Stem Cell Discovery
Vol.3 No.1(2013), Article ID:26956,8 pages DOI:10.4236/scd.2013.31010
Reprogramming mouse ear mesenchymal stem cells (EMSC) expressing the Dlk1-Dio3 imprinted gene cluster
![]()
1Stem Cell Biology Laboratory, Pennington Biomedical Research Center, Baton Rouge, USA
2NuPotential, Inc., Baton Rouge, USA; *Corresponding Author: Kenneth.Eilertsen@pbrc.edu
3Institute of Animal Reproduction and Food Research of Polish Academy of Sciences, Olsztyn, Poland
4Department of Anatomy & Cell Biology, Carver College of Medicine, University of Iowa, Iowa City, USA
Received 22 October 2012; revised 23 November 2012; accepted 25 December 2012
Keywords: iPSCs; Reprogramming; Pluripotent; Stem Cell
ABSTRACT
The identification of a single, early marker for full developmental potential of induced pluripotent stem (iPS) cells has proven elusive. Recently, however, activation of the imprinted gene cluster, Dlk1-Dio3 has emerged as a viable candidate in the mouse. To explore the relationship between Dlk1-Dio3 expression and developmental potential more fully, we used murine ear mesenchymal stem cells (mEMSC) for iPS cell induction. Mouse EMSC are easily obtained and share functional characteristics with embryonic stem (ES) cells and therefore, may be a reliable non-embryonic source for iPS cell production. We report that mEMSC express high levels of Gtl2, a maternally expressed gene within the Dlk1-Dio3 imprinted cluster. Moreover, mEMSC produce Gtl2 expressing (Gtl2on) iPSC clones that share functional characteristics with ES cell clones. The production of Gtl2on iPS cell clones from mEMSC provides a new model with which to investigate the regulation of Dlk1-Dio3 cluster activity during direct cell reprogramming.
1. INTRODUCTION
Induced pluripotent stem (iPS) cells have been produced from murine and human somatic cells by overexpression of defined factors [1-3]. These iPS cells share certain structural and functional characteristics with embryonic stem (ES) cells including similar epigenetic patterns [4,5], differentiation potential [2,6] and full-term development, as demonstrated by tetraploid complementation [7-9]. Taken together, these data suggest that iPS cells, like ES cells, hold tremendous clinical promise for the treatment of disease, repair of damaged tissues and for drug discovery [10].
Despite these similarities, there is growing evidence that iPS cells also differ significantly from ES cells with respect to mRNA and microRNA (miRNA) expression patterns [11-13], chimera development and postnatal viability [3,14,15] and immunogenicity [16]. These observations indicate that direct reprogramming of somatic cells may induce abnormalities in iPS cells and that these abnormalities may be incompatible with therapeutic applications (for review see [17]). Therefore, in order to realize the full clinical potential of iPS cells, it is imperative to more fully understand the mechanisms that drive somatic cell reprogramming with a view toward improving iPS cell production technology.
Ear mesenchymal stem cells (EMSC) were first isolated as a primary culture from the external ear tissue of adult mice and shown to possess the capacity for selfrenewal and differentiation into chondrocytes, osteocytes, adipocytes [18] and myocytes [19]. EMSC also serve as a simple model for adipogenesis [20] with potentially broader applications for regenerative medicine [21]. Human mesenchymal stem cells (MSCs) are also capable of undergoing adipogenesis; a process that is in part, regulated by the delta-like 1 (Dlk-1) gene [22]. Dlk-1, also called preadipocyte factor 1 (Pref-1) is a paternally expressed imprinted gene that plays a key role in maintaining preadipocytes in their undifferentiated state both in vitro [23-25] and in vivo [26-28].
The Dlk1-Dio3 imprinted gene cluster is located on mouse chromosome 12qF1 and this region also plays an important role in establishing cell pluripotency and in regulating preand post-natal development [29]. Indeed, the activation state of this single gene cluster was recently shown to distinguish mouse iPS cells from ES cells and to predict the developmental potential of iPS cell clones [29,30]. By comparing genetically matched iPS and ES cells, Stadtfeld and colleagues revealed two groups of iPS cell clones, those with active Dkl1-Dio3 expression (Gtl2on) and those in which the Dkl1-Dio3 cluster is aberrantly silenced (Gtl2off). ES cells were found to be Gtl2on while most iPS cell clones were Gtl2off. The lack of Dlk1-Dio3 expression in iPS cell clones correlated with poor chimera formation and failed post-natal development. By contrast, Gtl2on iPS cell clones, like ES cell clones, produced high-grade chimeras and viable mice. Interestingly, the few Gtl2on iPS cell clones observed were all derived from fibroblast cultures expressing high levels of Gtl2 while haematopoietic cells, which express low levels of Gtl2, consistently produced Gtl2off iPS cell clones [30]. This pivotal work raises the interesting possibility that the endogenous level of Gtl2 expression in somatic cells may serve as a predictor of their developmental potential when used to generate iPS cell clones.
MSCs isolated from adult mouse bone marrow are capable of efficiently producing high-quality iPS cell clones that closely resemble ES cells [31]. However, the expression status of the Dlk1-Dio3 gene cluster is unknown in these MSCs and in the resultant MSC-derived iPS cell clones. Thus, it is currently unclear if Gtl2on iPS cell clones can be produced from MSCs. To investigate the status of Gtl2 expression in successful MSC-based iPS cell production, we used murine EMSC to produce iPS cell clones and assessed their level of Dlk1-Dio3 gene expression and developmental potential.
2. MATERIALS AND METHODS
2.1. Cell Culture
Mouse ear mesenchymal stem cells (mEMSC) were isolated using previously published procedures [32]. The cells were cultured in DMEM/Ham’s F-12 medium (Invitrogen), supplemented with 100 U/ml penicillin, 100 μg/ml streptomycin and 15% FBS (Invitrogen).
Mouse ES cells and induced pluripotent stem (iPS) cells were maintained in complete ES cell media with 15% FBS and LIF (Millipore) either on mitomycin C (Sigma-Aldrich)-treated MEFs or 0.1% gelatin-coated plates. The established miPS cell lines were treated with 0.05% trypsin/0.53 mM EDTA (Life Technologies) for 5 min at room temperature and split at 1:6 to 1:10.
2.2. Lentiviral Transduction and Derivation of Mouse iPS Cell Clones
The reprogramming of mEMSCs was induced with a single constitutive polycistronic lentivirus (EF1α-STEM CCA, Millipore), which encodes mouse cDNAs for Oct4, Klf4, Sox2 and c-Myc. EMSCs (passage 0) were seeded at 1 × 105 cells in one well of a six-well plate one day before transduction. The cells were infected with the lentivirus in the presence of 5 μg/ml polybrene (Millipore) overnight. Two days after the infection, cells were transferred onto an inactivated MEF feeder layer and then cultured with complete ES cell medium. ES media was changed every other day. Cell morphology and iPS colony formation were monitored daily until the colonies could be picked and passaged after trypsin treatment 16 days post-transduction.
2.3. Karyotyping and Bisulfite Genomic Sequencing
G-banding chromosomal analysis was performed on one mouse iPS (miPS) cell line (Cell Line Genetics, Madison, WI). Genomic DNA was isolated from EMSC, mESCs, and two miPS cell lines and purified using the DNeasy Tissue Kit (Qiagen). Bisulfite treatment of genomic DNA was carried out with the EZ DNA Methylation Kit (Zymo Research). The promoter regions of mouse Oct4 and Nanog genes were amplified by PCR using the primer sequences as follows. For Oct4, the forward primer was 5’-gattttgaaggttgaaaatgaagg and the reverse primer was 5’-aaatctaaaaccaaatatccaacc. For Nanog, the forward primer was 5’-gattttgtaggtgggattaactgtgaattt and the reverse primer was 5’-accaaaaaaacccacactcatatcaatata. PCR products were subcloned into the pCRII-TOPO cloning vector (Invitrogen) and sequenced on an Applied Biosystems 377 DNA sequencer.
2.4. Alkaline Phosphatase Staining and Immunofluorescence
Alkaline phosphatase (AP) activity in miPS cells was determined using the Alkaline Phosphatase Staining Kit II (Stemgent). For immunofluorescent (IF) staining, cells were fixed in 4% paraformaldehyde for 15 min, rinsed in PBS, and blocked with 10% normal goat serum in PBS containing 0.1% Tween 20 for 60 min at room temperature. After incubation with primary antibodies against Oct4 (1:200, Abcam), Sox2 (1:200, Abcam), Nanog (1: 200, Abcam), SSEA-1 (1:100, Abcam), Map2 (1:200, Abcam), alpha smooth muscle actin (SMA, 1:100, Abcam), Lin28 (1:100, Cell Signaling), or heaptocyte nuclear factor-3 beta (HNF 3β, 1:200, Millipore) overnight at 4˚C, cells were washed and incubated with fluorescence-labeled secondary antibodies (Alexa Fluor 488 or 594, Molecular Probe). Cells were counter-stained by DAPI (4’,6- diamidino-2-phenyl-indole, Vector lab) and analyzed on a fluorescence microscope with a Axiocam camera (Zeiss).
2.5. RNA Extraction and Quantitative RT-PCR
Total RNA was isolated using TRIzol reagent (Invitrogen) and purified with the RNeasy Mini Kit (Qiagen). RNA (200 ng) was reverse transcribed using the High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems). Real-time PCR analysis was done in an ABI Prism 7900 sequence detection system (Applied Biosystems) using the following Taqman primers/probes: GAPDH (Mm99999915_g1), Oct4 (Mm00658129_gH), Sox2 (Mm00488369_s1), Nanog (Mm01617762_g1), Lin28 (Mm00524077_m1), Klf4 (Mm00516104_m1), c-Myc (Mm00487804_m1), Gtl2 (Mm00522599_m1), Rian (Mm01325839_g1), Dlk1 (Mm00494477_m1), HNF 3β (Mm01976556_s1), Map2 (Mm00485230_m1), Desmin (Mm00802455_m1), SMA (Mm00725412_s1). Reactions were performed in duplicate for each sample and repeated twice, and each run included a standard curve. Gene expression was normalized to GAPDH in all samples. To determine the expression of viral transgenes in reprogrammed cells, the Viral Gene Detection qPCR Kit (Millipore) was used in a Roche LightCycler 480 system.
2.6. In Vitro Differentiation of Mouse iPS Cells
Mouse iPS cells were dissociated with 0.05% trypsin/ 0.53 mM EDTA and plated in Petri dishes to allow embryoid body (EB) formation in ESGRO complete basal medium (Millipore) with 15% FBS. After 8 days, EBs were plated in gelatin-coated plates with serum-free ITS Fn (Insulin-Transferrin-Selenium-Fibronectin, Sigma) medium and incubated for 20 days.
2.7. Teratoma Formation and Immunohistology
Mouse iPS cells (4 × 106 cells from clones 1 and 2) were resuspended in a mixture of DMEM/F12 and Matrigel (BD), and injected subcutaneously by a 20G needle in the dorsal region of athymic nude mice. Four weeks after injection, tumors were dissected, fixed in Bouin’s solution (Sigma), and embedded in paraffin. Sections of teratomas were analyzed by H&E staining and immunohistology.
3. RESULTS
3.1. Characterization of Mouse iPS Cells Generated from Ear Mesenchymal Stem Cells with a Single Constitutive Polycistronic Vector
Mouse ear mesenchymal stem cells (mEMSC) (Figure 1(a)) were reprogrammed by overexpressing the four
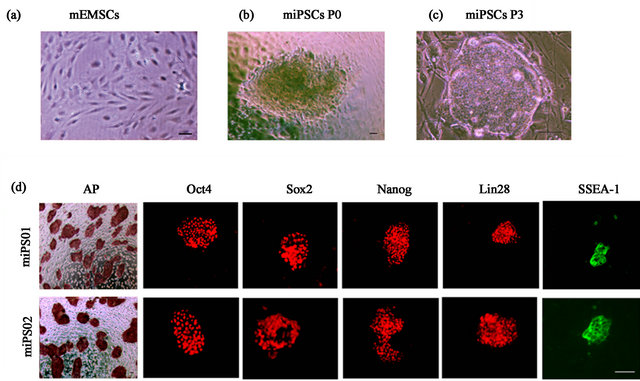
Figure 1. Generation of mouse iPS cells from mouse ear mesenchymal stem cells. (a) Mouse ear mesenchymal stem cells (mEMSC) in culture; (b-c) One representative mouse iPS cell colony generated from mEMSCs at passage 0, and passage 3, separately; (d) Alkaline phosphatase (AP) staining and immunostaining for Oct4, Sox2, Nanog, Lin28 and SSEA1 in miPS01 and miPS02 EMSCderived colonies.
pluripotency factors Oct4, Klf4, Sox2, and c-Myc (OKSM) in a single lentivirus vector. Several colonies with ES cell-like morphology first became visible 8 days after transduction (Figure 1(b)). On day 17, mouse iPS cell colonies were handpicked and expanded. An average of 30 iPS colonies were detected from 1 × 105 EMSCs that were seeded in two independent experiments indicating a reprogramming efficiency of 0.03%. After two additional passages, the colonies exhibited a more distinct appearance with compact cells displaying high nuclear to cytoplasmic ratios and prominent nucleoli (Figure 1(c)). Ten miPS cell clones were established and the characterization of two representative lines, miPS01 and miPS02, are the basis for this study. Both miPS01 and miPS02 stained positive for the pluripotency markers alkaline phosphatase (AP), Oct4, Sox2, Nanog, Lin28 and SSEA-1 (Figure 1(d)).
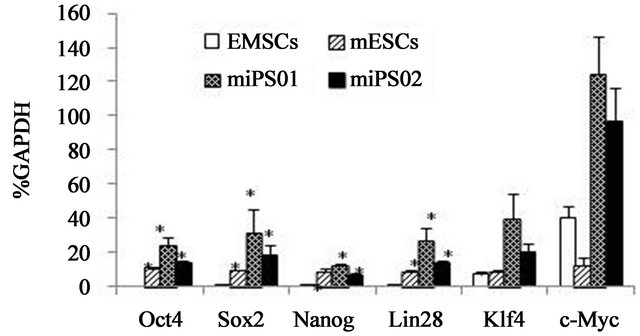
Quantitative RT-PCR analysis revealed similar mRNA levels for the pluripotency genes Oct4, Sox2, Nanog and Lin28 in the miPS01 and miPS02 cell clones and in mouse ES cells. Expression of these markers was not detected in the parental mEMSCs (Figure 2(a)). However, parental EMSCs did express the Klf4 and c-Myc genes with similar (Klf4) and highly elevated (c-Myc) expression of these genes also observed in both of the miPS cell clones (Figure 2(a)).
The generated miPS cells also had a normal karyotype following 8 weeks of culture (Figure 2(c)) indicating that at least a gross level of genetic stability is maintained during and subsequent to, reprogramming. The DNA methylation status of CpG dinucleotides from both the Oct4 and Nanog promoters was also investigated by bisulfite sequencing (Figure 2(b)). These regions are heavily methylated in the parental EMSC (Figure 2(b)), consistent with the lack of transcription we observed by RT-PCR (Figure 2(a)). By contrast, these promoter regions within both iPS cell clones were primarily unmethylated (miPS- 1 and miPS-2, Figure 2(b)), again consistent with the elevated expression levels observed (Figure 2(a)). Oct4 and Nanog promoter demethylation is a hallmark of mES cells (Figure 2(b)) and an essential marker for successful iPS cell production.
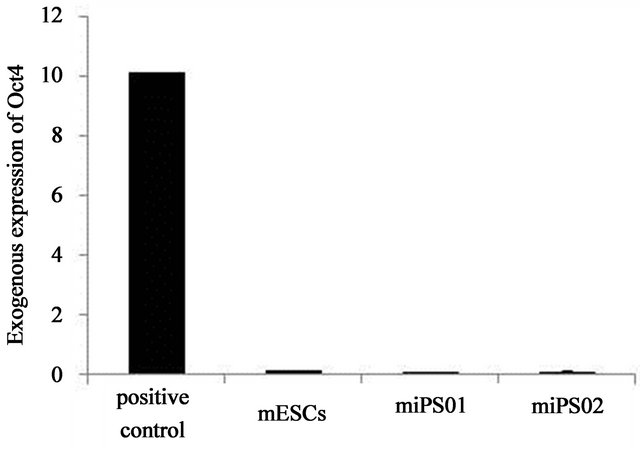
Partially reprogrammed cells are characterized by the continued expression of viral transgenes [33,34] and the aberrant silencing of the imprinted Dlk1-Dio3 gene cluster [29,30]. The polycistronic vector we used in this study results in the transcription of all four transgenes as a single molecule of mRNA. Thus, the absence of expression of any single transgene should indicate the absence of expression for all transgenes in the cassette. Operating under this premise, we found no viral Oct4 expression in either of our two mouse iPS cell lines (Figure 3) indicating silencing of all viral transgenes within the cassette.
Murine Gtl2 and Rian are maternally-expressed, im-
 (a)
(a)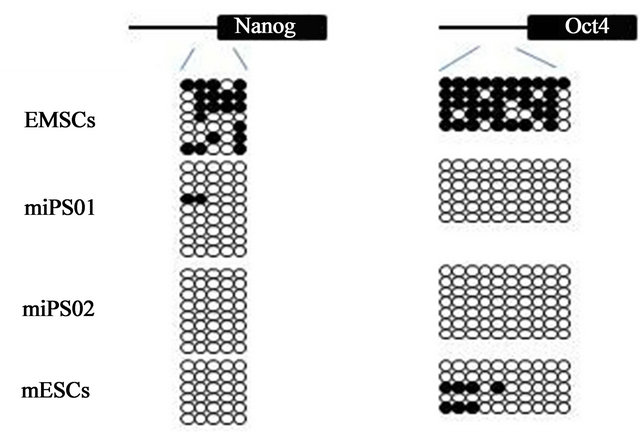 (b)
(b)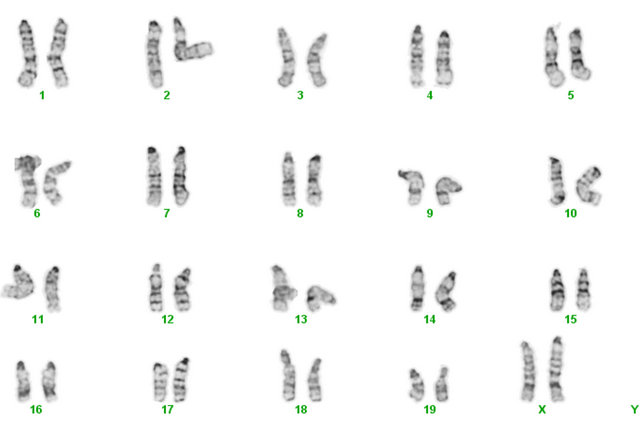 (c)
(c)
Figure 2. Characterization of mouse iPS cells generated from mEMSC. (a) Relative gene expression levels of Oct4, Sox2, Nanog, Lin28, Klf4 and c-Myc in mouse EMSCs, mouse ES cells and two mouse iPS cell lines generated from mEMSCs; (b) DNA methylation status of Oct4 and Nanog promoter regions from mouse EMSCs (top), two mEMSC-derived iPS cell lines (miPS-1 and miPS-2) and mES cells (bottom). Unmethylated CpG dinucleotides (open circles); Methylated CpG dinulceotides (closed circles); (c) Karyotype from a mEMSC-derived iPS cell.
printed genes that reside within the Dlk1-Dio3 gene cluster. The importance of these genes is highlighted by recent reports demonstrating that the expression of Gtl2 reflects full developmental potential in iPS cell clones [29, 30]. Therefore, we used quantitative PCR to determine the expression status of Gtl2, Rian and Dlk-1 genes in the mEMSC and in our two miPS cell clones. These data are shown in Figure 3(b). We found that both Gtl2 and Rian genes were more highly expressed in the parental mEM SCs than they were in mouse ES cells. This was particularly evident regarding the expression level of Rian. Following reprogramming of the mEMSC, Gtl2 expression remained high while the level of Rian expression decreased nearly 5-fold in the iPS cell clones (Figure 3(b), miPS01 and miPS02). Nevertheless, Rian expression in these clones was still higher than it was in the mouse ES cells. Expression of the paternally imprinted gene Dlk1 was minimal among all the cell lines tested (Figure 3(b)). Taken together, these data are consistent with the previous reports regarding the transcriptional activity of Gtl2 and Rian genes in fully pluripotent iPS cell clones.
 (a)
(a)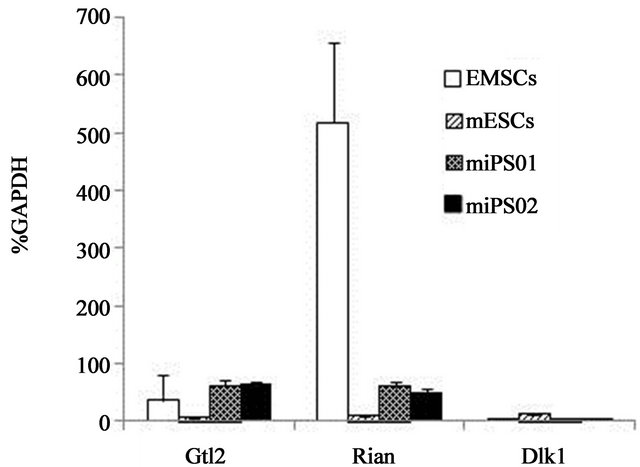 (b)
(b)
Figure 3. Expression of the Dlk-1/Dio3 imprinted gene cluster in mEMSC-derived iPS cell clones. (a) Relative expression level of exogenous Oct4 in mouse ES cells and in miPS01 and miPS02 cell lines; (b) Relative expression levels of Gtl2, Rian and Dlk1 genes in mouse EMSCs, mES cells and in miPS01 and miPS02 cell lines.
3.2. In Vitro Differentiation of mEMSC Derived iPS Cells
To investigate the differentiation potential of our mEM SC-derived iPS cell lines, we first attempted to induce embryoid body (EB) formation (see Materials and Methods). EB-like structures first appeared after 8 days in culture (Figure 4(a)).
Subsequent dissociation of EBs into differentiation medium resulted in the formation of all 3 germ layers at 28 days. The presence of ectoderm was confirmed by the expression of the neuronal marker, microtubule-associated protein 2 (MAP2), mesoderm by the muscle marker, smooth muscle actin (SMA) and endoderm, by the hepatic nuclear factor-3 beta (HNF-3β) protein (Figure 4(b)). Quantitative RT-PCR of mRNA isolated from these differentiated cells (diff-D28) also showed a decrease in expression of the pluripotency genes Oct4, Sox2, Nanog, Lin28, Klf4, and cMyc relative to the iPS cell clones (Figure 5(a)).
Expression of the Gtl2, Rian and Dlk1 genes all increased following differentiation (Figure 5(a), diff-D28 versus miPSCs). Coincident with the increased expression of the Dlk1/Dio3 gene cluster genes was ans increase in expression of the differentiation-specific MAP2, SMA, Desmin and HNF-3β genes in diff-D28 cells relative to mES and miPS cell clones (Figure 5(b)). Interestingly, diff-D28 cells expressed higher levels of MAP2 and HNF-3β genes but lower levels of Desmin and SMA genes, than mEMSCs (Figure 5(b)).
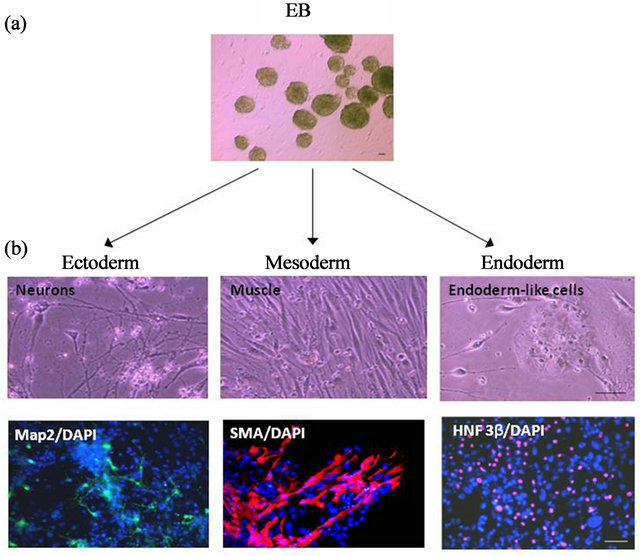
Figure 4. In vitro differentiation of mEMSC-derived iPS cell clones. (a) Representative embryoid bodies generated from mEMSCs; (b) Top panel: morphology of mouse iPS cells following differentiation into ectoderm, mesoderm and endoderm lineages. Bottom panel: corresponding ectoderm (MAP2), mesoderm (SMA) and endoderm (HNF3β) markers detected in differentiated tissues by immunofluorescence.
3.3. Teratoma Formation from miPS Clones
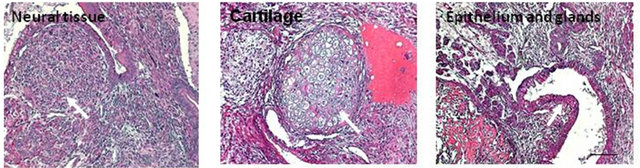
We next determined the capacity of our miPS clones to form teratoma, a hallmark of pluripotency. Mouse iPS cells were resuspended and injected subcutaneously into the dorsal region of athymic nude mice. Tumors were observed from both mEMSC-derived iPS clones after 26 days. They were removed and stained with H&E to reveal gross tissue structures (Figure 6(a), as indicated) and with specific antibodies for the ectoderm, mesoderm and endoderm markers, MAP2, SMA and HNF-3, respectively (Figure 6(b)). These analyses demonstrated that all 3 germ layers were present in the teratoma produced.
4. DISCUSSION
The establishment of induced pluripotent stem (iPS)
 (a)
(a)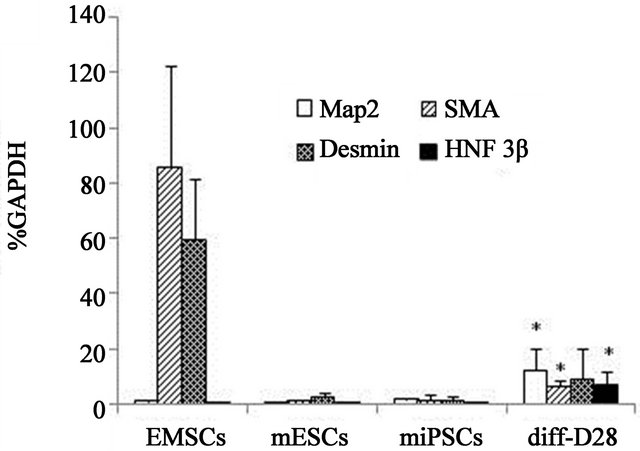 (b)
(b)
Figure 5. Gene expression before and after in vitro differentiation of miPS cell clones. (a) Relative expression levels of Oct4, Sox2, Nanog, Lin28, Klf4, c-Myc, Gtl2, Rian and Dlk genes in mEMSC-derived iPS cells before and after in vitro differentiation; (b) Relative expression levels of MAP2, Desmin, smooth muscle actin (SMA) and HNF3β genes in mEMSC, mES cells, EMSC-derived miPS cells and differentiated miPS cells (diffD28) following 28 days in culture.
cells is a complex and inefficient process that currently requires considerable cell manipulation. Moreover, once iPS cells are produced, they must be validated through labor-intensive experiments demonstrating their capacity for differentiation. This authentication includes embryoid body (EB) formation, the expression of pluripotency and pluripotency-related genes and teratoma formation [35]. However, such analyses reveal that many iPS cell lines produced are not fully pluripotent and thus, are of limited value. Finally, only a small number of parental cell types are known to produce high quality iPS cells and the most appropriate cell type remains poorly defined [31]. The availability of an easily obtainable cell type that is capable of efficiently producing high-quality iPS cells would therefore, be an important advance for the field. In addition, the identification of an early, reliable marker for full pluripotency in this cell type would also streamline the production process by limiting extensive validation efforts to those select iPS cell lines with a high probability for success.
Two recent reports may provide an important clue regarding such an early marker by elucidating the role that activation of the Dlk-1/Dio3 imprinted gene cluster plays in determining the developmental potential of mouse iPS cells clones [29,30]. Salient to our work described here, Stadtfeld et al. found that somatic cells expressing high levels of the Dlk-1/Dio3 cluster gene, Gtl2, produced developmentally competent (Gtl2on) iPS cells while somatic cells expressing low levels of Gtl2 gave rise to developmentally incompetent (Gtl2off) iPS cells clones. Thus, there appears to be a clear positive correlation between Gtl2 activity in somatic cells and the developmental potential of the resultant iPS cell clones. We found that mouse EMSC express high levels of two Dlk-1/Dio3 cluster genes, Gtl2 and Rian, and that Gtl2 expression remained high in both of the EMSC-derived iPS cell clones we produced (Gtl2on) (Figure 3(b)). Moreover, these Gtl2on iPS cell clones were fully competent for both in vitro and in vivo differentiation (Figures 4-6). Taken together, these data provide new evidence in support of the notion that Dlk-1/Dio3 cluster activity is an indicator for full developmental competence of iPS cell clones.
Somewhat surprisingly, we found that Gtl2 expression was higher in our iPS cells than in mouse ES cells (Figure 3(b)) raising the interesting possibility that EMSC may be a viable alternative to mouse fibroblasts, the current standard for iPS cell production [30]. Future studies specifically designed to compare the developmental potential of iPS cells produced from a broader range of Gtl2 expressing and non-expressing somatic cell types will be of considerable value in this regard. Targeting the Dlk-1/Dio3 imprinted gene cluster for activation in nonexpressing somatic cell lines, through epigenetic modifiEctoderm Mesoderm Endoderm
 (a)
(a)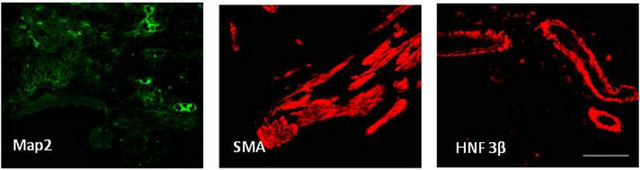 (b)
(b)
Figure 6. Teratoma formation and analysis. (a) All three embryonic germ layers found in excised teratomas (H&E staining); (b) Lineage confirmation by marker detection; MAP2 (ectoderm), smooth muscle actin (SMA) (mesoderm), and HNF3β (ectoderm) (Immunofluorescence).
cations that reverse the silencing of these genes [30], may also broaden the repertoire of usable cell lines for highly efficient iPS cell production. Finally, the identification of a human equivalent to mEMSC could eventually provide an avenue for more efficient human iPS cell production with broad implications for future research and clinical applications for this technology.
5. ACKNOWLEDGEMENTS
The authors thank Dr. David Burk and Ms. Susan Newman for technical advice. This work was funded by the Louisiana Board of Regents Industrial Ties Program Grant LEQSF (2008-11)-RD-B-06 and NIH SBIR grant 1 R43 HL104977. This project also used Genomics, Cell Biology and Imaging core facilities that are supported in part by COBRE (NIH-8 P20 GM103528) and NORC (NIH 1P30-DK072476) center grants from the National Institutes of Health.
![]()
![]()
REFERENCES
- Takahashi, K. and Yamanaka, S. (2006) Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell, 126, 663-676. doi:10.1016/j.cell.2006.07.024
- Wernig, M., Meissner, A., Foreman, R., Brambrink, T., Ku, M., Hochedlinger, K., Bernstein, B.E. and Jaenisch, R. (2007) In vitro reprogramming of fibroblasts into a pluripotent ES-cell like state. Nature, 448, 318-324. doi:10.1038/nature05944
- Hanna, J., Markoulaki, S., Schorderet, P., Carey, B.W., Beard, C., Wernig, M., Creyghton, M.P., Steine, E.J., Cassady, J.P., Foreman, R., Lengner, C.J., Dausman, J.A. and Jaenisch, R. (2008) Direct reprogramming of terminally differentiated mature B lymphocytes to pluripotency. Cell, 133, 250-264. doi:10.1016/j.cell.2008.03.028
- Maherali, N., Ahfeldt, T., Rigamonti, A., Utikal, J., Cowan, C. and Hochedlinger, K. (2008) A high-efficiency system for the generation and study of human induced pluripotent stem cells. Cell Stem Cell, 3, 340-345. doi:10.1016/j.stem.2008.08.003
- Mikkelsen, T.S., Hanna, J., Zhang, X., Ku, M., Wernig, M., Schorderet, P., Bernstein, B.E., Jaenisch, R., Lander, E.S. and Meissner, A. (2008) Dissecting direct reprogramming through integrative genomic analysis. Nature, 454, 49-55. doi:10.1038/nature07056
- Okita, K., Ichisaka, T. and Yamanaka, S. (2007) Generation of germline-competent induced plu-ripotent stem cells. Nature, 448, 313-317. doi:10.1038/nature05934
- Boland, M.J., Hazen, J.L., Nazor, K.L., Rodriguez, A.R., Gifford, W., Martin, G., Kupriyanov, S. and Baldwin, K.K. (2009) Adult mice generated from induced plu-ripotent stem cells. Nature, 461, 91-94. doi:10.1038/nature08310
- Zhao, X.-Y., Li, W., Lv, Z., Liu, L., Tong, M., Hai, T., Hao, J., Guo, C.-L., Ma, Q.-W., Wang, L., Zeng, F. and Zhou, Q. (2009) iPS cells produce viable mice through tetraploid complementation. Nature, 461, 86-90. doi:10.1038/nature08267
- Kang, L., Wang, J., Zhang, Y., Kou, Z. and Gao, S. (2009) iPS cells can support full-term development of tetraploid blastocyst-complemented embryos. Cell Stem Cell, 5, 135-138. doi:10.1016/j.stem.2009.07.001
- Rubin, L.L. and Haston, K.M. (2011) Stem cell biology and drug discovery. BMC Biology, 7, 9-42.
- Chin, M.H., Mason, M.J., Xie, W., Volinia, S., Singer, M., Peterson, C., Ambartsumyan, G., Aimiuwu, O., Richter, L., Zhang, J., Khvorostov, I., Ott, V., Grunstein, M., La-von, N., Benvenisty, N., Croce, C.M., Clark, A.T., Baxter, T., Pyle, A.D., Teitell, M.A., Pelegrini, M., Plath, K. and Lowry, W.E. (2009) Induced pluripotent stem cells and embryonic stem cells are distinguished by gene expression signatures. Cell Stem Cell, 5, 111-123. doi:10.1016/j.stem.2009.06.008
- Marchetto, M.C., Yeo, G.W., Kainohana, O., Marsala, M., Gage, F.H. and Muotri, A.R. (2009) Transcriptional signature and memory retention of human induced pluripotent stem cells. PLoS One, 4, e7076. doi:10.1371/journal.pone.0007076
- Wilson, K.D., Venkatasubrahmanyam, S., Jia, F., Sun, N., Butte, A.J. and Wu, J.C. (2009) MicroRNA profiling of human-induced pluripotent stem cells. Stem Cells and Development, 18, 749-758. doi:10.1089/scd.2008.0247
- Kim, J.B., Zaehres, H., Wu, G., Gentile, L., Ko, K., Sebastiano, V., Araúzo-Bravo, M.J., Ruau, D., Han, D.W., Zenke, M. and Schöler, H.R. (2008) Pluripotent stem cells induced from adult neural stem cells by re-programming with two factors. Nature, 454, 646-650. doi:10.1038/nature07061
- Meissner, A., Wernig, M. and Jaenisch, R. (2007) Direct reprogramming of genetically unmodified fibroblasts into pluripotent stem cells. Nature Biotechnology, 25, 1177- 1181. doi:10.1038/nbt1335
- Zho, T., Zhang, Z.N., Rong, Z. and Xu, Y. (2011) Immuno-genicity of induced pluripotent stem cells. Nature, 474, 212-215.
- Pera, M.F. (2011) Stem cells: The dark side of induced pluripotency. Nature, 471, 46-47. doi:10.1038/471046a
- Gawronska-Kozak, B. (2004) Regeneration in the ears of immunodeficient mice: Identification and lineage analysis of mesenchymal stem cells. Tissue Engineering, 10, 1251-1265.
- Gawronska-Kozak, B., Manuel, J.A. and Prpic, V. (2007) Ear mesenchymal stem cells (EMSC) can differentiate into spontaneously contracting muscle cells. Journal of Cellular Biochemistry, 102, 122-135. doi:10.1002/jcb.21286
- Rim, J.S., Mynatt, R.L. and Gawronska-Kozak, B. (2005) Mesenchymal stem cells from the outer ear: a novel adult stem cell model system for the study of adipogenesis. The FASEB Journal, 19, 1205-1207.
- Staszkiewicz, J., Frazier, T.P., Rowan, B.G., Bunnell, B.A., Chiu, E.S., Gimble, J.M. and Gawronska-Kozak, B. (2010) Cell growth characteristics, differentiation frequency, and immunophenotype of adult ear mesenchymal stem cells. Stem Cells and Development, 19, 83-92. doi:10.1089/scd.2009.0051
- Abdallah, B.M., Jensen, C.H., Gutierrez, G., Leslie, R.G. Q., Jensen, T.G. and Kassem, M. (2004) Regulation of human skeletal stem cells differentiation by Dlk1/Pref1. Journal of Bone and Mineral Research, 19, 841-852. doi:10.1359/jbmr.040118
- Smas, C.M., Green, D. and Sul, H.S. (1993) Structural characterization and alternate splicing of the gene encoding the preadipocyte EGF-like protein pref-1. Biochemistry, 33, 9257-9265.
- Smas, C.M. and Sul, H.S. (1993) Pref-1, a protein containing EGF-like repeats, inhibits adipocyte differentiation. Cell, 73, 725-734. doi:10.1016/0092-8674(93)90252-L
- Sul, H.S., Smas, C., Mei, B. and Zhou, L. (2000) Function of pref-1 as an inhibitor of adipocyte differentiation. International Journal of Obesity, 24, S15.
- Moon, Y.S., Smas, C.M., Lee, K., Villena, J.A. and Kim, K.H. (2002) Mice lacking paternally expressed Pref-1/ Dlk1 display growth retardation and accelerated adiposity. Mol. The Journal of Cell Biology, 22, 5585-5592. doi:10.1128/MCB.22.15.5585-5592.2002
- Lee, K., Villena, J.A., Moon, Y.S., Kim, K.H., Lee, S., Kang, C. and Sul, H.S. (2003) Inhibition of adipogenesis and development of glucose intolerance by soluble preadipocyte factor-1 (Pref-1). Journal of Clinical Investigation, 111, 453-461.
- Wang, Y., Kim, K.-A., Kim, J.-H. and Sul, H.S. (2006) Pref-1, a preadipocyte secreted factor that inhibits adipogenesis. Journal of Nutrition, 136, 2953-2956.
- Liu, L., Luo, G.-Z., Yang, W., Zhao, X., Zheng, Q., Lv, Z., Li, W., Wu, H.-J., Wang, L., Wang, X.-J. and Zhou, Q. (2010) Activation of the imprinted Dlk1-Dio3 region correlates with pluripotency levels of mouse stem cells. JBC, 285, 19483-19490. doi:10.1074/jbc.M110.131995
- Stadtfeld, M., Apostolou, E., Akutsu, H., Fukuda, A., Follett, P., Natesan, S., Kono, T., Shioda, T. and Hochedlinger, K. (2010) Aberrant silencing of imprinted genes on chromosome 12qF1 in mouse induced pluripotent stem cells. Nature, 465, 175-183. doi:10.1038/nature09017
- Niibe, K., Kawamura, Y., Araki, D., Morikawa, S., Miura, K., Suzuki, S., Shimmura, S., Sunabori, T., Mabuchi, Y., Nagai, Y., Nakagawa, T., Okano, H. and Matsuzaki, Y. (2011) Purified mesenchymal stem cells are an efficient source for ips cell induction. PLoS One, 6, 1-9. doi:10.1371/journal.pone.0017610
- Staszkiewicz, J., Gimble, J.M., Manuel, J.A. and Gawronska-Kozak, B. (2008) IFATS collection: Stem cell antigen-1-positive ear mesenchymal stem cells display enhanced adipogenic potential. Stem Cells, 26, 2666-2673. doi:10.1634/stemcells.2008-0270
- Hotta, A. and Ellis, J. (2008) Retroviral vector silencing during iPS cell induction: An epigenetic beacon that signals distinct pluripotent states. Journal of Cellular Biochemistry, 105, 940-948. doi:10.1002/jcb.21912
- Silva, J., Barrandon, O., Nichols, J., Kawaguchi, J. and Thorold, W. (2008) Theunissen, Austin Smith. Promotion of reprogramming to ground state pluripotency by signal inhibition. PLoS Biology, 60, 2237-2247.
- Chan, E.M., Ratanasirintrawoot, S., Park, I.-H., Manos, P.D., Loh, Y.-H., Huo, H., Miller, J.D., Hartung, O., Rho, J., Ince, T.A., Daley, G.Q. and Schlaeger, T.M. (2009) Live cell imaging distinguishes bona fide human iPScells from partially reprogrammed cells. Nature Biotechnology, 27, 1033-1037. doi:10.1038/nbt.1580

