American Journal of Plant Sciences
Vol.3 No.3(2012), Article ID:17905,10 pages DOI:10.4236/ajps.2012.33036
Potential Inbreeding in a Small Population of a Mass Flowering Species, Xanthorrhoea johnsonii (Xanthorrhoeaceae): Is Your Mother My Father?
![]()
1Department of Mathematics and Computing, Faculty of Sciences, University of Southern Queensland, Toowoomba, Australia; 2School of Environment, Griffith University, Nathan, Australia.
Email: *J.Zalucki@griffith.edu.au
Received November 23rd, 2011; revised December 26th, 2011; accepted January 11th, 2012
Keywords: Xanthorrhoeaceae; inbreeding; mass flowering; microsatellite; paternity; outcrossing; pollen dispersal; genetic structure; isolated population; Xanthorrhoea johnsonii
ABSTRACT
Xanthorrhoea johnsonii is a long lived slow growing perennial understorey species, that produces a large quantity of passively dispersed seed every 3 - 5 years. Reproductive maturity is not reached until 20 - 30 years of age. The temporal asynchrony of the flowering event in this population was analogous to geographic isolation through fragmentation. A small population of plants flowering in isolation provided the opportunity to examine outcrossing rates, genetic diversity and the paternity of progeny at a small spatial scale (0.2 ha). The geographic location and physical characteristics of the adult plants were recorded, and both adults and their seed were sampled for genetic analysis. Four microsatellite loci were screened for genetic diversity and spatial structure analysis. A population outcrossing rate was estimated, as well as the number of paternal parents required to resolve the progeny multilocus genotypes. High genetic diversity was found in both adults and progeny with an estimated 97% outcrossing rate. All maternal lines required several paternal contributors, with no evidence of dominant paternal genotypes. Pollen transfer occurred between both geographically close and distant plants.
1. Introduction
Small fragmented populations can be vulnerable to loss of genetic diversity due to genetic drift and inbreeding [1, 2]. This scenario is all too common for many plant populations, where large areas of natural landscapes have become fragmented and interspersed with various types of anthropogenic land use activities. The reduction in the number of potential breeding pairs in these smaller populations can lead to increased selfing or biparental inbreeding [3-6]. In the long term smaller effective population size can also lead to a loss of genetic diversity in successive generations due to genetic drift [5,7,8]. Plants that have a mixed mating system, and have maintained high outcrossing rates could be vulnerable to a loss of genetic variability if selfing rates and levels of bi-parental inbreeding, increase due to habitat fragmentation [9,10].
The spatial arrangement of related individuals within a population can influence the potential for inbreeding [7]. The location of individual plants is the result of past seed dispersal events and recruitment patterns. Local genetic structuring occurs when genetically related seeds are deposited in close proximity, germinate, establish and grow into mature plants [11]. Structuring is reinforced in subsequent generations if pollen is then moved among these related individuals (biparental inbreeding), or within the same individual (selfing). Both selfing and biparental inbreeding will produce seed along local pedigree lines [12]. Long-lived perennial plants often have an extended flowering period and individual plants may not exhibit synchronous flowering within this time period [13].
The movement of pollen within a plant can influence the potential for inbreeding. For plants that produce a spike inflorescence, with protandrous flowers, it is likely that there are both male and female phase flowers present at any one time on an individual plant [14,15]. Depending on how the pollinators move among the flowers within an individual spike there is the opportunity for selfing via geitonogamous pollination. At the population level, flowering density may impact the distances pollinators’ travel among flowering plants and the amount of time pollinators spend foraging on individual plants [16, 17].
Each plant can potentially contribute genes to the next generation by either (or both) maternal and paternal roles. Genetic diversity in the seed generation is higher if all flowering plants (rather than just a few individuals) contribute genes to the seed generation. Failure of any plant to be a parent (either in a maternal or paternal role) will reduce the level of genetic diversity in the seed generation. Unequal representations of certain genotypes in the seed generation is the result of some plants either setting more seed, or fathering more seeds than other plants. Ultimately then, the genetic relatedness of the viable seed produced at the end of each flowering season is the result of the effects of the breeding system, pollen movements and mating patterns for that particular flowering season [18-20].
Using genetic markers to assess the outcrossing rates of seed is one way of examining levels of inbreeding and genetic diversity. Microsatellite markers are often the marker of choice for population studies as they are highly polymorphic, considered to be selectively neutral and reflect unbiased patterns of mutation gene flow and genetic drift [21]. They can be used to explore population structure, outcrossing rates and relatedness within a population [22-24] .
Xanthorrhoea johnsonii is a slow-growing, long-lived perennial that does not reach reproductive maturity until 20 - 30 years of age and then reproduces every 3 - 5 years. These plants are self-compatible and produce viable seeds. Insects, birds, lizards, and possums visit the spike inflorescences but the most likely pollinators are bees, specifically Trigona carbonaria and Apis mellifera. Each plant can produce thousands of seeds and these have been observed in thick deposits on the surface of the soil/litter layer. Primary seed dispersal is passive and secondary seed dispersal agents are unknown. Whatever the fate of the seed, it is likely that large quantities of closely related seeds end up within short distances of the maternal parent. A previous study of several larger populations during normal breeding seasons showed consistently high outcrossing rates despite varying flowering densities [25].
Xanthorrhoea johnsonii is a dominant understorey species, is fire tolerant, with fire events often stimulating flowering in some individuals [26]. A controlled burn to reduce fuel load (within an area of approx 0.2 ha) triggered an isolated flowering event among approximately 30 individual X. johnsonii plants. This created a small, temporally isolated/fragmented population and provided a unique opportunity to assess the outcrossing rates and genetic variability of the progeny from one breeding season. In this effectively closed population the seed or progeny produced must have both maternal and paternal parents sourced from this location. This provided an excellent opportunity to quantify selfing rates and biparental inbreeding in this population.
The specific aims of the project were to use microsatellite markers previously developed for this species to: 1) investigate the genetic variability in the adult and progeny populations, and any possible spatial patterns in relatedness; 2) estimate outcrossing rates, and 3) examine cohorts of seed from several maternal lines and estimate the number of different pollen donors in this population.
2. Materials and Methods
2.1. Study Species and Sample Site
Xanthorrhoea johnsonii is a species endemic to the Australian east coast [27] and is the most common and widespread species of Xanthorrhoea in Queensland (QLD) and New South Wales (NSW) (Figure 1). Typically a plant has an unbranched trunk between 10 cm to 5 m in height. Each flowering individual produces a single indeterminate spike inflorescence that varies from 75 cm to 2 m in length. Flowering usually occurs between May and November [28].
The length of time of a flowering event varies from 10 days in warmer temperatures, to approximately 3 weeks in cooler conditions (for each individual plant). Individual flowers open in sequence from the bottom of the spike towards the top and individual flowers appear to be protandrous with stigmas maturing after approximately 3 days [25]. Because of the sequential, progressive nature of flowering, both male and female phase flowers can be present on the same inflorescence.
Each inflorescence is capable of producing in excess of one thousand flowers (depending on spike length) and each flower has 6 stamens and 3 ovules [25,29]. Therefore, each plant is capable of setting several thousand seeds each time it flowers. Primary seed dispersal is passive and nothing is known about secondary dispersal agents.
Toohey Forest covers approx 681 hectares (LAT 27˚32'S LONG 153˚43'E) of eucalypt or open sclerophyll forest. The out of season flowering event (January) encompassed an area of approximately 0.2 hectares and included 30 flowering plants. The GPS positions (lat/ longs), plant heights (cm) and spike lengths (cm) were recorded for all 30 plants. Leaves and seed were sampled from a subset (seven) of these plants for genetic analysis of adults and progeny. The smallest distance between any two plants sampled was 1.85 m and the greatest distance was 46 m. Samples consisted of 2 to 3 cm of leaf tissue taken from the growing tips of the plants and were immediately placed on ice. These samples were frozen at
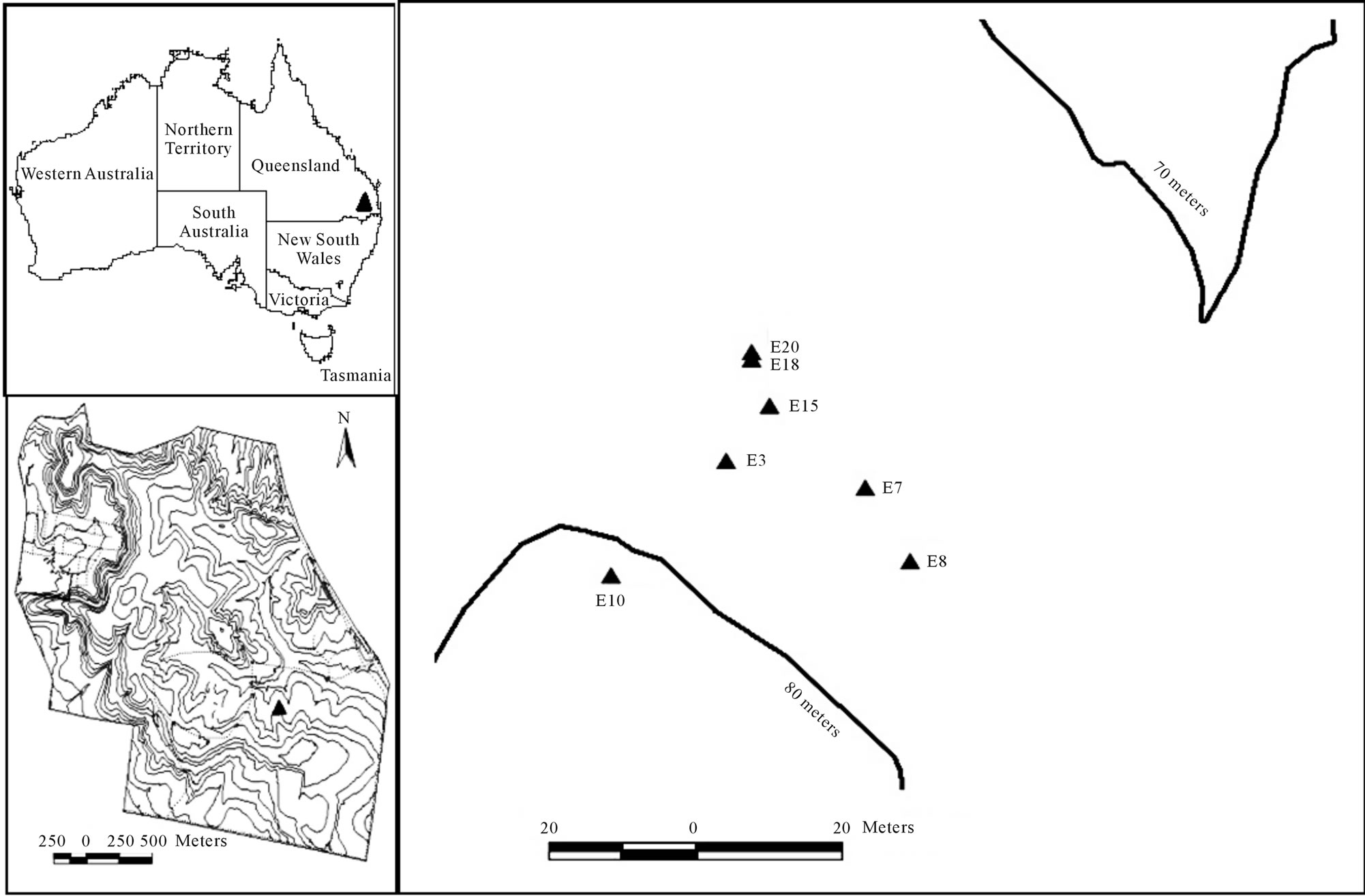
Figure 1. Location of the study population and the seven sampled adult plants within.
−80˚C within 30 minutes of sampling. Sampled plants were tagged for future seed collection.
Once mature, a random sample of approximately 25 seeds per plant was collected. Seed was germinated on 0.6% agar plates and incubated at 25˚C in the dark for 7 to 10 days. Two millimetres of the radical was removed for genetic analysis from 20 germinated seed per parent plant and frozen at −80˚C.
2.2. Molecular Methods
Molecular analysis of the leaf samples from the adult flowering plants was used to determine maternal genotypes. Total genomic DNA was extracted from the leaf samples using the QIAGEN DNeasy Plant Mini-kit (QIAGEN Industries) after first grinding 100 mg of leaf tissue in liquid nitrogen. Poly-vinyl pyrolidone (PVP) powder (5 mg) was included in the grinding process to assist in removing DNA-degrading phenolic compounds [30].
Analysis of the seedling radicals was used to identify the progeny genotypes. DNA extraction from all seedling tissue followed a modification of a Hair Lysis Extraction protocol [31]. Small amounts (2 - 3 mm) of unfrozen seedling tissue were combined with 100 mL of lysis extraction buffer and 5 mL Proteinase K (20 mg/mL). The extraction buffer included 1% 10 × PCR buffer (FisherBiotech), 2.5 mM MgCL2 and 0.5% TWEEN-20. All samples were then incubated for 45 min at 60˚C then stored at −20˚C until used in polymerase chain reaction (PCR) amplification.
Five sets of X. johnsonii microsatellite primers (X1, X4, X5, X8 and X9) developed by (J. Zalucki unpubl. res.) were amplified for all seven maternal parents (E3, E7, E8, E10, E15, E18 and E20) and 140 progeny (20 per maternal parent). PCR reactions were performed using 12.5 mL reaction volume containing: 1.25 mL 10 × Taq polymerase buffer, 2 mM MgCL2 0.2 mM dNTP’s, 0.4 mM of forward and reverse primers, 0.25 Units of Biotech Taq polymerase and 10 - 15 ng of DNA template. Temperature cycling conditions were: 3 minutes denaturation at 95˚C, then 30 cycles of 30 seconds at 94˚C, 30 seconds at 54˚C, and 30 seconds at 72˚C; followed by a final extension phase of 7 minutes at 72˚C.
A high rate of failures during initial PCR amplifications was due to highly variable DNA yield from the Hair Lysis Extraction process. The PCR success rate was improved considerably by increasing or decreasing the amount of DNA template used for each individual. Annealing temperatures were reduced by 2˚C - 4˚C for some individuals. Initial annealing temperatures (˚C) for X1, X4, X5, X8 and X9 were 61, 51, 59, 59 and 59 respectively. PCR amplification was not successful at all loci for all progeny; however, the minimum number of progeny successfully screened at any of the five loci was 15.
One locus, X8, was excluded from further analysis due to the likely occurrence of null alleles (high levels of homozygote excess and mismatches between the progeny and the known maternal parent). Significant null allele estimates were found at this locus using Micro-Checker [32]. This program estimates null allele frequencies using both methods by Chakraborty et al. [33] and Brookfield [34]. For all four dinucleotide, polymorphic loci included in the analysis, all alleles found in the maternal parents were also found in the progeny and there were no further mismatches between a known maternal parent and its progeny.
2.3. Statistical Analysis
Expected heterozygosity (HE) for each locus, exact tests for deviations from Hardy-Weinberg Equilibrium (HWE) for each locus and across all loci (1000 iterations), and linkage disequilibrium for all locus pairs (1000 iterations), were calculated using the program GENEPOP (Raymond and Rousset [35]; web version at http://genepop.curtin.edu.au/ 27 April 2010). Weir and Cockerham [36] FIS values (1000 iterations) were also calculated for each locus and across all loci using GENEPOP [35].
The average pairwise relatedness between adult plants and also between each maternal parent its own progeny and the progeny of each of the other six maternal parents, was calculated in the program GenAlEx 6 [39]. The relatedness coefficient “r” was calculated using the Queller and Goodnight [37] estimator.
Pairwise genetic and geographic distances [38] were calculated and spatial autocorrelation performed on the progeny using GenAlEx 6 [39]. Progeny were assigned the spatial coordinates (lat/longs) of their maternal parents for the purpose of this analysis. A minimum distance class of 5 m was used. Following Peakall et al., [40], GenAlEx tests for statistical significance based on the random permutation of “r” (relatedness co-efficient) under the assumption of no spatial structure. From 1000 random and then ranked permutations of all individuals among the geographic locations the 95% confidence interval was determined. If the calculated r-value falls outside this confidence region, then significant positive or negative spatial genetic structure was inferred. An autocorrelogram showing the autocorrelation coefficient “r” as a function of the nominated distance classes and the 95% confidence interval was plotted in GenAlEx.
The program Cervus [41] was used to identify possible paternal parents for each of the progeny and estimate the degree of selfing in the population. When given the multilocus genotypes of both the progeny and the known maternal parent, Cervus calculates the progeny’s likelihood of decent (LOD) for all nominated candidate paternal parents. An error rate of 1% was specified to allow for possible mistypings, null alleles and mutations, as recommended by Marshall et al., [41]. Consequently, the highest LOD score is not necessarily assigned to the candidate paternal parent simply expressing the lowest number of mismatches. The allele frequencies of all sampled individuals are also taken into account and the highest LOD score therefore reflects the candidate parent statistically most likely to be the true paternal parent. Delta scores (the difference in LOD scores between the first and second most likely candidate parents) are also calculated by CERVUS and are a measure of the reliability of assigning parentage to the most likely candidate. Based on the simulation of a large number of parentage tests, strict (95%) and relaxed (80%) confidence levels were applied to the delta values to indicate the probability that true parentage has been assigned, while taking into account the genotype of the known maternal parent [41]. All other sampled adults (6) were included as possible candidate paternal parents for all progeny sampled (140). In addition the known maternal parents were also included as possible paternal parents, to allow for possible selfing. Therefore, for each of the 140 progeny sampled, there were 7 possible paternal parents screened. The significance values of the delta scores were calculated based on an estimate that 23% of the adult flowering plants in the natural population were sampled (7 out of 30 recorded at the time of field collection).
The software Gerud version 2.0 [42] was used to estimate the minimum number of paternal parents contributing to the multi-locus progeny arrays. The progeny from each known maternal parent was analysed separately. Based on the multi-locus genotype of the known maternal parent and all its progeny, the program simulates all possible paternal genotypes. The program then constructs sets of these genotypes that require the minimum number of individuals acting as paternal parents and identifies the number of progeny fathered by each potential paternal parent. In cases where multiple minimum-father solutions are possible, the program lists all paternal genotypic combinations and ranks the solutions based on relative probability. The program can identify at most six paternal parents contributing to the progeny genotypes, however detection of each additional paternal genotype and the inclusion of each additional locus increases the computational requirements of the program exponentially.
Given the large increase in the number of alleles identified in the progeny in comparison to their known maternal parent (Table 1), we expected that several paternal parents would be contributing to each family group. We therefore limited the number of loci used in this analysis (as recommended in Jones [42]) to the two loci (X1 and X4) with the highest exclusion probabilities calculated in Gerud. Because Gerud is unable to deal with missing data, where there was failed allele amplification at either locus X1 or X4 for any of the progeny, that individual was removed from the analysis.
3. Results
The heights of the seven adult plants sampled, ranged from 25 cm to 86 cm. (i.e. ~25 to 86 years old [43]). Spike length ranged from 36 cm to 108 cm in length and circumference from 13.5 to 18 cm. Using an average of 2.1 seeds per cm2 [25] seed set in these plants ranged from just over 1000 seeds to approximately 4000 seeds.
The average number of alleles across all four loci was 6.75 for the maternal parents and 14 for the progeny. While all alleles found in the maternal parents were also found in progeny, the number of alleles found in the progeny was 47% to 67% times greater than the number found in the maternal parents (Table 1). Two of the four loci (X4 and X9) were found to significantly deviate from HWE (p < 0.01). Observed heterozygosity was consistently lower than expected across all four loci and FIS values were small and positive (Table 1). After adjustment for multiple comparisons (n = 6, alpha = 0.05) Locus X4 showed significant linkage disequilibrium with loci X5 and X9. All other locus pairs were in linkage equilibrium. Pairwise relatedness “r” [37] values between adult plants ranged from 0 to 0.3. As expected the mean relatedness “r” [37] was greater between a known maternal parent and its progeny (0.44 - 0.55), than between each maternal parent and the progeny of another (0.23 to –0.24).
Significant positive spatial structure was found among the progeny within the 5 m distance class and significant negative spatial structure was found at eight of the remaining nine distance classes (Figure 2). Measures of genetic similarity (r) between pairs of individual progeny ranged from 0.197 in the 5 m distance class to –0.079 in the 45 m distance class.
Six of the 140 progeny were assigned one of the seven adult plants as a likely paternal parent at a 95% confidence level, based on the delta scores calculated in Cer-
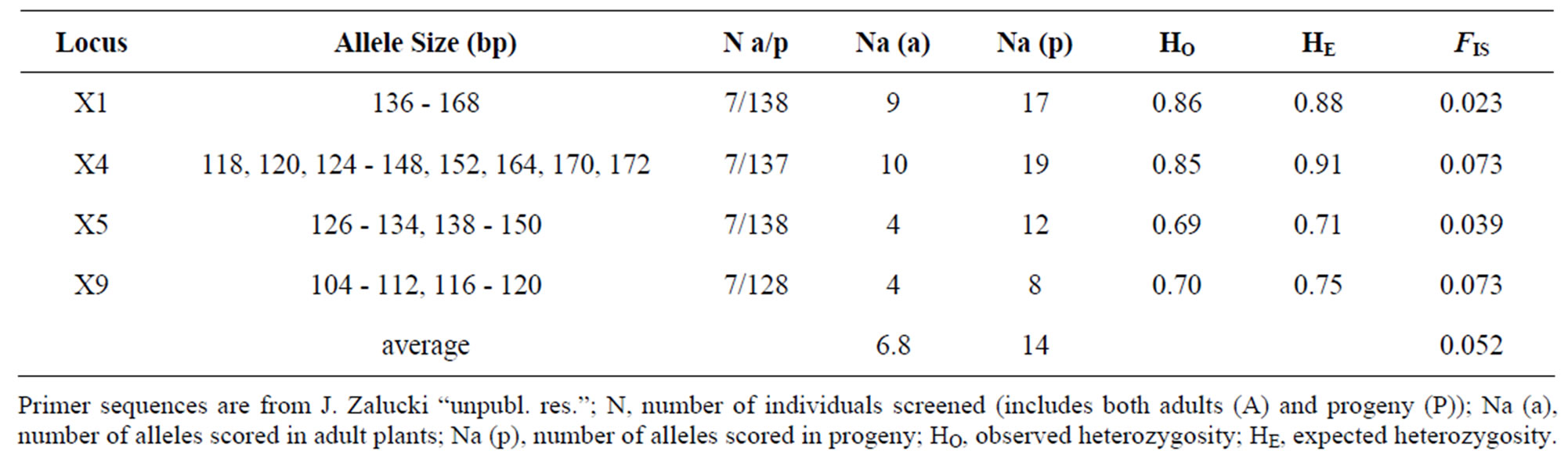
Table 1. Characteristics of four dinucleotide microsatellite loci.
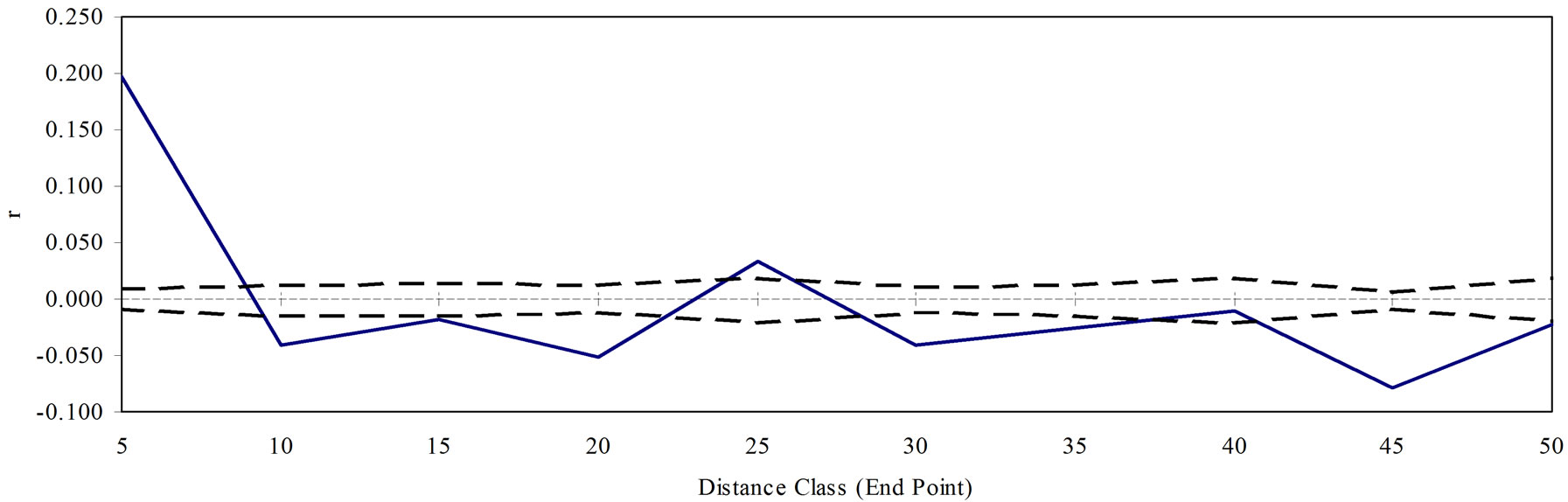
Figure 2. Autocorrelogram showing the spatial genetic structure among progeny within successive 5 m distance classes. The dashed horizontal line (with distance tic marks) represents no spatial genetic structure (r = 0), the two dashed lines represent the upper and lower confidence limits that bound the 95% confidence interval about the null hypothesis of no spatial structure for the combined data set as determined by 1000 permutations. The solid line represents the data set autocorrelation.
vus [41]. Another seven progeny were assigned a paternal parent (from the seven sampled adults) at an 80% confidence level (Table 2). Therefore, for 127 of the 140 progeny sampled there was less than an 80% chance that any of the adult plants sampled was more likely than another to be the paternal parent (Table 2) suggesting outcrossing is high. All 13 progeny assigned a paternal parent had no allelic mismatches at any loci between progeny, the known maternal parent and the first paternal candidate. Zero or one mismatches were associated with the second paternal candidate.
No progeny from E15 was assigned a paternal parent from the seven adult plants sampled (Table 2). In addition, E15 was not assigned as a likely paternal parent for any of the assigned progeny. E10 was identified as a paternal parent for two of its own seed (i.e. selfed), however it was not assigned as a likely paternal parent to any of the other progeny. Pollen was moved among all other adult plants (Table 2).
Of the 140 progeny analysed four showed evidence of potential selfing. Two of these were at 95% confidence and both from maternal parent E10, and two at 80% confidence, both from maternal parent E7. This suggests that a conservative estimate of the populations outcrossing rate would be around 97% for this mating season.
The program Gerud [42] reconstructs the paternal genotypes needed to account for the progeny genotypes, given the known maternal parent genotype. When reconstructing paternal genotype solutions, some progeny may potentially be descended from more than one paternal parent. Therefore the sum of the progeny distributed among putative paternal parents may be higher than the number of progeny included in the analysis. Several equally likely solutions may be found, many varying
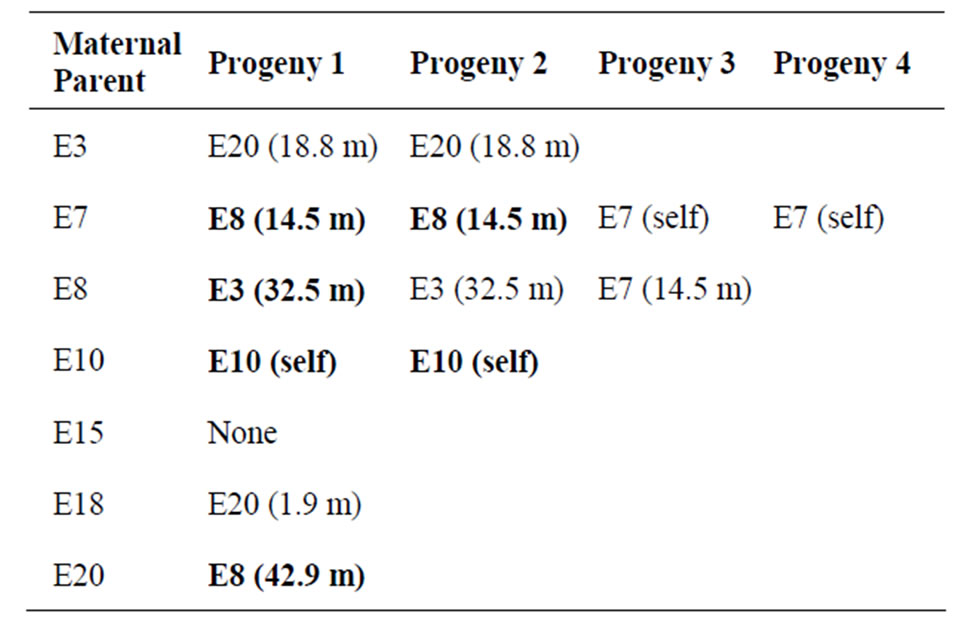
Table 2. Details of the 13 progeny assigned a paternal parent from the seven sampled adult plants (the known maternal parents). Geographic distances between assigned paternal parents and the known maternal parent are given in parentheses and those assigned with 90% confidence are shown in bold.
only in the assigning of some putative fathers as homozygotes rather than heterozygotes.
A minimum of five paternal parents were required to account for the genotypic diversity identified in the progeny from both E10 and E18 maternal parents (Table 3). In each case one putative paternal parent did not dominate in their contribution to progeny genotypes. Six paternal parent genotypes were identified for the progeny for both E8 and E15 maternal parents (Table 3). One putative paternal genotype identified for the progeny of E8, was included in all 15 of the most likely solutions and consistently contributed to more progeny than the other five potential parent genotypes. More than six paternal parents were needed to resolve the genotypic variation identified in the progeny from maternal parents E3, E7 and E20. The Gerud software was unable to reconstruct possible genotypic solutions in these cases.
4. Discussion
The asynchronous flowering of a small number of Xanthorrhoea johnsonii occurring within a much larger population of reproductively mature plants provided a unique opportunity to examine the mating patterns of this species within a closed population for one mating season. The reproductive isolation of the population in this particular season determined that both the maternal and paternal parents for each seed produced in this season could only come from within the closed population. The potential for inbreeding, via either selfing and/or bi-parental inbreeding, was potentially amplified under these conditions, and would be evident in genetic analysis of the progeny.
Xanthorrhoea johnsonii has been shown to be capable of selfing in previous genetic analysis [25]. The potential
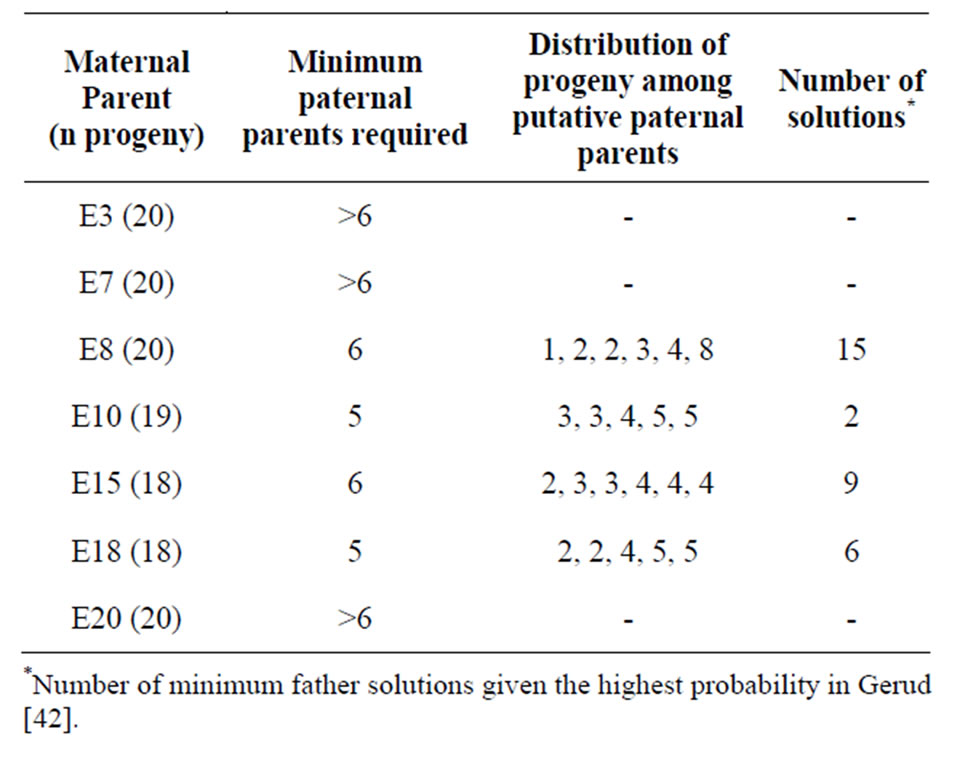
Table 3. The minimum number of paternal parents contributing to the multi-locus progeny arrays.
for selfing may be heightened if pollinators spend more time on single plants due to the fewer numbers of flowering plants available [18] . Bi-parental inbreeding is possible if some spatial genetic structuring among adult plants already exists due to historic mating patterns, dispersal and recruitment events. Plant dispersal can occur during two independent life cycle stages: 1) pollen dispersal before fertilisation, and 2) seed dispersal after fertilisation and embryo development. Gene flow can therefore occur due to the immigration of paternal genotypes through pollen dispersal and/or the immigration of both maternal and paternal genotypes via seed dispersal/immigration [44,45].
In species that exhibit only localised seed dispersal, fine scale differentiation between family groups often occurs. This localised spatial genetic structuring can occur in conjunction with long-range pollen dispersal that can result in simultaneous broad scale homogenisation of variation between geographically distant populations [44, 45]. Little is known about the degree of spatial genetic structure that exists in adult populations of X. johnsonii. However, this species does exhibit a seemingly passive mode of seed dispersal, [26] where large quantities of full and/or half sib seeds are initially shed in close proximity to the maternal parent plant (localised seed dispersal), which suggests that some spatial genetic structure among the adult plants could be expected. Under conditions where neighbouring plants are more likely to be closely related, the potential for bi-parental inbreeding would be further enhanced in a closed population where pollen could only be sourced from among those neighbouring plants, as was the case in this study. In this study any effect of immigrant seed into the population was avoided as all seed was collected directly from the maternal plants inflorescence.
4.1. Genetic Variability and Spatial Patterns
The number of alleles across the four loci in the adult plants ranged from four to nine alleles. This suggests that the sampled adult plants were quite diverse and unrelated. Plants were distributed throughout 0.2 ha and reflected a range of geographic proximities to one another. Plant E18 showed a potential half sib relationship (r ~ 0.25) with plant E20 (1.8 m apart) and plant E7 27.2 m apart. However all other pairwise relatedness values indicated that adult plants were unrelated to each other (r ~ 0) regardless of geographic proximity. Observed and expected heterozygosity was not calculated for the adult plants alone. However, in a previous genetic study of four populations of X. johnsonii, observed/expected heterozygosities (based on allozyme data) ranged from 0.33/ 0.39 to 0.42/0.53, with no significant deviations from Hardy-Weinberg equilibrium [25].
The maintenance of high levels of genetic diversity among the adult plants within close proximity to each other, and across the population as a whole, is likely the result of several different population processes facilitateing historic gene flow through seed and pollen dispersal: 1) it suggests that historic pollen movement must have consistently covered distances at least greater than nearest neighbour; 2) secondary seed dispersal may be more prevalent than currently understood; and 3) although a large number of halfand full-sib seeds are passively deposited within close proximity to a maternal plant each season, only a very small number of these establish during a recruitment event. Temporally patchy recruitment in this way would limit the formation of family group structure, as found in this population.
The age of individual X. johnsonii can be estimated using the height of the plant as a surrogate measure, and both the sampled and unsampled plants in this population reflected a wide range of ages (~25 to 86 years). The varied age of the plants within the population suggests that recruitment events are sporadic over a number of years and are not the result of a mass recruitment event within one season. The latter event would produce an even age stand of individuals which is not evident in this or other species within the genus [26]. The lack of relatedness among the sampled adult plants of various ages suggests that no single pollen donor consistently contributed to seed/progeny across several seasons. No full sib relationships were identified among the adult plants in this study.
The average number of alleles (14) found in the progeny was twice that found in the sampled adults (6.72). The only viable source of pollen was from plants within the population and therefore these additional alleles must have been contributed to the progeny from unsampled adult plants acting as pollen donors/fathers. This indicates that the extant adult population is likely even more genetically diverse than reflected in our limited sample of adults. Although the observed heterozygosity was lower than expected at all four loci (Table 2), the very small FIS values suggests a lack of certain heterozygote genotypes, rather than an excess of homozygotes caused by inbreeding or null alleles. A lack of some allele combinations in heterozygote progeny is expected when several paternal parents contribute new and different alleles to the progeny array from one maternal parent.
Although no evidence of inbreeding was found among the sampled adult plants, both bi-parental inbreeding and selfing were potentially heightened within the sampled progeny due to the effective isolation of the population during the flowering season. The unseasonable timing of the flowering event provided potential pollinators with a concentrated and limited resource of both pollen and nectar to contribute to the progeny/seed. Our analysis of the selfing rates and spatial autocorrelation of progeny arrays allowed us to test our hypothesis of potential genetic structuring in this isolated population.
The estimated outcrossing rate for this population was quite high at 97%. Only four of the 140 progeny were identified as possible selfing events (two individuals at 95% and two individuals at 80% confidence) (Table 2). These progeny were from only two maternal parents. In populations with low genetic diversity, true selfing may be difficult to distinguish from bi-parental inbreeding between closely related individuals. This is unlikely to be the case in this study. A previous study of this species based on allozyme data also found high outcrossing rates [25]. During normal flowering seasons in large populations with high flowering density, outcrossing rates were on average 99%. In populations where the individual plants were more isolated (approximately 50 m between nearest neighbours) and flowering density was therefore lower, outcrossing rates decreased slightly to 97% [25].
A strong positive genetic structure within the first two distance classes (up to 10 metres) was found in the progeny analysis (Figure 2). This was expected within the first distance class (up to 5 m) as all progeny from a given maternal parent were assigned that maternal plants geographic co-ordinates. Individuals within the first distance class of 5 m were therefore predominantly from the same maternal parent and would reflect at least half-sib relationships which would generate significant positive spatial structure in the smallest distance classes of the autocorrelation analysis. However, two maternal plants E18 and E20, occurred only 1.85 m apart. Therefore, comparisons of individuals between these two family groups were included within the first distance class (5 m). This suggests some positive genetic structure beyond the half-sib relationships of progeny from the same maternal parent.
Any family group structure in the progeny disappeared within only 10 m (Figure 2). This reflects the high levels of diversity found in the seed generation as a whole, but also highlights the lack of any apparent nearest-neighbour pattern of pollen transfer. Although seed dispersal may be limited, pollen movement must be more widespread (i.e. not only nearest-neighbour) in order to maintain high diversity and limit family spatial structure. The isolation of this population in this particular breeding season could have promoted inbreeding in the progeny, however this was not the case. The lack of genetic structure found in the progeny (in distance classes > 10 m) also shows that the limited pollen pool and small population size in this particular breeding season has not promoted inbreeding.
4.2. Pollen Donors and Movement
Given that a 47% - 67% increase in allele numbers was found in the progeny compared to the maternal parents, it was apparent that more than one father had contributed to the progeny of each maternal parent. As maternal genotypes were known, the number of paternal parents needed to account for each family progeny array was then assessed. The minimum number of pollen donors required for any of the seven family groups (of 20 seeds) was five, with three of these families requiring at least six paternal parents (Table 3). The distribution of progeny assigned to each of the putative paternal parent genotypes was relatively even i.e. no single paternal multilocus genotype was a dominant contributor to the progeny (Table 3). Due to the 3 - 5 year flowering cycle exhibited by this species, it is likely that, historically, it has not been the same combinations of plants contributing to seed set each year. This will help mitigate against loss of genetic variability in this species.
The need for multiple paternal parents to resolve the multi-locus genotypes of the progeny within each family group suggests that many plants within the population are acting independently as both maternal and paternal parents. From the seven sampled maternal parents in this study, five (71%) were identified as probable paternal parents for at least one progeny from another sampled adult plant (Table 2). Pollen transfer in these cases occurred across a range of distances, from as little as 1.9 m from plant E20 to plant E18, up to 43 m from plant E8 to plant E20 (Table 2). Given the assumed limited and passive mode of seed dispersal in this species, multiple pollen donors donating pollen over a wide range of distances is likely the predominant process maintaining high genetic diversity, and limiting family group sub-structuring. The need for multiple paternal parents also suggests that either a few individual pollinators are transferring mixed pollen loads, or numerous pollinators are involved in moving pollen among the plants.
As populations become smaller and more geographically isolated from one another through increased habitat fragmentation in urbanised areas, the potential for inbreeding and subsequent loss of genetic diversity is potentially heightened [3-6] . Prior to this study, the only previous estimates of outcrossing rates for this species were calculated for several large populations [25]. Our estimates of high levels of genetic diversity and a high outcrossing rate corroborate these findings but are based on a much smaller spatial scale (0.2 hectare). Our study suggests that family group substructure may not be a feature of X. johnsonii populations, that individual plants produce genetically diverse progeny within a breeding season and that there is no evidence of single plants acting as principle pollen donors. This is an encouraging finding for this species as it is constantly under threat due to habitat fragmentation. As this study focused on only a single population, and a small number of adults, future study of paternal parentage (pollen donors) within geographically fragmented populations warrants study. The sporadic and seemingly low recruitment rates could be both a help and a hindrance for this species in the future. The low recruitment rates keep the genetic diversity of the population high, with few full or half sibs being established in close proximity. But the low recruitment rates could become a problem if their frequency declines. Changes in micro-climatic conditions could change germination rates and recruitment events. As X. johnsonii is long-lived, this problem may not be evident in the short term. It would become a problem for the species if recruitment were to fail over the course of several decades.
5. Acknowledgements
The molecular work for this study was done in the Molecular Ecology Lab at Griffith University and we thank Professor Jane Hughes for her guidance. David Gopurenko and Jemma Sommerville were responsible for the development of this library and running the samples. Special thanks to Professor Meron Zalucki for helpful comments on this manuscript. Valuable comments from four anonymous referees improved this manuscript.
REFERENCES
- I. Bartish Peterson and J. Peterson, “Effects of Population Size on Genetic Diversity, Fitness and Pollinator Community Composition in Fragmented Populations of Anthericum liliago L,” Plant Ecology, Vol. 198, No. 1, 2008, pp. 101-110. doi:10.1007/s11258-007-9388-4
- N. J. Ouborg, P. Vergeer and C. Mix, “The Rough Edges of the Conservation Genetics Paradigm for Plants,” Journal of Ecology, Vol. 94, No. 6, 2006, pp. 1233-1248. doi:10.1111/j.1365-2745.2006.01167.x
- G. Caughley, “Directions in Conservation Biology,” Journal of Animal Ecology, Vol. 63, No. 2, 1994, pp. 215-244. doi:10.2307/5542
- P. Crnokrak and D. A. Roff, “Inbreeding Depression in the Wild,” Heredity, Vol. 83, No. 3, 1999, pp. 260-270. doi:10.1038/sj.hdy.6885530
- N. C. Ellstrand and D. R. Elam, “Population Genetic Consequences of Small Population-Size—Implications for Plant Conservation,” Annual Review of Ecology and Systematics, Vol. 24, No. 1, 1993, pp. 217-242. doi:10.1146/annurev.es.24.110193.001245
- D. A. Falk and K. E. Holsinger (Eds.), “Genetics and Conservation of Rare Plants,” Oxford University Press, New York, 1991.
- R. Zhao, H. Xia and B. R. Lu, “Fine-Scale Genetic Structure Enhances Biparental Inbreeding by Promoting Mating Events between More Related Individuals in Wild Soybean (Glycine Soja; Fabaceae) Populations,” American Journal of Botany, Vol. 96, No. 6, 2009, pp. 1138- 1147. doi:10.3732/ajb.0800173
- S. C. H. Barrett and J. R. Kohn, “Genetic and Evolutionary Consequences of Small Population Size in Plants: Implications for Conservation,” In: D. A. Falk and K. E. Holsinger, Eds., Genetics and Conservation of Rare Plants Oxford University Press, New York, 1991, pp. 3-30.
- O. Honnay and H. Jacquemyn, “Susceptibility of Common and Rare Plant Species to the Genetic Concequences of Habitat Fragmentation,” Conservation Biology, Vol. 21, No. 3, 2006, pp. 823-831. doi:10.1111/j.1523-1739.2006.00646.x
- A. Young, T. Boyle and T. Brown, “The Population Genetic Consequences of Habitat Fragmentation for Plants,” Trends in Ecology and Evolution, Vol. 11, No. 10, 1996, pp. 413-418. doi:10.1016/0169-5347(96)10045-8
- Q. F. Geng, C. L. Lian, S. Goto, J. M. Tao, M. Kimura, M. D. S. Islam and T. Hogetsu, “Mating System, Pollen and Propagule Dispersal, and Spatial Genetic Structure in a High-Density Population of the Mangrove Tree Kandelia Candel,” Molecular Ecology, Vol. 17, No. 21, 2008, pp. 4724-4739. doi:10.1111/j.1365-294X.2008.03948.x
- Y. Isagi, D. Saito, H. Kawaguchi, R. Tateno and S. Watanabe, “Effective Pollen Dispersal Is Enhanced by the Genetic Structure of an Aesculus Turbinata Population,” Journal of Ecology, Vol. 95, No. 5, 2007, pp. 983-990. doi:10.1111/j.1365-2745.2007.01272.x
- T. Kenta, Y. Isagi, M. Nakagawa, M. Yamashita and T. Nakashizuka, “Variation in Pollen Dispersal between Years with Different Pollination Conditions in a Tropical Emergent Tree,” Molecular Ecology, Vol. 13, No. 11, 2004, pp. 3575-3584. doi:10.1111/j.1365-294X.2004.02345.x
- C. F. Williams, “Effects of Floral Display Size and Biparental Inbreeding on Outcrossing Rates in Delphinium barbeyi (Ranunculaceae),” American Journal of Botany, Vol. 94, No. 10, 2007, pp. 1696-1705. doi:10.3732/ajb.94.10.1696
- M. B. Routley and B. C. Husband, “The Effect of Protandry on Siring Success in Chamerion angustifolium (Onagraceae) with Different Inflorescence Sizes,” Evolution, Vol. 57, No. 2, 2003, pp. 240-248.
- G. H. Pyke, “Optimal Foraging: Movement Patterns of Bumblebees between Infloresences,” Theoretical Population Biology, Vol. 13, 1978, pp. 72-98. doi:10.1016/0040-5809(78)90036-9
- H. S. Ishii, Y. Hirabayashi and G. Kudo, “Combined Effects of Inflorescence Architecture, Display Size, Plant Density and Empty Flowers on Bumble Bee Behaviour: Experimental Study with Artificial Inflorescences,” Oecologia, Vol. 156, No. 2, 2008, pp. 341-350. doi:10.1007/s00442-008-0991-4
- J. D. Karron, K. G. Holmquist, R. J. Flanagan and R. J. Mitchell, “Pollinator Visitation Patterns Strongly Influence among-Flower Variation in Selfing Rate,” Annals of Botany, Vol. 103, No. 9, 2009, pp. 1379-1383. doi:10.1093/aob/mcp030
- M. O. Johnston, E. Porcher, P. Ä. Cheptou, C. G. Eckert, E. Elle, M. A. Geber, S. Kalisz, J. K. Kelly, D. A. Moeller, M. Vallejo-Marín and A. A. Winn, “Correlations among Fertility Components Can Maintain Mixed Mating in Plants,” The American Naturalist, Vol. 173, No. 1, 2009, pp. 1-11. doi:10.1086/593705
- J. P. Gonzalez-Varo, R. G. Albaladejo and A. Aparicio, “Mating Patterns and Spatial Distribution of Conspecific Neighbours in the Mediterranean Shrub Myrtus communis (Myrtaceae),” Plant Ecology, Vol. 203, No. 2, 2009, pp. 207-215. doi:10.1007/s11258-008-9534-7
- J. C. Avise, “Molecular Markers, Natural History and Evolution,” Chapman & Hall, New York, 1994, p. 511. doi:10.1007/978-1-4615-2381-9
- S. G. Michalski and W. Durka, “High Selfing and High Inbreeding Depression in Peripheral Populations of Juncus Atratus,” Molecular Ecology, Vol. 16, No. 22, 2007, pp. 4715-4727. doi:10.1111/j.1365-294X.2007.03547.x
- O. J. Hardy, S. C. Gonzalez-Martinez, B. Colas, H. Freville, A. Mignot and I. Olivieri, “Fine-Scale Genetic Structure and Gene Dispersal in Centaurea corymbosa (Asteraceae). II. Correlated Paternity within and among Sibships,” Genetics, Vol. 168, No. 3, 2004, pp. 1601-1614. doi:10.1534/genetics.104.027714
- M. Gaudeul and I. Till-Bottraud, “Low Selfing in a MassFlowering, Endangered Perennial, Eryngium alpinum L. (Apiaceae),” American Journal of Botany, Vol. 90, No. 5, 2003, pp. 716-723. doi:10.3732/ajb.90.5.716
- J. M. Zalucki, “A Study of the Repoductive Biology of Xanthorrhoea johnsonii (Xanthorrhoaceae) in Toohey Forest,” Ph.D. Thesis, Griffith University, Nathan, 1998.
- B. B. Lamont, R. Wittkuhn and D. Korczynskyj, “Turner Review No. 8 Ecology and Ecophysiology of Grasstrees,” Australian Journal of Botany, Vol. 52, No. 5, 2004, pp. 561-582. doi:10.1071/BT03127
- A. T. Lee, “Notes on Xanthorrhoea in Eastern Australia,” Contributions from the N.S.W. National Herbarium, Vol. 4, 1966, pp. 35-54.
- D. J. Bedford, A. T. Lee, T. D. Macfarlane, R. J. F. Henderson and A. S. George, “Xanthorrhoeaceae,” In: A. E. Orchard, Ed., Flora of Australiam, Vol. 46, Australian Government Publishing Service, Canberra, 1986, pp. 88-171.
- I.A. Staff, “The Fruits and Seed Productivity in Xanthorrhoea Species,” Proceedings of the Linnean Society of N.S.W., Vol. 100, 1975, pp. 95-102.
- C. R. Loomis, G. G. Shipley and D. M. Small, “PhaseBehavior of Hydrated Cholesterol,” Journal of Lipid Research, Vol. 20, No. 4, 1979, pp. 525-535.
- S. D. Song, R. A. I. Drew and J. M. Hughes, “Multiple Paternity in a Natural Population of a Wild Tobacco Fly, Bactrocera cacuminata (Diptera: Tephritidae), Assessed by Microsatellite DNA Markers,” Molecular Ecology, Vol. 16, No. 11, 2007, pp. 2353-2361. doi:10.1111/j.1365-294X.2007.03277.x
- C. van Oosterhout, W. F. Hutchinson, D. P. M. Wills and P. F. Shipley, “Micro-Checker User Guide,” The University of Hull, Hull, 2003.
- R. Chakraborty, M. Deandrade, S. P. Daiger and B. Budowle, “Apparent Heterozygote Deficiencies Observed in DNA Typing Data and Their Implications in Forensic Applications,” Annals of Human Genetics, Vol. 56, 1992, pp. 45-57. doi:10.1111/j.1469-1809.1992.tb01128.x
- J. F. Y. Brookfield, “A Simple New Method for Estimating Null Allele Frequency from Heterozygote Deficiency,” Molecular Ecology, Vol. 5, No. 3, 1996, pp. 453- 455.
- M. Raymond and F. Rousset, “Genepop (Version 1.2): Population Genetics Software for Exact Tests and Ecumenicism,” Journal of Heredity, Vol. 86, No. 3, 1995, pp. 248-249.
- B. S. Weir and C. C. Cockerham, “Estimating F-Statistics for the Analysis of Population Structure,” Evolution, Vol. 38, No. 6, 1984, pp. 1358-1370. doi:10.2307/2408641
- D. C. Queller and K. F. Goodnight, “Estimating Relatedness Using Genetic-Markers,” Evolution, Vol. 43, No. 2, 1989, pp. 258-275. doi:10.2307/2409206
- P. E. Smouse and R. Peakall, “Spatial Autocorrelation Analysis of Individual Multiallele and Multilocus Genetic Structure,” Heredity, Vol. 82, No. 5, 1999, pp. 561-573. doi:10.1038/sj.hdy.6885180
- R. Peakall and P. E. Smouse, “Genalex 6: Genetic Analysis in Excel. Population Genetic Software for Teaching and Research,” Molecular Ecology Notes, Vol. 6, No. 1, 2006, pp. 288-295. doi:10.1111/j.1471-8286.2005.01155.x
- R. Peakall, D. Ebert, L. J. Scott, P. F. Meagher and C. A. Offord, “Comparative Genetic Study Confirms Exceptionally Low Genetic Variation in the Ancient and Endangered Relictual Conifer, Wollemia nobilis (Araucariaceae),” Molecular Ecology, Vol. 12, No. 9, 2003, pp. 2331-2343. doi:10.1046/j.1365-294X.2003.01926.x
- T. C. Marshall, J. Slate, L. E. B. Kruuk and J. M. Pemberton, “Statistical Confidence for Likelihood-Based Paternity Inference in Natural Populations,” Molecular Ecology, Vol. 7, No. 5, 1998, pp. 639-655. doi:10.1046/j.1365-294x.1998.00374.x
- A. G. Jones, “Gerud 2.0: A Computer Program for the Reconstruction of Parental Genotypes from Half-Sib Progeny Arrays with Known or Unknown Parents,” Molecular Ecology Notes, Vol. 5, No. 3, 2005, pp. 708-711. doi:10.1111/j.1471-8286.2005.01029.x
- A. Bulow-Olsen, J. Just and M. J. Liddle, “Growth and Flowering History of Xanthorrhoea johnsonii Lee (Liliaceae) in Toohey Forest Queensland,” Botanical Journal of the Linnean Society, Vol. 84, No. 3, 1982, pp. 195-207. doi:10.1111/j.1095-8339.1982.tb00534.x
- J. L. Hamrick, D. A. Murawski and J. D. Nason, “The Influence of Seed Dispersal Mechanisms on the GeneticStructure of Tropical Tree Populations,” Vegetatio, Vol. 108, No. 1, 1993, pp. 281-297.
- J. L. Hamrick and J. D. Nason, “Gene Flow in Forest Trees,” In: A. Young, D. Boshier and T. Boyle, Eds., Forest Conservation and Genetics: Principles and Practice, CSIRO Pub., Collingwood, 2000, pp. 81-90.
NOTES
*Corresponding author.

