Advances in Bioscience and Biotechnology
Vol. 3 No. 7 (2012) , Article ID: 24500 , 9 pages DOI:10.4236/abb.2012.37101
Isolation, cytotoxic activity and phylogenetic analysis of Bacillus sp. bacteria associated with the red sea sponge Amphimedon ochracea
![]()
1Marine Chemistry Laboratory, The National Institute of Oceanography and Fisheries, Alexandria, Egypt
2Department of Biochemistry, Faculty of Science, Alexandria University, Alexandria, Egypt
3Department of Marine Zoology, Netherlands Centre for Biodiversity, Leiden, The Netherlands
Email: *haiam_morsy@yahoo.com, m_shreadah@yahoo.com, nihadabdelmonem@yahoo.com, galila_69@yahoo.com, Rob.vanSoest@ncbnaturalis.nl
Received 18 August 2012; revised 23 September 2012; accepted 17 October 2012
Keywords: Sponge; Associated Bacteria; Bacillus sp.; Bioactive Metabolites; Cytotoxic Activity; Phylogenetic Analysis
ABSTRACT
Most of marine sponges harbor dense and diverse microbial communities of bioactivity importance. Four Gram positive bacterial cultures (HA-21, HA-68, HAMS-105 and HA-MS-119) were isolated from the sponge Amphimedon ochracea, collected from the Red Sea coast of Egypt. Bacterial species were identified based on the phylogenetic analysis of the nucleotide sequences of their 16S rDNA genes. The Sequences similarity values of 98% - 100% to other strains in the NCBI database showed strong similarities with the 16S rDNA genes of firmicutes (Bacillus sp.). The four bacterial species were submitted to the GenBank database and had accession numbers of: HA-21 [JQ- 768238]; HA-68 [JQ751264]; HA-MS-105 [JQ768239]; HAMS-119 [JQ768240]. The cytotoxic activities of the bacterial isolates were tested against three established human cancer cell lines; HepG2 (hepatocellular carcinoma), HCT (colon carcinoma) and MCF-7 (breast carcinoma). The inhibitory effect on these cell lines, measured by MTT cell assay protocol, revealed promising cytotoxic activity of the four isolates (IC50 values (µg/mL) were: HA-21: 13.2, 9.3 and 12.2; HA-68: 10.42, 4.3 and 5.5; HA-MS-105: 46.9, 28.6 and 21.3; HAMS-119: 10.42, 6.3 and 22.1; respectively). The recovery of bacterial strains with cytotoxic activity suggests that marine invertebrates remain a rich source for the isolation of culturable isolates capable of producing novel bioactive secondary metabolites.
1. INTRODUCTION
The potential of marine natural products has captivated many researchers over the years. Inspired by the vastness of our oceans, and almost incomprehensible level of biodiversity in the marine environment, researchers have enthusiastically pursued the pharmacological potential of secondary metabolites from marine organisms, with the attention of natural product chemists and pharmaceutical companies being focused firmly on anticancer drugs, with several promising sponge-derived compounds in clinical and preclinical cancer trials [1,2]. However, to the best of our knowledge, despite an enormous number of biologically active compounds being isolated from marine sponges, not a single compound obtained from a sponge has been approved as a drug to date, with a major hurdle on progress being the so-called supply problem [3]. As a significant portion of the bioactive metabolites, thought originally to be products of the source animal often synthesized by their symbiotic microbiota, researchers have paid more attention to microorganisms as a renewable source of bioactive natural products. More specifically, bacteria from the marine environment have shown great potential as suggested by the diversity of secondary metabolites [4].
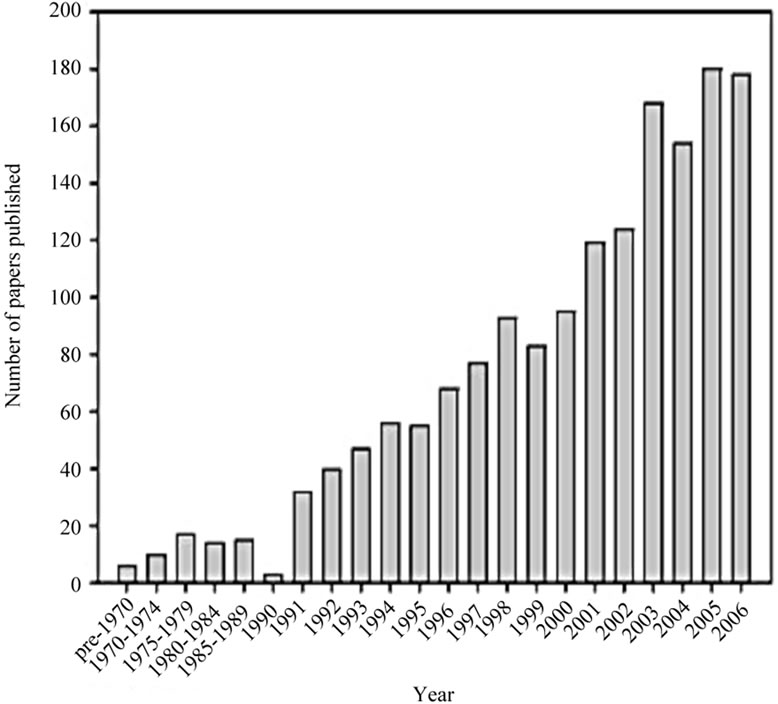
In contrast to the terrestrial environment where plants are the most prolific sources of natural products, this leading position in the sea is taken by invertebrates such as sponges, tunicates, bryozoans, or mollusks, with sponges being the most prolific sources of natural products [5]. The wealth of their secondary metabolites has been related to the role of chemical defense played by these constituents; such ecological function is believed to be crucial for the survival of the producer organisms, which are soft-bodied, sessile or slow-moving animals, lacking, in most cases, morphological defense structures such as shells or spines [6]. Sponges have been the focus of much recent interest (Figure 1) due to the following two main (and often interrelated) factors: 1) They form close associations with a wide variety of microorganisms and 2) they are a rich source of biologically active secondary metabolites. This increasing research interest has greatly improved our knowledge of sponge-microbe interactions [7]. With the development of the phylogenetic
 (a)
(a)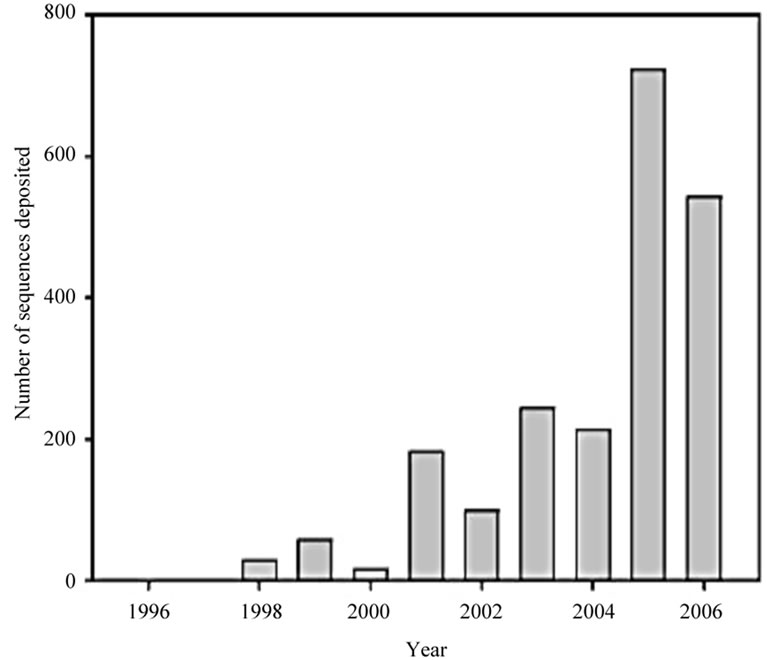 (b)
(b)
Figure 1. Increasing research interest in marine sponge-microorganism associations. (a) Number of publications retrieved from the ISI Web of Science database; (b) Number of sponge-derived 16S rRNA gene sequences deposited in GenBank per year (adopted from [7]).
analysis of 16S rRNA gene sequences, putative identification of bacteria by comparison with sequences from known closely related organisms can be easily achieved.
Sponges are filter-feeding, sessile multicellular organisms that live mainly in marine habitats. Because of their sessile lifestyle, they largely rely on chemical defense against potential predators. Thus, a great variety of novel biologically active metabolites have been isolated from sponges [8,9]. Some sponge-derived metabolites reveal striking similarity to known microbial metabolites, and it has been hypothesized that many natural products from marine invertebrates may be of microbial origin [10]. Microorganisms play a central role in sponge biology: they serve as food particles and are found to live associated with many sponges interand intracellularly [11,12]. The diversity of microbes known from sponges was categorized in 14 recognized bacterial phyla (and one candidate phylum), both major archaeal lineages, and assorted microbial eukaryotes. Members of bacterial phyla Acidobacteria, Actinobacteria, Bacteroidetes, Chloroflexi, Cyanobacteria, Deinococcus-Thermus, Firmicutes, Gemmatimonadetes, Nitrospira, Planctomycetes, Proteobacteria (Alpha, Beta, Delta, and Gammaproteobacteria), Spirochaetes, and Verrucomicrobia have been recovered from several marine sponges [13]. Several observations support the idea that bacteria synthesize sponge-specific compounds either completely or in the form of precursors completed subsequently by sponge metabolism [14-18]. Antimicrobial, cytotoxicity and other biological activities of associated bacteria may play a significant ecological role in sponge-bacteria associations. Furthermore, the isolation of sponge-associated bacteria producing bioactive metabolites, which were originally isolated from sponges, strongly supports the hypothesis of the microbial origin of the compounds formerly ascribed to sponges [19-21].
2. MATEIALS AND METHODS
2.1. Sponge Collection and Taxonomic Identification
Sponge sample was collected by SCUBA diving at depth 1.5 m in the Red Sea (El-Gona station; GPS: N: 27 22 39.98, E: 33 40 58.95) in May 2010 (Figure 2). The sponge sample was taxonomically identified as Amphimedon ochracea (Keller, 1889), Order Haplosclerida: Family Niphatidae (Figure 3). Taxonomic identification was performed by Prof. Rob. W. M. van Soest (University of Amsterdam, Netherlands). A voucher fragment is incorporated in the collections of the Zoological Museum of Amsterdam, now part of the Netherlands Centre for Biodiversity Naturalis (NCB Naturalis) at Leiden, The Netherlands, under registration number (ZMA Por. 22534). Another sponge sample was also deposited in the
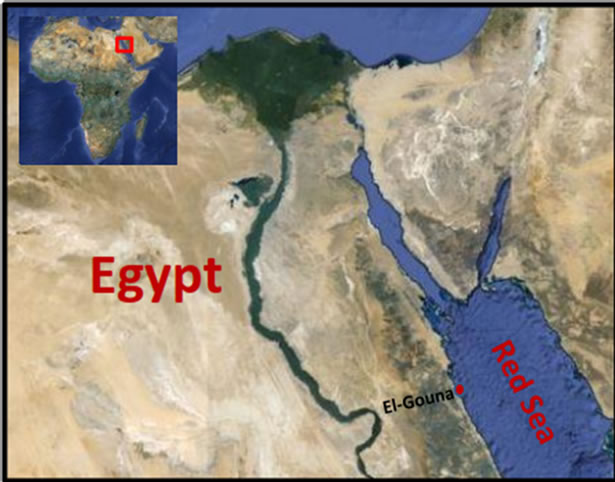
Figure 2. Egypt map showing the sponge sampling site (ElGouna, Egypt) on Red sea (site is indicated by red arrow).
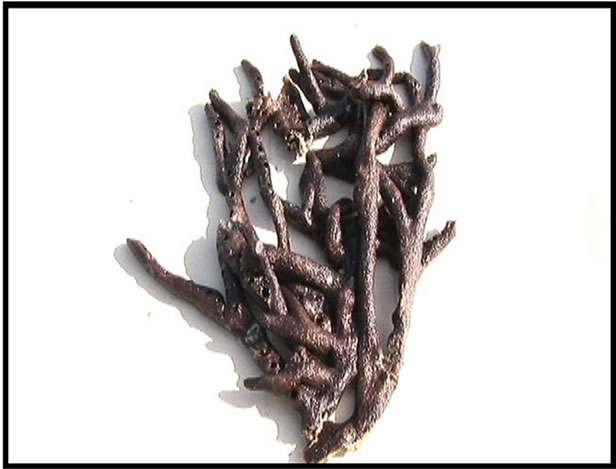

Figure 3. Sponge sample (Amphimedon ochracea).
Red Sea invertebrates’ collection at the Marine Biotechnology Unit; the National Institute of Oceanography and Fisheries, Alexandria.
2.2. Processing of the Sponge Sample
Sponge was transferred to plastic bags containing seawater and transported to the laboratory. Sponge specimen were rinsed in sterile seawater, cut into pieces with sterile scalpels of ca. 1 cm and then thoroughly homogenized in a sterile mortar with 10 volumes of sterile seawater. The supernatant was diluted in ten-fold series (10–1, 10–2, and 10–3) using autoclaved environmental water. Fifty microliter aliquots of each dilution were plated on various agar plates’ solid media.
2.3. Isolation of Sponge Associated Bacteria
Two growth media were prepared to isolate a wide range of the sponge associated bacteria; Marine Agar [22] and ISP medium 2 [23] served as general rich media to grow many heterotrophic marine bacteria. All media were supplemented with 0.2 µm pore size filtered cycloheximide (100 µg/mL), nystatin (25 µg/mL) and nalidixic acid (25 µg/mL). Cycloheximide and nystatin inhibit fungal growth, while nalidixic acid inhibits many fast-growing Gram-negative bacteria [24]. All media contained Difco Bacto agar (18 g/L) and were prepared in 1 L artificial sea water (NaCl 234.7 g, MgCl2·6H2O 106.4 g, Na2SO4 39.2 g, CaCl2 11.0 g, NaHCO3 1.92 g, KCl 6.64 g, KBr 0.96 g, H3BO3 0.26 g, SrCl2 0.24 g, NaF 0.03 g and ddH2O to 10.0 L) [25]. The inoculated plates were incubated at 28˚C for 5 - 10 days. Distinct colony morphotypes were picked and re-streaked until visually free of contaminants. Isolates were inoculated into liquid media ISP medium 2. The isolates were maintained on plates for short-term storage and long-term strain collections were set up in medium supplemented with 30% glycerol at −80˚C [26].
2.4. Extract Preparation and Cytotoxic Activity Screening
Bacterial isolates were cultured in 500 mL Erlenmeyer flasks containing 200 mL of ISP2 medium. The liquid cultures were grown for 7 - 14 days depending on their growth rate at 30˚C while shaking at 150 rpm. Crude extracts were prepared from whole cultures; containing cells and broths by ethyl acetate equal volume/three times, evaporated, lyophilized and dissolved in 1:1 v/v DMSO/H2O and kept at 4˚C until use for cytotoxic assay. The MTT (3-(4,5Dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide) assay which Measuring the mitochondrial reductase enzyme activity of viable cells that Could reduce MTT to formazan, giving a purple color, was used for screening the cytotoxicity of the bacterial extracts [27]. The cytotoxic activity of the microbial crude extracts was determined against three established cancer cell lines; MCF-7, HepG2 and HCT cells. Based on the values of the IC50, the most potent four bacterial extracts which exhibited the lowest IC50 were chosen for further study of their phylogenetic analysis and identification.
2.5. Molecular Identification and Phylogenetic Analysis
The isolated bacteria were first identified to the species level by PCR amplification of the 16S rRNA gene, BLAST analysis, and comparison with sequences in the GenBank nucleotide database. Specifically, the 16S rRNA gene from the strain was amplified using universal primers 27f (5’-AGAGTTTGATCCTGG CTCAG-3’) and 1492r (5’-GGTTACCTTGTTACGACTT-3’). The PCR conditions used were the same as those described previously [28]. The PCR products were purified and sequenced by the GATC-Biotech. Company (Tübingen, Germany). The sequences were compared with known sequences in the GenBank nucleotide database and the species level was identified as the nearest phylogenetic neighbor with 98% - 100% similarity [29]. Sequence alignment and phylogenetic analysis were performed using the Mega-5 software package [30]. Tree construction was conducted using neighbor joining algorithm (Jukes-Cantor correction) with bootstrap values based on 1000 replications. The 16S rRNA gene sequences of the isolates were deposited in GenBank under the accession Numbers indicated in brackets: HA-21 [JQ768238]; HA- 68 [JQ751264]; HA-MS-105 [JQ768239] and HAMS- 119 [JQ768240].
3. RESULTS AND DISCUSSION
Screening of Microbial Extracts That Show Cancer Cytotoxicity
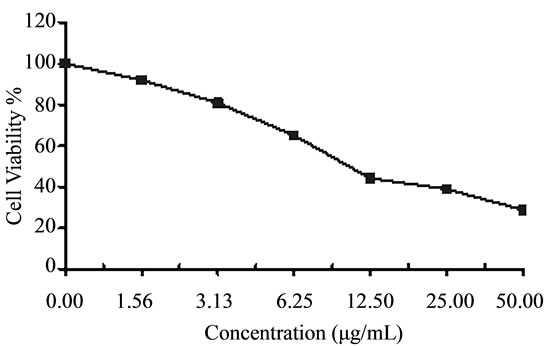
It has been estimated that over 99% of the marine sponge-associated microbes have yet to be cultured in the laboratory with bacteria isolated from the sponges containing diverse Bacillus species being one of the most divergent forms [31]. Crude extracts of four Bacillus sp. cultures isolated from the sponge Amphimedon ochracea were subjected to cytotoxicity screening against three established cancer cell lines; HepG2 (hepatocellular carcinoma), HCT (colon carcinoma) and MCF-7 (breast carcinoma). In vitro cytotoxicity assays are a potentially useful tool in the study of toxic compounds of complex mixtures. We found that the extracts obtained from the tested sponge associated bacteria were toxic against those carcinoma cells (Figures 4-6). The IC50 values of these extracts against cancer cell lines were in range of 4.3 - 46.9 µg/mL (Table 1). These values represent in all cases a comparable high toxicity to many other marine bacteria crude extracts. However, it was not possible to identify the active principle until now. These results suggest that the bacterial isolates HA-21, HA-68, HAMS-105 and HAMS-119 are good candidates for further activity-monitored fractionation to identify active principles.
All the bacteria isolated were identified using 16S ribosomal DNA sequences [32]. The tested isolates belong to the Bacillus sp. (firmicutes) (Figure 7). Interestingly, the firmicutes have been reported from diverse marine sponges irrespective of their taxonomic identity, geographic location, or natural products profile [33,34]. The isolates HA-68, HA-MS-105 and HAMS-119 revealed species-level similarity to Bacillus subtilis, Bacillus safensis and Lactobacillus murinus, respectively. However, the isolate HA-21 belonged to unidentified Bacillus sp. (Table 2).
 (a)
(a)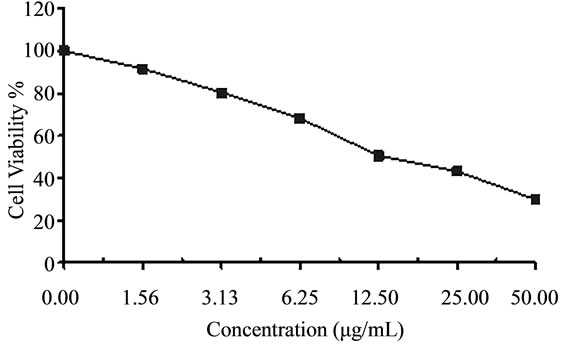 (b)
(b)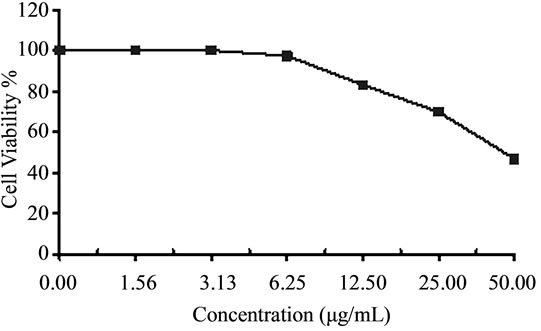 (c)
(c)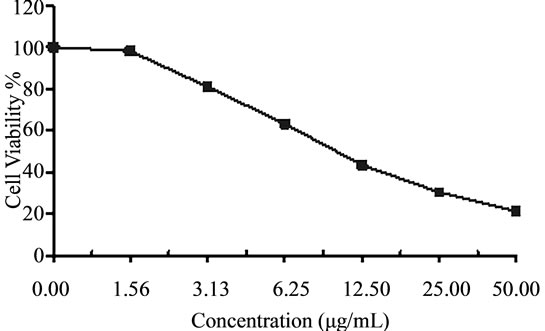 (d)
(d)
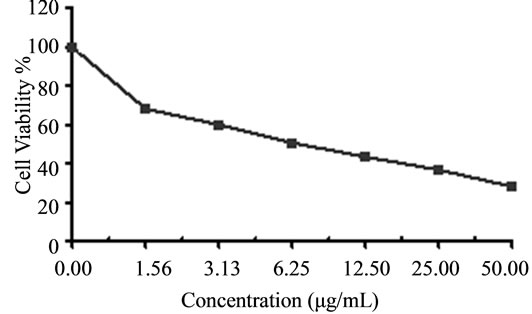
Figure 4. Cytotoxicity of (a) HAMS-119, (b) HA-21, (c) HA-MS-105 and (d) HA-68 bacterial extracts against Hepatocellular carcinoma cells (HepG-2) at 6 different concentrations. Cell viability was determined using the MTT bromide colorimetric assay.
 (a)
(a) (b)
(b)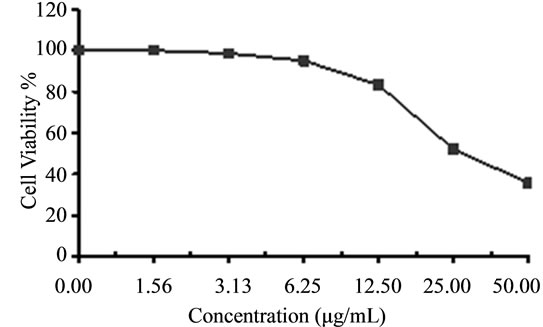 (c)
(c)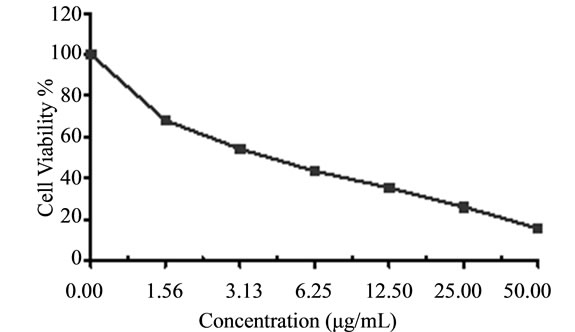 (d)
(d)
Figure 5. Cytotoxicity of (a) HAMS-119, (b) HA-21, (c) HA-MS-105 and (d) HA-68 bacterial extracts against colon carcinoma cells (HCT) at 6 different concentrations. Cell viability was determined using the MTT bromide colorimetric assay.
 (a)
(a)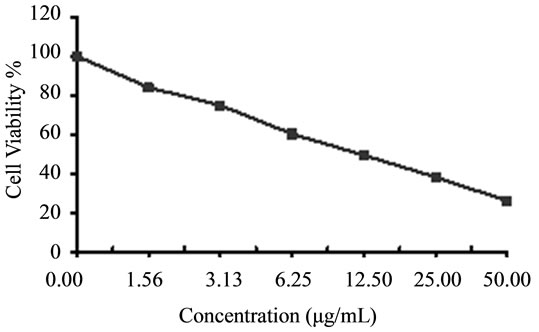 (b)
(b)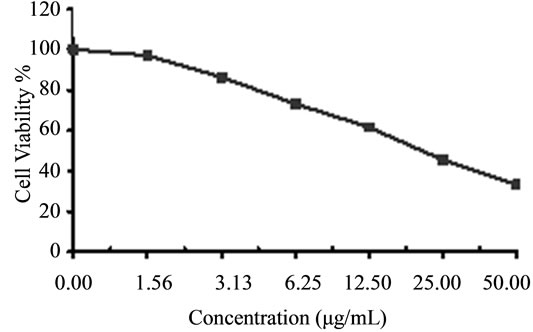 (c)
(c)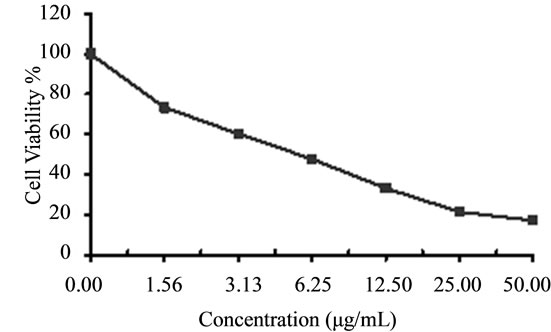 (d)
(d)
Figure 6. Cytotoxicity of (a) HAMS-119, (b) HA-21, (c) HA-MS-105 and (d) HA-68 bacterial extracts against breast carcinoma cells (MCF-7) at 6 different concentrations. Cell viability was determined using the MTT bromide colorimetric assay.
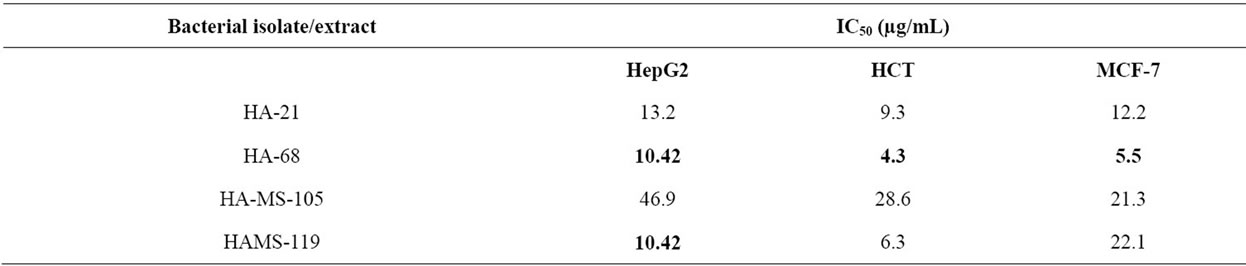
Table 1. The IC50 values of the bacterial extracts against HepG2, HCT and MCF-7 carcinoma cell lines. The lowest IC50 to each cancer type were represented in bold.
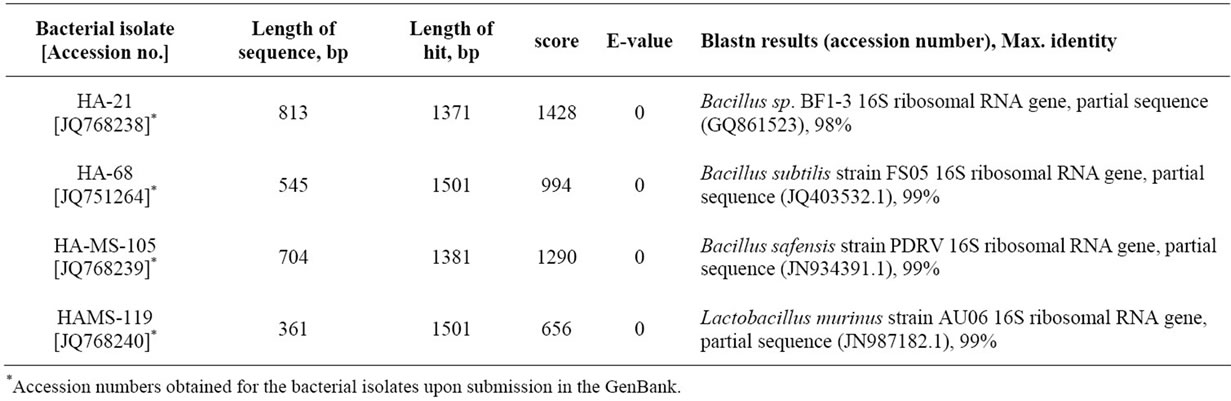
Table 2. Sequence analysis of the four tested bacterial isolates, using NCBI Blastn analysis tool, showing the maximum sequences similarity values of 98% - 100% to other strains in the NCBI database.
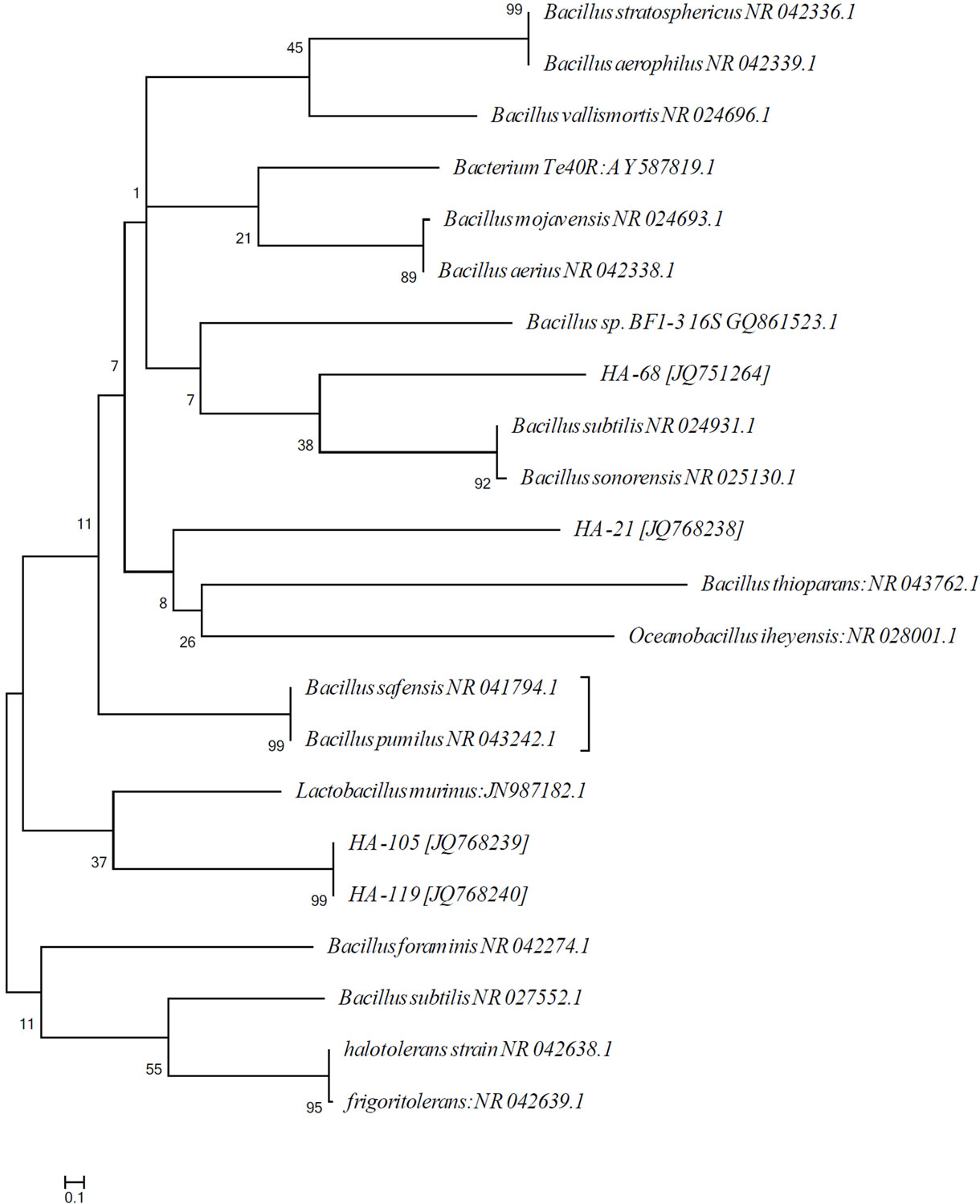
Figure 7. Molecular phylogenetic anaylsis by Maximum Likelihood method. The evolutionary history was inferred by using the Maximum Likelihood method based on the Tamura-Nei model [35,36]. The tree with the highest log likelihood (−1259.6498) is shown. The percentage of trees in which the associated taxa clustered together is shown next to the branches. Initial tree(s) for the heuristic search were obtained automatically as follows. When the number of common sites was <100 or less than one fourth of the total number of sites, the maximum parsimony method was used; otherwise BIONJ method with MCL distance matrix was used. The analysis involved 22 nucleotide sequences. Codon positions included were 1st + 2nd + 3rd + Noncoding. All positions containing gaps and missing data were eliminated. There were a total of 54 positions in the final dataset. The scale bar indicates 0.1 substitutions per nucleotide position.
4. CONCLUSION
The current study shows that bacterial isolates from the sponge (A. ochracea) possess bioactive properties. The tested isolates, HA-21, HA-68, HA-MS-105 and HAMS- 119 displayed potential cytotoxicity against three established cancer cell lines. Thus this investigation highlights the importance of bacteria associated with the sponge A. ochracea as a valuable resource for the discovery of novel bioactive molecules. Further chemical isolation and characterization of active compounds from these bacterial extracts is under investigation, and findings will be reported in due course.
5. ACKNOWLEDGEMENTS
We gratefully acknowledge Ms. Asmaa N. Ali and Mr. Mohamed Adel for the technical assistance in the laboratory. Financial support was provided by the National Institute of Oceanography and Fisheries (NIOF), Egypt.
![]()
![]()
REFERENCES
- Newman, D.J. and Cragg, G.M. (2004) Marine natural products and related compounds in clinical and advanced preclinical trials. Journal of Natural Products, 67, 1216- 1238. doi:10.1021/np040031y
- Simmons, T.L., Andrianasolo, E., McPhail, K., Flatt, P.M. and Gerwick, W.H. (2005) Marine natural products as anticancer drugs. Molecular Cancer Therapeutics, 4, 333- 342.
- Thoms, C. and Shrupp, P. (2005) Biotechnological potential of marine sponges and their associated bacteria as producers of new pharmaceuticals. Journal of International Biotechnology Law, 2, 257-264.
- Simmons, T.L., Coates, R.C., Clark, B.R., Eugene, N., Gonzalez, D., Esquenazi, E., et al. (2008) Biosynthetic origin of natural products isolated from marine organisms-invertebrate assemblages. Proceedings of the National Academy of Sciences of the United States of America, 105, 4587-4594. doi:10.1073/pnas.0709851105
- Livett, B.G., Gaylera, K.R. and Khalilb, Z. (2004) Drugs from the sea: Conopeptides as potential therapeutics. Current Medicinal Chemistry, 11, 1715-1723. doi:10.2174/0929867043364928
- Faulkner, D.J. (2000) Marine natural products. Natural Products Reports, 17, 7-55. doi:10.1039/a809395d
- Taylor, M.W., Radax, R., Steger, D. and Wagner, M. (2007) Sponge-associated microorganisms: Evolution, ecology, and biotechnological potential. Microbiology and Molecular Biology Reviews, 71, 295-347.
- Faulkner, D.J., Harper, M.K., Haygood, M.G., Salomon, C.E. and Schmidt, E.W. (2000) Symbiotic bacteria in sponges: Sources of bioactive substances. In: Fusetani, N., Ed., Drugs from the Sea, Karger, Basel, 107-119.
- Sipkema, D., Franssen, M.C.R., Osinga, R., Tramper, J. and Wijffels, R.H. (2005) Marine sponges as pharmacy. Marine Biotechnology, 7, 142-162. doi:10.1007/s10126-004-0405-5
- Faulkner, D.J. (2002) Marine natural products. Natural Products Reports, 19, 1-48. doi:10.1039/b009029h
- Friedrich, A.B., Merkert, H., Fendert, T., Hacker, J., Proksch, P. and Hentschel, U. (1999) Microbial diversity in the marine sponge Aplysina cavernicola (formerly Verongia cavernicola) analyzed by fluorescence in-situ hybridization (FISH). Marine Biology, 134, 461-470. doi:10.1007/s002270050562
- Vacelet, J. and Donadey, C. (1977) Electron microscope study of the association between some sponges and bacteria. Journal of Experimental Marine Biology and Ecolology, 30, 301-314. doi:10.1016/0022-0981(77)90038-7
- Taylor, M.W., Radax, R., Steger, D. and Wagner, M. (2007) Sponge-associated microorganisms: Evolution, ecology, and biotechnological potential. Microbiology and Molecular Biology Reviews, 71, 295-347. doi:10.1128/MMBR.00040-06
- Bewley, C.A., Holland, N.D. and Faulkner, D.J. (1996) Two classes of metabolites from Theonella swinhoei are localized in distinct populations of bacterial symbionts. Experientia, 52, 716-722. doi:10.1007/BF01925581
- Flowers, A.E., Garson, M.J., Webb, R.I., Dumdei, E.J. and Charan, R.D. (1998) Cellular origin of chlorinated diketopiperazines in the dictyoceratid sponge Dysidea herbacea (Keller). Cell and Tissue Research, 292, 597- 607doi:10.1007/s004410051089
- Proksch, P., Edrada, R.A. and Ebel, R. (2002) Drugs from the seas? Current status and microbiological implications. Applied Microbiology and Biotechnology, 59, 125-134. doi:10.1007/s00253-002-1006-8
- Schmidt, E.W., Obraztsova, A.Y., Davidson, S.K., Faulkner, D.J. and Haygood, M.G. (2000) Identification of the antifungal peptidecontaining symbiont of the marine sponge Theonella swinhoei as a novel δ-proteobacterium, BC and idatus entotheonella palauensis. Marine Biology, 136, 969- 977. doi:10.1007/s002270000273
- Unson, M.D., Holland, N.D. and Faulkner, D.J. (1994) A brominated secondary metabolite synthesized by the cyanobacterial symbiont of a marine sponge and accumulation of the crystalline metabolite in the sponge tissue. Marine Biology, 119, 1-11. doi:10.1007/BF00350100
- Oclarit, J.M., Okada, H., Ohta, S., Kaminura, K., Yamaoka, Y., Iizuka, T., Miyashiro, S. and Ikegami, S. (1994) Anti-bacillus substance in the marine sponge, Hyatella species produced by an associated Vibrio species bacterium. Microbios, 78, 7-16.
- Shigemori, H., Bae, M.A., Yazawa, K., Sasaki, T. and Kobayashi, J. (1992) Alteramide A, a new tetracyclic alkaloid from a bacterium Alteromonas sp. associated with the marine sponge Halichondria okadai. The Journal of Organic Chemistry, 57, 4317-4320. doi:10.1021/jo00041a053
- Stierle, A.C., Cardellina, J.H. and Singleton, F.L. (1988) A marine Micrococcus produces metabolites ascribed to the sponge Tedania ignis. Cellular and Molecular Life Sciences, 44, 1021. doi:10.1007/BF01939910
- Weiner, R.M., Segall, A.M. and Colwell, R.R. (1985) Characterization of a marine bacterium associated with Crassostrea virginica (the eastern oyster). Applied and Environmental Microbiology, 49, 83-90.
- Shirling, E.B. and Gottlieb, D. (1966) Methods for characterization of Streptomyces species. International Journal of Systematic Bacteriology, 16, 313-340. doi:10.1099/00207713-16-3-313
- Webster, N.S., Wilson, K.J., Blackall, L.L. and Hill, R.T. (2001) Phylogenetic diversity of bacteria associated with the marine sponge Rhopaloeides odorabile. Applied and Environmental Microbiology, 67, 434-444. doi:10.1128/AEM.67.1.434-444.2001
- Lyman, J. and Fleming, R. (1940) Composition of seawater. Journal of Marine Research, 3, 134-146.
- Sambrook, J., Fritsch, E.F. and Maniatis, T. (2001) Molecular cloning, a laboratory manual. Cold Spring Harbor Laboratory Press, New York.
- Acinas S.G., Anton J. and Rodriguez-Valera F. (1999) Diversity of free-living and attached bacteria in offshore western Mediterranean waters as depicted by analysis of genes encoding 16S rRNA. Applied and Environmental Microbiology, 65, 514-522.
- Thanomsub, B., Poomeechockchai, W., Limtrakul, A., Arunrattiyakorn, P., Petchleelaha, W., Nitida, T. and Kanzaki, H. (2006) Withdrawn: Chemical structures and biological activities of rhamnolipids produced by Pseudomonas aeruginosa B189 isolated from milk factory waste. Bioresource Technology, 98, 1149-1153. doi:10.1016/j.biortech.2005.10.045
- Hentschel, U., Schmid, M., Wagner, M., Fieseler, L., Gernert, C. and Hacker, J. (2001) Isolation and phylogenetic analysis of bacteria with antimicrobial activities from the mediterranean sponges Aplysina aerophoba and Aplysina cavernicola. FEMS Microbiology Ecology, 35, 305-312. doi:10.1111/j.1574-6941.2001.tb00816.x
- Tamura, K., Peterson, D., Peterson, N., Stecher, G., Nei, M. and Kumar, S. (2011) MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Molecular Biology and Evolution, 28, 2731-2739. doi:10.1093/molbev/msr121
- Kennedy, J., Baker, P., Piper, C., Cotter, P.D., Walsh, M., Mooij, M.J., Bourke, M.B., Rea, M.C., O’Connor, P.M., Ross, R.P., Hill, C., O’Gara, F., Marchesi, J.R. and Dobson, A.D. (2009) Isolation and analysis of bacteria with antimicrobial activities from the marine sponge Haliclona simulans collected from Irish waters. Marine Biotechnology, 11, 384-396. doi:10.1007/s10126-008-9154-1
- Thakur, N.L., Hentschel, U., Krasko, A., Pabel, C.T., Anil, A.C. and Müller, W.E.G. (2003) Antibacterial activity of the sponge Suberites domuncula and its primmorphs: Potential basis for epibacterial chemical defense. Aquatic Microbial Ecology, 31, 77-83. doi:10.3354/ame031077
- Hentschel, U., Schmid, M., Wagner, M., Fieseler, L., Gernert, C. and Hacker, J. (2001) Isolation and phylogenetic analysis of bacteria with antimicrobial activities from the Mediterranean sponges Aplysina aerophoba and Aplysina cavernicola. FEMS Microbiology Ecology, 35, 305-312. doi:10.1111/j.1574-6941.2001.tb00816.x
- Webster, N.S. and Hill, R.T. (2001) The culturable microbial community of the Great Barrier Reef sponge Rhopaloeides odorabile is dominated by an α-Proteobacterium. Marine Biology, 138, 843-851. doi:10.1007/s002270000503
- Tamura, K. and Nei, M. (1993) Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Molecular Biology and Evolution, 10, 512-526.
- Tamura, K., Peterson, D., Peterson, N., Stecher, G., Nei, M. and Kumar, S. (2011) MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Molecular Biology and Evolution, 28, 2731-2739.
NOTES
*Corresponding author.

