Advances in Bioscience and Biotechnology
Vol.05 No.14(2014), Article ID:52044,11 pages
10.4236/abb.2014.514120
Optimization of Protease Production by Bacillus mojavensis A21 on Chickpea and Faba Bean
Samiha Mhamdi1, Anissa Haddar1, Ibtissem Hamza Mnif1, Fakher Frikha2, Moncef Nasri1, Alya Sellami Kamoun1,2*
1Laboratoire de Génie Enzymatique et de Microbiologie, Université de Sfax, Ecole Nationale d’Ingénieurs de Sfax, Sfax, Tunisia
2Université de Sfax, Faculté des Sciences de Sfax, Sfax, Tunisia
Email: *kamoun_alya@tunet.tn
Copyright © 2014 by authors and Scientific Research Publishing Inc.
This work is licensed under the Creative Commons Attribution International License (CC BY).
http://creativecommons.org/licenses/by/4.0/



Received 25 September 2014; revised 26 October 2014; accepted 25 November 2014
ABSTRACT
Response surface methodology (RSM) was employed to optimize the medium composition and culture conditions for the production of alkaline proteases by Bacillus mojavensis A21 on uncom- mon substrates: chickpea (CF) and faba bean (FF) flours. A significant positive influence of tem- perature, CF, FF, incubation time and inoculums size on the protease production was evaluated by Plackett Burman Design. Among these, CF was the most influential factor. The enhancement of protease to 9127 U/ml was achieved with the optimization procedure on the medium composed of (g/l): CF, 40; FF 30, NaCl 2.0; KH2PO4 1; K2HPO4 1; CaCl2, 0.1; MgSO4 0.1. The cultures were con- ducted for 72 hours with an IS of 2%, at 30˚C, an agitation speed of 150 rpm and an initial pH of 8.0. More interestingly, the optimization was accomplished using two cheap and local fermentation substrates, CF and FF, which could result in a significant reduction in the cost of medium consti- tuents. The maximum alkaline protease production was 9127 U/ml after 72 h of incubation and showed 5-fold increase in protease production over the initial level.
Keywords:
Bacillus mojavensis, Protease, Chickpea, Faba Bean, Plackett Burman Design

1. Introduction
Microorganisms are essential sources for enzyme production for industry. Among these enzymes, proteases account for nearly 60% of the total industrial enzyme market [1] . Bacillus strains are the main producer for large proportion of commercially available proteases.
The use of alkaline proteases has increased remarkably in many industrial processes, including the production of detergents, food processing, in animal feed and for silver regeneration of X-ray films [2] . It is well known that extracellular protease production by microorganisms is greatly influenced by the media components, especially the carbon and nitrogen sources, but also including additional factors such as the temperature, pH, incubation time, agitation and inoculums density [3] . Since approximately 30% - 40% of the production cost in industrial enzyme production is due to medium growth, optimizing the medium composition is especially vital [4] .
In general, no defined medium could be carried out for the production of alkaline proteases from different microorganisms; each strain has its specific required conditions for maximum enzyme production [2] . The effect of environmental conditions on the production of extracellular proteolytic enzymes can play an important role in the induction or repression of the enzyme by specific compounds [5] . Protease production is dependent on the availability of both carbon and nitrogen sources within the medium and both of these parameters exert regulatory effects on enzyme synthesis [6] . At present, the cost of enzyme production is very high due to high cost of the substrates and microbiological media used. Therefore, development of novel processes to increase the yield of proteases with respect to their industrial requirements, coupled with lowering the production cost, is highly appreciable from a commercial point of view. Therefore, microbial enzymes producing industries are always in search of new and cheaper methods to enhance enzymes production and decrease the market price of the desired enzyme. In this way, efforts have been directed to search for alternatives explore different means to reduce protease production cost through improving the yield and using either cost-free or low-cost feed stocks or agricultural products as substrates for proteases production [7] [8] .
Optimization of media compounds by the traditional “one-variable at-a-time” strategy is the most frequently used method in biotechnology [9] . This strategy is extremely time consuming and expensive when a large number of variables need to be considered. Additionally, this method is unable to detect the interactions among different production factors true and then unable to find optimal conditions. Recently, the use of statistical approaches involving Plackett-Burman design has gained momentum in medium optimization [10] - [12] . This method is also useful for understanding interactions among various physico-chemical parameters using a minimal number of experiments. The Plackett-Burman design allows for the screening of the main factors from a large number of variables, and this information can be retained in further optimization.
Bacillus mojavensis A21 has been recently identified as a producer of extracellular bleaching-stable alkaline proteases. Haddar et al., 2009 [13] [14] had purified three detergent stable alkaline serine-proteases from the culture supernatant of B. mojavensis A21. Otherwise, B. mojavensis A21 proteases were used to produce hydrolysates of various proteins [15] - [17] . Haddar et al. [18] optimized for B. mojavensis A21 culture conditions, by Plackett-Burman and Central Composite Design and the protease production was enhanced by 14 folds on hulled grain of wheat and sardinella peptone based medium.
In view of the promising applicability of the alkaline proteases as a builder for detergents, there is an interest in producing these enzymes in the highest yields with the lowest cost fermentation media. Herein, we report a statistical optimization of culture conditions for the production of proteases by B. mojavensis A21, on uncommon fermentation substrates such as faba bean and chickpea flours.
2. Material and Methods
2.1. Microorganism
B. mojavensis A21 producing bleach-stable alkaline proteases was isolated from a marine water sample. It was identified on the basis of 16 S rRNA gene sequencing [10] .
2.2. Carbon and Nitrogen Substrates
Seeds of chickpea (Cicer arietinum), faba bean (Vicia faba), lens (Lens culinaris), corn (Zea mays), pea (Pisum sativum) and mil (Pennisetum glaucum) were purchased in local market and minced in our laboratory to obtain the corresponding flours: CF, FF, LF, CNF, PF and MF. Sardinella peptone (SP) [19] , combined heads and viscera sardinella powder (CHVSP) [20] and hydrolyzed feathers (HF) [21] were prepared in LGEM laboratory.
2.3. Culture Conditions
B. mojavensis A21 was maintained on Luria-Bertani (LB) medium [22] . The proteolytic enzymes were initially produced on a basal medium composed of g/l: Yeast extract (YE) 1, NaCl 2, K2HPO4 1, KH2PO4 0.3, MgSO4 1, and supplemented with a carbon source at 30 g/l. The cultures were conducted at pH 8, 30˚C and 200 rpm for 48 hours in 250 ml Erlenmeyer flasks with a working volume of 25 ml. The culture medium was centrifuged at 10,000 g for 15 min at 4˚C and the cell free supernatant was used for estimation of proteolytic activity.
2.4. Protease Assay
Protease activity was measured according to the method of Kembhavi et al. (1993) [23] using casein as a substrate. A sample of 0.5 ml suitably diluted enzyme, was mixed with 0.5 ml of 100 mM glycine-NaOH, pH 10.0 containing casein at 1% (w/v) and incubated for 15 min at 60˚C. The reaction was stopped with the addition of 0.5 ml of trichloroacetic acid (20%; w/v). The mixture was allowed to stand at room temperature for 15 min and then centrifuged at 10,000 g for 15 min to remove the precipitate. The absorbance of the soluble fraction was measured at 280 nm. A standard curve was generated using solutions of 0 - 50 mg/l of tyrosine. One unit of protease activity was defined as the amount of enzyme required to liberate 1 μg of tyrosine per min under the experimental conditions used.
2.5. Experimental Designs
A Plackett-Burman design is used for rapid screening multifactor to find the most significant independent factors [5] [24] [25] . The independent variables of the protease production were Chikpea flour (CF), Faba bean flour (FF), K2HPO4, KH2PO4, CaCl2, NaCl, MgSO4, incubation temperature, speed of agitation, incubation time and inoculums size (IS). Then the eleven factors were investigated using the Plackett-Burman design with a first-order polynomial equation. Each factor was tested at low (−1) and high (+1) levels (Table 1). Eleven variables were screened in 16 experimental runs in which 4 runs were repeated at the central level (0) (Table 2). The fitted first-order model is: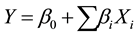 ; Y is the predicted response,
; Y is the predicted response, 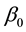 and
and 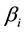 are constant coefficients, and Xi is the coded independent factors.
are constant coefficients, and Xi is the coded independent factors.
Box-Behnken design of RSM (Response Surface Methodology) [25] [26] was employed to optimize levels of the five selected factors (temperature, CF, FF, incubation time and inoculums size) for enhancing the protease production. The five independent factors were investigated at three different levels (−1, 0, +1) (Table 3) and the experimental design used for the study is shown in Table 4.
The protease production was fitted using a second-order polynomial equation and a multiple regression of the data was carried out for obtaining an empirical model related to the factors. The general form of the second-order polynomial equation is:
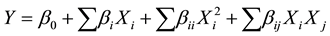
where Y is the predicted response, 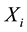 and
and 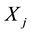 are independent factors,
are independent factors, 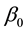 is the intercept,
is the intercept, 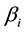 is the linear
is the linear
Table 1. Levels of the factors tested in Plackett-Burman design.
CF: chickpea flour; FF: faba bean flour and IS: inoculums size.
Table 2. Plackett-Burman design for screening of significant factors affecting protease production levels.
Variables X1 to X11 are described in Table 1.
Table 3. Levels of the factors tested in Box-Behnken design.
coefficient, 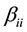 is the quadratic coefficient and
is the quadratic coefficient and 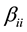 is the interaction coefficient.
is the interaction coefficient.
Design-expert, version 7.0 (STAT-EASE Inc., Minneapolis, USA) was used for the experimental designs and statistical analysis of the experimental data. The analysis of variance (ANOVA) was used to estimate the statistical parameters.
2.6. Chemical Composition of CF and FF
Dry weights of chickpea and faba bean flours were determined after heating samples at 105˚C to constant weight, and then ash content was determined after heating dried samples at 600˚C for 2 h. Total nitrogen was determined using the Kjeldahl method and then protein content was estimated. Crude fat was determined gravimetrically after Soxhlet extraction of dried samples with diethyl ether. Starch and cellulose were estimated as described by Ewers [27] and Gautier [28] respectively.
Table 4. The Box-Behnken design of RSM for optimization of the protease production by B. mojavensis A21.
3. Results and Discussion
3.1. Effect of Carbon Sources on Protease Production
B. mojavensis A21 strain was cultivated on initial basal medium containing YE at 1 g/l as nitrogen source and different complex sources of carbon (30 g/l) such as chickpea, faba bean, lens, corn, pea and mil. As shown in Figure 1, B. mojavensis A21 was found to produce proteolytic enzymes in the presence of complex organic sources. Among these, chickpea flour (CF) and faba bean flour (FF) gave the highest proteolytic activities of 2995 U/ml and 2350 U/ml respectively. Various concentrations of CF and FF were then tested for the proteases production by B. mojavensis A21. It was shown that the proteolytic activity was highly stimulated when increasing the concentrations of CF. This production reached 4336 U/ml on CF at 60 g/l and then it was stabilized (Figure 2).
Our results are in concordance with other reports regarding proteases production, in which it has been observed that complex carbon sources such hulled grain of wheat [12] , soybean meal [29] or fish powder [30] , were efficiently used for protease production by microorganisms. Such products constitute better substrates for enzyme production than simple sugars like glucose which can induce catabolic repression [31] .
Figure 1. Effect of different carbon sources on protease production by B. mojavensis A21 at 37˚C in the basal medium (pH 8.0) supplemented with different carbon sources at 30 g/l. Flours were obtained from chickpea: CF, faba bean: FF, lens: LF, corn: CNF, pea: PF and mil: MF.
Figure 2. Effect of increasing concentration of CF on protease production by B. mojavensis A21 in the basal medium.
3.2. Effect of Nitrogen Sources
In a second experiment, protease production by B. mojavensis A21 was checked in the basal medium containing CF at 30 g/l and supplemented with 1 g/1 of various nitrogen sources YE: yeast extract, SP: sardinella peptone, CHVSP: combined heads and viscera sardinella powder, HF: hydrolyzed feathers. As shown in Figure 3, the bacteria exhibited high levels of enzyme activity of 3000 U/ml and 2850 U/ml when the medium was supplemented with YE or SP respectively. The supplementation of medium with enzymatically hydrolyzed proteins CHVSP or HF had a negative influence on the enzyme production which drops to 1081 and 1000 U/ml respectively.
In order to substitute YE by a low cost and available nitrogen source, FF was assayed at different concentrations from 10 to 40 g/l in basal medium containing 40 g/l of CF as carbon source. Protease production was enhanced with the increase of FF concentration up to 30 g/l (Figure 4), reached 4645 U/ml, and then decreased beyond.
In this study, two kinds of legume seeds were shown to be excellent carbon and nitrogen sources for B. mojavensis A21 proteases production which was maximized in the presence of CF. This can be explained by the fact that the two respective flours contain high amounts of proteins.
Figure 3. Effect of different nitrogen sources on protease production by B. mojavensis A21 Cultures were conducted at 37˚C (pH 8.0) in the basal medium containing CF at 30 g/l and supplemented (or not) with one of the following nitrogen sources (30 g/l): YE: yeast extract, SP: sardinella peptone, CHVSP: combined heads and viscera sardinella powder, HF: hydrolyzed feathers.
Figure 4. Protease production by B. mojavensis A21 on medium containing CF at 40 g/l supplemented with various concentrations of FF.
3.3. Screening of Significant Variables Using Plackett-Burman Design
The Plackett-Burman design is a powerful method for screening significant factors. Sixteen runs were carried out to analyze the effect of 11 variables on protease production and the results are demonstrated in Table 2. The t-test was used to identify the effect of every factor on protease production. Figure 5 shows that CF, FF, temperature, incubation time and IS are the significant factors (p < 0.05). According to the model, CF, FF, NaCl, Speed agitation and incubation time exerted a positive effect. However, CaCl2, KH2PO4, K2HPO4, MgSO4, temperature and IS exerted negative effects on the protease production. CF, FF, temperature, incubation time and IS were selected for further optimization to obtain a maximum response. CaCl2, KH2PO4, K2HPO4 and MgSO4 have been fixed to the low level corresponding to 0.5 g/l, 0.1 g/l, 0.1 g/l and 0.1 g/l, respectively. The NaCl and speed agitation have been fixed to the levels corresponding to 2 g/l and 250 rpm, respectively.
3.4. Optimization of Significant Variables Using Variance (ANOVA) Analysis
A response surface design is further applied when the optimal region for running the process has been identified. Based on the Plackett-Burman design, RSM using Box-Behnken design was applied to determine the optimal levels of the five selected variables (CF, FF, temperature, incubation time and IS) which significantly influenced the protease production. The respective low and high levels with the coded levels for the five variables are defined in Table 3. A total of 46 runs with different combination of CF, FF, temperature, incubation time and IS were performed (Table 4). The experimental resultants were analyzed by standard ANOVA (Table 5) and the Box-Behnken design was fitted with the second-order polynomial equation (in coded factor):
Y = 1761(±52.93) + 723.375(±89.75) CF + 413.1875(±89.75) FF + 156.375(±89.75) Time − 378.3125(±89.75) Temperature + 466(±179.50) CF × Time − 375(±179.50) Time × Temperature
where Y the protease production by B. mojavensis A21.
The statistical significance of the model equation was evaluated by the F-test for ANOVA. The model F-value of 19.68 implies the model is significant. There was only a 0.01% chance that the model F-value could occur due to noise. The lack of fit F-value of 2.16 implies that was not significant relative to the pure error and that was a 19.83 % chance that the lack of fit F-value could occur due to noise. The adequate precision measures the
Figure 5. Identification of significant variables for proteases production by B. mojavensis A21 using Plackett-Burman design (Pareto chart); where A is CF, B is FF, H is temperature, K is time and L is inoculums size.
Table 5. Analysis of variance (ANOVA) for response surface reduced 2FI Model.
Standard Deviation = 358.99; R2 = 0.752, adjust R2 = 0.714; *Statistically significant at 95% of confidence level.
signal to noise ratio. A ratio greater than 4 is desirable. Adequate precision which was calculated to be 17.73 indicates an adequate signal. This model can be used to navigate the design space.
According to the model, we can notice that the increasing of the temperature causes to a decrease in the protease production. Table 5 shows that only CF, FF, temperature, incubation time, the interaction CF × Time and the interaction Time × Temperature are significant however, IS is not significant. The interaction CF × Time exerted a positive effect on the protease production, whereas the interaction Time × Temperature exerted a negative effect on the protease production. Among the five significant variables (CF, FF, incubation time, temperature and IS), CF concentration was found to have the greatest effect on the production of alkaline proteases. The optimal values of the five variables predicted by the model are CF = 38.08 g/l, FF = 27.49 g/l, time = 67.68 h, temperature = 30.26˚C and IS = 3.37. The maximum predicted protease production was 3798.67 ± 435.63 U/ml.
In a first attempt to optimize B. mojavensis A21 proteases production by statistical design on wheat bran flour and sardinella peptone, Haddar [13] obtained 1783 U/ml. Herein, we showed that protease production by B. mojavensis A21 was enhanced by 5 folds when cultivated on the optimized medium based on chickpea and faba bean flours used as carbon and nitrogen sources. Similarly, several works related the optimization of proteases production by statistical design for various Bacillus strains [30] [32] - [35] .
Using cost-effective media formulation and optimizing this media to determine its minimum requirements for maximum enzyme production is extremely important in industrial-scale protease production for economic reasons. Therefore, using common and local low cost substrates such as chickpea and faba bean, will serve as a potential example for the applications in industrial microbial fermentations. The simple expedient of replacing YE in the medium with CF or FF could significantly lower the production cost.
Maximum proteases production of 9127 U/ml was achieved at the optimized culture conditions: 40 g/l CF, 30 g/l FF, NaCl 2 g/l; KH2PO4 1 g/l; K2HPO4 1 g/l; CaCl2, 0.5 g/l; MgSO4 0.1 g/l and IS 2% (v/v). In addition other factors were taken at suitable levels, as shown in the Plackett-Burman design matrix: a temperature of 30˚C, a speed of agitation at 150 rpm, incubation time of 72 h and an initial pH of 8.0.
3.5. Time Courses of Protease Production and Cell Growth
The time courses of protease activity and the growth of B. mojavensis A21 under the optimized conditions are shown in Figure 6. The kinetics showed that the secretion of the proteases was associated with the bacterial growth. Protease activity increased during the exponential growth phase and reached a proteolytic activity of 9127 U/ml at the end of the stationary phase, after 72 h of growth. The kinetics of growth and protease production showed that the majority of alkaline proteases (7500 U/ml) were secreted by B. mojavensis A21 at the end of the exponential growth phase of the bacteria.
Figure 6. Kinetics of growth and alkaline proteases production by B. mojavensis A21 in the optimized medium consisting of (g/l): CF 40; FF 30, NaCl 2.0; KH2PO4 1.0; K2HPO4 1; CaCl2 0.1; MgSO4 0.1; pH 8.0 and IS 2%. (

Table 6. Chemical composition of CF (chickpea flour) and FF(faba bean flour).
3.6. Chemical Compositions of CF and FF
The chemical compositions of CF and FF were determined (Table 6). Both FF and CF exhibited high content of proteins, 29.08% and 23.64%, and starch 42.24% and 46.91% respectively. While FF is richer in cellulose and had a lower fat content than CF. Though, proteins and starch represent both more than 70% of raw material for the two pulses.
The two pulses contained important quantities of proteins and starch which are higher than in hulled grain of wheat (HGW) 8% - 12% and 50% - 60% respectively [12] . Our results and those of Haddar et al. [13] clearly indicated that proteins and starch are useful for protease production. Herein the proteases production by B. mojavensis A21 strain, was amplified by the use of highly containing proteins substrates such FF and CF. This enhancement could be explained by the higher content of proteins compared to the previously used raw material: HGW. More proteins probably stimulate more proteases synthesis.
Thus, the use of CF and FF as carbon and nitrogen sources may result in a more cost-effective process, and so the production of detergent proteases by the A21 strain will be effectively economic.
4. Conclusion
Due to the increasing economic relevance of alkaline proteases, this study was conducted in an attempt to optimize a variety of fermentation parameters, including medium compositions and culture conditions, for maximal alkaline protease production. Eleven variables were tested using the Plackett-Burman design, and five variables (CF, FF, incubation time, temperature and IS) were selected as the most influential factors on enzyme production. The maximum alkaline protease production was amplified by 5 folds over the initial level and reached 9127 U/ml after 72 h of incubation. The optimized medium established in this work might result in a significant reduction in the cost of medium constituents and would thus offer advantages for large-scale fermentations.
Acknowledgements
The authors are thankful to Mrs Imen Khouni from the Centre de Biotechnologie Borj Cédria (Tunisia) for starch and cellulose determination.
The present work was supported by the Ministère de l’Enseignement Supérieur et de la Recherche Scienti- fique, Tunisia.
References
- Banerjee, U.C., Sani, R.K., Azmi, W. and Soni, R. (1999) Thermostable Alkaline Protease from Bacillus brevis and Its Characterization as a Laundry Detergent Additive. Process Biochemistry, 35, 213-219. http://dx.doi.org/10.1016/S0032-9592(99)00053-9
- Gupta, R., Beg, Q.K. and Lorenz P. (2002) Bacterial Alkaline Proteases: Molecular Approaches and Industrial Applications. Applied Microbiology and Biotechnology, 59, 15-32. http://dx.doi.org/10.1007/s00253-002-0975-y
- Johnvesly, B. and Naik, G.R. (2001) Studies on Production of Thermostable Alkaline Protease from Thermophilic and Alkaliphilic Bacillus sp. JB-99 in a Chemically Defined Medium. Process Biochemistry, 37, 139-144. http://dx.doi.org/10.1016/S0032-9592(01)00191-1
- Kirk, O., Borchert, T.V. and Fuglsang, C.C. (2002) Industrial Enzyme Applications. Current Opinion in Biotechnology, 13, 345-351. http://dx.doi.org/10.1016/S0958-1669(02)00328-2
- Wang, Q., Hou, Y., Xu, Z., Miao, J. and Li, G.O. (2007) Optimization of Cold-Active Protease Production by the Psychrophilic Bacterium Colwellia sp. NJ341 with Response Surface Methodology. Bioresource Technology, 99, 1926- 1931. http://dx.doi.org/10.1016/j.biortech.2007.03.028
- Chu, I.M., Lee, C. and Li, T.S. (1992) Production and Degradation of Alkaline Protease in Batch Cultures of Bacillus subtilis ATCC 14416. Enzyme and Microbial Technology, 4, 55-61.
- Sandhya, C., Sumantha, A., Szakacs, G. and Pandey, A. (2005) Comparative Evaluation of Neutral Protease Production by Aspergillus oryzae in Submerged and Solid-State Fermentation. Process Biochemistry, 40, 2689-2694. http://dx.doi.org/10.1016/j.procbio.2004.12.001
- Prakasham, R.S., Rao, Ch.S. and Sarma, P.N. (2006) Green Gram Husk an Inexpensive Substrate for Alkaline Protease Production by Bacillus sp. in Solid-State Fermentation. Bioresource Technology, 97, 1449-1454. http://dx.doi.org/10.1016/j.biortech.2005.07.015
- Haaland, P.D. (1989) Statistical Problem Solving. In: Haaland, P.D., Ed., Experimental Design in Biotechnology, Marcel Dekker, New York, 1-18.
- Reddy, L.V.A., Wee, Y.J., Yun, J.S. and Ryu, H.W. (2008) Optimization of Alkaline Protease Production by Batch Culture of Bacillus sp. RKY3 through Plackett-Burman and Response Surface Methodological Approaches. Bioresource Technology, 99, 2242-2249. http://dx.doi.org/10.1016/j.biortech.2007.05.006
- Bhunia, B. and Dey, A. (2012) Statistical Approach for Optimization of Physicochemical Requirements on Alkaline Protease Production from Bacillus licheniformis NCIM 2042. Enzyme Research, 2012, Article ID: 905804. http://dx.doi.org/10.1155/2012/905804
- Kumar, R.S., Ananthan, G. and Prabhu, A.S. (2014) Optimization of Medium Composition for Alkaline Protease Production by Marinobacter sp. GACAS9 Using Response Surface Methodology―A Statistical Approach. Biocatalysis and Agricultural Biotechnology, 3, 191-197.
- Haddar, A., Bougatef, A., Agrebi, R., Sellami-Kamoun, A. and Nasri, M. (2009) A Novel Surfactant-Stable Alkaline Serine-Protease from a Newly Isolated Bacillus mojavensis A21. Purification and Characterization. Process Biochemistry, 44, 29-35. http://dx.doi.org/10.1016/j.procbio.2008.09.003
- Haddar, A., Agrebi, R., Bougatef, A., Hmidet, N., Sellami-Kamoun, A. and Nasri, M. (2009) Two Detergent Stable Alkaline Serine-Proteases from Bacillus mojavensis A21: Purification, Characterization and Potential Application as a Laundry Detergent Additive. Bioresource Technology, 100, 3366-3373. http://dx.doi.org/10.1016/j.biortech.2009.01.061
- Hmidet, N., Balti, R., Nasri,R., Sila, A.,Bougatef, A. and Nasri, M. (2011) Improvement of Functional Propreties and Antioxidant Activities of Cuttlefish (Sepia officinalis) Muscle Proteins Hydrolyzed by Bacillus mojavensis A21 Proteases. Food Research International, 44, 2703-2711. http://dx.doi.org/10.1016/j.foodres.2011.05.023
- Younes, I., Ghorbel-Bellaaj, O., Nasri, R., Chaabouni, M., Rinaudo, M. and Nasri, M. (2012) Chitin and Chitosan Preparation from Shrimp Shells Using Optimized Enzymatic Deproteinization. Process Biochemistry, 47, 2032-2039. http://dx.doi.org/10.1016/j.procbio.2012.07.017
- Nasri, R., Chataigné, G., Bougatef, A., Karra Chaabouni, M., Dhulser, P., Nasri, M. and Nedjar-Arroume, N. (2013) Novel Angiotensin I-Converting Enzyme Inhibitory Peptides from Enzymatic Hydrolysates of Goby (Zosterisessor ophiocephalus) Muscle Proteins. Journal of Proteomics, 91, 444-452. http://dx.doi.org/10.1016/j.jprot.2013.07.029
- Haddar, A., Fakhfakh-Zouari, N., Hmidet, N., Frikha, F., Nasri, N. and Sellami Kamoun, A. (2010) Low-Cost Fermentation Medium for Alkaline Protease Production by Bacillus mojavensis A21 Using Hulled Grain of Wheat and Sardinella Peptone. Journal of Bioscience Bioengineering, 110, 288-294. http://dx.doi.org/10.1016/j.jbiosc.2010.03.015
- Souissi, N., Bougatef, A., Triki-Ellouz, Y. and Nasri, M. (2007) Biochemical and Functional Properties of Sardinella (Sardinella aurita) By-Product Hydrolysates. Food Technology and Biotechnology, 45, 187-194.
- Triki-Ellouz, Y., Bayoudh, A., Kammoun, S., Gharsallah, N. and Nasri, M. (2001) Production of Protease by Bacillus subtilis Grown on Sardinelle Heads and Viscera Flour. Bioresource Technology, 80, 49-51. http://dx.doi.org/10.1016/S0960-8524(01)00057-8
- Fakhfakh, N., Ktari, N., Haddar, A., Hamza Mnif, I., Dahmen, I. and Nasri, M. (2011) Total Solubilisation of the Chicken Feathers by Fermentation with a Keratinolytic Bacterium, Bacillus pumilus A1, and the Production of Protein Hydrolysate with High Antioxidative Activity. Process Biochemistry, 46, 1731-1737. http://dx.doi.org/10.1016/j.procbio.2011.05.023
- Miller, J.H. (1972) Experiments in Molecular Genetics. Cold Spring Harbor Laboratory, Cold Spring Harbor, New York.
- Kembhavi, A.A., Kulkarni, A. and Pant, A.A. (1932) Salt-Tolerant and Thermostable Alkaline Protease from Bacillus subtilis NCIM No 64. Applied Biochemistry and Biotechnology, 38, 83-92. http://dx.doi.org/10.1007/BF02916414
- Plackett, R.L. and Burman, J.P. (1946) The Design of Optimum Multifactorial Experiments. Biometrika, 33, 305-325. http://dx.doi.org/10.1093/biomet/33.4.305
- Box, G.E.P., Hunter, W.G. and Hunter, J.S. (1978) Statistics for Experiments: An Introduction to Design Data Analysis and Model Building. John Wiley, New York.
- Khuri, A.I. and Cornell, J.A. (1987) Response Surfaces: Design and Analysis. Marcel Decker Inc., New York.
- Li, X., Xu, T., Ma, X., Guo, K., Kai, L., Zhao, Y., Jia, X. and Ma, Y. (2008) Optimization of Culture Conditions for Production of Cis-Epoxysuccinic Acid Hydrolase Using Response Surface Methodology. Bioresource Technology, 99, 5391-5396. http://dx.doi.org/10.1016/j.biortech.2007.11.017
- Ewers, E. (1965) Determination for Starch by Extraction and Dispersion with Hydrochloric Acid. International Organisation for Standardization (ISO/TC 93/WGL).
- Joo, H.S., Kumar, C.G., Park, G.C., Kim, K.T., Paik, S.R. and Chang, C.S. (2002) Optimization of the Production of an Extracellular Alkaline Protease from Bacillus horikoshii. Process Biochemistry, 38, 155-159. http://dx.doi.org/10.1016/S0032-9592(02)00061-4
- Sellami-Kamoun, A., Ghorbel-Frikha, B., Haddar, A. and Nasri, M. (2011) Enhanced Bacillus cereus BG1 Protease Production by the Use of Sardinelle (Sardinella aurita) Powder. Annals of Microbiology, 61, 273-280. http://dx.doi.org/10.1007/s13213-010-0134-0
- Hanlon, G.W., Hodges, N.A. and Russel, A.D. (1982) The Influence of Glucose, Ammonium and Magnesium Availability on the Production of Protease and Bacitracin by Bacillus licheniformis. Journal of Genetic and Microbiology, 128, 845-851.
- Puri, S., Khali, O. and Gupta, R. (2002) Optimization of Alkaline Protease Production from Bacillus sp. by Response Surface Methodology. Current Microbiology, 44, 286-290. http://dx.doi.org/10.1007/s00284-001-0006-8
- Tari, C., Genckal, H. and Tokatli, F. (2006) Optimization of a Growth Medium Using a Statistical Approach for the Production of an Alkaline Protease from a Newly Isolate Bacillus sp. L21. Process Biochemistry, 41, 659-665.
- Oskouie, S.F.G., Tabandeh, F., Yakhchali, B. and Eftekhar, F. (2008) Response Surface Optimization of Medium Composition for Alkaline Protease Production by Bacillus clausii. Biochemical Engineering Journal, 39, 37-42. http://dx.doi.org/10.1016/j.bej.2007.08.016
- Agrebi, R., Haddar, A, Hajji, M., Frikha, F., Manni, L., Jellouli, K. and Nasri, M. (2009) Fibrinolytic Enzymes from a Newly Isolated Marine Bacterium Bacillus subtilis A26: Characterization and Statistical Media Optimization. Canadian Journal of Microbiology, 55, 1049-1061. http://dx.doi.org/10.1139/W09-05
NOTES
*Corresponding author.









