV. There are few ORFs which are distantly related with amino acid identity approximately up to 65%. Among thiese 30 distantly related ORF 10 are found as unique to 12 are unique to PPVs (ORFs 002, 005, 012, 013, 068, 113, 115, 116, 119, 120, 121 and 124). There are two ORFs 58, 57 that encode ankyrin repeat- containing proteins (ARPs) are observed only 50% identical between BPSV and ORFV. However, BPSV contains two (ORFs 003 and 004) additional ARPs in the left terminal genomic region which are not present in ORFV.
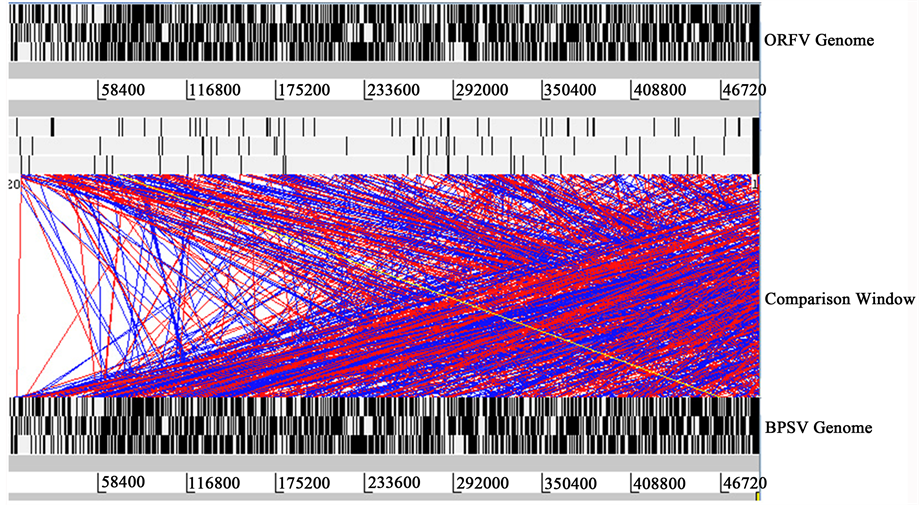
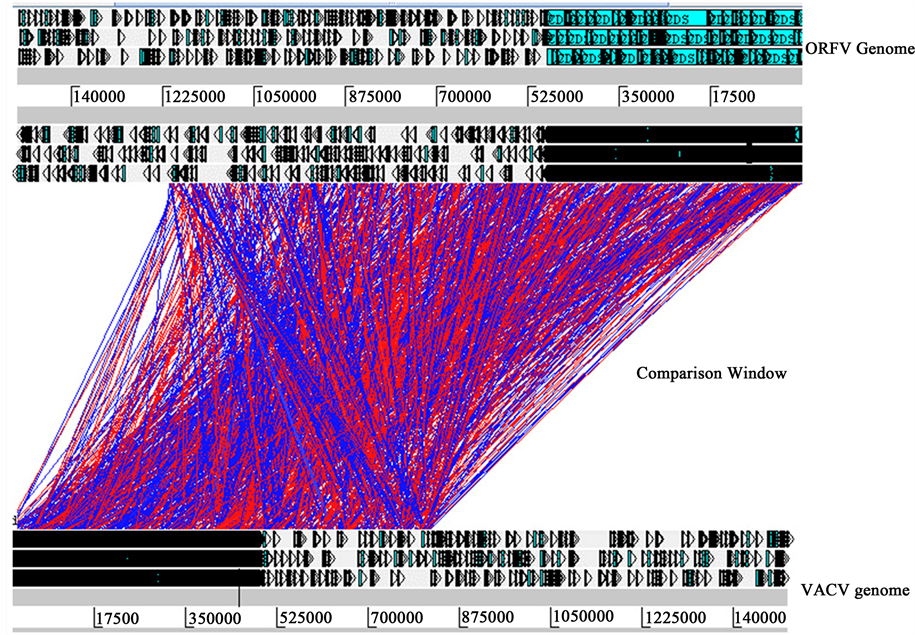
Figure 3. Comparative genomics of ORFV with other two closely related viruses BPSV and VACV. The red and blue bars indicate regions of similarity with red bars indicating corresponding regions that are oriented similarly and blue bars indicating regions oriented in opposite directions. Comparison of whole genome by ACT. The top view shows the subject sequence ORFV and the bottom view shows the query sequence: (a) Comparison of BPSV with ORFV; (b) Comparison of VACV with ORFV.
3.6. Comparison of VACV with ORFV
While compare with VACV genome with ORFV geneome, 13 ORFs (ORFs 061, 080, 088, 103, 109, 110, 112, 126, 128, 009, 016, 122 and 129) of ORFV show homology with VACV. Among these 13 homologue 10 genes have known function. Interestingly, ORFV lack homologues to VACV D9R. VACV D9R gene contains a mutT motif which is also present in VACV D10R that encodes a viral transcription protein and DNA ligase encoding gene VACV A50R. That suggest some other gene of ORFV which are not yet functionally characterized might have done the works for these genes. ORF 80 has homology with both VACV and BPSV. Besides, ORF 088, 109, 110 and 138 is orthologous with VACV.
3.7. Phylogenetic Analysis
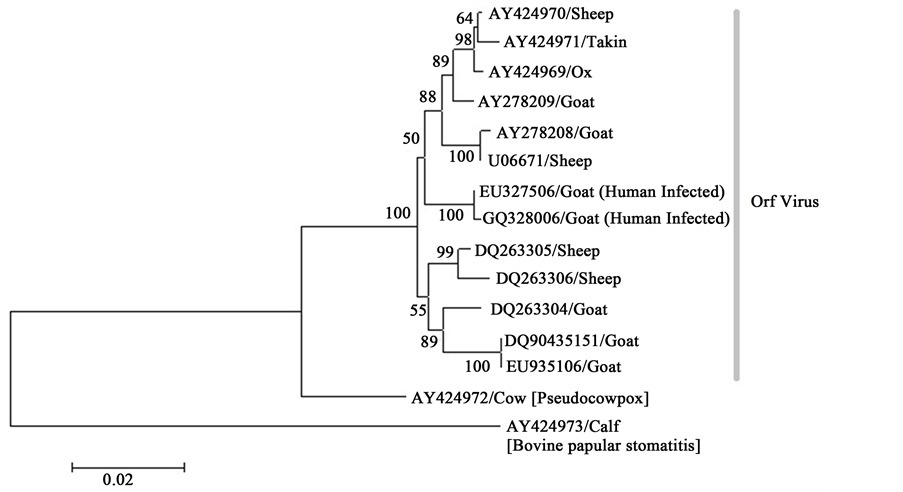
In phylogenetic analysis based on the complete B2L gene, the ORF/09/Korea strain was closer to the Taiping isolate from Taiwan. ORF-ca1 and NZ-2-1 are closely related despite of different origin. Other ORF virus of sheep and Goat origin are less similar. Pseudocowpox and BPSV are distantly related with ORF virus (Figure 4).
4. Discussion
ORF virus shares specific genomic features with other poxviruses as VACV and BPSV, in terms of genome organization and gene content. Comparative genome sequences with other two closely related viruses VACV and BPSV here provide a comparative view of Parapoxvirus (PPV) genomics and basic knowledge of viral functions associated with virus replication and manipulation of cellular responses. Based on comparative genomic analysis, the genomes of BPSV and ORFV differ significantly which may be responsible for differences in host range. Modern genome analysis tools, such as Artemis Comparison Tools (ACT), promptus to understand the PPV biology which will permit the engineering of novel vaccine viruses and expression vectors with enhanced efficacy and greater versatility [30] . Nevertheless, we have identified four unique genes with potential importance as virulence factors and thus they could be vaccine candidates in the future. These genes should not correspond to horizontal gene transfer and their characteristic features may be consequences of the specific evolutionary cycle that shapes the ORFV gene repertoires in the context of their parasitic lifestyles [31] . ORF 039 with signal peptides and transmembrane domains may be directly toxic or confer association with the host. Therefore, further focus can be placed on subset of this for functional analysis. The function, subcellular location, average of
Figure 4. Phylogenetic analysis of different parapoxviruses. Phylogenetic tree is constructed based on viral B2L gene. Bootstrap values (derived from 1000 replicate neighbor-joining (NJ) trees estimated under the ML substitution model) are shown for key nodes > 50%.
hydrophobicity and protein regions that share a significant degree of sequence similarity with known protein family can be detected by using computational approach. Primarily 2D-PAGE might be used for membrane protein analysis for ORF 039 [32] [33] . Besides, Combination of nano liquid chromatography (NanoLC) and mass spectrometry (MS) can be used to detect the transmembrane domian of ORF 039 [33] . ORF 039 can be used for Insilco protein homology modeling. Identification of certain protein ligand of this protein may open a new door for the development of antibody for ORFV and also for potential drug target. Insilco drug target analysis may be carried out to find out the potentiality of the genes for further analysis.
Similar Insilco studies by the following different approach have been carried out to identify potential genes for therapeutic targets [34] [35] . Unique genes or proteins that are involved in a certain pathway or having certain characteristics like outer membrane protein, presence of unique protein ligand family, etc. are always an ideal candidate for drug target [35] - [37] . Insilco identification of target genes and prediction of drug candidate is a well-established methodology in drug discovery. Screening out the target genes and their corresponding drug by Insilco approach makes the drug discovery procedure more robust, quick and economically feasible [36] .
The development of novel vaccine strain will control contagious pustular dermatitis in a small ruminant, potential source of leather, meat and milk which have a significant role in the economy of a country [13] . Therefore, control of such ORF related disease will have a significant role in economic development of a country. This study will be very much useful for further study of the evolution of the ORFV that may provide an encouragement for the development of new diagnostic tools and medicines.
Acknowledgements
The authors wish to thank Ministry of Education (17/10, M-15/2007/226 and HEQEP, CP-W1-3413), Bangladesh for partially funding this research work.
References
- Delcher, A.L., Harmon, D., Kasif, S., White, O. and Salzberg, S.L. (1999) Improved Microbial Gene Identification with GLIMMER. Nucleic Acids Research, 27, 4636-4641. http://dx.doi.org/10.1093/nar/27.23.4636
- Besemer, J., Lomsadze, A. and Borodovsky, M. (2001) GeneMarkS: A Self-Training Method for Prediction of Gene Starts in Microbial Genomes. Implications for Finding Sequence Motifs in Regulatory Regions. Nucleic Acids Research, 29, 2607-2618. http://dx.doi.org/10.1093/nar/29.12.2607
- Stothard, P. (2000) The Sequence Manipulation Suite: JavaScript Programs for Analyzing and Formatting Protein and DNA Sequences. Biotechniques, 28, 1102-1104.
- Schiex, T., Gouzy, J., Moisan, A. and Oliveira, Y. (2003) Frame D: A Flexible Program for Quality Check and Gene Prediction in Prokaryotic Genomes and Noisy Matured Eukaryotic Sequences. Nucleic Acids Research, 31, 3738-3741. http://dx.doi.org/10.1093/nar/gkg610
- Neumann, R.S., Kumar, S. and Tabrizi, K.S. (2014) BLAST Output Visualization in the New Sequencing Era. Brief Bioinformatics, 15, 484-503. http://dx.doi.org/10.1093/bib/bbt009
- Yandell, M. and Ence, D. (2012) A Beginner’s Guide to Eukaryotic Genome Annotation. Nature Reviews Genetics, 13, 329-342. http://dx.doi.org/10.1038/nrg3174
- Robinson, A.J. and Balassu, T.C. (1981) Contagious Pustular Dermatitis (Orf). Veterinary Bulletin, 51, 771-782.
- Haig, D.M. and Mercer, A.A. (1998) Ovine Diseases. Orf. Veterinary Research, 29, 311-326.
- Fleming, S.B., Blok, J., Fraser, K.M., Mercer, A.A. and Robinson, A.J. (1993) Conservation of Gene Structure and Arrangement between Vaccinia Virus and Orf Virus. Virology, 195, 175-184. http://dx.doi.org/10.1006/viro.1993.1358
- Rziha, H.J., Henkel, M., Cottone, R., Meyer, M., Dehio, C. and Büttner, M. (1999) Parapoxviruses, Potential Alternative Vectors for Directing the Immune Response in Permissive and Non-Permissive Hosts. Journal of Biotechnology, 73, 235-242. http://dx.doi.org/10.1016/S0168-1656(99)00141-8
- Chi, X., Zeng, X., Hao, W., Li, M., Li, W., Huang, X.H., et al. (2013) Heterogeneity among Orf Virus Isolates from Goats in Fujian Province, Southern China. PLoS ONE, 8, 1-12. http://dx.doi.org/10.1371/journal.pone.0066958
- Kumar, N., Wadhwa, A., Chaubey, K.K., Singh, S.V., Gupta, S., Sharma, S., et al. (2014) Isolation and Phylogenetic Analysis of an Orf Virus from Sheep in Makhdoom, India. Virus Genes, 48, 312-319. http://dx.doi.org/10.1007/s11262-013-1025-9
- Robinson, A.J. and Lyttle, D.J. (1992) Parapoxviruses. Their Biology and Potential as Recombinant Vaccines. Recom- binant Poxviruses, 28, 285-327.
- Lyttle, D.J., Fraser, K.M., Fleming, S.B., Mercer, A.A. and Robinson, A.J. (1994) Homologs of Vascular Endothelial Growth Factor Are Encoded by the Poxvirus Orf Virus. Journal of Virology, 68, 84-92.
- Savory, L.J., Stacker, S.A., Fleming, S.B., Niven, B.E and Mercer, A.A. (2000) Viral Vascular Endothelial Growth Factor Plays a Critical Role in Orf Virus Infection. Journal of Virology, 74, 699-706. http://dx.doi.org/10.1128/JVI.74.22.10699-10706.2000
- Fleming, S.B., McCaughan, C.A., Andrews, A.E., Nash, A.D. and Mercer, A.A. (1997) A Homolog of Interleukin-10 Is Encoded by the Poxvirus Orf Virus. Journal of Virology, 71, 4857-4861.
- McInnes, C.J., Wood, A.R. and Mercer, A.A. (1998) Orf Virus Encodes a Homolog of the Vaccinia Virus Interferon- Resistance Gene E3L. Virus Genes, 17, 107-115. http://dx.doi.org/10.1023/A:1026431704679
- Delhon, G., Tulman, E.R., Afonso, C.L., Lu, Z., de la Concha-Bermejillo, A., Lehmkuhl, H.D., et al. (2004) Genomes of the Parapoxviruses Orf Virus and Bovine Papular Stomatitis Virus. Journal of Virology, 78, 168-177. http://dx.doi.org/10.1128/JVI.78.1.168-177.2004
- Rutherford, K., Parkhill, J., Crook, J., Horsnell, T., Rice, P., Rajandream, M.A. and Barrell, B. (2000) Artemis, Sequence Visualization and Annotation. Bioinformatics, 16, 944-945. http://dx.doi.org/10.1093/bioinformatics/16.10.944
- DeGrado, W.F., Gratkowski, H. and Lear, J.D. (2003) How Do Helix-Helix Interactions Help Determine the Folds of Membrane Proteins? Perspectives from the Study of Homo-Oligomeric Helical Bundles. Protein Science, 12, 647-665. http://dx.doi.org/10.1110/ps.0236503
- Delcher, A.L., Harmon, D., Kasif, S., White, O. and Salzberg, S.L. (1998) Improved Microbial Gene Identification with GLIMMER. Nucleic Acids Research, 27, 36-41.
- Durbin, R., Eddy, S.R., Krogh, A. and Mitchison, G.J. (1998) Biological Sequence Analysis, Probabilistic Models of Proteins and Nucleic Acids. Cambridge University Press, Cambridge.
- Gershenzon, N.I. and Ioshikhes, I.P. (2006) Synergy of Human Pol II Core Promoter Elements Revealed by Statistical Sequence Analysis. Bioinformatics, 21, 295-300.
- Fatemi, M., Pao, M.M., Jeong, S., Gal-Yam, E.N., Egger, G., Weisenberger, D.J. and Jones, P.A. (2005) Footprinting of Mammalian Promoters, Use of a CpG DNA Methyltransferase Revealing Nucleosome Positions at a Single Molecule Level. Nucleic Acids Research, 33, 176-188. http://dx.doi.org/10.1093/nar/gni180
- Guigó, R., Knudsen, S., Drake, N. and Smith, T. (1992) Prediction of Gene Structure. Journal of Molecular Biology, 226, 141-157. http://dx.doi.org/10.1016/0022-2836(92)90130-C
- Himmelreich, R., Hilbert, H., Plagens, H., Pirkl, E., Li, B.C. and Herrmann, R. (1996) Complete Sequence Analysis of the Genome of the Bacterium Mycoplasma Pneumoniae. Nucleic Acids Research, 24, 4420-4449. http://dx.doi.org/10.1093/nar/24.22.4420
- Nakai, K., Kanehisa, M. and Doolittle, R.F. (1991) Signal Peptide and Cytokine Receptor, Function and Regulation. Proteins Structure Function Genetics, 11, 95-110. http://dx.doi.org/10.1002/prot.340110203
- Zhang, K., Liu, Y., Kong, H., Shang, Y. and Liu, X. (2014) Comparison and Phylogenetic Analysis Based on the B2L Gene of Orf Virus from Goats and Sheep in China during 2009-2011. Archives of Virology, 159, 1475-1479. http://dx.doi.org/10.1007/s00705-013-1946-6
- Amann, R., Rohde, J., Wulle, U., Conlee, D., Raue, R., Martinon, O. and Rziha, H.J. (2013) A New Rabies Vaccine Based on a Recombinant ORF Virus (Parapoxvirus) Expressing the Rabies Virus Glycoprotein. Journal of Virology, 87, 1618-1630. http://dx.doi.org/10.1128/JVI.02470-12
- Zhao, K., He, W., Gao, W., Lu, H.J., Han, T., Li, J., et al. (2011) Orf Virus DNA Vaccines Expressing ORFV 011 and ORFV 059 Chimeric Protein Enhances Immunogenicity. Journal of Virology, 8, 562. http://dx.doi.org/10.1186/1743-422X-8-562
- Ogata1, H. and Claverie, J.M. (2007) Unique Genes in Giant Viruses: Regular Substitution Pattern and Anomalously Short Size. Genome Research, 17, 1353-1361. http://dx.doi.org/10.1101/gr.6358607
- Renesto, P., Abergel, C., Decloquement, P., Moinier, D., Azza, S., Ogata, H., Fourque, P., Gorvel, J.P. and Claverie, J.M. (2006) Mimivirus Giant Particles Incorporate a Large Fraction of Anonymous and Unique Gene Products. Journal of Virology, 80, 11678-11685. http://dx.doi.org/10.1128/JVI.00940-06
- Dung, N.T., Chi, D.H., Thao, L.T., Dung, T.K., Nhi, N.B. and Chi, P.V. (2013) Identification and Characterization of Membrane Proteins from Mouse Brain Tissue. Journal of Proteomics Bioinformatics, 6, 142-147.
- Chakraborty, S. (2014) In Silico Analysis Identifies Genes Common between Five Primarygastrointestinal Cancer Sites with Potential Clinical Applications. Annals of Gastroenterology, 27, 231-236.
- Dutta, A., Singh, S.K., Ghosh, P., Mukherjee, R., Mitter, S. and Bandyopadhyay, D. (2006) In Silico Identification of Potential Therapeutic Targets in the Human Pathogen. In Silico Biology, 6, 43-47.
- Wang, T.T., Mendoza, L.T., Laperriere, D., Libby, E., MacLeod, N.B., Nagai, Y., Bourdeau, V., Konstorum, A., Lallemant, B., Zhang, R., et al. (2005) Large-Scale in Silico and Microarray-Based Identification of Direct 1,25- Dihydroxyvitamin D3 Target Genes. Molecular Endocrinology, 19, 2685-2695. http://dx.doi.org/10.1210/me.2005-0106
- Das, S. and Chaudhuri, K. (2003) Identification of a Unique IAHP (IcmF Associated Homologous Proteins) Cluster in Vibrio Cholerae and Other Proteobacteria through in Silico Analysis. In Silico Biology, 3, 287-300.
Supplementary
Table S1. Open reading frame predicted by GLIMMER.
NOTES
*Corresponding author.