Food and Nutrition Sciences
Vol.4 No.6A(2013), Article ID:33008,7 pages DOI:10.4236/fns.2013.46A001
Genetic Engineering Peanut for Higher Droughtand Salt-Tolerance
![]()
1Department of Biological Sciences, Texas Tech University, Lubbock, USA; 2Zhejiang Academy of Agricultural Sciences, Hangzhou, China.
Email: *hong.zhang@ttu.edu
Copyright © 2013 Li Sun et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Received February 4th, 2013; revised March 21st, 2013; accepted April 5th, 2013
Keywords: Drought Tolerance; Peanut Transformation; Salt Resistance; Transgenic Plants; Yield Improvement
ABSTRACT
Peanut (Arachis hypogaea L.) is one of the major oilseed crops, mainly grown in tropical and sub-tropical regions of the world. It is also rich in proteins, vitamins and ions, therefore it constitutes an important portion of food nutrition for people in these regions. The production of peanut is being threatened by the changing environments as the major peanut producing counties such as China, India, and USA are facing severe water shortage for peanut irrigation. The yield and quality of peanut are negatively affected by drought and salinity. Making peanut more droughtand salt-tolerant will likely sustain peanut production in countries where water shortage or saline soil are already problems. Efforts were made to genetically engineer peanut for higher tolerance to drought and salt. Analysis of these transgenic peanut plants indicated that the agronomic traits such as peanut yields were the same between wild-type and transgenic peanut plants under normal growth conditions, yet the yields of transgenic peanut plants were much higher than wild-type peanut plant under reduced irrigation conditions. Other traits such as protein content and fatty acid compositions in the seeds of transgenic peanut plants were not altered under both normal and drought conditions, indicating that the genetic manipulation of peanut for stress tolerance did not affect chemical compositions of peanut seeds in transgenic peanut plants, only increased seed yields under stress conditions.
1. Introduction
Environmental stresses such as drought and salinity are major factors that limit peanut production in the world [1,2]. It was estimated that drought stress alone is responsible for over $500 million loss in peanut production annually [3]. The major peanut producing countries such as China, India, Nigeria, and USA are all facing severe water shortage for crop irrigation, therefore it is conceivable that the world peanut production will be severely affected in the future. The climate change prediction indicates that more extreme weathers, particularly drought conditions, will occur more often in the tropical and sub-tropical regions of the world [4-6], which will make peanut production extremely challenging in the future. As one of the most nutritious foods for oil and protein source, peanut is very popular among counties in Africa, Asia, and North and South America [7]. Reduced peanut production will affect prices of peanut and peanut derived products such as peanut butter, which will add hardships to people in many countries. Now the great challenge ahead is how to sustain peanut production, and perhaps even increase peanut production for our growing population at the time the condition for peanut production is deteriorating.
Traditional approach in breeding for droughtand salt-tolerant peanut has been slow, due to the rareness of alleles for the drought and salt tolerance in the exiting peanut germplasms [8]. Furthermore, even if a gene for drought or salt tolerance is found in a wild relative of peanut, it will be difficult to introgress the gene into the cultivated peanut cultivars due to the reproductive barrier, not to mention the long time required for many generations of back-crossing. With the advent of molecular biology, biotechnology approach offers a powerful alternative in creating droughtand salt-tolerant peanut in a more efficient way [9-11]. Indeed, over the last 20 years, many genes that confer drought tolerance and salt tolerance were introduced into various crops and tested in laboratories and field, and a few of those genes showed great promises for commercial release [11,12]. For example, the transcription factor genes such as the DREB/ CBF family have been widely used in improving stress tolerance in several crops [13,14]. This class of genes may be useful in creating heatand drought-tolerant peanut in the future (see the example below). In addition to the transcription factor genes, other types of genes might be useful in peanut improvement as well. These genes include those that encode enzymes of ABA biosynthesis or cytokinin biosynthesis pathways, antioxidation metabolisms, and stress signal transduction pathways [8,11]. Some of these functional genes might be useful in improving stress tolerance in peanut in the future.
2. Recent Progress in Peanut Biotechnology
2.1. Peanut Transformation Technologies
To improve peanut’s stress tolerance via biotechnology approach, we must be able to transform peanut. Earlier efforts in transforming peanut using Agrobacterium-mediated gene transfer or biolistic bombardment were largely successful [15-23]. However, the transformation efficiencies of these efforts were very low, ranging from 0.3% to less 10%. Sharma and Anjaiah [7] established a peanut transformation procedure that drastically increased peanut transformation efficiency to 55% or higher, which made peanut transformation relatively easy and highly reproducible. More importantly, this new procedure, although still using Agrobacterium as a vehicle, appears to be applicable to many ecotypes of peanut. We were able to use this method to transform both New Mexico Valencia and Texas Runner types at equal frequencies, i.e. ~50% [24-26].
2.2. Creation of Drought-Tolerant Peanut
The first report on engineering peanut for increased drought tolerance was from Sharma’s group in India [8]. They introduced an Arabidopsis transcription factor gene called AtDREB1A into a drought sensitive peanut line and analyzed transgenic peanut plants for transpiration efficiency under well watered and water-deficit conditions. The transcription factor AtDREB1A binds to the dehydration responsive element (DRE) that is located upstream of many stress responsive genes, thereby activates expression of DRE-containing genes under stressful conditions [27]. Because AtDREB1A was under the control of a stress responsive promoter, Bhatnagar-Mathur et al. [8] were able to obtain transgenic peanut plants that demonstrated similar phenotype to wild-type plants under well watered conditions, yet these transgenic plants showed higher transpiration efficiency under water-deficit conditions. Although no evidence was given in that paper, the AtDREB1A-trasngenic peanut plants are expected to be more drought-, salt-, and low temperature-tolerant, based on known information regarding other AtDREB1A-transgenic plants [28,29].
The second report in literature on genetically engineering peanut for improved drought tolerance was from our effort [24] in which we introduced a cytokinin biosynthetic gene IPT into peanut. The IPT gene encodes isopentenyltransferase, an enzyme that plays a critical role in a rate-limiting step of cytokinin biosynthesis [30]. Drought stress typically causes early senescence in plants, which is advantageous for plants to survive under severe drought conditions in nature as they can quickly finish their life cycle. But this trait is bad for crops, as the yield is severely compromised. If senescence can be delayed under drought conditions, the yield penalty could be reduced. Richmond and Lang [31] demonstrated that exogenous application of cytokinin could delay senescence in plants, and this concept was further proved by later studies that increased cytokinin production by expressing an IPT gene in transgenic plants could indeed delay senescence [32-34]. However, the delayed senescence might not lead to increased seed yield as timing and place of IPT expression in transgenic plants are also important for reducing the yield penalty under drought conditions. Rivero et al. [35] provided an exciting example that regulated expression of IPT by a water-deficit inducible promoter PSARK could significantly reduce the yield penalty caused by drought in transgenic tobacco plants. This strategy worked very well for rice [36], as well as for peanut [24]. The PSARK::IPT-transgenic peanuts plants performed much better than wild-type plants under reduced irrigation conditions in greenhouse, growth chamber, and field conditions. For example, they produced much larger root systems under reduced irrigation in greenhouse conditions, which allowed them to use water more efficiently [24]. They maintained higher photosynthetic rates and stomatal conductance, produced significantly more biomass under reduced irrigation conditions in greenhouse and field conditions (Figures 1(A) and (B)). The yields of PSARK::IPT-transgenic peanuts plants were 30% - 35% higher than that of wild-type plants based on two years of field data [24]. Furthermore, PSARK::IPT-transgenic peanuts plants appeared to produce larger and more multipleseed pods than wild-type peanut plants do (Figure 2).
2.3. Creation of Salt-Tolerant Peanut
As a glycophytic plant, peanut growth is very sensitive to salt, therefore making peanut salt tolerant will improve peanut production in many parts of the world where soils are saline. In 1999, Blumwald’s group demonstrated that overexpression of AtNHX1 that encodes the vacuolar membrane-bound sodium/proton (Na+/H+) antiporter in
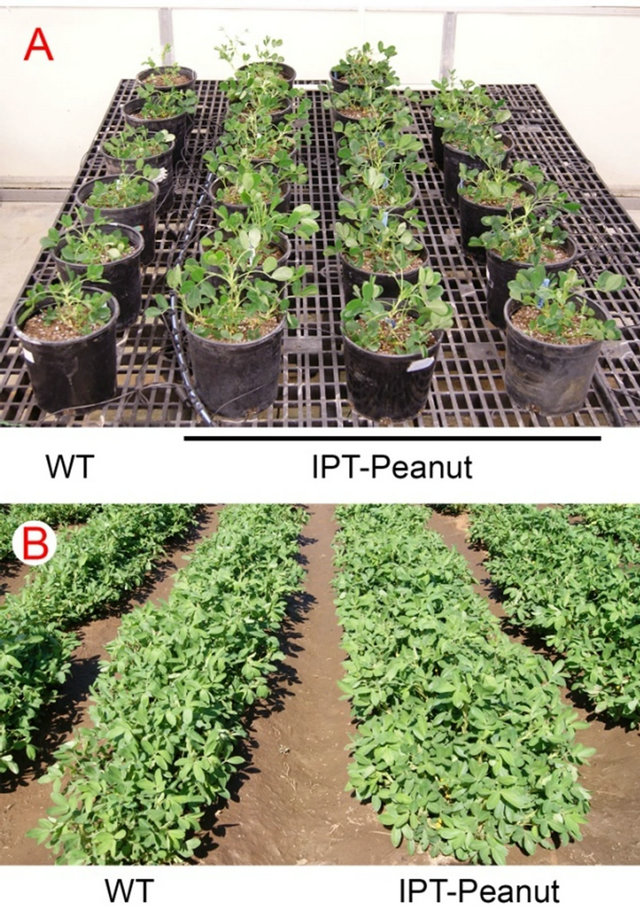

Figure 1. Wild-type and IPT-transgenic peanut plants in greenhouse (A) and the field (B) under reduced irrigation conditions.
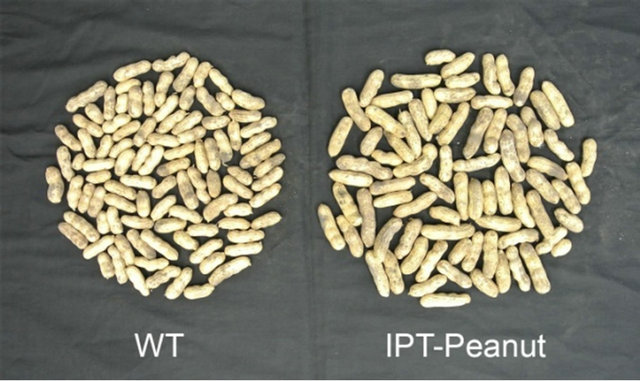
Figure 2. The phenotype of seeds from wild-type and IPTtransgenic peanut plants grown under reduced irrigation conditions.
Arabidopsis could improve salt tolerance in transgenic plants [37]. The increased Na+/H+ antiporter activity could lead to increased Na+ sequestration into vacuole, which reduces Na+ toxicity in cytoplasm and at the same time reducing water potential in the vacuole, leading to increased salt tolerance in AtNHX1-overexpressing plants [38]. This approach was successfully used to increase salt tolerance in other plant species such as tomato [39], rapeseed [40], cotton [41], and soybean [42]. We introduced AtNHX1 into peanut and demonstrated that the salt tolerance of peanut could also be improved [26]. AtNHX1- overexpressing peanut plants could tolerate up to 150 mM NaCl in soil, produce higher amounts of biomass, and maintain higher photosynthetic rates and stomatal conductance during salt treatment [26]. Our greenhouse experiments indicated that overexpression of AtNHX1 could confer increased salt tolerance in peanut, however, we have not tested how these AtNHX1-overexpressing peanut plants would behave under field conditions.
2.4. Creation of Droughtand Salt-Tolerant Peanut
Recently, we demonstrated that by overexpressing an Arabidopsis vacuolar pyrophosphatase gene AVP1, we could increase drought and salt tolerance simultaneously in transgenic peanut plants [25]. The vacuolar pyrophosphatase is a proton pump that generates proton chemical gradient across vacuolar membranes. Many secondary transporters such as the Na+/H+ antipoter (i.e. AtNHX1) depend on the proton chemical gradient generated by proton pump like AVP1 for activities [38,43]. Increasing proton pump activities could increase Na+/H+-antipoter activities, thereby increasing salt tolerance [43]. Gaxiola’s group demonstrated that overexpression of AVP1 in transgenic plants could increase both droughtand salt-tolerance [44,45]. The increased drought tolerance in the AVP1- overexpresing plants was due to robust root development, which is caused by increased auxin polar transport in transgenic plants [46]. Since the pioneering work by Gaxiola et al. [44], AVP1 and its homologs have been overexpressed in many plants, and in all cases, increased droughtand salt-tolerance have been observed [45,47- 51]. AVP1-overexpressing peanut plants were both droughtand salt-tolerant in greenhouse conditions, as they produced higher amounts of biomass, maintained higher photosynthetic rates and transpiration rates under reduced irrigation and saline conditions [25]. These peanut plants also produced higher yields under reduced irrigation conditions in two-years of field trials [25].
Even though the seed pods from IPT-transgenic peanut appear larger than those from wild-type peanut plants (Figure 2), there were no major changes in the oil contents between IPT-transgenic peanut seeds and wild-type peanut seeds [24]. Similarly, there were no major changes in the oil contents between AVP1-transgenic and wildtype peanut seeds as well [25]. The contents of major fatty acids such as palmitic acid (C16:0), oleic acid (C18:1) and linoleic acid (C18:2) were similar between wild-type and IPT-transgenic and AVP1-transgenic peanut plants. The minor fatty acids varied a little between transgenic peanut plants and wild-type plants, but these differences were also found between wild-type and the segregated non-transgenic lines [24,25], suggesting that they might be due to somatic variation resulting from tissue cultures. Furthermore, these differences are likely found among different peanut ecotypes, which show little nutritional differences. It is apparent that overexpression of AVP1 and regulated expression of IPT in peanut do not affect the oil content and major fatty acid compositions. Whether or not PSARK::IPT-transgenic peanut and AVP1-expressing peanut plants would increase peanut yield under field conditions in large-scale trials is not known, but the data we have from laboratory experiments as well as small-scale field experiments are very promising, which strongly suggest that IPT and AVP1 are excellent target genes that could improve peanut production in areas where water is limited.
3. Future Challenges
3.1. Improving Peanut Transformation Efficiency
Although the procedure developed by Sharma and Anjaiah [7] is quite good in transforming peanut, the false positive rate of over 40% among the regenerated peanut lines is still too high, as sorting out false positives is time-consuming and resource-wasting. Kanamycin was used as the selection marker in transformation, but peanut appears to have some resistance to kanamycin, which leads to a large numbers of escapers in the regeneration process. Because of the resistance to kanamycin, transgenic peanut cannot be screened directly at seedling stage in kanamycin media, instead transgenic peanut plants have to be screened by DNA blot, PCR or RNA blot analysis, which makes characterization of transgenic peanut plants labor-intensive and time-consuming. Therefore it is necessary to develop a more efficient system for peanut transformation, perhaps a better antibiotics that is more effective in killing non-transgenic peanut plants needs to be identified as the selective marker.
3.2. Finding More Efficient Genes for Peanut Improvement
The effect of expressing AtDREB1A in peanut under the control of a stress responsive promoter on peanut yield in field conditions has yet to be reported, it is difficult to estimate the impact of this gene on improving peanut production in the semi-arid regions of the world. Based on our own experiences on working with IPT-transgenic peanut and AVP1-transgenic peanut, we have the following observations: IPT-transgenic peanut plants consistently produced higher yields than AVP1-trasngenic plants, and both consistently outperformed wild-type peanut plants (on averages 30% and 20% better than wild-type plants, respectively) under reduced irrigation conditions in the semi-arid regions of West Texas [24,25]. However, IPTtransgenic peanut plants are not salt-tolerant, yet AVP1- transgenic peanut plants are. Consequently, each has its own strength and weakness. Other genes that prove to be effective in conferring drought tolerance in other transgenic crops have not been tested in peanut, their potential use in peanut might be worth of trying. For example, the SNAC1 gene from rice was shown to confer 23% - 34% higher yields when overexpressed in transgenic rice plants under drought conditions [52]. The yield differences between SNAC1-transgenic rice and wild-type rice are similar to the differences found between IPT-transgenic peanut and wild-type peanut plants under water-deficit conditions. Recently a gene named OsSIZ1 was shown to significantly increase both droughtand heat-tolerance when overexpressed in transgenic creeping bentgrass [53], and similar observation was made in transgenic cotton plants expressing the OsSIZ1 transgene (our unpublished data), suggesting that OsSIZ1 would be an excellent gene that could be used for improving peanut. There might be more genes out there that confer similar or higher tolerance to drought stress, we need to find an efficient way to test them in transgenic peanut plants in the future.
3.3. GMO Issues
Perhaps the biggest challenge that we face today in plant biotechnology is not a technical issue, but a political or philosophic issue: should genetically modified organisms (GMO) or genetic engineering products such as transgenic peanut plants be developed and marketed? One of the major obstacles for the adoption of GMO in Europe and many other countries is the perception that GMO is not safe. Although the safety issue has been carefully addressed for each GMO product that is approved for commercial release, there is still strong opposition against GMO products. We have to admit that we did not do a good job in educating general public about the nature of changes in each GMO product, which led to misunderstanding of the GMO products by the general public. There are a number of GMO products on market and these GMO products contain either herbicide tolerance gene or insect resistance gene, which were inserted into crops either as a single trait or stacked traits [54]. The benefits of having crops with these genes far outweigh the potential harm that these crops might bring, in fact after nearly 20 years of growing crops with these traits and more than 134 million hectares of transgenic crops grown by 2009 [55], there was not a single report on the adverse impact of GMO on food safety or environmental damage [56]. In US, three governmental agencies oversee the testing and approval of genetically engineered products: the US Department of Agriculture (USDA), Food and Drug Administration (FDA), and Environmental Protection Agency (EPA) [56]. USDA has strict regulations for testing all GMOs before their release, and is responsible for protecting US agriculture and environment from potential pests. FDA is responsible for ensuring the safety of GMO as human’s food or animal feed, and EPA is responsible for ensuring that genetically engineered pesticides pose no threats to human and our environment [56]. All GMO products on the market had been thoroughly tested and are continuingly to be carefully scrutinized. If any potential harm to human health or to our environment is found, they will be removed from market immediately. The benefits of using GMOs have been enormous: reduction of 17% pesticides usage from 1996 to 2007 in the US alone (about 359,000 tons of chemicals were not applied in the US soil) and over $19 billion more revenue generated for US farmers during this period [54].
Another misperception about GMO is that man should not change the genetic make-up of crops, as crop plants have evolved for so long to reach today’s state, there must be a reason why crops should be the way they are. People who think this way do not realize that in fact there were no crops that would look like what they are today even just 100 years ago. All crop plants we have today were derived from many years’ selection and breeding by people, the crops’ genomes had changed dramatically over the last century. The yields of crops that existed several thousand years ago were extremely low, usually less than 5% of today’s yields. For example, corn’s wild relatives today produce ears that are about 2 cm long with tiny kernels. Furthermore, the wild relatives of corn or rice not only have low yields, but low quality as well. They usually taste bad, with low nutritional values, and many even have high levels of allergens. Over the years farmers had selected the best varieties to breed, and especially since the discovery of Mendelian’s rules of genetics in early 1910s, scientists had bred crops with the best agronomic traits and created modern varieties of today’s crops. People never questioned crops created by traditional breeding methods in the last 100 years, but in fact traditional breeding involves manipulating hundreds, perhaps thousands of genes during crossing between different parental lines or lines from exotic germplasms. Whereas, GMO products usually involve one or a few genes whose biochemistry has been studied extensively, and whose potential being allergens or toxicants is very small, if existent [56]. In summary, given the strict regulations from government and thorough studies from academia, the GMO resulted from biotechnology should be safer, not the other way around [57,58]. We need to pass this information to general public as one of our missions, if we want to continue to succeed in creating better and safer biotechnology products in the future.
4. Acknowledgements
Our research was supported by grants from Texas Peanut Producers Board, US National Peanut Board, and USDAOgallala Aquifer Program.
REFERENCES
- J. R. Stansell and J. E. Pallas, “Yield and Quality Response of Florunner Peanut to Applied Drought at Several Growth Stages,” Peanut Science, Vol. 12, No. 2, 1985, pp. 64-70. doi:10.3146/pnut.12.2.0005
- M. C. Lamb, J. I. Davidson, J. W. Childre and N. R. Martin, “Comparison of Peanut Yield, Quality, and Net Returns between Nonirrigated and Irrigated Production,” Peanut Science, Vol. 24, No. 2, 1997, pp. 97-101. doi:10.3146/i0095-3679-24-2-7
- K. K. Sharma and M. Lavanya, “Recent Developments in Transgenics for Abiotic Stress in Legumes of the SemiArid Tropics,” In: M. Ivanaga, Ed., Genetic Engineering of Crop Plants for Abiotic Stress, JIRCAS Working Report No. 23, JIRCAS, Tsukuba, 2002, pp. 61-73.
- D. B. Lobell, M. B. Burke, C. Tebaldi, M. D. Mastrandrea, W. P. Falcon and R. L. Naylor, “Prioritizing Climate Change Adaptation Needs for Food Security in 2030,” Science, Vol. 319, No. 5863, 2008, pp. 607-610. doi:10.1126/science.1152339
- D. S. Battisti and R. L. Naylor, “Historical Warnings of Future Food Insecurity with Unprecedented Seasonal Heat,” Science, Vol. 323, No. 5911, 2009, pp. 240-244. doi:10.1126/science.1164363
- S. P. Long and D. R. Ort, “More than Taking the Heat: Crops and Global Change,” Current Opinion in Plant Biology, Vol. 13, No. 3, 2010, pp. 241-248. doi:10.1016/j.pbi.2010.04.008
- K. Sharma and V. Anjaiah, “An Efficient Method for the Production of Transgenic Plants of Peanut (Arachis hypogaea L.) through Agrobacterium tumefaciens-Mediated genetic Transformation,” Plant Science, Vol. 159, No. 1, 2000, pp. 7-19. doi:10.1016/S0168-9452(00)00294-6
- P. Bhatnagar-Mathur, et al., “Stress-Inducible Expression of AtDREB1A in Transgenic Peanut (Arachis hypogaea L.) Increases Transpiration Efficiency under Water-Limiting Conditions,” Plant Cell Reports, Vol. 26, No. 12, 2007, pp. 2071-2082. doi:10.1007/s00299-007-0406-8
- R. Mittler and E. Blumwald, “Genetic Engineering for Modern Agriculture: Challenges and Perspectives,” Annual Review of Plant Biology, Vol. 61, 2010, pp. 443-462. doi:10.1146/annurev-arplant-042809-112116
- R. K. Varshney, K. C. Bansal, P. K. Aggarwal, S. K. Datta and P. Q. Craufurd, “Agricultural Biotechnology for Crop Improvement in a Variable Climate: Hope or Hype?” Trends in Plant Science, Vol. 16, No. 7, 2011, pp. 363- 371. doi:10.1016/j.tplants.2011.03.004
- J. Deikman, M. Petracek and J. E. Heard, “Drought Tolerance through Biotechnology: Improving Translation from the Laboratory to Farmers’ Fields,” Current Opinion in Biotechnology, Vol. 23, No. 2, 2012, pp. 243-250. doi:10.1016/j.copbio.2011.11.003
- S. Yang, B. Vanderbeld, J. Wang and Y. Huang, “Narrowing Down the Targets: Towards Successful Genetic Engineering of Drought-Tolerant Crops,” Molecular Plant, Vol. 3, No. 3, 2010, pp. 469-490. doi:10.1093/mp/ssq016
- K. Century, T. L. Reuber and O. J. Ratcliffe, “Regulating the Regulators: The Future Prospects for TranscriptionFactor-Based Agricultural Biotechnology Products,” Plant Physiology, Vol. 147, No. 1, 2008, pp. 20-29. doi:10.1104/pp.108.117887
- D. Todaka, N. Kazuo, S. Kazuo and Y.-S. Kazuko, “Toward Understanding Transcriptional Regulatory Networks in Abiotic Stress Responses and Tolerance in Rice,” Abiotic Stress in Rice, Vol. 5, 2012, p. 6. http://www.thericejournal.com/content/5/1/6
- P. Ozias-Akins, et al., “Regeneration of Transgenic Peanut Plants from Stably Transformed Embryogenic Callus,” Plant Science, Vol. 93, No. 1-2, 1993, pp. 185-194. doi:10.1016/0168-9452(93)90048-5
- M. Cheng, D. C. H. His and G. C. Phillips, “Recovery of Transformants of Valencia-Type Peanut Using Agrobacterium tumefaciens,” Peanut Science, Vol. 21, No. 2, 1994, pp. 84-88. doi:10.3146/i0095-3679-21-2-3
- M. Cheng, R. L. Jarret, Z. Li, A. Xing and J. W. Demski, “Production of Fertile Transgenic Peanut (Arachis hypogaea L.) Plants Using Agrobacterium tumefaciens,” Plant Cell Reports, Vol. 15, No. 9, 1996, pp. 653-657. doi:10.1007/BF00231918
- S. Eapen and L. George, “Agrobacterium tumefaciens Mediated Gene Transfer in Peanut (Arachis hypogaea L.),” Plant Cell Reports, Vol. 13, No. 10, 1994, pp. 582-586. doi:10.1007/BF00234516
- D. M. Livingstone and R. G. Birch, “Plant Regeneration and Microprojectile-Mediated Gene Transfer in Embryonic Leaflets of Peanut (Arachis hypogaea L.),” Australian Journal of Plant Physiology, Vol. 22, No. 4, 1995, pp. 585-591. doi:10.1071/PP9950585
- C. Singsit, et al., “Expression of a Bacillus Thuringiensis cryIA(c) Gene in Transgenic Peanut Plants and Its Efficiency against Lesser Cornstalk Borer,” Transgenic Research, Vol. 6, No. 2, 1997, pp. 169-176. doi:10.1023/A:1018481805928
- H. Yang, C. Singsit, A. Wang, D. Gonsalves and P. OziasAkins, “Transgenic Peanut Plants Containing a Nucleocapsid Protein Gene of Tomato Spotted Wilt Virus Show Divergent Levels of Gene Expression,” Plant Cell Reports, Vol. 17, No. 9, 1998, pp. 693-699. doi:10.1007/s002990050467
- A. H. McKently, G. A. Moore, H. Doostdar and R. P. Niedz, “Agrobacterium-Mediated Transformation of Peanut (Arachis hypogaea L.) Embryo Axes and the Development of Transgenic Plants,” Plant Cell Reports, Vol. 14, No. 11, 1995, pp. 699-703. doi:10.1007/BF00232650
- V. K. Rohini and K. S. Rao, “Transformation of Peanut (Arachis hypogaea L.): A Non-Tissue Culture Based Approach for Generating Transgenic Plants,” Plant Science, Vol. 150, No. 1, 2000, pp. 41-49. doi:10.1016/S0168-9452(99)00160-0
- H. Qin, et al., “Regulated Expression of an Isopentenyltransferase Gene (IPT) in Peanut Significantly Improves Drought Tolerance and Increases Yield under Field Conditions,” Plant and Cell Physiology, Vol. 52, No. 11, 2011, pp. 1904-1914. doi:10.1093/pcp/pcr125
- H. Qin, et al., “Expression of the Arabidopsis Vacuolar H+-Pyrophosphatase Gene AVP1 in Peanut to Improve Drought and Salt Tolerance,” Plant Biotechnology Reports, 2012. doi:10.1007/s11816-012-0269-5
- M. Banjara, L. Zhu, G. Shen, P. Payton and H. Zhang, “Expression of an Arabidopsis Sodium/Proton Antiporter Gene (AtNHX1) in Peanut to Improve Salt Tolerance,” Plant Biotechnology Report, Vol. 6, 2011, pp. 59-67.
- K. Yamaguchi-Shinozaki and K. Shinozaki, “Organizantion of Cis-Acting Regulatory Elements in Osmoticand Cold-Stress-Responsive Promoters,” Trends in Plant Science, Vol. 10, 2005, pp. 88-94. doi:10.1016/j.tplants.2004.12.012
- Y. Ito, et al., “Functional Analysis of Rice DREB1/CBFType Transcription Factors Involved in Coldresponsive Gene Expression in Transgenic Rice,” Plant Biotechnology Reports, Vol. 47, No. 1, 2006, pp. 141-153. doi:10.1093/pcp/pci230
- K. Datta, N. Baisakh, M. Ganguly, S. Krishnan, K. Yamaguchi-Shinozaki and S. K. Datta, “Overexpression of Arabidopsis and Rice Stress Genes’ Inducible Transcription Factor Confers Drought and Salinity Tolerance to Rice,” Plant Biotechnology Journal, Vol. 10, No. 5, 2012, pp. 579-586. doi:10.1111/j.1467-7652.2012.00688.x
- H. Sakakibara, “CYTOKININS: Activity, Biosynthesis, and Translocation,” Annual Review of Plant Biology, Vol. 57, 2006, pp. 431-449. doi:10.1146/annurev.arplant.57.032905.105231
- A. E. Richmond and A. Lang, “Effect of Kinetin on Protein Content and Survival of Detached Xanthium Leaves,” Science, Vol. 125, No. 3249, 1957, pp. 650-651. doi:10.1126/science.125.3249.650-a
- C. M. Smart, S. R. Scofield, M. W. Bevan and T. A. Dyer, “Delayed Leaf Senescence in Tobacco Plants Transformed with tmr, a Gene for Cytokinin Production in Agrobacterium,” Plant Cell, Vol. 3, 1991, pp. 647-656.
- B. Martineau, C. M. Houck, R. E. Sheehy and W. R. Hiatt, “Fruit-Specific Expression of the A. tumefaciens Isopentenyl Transferase Gene in Tomato: Effects on Fruit Ripening and Defense-Related Gene Expression in Leaves,” Plant Journal, Vol. 5, No. 1, 1994, pp. 11-19. doi:10.1046/j.1365-313X.1994.5010011.x
- S. Gan and R. M. Amasino, “Inhibition of Leaf Senescence by Autoregulated Production of Cytokinin,” Science, Vol. 270, No. 5244, 1995, pp. 1986-1988. doi:10.1126/science.270.5244.1986
- R. M. Rivero, et al., “Delayed Leaf Senescence Induces Extreme Drought Tolerance in a Flowering Plant,” Proceedings of National Academy of Sciences of the United States of America, Vol. 104, No. 49, 2007, pp. 19631- 19636. doi:10.1073/pnas.0709453104
- Z. Peleg, M. Reguera, H. Walia and E. Blumwald, “Cytokinin Mediated Source-Sink Modifications Improve Drought Tolerance and Increases Grain Yield in Rice under Water Stress,” Plant Biotechnology Journal, Vol. 9, No. 7, 2011, pp. 747-758. doi:10.1111/j.1467-7652.2010.00584.x
- M. P. Apse, G. S. Aharon, W. A. Snedden and E. Blumwald, “Salt Tolerance Conferred by Overexpression of a Vacuolar Na+/H+ Antiport in Arabidopsis,” Science, Vol. 285, No. 5431, 1999, pp. 1256-1258. doi:10.1126/science.285.5431.1256
- E. Blumwald, “Salt Transport and Salt Resistance in Plants and Other Organisms,” Current Opinion in Cell Biology, Vol. 12, No. 4, 2000, pp. 431-434. doi:10.1016/S0955-0674(00)00112-5
- H. X. Zhang and E. Blumwald, “Transgenic Salt-Tolerant Tomato Plants Accumulate Salt in Foliage but Not in Fruit,” Nature Biotechnology, Vol. 19, 2001, pp. 765-768. doi:10.1038/90824
- H. X. Zhang, J. N. Hodson, J. P. Williams and E. Blumwald, “Engineering Salt-Tolerant Brassica Plants: Characterization of Yield and Seed Oil Quality in Transgenic Plants with Increased Vacuolar Sodium Accumulation,” Proceedings of National Academy of Sciences of the United States of America, Vol. 98, No. 22, 2001, pp. 12832- 12836. doi:10.1073/pnas.231476498
- C. He, et al., “Expression of an Arabidopsis Vacuolar Sodium/Proton Antiporter Gene in Cotton Improves Photosynthetic Performance under Salt Conditions and Increases Fiber Yield in the Field,” Plant Biotechnology Reports, Vol. 46, No. 11, 2005, pp. 1848-1854. doi:10.1093/pcp/pci201
- T. X. Li, et al., “Stable Expression of Arabidopsis Vacuolar Na+/H+ Antiporter Gene AtNHX1, and Salt Tolerance in Transgenic Soybean for over Six Generations,” Chinese Science Bulletin., Vol. 55, No. 12, 2010, pp. 1127- 1134. doi:10.1007/s11434-010-0092-8
- R. A. Gaxiola, G. R. Fink and K. D. Hirschi, “Genetic Manipulation of Vacuolar Proton Pumps and Transporters,” Plant Physiology, Vol. 129, No. 3, 2002, pp. 967- 973. doi:10.1104/pp.020009
- R. A. Gaxiola, et al., “Droughtand Salt-Tolerant Plants Result from Overexpression of the AVP1 H+-Pump,” Proceedings of National Academy of Sciences of the United States of America, Vol. 98, No. 20, 2001, pp. 11444- 11449. doi:10.1073/pnas.191389398
- S. Park, et al., “Up-Regulation of a H+-Pyrophosphatase (H+-PPase) as a Strategy to Engineer Drought-Resistant Crop Plants,” Proceedings of National Academy of Sciences of the United States of America, Vol. 102, No. 52, 2005, pp. 18830-18835. doi:10.1073/pnas.0509512102
- J. Li, et al., “Arabidopsis H+-PPase AVP1 Regulates AuxinMediated Organ Development,” Science, Vol. 310, No. 5745, 2005, pp. 121-125. doi:10.1126/science.1115711
- F. Y. Zhao, X. J. Zhang, P. H. Li, Y. X. Zhao and H. Zhang, “Co-Expression of the Suaeda salsa SsNHX1 and Arabidopsis AVP1 Confer Greater Salt Tolerance to Transgenic Rice than the Single SsNHX1,” Molecular Breeding, Vol. 17, No. 4, 2006, pp. 341-353. doi:10.1007/s11032-006-9005-6
- B. Li, A. Wei, C. Song, N. Li and J. Zhang, “Heterologous Expression of the TsVP Gene Improves the Drought Resistance of Maize,” Plant Biotechnology Journal, Vol. 6, No. 2, 2008, pp. 146-159. doi:10.1111/j.1467-7652.2007.00301.x
- S. Lv, K. Zhang, Q. Gao, L. Lian, Y. Song and J.-R. Zhang, “Overexpression of an H+-PPase from Thellungiella halophila in Cotton Enhances Salt Tolerance and Improves Growth and Photosynthetic Performance,” Plant Biotechnology Reports, Vol. 49, No. 8, 2008, pp. 1150-1164. doi:10.1093/pcp/pcn090
- S. Lv, L.-J. Lian, P. L. Tao, Z.-X. Li, K.-W. Zhang and J.-R. Zhang, “Overexpression of Thellungiella halophila H+-PPase (TsVP) in Cotton Enhances Drought Stress Resistance of Plants,” Planta, Vol. 229, No. 4, 2009, pp. 899-910. doi:10.1007/s00425-008-0880-4
- V. Pasapula, et al., “Expression of an Arabidopsis Vacuolar H+-Pyrophosphatase Gene (AVP1) in Cotton Improves Droughtand Salt-Tolerance and Increases Fiber Yield in the Field Conditions,” Plant Biotechnology Journal, Vol. 9, No. 1, 2011, pp. 88-99. doi:10.1111/j.1467-7652.2010.00535.x
- H. Hu, et al., “Overexpressing a NAM, ATAF, and CUC (NAC) Transcription Factor Enhances Drought Resistance and Salt Tolerance in Rice,” Proceedings of National Academy of Sciences of the United States of America, Vol. 103, No. 35, 2006, pp. 12987-12992. doi:10.1073/pnas.0604882103
- Z. Li, et al., “Heterologous Expression of OsSIZ1, a Rice SUMO E3 Ligase, Enhances Broad Abiotic Stress Tolerance in Transgenic Creeping Bentgrass,” Plant Biotechnology Journal, Vol. 11, No. 4, 2012, pp. 432-445. doi:10.1111/pbi.12030
- J. R. Park, I. McFarlane, R. H. Phipps and G. Ceddia, “The Role of Transgenic Crops in Sustainable Development,” Plant Biotechnology Journal, Vol. 9, No. 1, 2011, pp. 2-21. doi:10.1111/j.1467-7652.2010.00565.x
- C. James, “Global Status of Commercialised Biotech/GM Crops,” ISAAA Brief 41, Executive Summary, 2009. http://www.isaaa.org/resources/publications/briefs/41/executivesummary/default.asp
- A. McHughen and S. Smyth, “US Regulatory System for Genetically Modified [Genetically Modified Organism (GMO), rDNA or Transgenic] Crop Cultivars,” Plant Biotechnology Journal, Vol. 6, No. 1, 2008, pp. 2-12.
- P. G. Lemaux, “Genetically Engineered Plants and Foods: A Scientist’s Analysis of the Issues (Part I),” Annual Review of Plant Biology, Vol. 59, 2008, pp. 771-812. doi:10.1146/annurev.arplant.58.032806.103840
- P. G. Lemaux, “Genetically Engineered Plants and Foods: A Scientist’s Analysis of the Issues (Part II),” Annual Review of Plant Biology, Vol. 60, 2009, pp. 511-559. doi:10.1146/annurev.arplant.043008.092013
NOTES
*Corresponding author.

