American Journal of Plant Sciences
Vol.4 No.8A(2013), Article ID:35851,10 pages DOI:10.4236/ajps.2013.48A001
Asymmetric Somatic Plant Hybridization: Status and Applications
![]()
1Institute for Agricultural and Fisheries Research (ILVO), Plant Sciences Unit, Applied Genetics and Breeding, Melle, Belgium; 2Department of Plant Production, Faculty of Bioscience Engineering, Ghent University, Ghent, Belgium.
Email: johan.vanhuylenbroeck@ilvo.vlaanderen.be
Copyright © 2013 Lakshmanan Prabhu Shankar et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Received May 12th, 2013; revised June 13th, 2013; accepted July 15th, 2013
Keywords: Asymmetric Somatic Fusion; Genome Fragmentation; Protoplast
ABSTRACT
To create asymmetric somatic hybrids, the genome of the so-called donor protoplast is fragmented prior to protoplast fusion. As a result, only a limited amount of the donor genome is transferred to the fusion product. This technique can circumvent some commonly observed problems related to symmetric fusion and offers a practical breeding tool for asexual hybridization. Genomes are typically fragmented by irradiation, microprotoplast production or application of metabolic inhibitors such as iodoacetamide. Irradiation and microprotoplast production fragment the nuclear genome, whereas iodoacetamide inactivates the cytoplasmic genome. It can therefore be used to introduce cytoplasmic male sterility, an important practical application. For hybrid verification and genome characterization, molecular markers and cytogenetic techniques are applied. This review highlights and discusses progress made during the last decade in spermatophytes asymmetric protoplast fusion.
1. Introduction
Plant cells from which the cell wall has been enzymatically or mechanically removed are called protoplasts. Theoretically, protoplasts are totipotent, meaning that after their isolation and subsequent culture they have the capability to dedifferentiate, re-start the cell cycle, go through repeated mitotic divisions and then proliferate or regenerate into various organs. Fusion of protoplasts from different species can therefore be a practical breeding tool [1] and circumvents sexual hybridization related prezygotic or postzygotic barriers [2]. Somatic hybridization differs from other techniques in many respects. When comparing somatic hybridization to transgenic approaches, somatic hybridization enables broadening of the germplasm base, allows the transfer of uncloned multiple genes and generates products that are not subjected to the same legal regulations as transgenic lines [3,4]. Also, it transfers both monoand polygenic traits [5]. The first protoplast fusion was described more than a century ago by Küster [6]. Tobacco was the first crop in which successful interspecific somatic hybridization was reported [7]. Since then, improvements and somatic hybrids have been made in many species and complete plant regeneration was accomplished. A detailed historical overview of breakthroughs in protoplast related research is incorporated in Gamborg [8] and Davey et al. [9].
Protoplast fusion can be either symmetric or asymmetric depending on the nature of genetic contribution of the fusion partners. A somatic breeding protocol can typically be subdivided into the following steps: isolation, fragmentation (in case of asymmetric hybridization), fusion, regeneration and selection [10,11] (Figure 1). In symmetric fusion the complete genomes of both parent protoplasts are fused. However, when two complete genomes fuse, a phenomenon called “gene conflict” may arise, because certain chromosomes repel one another. Moreover, the technique introduces a significant amount of unwanted genetic material. These limitations result in abnormal growth, regeneration of hybrids with low fertility, non-rooted shoots, slow hybrid growth, and recalcitrant calli or microcalli [2].
In asymmetric fusion, after genome fragmentation only a limited amount of one genome is transferred to the
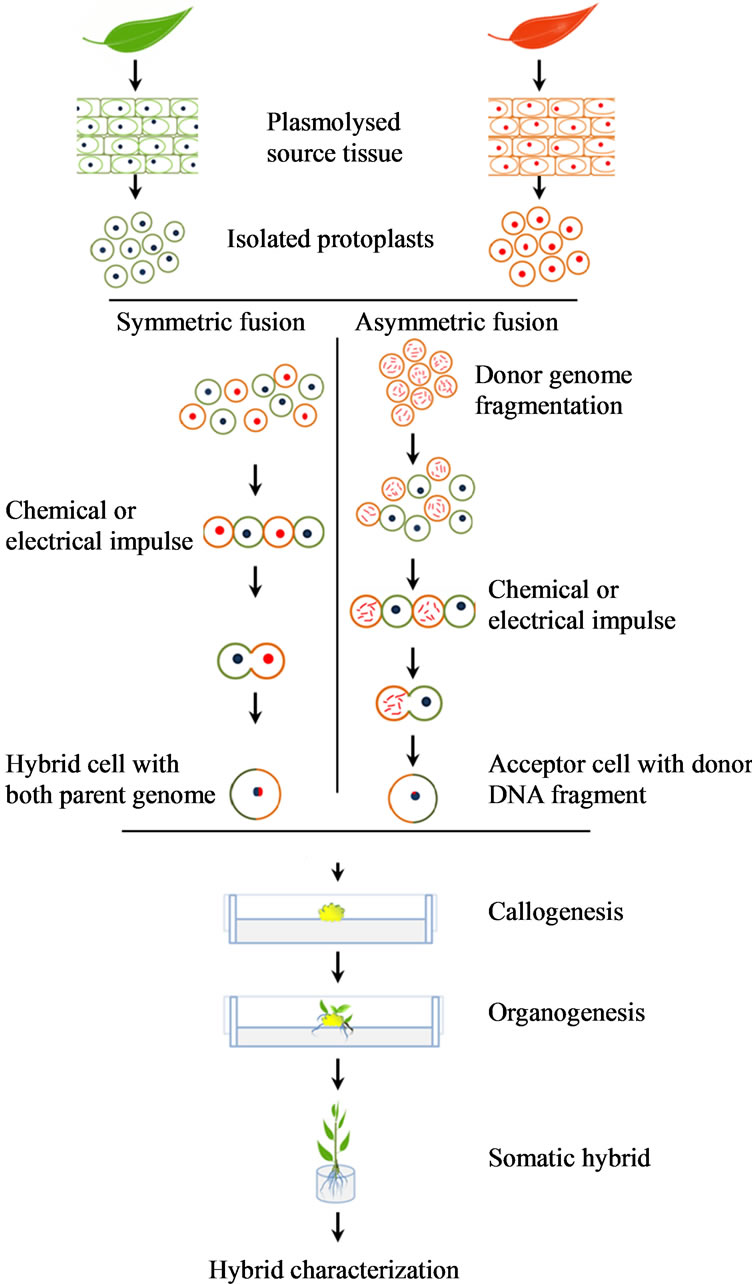
Figure 1. Schematic representation of symmetric and asymmetric somatic hybridization techniques.
fusion product [12,13] (Figure 1). Cytoplasmic genomes can also be recombined with nuclear genomes for applications such as cytoplasmic male sterility (CMS) introduction [11]. The technique is being applied to circumvent the above mentioned barriers in symmetric fusion. For example, symmetric hybrids between Brassica napus and Lesquerella fendleri are self-sterile, whereas asymmetric hybrids between the same fusion parents are selffertile [14]. Similarly, symmetric fusion between Orychophragmus violaceus and B. napus yields sterile hybrids, whereas asymmetric hybrids are fertile and can set seeds [15]. By introgressing fewer genes than after sexual crossing or symmetric somatic fusion, the number of backcrosses can also be significantly reduced. Genome fragmentation of the donor parent encourages the elimination of much of its redundant genetic material in the somatic hybrid. Moreover, in asymmetric fusions, most karyotype instability causing donor genes are eliminated during the first post-fusion mitoses, as opposed to symmetrical fusions after which eliminations can occur up to the first sexually derived generation [16]. In other words, not only does asymmetric fusion introduce fewer genes in a recipient genome after fragmenting the donor genome, but elimination of disadvantageous genes or chromosomes also proceeds quicker. Nevertheless, chromosomes and chromosome fragments can still be lost during meiosis due to rearrangements [17].
The objective of this review is to provide an overview of the development of technologies for fragmentation and screening. Besides the progress made in spermatophytes asymmetric protoplast fusion related research during the last decade is highlighted and discussed.
2. Fragmentation Techniques
In order to obtain asymmetric fusions, only partial genomes are transferred. Several techniques can be used for genome fragmentation of the so-called “donor” genome: irradiation, microprotoplasts (MPPs), or metabolic inhibitors as iodoacetamide (IOA). In some cases asymmetric fusions were realized without fragmentation treatment. The production of these asymmetric hybrids decreased possible long term irradiation effects on hybrid growth and development [18].
2.1. Irradiation
Genome fragmentation by irradiating can be achieved by using either ionizing radiation techniques (X or gamma rays) or non ionizing radiation (UV rays). Irradiation often induces random chromosome breakage, but also gene deletion and rearrangement and can be responsible for hybrid sterility [19]. The first application of X-rays to obtain asymmetric hybrids was performed in parsley [20]. UV light used to create asymmetric hybrids was used for the first time on Nicotiana donor protoplasts [21]. Earlier, X or gamma rays were more frequently used for donor protoplast fragmentation, but nowadays UV treated protoplasts are more widely applied. Both irradiation types efficiently induce asymmetric somatic hybrids in a dosedependent manner, e.g., between Brassica napus and Arabidopsis thaliana [22]. Nonetheless, within a single species susceptibility towards different irradiation types can vary strongly [23]. Hall et al. [24] investigated whether UV radiation could be used as an alternative for ionizing radiation techniques. They found that UV had a detrimental effect on sugar beet protoplasts: resynthesis of a cell wall, cell growth and cell division were partially or totally eliminated. However, protoplast viability had not decreased after 6 days culture, but after 14 days, the UV treated cells died. On the other hand, a significant advantage of UV radiation over ionizing radiation was its easy application and high reproducibility. Similar observations were made after exposure of Cichorium protoplasts to UV [25]. In cucumber, the negative effect of UV-C irradiation on cell wall regeneration, protoplast viability and the intensity of the nuclei after DAPI staining was also demonstrated [26].
A general problem is the quantification of DNA damage after an irradiation treatment. Abas et al. presented Comet assay single cell gel electrophoresis (SCGE) as a reliable tool to observe single and double strand breaks in mesophyll protoplasts of Nicotiana plumbaginifolia [27]. Xu et al. revealed extensive DNA fragmentation in UV irradiated Citrus unshiu protoplasts with the terminal deoxynucleotidyl transferase biotin-dUTP nick end labeling assay [28]. However, both methods are not generally applicable.
2.2. Microprotoplast Mediated Chromosome Transfer
Next to donor protoplast irradiation, micronuclei and microprotoplasts mediated chromosome transfer (MMCT), which was originally developed for mammalian cells, has been considered as an alternative method for partial genome transfer [29].
Mass induction of micronucleation and efficient isolation of microcells are key steps in any MMCT for successful transfer of partial genomes [30]. Microtubules are involved in several processes such as chromosome migration, cell structure, cellulose microfibrils guidance and arrangement, cell wall formation, intracellular movement and cell differentiation. Toxic substances as antimitotic herbicides or colchicine prevent their normal polymerization [31]. Application of these spindle toxins to synchronized cells generally blocks cells in metaphase and scatters chromosomes in the cytoplasm; subsequently, those decondense into micronuclei. Subsequently, these micronucleated cells are stripped of their cell wall, and the resulting MPPs are ultracentrifuged to subdivide them into classes [32]. These can be further filtered through sequential filters of smaller pore width. Some recent examples of suspension cell derived MPPs are Citrus unshiu [33] and Beta vulgaris [34].
In developing microspores of ornamental species like Lilium and Spathiphyllum, micronuclei were induced through the action of mitosis arresting chemicals, without synchronization requirement [35,36]. By using microspores instead of suspension cultures the risk of mutation accumulation in suspension cells can be avoided [37].
Regardless of the source material, the efficiency of spindle toxins depends on their type, dose, incubation period and the plant genotype. Those parameters can be optimized, as recently demonstrated for Spathiphyllum wallisii [36]. For this crop, the highest micronuclei indices were obtained after microspore treatment with 10 µM oryzalin for 72 h or 20 µM chlorpropham for 48 h for a particular model genotype. The maximal number of micronuclei observed was 12, while the haploid chromosome number amounts 15. Oryzaline is the most widely used mitosis inhibitor, but its efficiency varies based on the plant species [38,39]. Famelaer et al. quantified genome fragmentation in Beta microprotoplasts through flow cytometry and confocal microscopy [34].
2.3. Cytoplasmic Inactivation
Metabolic inhibitors, such as IOA and rhodamine 6-G can be used to obtain asymmetric fusions. The exact mode of action of IOA has not yet properly been described, the compound however inhibits protoplast division by irreversibly inactivating the cytoplasm. In red chicory mesophyll protoplasts division was totally inhibited after treatment with 2 - 4 mM iodoacetate [40]. When using IOA, a lower optimal concentration of 1.625 mM was found [25]. The lower optimal IOA concentrations compared to iodoacetate might be explained by better cell penetration of IOA. Similar optimal IOA concentration were found in Musa [41]. Lower IOA concentrations (0.5 mM) stopped growth of Gossypium hirsutum protoplasts, whereas 3 mM and 7.5 mM IOA were required to stop cell proliferation in Citrus [42] and Bupleurum scorzonerifolium [43], respectively.
Fusion of IOA-treated recipient parental protoplasts with irradiated donor protoplasts could produce cybrids. In Cichorium, successful asymmetric protoplast fusion has been performed between γ-rays-irradiated sunflower protoplasts and iodoacetate-treated red chicory protoplasts [40]. IOA treatments prevent cell division, but fusion with non-IOA treated protoplasts restores cell division ability, thus opening ways towards heterokaryon or cybrid selection.
3. Asymmetric Hybridization
In earlier protoplast reviews, the concept of asymmetric hybridization is well explained, but has not been the focus of the review [11,12]. Fragmentation techniques have been highlighted, however, [24,29] also in practical manuals [44]. In the review by Xia [13], wheat is used as an example to discuss chromosome engineering through asymmetric fusion.
Over the last decade, asymmetric fusion techniques have been widely applied and several new asymmetric hybrids were obtained (Table 1). The most studied families were Brassicacae and Poaceae, followed by Rutaceae. The number of asymmetric hybridization realized through PEG fusion was 4-fold the number of fusions generated by electrical fusion.
Asymmetric hybridization has allowed new genome combinations that would be more difficult, if not impossible, to realize through classical symmetric fusion or sexual crossing. For the first time, an asymmetric hybrid
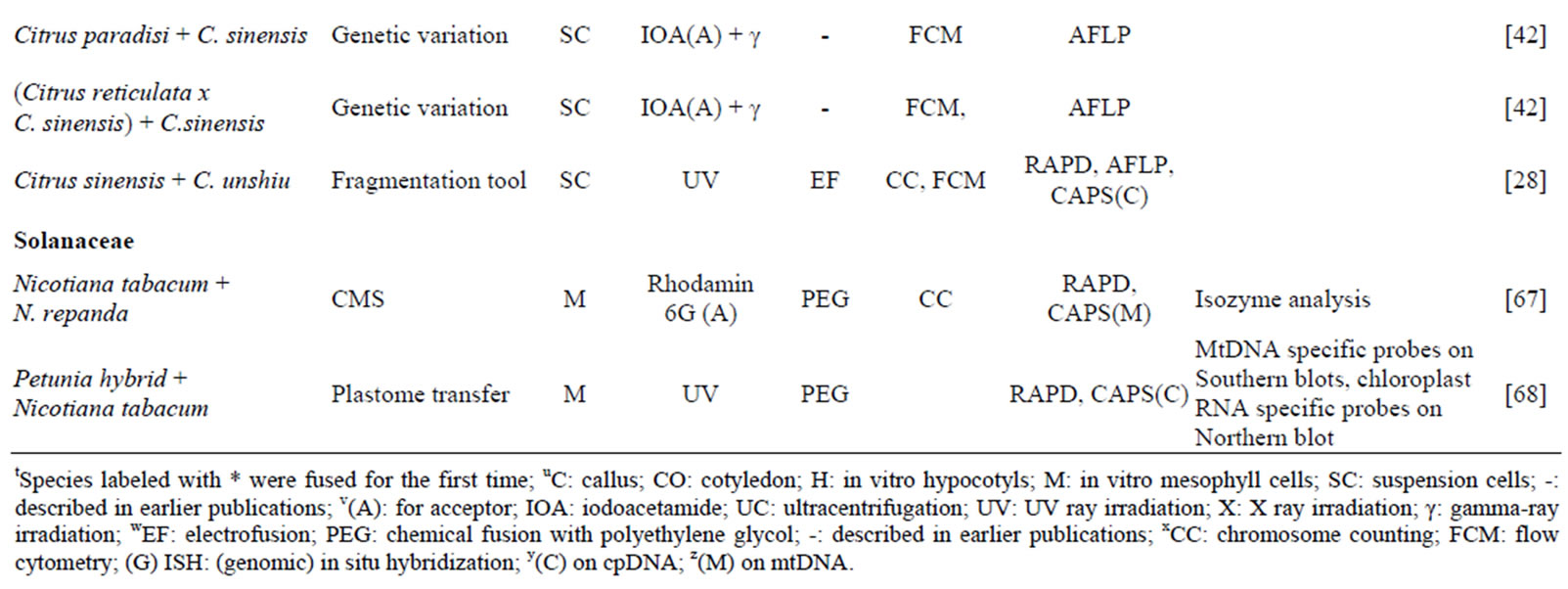

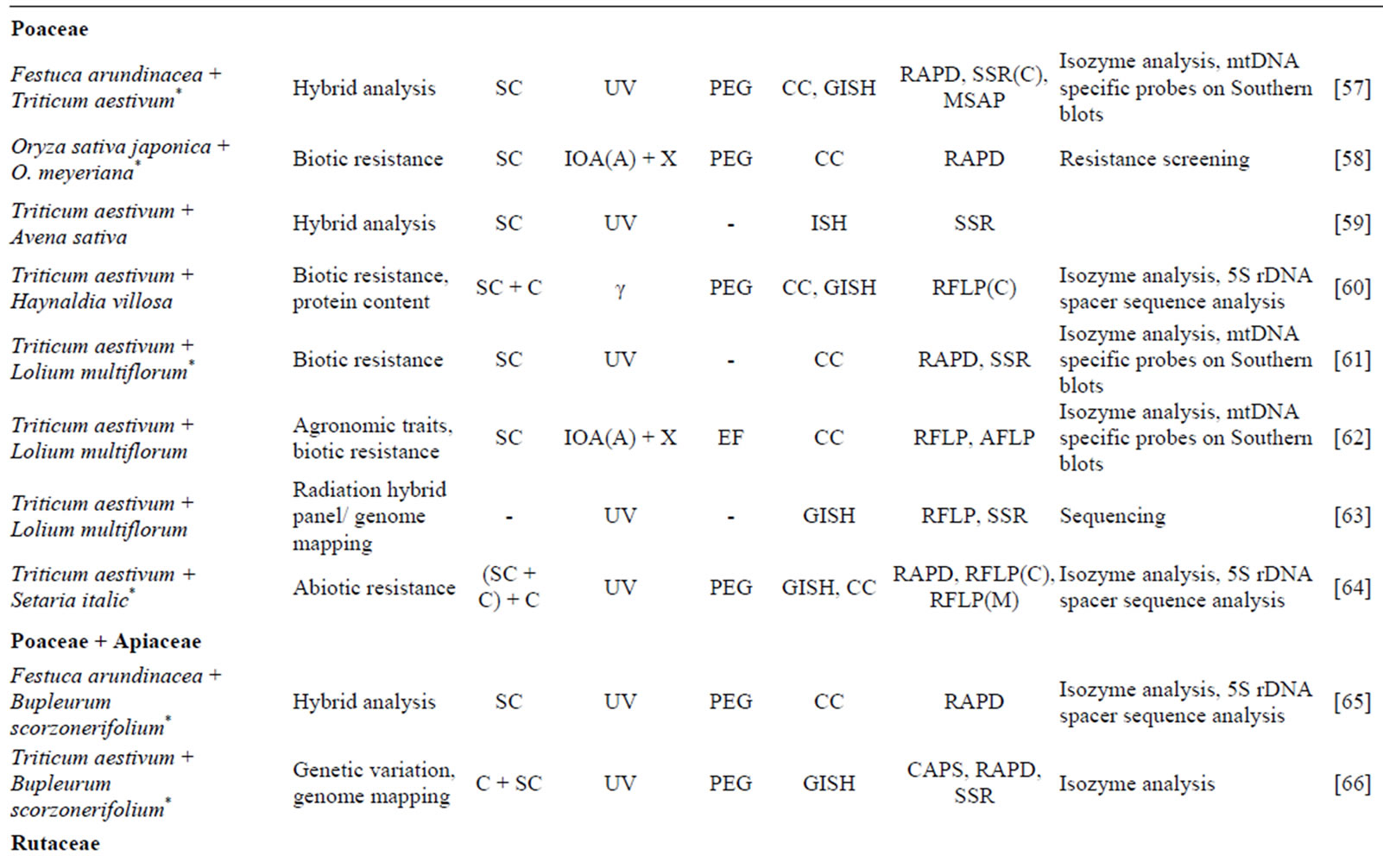
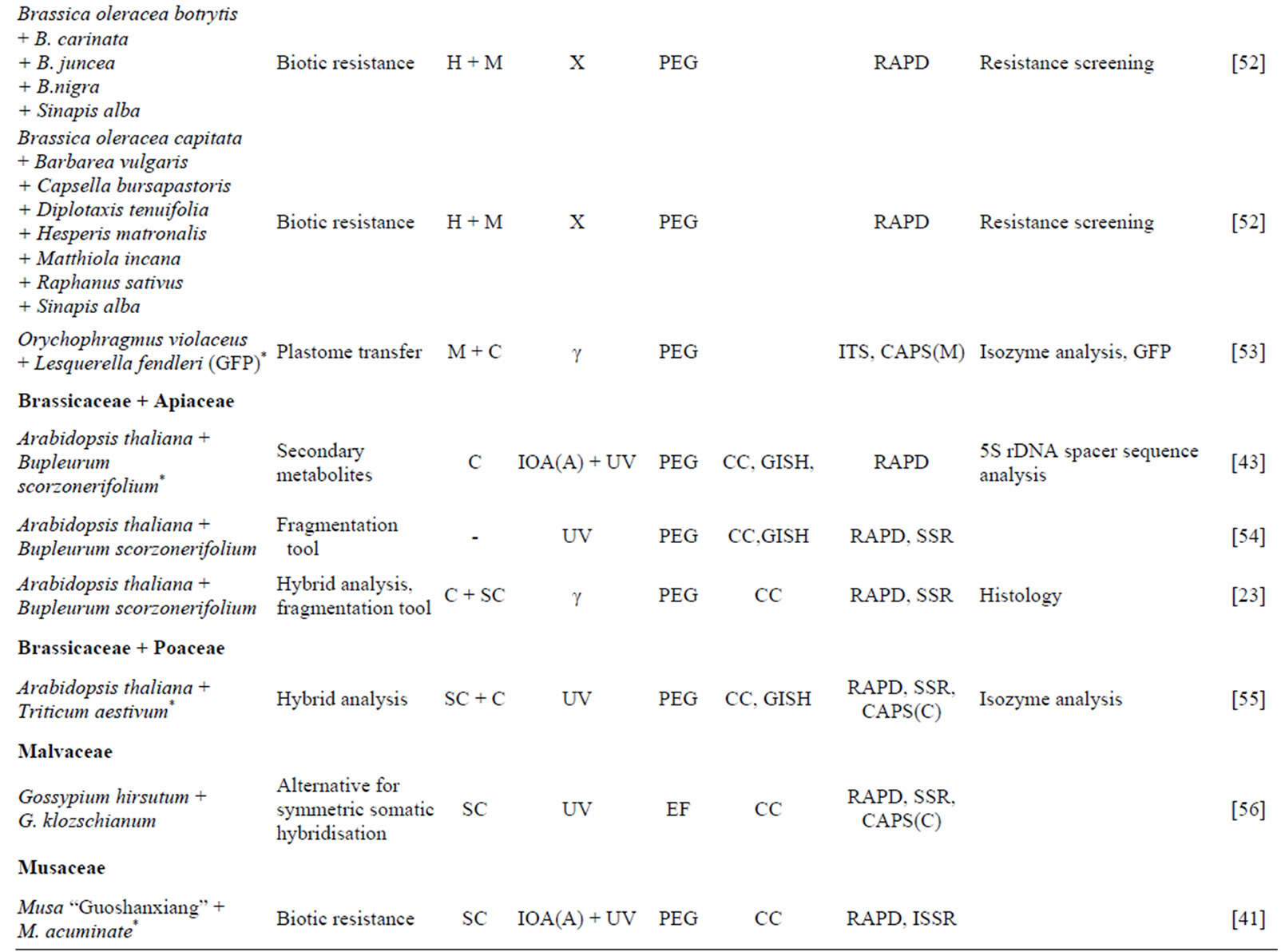

Table 1. Progress in asymmetric protoplast fusion in different plant families during last decade (2004-2013).
tSpecies labeled with * were fused for the first time; uC: callus; CO: cotyledon; H: in vitro hypocotyls; M: in vitro mesophyll cells; SC: suspension cells; -: described in earlier publications; v(A): for acceptor; IOA: iodoacetamide; UC: ultracentrifugation; UV: UV ray irradiation; X: X ray irradiation; γ: gamma-ray irradiation; wEF: electrofusion; PEG: chemical fusion with polyethylene glycol; -: described in earlier publications; xCC: chromosome counting; FCM: flow cytometry; (G) ISH: (genomic) in situ hybridization; y(C) on cpDNA; z(M) on mtDNA.
was reported in banana [41]. Interfamilial asymmetric hybrids have been produced between the dicot Arabidopsis thaliana and the monocot common wheat [55]. Fusion between phylogenetically remote tall fescue, Italian ryegrass and common wheat was achieved [57,61,62]. In cotton, asymmetric hybrids were obtained as an alternative for symmetric hybrids [56]. New somatic hybrids were obtained between monocot Festuca arundinacea and dicot Bupleurum scorzonerifolium through symmetric as well as asymmetric fusions [65]. The first successful somatic hybrid regeneration between Oryza sativa L. ssp. japonica and O. meyerina L. was reported [58]. Scholze et al. produced the first Raphanus-Brassica somatic hybrids with fungal and virus disease resistance [52]. Cybrids were produced between chloroplast transformant tobacco and petunia [68]. Using UV irradiated asymmetric hybrids a radiation hybrid panel was established for Lolium multiflorum [63]. Taski-Adjukovic et al. [48] regenerated an asymmetric hybrid between sunflower and Helianthus maximiliani for the first time.
Acceptor protoplast sources for asymmetric hybridization existed mainly of suspension cell cultures, mesophyll, callus and hypocotyls. The donor protoplast source differed in about 30% of the cases from the one for the acceptor. Brassicaceae and Asteraceae hypocotyl acceptor protoplasts were combined with mesophyll donor protoplasts [48,51,52].
Biotic resistance introduction, genetic variation, agronomic traits such as seedless fruits, hybrid analysis, fragmentation technology development and secondary metabolite production were the most important recent aims for asymmetric hybridization. Abiotic resistance introduction, hybridization, genome mapping and the establishment of chromosome addition lines were rare objectives (Table 1). In Bupleurum scorzonerifolium, asymmetric hybrids were obtained after protoplast fusion of UV treated B. scorzonerifolium and wheat protoplasts. However, instead of generating wheat carrying B. scorzonerifolium chromosome fragments, the reverse was found. This study can be of major interest for the construction of physical maps of the wheat genome [66]. The same was observed when untreated Arabidopsis thaliana protoplasts were fused with UV treated Bupleurum protoplasts [43].
Other motives were plastome and/or CMS transfer [67]. The latter is an important practical application of new genome/cytoplasmome combinations. Fitter et al. [69] demonstrated the possibility of introgressing CMS carried by mtDNA from a wild species into the cultivated crop. In Cichorium, CMS was introduced after asymmetric fusion with sunflower [40]. Sheahan et al. [70] reported the phenomenon of massive mitochondrial fusion (MMF) which leads to near-complete mixing of the mitochondrial population within 24 h. MMF appears specific to dedifferentiation, since it also occurs in mesophyll protoplasts of Arabidopsis and Medicago but not in protoplasts from already dedifferentiated cells such as tobacco BY-2 or callus cultures. These results allow a clearer interpretation of how novel mitochondrial genotypes develop following cell fusion. In other investigations, Sytnik et al. [71] demonstrated that also chloroplasts can be transferred to remote species by protoplast fusion.
4. Characterization of Asymmetric Somatic Hybrids and Genome Stability
Apart from morphological characterization, the majority of the publications reporting on complete plant regeneration describe the use of molecular tools to unravel the genomic constitution of the alleged hybrids (Table 1). DNA markers were sometimes complemented with isozyme analysis, sodium dodecyl sulphate polyacrylamide gel electrophoresis or sequence analysis. The most frequently employed molecular markers were Random Amplification of Polymorphic DNA (RAPD), Simple Sequence Repeats (SSR), Amplification Fragment Length Polymorphism (AFLP), Restriction Fragment Length Polymorphism (RFLP) and Cleaved Amplified Polymorphic Sequence (CAPS). PCR-RFLP and CAPS analysis using mitochondrial or chloroplast universal primer pairs were efficient and reliable methods for characterizing the cytoplasmic genome. This technique was applied for both chloroplast and mitochondria screening, whereas SSR was only used once for chloroplast evaluation. Southern blotting for cpDNA and mtDNA was employed to screen cytoplasmic DNA, whereas Northern blotting was used once for chloroplast evaluation. Compared to RFLP with labeled probes, CAPS is simpler, more rapid and less expensive [72]. Chloroplast SSR is even more convenient and efficient since enzyme cutting following PCR reaction is not needed [73]. Also sequencing of common bands and searching for restriction endonuclease sites could be cheaper and more convenient than actual CAPS analysis (though after sequencing CAPS could be used to confirm the results).
Besides molecular markers, cytogenetic tools as chromosome counting, flow cytometry and genomic in situ hybridization (GISH) can be used to distinguish asymmetric hybrids (Table 1). Especially the latter technique enables to visualize the hybrid genomic constitution and to follow genomic stabilization. After a symmetric fusion of two Triticum genotypes with Psathyrostachys, GISH analysis showed that the hybrids were highly asymmetric and contained only wild rye chromosome fragments, whereas the strong relationship of the hybrids and wheat was demonstrated by SSR markers [18]. Also the chloroplasts of the hybrids and wheat were identical. Apparently the genetic complementation of 2 Triticum parents stimulates the rejection of wild rye donor chromosomes. In the asymmetric Triticum aestivum + Setaria italica fusion, genome complementation can be used as a selection tool. The only regenerative callus type has 5 recombinant chromosomes and a chromosome count of 48, exceeding the 42 of normal wheat [64]. In non-regenerative calli, only 0 - 2 Setaria chromosomes were present. Symmetric [74] as well as asymmetric [43] Arabidopsis thaliana + Bupleurum scorzonerifolium somatic fusions were made. The symmetric hybrids contain the complete B.s. chromosome set, 0 - 2 A.t. chromosomes and some nuclear or cytoplasmic A.t. fragments. Also after B.s. UV irradiation, surprisingly A.t. genes were introgressed in the B.s. genome whereas the opposite was expected. In other words, A.t. chromatine is preferentially eliminated, the type of cross merely defines whether full chromosomes or DNA fragments were integrated.
Like nuclear genomes, cytoplasmic genomes are not always stable upon fusion. Intergenomic chloroplast recombination is a rare event in higher plants in contrast to mitochondrial genomes that show high recombination levels [75]. Sequencing, used for searching restriction endonuclease sites, can be efficiently combined with CAPS to demonstrate mtDNA recombination. MtDNA recombination was proven in Triticum aestivum + Setaria italica [64] and Arabidopsis thaliana + Brassica oleracea [76]. The latter authors proposed mitochondrial recombination as a tool for CMS introduction in cabbage. Although more rarely occurring, cpDNA recombination in hybrids has been demonstrated. In Triticum aestivum + Setaria italica hybrids, cpDNA coexistence as well as recombination occur [64]. It was also observed in Bupleurum scorzonerifolium + Swertia mussottii [46].
5. Conclusions
Recently, interest in protoplast related research has been renewed. Fusion of protoplasts allows us to move from traditional plant breeding methods to asexual methods. Compared to symmetric somatic hybridization, asymmetric fusion has some important advantages as it may limit genome conflicts. As shown in recent publications asymmetric fusion can create genomic variation for plant quality and yield improvement in agricultural or industrial crops. Especially the possibility to introduce CMS in important agricultural crops offers the breeders a tool towards hybrid seed production.
Interesting opportunities for further research may be found in the development of advanced methods for genome fragmentation including the combination of multiple techniques, such as the irradiation of MPPs, or the creation of MPPs from unreduced gametes formed by interspecific hybrids. The latter might open the possibility to transfer recombined chromosomes in a single step. Moreover, MPPs could be selected based on filtration properties, which would result in different genome types, enabling researchers to attribute plant traits to particular chromosomes and further utilize MPPs accordingly. Furthermore, the fast evolution in marker development will allow more profound studies on genome stability. Furthermore, the contribution of GISH to genome characterization studies after fusion will further increase and can be used as a tool for monitoring genomic stabilization.
REFERENCES
- Johnson and R. Veilleux, “Somatic Hybridization and Application in Plant Breeding,” In: J. Janick, Ed., Plant Breeding Rev 20, John Wiley & Sons, New York, 2001, pp. 167-225.
- T. Eeckhaut, K. Van Laere, J. De Riek and J. Van Huylenbroeck, “Overcoming Interspecific Barriers in Ornamental Plant Breeding,” In: J. Teixeira da Silva, Ed., Floriculture, Ornamental and Plant Biotechnology: Advances and Topical Issues, Global Science Books, London, 2006, pp. 540-551.
- J. W. Grosser and F. G. Gmitter, “2004 SIVB Congress Symposium Proceedings of Thinking Outside the Cell: Applications of Somatic Hybridization and Cybridization in Crop Improvement, with Citrus as a Model,” In Vitro Cellular & Developmental Biology Plant, Vol. 41, No. 3, 2005, pp. 220-225. doi:10.1079/IVP2004634
- J. Grosser and F. Gmitter, “Protoplast Fusion for Production of Tetraploids and Triploids: Applications for Scion and Rootstock Breeding in Citrus,” Plant Cell, Tissue and Organ Culture, Vol. 104, No. 3, 2011, pp. 343-357. doi:10.1007/s11240-010-9823-4
- R. Thieme, U. Darsow, L. Rakosy-Tican, Z. Kang, T. Gavrilenko, O. Antonova, U. Heimbach and T. Thieme, “Use of Somatic Hybridization to Transfer Resistance to Late Blight and Potato Virus Y (PVY) into Cultivated Potato,” Plant Breeding and Seed Science, Vol. 50, No. 1, 2004, pp. 113-118.
- E. Küster, “Über die Verschmelzung Nachter Protoplasten,” Berichte der Deutschen Botanischen Gesellschaft, Vol. 27, No. 10, 1909, pp. 589-598.
- P. S. Carlson, H. H. Smith and R. D. Dearing, “Parasexual Interspecific Plant Hybridization,” Proceedings of the National Academy of Sciences of the United States of America, Vol. 69, No. 8, 1972, pp. 2292-2294. doi:10.1073/pnas.69.8.2292
- O. L. Gamborg, “Plant Tissue Culture. Biotechnology. Milestones,” In Vitro Cellular & Developmental Biology Plant, Vol. 38, No. 2, 2002, pp. 84-92. doi:10.1079/IVP2001281
- M. Davey, P. Anthony, J. Power and K. Lowe, “2004 SIVB Congress Symposium Proceedings of Thinking Outside the Cell: Plant Protoplast Technology: Status and Applications,” In Vitro Cellular & Developmental Biology Plant, Vol. 41, No. 3, 2004, pp. 202-212. doi:10.1079/IVP2004633
- M. Razdan, “Somatic Hybridization,” In: Introduction to Plant Tissue Culture, Science Publishers, Enfield, 2003, pp. 147-175.
- J. Liu, X. Xu and X. Deng, “Intergeneric Somatic Hybridization and Its Application to Crop Genetic Improvement,” Plant Cell, Tissue and Organ Culture, Vol. 82, No. 1, 2005, pp. 19-44. doi:10.1007/s11240-004-6015-0
- S. Waara and K. Glimelius, “The Potential of Somatic Hybridization in Crop Breeding,” Euphytica, Vol. 85, No. 2, 1995, pp. 217-233. doi:10.1007/BF00023951
- G. Xia, “Progress of Chromosome Engineering Mediated by Asymmetric Somatic Hybridization,” Journal of Genetics and Genomics, Vol. 36, No. 9, 2009, pp. 547-556. doi:10.1016/S1673-8527(08)60146-0
- M. Skarzhinskaya, M. Landgren and K. Glimelius, “Production of Intertribal Somatic Hybrids between Brassica napus L. and Lesquerella fendleri (Gray) Wats,” Theoretical and Applied Genetics, Vol. 93, No. 8, 1996, pp. 1242-1250. doi:10.1007/BF00223456
- Q. Hu, L. Hansen, J. Laursen, C. Dixelius and S. Andersen, “Intergeneric Hybrids between Brassica napus and Orychophragmus violaceus Containing Traits of Agronomic Importance for Oilseed Rape Breeding,” Theoretical and Applied Genetics, Vol. 105, No. 6-7, 2002, pp 834-840. doi:10.1007/s00122-002-1017-y
- H. Cui, Z. Yu, J. Deng, X. Gao, Y. Sun and G. Xia, “Introgression of Bread Wheat Chromatin into Tall Wheatgrass via Somatic Hybridization,” Planta, Vol. 229, No. 2, 2009, pp. 323-330. doi:10.1007/s00425-008-0832-z
- C. Rambaud, J. Dubois and J. Vasseur, “Male-Sterile Chicory Cybrids Obtained by Intergeneric Protoplast Fusion,” Theoretical and Applied Genetics, Vol. 87, No. 3, 1993, pp. 347-352. doi:10.1007/BF01184922
- C. Li, G. Xia, F. Xiang, C. Zhou and A. Cheng, “Regeneration of Asymmetric Somatic Hybrid Plants from the Fusion of Two Types of Wheat with Russian Wildrye,” Plant Cell Reports, Vol. 23, No. 7, 2004, pp. 461-467. doi:10.1007/s00299-004-0834-7
- Y. Gleba, S. Hinnisdaels, V. Sidorov, V. Kaleda, A. Parokonny, N. Boryshuk, N. Cherep, I. Negrutiu and M. Jacobs, “Intergeneric Asymmetric Hybrids between Nicotiana plumbaginifolia and Atropa belladonna Obtained by Gamma-Fusion,” Theoretical and Applied Genetics, Vol. 76, No. 5, 1988, pp. 760-766. doi:10.1007/BF00303523
- D. Dudits, O. Fejer, G. Hadlaczky, C. Koncz, G. Lazar and G. Horvath, “Intergeneric Gene Transfer Mediated by Plant Protoplast Fusion,” Molecular and General Genetics, Vol. 179, No.2, 1980, pp. 283-288. doi:10.1007/BF00425455
- D. Dimanov and A. Atanassov, “A Study of the Possibility to Apply UV-Light in Experiments of Somatic Hybridization in Protoplast Cultures of Nicotiana alata and N. tabacum (Line Virginia 89),” Genetics and Breeding, Vol. 22, No 1, 1989, pp. 40-44.
- J. Forsberg, C. Dixelius, U. Lagercrantz and K.Glimelius, “UV Dose-Dependent DNA Elimination in Asymmetric Somatic Hybrids between Brassica napus and Arabidopsis thaliana,” Plant Science, Vol. 131, No. 1, 1998, pp. 65-76. doi:10.1016/S0168-9452(97)00242-2
- M. Wang, Z. Peng, S. Hong, D. Zhi and G. Xia, “Hybrid Inflorescences Derived from Gamma-Fusion of Arabidopsis thaliana with Bupleurum scorzonerifolium,” Protoplasma, Vol. 249, No. 1, 2012, pp. 197-205. doi:10.1007/s00709-011-0278-1
- R. Hall, F. Krens and G. Rouwendal, “DNA RadiationDamage and Asymmetric Somatic Hybridization: Is Uv a Potential Substitute or Supplement to Ionizing-Radiation in Fusion Experiments,” Physiologia Plantarum, Vol. 85, No. 2, 1992, pp. 319-324. doi:10.1111/j.1399-3054.1992.tb04741.x
- D. Deryckere, “Development of Asymmetric Somatic Hybridization Technology in Industrial Chicory (Cichorium intybus L.),” PhD Thesis, Ghent University, Ghent, 2013.
- B. Navratilova, L. Luhova and M. Petrivalsky, “Effect of UV-C Irradiation on Mesophyll Protoplasts of Cucumis sativus,” Plant Cell, Tissue and Organ Culture, Vol. 94, No. 3, 2008, pp. 313-318. doi:10.1007/s11240-008-9416-7
- Y. Abas, N. Touil, M. Kirsch-Volders, G. Angenon, M. Jacobs and I. D. H. Famelaer, “Evaluation of UV Damage at DNA Level in Nicotiana plumbaginifolia Protoplasts using Single Cell Gel Electrophoresis,” Plant Cell, Tissue Organ Culture, Vol. 91, No. 2, 2007, pp. 145-154. doi:10.1007/s11240-007-9257-9
- X. Y. Xu, Z. Y. Hu, J. F. Li, J. H. Liu and X. X. Deng, “Asymmetric Somatic Hybridization between UV-Irradiated Citrus unshiu and C. sinensis: Regeneration and Characterization of Hybrid Shoots,” Plant Cell Reports, Vol. 26, No 8, 2007, pp. 1263-1273. doi:10.1007/s00299-007-0350-7
- A. Yemets and Y. Blume, “Antimitotic Drugs for Microprotoplast-Mediated Chromosome Transfer,” In: Y. Blume, W. Baird, A. Yemets and Breviario, Eds., Plant Genomics, Cell Engineering and Breeding. The Plant Cytoskeleton: A Key Tool for Agro-Biotechnology, Springer, Dordrecht, 2009, pp. 419-434.
- K. S. Ramulu, P. Dijkhuis, I. Famelaer, T. Cardi and H. A. Verhoeven, “Isolation of Sub-Diploid Microprotoplasts for Partial Genome Transfer in Plants: Enhancement of Micronucleation and Enrichment of Microprotoplasts with One or a Few Chromosomes,” Planta, Vol. 190, No. 2, 1993, pp. 190-198. doi:10.1007/BF00196611
- K. Vaughn, “Anticytoskeletal Herbicides,” In: P. Nick, Ed., Plant Microtubules, Potential for Biotechnology, Springer, Berlin, 2000, pp. 193-205. doi:10.1007/978-3-662-22300-0_9
- K. S. Ramulu, P. Dijkhuis, E. Rutgers, J. Blaas, W. H. J. Verbeek, H. A. Verhoeven and C. M. Colijn-Hooymans, “Microprotoplast Fusion Technique: A New Tool for GeneTransfer between Sexually-Incongruent Plant Species,” Euphytica, Vol. 85, No. 1-3, 1995, 255-268. doi:10.1007/BF00023954
- Q. Zhang, J. Liu and X. Deng, “Isolation of Microprotoplasts from a Partially Synchronized Suspension Culture of Citrus unshiu,” Journal of Plant Physiology, Vol. 163, No. 11, 2006, pp. 1185-1192. doi:10.1016/j.jplph.2005.09.003
- I. Famelaer, H. A. Verhoeven, P. Dijkhuis and K. S. Ramulu, “A Study of the Process of Synchronisation and Micronucleation in Beta vulgaris and the Monitoring of an Isolation Procedure for Micro-Nuclei and Micro-Protoplasts by Confocal Laser Scanning Microscopy and Flow Cytometry,” Plant Cell, Tissue and Organ Culture, Vol. 90, No. 2, 2007, pp. 169-179. doi:10.1007/s11240-007-9225-4
- H. Saito and M. Nakano, “Isolation and Characterization of Gametic Microprotoplasts from Developing Microspores of Lilium longiflorum for Partial Genome Transfer in the Liliaceous Ornamentals,” Sexual Plant Reproduction, Vol. 15, No. 4, 2002, pp. 179-185. doi:10.1007/s00497-002-0153-5
- P. Lakshmanan, T. Eeckhaut, J. Van Huylenbroeck and E. Van Bockstaele, “Micronucleation by Mitosis Inhibitors in Developing Microspores of Spathiphyllum wallisii Regel,” Plant Cell Reports, Vol. 32, No. 3, 2013, pp. 369- 377. doi:10.1007/s00299-012-1370-5
- W. Guo and J. W. Grosser, “Somatic Hybrid Vigor in Citrus: Direct Evidence from Protoplast Fusion of an Embryogenic Callus Line with a Transgenic Mesophyll Parent Expressing the GFP Gene,” Plant Science, Vol. 168, No. 6, 2005, pp. 1541-1545. doi:10.1016/j.plantsci.2005.02.029
- P. C. Binsfeld, R. Wingender and H. Schnabl, “Characterization and Molecular Analysis of Transgenic Plants Obtained by Microprotoplast Fusion in Sunflower,” Theoretical and Applied Genetics, Vol. 101, No. 8, 2000, pp. 1250-1258. doi:10.1007/s001220051604
- F. Saito and M. Nakano, “Partial Synchronization of Cell Division and Micronucleation in Suspension-Cultured Cells of Hemerocallis hybrida: The Effects of Hydroxyurea and Various Spindle Toxins,” Breeding Science, Vol. 51, No. 4, 2001, pp. 285-291. doi:10.1270/jsbbs.51.285
- S. Varotto, E. Nenz, M. Lucchin and P. Parrini, “Production of Asymmetric Somatic Hybrid Plants between Cichorium intybus L. and Helianthus annuus L.,” Theoretical and Applied Genetics, Vol. 102, No. 6-7, 2001, pp. 950-956. doi:10.1007/s001220000485
- W. Xiao, X. Huang, Q. Gong, X. Dai, J. Zhao, Y. Wei and X. Huang, “Somatic Hybrids Obtained by Asymmetric Protoplast Fusion between Musa Silk cv. Guoshanxiang (AAB) and Musa acuminata cv. Mas (AA),” Plant Cell, Tissue and Organ Culture, Vol. 97, No. 3, 2009, pp. 313-321. doi:10.1007/s11240-009-9530-1
- C. De Bona, J. Gould, J. Miller, D. Stelly and E. Louzada, “Citrus Asymmetric Somatic Hybrids Produced via Fusion of Gamma-Irradiated and Iodoacetamide-Treated Protoplasts,” Pesquisa Agropecuaria Brasileira, Vol. 44, No. 5, 2009, pp. 454-462. doi:10.1590/S0100-204X2009000500004
- M. Wang, G. Xia and Z. Peng, “High UV-Tolerance with Introgression Hybrid Formation of Bupleurum scorzonerifolium Willd,” Plant Science, Vol. 168, No. 3, 2005, pp. 593-600. doi:10.1016/j.plantsci.2004.09.017
- J. Grosser, M. Calovic and E. Louzada E, “Protoplast Fusion Technology: Somatic Hybridization and Cybridization,” In: M. Davey and P. Anthony, Eds., Plant Cell Culture: Essential Methods, John Wiley & Sons, New York, 2010, pp. 175-198. doi:10.1002/9780470686522
- J. Wang, C. Zhao, C. Liu, G. Xia and F. Xiang, “Introgression of Swertia mussotii Gene into Bupleurum scorzonerifolium via Somatic Hybridization,” BMC Plant Biology, Vol. 11. No. 71, 2011. http://www.biomedcentral.com/1471-2229/11/71
- L. Jiang, Y. Cai Y, G. Xia and F. Xiang, “Introgression of the Heterologous Nuclear DNAs and Efficacious Compositions from Swertia tetraptera Maxim into Bupleurum scorzonerifolium Willd via Somatic Hybridization,” Protoplasma, Vol. 249, No. 3, 2012, pp. 737-745. doi:10.1007/s00709-011-0317-y
- Y. Yu, Z. Li, P. Wang and F. Xiang, “Genetic and Biochemical Characterization of Somatic Hybrids between Bupleurum scorzonerifolium and Gentianopsis paludosa,” Protoplasma, Vol. 249, No.4, 2012, pp. 1029-1035. doi:10.1007/s00709-011-0336-8
- K. Taski-Ajdukovic, D. Vasic and N. Nagl, “Regeneration of Interspecific Somatic Hybrids between Helianthus annuus L. and Helianthus maximiliani (Schrader) via Protoplast Electrofusion,” Plant Cell Reports, Vol. 25, No.7, 2006, pp. 698-704. doi:10.1007/s00299-006-0134-5
- X. Du, X. Ge, X. Yao, Z. Zhao and Z. Li, “Production and Cytogenetic Characterization of Intertribal Somatic Hybrids between Brassica napus and Isatis indigotica and Backcross Progenies,” Plant Cell Reports, Vol. 28, No. 7, 2009, pp. 1105-1113. doi:10.1007/s00299-009-0712-4
- Z. G. Zhao, T. T. Hu, X. H. Ge, X. Z. Du, L. Ding and Z. Y. Li, “Production and Characterization of Intergeneric Somatic Hybrids between Brassica napus and Orychophragmus violaceus and their Backcrossing Progenies,” Plant Cell Reports, Vol. 27, No. 10, 2008, pp. 1611-1621. doi:10.1007/s00299-008-0582-1
- G. X. Wang, Y. Tang, H. Yan, X. G. Sheng, W. W. Hao, L. Zhang, K. Lu and F. Liu, “Production and Characterization of Interspecific Somatic Hybrids between Brassica oleracea var. botrytis and B. nigra and their Progenies for the Selection of Advanced Pre-Breeding Materials,” Plant Cell Reports, Vol. 30, No. 10, 2011, pp. 1811-1821. doi:10.1007/s00299-011-1088-9
- P. Scholze, R. Kramer, U. Ryschka, E. Klocke and G. Schumann, “Somatic Hybrids of Vegetable Brassicas as Source for New Resistances to Fungal and Virus Diseases,” Euphytica, Vol. 176, No. 1, 2010, pp. 1-14. doi:10.1007/s10681-010-0205-0
- O. Ovcharenko, V. Momot, N. Cherep, Y. Sheludko, I. Komarnitsky, V. Rudas and N. Kuchuk, “Transfer of Transformed Lesquerella fendleri (Gray) Wats Chloroplasts into Orychophragmus violaceus (L.) OE Schulz by Protoplast Fusion,” Plant Cell, Tissue and Organ Culture, Vol. 105, No. 1, 2011, pp. 21-27. doi:10.1007/s11240-010-9833-2
- M. Wang, Z. Peng, M. Wang, W. Guo, J. Zhao, D. Zhi and G. Xia, “The Behavior of Bupleurum scorzonerifolium as a Parent of Somatic Hybrid Introgressed Lines is Associated with UV Resistance of its Chromosomes,” Russian Journal of Plant Physiology, Vol. 58, No. 4, 2011, pp. 615-621. doi:10.1134/S102144371104025X
- J. Deng, H. Cui, D. Zhi, C. Zhou and G. Xia, “Analysis of Remote Asymmetric Somatic Hybrids between Common Wheat and Arabidopsis thaliana,” Plant Cell Reports, Vol. 26, No. 8, 2007, pp. 1233-1241. doi:10.1007/s00299-007-0345-4
- X. Yang, X. Zhang, S. Jin, L. Fu and L. Wang, “Production and Characterization of Asymmetric Hybrids between Upland Cotton Coker 201 (Gossypium hirsutum) and Wild Cotton (G. klozschianum Anderss), “ Plant Cell Tissue and Organ Culture, Vol. 89, No. 2-3, 2007, pp. 225-235. doi:10.1007/s11240-007-9245-0
- Y. Cai, F. Xiang, D. Zhi, H. Liu and G. Xia, “Genotyping of Somatic Hybrids between Festuca arundinacea Schreb. and Triticum aestivum L.,” Plant Cell Reports, Vol. 26, No. 10, 2007, pp. 1809-1819. doi:10.1007/s00299-007-0397-5
- C. Yan, K. Qian, Q. Yan, X. Zhang, G. Xue, W. Huangfu, Y. Wu, Y. Zhao, Z. Xue, J. Huang, G. Xu and P. Wu, “Use of Asymmetric Somatic Hybridization for Transfer of the Bacterial Blight Resistance Trait from Oryza meyeriana L. to O. sativa L. sp japonica,” Plant Cell Reports, Vol. 22, No. 8, 2004, pp. 569-575. doi:10.1007/s00299-003-0732-4
- F. Xiang, J. Wang, C. Xu and G. Xia, “The Chromosome Content and Genotype of Two Wheat Cell Lines and of their Somatic Fusion Product with Oat,” Planta, Vol. 231, No. 5, 2010, pp. 1201-1210. doi:10.1007/s00425-010-1113-1
- [61] A. Zhou and G. Xia, “Introgression of the Haynaldia villosa Genome into Gamma-Ray-Induced Asymmetric Somatic Hybrids of Wheat,” Plant Cell Reports, Vol. 24, No. 5, 2005, pp. 289-296. doi:10.1007/s00299-005-0922-3
- [62] A. Cheng and G. Xia, “Somatic Hybridisation between Common Wheat and Italian Ryegrass,” Plant Science, Vol. 166, No. 5, 2004, pp. 1219-1226. doi:10.1016/j.plantsci.2003.12.038
- [63] T. Ge, X. Lin, F. Qin, S. Yu and Y. Yu, “Protoplast Electrofusion between Common Wheat (Triticum aestivum L.) and Italian Ryegrass (Lolium multiflorum Lam.) and Regeneration of Mature Cybrids,” In Vitro Cellular & Developmental Biology-Plant, Vol. 42, No. 2, 2006, pp. 179- 187. doi:10.1079/IVP2005742
- [64] A. Cheng, H. Cui and G. Xia, “Construction of a Primary RH Panel of Italian Ryegrass Genome via UV-Induced Protoplast Fusion. Plant Biology, Vol. 8, No. 5, 2006, pp. 673-679. doi:10.1055/s-2006-924276
- [65] F. Xiang, G. Xia, D. Zhi, J. Wang, H. Nie and H. Chen, “Regeneration of Somatic Hybrids in Relation to the Nuclear and Cytoplasmic Genomes of Wheat and Setaria italica,” Genome, Vol. 47, No. 4, 2004, pp. 680-688. doi:10.1139/g04-023
- [66] M. Wang, Z. Peng, L. Wang, J. Zhao, J. Che and G. Xia, “Different Rates of Chromosome Elimination in Symmetric and Asymmetric Somatic Hybridization between Festuca arundinacea and Bupleurum scorzonerifolium,” Russian Journal of Plant Physiology, Vol. 58, No. 1, 2011, pp. 133-141. doi:10.1134/S1021443710061068
- [67] C. Zhou, G. Xia, D. Zhi and Y. Chen, “Genetic Characterization of Asymmetric Somatic Hybrids between Bupleurum scorzonerifolium Willd and Triticum aestivum L.: Potential Application to the Study of the Wheat Genome,” Planta, Vol. 223, No. 4, 2006, pp. 714-724. doi:10.1007/s00425-005-0127-6
- [68] Y. Sun, Q. Xue, C. Ding, X. Zhang, L. Zhang, W. Wang and S. Ali, “Somatic Cybridization between Nicotiana tabacum and N. repanda Based on a Single Inactivation Procedure of Nuclear Donor Parental Protoplasts,” Plant Science, Vol. 168, No. 2, 2005, pp. 303-308. doi:10.1016/j.plantsci.2004.05.026
- [69] A. Sigeno, S. Hayashi, T. Terachi and H. Yamagishi, “Introduction of Transformed Chloroplasts from Tobacco into Petunia by Asymmetric Cell Fusion,” Plant Cell Reports, Vol. 28, No. 11, 2009, pp. 1633-1640. doi:10.1007/s00299-009-0763-6
- [70] J. T. Fitter, M. R. Thomas, C. Niu and R. J. Rose, “Investigation of Nicotiana tabacum (+) N. suaveolens Cybrids with Carpelloid Stamens,” Journal of Plant Physiology, Vol. 162, No.2, 2005, pp. 225-235. doi:10.1016/j.jplph.2004.02.006
- [71] M. B. Sheahan, D. W. McCurdy and R. J. Rose, “Mitochondria as a Connected Population: Ensuring Continuity of the Mitochondrial Genome during Plant Cell Dedifferentiation through Massive Mitochondrial Fusion,” The Plant Journal, Vol. 44, No 5, 2005, pp. 744-755. doi:10.1111/j.1365-313X.2005.02561.x
- [72] E. Sytnik, I. Komarnytsky, Y. Gleba and N. Kuchuk, “Transfer of Transformed Chloroplasts from Nicotiana tabacum to the Lycium barbarum Plants,” Cell Biology International, Vol. 29, No. 1, 2005, pp. 71-75. doi:10.1016/j.cellbi.2004.11.013
- [73] W. W. Guo, D. Prasad, Y. J. Cheng, P. Serrano, X. X. Deng and J. W. Grosser, “Targeted Cybridization in Citrus: Transfer of Satsuma Cytoplasm to Seedy Cultivars for Potential Seedlessness,” Plant Cell Reports, Vol. 22, No. 10, 2004, pp. 752-758. doi:10.1007/s00299-003-0747-x
- [74] Y. Cheng, M. C. de Vicente, H. Meng, W. Guo, N. Tao and X. Deng, “A Set of Primers for Analyzing Chloroplast DNA Diversity in Citrus and Related Genera,” Tree Physiology, Vol. 25, No. 6, 2005, pp. 661-672. doi:10.1093/treephys/25.6.661
- [75] M. Wang, J. Zhao, Z. Peng, W. Guo, Y. Wang, L. Wang and G. Xia, “Chromosomes Are Eliminated in the Symmetric Fusion between Arabidopsis thaliana L. and Bupleurum scorzonerifolium Willd., “ Plant Cell, Tissue and Organ Culture, Vol. 92, No. 2, 2008, pp. 121-130. doi:10.1007/s11240-007-9308-2
- [76] S. Trabelsi, R. Gargouri-Bouzid, F. Vedel, A. Nato, L. Lakhoua and N. Drira, “Somatic Fybrids between Potato Solanum tuberosum and Wild Species Solanum vernei Exhibit a Recombination in the Plastome,” Plant Cell Tissue and Organ Culture, Vol. 83, No. 1, 2005, pp. 1-11. doi:10.1007/s11240-005-3667-3
- [77] H. Yamagishi, S. Nakagawa, D. Kinoshita, A. Ishibashi and Y. Yamashita, “Somatic Hybrids between Arabidopsis thaliana and Cabbage (Brassica oleracea L.) with All Chromosomes Derived from A.thaliana and Low Levels of Fertile Seed,” Journal of the Japanese Society for Horticultural Science, Vol. 77, No. 3, 2008, pp. 277-282. doi:10.2503/jjshs1.77.277

