American Journal of Plant Sciences
Vol.3 No.1(2012), Article ID:16625,11 pages DOI:10.4236/ajps.2012.31001
New Pineapple Somaclonal Variants: P3R5 and Dwarf
![]()
Laboratory for Plant Breeding, Centro de Bioplantas, Universidad de Ciego de Ávila, Ciego de Ávila, Cuba.
Email: pfa_guillermo@agronomia.unica.cu
Received July 5th, 2011; revised August 2nd, 2011; accepted September 9th, 2011
Keywords: Ananas comosus (L.) Merr.; Genotype Variation; Phenotype Variation
ABSTRACT
The Food and Agriculture Organization has highlighted pineapple as one of the most important tropical fruits. Since classical pineapple breeding is difficult, biotechnology has emerged as an attractive instrument. We obtained two new pineapple somaclonal variants derived from in vitro culture of cv. Red Spanish Pinar: P3R5 and Dwarf. The AFLP analysis revealed an existing genetic distance. So far 44 phenotype indicators selected due to their relation to a wide range of important agricultural, morphological and physiological processes have been evaluated. P3R5 differed from the donor in 19 variables (19/44; 43.18%), while Dwarf varied in 31 indicators (31/44; 70.45%). The number of shoots was significantly different among the three plant materials. Dwarf showed two shoots per plant while P3R5 and the donor did not form any shoots. We also observed that water use efficiency, chlorophyll b concentration, total chlorophyll concentration, transpiration rate, chlorophyll a concentration, thickness of leaf photosynthetic parenchyma, fruit mass with crown, content of free phenolics and superoxide dismutase specific activity were also very different among the three plant materials. The Euclidean distances of each somaclonal variant to the donor plant material taking into consideration the genotype (AFLP) and the phenotype evaluations were also calculated. Regarding the genotype information, P3R5 is separated from cv. Red Spanish Pinar by 2.83 units of Euclidean distance, and Dwarf by 3.00 units. However, the phenotype indicators revealed higher differences: 3.74 in P3R5 and 4.71 in Dwarf. To our knowledge, this is the first report of a comprehensive analysis of pineapple somaclonal variants.
1. Introduction
Pineapple belongs to the Bromeliaceae family and is one of the most economically important tropical fruit. The interest for its production is due to the high cost that reaches in the fresh fruit and industrialized markets, its great food value, its pleasant taste and its beauty for commercialization. The worldwide production in 2008 was 19.16 million tons [1]. Because of this, several research groups are developing basic and applied studies to create new varieties with better agronomic performance. Pineapple breeding using naturally occurring genetic variation and conventional methods has succeeded in several countries.
In 1914, the Pineapple Growers Association of Hawaii started one of earliest and most concerted efforts in pineapple improvement. One of the main objectives was to develop pest and disease resistance in Smooth Cayenne [2]. Many other countries have also started hybridization programmes to develop high-yielding varieties with specific adaptation to their own environments, for instance, in Taiwan [3,4], Malaysia [5-7], The Philippines, Cote d’Ivoire, Puerto Rico, Cuba and Australia [8- 15]. The varieties obtained recently by hybridization programs in Brazil (Ajubá, Imperial), Australia (Aus-Carnival and Aus-Jubilee), USA (Honey Gold and MD2), Martinique (FLHORAN41) and Malaysia (Josapine) [16].
As classical pineapple breeding is extremely laborious and time-consuming [17], biotechnology is an attractive tool for improving elite clones [18-23]. In this context, some results have been obtained with somaclonal variation.
Genetic variation is very important in crop improvement and forms the basis of development of new varieties. Somaclonal variation is a valuable tool in plant breeding wherein variation in tissue culture regenerated plants from somatic cells can be used in the development of crops with novel traits [24]. Larkin and Scowcroft [25] were the first researchers to demonstrate and coin the term somaclonal variation. Variations may pre-exist in the natural population of plants from field collection or genebank or it may develop as a result of tissue culture conditions [26].
In recent years a number of studies have measured, through molecular markers, the extent of somaclonal variation in plants [27]. Lack of polymorphisms associated with in vitro regeneration was reported in tomato [28], Norway spruce [29], oil palm [30], begonia [31], almond [32], and potato [33,34] using RAPD, ISSR and AFLP markers. By contrast, major differences were found in alfalfa [35], in Codonopsis lanceolata [36] and wild pear [37] using RAPD and ISSR markers.
Pineapple somaclonal variations have been previously characterized [38-42]. However, all these studies showed only few characters and were not studied in detail as only the number of leaves per plant; the number of thorns per leaf and leaf color were reported.
The present study culturing in vitro pineapple axillary buds with naphthalene acetic acid and 6-benziladenine for micropropagation; kinetin to induce callus formation; and indole-3-butyric acid and gibberellic acid for plant regeneration, was carried out to check possible genetic alterations in the plants produced. These growth regulators were the only putative mutagenic agents we used.
We obtained two new pineapple somaclonal variants derived from in vitro culture of cv. Red Spanish Pinar [43,44]. This paper shows a broad genotypic and phenotypic analysis of P3R5 and Dwarf somaclonal variants.
2. Materials and Methods
2.1. Plant Material, Media and Culture Conditions Fifty pineapple buds (cv. Red Spanish Pinar, donor) were collected from field-grown plants and cultured following the protocol described by Daquinta and Benegas [45]. Explants were placed in conventional plant containers for micropropagation (300 ml) where 25 ml liquid culture medium fed five explants. The culture medium included Murashige and Skoog salts [46], 100 mg·l–1 myo-inositol, 0.1 mg·l–1 thiamine-HCl, 30 g·l–1 sucrose, 4.4 μM 6- benzyladenine and 5.3 μM naphthalene-acetic acid. Fortythree young pineapple shoots were obtained after 42 d of bud implantation.
2.2. Multiplication Culture Medium
Shoots were transferred to the multiplication culture medium (as described above except: 9.3 μM 6-benziladenine and 1.6 μM naphthalene-acetic acid). They were subcultured at 42 d intervals for 168 d. Twenty-four thousand seven hundred and sixty-eight shoots were then obtained.
2.3. Callus Induction
Three hundred young leaves were randomly selected as explants for callus formation. The culture medium included: Murashige and Skoog salts [46], 100 mg·l–1 myoinositol, 0.1 mg·l–1 thiamine-HCl, 30 g·l–1 sucrose, 29.0 μM naphthalene-acetic acid and 9.7 μM kinetin. The calli were proliferated for 4 months with subcultures every 30 d.
2.4. Plant Regeneration
Five hundred calli (Ø: 3 mm) were randomly selected and transferred to the plantlet regeneration medium: Murashige and Skoog salts [46], 100 mg·l–1 myo-inositol, 0.1 mg·l–1 thiamine-HCl, 30 g·l–1 sucrose, 0.9 μM indole- 3-butyric acid, 1.1 μM 6-benzyladenine and 0.09 μM gibberellic acid. Four hundred and twenty-seven in vitroplantlets were obtained and later hardened in ex vitro for 6 months [47].
2.5. Hardening and Field Conditions
For ex vitro hardening, plantlets were placed in plastic trays containing 82 cm3 of a mixture zeolite + filter cake (1:1). Microject automated irrigations for 25 s every 30 min were applied. Plantlets were kept under a photosynthetic photon flux density of 458 μmol·m–2·s–1. Standard phytosanitary controls were applied. After hardening of in vitro-plantlets, 387 plantlets were transferred to the field environment and asexually propagated for two generations (30 months). The donor cultivar was used as a control. Two phenotype variants were then identified: P3R5 and Dwarf. A more detailed study was carried out to compare these two variants with the donor plant (cv. Red Spanish Pinar, plant material never cultured in vitro). The experiment was developed in the Field Experimental Station at the Bioplant Centre. A random block design was implemented (80 plants/clone). Field management was performed according to instructions recommended by the Cuban Ministry for Agriculture. The most important pineapple phenotypic traits were recorded during 18 months. The agricultural evaluations were made in field conditions.
2.6. Plant Material and DNA Extraction
In the second generation, in order to perform the AFLP characterization, young leaves of the donor genotype (cv. Red Spanish Pinar) and the two variants (P3R5 and Dwarf) were collected. Samples were stored at –20˚C until DNA extraction. Extraction started from 250 mg fresh mass that were finely grounded in liquid nitrogen. Extraction buffer (650 μl) was then added. It included: Tris-Cl (pH 7.5, 50 mmol·l–1), ethylene-diamine-tetraacetic acid (20 mmol·l–1), sodium chloride (0.3 mmol·l–1), sarcosil (2%), sodium dodecyl sulfate (0.5%) and urea (4.8 mmol·l–1). A mixture (650 μl) of phenol:chloroform: isoamyl alcohol (25:24:1, v:v:v) was added. Samples were centrifuged for 15 min at 12,000 rpm at room temperature. Supernatants were collected and the pellets, discarded. Isopropanol (0.8 volumes) was added. Samples were shortly shaken in a vortex and incubated for 10 min at room temperature. Samples were centrifuged for 10 min at 12,000 rpm. Supernatant was discarded and the pellet was washed with ethanol (70%). DNA was dried (vacuum) and dissolved in 50 μl water supplemented with 10 μg·ml–1 RNase A. DNA integrity and purity was checked by electrophoresis in agarose gels (0.8%) stained with ethidium bromide. Concentration was estimated visually by comparison with standards (100 - 1000 ng·μl–1). Concentrations of DNA samples were adjusted to 500 ng·μl–1.
2.7. AFLP Analysis
AFLP technique was carried out [48]. The digestion of genome DNA, the pre-amplification with a selective base and the selective amplification was performed [47]. Autoradiographs were analyzed visually to build a dicotomic numerical matrix: the number one was assigned when the band was present while zero was assigned when absent. We disregarded all weak and low peak AFLP bands. The matrix was processed with the NTSYSpc software [49]. The simple matching index was used to create a matrix of similarity. From this matrix, a matrix of genetic distance was obtained. The UPGMA (Unweighted Pair Group Method with Arithmetic Averages) method was used to generate a dendogram.
2.8. Morphological, Physiological and Biochemical Determination
In a subsequent procedure, the three plant materials were transferred to the Pineapple Germplasm Bank at the Bioplant Centre in a random block design. Plants grew for 6 months and then D leaves [50] of Red Spanish Pinar (donor), P3R5 and Dwarf were collected. Ten plants per genotype were studied (one leaf per plant). The stoma diameter, number of stomata per mm2, diameter of leaf vascular tissue, thickness of the leaf aquiferous parenchyma, and thickness of the leaf photosynthetic parenchyma were measured [51]. The photosynthetic rate, the transpiration rate, the water use efficiency, and the internal leaf CO2 concentration were recorded using a Portable CIRAS-2 Photosynthesis System (Europe, PP Systems, UK); covering with the leaf, the whole area of the cuvette (PLC6, 2.5 cm2). The carbon dioxide concentration and the relative humidity of the air entering the cuvette were 375 μmol·mol−1 and 80% respectively, under environmental temperature (25˚C - 27˚C). Prior to obtaining the experimental data, we measured the maximum light intensity at which photosynthesis was stable which was attained at 600 μmol·m−2·s−1.
To determine the levels of chlorophyll pigments (a, b, total), leaves were thinly grounded in liquid nitrogen. Evaluations were made [52]. Extraction was carried out with 5.0 ml acetone (80%, v:v). The samples were centrifuged (12,000 rpm, 4˚C, 15 min), supernatants collected and absorbances at 647 and 664 nm were recorded.
Contents of malondialdehyde and other aldehydes [53]; and phenolics (cell wall-linked, free, and total were determined [54]. Total protein contents were recorded [55]. Enzymatic activities and specific activities of phenylalanine ammonia-lyase [56], superoxide dismutase [57], and phosphoenol pyruvate carboxylase [58,59] were also measured.
2.9. Data Analysis
The Statistical Package for Social Sciences (Version 8.0 for Windows, SPSS Inc.) was used to perform One-Way ANOVA and Tukey tests (P ≤ 0.05). The Euclidean distances of each somaclonal variant to the donor plant material were calculated. Data were standardized to vary from 0 to 1 [60].
3. Results and Discussion
3.1. Agricultural and AFLP Characterization
Only two plant materials were found to be “solid” somaclonal variants after studying three vegetative generations (P3R5 and Dwarf) and the AFLP analysis, which represents 0.52% (2 somaclonal variants/387 plants transferred to the field). The dendogram generated with the AFLP information revealed an existing genetic distance among the somaclonal variants and the donor plant [43]. The genetic distances among the three plant materials are not too significant. However, as they have different banding patterns, they are different at the genetic level.
The agricultural characterization of the third vegetative generation in Tables 1(a)-(b) and Figure 1 shows that, in comparison with the donor plant (cv. Red Spanish Pinar), the variant P3R5 showed differences in the number of slips and suckers, and in the presence of thorns in the leaves and in the fruit crowns. The somaclonal variant Dwarf was different from the donor plant in regard to the plant height; the peduncle diameter; the number of shoots, slips and suckers; the fruit mass with crown; the number of eyes in the fruit; the fruit height and diameter; the leaf color; the plant architecture; the length of plant generation cycle; and the fruit color and shape (Table 1(a), agricultural characterization of the third vegetative generation).
3.2. Morphological, Physiological and Biochemical Characterization
The morphological, physiological and biochemical characterization of D leaves in Table 1(b) shows that, in comparison with the donor plant (cv. Red Spanish
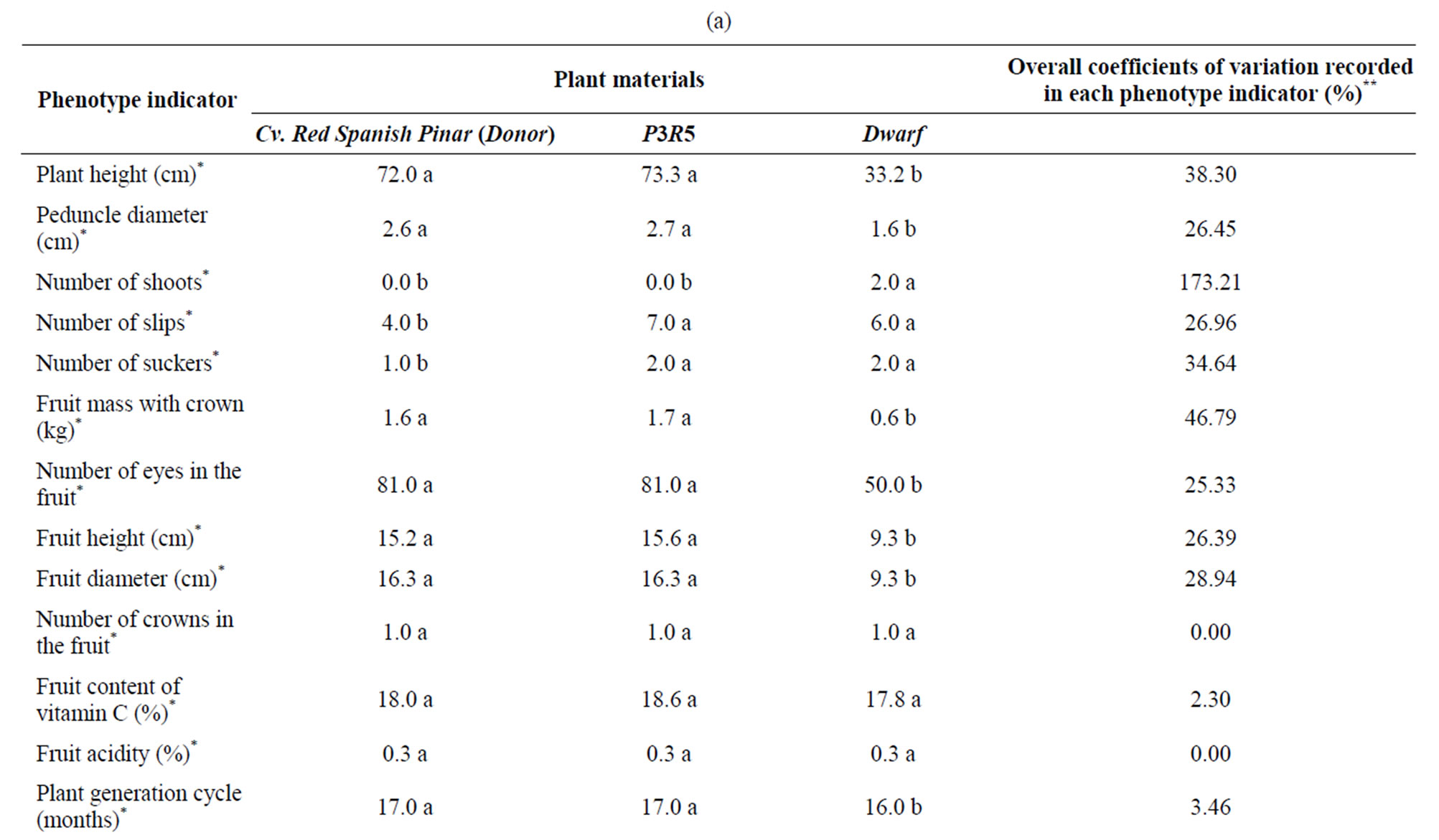
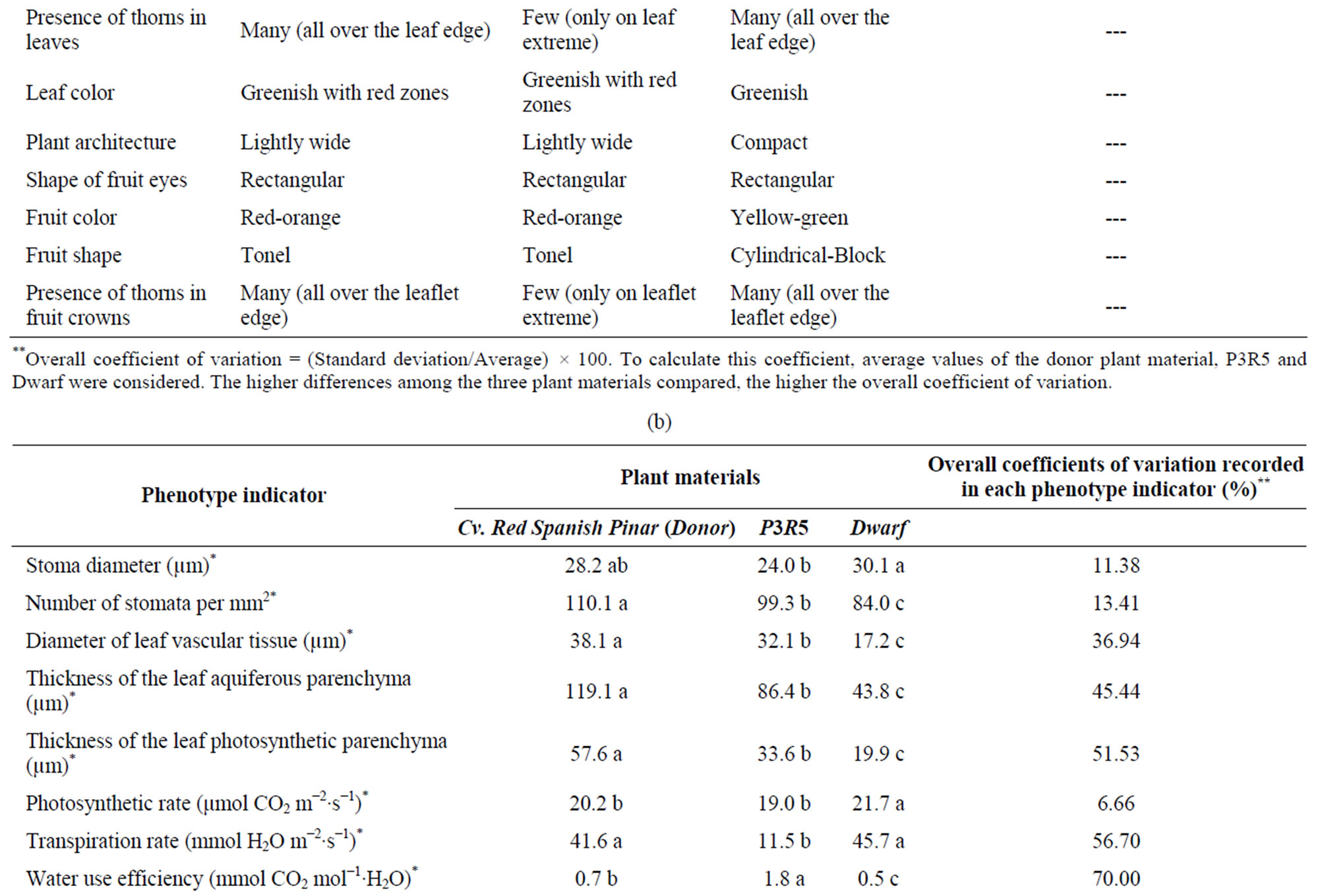
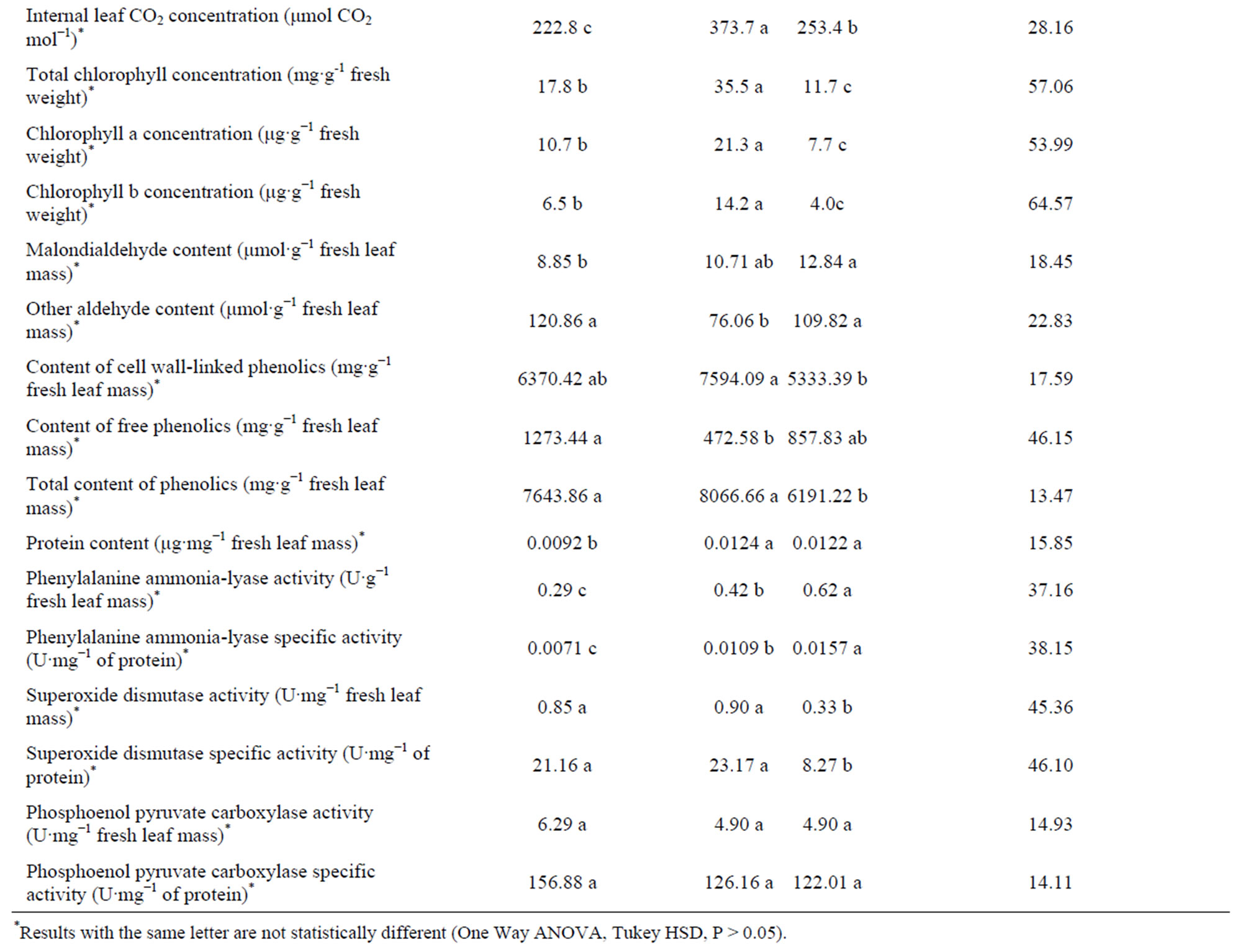
Table 1. Phenotypic characterization of pineapple somaclonal variants. (a) Agricultural characterization of the third vegetative generation; (b) Morphological, physiological and biochemical characterization of D leaves.
Pinar), the variant P3R5 showed statistically significant differences in 15 indicators while Dwarf in 17 variables. Compared to the donor plant, P3R5 somaclonal variant showed significant low values in several aspects, but mainly in the transpiration rate that only reached 28% of the rate in the donor (11.5 mmol H2O m−2·s−1/41.6 mmol H2O m−2·s−1). Moreover, content of free phenolics in P3R5 merely represented 37% (472.58 mg·g−1 fresh leaf mass/1273.44 mg·g−1 fresh leaf mass). Significant increases were also recorded in P3R5 in comparison with cv. Red Spanish Pinar. For instance, the donor showed 39% of the water use efficiency evaluated in P3R5 (0.7 mmol CO2 mol−1 H2O/1.8 mmol CO2 mol−1 H2O) (Table 1).
Comparing Dwarf somaclonal variant with the donor, it only reached 35% of the thickness of the photosynthetic parenchyma of D leaf recorded in the donor (19.9 µm/57.6 µm), 37% of the thickness of the leaf aquiferous parenchyma (43.8 µm/119.1 µm), and 39% of the superoxide dismutase activity and specific activity (0.33 U·mg−1 fresh leaf mass/0.85 U·mg−1 fresh leaf mass, 8.27 U·mg−1
of protein/21.16 U·mg−1 of protein, respectively). On the other hand, the donor plant material only showed about 46% of the phenylalanine ammonia-lyase activity and specific activity (0.29 U·mg−1 fresh leaf mass/0.62 U·mg−1 fresh leaf mass, 0.0071 U·mg−1 of protein/0.0157 U·mg−1 of protein) (Table 1).
Changes in the above mentioned morphological, physiological and biochemical indicators have been frequently studied when plants have been submitted to different sources of stress. Reference [61], recorded the optimization of CO2 gain through stomatal aperture while minimizing water loss in rice. The effects of flooding and drought stress on citrus seedlings physiology were measured [62]. The response of cucumber seedlings to drought stress also were measured [63]. However, to our knowledge, the effects of somaclonal variation on plant physiology have not been deeply studied. Further studies are required to elucidate the mechanisms that explain the differences observed in P3R5 and Dwarf somaclonal variants.
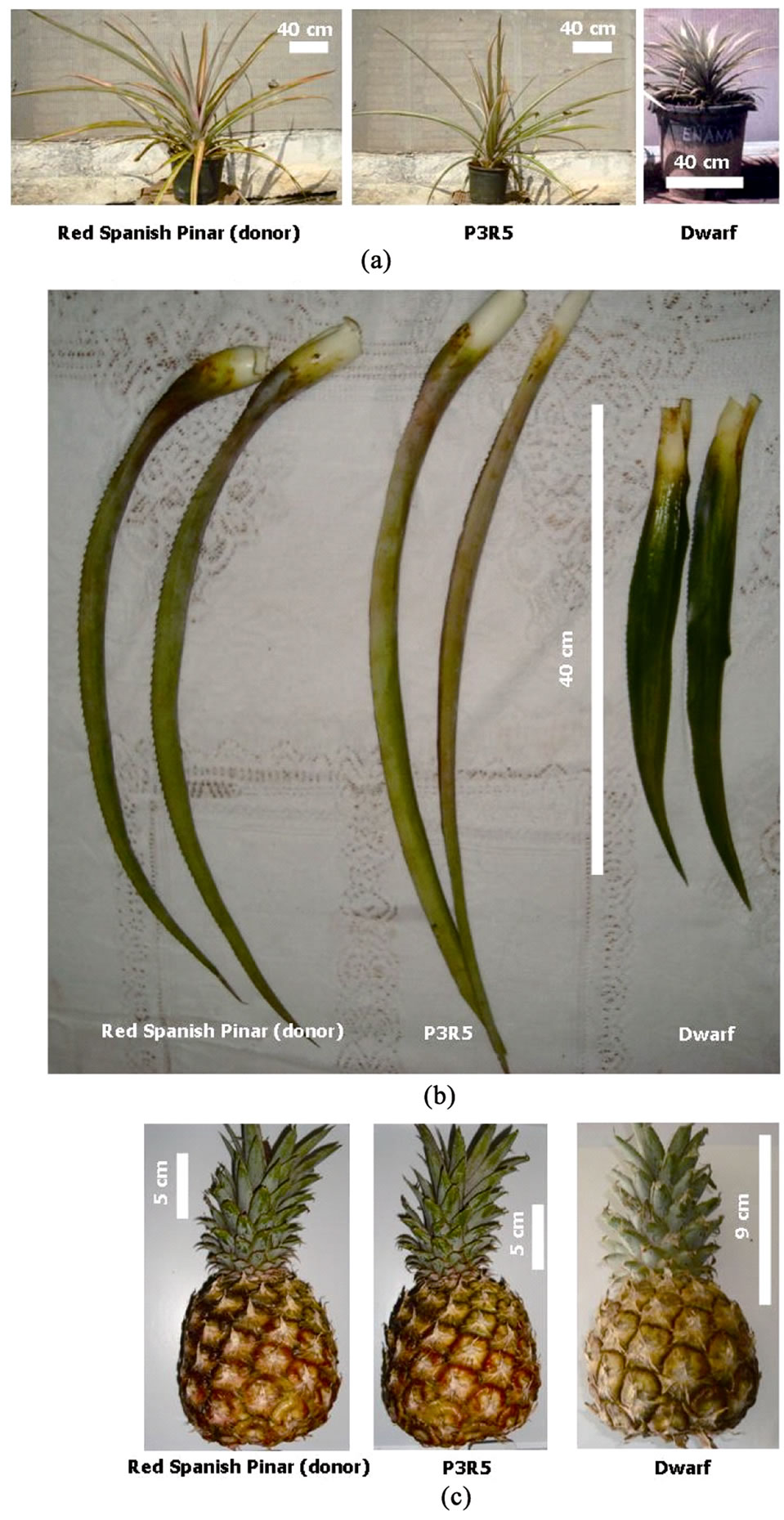

Figure 1. Pineapple material of Red Spanish Pinar (Donor), P3R5 and Dwarf somaclonal variants. (a) Plants at 6 months of growth under controlled environment; (b) D leaves of 6 month-old plants grown under the field environment; (c) Fruits just harvested in the field.
3.3. General Variation: Overall Coefficients of Variation and Euclidean Distances
The overall coefficients of variation in Table 1 indicate that the number of shoots was significantly different among the three plant materials (173.21%).
Dwarf showed two shoots per plant while P3R5 and the donor did not form any shoot. We classified the overall coefficients of variation of the other phenotype indicators in three categories: less than 23%, between 23 and 46%, and over 46%. Then we observed that water use efficiency, chlorophyll b concentration, total chlorophyll concentration, transpiration rate, chlorophyll a concentration, thickness of the leaf photosynthetic parenchyma, fruit mass with crown, free phenolics content and superoxide dismutase specific activity were also very different among the three plant materials. However, other aldehyde content, malondialdehyde content, content of cell wall-linked phenolics, protein content, phosphoenol pyruvate carboxylase activity, phosphoenol pyruvate carboxylase specific activity, total content of phenolics, number of stomata per mm2, stoma diameter, photosynthetic rate, plant generation cycle, fruit content of vitamin C, number of crowns in the fruit and fruit acidity showed low variability.
Table 2 summarizes the phenotypic changes of P3R5 and Dwarf somaclonal variants with respect to the donor plant material. We have used 44 indicators based on a wide range of horticultural and physiological traits. These data clearly show the various aspects where somaclonal variation can occur in pineapple. P3R5 differed from the donor in 19 variables (19/44; 43.18%), while Dwarf in 31 indicators (31/44; 70.45%; Table 2).
Figure 2 shows the Euclidean distances of each somaclonal variant to the donor plant material taking into consideration the genotype (AFLP) and the phenotype evaluations. Regarding the genotype information, P3R5 is separated from cv. Red Spanish Pinar by 2.83 units of Euclidean distance, and Dwarf by 3.00 units. However, the phenotype indicators revealed bigger differences: 3.74 in P3R5 and 4.71 in Dwarf. These figures support the impressive effects on phenotype of small genetic modifications caused by in vitro culture. Authors studied several Syngonium podophylum somaclonal variants within which small genetic differences and significant phenotype modifications were also observed [64]. Similar results were recorded in Actinidia deliciosa somaclonal variants [65].
References [66,67] summarized that 22 cultivars had been released from somaclonal variation with improved traits including yield; plant architecture; colour; pest resistance; salt, heat and freezing tolerance. However, considering that pineapple culture through in vitro derived plants is in practice for a long time (over 20 years in Cuba), we have only these two (P3R5 and Dwarf) variants that are stable and thus somaclonal variation in this crop should be considered a rare event.
Somaclonal variation has been associated with changes in chromosome number and structure, point mutations, DNA methylation [68], transposon activation, deletion, genome rearrangement, polyploidy, or nucleotide substitution [69]. However, not much has been published about the effects of somaclonal variation at morphological and physiological levels.
At this point of our investigation, it is difficult to say
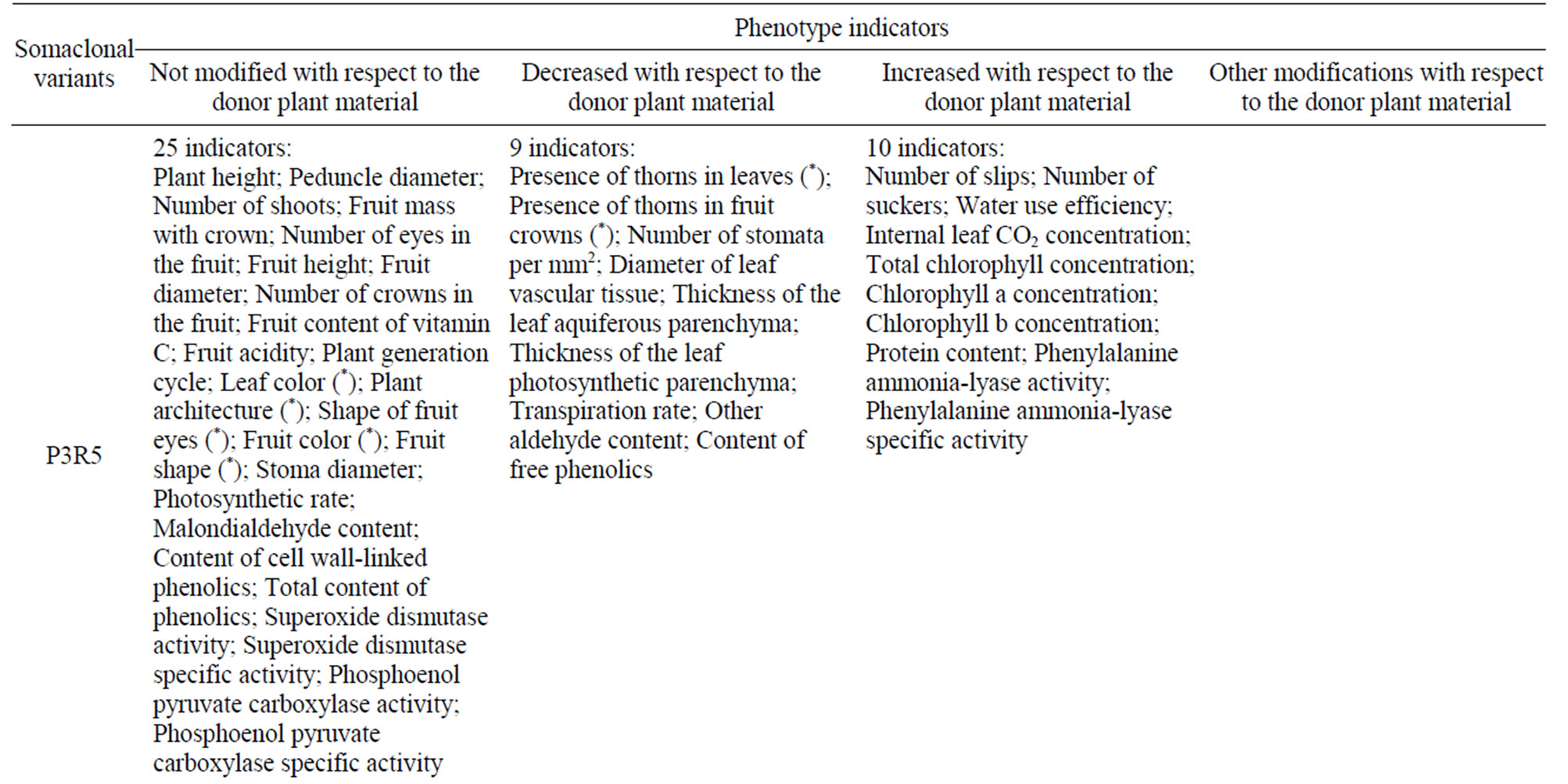
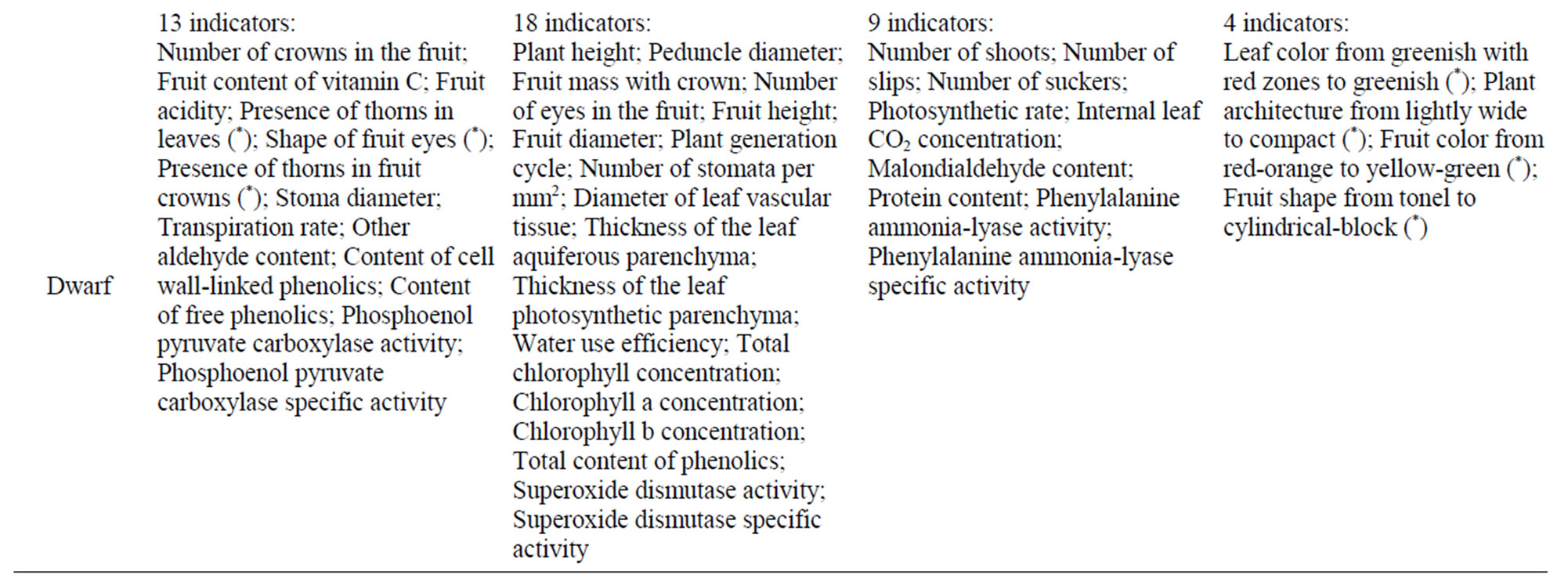


Table 2. Summary of phenotypic modifications of P3R5 and Dwarf somaclonal variants. Classification supported by One Way ANOVA and Tukey HSD (P = 0.05) but asterisks indicate qualitative analysis (Table 1).
which genes are involved in the morphological and physiological changes that were observed in this study. This paper differs from our previous one [43] largely in the addition of a large amount of phenotypic data. The additional data make it less likely that AFLP comparisons of the wild-type line and the two mutants will lead to tagging of the genes for thorn production and dwarf stature. On the other hand, AFLP-based gene tagging will require sexual hybridization between the two mutants. Due to the well documented pineapple self-incompatibility, we do not expect that the mutants set seed after pollination.
In Ananas comosus (L.) Merr., the self-incompatibility is brought about by inhibition of pollen tube growth in the upper third of the style [70,71]. It is gametophytically controlled by a single locus with multiple alleles [72]. The self-rejection reaction is variable in intensity and generally stronger in the cultivated varieties, which is probably a result of the domestication process and selection for seedless fruits [73]. Therefore, other strategies are in progress in our laboratory such as the linkage disequilibrium-based association mapping according to [74] to identify the genes for thorn production and dwarf stature. Studies about the sterility ratios of the new somaclonal variants, as well as, possibilities for somatic cell fusion are also being carried out.
4. Conclusions
We report here the identification of an agriculturally useful mutant (P3R5), with less thorny leaves and there-
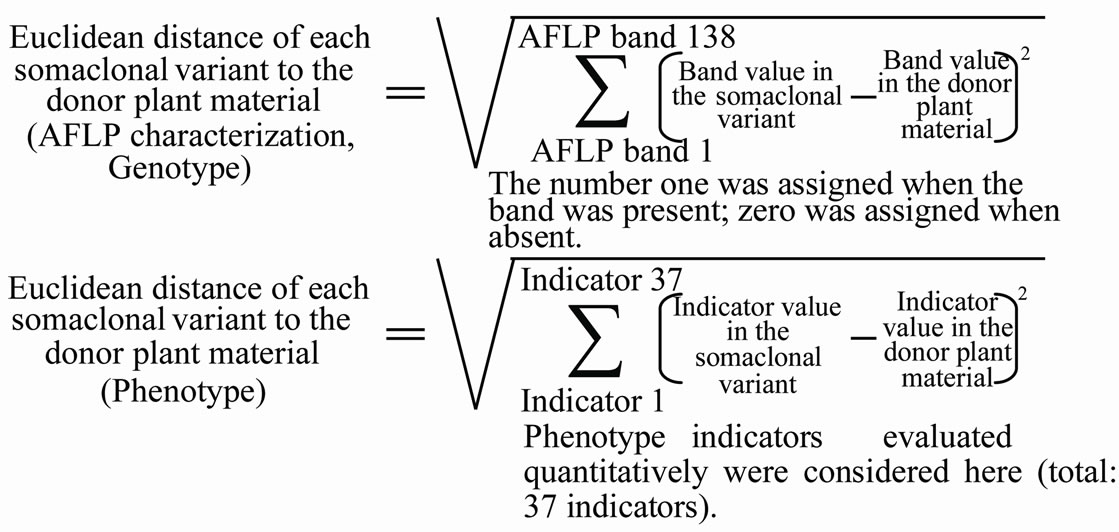 (a)
(a)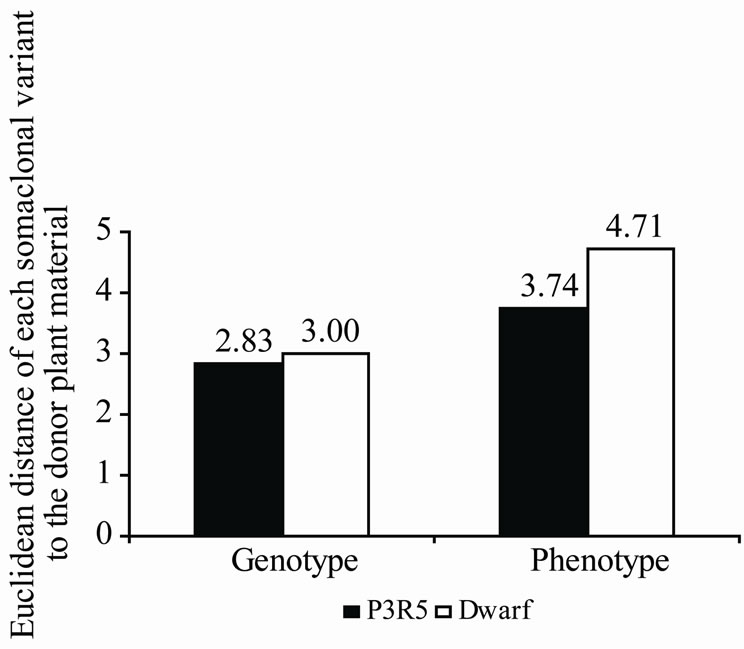 (b)
(b)
Figure 2. Euclidean distances of each somaclonal variant to the donor plant material (cv. Red Spanish Pinar). (a) Formulae; (b) Euclidean distances recorded.
fore, easier to manage in the field. Another mutant with ornamental value (Dwarf) was also obtained. Taking into consideration the phenotype, somaclonal variant Dwarf is more different from the donor plant than P3R5. The AFLP characterization supports these phenotype differences at genome level [43,44]. As far as we know, this is the first report of a comprehensive analysis of pineapple somaclonal variants.
5. Acknowledgements
This research was supported by the Cuban Ministry for Science, Technology and the Environment. We are grateful to Mrs. Julia Martínez, Mrs. Alitza Iglesias and Mrs. Mayda Arzola for their excellent technical assistance. We express gratitude to Ms. Taletha Laudat and Mrs. Glyn Jabbour for the professional language editing.
REFERENCES
- FAOSTAT, “FAO Statistic Division,” 2010. http://faostat.fao.org/site/567/DesktopDefault.aspx?PageID=567
- D. D. Williams and H. Fleich, “Historical Review of Pineapple Breeding in Hawaii,” Acta Horticulturae, Vol. 334, 1993, pp. 67-76.
- M. Fitchet, “Observations on Pineapple Improvement in Taiwan, Republic of China,” Subtropica, Vol. 10, No. 11, 1989, pp. 10-12.
- Y. K. Chan, “Performance and Stability of Pineapple Hybrids in G × E Trial in Malaysia,” Acta Horticulturae, Vol. 425, 1997, pp. 201-211.
- Y. C. Wee, “The Masmerah Pineapple: A New Cultivar for the Malaysian Pineapple Industry,” Wold Crops, Vol. 26, No. 2, 1974, pp. 67-76.
- Y. K. Chan and C. K. Lee, “The Hybrid 1 Pineapple: A New Canning Variety Developed at MARDI,” Teknologi Buah-buahan, Vol. 1, 1985, pp. 24-30.
- Y. K. Chan and H. K. Lee, “Josapine: A New Pineapple Hybrid Developed at MARDI,” Malaysia, 1996, pp. 217- 220.
- O. D. Ramírez, H. Gandía and F. H. Vélez, “P.R.1-67 a New Pineapple Selection,” Fruit Varieties Horticultural Digest, Vol. 26, No. 1, 1972, pp. 13-15.
- C. Loison-Cabot, “Practice of Pineapple Breeding,” Acta Horticulturae, Vol. 196, 1987, pp. 25-36.
- V. M. Villegas, R. B. Pementel, S. V. Siar and E. R. Barile, “Preliminary Evaluation of Promising Pineapple Hybrids,” CIRAD-ISMS, Martinique, 1995.
- J. R. S Cabral, A. P. de Matos and G. Da Cunha, “Selection of Pineapple Cultivars Resistant to Fusariose,” Acta Horticulturae, Vol. 334, 1993, pp. 53-58.
- J. R. S Cabral, A.P. de Matos and G. Coppens d’Eeckenbrugge, “Segregation for Resistant to Fusariose Leaf Colour and Leaf Margin Type from the EMBRAPA Pineapple Hibridization Programme,” Acta Horticulturae, Vol. 425, 1997, pp. 153-200.
- R. Benega, A. Cisneros, M. Hidalgo, J. Martínez, E. Arias, M. Arzola, C. Carvajal and M. Isidrón, “Hybridization in Pineapple: Results and Strategies to Save Time for Obtaining and Releasing New Hybrid Varieties for Growers,” Third International Pineapple Symposium, Thailand, 1998, pp. 15-16.
- G. M. Sanewski, “The Australian Pineapple Fresh Market Breeding Program,” Third International Pineapple Symposium, Thailand, 1998, p.51.
- G. Coppens d’Eeckenbrugge and M. F. Duval, “Pineapple Breeding at CIRAD. II. Evaluation of ‘Scarlet’, a New Hybrid for the Fresh Fruit Market, as Compared to Smooth Cayenne,” Acta Horticulturae, Vol. 529, 2000, pp. 155-163.
- D. P. Bartholomew, G. Coppens d’Eeckenbrugge and C. C. Chen, “Register of New Fruit and Nut Cultivars,” HortScience, Vol. 45, No. 5, 2010, pp. 740-742.
- C. Cabot and J. C. Lacoevilhe, “A Genetic Hybridization Programme for Improving Pineapple Quality,” Acta Horticulturae, Vol. 275, 1990, pp. 395-400.
- S. Sripaoraya, R. Marchant, J. B. Power and M. R. Davey, “Herbicide-Tolerant Transgenic Pineapple (Ananas comosus) Produced by Microprojectile Bombardment,” Annals of Botany, Vol. 88, No. 4, 2001, pp. 597-603. doi:10.1006/anbo.2001.1502
- S. Sripaoraya, S. Keawsompong, P. Insupa, M. R. Davey, J. B. Power and P. Srinives, “Evaluation of Transgene Stability, Gene Expression and Herbicide Tolerance of Genetically Modified Pineapple under Field Conditions,” Acta Horticulturae, Vol. 702, 2006, pp. 37-40.
- P. Espinosa, J. C. Lorenzo, A. Iglesias, L. Yabor, E. Men- éndez, J. Borroto, L. Hernández and A. D. Arencibia, “Production of Pineapple Transgenic Plants Assisted by Temporary Immersion Bioreactors,” Plant Cell Report, Vol. 21, No. 2, 2002, pp. 136-140. doi:10.1007/s00299-002-0481-9
- J. R. Botella and D. J. Fairbairn, “Present and Future Potential of Pineapple Biotechnology,” Acta Horticulturae, Vol. 666, 2005, pp. 23-28.
- J. D. Carlier, G. Coppens d’Eeckenbrugge and J. M. Leitão, “Pineapple. Genome Mapping and Molecular Breeding in Plants Vol. 4. Fruits and Nuts,” In: C. Kole, Ed., Springer-Verlag, Berlin, 2007, pp. 331-342.
- L. Yabor, B. Valle, C. Carvajal, C. Aragón, M. Hernández, J. González, M. Daquinta, A. Arencibia and J. C. Lorenzo, “Characterization of a Field-Grown Transgenic Pineapple Clone Containing the Genes Chitinase, AP24, and Bar,” In Vitro Cell Developmental Biology—Plant, Vol. 46, No. 1, 2009, pp. 1-7.
- V. Cardoza and N. C. Stewart, “Brassica Biotechnology: Progress in Cellular and Molecular Biology,” In Vitro Cell Developmental Biology—Plant, Vol. 40, 2004, pp. 542-551.
- P. J. Larkin and W. R. Scowcroft, “Somaclonal Variationa Novel Source of Variability from Cell Culture for Plant Improvement,” Theoretical Applied Genetic, Vol. 60, No. 4, 1981, pp. 197-214. doi:10.1007/BF02342540
- N. F. Castillo, N. V. Bassil, S. Wada and B. M. Reed, “Genetic Stability of Criopreserved Shoot Tips of Rubus Germplasm,” In Vitro Cell Developmental Biology— Plant, Vol. 46, 2010, pp. 246-256.
- R. Aversano, S. Savarese, J. M. De Nova, L. Frusciante, M. Punzo and D. Carputo, “Genetic Stability at Nuclear and Plastid DNA Level in Regenerated Plants of Solanum Species and Hybrids,” Euphytica, Vol. 165, No. 2, 2009, pp. 353-361. doi:10.1007/s10681-008-9797-z
- M. J. Smulders, W. Rus-Kortekaas and L. J. W. Gilissen, “Natural Variation in Patterns of Polysomaty among Individual Tomato Plants and Their Regenerated Progeny,” Plant Science, Vol. 106, No. 2, 1995, pp. 129-139. doi:10.1016/0168-9452(95)04082-6
- J. L. Fourré, P. Berger, L. Niquet and P. André, “Somatic Embryogenesis and Somaclonal Variation in Norway Spruce: Morphogenetic, Cytogenetic and Molecular Approaches,” Theoretical Applied Genetic, Vol. 94, No. 2, 1997, pp. 159-169. doi:10.1007/s001220050395
- B. Rival, L. Bertrand, T. Beale, M. C. Combes, P. Touslot and P. Leshermes, “Suitability of RAPD Analysis for Detection of Somaclonal Variation in Oil Palm (Elaeis guineensis Jacq.),” Plant Breeding, Vol. 117, No. 1, 1998, pp. 73-76. doi:10.1111/j.1439-0523.1998.tb01451.x
- H. Bouman and G. J. De Klerk “Measurement of the Extent of Somaclonal Variation in Begonia Plants Regenerated under Various Conditions. Comparison of Three Assays,” Theoretical Applied Genetic, Vol. 102, No. 1, 2001, pp. 111-117. doi:10.1007/s001220051625
- M. Martins, D. Sarmento and M. M. Oliveira, “Genetic Stability of Micropropagated Almond Plantlets, as Assessed by RAPD and ISSR Markers,” Plant Cell Report, Vol. 23, No. 7, 2004, pp. 492-496. doi:10.1007/s00299-004-0870-3
- L. Barandalla, E. Ritter and J. I. Ruiz, “Oryzalin Treatment of Potato Diploids Yields Tetraploid and Chimeric Plants from which Euploids Could Be Derived by Callus Induction,” Potato Research, Vol. 10, 2006, pp. 143-154.
- S. K. Sharma, G. J. Bryan, M. O. WinWeld and S. Millam, “Stability of Potato (Solanum tuberosum L.) Plants Regenerated via Somatic Embryos, Axillary Bud Proliferated Shoots, Microtubers and True Potato Seeds: A Comparative Phenotypic, Cytogenetic and Molecular Assessment,” Planta, Vol. 226, No. 6, 2007, pp. 1449-1458. doi:10.1007/s00425-007-0583-2
- E. Piccioni, G. Barcaccia, M. Falcinelli and A. Standardi, “Estimating Somaclonal Variation in Axillary Branching Propagation and Indirect Somatic Embryogenesis by RAPD Fingerprinting,” International Journal Plant Science, Vol. 158, No. 5, 1997, pp. 556-562. doi:10.1086/297467
- W. L. Guo, L. Gong, Z. F. Ding, Y. D. Li, F. X. Li, S. P. Zhao and B. Liu, “Genomic Instability in Phenotypically Normal Regenerants of Medicinal Plant Codonopsis lanceolata Benth. et Hook. as Revealed by ISSR and RAPD Markers,” Plant Cell Report, Vol. 25, No. 9, 2006, pp. 896-906.
- M. A. Palombi, B. Lombardo and E. Caboni, “In Vitro Regeneration of Wild Pear (Pyrus pyraster Burgsd) Clones Tolerant to Fechlorosis and Somaclonal Variation Análisis by RAPD Markers,” Plant Cell Report, Vol. 26, No. 4, 2007, pp. 489-496. doi:10.1007/s00299-006-0256-9
- K. Wakasa, “Use of Tissue Culture for Propagation and Mutant Induction in Ananas comosus,” National Institution Agriculture Science Annual Report, Tokyo, 1977.
- K. Wakasa, “Pineapple (Ananas comosus L. Merr ),” In: Y. P. S. Bajaj, Ed., Biotechnology in Agriculture and Forestry, Springer Verlag, Berlin, 1989, pp. 13-29.
- M. G. Dewald, G. A. Moore, W. B. Sherman and M. H. Evans, “Production of Pineapple Plants in Vitro,” Plant Cell Report, Vol. 7, No. 7, 1988, pp. 535-537. doi:10.1007/BF00272750
- J. L. Lii, E. Rosa-Márquez and E. Lizard, “Smooth Leaf (Spineless) Red Spanish Pineapple (Ananas comosus) Propagated in Vitro,” Journal of Agriculture University Puerto Rico, Vol. 73, 1989, pp. 15-17.
- S. Feuser, M. Kelen, M. Daquinta and R. Onodari, “Genotypic Fidelity of Micropropagated Pineapple (Ananas comosus) Plantlets Assessed by Isozyme and RAPDs,” Plant Cell Tissue Organ Culture, Vol. 72, No. 3, 2003, pp. 221-227. doi:10.1023/A:1022321405683
- G. Pérez, E. Yanez, M. Isidrón, J. C. Lorenzo, “Phenotypic and AFLP Characterization of Two New Pineapple Somaclones Derived from in Vitro Culture,” Plant Cell Tissue Organ Culture, Vol. 96, No. 1, 2009, pp. 113-116. doi:10.1007/s11240-008-9463-0
- G. Pérez, A. Mbogholi, F. Sagarra, C. Aragón, J. González, M. Isidrón and J. C. Lorenzo, “Morphological and Physiological Characterization of Two New Pineapple Somaclones Derived from in Vitro Culture,” In Vitro Cell Developmental Biology—Plant, Vol. 47, No. 3, 2011, pp. 428-433. doi:10.1007/s11627-011-9342-y
- M. Daquinta and R. Benega, “Brief Review of Tissue Culture of Pineapple,” Pineapple News, Vol. 3, 1997, pp. 7-9.
- T. Murashige and F. Skoog, “A Revised Medium for Rapid Growth and Bioassays with Tobacco Tissue Cultures,” Physiologia Plantarum, Vol. 15, No. 3, 1962, pp. 473-497. doi:10.1111/j.1399-3054.1962.tb08052.x
- E. Yanez, K. Gil, L. Rebolledo, D. Uriza, O. Martínez, M. Isidrón and J. Simpson, “AFLP Characterization of the Mexican Pineapple Collection,” Journal American Society Horticultural Sciences, Vol. 18, 2004, pp. 36-41.
- P. Vost, R. Hgogers, R. Bleeker, T. Van de Lee and M. Homes, “AFLP: A New Technique for DNA Fingerprinting,” Nucleic Acids Research, Vol. 23, No. 21, 1995, pp. 4407-4414. doi:10.1093/nar/23.21.4407
- F. J. Rohlf, “NTSYS-Pc: Numerical Taxonomy and Multivariate Analysis System (Version 2.1),” Exeter Software, Setauket, 2000.
- C. Py, J. J. Lacoeuilhe and C. Teisson, “The Pineapple: Cultivation and Uses,” Maisonneuve & Larose, Paris, 1987.
- D. A. Johansen, “Plant Microtechnique,” McGraw-Hill Book, New York. 1949.
- R. J. Porra, “The Chequered History of the Development and Use of Simultaneous Equations for the Accurate Determination of Chlorophylls a and b,” Photosynthesis Research, Vol. 73, No. 1-3, 2002, pp. 149-156. doi:10.1023/A:1020470224740
- R. L. Heath and J. Packer, “Photoperoxidation in Isolated Chloroplast: I. Kinetics and Stoichiometry of Fatty Acid Peroxidation,” Archives of Biochemistry and Biophysics, Vol. 125, No. 1, 1968, pp. 189-198. doi:10.1016/0003-9861(68)90654-1
- S. I. Gurr, J. McPherson and D. J. L. Bowles, “Lignin and Associated Phenolic Acids in Cell Walls,” In: D. L. Wilkinson, Ed., Molecular Plant Pathology, Oxford, 1992, pp. 51-56.
- M. M. Bradford, “A Rapid and Sensitive Method for the Quantification of Microgram Quantities of Protein Utilising the Principle of Protein-Dye Binding,” Analytical Biochemistry, Vol. 72, No. 1-2, 1976, pp. 248-254. doi:10.1016/0003-2697(76)90527-3
- J. Jorrin and R. A. Dixon, “Stress Responses in Alfalfa (Medicago sativa L.). II. Purification, Characterization, and Induction of Phenylalanine Ammonia-Lyase Isoforms from Elicitor-Treated Cell Suspension Cultures,” Plant Physiology, Vol. 92, No. 2, 1990, pp. 447-455. doi:10.1104/pp.92.2.447
- J. McCord and I. Fridovich, “Superoxide Dismutase: An Enzymic Function for Erythrocuprein,” Journal Inorganic and Biochemestry, Vol. 244, 1969, pp. 6049-6055.
- G. Siegel and M. Stitt, “Partial Purification of Two Forms of Spinach Leaf Sucrosephosphate Synthase Which Differ in Their Kinetic Properties,” Plant Science, Vol. 66, No. 2, 1990, pp. 205-210. doi:10.1016/0168-9452(90)90205-3
- L. Van Quy, T. Lamaze and M. L. Champigny, “Effect of Light and NO3 on Wheat Leaf Phosphoenolpyruvate Carboxylase Activity. Evidence for Covalent Modulation of the C3 Enzyme,” Physiologia Plantarum, Vol. 97, No. 4, 1991, pp. 1476-1482. doi:10.1104/pp.97.4.1476
- M. Kantardzic, “Data Mining: Concepts, Models, Methods, and Algorithms”, Wiley, New York, 2003.
- A. Price, J. Cairns, P. Horton, H. G. Jones and H. Griffiths, “Linking Drought-Resistance Mechanisms to Drought Avoidance in Upland Rice Using a QTL Approach: Progress and New Opportunities to Integrate Stomatal and Mesophyll Responses,” Journal Experimental Botany, Vol. 53, No. 371, 2002, pp. 989-1004. doi:10.1093/jexbot/53.371.989
- F. García-Sánchez, J. P. Syvertsen, V. Gimeno, P. Botía and J. G. Perez-Perez, “Responses to Flooding and Drought Stress by Two Citrus Rootstock Seedlings with Different Water-Use Efficiency,” Physiologia Plantarum, Vol. 130, No. 4, 2007, pp. 532-542. doi:10.1111/j.1399-3054.2007.00925.x
- L. Qing-Ming, L. Bin-Bin, W. Yang and Z. Zhi-Rong, “Interactive Effects of Drought Stresses and Elevated CO2 Concentration on Photochemistry Efficiency of Cucumber Seedlings,” Journal of Integrative Plant Biology, Vol. 50, No. 10, 2008, pp. 1307-1317. doi:10.1111/j.1744-7909.2008.00686.x
- J. Chen, R. J. Henny, P. S. Devanand and C. T. Chao “AFLP Analysis of Nephthytis (Syngonium podophyllum Schott) Selected from Somaclonal Variants,” Plant Cell Report, Vol. 24, No. 12, 2006, pp. 743-749. doi:10.1007/s00299-005-0032-2
- M. J. Prado, M. V. González, S. Romo and M. T. Herrera, “Adventitious Plant Regeneration on Leaf Explants from Adult Male Kiwifruit and AFLP Analysis of Genetic Variation,” Plant Cell Tissue Organ Culture, Vol. 88, No. 1, 2007, pp. 1-10. doi:10.1007/s11240-006-9116-0
- S. M. Jain, “Tissue Culture-Derived Variation in Crop Improvement,” Euphytica, Vol. 118, No. 2, 2001, pp. 153-166. doi:10.1023/A:1004124519479
- R. Li, R. Qu, A. H. Bruneau and D. P. Livingston, “Selection for Freezing Tolerance in St. Augustinegrass through Somaclonal Variation and Germplasm Evaluation,” Plant Breeding, Vol. 129, 2010, pp. 417-421.
- P. T. H. Brown, F. D. Lange, E. Kranz and H. Lorz, “Analysis of Single Protoplasts and Regenerated Plants by PCR and RAPD Technology,” Molecular and General Genetics, Vol. 237, No. 3, 1993, pp. 311-317.
- P. Bhatia, N. Ashwath, T. Senaratna and S. Krauss, “Genetic Analysis of Cotyledon-Derived Regenerants of Tomato Using AFLP Markers,” Current Science, Vol. 88, 2005, pp. 280-284.
- K. R. Kerns, “Concerning the Growth of Pollen Tubes in Pistils of Cayenne Flowers,” Pineapple Quarterly, Vol. 1, 1932, pp. 133-137.
- S. K. Majumder, K. R. Kerns, J. L. Brewbaker and G. A. Johansmessen, “Assessing Self-Incompatibility by a Pollen Fluorescence Technique,” Proceedings of the American Society for Horticultural Science, Vol. 84, 1964, pp. 217-223.
- J. L. Brewbaker and D. D. Gorrez, “Genetics of SelfIncompatibility in the Monocot Genera, Ananas (Pineapple) and Gasteria,” American Journal of Botany, Vol. 54, No. 5, 1967, pp. 611-616. doi:10.2307/2440666
- G. Coppens d’Eekenbrugge, M. F. Duval and F. Van Miegroet, “Fertility and Self-Incompatibility in the Genus Ananas, ” Acta Horticulturae, Vol. 334, 1993, pp. 45-51.
- S. A Flint-Garcia, J. M. Thornsberry and E. S. Buckler, “Structure of Linkage Disequilibrium in Plants,” Annual Review Plant Biology, Vol. 54, 2003, pp. 357-374.

