Advances in Microbiology
Vol.3 No.1(2013), Article ID:29101,13 pages DOI:10.4236/aim.2013.31001
Diversity and Bioactivity of Cultivable Animal Fecal Actinobacteria
1Yunnan Institute of Microbiology, Yunnan University, Kunming, China
2Lab of Metabolic Disease Research and Drug Development, China Medical University, Shenyang, China
3Sanqi Research Institute of Wenshan, Wenshan, China
4Yunnan Wild Animal Park, Kunming, China
Email: *jiangyikm@hotmail.com, lihan@mail.cmu.edu.cn, chenxiu2007@126.com, ym3612003@163.com, Danzheng@mail.cmu.edu.cn, ws-wangyong37@163.com, 1811124955@qq.com, xueshi_huang@yahoo.com.cn
Received November 2, 2012; revised December 4, 2012; accepted January 5, 2013
Keywords: Actinobacteria; Diversity; Bioactivity; Animal Feces
ABSTRACT
Microbial symbionts play important roles in food digestion and absorption, immunity, pathogens resistance, and health maintaining of their hosts by co-evolution. To provide new sources for discovering new leader compounds of drugs, the diversity and bioactivities of cultivable actinobacteria from animal feces have been studied. 31 species of animal fecal samples were collected from Yunnan Wild Animal Park. The purified cultures of actinobacteria were isolated from these samples by using 5 media. The 16S rRNA gene sequences of 528 selected strains were determined, the phylogenetic analysis was carried out, and anti-microbial and anti-tumor activities were determined. 35 genera (including a new genus, Enteractinococcus) of actinobacteria from the 31 species of animal feces were identified. Some strains had high anti-tumor and antimicrobial activities. More than 50 secondary metabolites were isolated and identified, a novel bioactive macrolactam polyketide glycoside, Sannastatin, was found. Nine fecal streptomycete strains were fermented respectively, blended to the microbial manure, and used to prophylaxis and treatment of soil-borne disease of notoginseng in field. The incidence rate of the disease was lower 81% than agricultural chemicals. Fecal actinobacteria, a possibility as a new source for discovering drug leader, agricultural chemicals and other industry products, will be discussed.
1. Introduction
Animal intestinal and fecal microorganisms (Fecal Microbiota) have been studied for decades [1]. A large number of microbial kinds exist in animal gastrointestinal tract and feces. The intestinal microbial community which composed of 1013 to 1014 microorganism outnumbers the somatic and herm cells by at least an order of magnitude [2]. However, the most part of these microorganisms are uncultivable yet [2-8]. In the cause of coevolution of microorganisms and their hosts, the former play an important role in digestion and absorption of food, immunity, resistance to pathogens, and maintaining health of host. But the relationship between microorganisms and their hosts remains largely unknown due to the complexity of the internal ecological system [9,10]. Some of intestinal microbes are pathogen, and some are beneficial. How to explore and utilize the enormous beneficial microbial resource is a very tempting challenge. Probiotics are dietary supplements containing friendly bacteria, and applying wildly for recovering balance of microbial system, improving intestinal and overall health and helping guard to against disease [11,12].
Market demand for new drugs in whole world is extremely urgent and extensive due to fast extension of stubborn disease (cancer, AIDS, HIV) and common ailment (hypertension, diabetes and Hyperlipidemia), germination of new disease can not been seeing the cause, and the fast spreading of antibiotic resistant pathogens [13,14].
Actinomycetes (Actinobacteria) have been paid a great attention owing to their production of various natural drugs and other bioactive metabolites including antibiotics, enzyme inhibitors and enzymes. Over 22,000 bioactive secondary metabolites (including antibiotics) were published in the scientific and patent literature, and about a half of them were produced by actinomycetes. About 150 antibiotics have being applied in human therapy and agriculture now; 100 - 120 of them were produced by actinomycetes [15]. Actinomycete is still an important source for new natural drugs development. So Baltz showed a proposition of “Renaissance in antibacterial discovery from actinomycetes” [16]. However, the development of new drugs from actinomycetes in common habitats is more and more difficult [17]. In order to overcome these challenges, some new concept based on genome was described, that is “new habitats, new methods, new species, new gene cluster, new products and new use” [17-19]. In other words, novel microbial kind should contain new gene cluster synthesizing new secondary metabolites, so far as getting new kind is an important premise for obtaining new compounds [17]. Many companies and laboratories focused on new actinomycete resources from new habitats, such as oceans, extreme environment and plants, for development of new drugs. In our view, making the uncultured to pure cultured microorganisms is one new hope for getting new drug leads.
Actinomycete, as a pathogen of human and animal, had been studied widely before [20]. But up to now, the research work on actinomycetes as a source for discovery of novel drug leads is very few in the world. In order to get much more unknown actinomycetes for discovering new bioactive metabolites, 31 species of animal which contain primate, mammality, birds, amphibian and insect; perissodactyla, artiodactyla and ruminant; carnivore, herbivorous and omnivorous, were selected. The actionmycetes in the feces samples of these animals were isolated, cultivated and identified. Anti-microbial activities, enzyme activity, and synthesis genes of five antibiotics of some selected strains were determined. Some results are reported here.
2. Materials and Methods
2.1. Collection and Preparation of Samples
Fresh fecal samples were collected from selected 31 species of animals which live in the Yunnan Wild Animal Park, Kunming, China. The samples were put in sterile dish immediately, and dried for 10 days at 28˚C. 2 g of each dried sample were pre-treated at 80˚C for 1 hour, put in 18 ml sterile water with 0.1% Na4P2O5, and shaken for 60 min at 220 rpm/min. The suspension was diluted from 10−1 to 10−8.
2.2. Isolation Medium of Actinobacteria
Five media [21] were used for isolating actinobacteria in fecal samples.
All of media were supplemented with filter 4 groups of sterilized mixture or single solutions: 1) 50 mg cycloheximide, 50 mg nystatin, 20 mg nalidixic acid, 3 mg penicillin; 2) 100 mg cycloheximide, 100 mg nystatin, 40 mg nalidixic acid, 5 mg penicillin; 3) 50 mg K2Cr2O7, 5 mg penicillin; 4) 75 mg K2Cr2O7, 5 mg penicillin for 1000 ml medium, as inhibitors against fungi and Gram negative bacteria.
Plate dilution method was used for isolating actionbacteria from the sample suspension, then the plates were cultivated for 7 to 35 days at 28˚C, then take count of colonies, and pick up actinobacteria to slant of the same isolation medium.
2.3. Identification of Pure Cultivated Actinobacteria
Total 2049 pure strains were isolated from the 31 animal feces samples, and were cultured on ISP media 2 and 3, at 28˚C for 7 - 14 days, then observed with light microscope one by one inspection. 528 strains of them were selected after throwing out the duplicates strains based on the same morphological and cultural characteristics [22]. The DNA of pure strains was extracted for 16S rRNA analysis (Orsini and V. Romano-Spica, 2001). 16S rRNA was amplifed by PCR using TaKaRa Ex Taq (TaKaRa Biotechnology) and the forward primer F8 (8 ± 27), 5’-GAG AGT TTG ATC CTG GCT CAG-3’ and the reverse primer (1510 ± 1492), 5’-GGT TAC CTT GTT ACG ACT T-3’ were used. The conditions used for thermal cycling were as follows: denaturation at 95˚C for 5 min, followed by 35 cycles of denaturation at 95˚C for 1 min, annealing at 56˚C for 1 min and extension at 72˚C for 3 min. At the end of the cycles, the reaction mixture was kept at 72˚C for 5 min and then cooled to 4˚C. The 1 ± 5 kb amplified 16S rDNA fragment was separated by agarose gel electrophoresis and purified by using a Watson gel extraction kit. The purified fragment was sequenced directly by using the Big Dye terminator cycle sequencing ready reaction kit (Perkin-Elmer) and was analyzed with an ABI PRISM 377 DNA sequencer [23]. The resultant sequences were manually aligned with available sequences from public databases. Phylogenetic trees were inferred by using the neighbour-joining [24] and maximum-likelihood methods [25]. All of pure cultivated strains were identified at a genus level.
2.4. Determination of Anti-Microbial Activity
Strains were fermented using the broth (YIM 61: soybean meal 20 g, glucose 10 g, peptone 4 g, K2HPO4 1 g, MgSO4·7H2O 0.5 g, NaCl 1 g, CaCO3 2 g, water 1000 ml, pH 7.8), shaking for 7 days at 28˚C. Agar diffusion method was used for determining the anti-microbial activities against Bacillus subtills (DSM 3258T), Staphylococcus aureus (DSM 30501T), Mycobacterium tuberculosis avium (un-pathogen from Dr. Lixin Zhang), Candida glabrata (DSM 24506T) and Aspergillus niger (IAM 190).
Activities of 19 enzymes were determined by using API ZYM kit (biomèrieux). Hydrolyzation of cellulose and chicken hair was determined with Shirling and Gottlieb’s methods.
2.5. Determination of Biosynthesis Enzyme Genes of Five Antibiotics
Biosynthesis genes of type I and II polyketide synthases (PKS), nonribosomal peptide synthase (NRPS) and polyene cytochrome P450 hydroxylase (CYP) were determined by PCR [26,27]. 3,5-amino-hydroxyl-Benzoic acid biosynthesis gene (AHBA) was determined using the method described by Zhang et al. [28]. Five sets of PCR primers were used: A3F (5’-GCSTACSYSATSTACACSTCSGG-3’) and A7R (5’-SASGTCVCCSGTSCGGTAS-3’) targeting NRPS sequences; K1F (5’-TSAAGTCSAACATCGGBCA-3’) and M6R (5’CGCAGGTTSC SGTACC-AG TA-3’) targeting PKS-I sequences; KSa and KSb (5’-TSGRCTACRTCAACGGSCACGG-3’) and (5’-TACSAGTCSWTCGCCTGGTTC-3’) targeting PKS-П sequences; PEH-1, [5’-TGGATCGGCGACGACC-G(G/C)(A/G/C)(T/C)CGT-3’] and PEH-2, [5’-CCG(T/A)A(G/C)AG(G/C)A(T/C)(G/C)CCGTCGTACTT-3’] targeting CYP genes; 755a (5’-AGAGGATCCTTCGAGCRSGAGTTCGC-3’) and 755b (5’-GCAGGATCCGGAMCATSGCCATGTAG-3’) targeting AHBA genes.
DNA preparations were used as template DNA for Taq Polymerase. Reactions were performed in a final volume of 50 μl containing 0.4 μM of each primer, 0.2 mM of each of the four dNTPs (Roche), 5 μl of extracted DNA, 1 U Taq polymerase (Appligene) with its recommended reaction buffer and 10% of DMSO. NRPS, PKS-I and PKS-П amplifications were performed in a Peltier Thermal Cycler PTC-200, according to the following profile: 5 min at 95˚C and 35 Cycles of 96˚C 1 min, 60˚C 30 s, 72˚C 45 s, and 5 min at 72˚C, followed by 10 min at 72˚C. The amplification products were analyzed by electrophoresis in 1% (w/v) agarose gels stained with ethidium bromide.
3. Results
3.1. Selective Isolation Effect of Five Media for Actinobacteria
Ten species of animal feces samples were used to test the selective isolation effect of five media for actinobacteria. As a result, medium YIM 212, YIM 171 and HV showed better isolation effect, which obtained 176, 169 and 164 strains of actinobacteria respectively. Totally, 746 pure cultivated strains of actinobacteria were isolated. The strain richness of each samples are quite different, 156 and 140 strains were isolated from Vicugna pacos and Rhinoceros sondaicus samples respectively, but only 20 and 36 strains were isolated respectively from Testudo elephantopus and Viverra zibetha (Table 1).
The combination and concentration of different kinds of inhibitors were tested several times for isolating actinobacteria from feces. As a result, media containing 50 mg/L K2Cr2O7 and 5 mg/L penicillin sodium or 50 mg/L nystatin, 20 mg/L nalidixic acid and 5 mg/L penicillin sodium showed better inhibiting effect, on which most Gram negative bacteria were inhibited and no fungi grown. Different dilutions of sample suspension were tested with YIM 171 medium many times. The optimum dilutions of fecal suspension for isolating actinobacteria were 10−5, 10−6, and 10−7, in which about 17 to 233 colonies grown on the plates, and it was very easy to pick up single colony (Table 2). However, the optimum concentration to each animal fecal sample should be tested before and all alone.
Table 1. Effect of selective isolation for actinobacteria from fecal samples of 10 species of animals with five media (Amount of strains obtained).
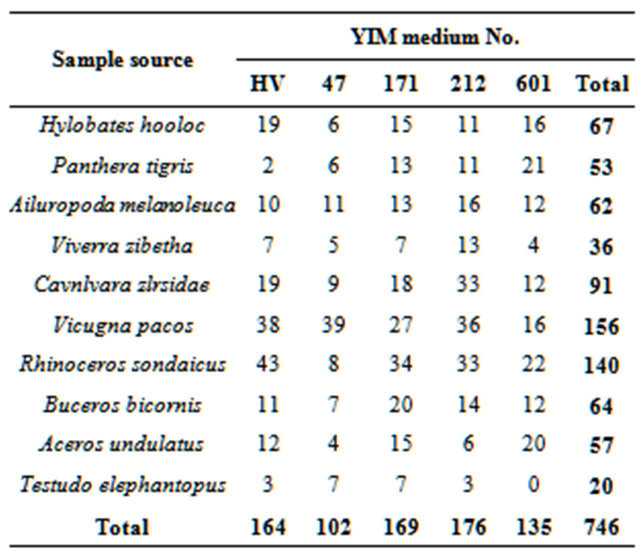
Table 2. Isolation effect for actinobacteria from fecal samples of 10 species of animals with YIM 171 at different dilution (cfu/g) dried sample on medium plate.
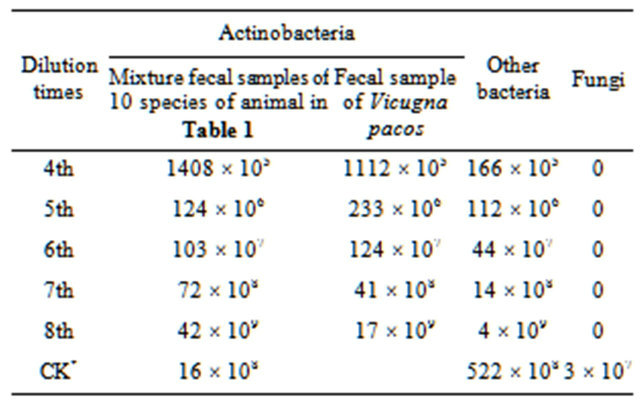
CK = without inhibitors at dilution 7th; *Can not pick up the single colony of actinomycetes.
3.2. Diversity of Actinobacteria
16S rDNA sequences of 528 pure cultivated strains from fecal samples of 31 species of animals were determined. The phylogenetic analysis was carried out and the strains were identified at a genus level. Totally, 35 genera of actinobacteria were identified. They were Agrococcus, Arthrobacter, Cellulosimicrobium, Cellulomonas, Citricoccus, Corynebacterium, Curtobacterium, Dietzia, Enteractinococcus, Gordonia, Isoptericola, Janibacter, Jiangella, Kocuria, Labedella, Leucobacter, Microbacterium, Micrococcus, Micromonospora, Mycobacterium, Nocardia, Nocardiopsis, Oerskovia, Patulibacter, Promicromonospora, Pseudonocardia, Rhodococcus, Saccharomonospora, Salinibacterium, Sanguibacter, Streptomyces, Tsukamurella, Verrucosispora, Williamsia and Yaniella. The 35 genera of actinobacteria belong to 18 Families of two orders, Order Actinomycetales and Order Solirubrobacterales. Order Actinomycetales included 7 Suborders, Micrococcineae (17 genera), Corynebacterineae (9 genera), Pseudonocardineae (2 genera), Streptomycineae (1 genus), Streptosporangineae (1 genus), Micromonosporineae (2 genera), and Jiangellineae (1 genus). No Suborder Actinomycineae, Propionibacterineae, Frankineae and Glycomycineae were isolated. Table 3 summarizes the composition of actinobacteria and other bacteria in each animal feces.
Table 3. Composition of actinobacteria from 31 species of animal feces.
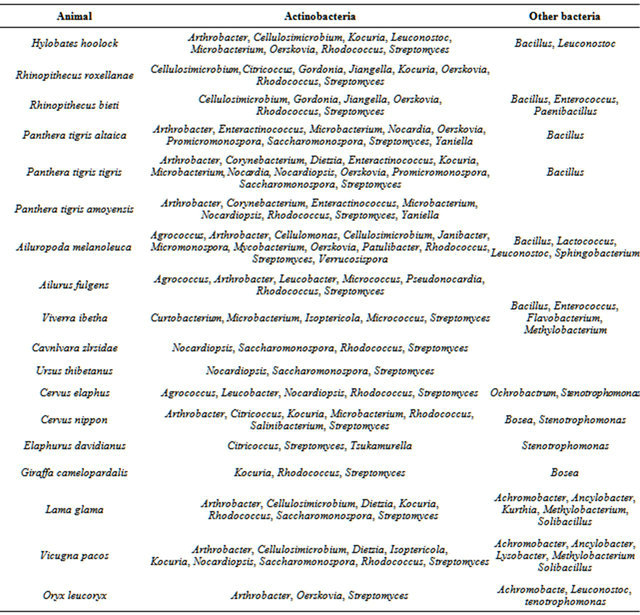
Continued

Members of Genus Streptomyces, the first preponderant microbes, were isolated from all samples, and cfu (colony-forming units)/g dried sample were 2 × 105 to 176 × 107 in different fecal samples. Thirty-nine species of the genus were identified, and 21 un-identified. Streptomyces albus, S. albidoflavus, S. griseus, S. hygroscopicus, S. rutgersensis, S. tendae, and S. violaceoruber etc. occurred at a high frequency.
Members of Rhodococcus, the second preponderant microbes, were isolated from 22 species of animal fecal samples, and they are the most at amount. Rhodococcus coprophilus which is an emerging tool in the microbial source tracking “tool-box”, Rh. corynebacterioides, Rh. corynebacterioides, Rh. equi, Rh. pyridinivorans and Rh. zopfii were occurred at a high frequency.
Members of Arthrobacter and Microbacterium were identified from 11 and 10 species of animal feces respectively. Kocuria was isolated from 5 species of animal feces. Micromonospora was isolated from feces of Ailuropoda melanoleuca, Elephas maximus and Anser anser domesticus; Nocardia only from Panthera tigris altaica.
Twelve genera of actinomycetes were isolated and identified in feces samples collected from Panthera tigris tigris. They belonged to ten Families, Cellulomonadaceae, Corynebacteriacea, Dietziaceae, Microbacteriaceae, Micrococcaceae, Nocardiacea, Nocardiopsaceae, Promicromonosporaceae, Pseudonocardineae and Streptomycetaceae. Streptomycetaceae occupied 64%, was preponderant, and secondly Micrococcaceae, 7% (Figures 1 and 2). Twelve genera of Actinobacteria were identified in feces samples of Ailuropoda melanoleuca, the genus Patulibacter belongs to Order Solirubrobacterales [29], and the genus Verrucosispora was found in peat bog near Gifhorn, Germany before [30] (Table 3).
Only two genera of actinomycetes were isolated from Sttruthio camelus, Tragelaphus buxtoni, Python reticulates, and Xylocopa dissimilis. No members of Actinoplanes, Actinomadura, Streptosporangium and Thermamonospora which are distributed widely in soil and lake were isolated from the 31 feces.
Some members of rare actinobacteria, such as the genus Yaniella [31] were identified respectively from feces of two species of tigers. A strain (YIM 100708) of Jiangella [32], a genus wildly distributing in saline and alkaline soil, desert, wall material of an indoor environment, cave and plant stem, were isolated from feces of Rhinopithecus bieti. Members of the genus Enteractinococcus, a novel genus, belong to Micrococcaceae, were isolated and characterized from three species of tigers [33]. It is worth to show that the 16S rDNA sequence similarities of 32 of 528 sequenced strains with valid published species were below 98.5%. In other words, nearly 6% pure cultivated strains were un-known, and they were possible novel species [19]. Figures 1 and 2 as two examples, showed the composition of actinomycetes in fecal samples of Panthera tigris tigris.
Twenty-five genera of other bacteria were isolated from these animal feces. They are Achromobacter, Ancylobacter, Bacillus, Bosea, Brevundimonas, Brochothrix, Comamonas, Enterococcus, Flavobacterium, Kluyvera, Klebsiella, Kurthia, Lactcus, Lysobacter, Methylobacterium, Ochrobactrum, Paenibacillus, Planococcus, Planomicrobium, Pseudomonas, Rhizobium, Solibacillus, Sphaerobacter, Sphingobacterium, Stenotrophomonas (Table 3).
3.3. Anti-Microbial Activities
Antimicrobial activities of 384 strains isolated from 22 species of animal feces were determined with agar diffusion method (Table 4). 84 strains (22%) showed inhibition activities against Bacillus subtills; 52 strains (13%)
against Staphylococcus aureus; 23 strains (6%) against Mycobacterium tuberculosis; 76 strains (20%) against Candida albicans; 61 strains (16%) against Aspergillus niger. 13% to 55% of 47 strains, mainly Streptomyces, from Python reticulates had higher inhibition activities to all of the five tested microbes. Strains of anti-Mycobacterium tuberculosis were fewer, only 6%. Each 3 strains from Helarctos malayanus, Aceros undulatus and Pavo
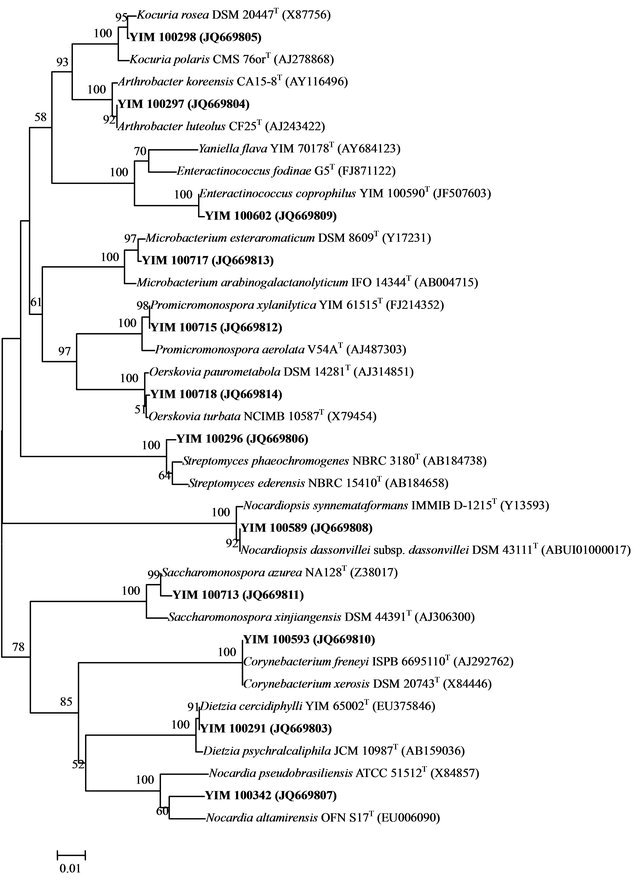
Figure 1. Neighbour-joining tree showing the phylogenetic relationships based on 16S rRNA gene sequences of culturable Actinomycetes from fecal samples of Panthera tigris tigris. Sequences obtained in this work are in bold. Bootstrap values (expressed as percentages of 1000 replications) greater than 50% are given at the nodes. Bar, 1 nt substitution per 100 nt.
Table 4. Anti-microbial activities of fecal actinobacteria.
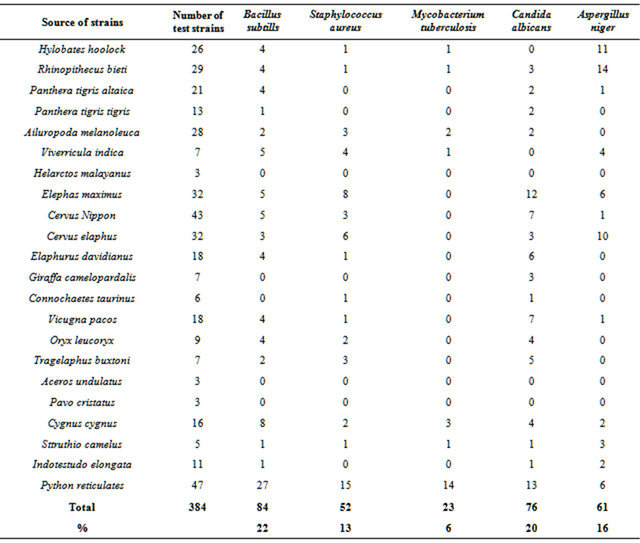
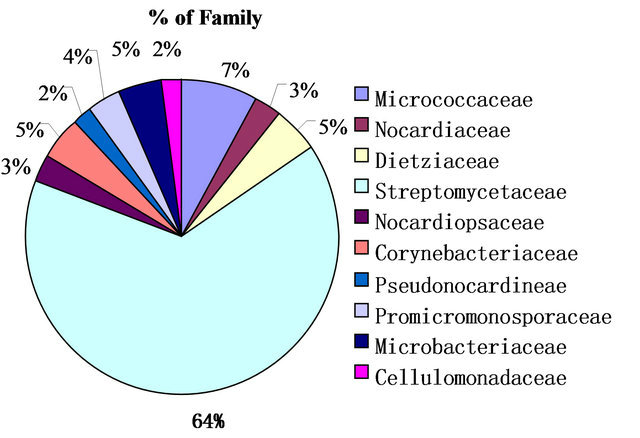
Figure 2. Composition of actinobacteria in Panthera tigris tigris feces.
cristatus feces have anti-microbial activities. These results showed that actinobacteria from animal feces have wide anti-microbial activities.
3.4. Antitumor Activities
Antitumor activities of 238 fecal actinobacterial strains were determined by using HL60, HepG-2, Skov-3, A431, and K562 cell lines in vitro. A part of results were showed in Table 5. 33% and 30% of tested strains showed K562 and HL60 cell line inhibition activity respectively, and more than 50% strains from Rhinopithecus bieti could inhibit k562 and HL 60. The IC50 of crude extracts from some strains were below 4 μg/ml. Large portion of the tested strains showed anti-tumor activities highlighted the distinct feature of actinomycetes from animal feces.
3.5. Activities of Enzymes
Enzyme activities of 233 strains were determined by using API ZYM Kit, biomèrieux (Figure 3). More than 90% of tested strains showed five enzymes activitiesincluding alkaline phosphatase, acid phosphatase, leucine arylamidase, naphthol-AS-BI-phosphohydrolase and β- glucosidase. α-glucosidase. 10% to 90% of strains possess of 12 other enzymes activities. Strains which can produce α-mannosidase and α-fucosidase were less than 10%. No strains showed β-glucuronidase activity (Figure 3). All of strains from 13 species of animal feces were able to hydrolyze cellulose and chicken hair at the same time, except those from Panthera tigris altaica, a carnivorous animal, couldn’t hydrolyze cellulose. More than
Table 5. Antitumor activities of fecal actinomycete strains.
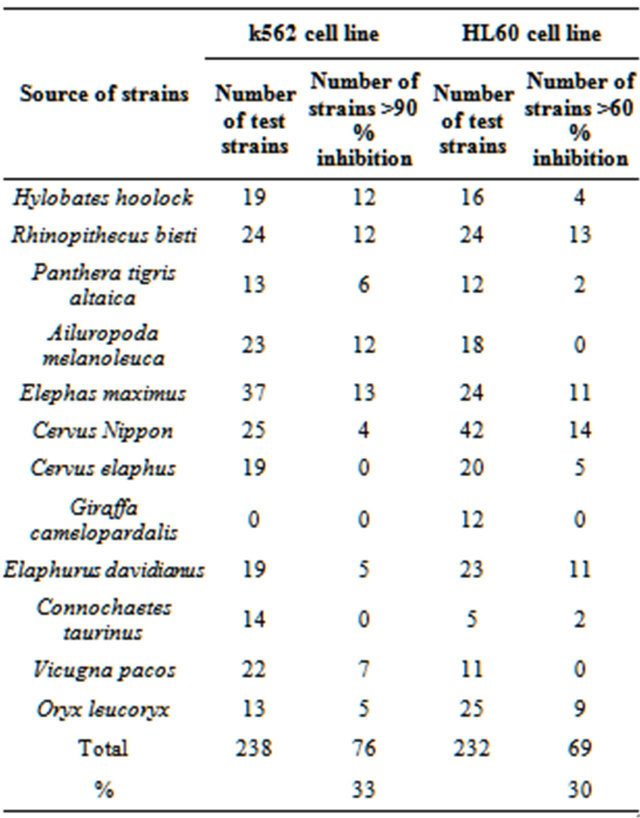


Figure 3. Enzyme activities of actinomycetes from animal feces; A = Alkaline phosphatase, B = Esterase (C4), C = Esterase lipase (C8), D = Lipase (C14), E = Leucine arylamidase, F = Valine arylamidase, G = Cystine arylamidase, H = Trypsinase, I = α-chymotrypsinase, J = acid phosphatase, K = naphthol-AS-BI-phosphohydrolase, L = α-galactosidase, M = β-galactosidase, N = β-glucuronidase, O = α-glucosidase, P = β-glucosidase, Q = N-acetyl-β-glucosaminidase, R = α-mannosidase, S = α-fucosidase.
90% of strains from Cervus elaphus, Giraffa camelopardalis, Vicugna pacos, Elaphurus davidianus and Oryx leucoryx were able to hydrolyze chicken hair. About 80% of strains from Giraffa camelopardalis, Equus burchelli and Oryx leucoryx were able to hydrolyze cellulose. Chicken hair hydrolyzation activity of the strains from Panthera tigris altaica and Ailuropoda melanoleuca, and cellulose hydrolyzation of the strains from Hypoblasts hoolock, Rhinopithecus bieti, Ailuropoda melanoleuca amd Elephas maximus were weaker (Figure 4).
3.6. Biosynthetic Enzyme Genes of Five Antibiotics
Biosynthetic enzyme genes of five metabolites of 201 strains from 15 species of animal feces were analyzed by using specific primers. Amount 101 strains, 19, 15, 34 and 22 strains contained PKS I, PKS II, NRPS and CYP genes respectively. Large portion of the strains from Rhinopithecus bieti, Panthera tigris altaica, Cavnlvara zlrsidae, Vicugna pacos, Rhinoceros sondaicus and Pavo cristatus exist these four genes. None of these four genes had been detected in strains from Cervus Nippon and Equus burchelli. A biosynthetic enzyme gene of 3-amino- 5-hydroxybenzoic acid (AHBA) for ansamycins biosynthesis was determined, but no positive strains were detected. These results showed that fecal actinobacteria have biosynthetic enzyme gene of four metabolites, but not AHBA (Figure 5).
4. Discussion
4.1. Comparison of Actinomycete Diversity in Soil, Sea and Feces
In our previous studies of pure cultivate actinobacteria, 17 genera were isolated from soil samples which collected from primeval forest in Grand Shangri-La, southwest China [34], 13 genera from subtropical every green forest in Gulin, Sichuan [35], 26 genera from soil samples
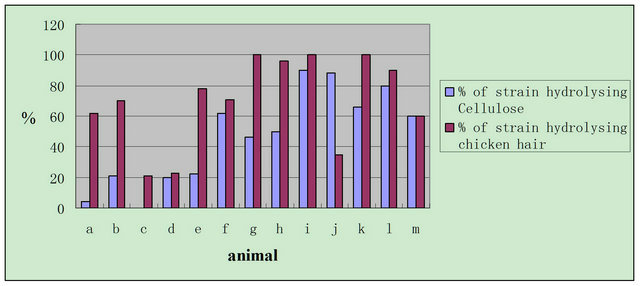
Figure 4. Hydrolyzation of actinomycetes to cellulose and chicken hair; a = Hypoblasts hoolock, b = Rhinopithecus bieti, c = Panthera tigris altaica, d = Ailuropoda melanoleuca, e = Elephas maximus, f = Cervus Nippon, g = Cervus elaphus, h = Elaphurus davidianus, I = Giraffa camelopardalis, j = Equus burchelli, k = Vicugna pacos, l = Oryx leucoryx, m = Tragelaphus buxtoni.
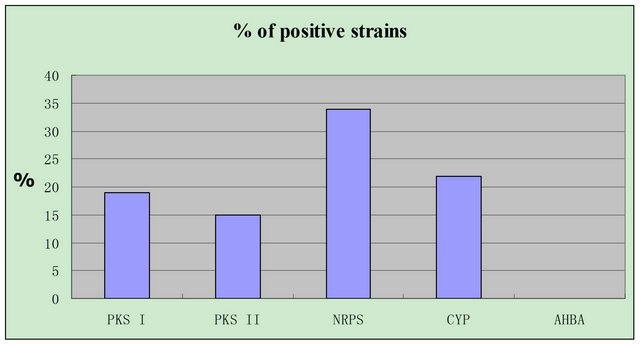
Figure 5. Biosynthetic enzyme genes of five compounds produced fecal actinomycetes.
from tropical rain forests in Xishuangbanna, southwest China [36], 16 genera from hypersaline soil in Qinghai, west China [37] (Table 6). Most part of these actinobacteria belong to cell Chemotype I, П, III and IV [38,39]. Their aerobic mycelium is abundant. 15 genera of actinomycetes in Baltic Sea were identified, and members of Micromonospraceae are the most [36]. Up to now, about 48 genera of actinobacteria were isolated and identified from some marine habitats in the world [40].
In this study, 35 genera of actinobacteria belong to two orders (covering 7 suborders) and 23 of other bacteria were isolated and identified from only 31 species of animal feces. Members of Streptomyces was the first preponderant microbe, cfu/g of dried samples were up to 109, and distributed in all of the 31 samples. Members of Rhodococcus, a kind of feces-phylic bacteria, were identified from 22 species of animal feces, and were the second widest distributions. Seventeen genera of Suborder Micrococcineae were isolated, which were the richest Suborder in diversity. These results indicated that, first, members of both genera Streptomyces and Rhodococcus were the widest distribution and the largest amount; second, composition of actinobacteria with Chemotype IV to IX [38,39] and globose and bacilliform shapes, specially suborder Microboccaceae, were the richest diversity, and occurred at a high frequency in most part of tested animal feces. These are distinct features of fecal actinobacterium community differing from those in soil and marine environment.
4.2. Bioactivities of Fecal Actinobacteria
In our earlier studies of anti-microbial activities of actinomycetes, five indicator strains which mentioned above had been used. As a result, 12% to 15% of Streptomycetes and 5% to 12% of rare actinomycetes showed anti-microbial activities in the study of soil actinomycetes from three areas in Yunnan and Sichuan, Southwest China [35], 4.3% to 21% actinomycetes from Baltic Sea showed anti-microbial activities. In this study, fecal actinobacteria showed wide-spectrum of anti-microbial activities. 486 strains have been detected, 15% to 27% of strains could inhibit Bacillus subtilis, 8% to 30% inhibited Staphylococcus arreus, 2% to 9% inhibited Mycobacterium tuberculosis avium, 11% to 20% inhibited Candida albicans, and 2% to 18% inhibited Aspergillus niger. Some strains could generated large zones of inhibition up to 60 mm. Fecal actinobacteria also showed wide-spectrum of anti-tumor abilities (Table 5). At least 30% of fecal actinomycete strains could inhibit two kinds of tumor cell lines simultaneously, and more than 50% strains from Rhinopithecus bieti could inhibit K562 and HL 60, the ferments crude extracts from some of them had high tumor cells inhibition activities with IC50 below 4 μg/ml. Fecal actinomycetes also containing various enzymes with high activities that could degrade difficult degradation substances, such as cellulose and chicken
Table 6. Diversity of soil actinomycetes and Baltic Sea.

hair. These active symbiotic fecal actinobacteria should provide enormous benefits to host, such as improving food digestion and absorption, maintaining the balance of microbial ecological system in intestinal tract, providing various resistances to pathogens and tumor, improving health of hosts and so on.
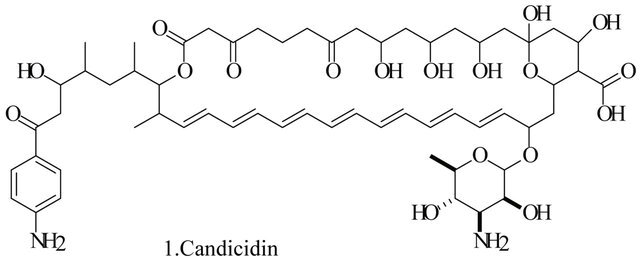
Up to the present, more than 60 bioactive secondary metabolites have been isolated and characterized from some fecal actinomycete strains, including abkhazomycin, AI 77B, akashin A, alazopeptin, apigenin, candicidin, cosmomycin, desertomycin, desferrioxamine E, discodermolide, emodin, enopetin A/B, erythromycins, favo-fungin, geldanamycin, kasugamycin, kidamycins, leucomycin, longestin, panosialin-wA, puromycin, rutamycin, rhodomycinone, stigmast-5-en-3-O-b-D-glucopranoside, tirandamycin, vicenistatin, polyene macrolides etc. A part of these compounds was showed in Figure 6. These compounds have complex structure and various activities. Several novel coumpounds, such as sannastatin, a novel toxic macrolactam polyketide glycoside, which produced by an un-identified Streptomyces sp. YIM 100282, had been found [41].
Nine streptomycete strains which showed the strongest inhibition activity against pathogens of notoginseng (Panax pseudo-ginseng var. Notoginseng) were selected from 2049 animal feces actinomycete strains. The nine strains were fermented respectively, blended to the microbial manure, and used to prophylaxis and treatment of soil-borne disease of notoginseng at field Wenshan, Yunnan for three years. The rate of incidence of the disease at the dosage of 30 g/m2 microbial manure was lower 81% than agricultural chemicals (Figure 7). The microbial manure can be widely used in large tracts of land.
 (a)
(a)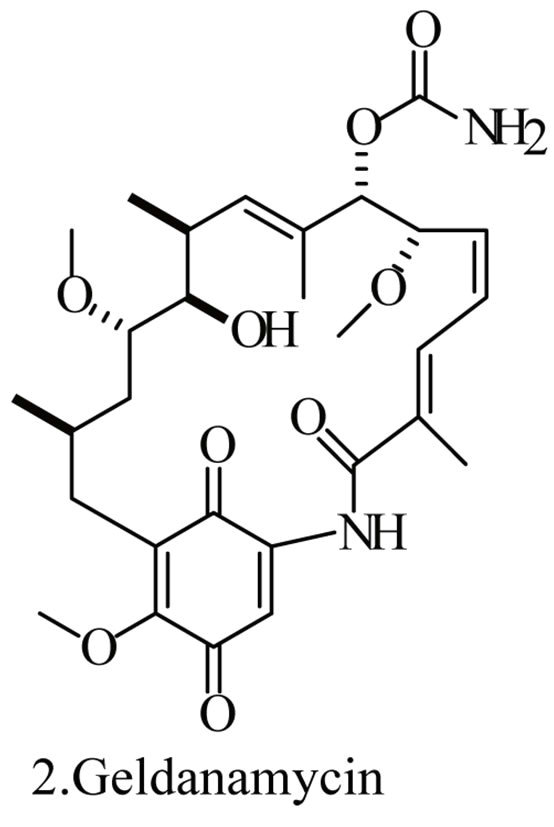 (b)
(b)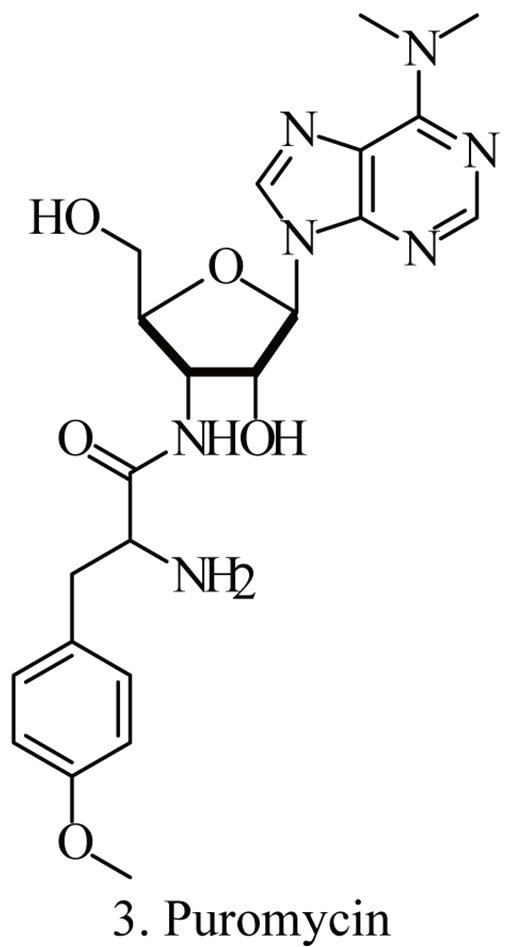 (c)
(c)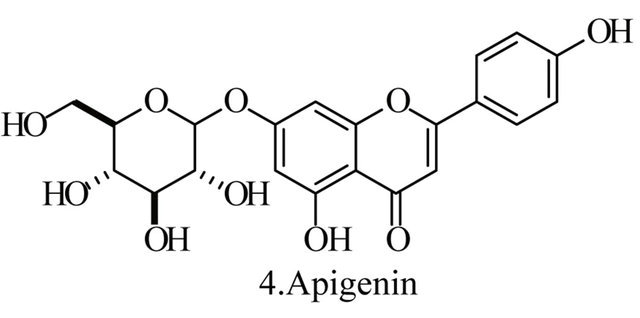 (d)
(d)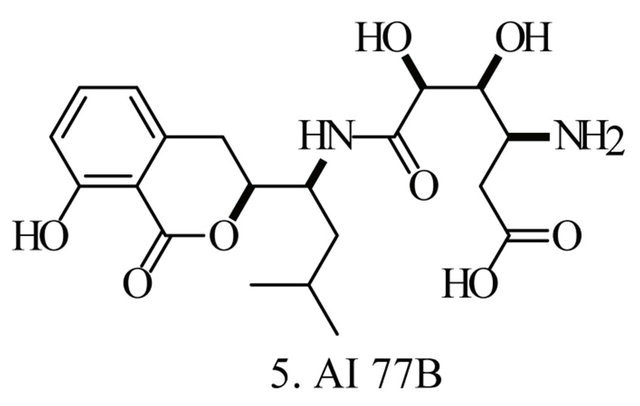 (e)
(e)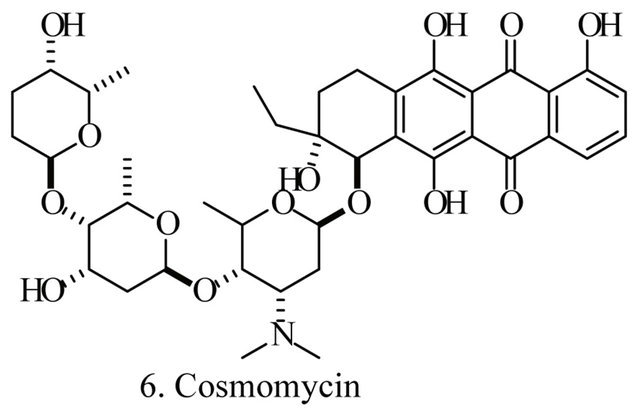 (f)
(f)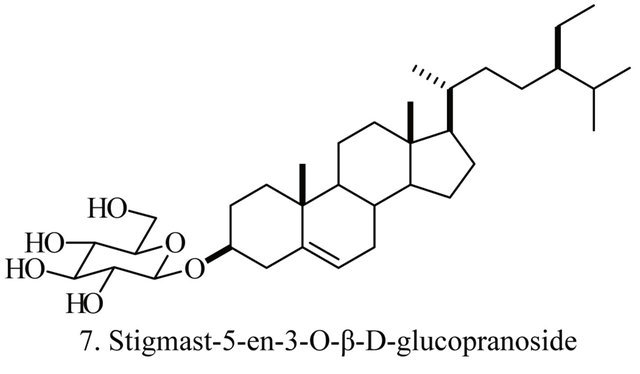 (g)
(g)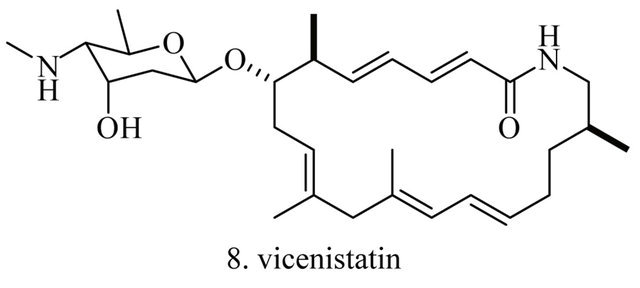 (h)
(h)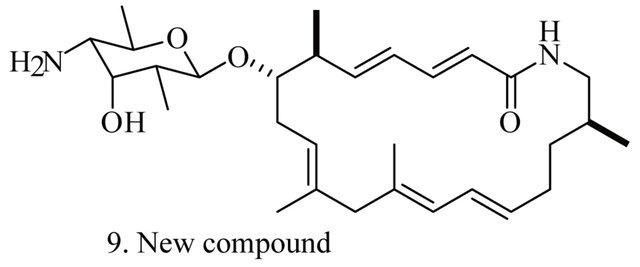 (i)
(i)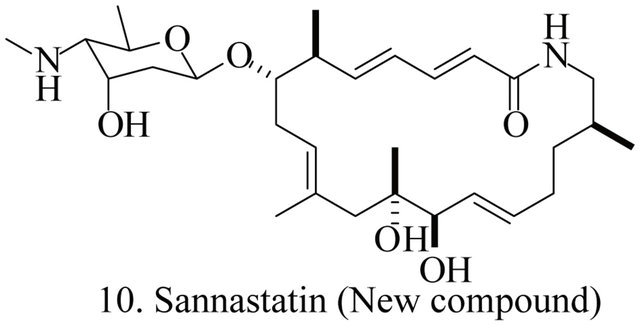 (j)
(j)
Figure 6. A part of bioactive compounds produced by actinomycete strains from animal feces.
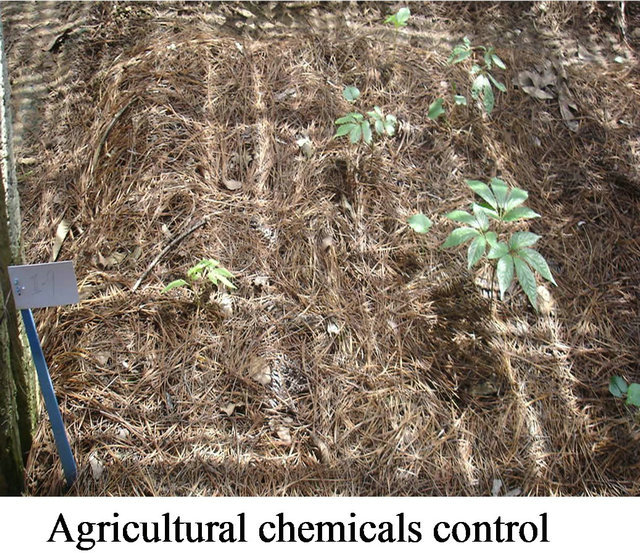
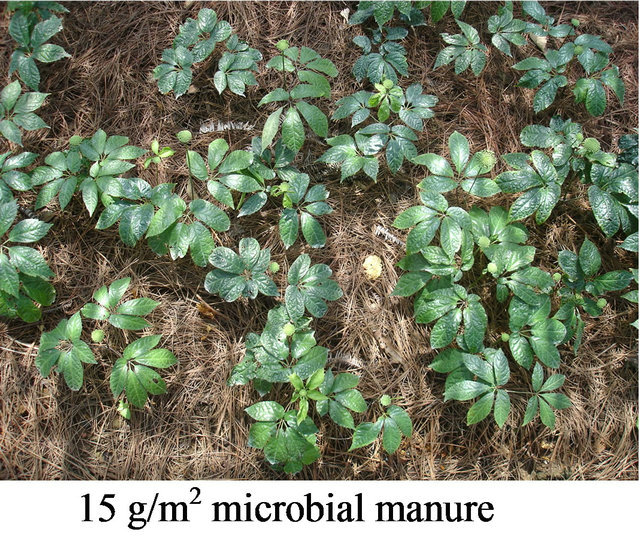
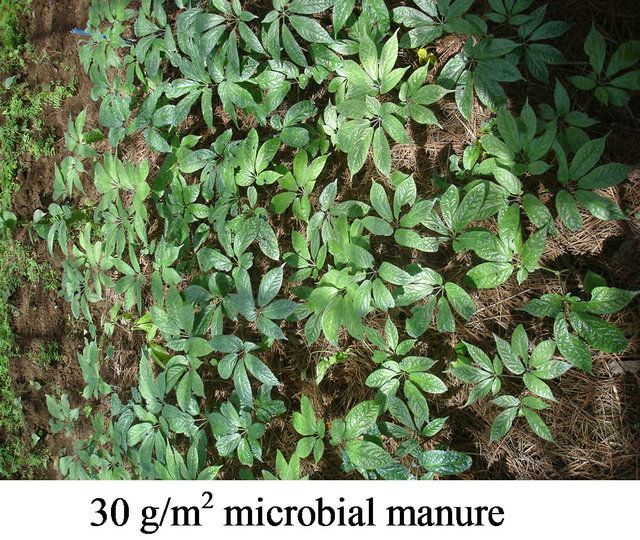



Figure 7. Effectiveness of prophylaxis and treatment of the microbial manure to soil-borne.
In conclusion, animal fecal actinobacteria, like those in soil, oceans, extreme environments and plants, has high diversity but different composition. Enormous unknown actinobacteria widely exist in animal feces. Therefore, animal fecal actinobacteria is important resources for developing novel antibiotics, anti-tumor agents, enzyme inhibitors, immunity inhibitor, agricultural chemicals, enzymes and other useful products. Both Streptomyces and Rhodococcus were the predominant genera and the widest distribution in animal feces. Genome sizes of these two genera are up to 9 × 107 base pairs, one of biggest genome in actinbacteria, and some species of them contains 20 or more natural product biosynthetic gene clusters [42-44]. We showed further a hypothesis that first, the function of actinomycetes in intestinal tract of hosts was played mainly through bioactive substances produced by members of the two genera; Second, secondary metabolites with bioactivities produced by fecal actionbacteria, except pathogen, should be no toxic or lower toxic to their hosts. Maybe these are the most important traits comparing with the microorganisms from other habitats.
4.3. Key of Isolating Actinobacteria from Animal Feces
Existence of Gram negative bacteria in a large number in animal feces is a main problem for isolation of fecal actinobacteria. In order to eliminate the trouble of Gram negative bacteria and fungi, and obtain much more unknown actinobacteria for discovering novel lead compounds, sampling and isolation methods are key points.
Based on many tests in our laboratory, first, fresh feces samples had to been dried at 25˚C - 28˚C for 7 to 10 days; second, pre-treatment of dried samples at 80˚C for 60 min has to be carried out before isolation; third, potassium bichromate 50 mg and 5 mg penicillin, or nystatin 50 mg, nalidixic acid 20 mg and 5 mg penicillin for 1000 ml medium, as inhibitors, have to be added in the isolation medium for inhibiting fungi and Gram negative bacteria; fourth, The dilution of samples should be 10−5, 10−6, and 10−7; fifth, YIM 212, YIM 171 and HV medium were better for isolation of actinobacteria from animal feces.
5. Acknowledgements
This research was supported by the National Natural Science Foundation of China (No. 30900002, 31270001 and 21062028), National Major scientific and technology special projects (2009ZX09302-003), National Institutes of Health USA (1P41GM086184-01A1). We thank Prof. H. Laatsch and Dr. S. X. Yang (University of Göttingen, Germany), Prof. G. L. Challis and Dr. L. J. Song (University of Warwick, UK) for analysis of bioactive metabolites. We thank Prof. Chenglin Jiang and Prof. M. Goodfellow for their guidance.
REFERENCES
- D. C. Savage, “Microbial Ecology of Gastrointestinal Tract,” Annual Review of Microbiology, Vol. 31, 1977, pp. 107-133. doi:10.1146/annurev.mi.31.100177.000543
- J. M. Simpson, B. Martineau, W. E. Jones, J. M. Ballam and R. I. Mackie, “Characterization of Fecal Bacterial Population in Canines: Effecs of Age, Breed and Dietary Fiber,” Microbial Ecology, Vol. 44, No. 2, 2002, pp. 186- 197. doi:10.1007/s00248-002-0001-z
- K. Daly, C. S. Stewart, H. J. Flint and S. P ShiraziBeechey, “Bacterial Diversity within the Equine Large Intestine as Revealed by Molecular Analysis of Cloned 16S rRNA Genes,” FEMS Microbiology Ecology, Vol. 38, No. 2-3, 2001, pp. 141-151. doi:10.1111/j.1574-6941.2001.tb00892.x
- L. M. Durso, G. P. Harhay, T. P. L. Smith, J. L. Bono, T. Z. DeSantis, D. M. Harhay, G. L. Anderson, J. E. Keen, W. W. Laegreid and M. L. Clawson, “Animal-to-Animal Variation in Fecal Microbial Diversity among Beef Cattle,” Applied and Environmental Microbiology, Vol. 76, No. 14, 2010, pp. 4858-4862. doi:10.1128/AEM.00207-10
- R. C. Gao. Z. X. Huang, C. F. Wu, B. Xu and X. Y. Wang, “Culture-Independent Analysis of Microflora in Gayals (Bos Frontalis) Feces,” African Journal of Biotechnology, Vol. 9, No. 19, 2010, pp. 2774-2788.
- H. J. Greetham, C. Giffard, R. A. Hutson, M. D. Collins and G. R. Gibson, “Becteriology of the Laborador Dog Gut: A Cultural and Genotypic Approach,” Journal of Applied Microbiology, Vol. 93, No. 4, 2002, pp. 640-646. doi:10.1046/j.1365-2672.2002.01724.x
- L. E. Ritchie, J. M. Steiner and J. S. Suchodoski, “Assessment of Microbial Diversity along the Feline Interatinal Trct Using 16S Rrna Gene Analysis,” FEMS Microbiology Ecology, Vol. 66, No. 3, 2008, pp. 590-598. doi:10.1111/j.1574-6941.2008.00609.x
- J. S. Suchodoski, J. Camacho and J. M. Steiner, “Analysis of Bacterial Diversity in the Canine, Duodenum, Jejunum, Ileum, and Colon by Comparative 16S rRNA Gene Analysis,” FEMS Microbiology Ecology, Vol. 66, No. 3, 2008, pp. 567-578. doi:10.1111/j.1574-6941.2008.00521.x
- M. M. Curtis and V. Sperandio, “A Complex Relationship: The Interaction among Symbiotic Microbes, Invading Pathogens, and Their Mammalian Host,” Mucosal Immunology, Vol. 4, No. 2, 2011, pp. 133-138. doi:10.1038/mi.2010.89
- L. V. Hooper and J. I. Gorden, “Commensal Host-Bacyrtial Relationship in the Gut,” Science, Vol. 292, No. 5519, 2001, pp. 1115-1118. doi:10.1126/science.1058709
- M. Falagas, G. I. Betsi and S. Athanasiou, “Probiotics for Prevention of Recurrent Vulvovaginal Candidiasis: A Review,” Journal of Applied Microbiology, Vol. 58, No. 2, 2006, pp. 266-272. doi:10.1093/jac/dkl246
- A. Meyer, M. Micksche, I. Herbacek and I. Elmadfa, “Daily Intake of Probiotic as Well as Conventional Yogurt has a Stimulating Effect on Cellular Immunity in Young Healthy Women,” Annals of Nutrition and Metabolism, Vol. 50, No. 3, 2006, pp. 282-289. doi:10.1159/000091687
- Y. Jiang, P. Xu, K. Lou, L. H. Xu and Z. H. Liu, “Problem and Countermeasure on Development of Pharmaceuticals from Actinomycete Resources,” Microbiology, Vol. 35, No. 2, 2008, pp. 272-274.
- D. J. Payne, J. Bradley, J. E. Edwards, D. G. Scheld and J. G. Bartlett, “Drugs from Bad Bugs: Confronting the Challenges of Antibacterial Discovery,” Nature Reviews: Drug Discovery, Vol. 6, No. 1, 2007, pp. 29-40. doi:10.1038/nrd2201
- J. Berdy, “Bioactive Microbial Metabolites,” The Journal of Antibiotics, Vol. 58, No. 1, 2005, pp. 1-26. doi:10.1038/ja.2005.1
- R. H. Baltz, “Renaissance in Antibacterial Discovery from Actinomycetes,” Current Opinion in Pharmacology, Vol. 8, No. 5, 2008, pp. 1-7. doi:10.1016/j.coph.2008.04.008
- Y. Jiang, Y. R. Cao, J. Wiese, K. Lou, L. X. Zhao, J. F. Imhoff and C. L. Jiang, “A New Approach of Research and Development on Pharmaceuticls from Actinomycetes,” Journal of Life Science US, Vol. 3, No. 7, 2009, pp. 52-56.
- P. R. Jensen, “Linking Species Concepts to Natural Product Discovery in the Post-Genomic Era,” Journal of Industrial Microbiology & Biotechnology, Vol. 37, No. 3, 2010, pp. 219-224. doi:10.1007/s10295-009-0683-z
- L. H. Xu, W. J. Li, Z. H. Liu and C. L. Jiang, “Actinomycete Taxonomy,” Acedemic Press, Beijing, 2007.
- B. L. Beman, M. Goodfellow, M. Mordarski and S. T. Williams, “Actinomycete Pathogen in the Biology of the Actinomyces,” Academic press, London, 1983.
- Y. Jiang, Y. R. Cao, L. Han, R. X. Jin, D. Zheng, W. X. He, Y. L. Li and X. S. Huang, “Diversity and Bioactivity of Culturable Actinobacteria from Animal Feces,” Acta Microbiol Sinica, Vol. 52, No. 10, 2012, pp. 1282-1289.
- E. H. Shirling and D. Gottlieb, “Methods for Characterization of Streptomyces Species,” International Journal of Systematic and Evolutionary Macrobiology, Vol. 16, No. 3, 1966, pp. 313-340. doi:10.1099/00207713-16-3-313
- X. L. Cui, P. H. Mao, M. Zeng, L. H. Xu and C. L. Jiang, “Streptomonospora Salina Gen. Nov., Sp. Nov., A New Member of the Family Nocardiopsaceae,” International Journal of Systematic and Evolutionary Macrobiology, Vol. 51, No. 2, 2001, pp. 357-363.
- N. Saitou and M. Nei, “The Neighbor-Joining Method: A New Method for Reconstructing Phylogenetic Trees,” Molecular Biology and Evolution, Vol. 4, No. 4, 1987, pp. 406-425.
- J. Felsenstein, “Evolutionary Trees from DNA Sequences: A Maximum Likelihood Approach,” Journal of Molecular Evolution, Vol. 17, No. 6, 1981, pp. 368-376. doi:10.1007/BF01734359
- I. González, A. Ayuso-Sacido, A. Anderson, O. Genilloud, “Actinomycetes Isolated from Lichens: Evaluation of Their Diversity and Detection of Biosynthetic Gene Sequences,” FEMS Microbiology Ecology, Vol. 54, No. 3, 2005, pp. 401-415. doi:10.1016/j.femsec.2005.05.004
- Y. B. Hwang, M. Y. Lee, H. J. Park, K. Han and E. S. Kim, “Isolation of Putative Polyene-Producing Actinomycetes Strains via PCR-Based Genome Screening for Polyene-Specific Hydroxylase Genes,” Process Biochemistry, Vol. 42, No. 1, 2007, pp. 102-107. doi:10.1016/j.procbio.2006.06.031
- H. T. Zhang, A. M. Liu, L. Z. Wu, G. Z. Sun, F. Han, Q. J. Gao, Y. Q. Zhang and Y. G. Wang, “Screening of AHBA Synthase Gene and Discovery of Actinomycin D Production in Streptomyces Violaceusniger 4353,” Journal of Chinese Antibiotics, Vol. 34, No. 1, 2009, pp. 30-33.
- S. N. R. Gundlapally and G. P. Ferran, “Description of Patulibacter Americanus Sp. Nov., Isolated from Biological Soil Crusts, Emended Description of the Genus Patulibacter Takahashi et al. 2006 and Proposal of Solirubrobacterales ord. nov. and Thermoleophilales ord. nov,” International Journal of Systematic and Evolutionary Macrobiology, Vol. 59, No. 1, 2009, pp. 87-94.
- H. Rheims, P. Schumann, M. Rohde and E. Stackebrandt, “Verrucosispora Gifhornensis Gen. Nov., sp. Nov., A New Member of the Actinobacterial Family Micromonosporaceae,” International Journal of Systematic and Evolutionary Macrobiology, Vol. 48, No. 4, 1998, pp. 1119- 1127. doi:10.1099/00207713-48-4-1119
- W. J. Li. X. Y. Zhi and J. P. Euzéby, “Proposal of Yaniellaceae Fam. Nov., Yaniella Gen. Nov. and Sinobaca Gen. Nov. as Replacements for the Illegitimate Prokaryotic Names Yaniaceae Li et al. 2005, Yania Li et al. 2004, Emend Li et al. 2005 and Sinococcus Li et al. 2006, Respectively,” International Journal of Systematic and Evolutionary Macrobiology, Vol. 58, No. 2, 2008, pp. 525- 527. doi:10.1099/ijs.0.65792-0
- S. K. Tang, X. Y. Zhi, Y. Wang, R. Shi, K. Lou, L. H. Xu and W. J. Li, “Haloactinopolyspora Alba gen. nov., sp. nov., A Halophilic Filamentous Actinomycete Isolated from a Salt Lake, with Proposal of Jiangellaceae fam. nov. and Jiangellineae Subord. Nov,” International Journal of Systematic and Evolutionary Macrobiology, Vol. 61, No. 1, 2011, pp. 194-200. doi:10.1099/ijs.0.021725-0
- Y. R. Cao, Y. Jiang, R. X. Jin, L. Han, W. X. He, Y. L. Li, X. S. Huang and Q. H. Xue, “Enteractinococcus Coprophilus gen. nov., sp. nov., of the Family Micrococcaceae Isolated from Panthera Tigris Amoyensis Feces, Transfer of Yaniella Fodinae Dhanjal et al. 2011 to the Genus Enteractinococcus as Enteractinococcus Fodinae comb. Nov.,” International Journal of Systematic and Evolutionary Macrobiology, Vol. 62, No. 11, 2012, pp. 2710-2716. doi:10.1099/ijs.0.034249-0
- Y. R. Cao, Y. Jiang and L. H. Xu, “Actinomycete Composition and Bioactivities in Grand Shangri-La,” Acta Microbiol Sinica, Vol. 49, No. 1, 2009, pp. 105-109.
- Y. R. Cao, Y. Jiang, Q. Wang, L. X. Zhao, R. X. Jin and C. L. Jiang, “Diversity and some Bioactivities of Cultured Actinomycetes in four Areas in Sichuan and Yunnan,” Acta Microbiol Sinica, Vol. 50, No. 8, 2010, pp. 995- 1000.
- Y. Jiang, J. Wiese, Y. R. Cao, L. X. Zhao, R. X. Jin and J. F. Imhoff, “Diversity of Cultured Actinomycete in the Baltic Sea,” Acta Microbiol Sinica, Vol. 51, No. 11, 2011, pp. 1461-1457.
- Y. Jiang, W. J. Li, P. Xu, S. K. Tang and L. H. Xu, “Study on Actinomycete Diversity under Salt and Alkaline Environments,” Acta Microbiol Sinica, Vol. 46, 2006, pp. 191-195.
- M. P. Lechevalier and H. A. Lechevalier, “Chemical Composition as a Criterion in the Classification of Aerobic Actinomycetes,” International Journal of Systematic and Evolutionary Macrobiology, Vol. 20, No. 4, 1970, pp. 435- 443. doi:10.1099/00207713-20-4-435
- M. P. Lechevalier and H. A. Lechevalier, “The Chemotaxonomy of Actinomycetes,” In: A. Dietz and D. W. Thayer, Eds., Actinomycete Taxonomy, Special Publications Society for Industrial Microbiology, Arlington, 1980, pp. 227-291.
- M. Goodfellow and H. P. Fiedler, “A Guide to Successful Bioprospecting: Informed by Actinobacterial Systematics,” Antonie van Leeuwenhoek, Vol. 98, No. 2, 2010, pp. 119-142. doi:10.1007/s10482-010-9460-2
- S. X. Yang, J. M. Gao. A. L. Zhang and H. Laatsch, “Sannastatin, a Novel Toxic Macrolactam Polyketide Glycoside Produced by Actinomycete Streptomyces Sannanensis,” Bioorganic & Medicinal Chemistry Letters, Vol. 21, No. 13, 2011, pp. 3905-3908. doi:10.1016/j.bmcl.2011.05.028
- S. Omura, H. Ikeda, J. Ishikawa, et al., “Genome Sequence of an Industrial Microorganism Streptomyces Avermitilis: Deducing the Ability of Producing Secondary Metabolites,” PNAS, Vol. 98, No. 21, 2001, pp. 12215-12220. doi:10.1073/pnas.211433198
- S. D. Bentley, K. F. Chater, A.-M. Cerdeno-Tarraga, et al., “Complete Genome Sequence of the Model Actinomycete Streptomyces Coelicolor A3(2),” Nature, Vol. 417, 2002, pp. 141-147. doi:10.1038/417141a
- P. M. Michael, L. W. Rene, W. L. H. William, et al., “The Complete Genome of Rhodococcus sp. RHA1 Provides Insights into a Catabolic Powerhouse,” PNAS, Vol. 103, No. 42, 2006, pp. 15582-15587. doi:10.1073/pnas.0607048103
NOTES
*Corresponding author.

