Natural Resources
Vol.3 No.4(2012), Article ID:25608,4 pages DOI:10.4236/nr.2012.34022
Heavy-Metal Tolerance and Antibiotic Susceptibility of Red Pigmented Bacteria Isolated from Marine Environment
![]()
1School of Bioscience and Biotechnology, Universiti Kebangsaan Malaysia, Bangi, Malaysia; 2School of Environmental Science and Natural Resources, Faculty of Science and Technology, Universiti Kebangsaan Malaysia, Bangi, Malaysia.
Email: *jafarzadem@yahoo.com
Received August 16th, 2012; revised September 22nd, 2012; accepted October 8th, 2012
Keywords: Antibiotic Susceptibility; Heavy Metal Resistance; Serratia sp; Marine Bacteria
ABSTRACT
This study was undertaken to determine heavy metal resistance and antibiotic susceptibility of three non-pathogenic red pigmented bacteria namely WPRA3, SM11-3j and SC-G18, isolated from marine environments of Malaysia. The bacteria isolates were identified by 16S rRNA sequencing and by biochemical and morphological tests. The 16S rRNA gene sequences of all isolates showed ≥96% similarity to Serratia spp. Antibiotic susceptibility test of isolates was assayed according to the Kirby-Bauer disc diffusion method. All isolates were highly resistant to beta-lactam antibiotics, but were susceptible to quinolone antibiotics. Minimum inhibitory concentration (MIC) of nine heavy metals (Ni2+, Co2+, Cr3+, Zn2+, Mn2+, Pb2+, Hg2+, Cd2+ and Cu2+) against the bacteria isolates were determined via the plate-dilution method. The isolates exhibited resistance to Ni2+, Co2+, Cr3+ and Zn2+. Isolates WPRA3 and SM11-3j showed higher multiple tolerances to heavy metals. The results obtained indicate that bacteria from marine environments of Malaysia present interesting metabolic activities, which should be studied and explored for potential biotechnological applications.
1. Introduction
The genus Serratia are gram negative bacteria, classified in the large family of Entrobacteriaceae of the class Gammaproteobacteria. Serratia species are widespread in the environment, occurring in soil, water, plants, vertebrates and humans. S. marcescens has been identified as the cause of many hospital epidemics [1]. The greatest impediment to the successful treatment of bacterial diseases is antibiotic resistance that is widespread and occurs in numerous bacterial genera [2]. Winshell and Neu [3] divided Serratia marcescens strains into three groups on the basis of antimicrobial resistance and pigment production. Group I, comprising non-pigmented strains, are resistant to most antibiotics. Group II, which are non-pigmented, and group III, which are pigmented, are susceptible to many antimicrobial agents. In general the pigmented strains are more susceptible to antibiotics.
Pollution of the marine environment by heavy metals has been recognized as one of the important problems that emerged from industrial, sewage treatment discharges and anti-fouling paints and natural sources include weathering of rocks, leaching of soils and volcanic eruptions [4,5]. Four classes of heavy metal can be differentiated in seawater depending on their concentration [6]. Class I, common elements with concentrations between 100 nM and 1 μM (Fe, Zn, Mo), class II, elements with concentrations between 10 nM and 100 nM (Ni, Cu, As, V, Mn, Sn, U), class III, rare elements (Co, Ce, Ag, Sb) and finally class IV, elements just below the 1 nM level (Cd, Cr, W, Ga, Zr, Th, Hg, Pb) [7]. Sn, Ce, Ga, Zr and Th have no biological influence because of the low solubility of the trior tetravalent cations [6]. Fe, Mo and Mn are important trace elements with low toxicity and Zn, Ni, Cu, V, Co, W and Cr are toxic elements with high to moderate importance as trace elements; As, Ag, Sb, Cd, Hg, Pb and U have limited beneficial function, but have to be considered as toxins [7]. In this study, we evaluated the heavy metal resistance and antibiotic susceptibility of three red pigmented Serratia sp. isolates designated WPRA3, SM11-3j and SC-G18.
2. Experimental
2.1. Bacterial Strain
Red pigmented bacteria designated as WPRA3, SM11-3j and SC-G18 were isolated from seawater of Port Dickson, mussel from Sungai Merbok (Kedah) and sea cucumber from Tinggi Island of Malaysia, respectively. Isolates were routinely grown in Marine Agar (MA; Difco). Identification of isolates was done by 16S rRNA sequence analysis. PCR amplification of the 16S rRNA gene was performed using two oligonucleotide primers, forward 5’-CTCCTACGGGAGGCAGCAG-3’ and reverse 5’-WATTACCGCGGCKGCTG-3’. The PCR programme was set using thermal minicycler (MJ Research, USA) as follows: initial denaturation was carried out for 5 min at 94˚C. It was followed by 35 cycles of denaturation at 94˚C for 45 sec, annealing at 55˚C for 45 sec and extension at 72˚C for 1.5 min with a further 10 min extension at 72˚C, using UniversAllTM tissue PCR kit. PCR product was purified using QIAquick PCR Purification Kit (QIAGEN, Germany) according to manufacturer’s instructions. Nucleotide sequences were analyzed by using BioEdit Sequence Alignment Editor. The 16S rRNA gene sequences of the isolates were compared with available 16S rRNA gene sequences in GenBank databases using the BLASTn search facility (http://www.ncbi.nlm.nih.gov). Morphological and biochemical tests were determined according to Bergey’s Manual using an isolated colony of bacteria from MA.
2.2. Antibiotic Susceptibility Assay
Antibiotic susceptibility test of the isolated bacteria was assayed according to the Kirby-Bauer disc diffusion method given by Bauer et al. [8]. Briefly, isolated bacteria were cultured in marine broth (MB; Difco) at 28˚C for 24 hours. Sterile swabs were immersed in the microbial suspensions (108 cells/mL) and applied to Muller Hinton Agar (MHA; Difco) plates immediately. Standard antibiotics (Oxoid) used were penicillin (10 U), ampicillin (10 µg), tetracycline (30 µg), chloramphenicol (10 µg, 30 µg), nalidixic acid (30 µg), streptomycin (10 µg), gentamicin (10 µg) and kanamycin (30 µg) were placed on the agar in the MHA plates. The plates were incubated at 28˚C for 24 hours. The result of inhibition zones of the isolates were interpreted as sensitive (S), intermediate (I) and resistant (R) according to the references to the standard provided by BD BBLTM Sensi-DiscTM Zone Interpretation Set (Oxoid). All of the experiments were performed in triplicate.
2.3. Heavy Metal Resistance
Isolated bacteria were tested to determine the minimal inhibitory concentrations (MICs) of the metals by the platedilution method as described by Malik and Jaiswal [9]. The metals Ni2+, Co2+, Cr3+, Zn2+, Mn2+, Pb2+, Hg2+, Cd2+ and Cu2+ were used as NiSO4, CoSO4·7H2O, Cr(NO2)3·9H2O, ZnSO4, MnSO4·H2O, Pb(NO3)2, HgCl2, CdSO4 and CuSO4 respectively. Stocks of the metal salts were prepared in deionised water and sterilized by filter membrane (0.2 μm) and stored at 4˚C. The heavy metal solutions were added to tryptone soy agar (TSA; Difco) in various concentrations ranging from 50 to 800 μg/mL. The plates were incubated at 28˚C for 48 hours. The concentration of the metal at which the isolate failed to grow on the plate was considered as the MIC. Colour changes of the isolate were also observed.
3. Results and Discussion
3.1. Bacterial Strains
Isolated bacteria were Gram-negative, facultative anaerobic, red pigmented and short rod-shaped bacteria. They were positive for catalase, the citrate utilization test, β-galactosidase, lysine decarboxylase and ornithine decarboxylase. Oxidase and indole tests were negative. Isolate WPRA3 was negative for the Voges Proskauer test, while SM11-3j and SC-G18 isolates were positive. Comparative 16S rRNA gene sequence analysis showed that all three isolates WPRA3, SM11-3j and SC-G18 shared ≥96% similarities with Serratia sp.
3.2. Antibiotic Susceptibility
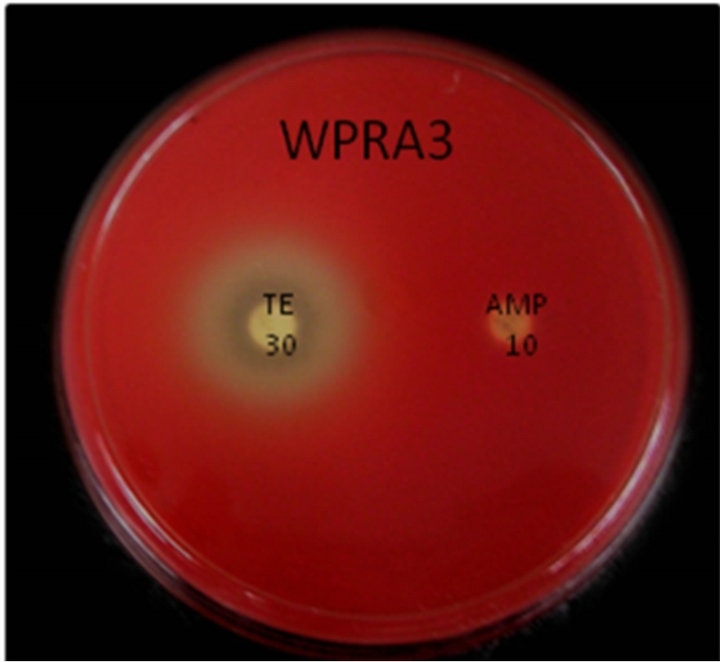
This assay revealed that isolates WPRA3, SC-G18 and SM11-3j were highly resistant to penicillin, ampicillin and tetracycline. Isolated bacteria showed high sensitivity to nalidixic acid and they were also susceptible to streptomycin, kanamycin and gentamicin. All isolate were susceptible to chloramphenicol. Antibiotic resistance of the isolate WPRA3, to tetracycline and ampicillin is shown in Figure 1(a), while antibiotic susceptibility of the isolates SM11-3j and SC-G18 to nalidixic acid is shown in Figures 1(b) and (c). The antibiotic susceptibility of the isolates WPRA3, SM11-3J and SC-G18 were similar to other non-pathogenic Serratia species such as S. marcescens subsp. sakuensis JCM 11315 [10], but they were more resistant to antibiotics than S. nemathodiphila DZ0503SBS1 [11], although in comparison with pathogenic Serratia species such as S. marcescens strain 08 [12] and S. marcescens [13] they were more susceptible to antibiotics. Table 1 shows the antibiotic susceptibility of isolates WPRA3, SM11-3J and SC-G18 in comparison with other Serratia species.
3.3. Heavy Metal Resistance
Isolate WPRA3 exhibited the highest heavy metal resistance as compared to isolates SM11-3j and SC-G18. The heavy metal resistance patterns of isolates WPRA3, SM11-3j and SC-G18 were Mn, Pb > Zn > Cr > Ni > Co, Mn > Cr > Zn > Ni, Co and Zn > Ni, Cr > Co respectively (Table 2). Isolates WPRA3, SM11-3j and SC-G18 were unable to grow in metal-agar media at any concentration
 (a)
(a)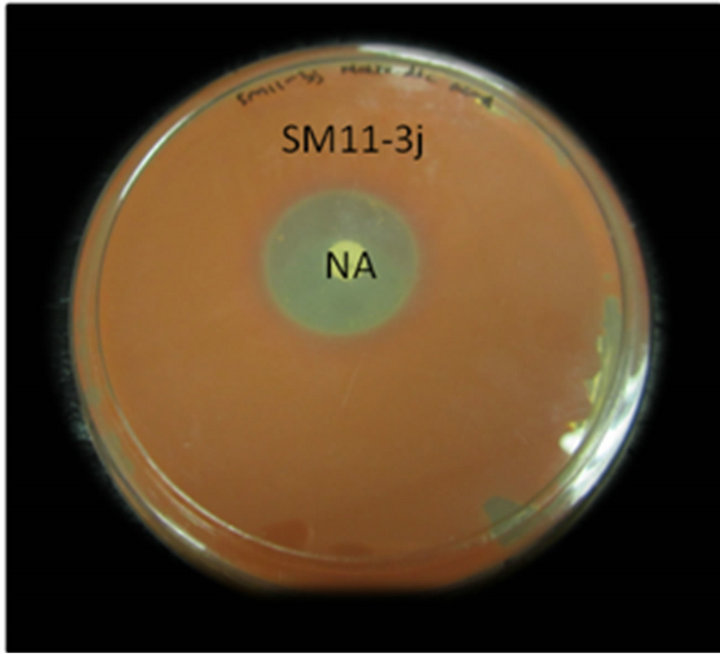 (b)
(b)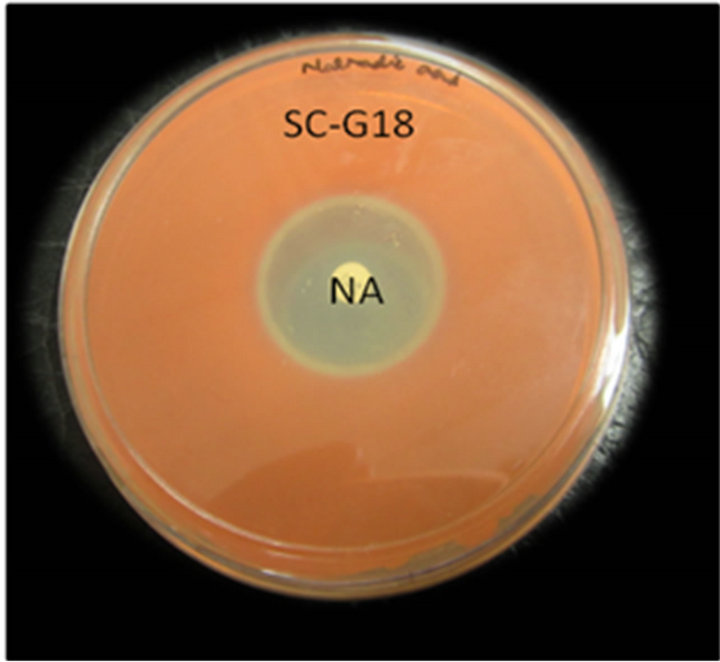 (c)
(c)
Figure 1. Antibiotic resistance of isolate WPRA3 to tetracycline (TE) and ampicillin (AMP) (a), antibiotic resistance of isolates SM11-3j (b) and SC-G18 (c) to nalidixic acid (NA).
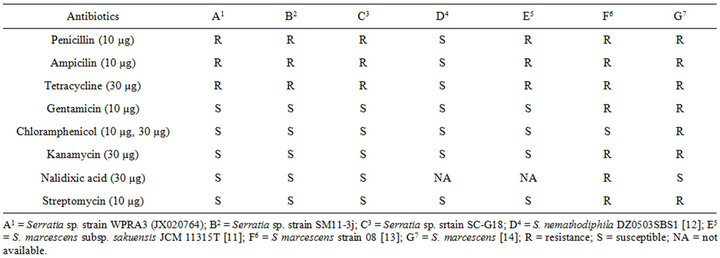
Table 1. Antibiotic susceptibility of isolated bacteria in comparison with other Serratia species.
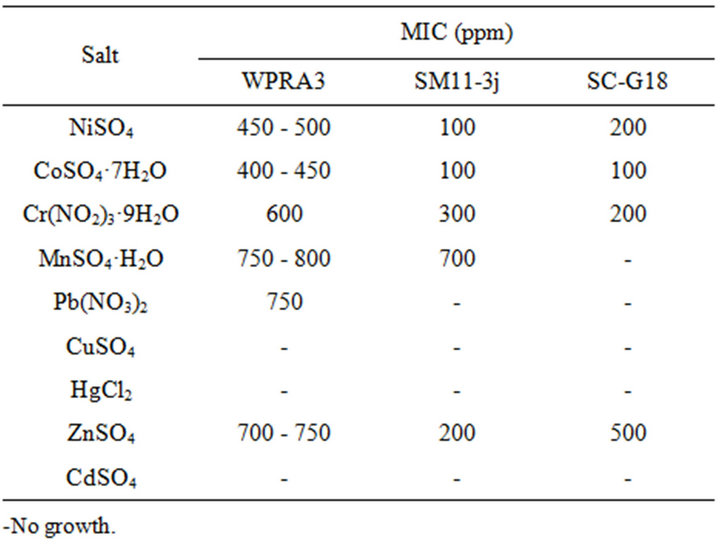
Table 2. Minimum inhibitory concentrations of heavy metal for isolated bacteria.
of copper, mercury and cadmium, may because of high toxicity of these metals [7]. Meanwhile, Isolates WPRA3 and SM11-3j showed the highest resistance to Mn, maydue to the lowest toxicity of Mn amongst other elements
[7]. In the presence of Ni, Pb and Co, isolate WPRA3 did not change its colony color and shape, while the color of the colony changed from red to pink in the presence of Zn. Isolate WPRA3 was unable to produce color in the presence of Mn and Cr. Color of colonies of isolates SM11-3j and SC-G18 were also changed from red to pink in the presence of Zn. Some Serratia species have also been reported as heavy metals tolerant, such as; Serratia sp. K1RP-49 could tolerate Pb, whereas it could not grow in the medium supplemented with Cd, Cr, Cu, Ni and Zn [14]. S. marcescens Strain C-1 was reported as Cobalt and Nickel resistant. The MIC values for strain C-1 were 30 mM Co (II) and 20 mM Ni (II) [15]. Serratia (PL18), isolated from a pitch lake located in Trinidad, was able to tolerate Pb (4 mM), Ni (5 mM), V (9 mM), Se (9 mM) and Zinc sulfate (9 mM) [16]. Moreover, some studies have evaluated the use of S. marcecsens for the degradation of Mo [17,18] and Cr [18,19].
4. Conclusion
Biochemical, morphological and molecular characteristics showed that the red pigmented isolates belonged to the genus Serratia. The Serratia sp. isolate WPRA3 (JX020764) and Serratia sp. isolate SM11-3j have broad ranges of multi heavy metal (Ni, Co, Cr, Zn, Mn and Pb both in broth and agar medium) and antibiotic (penicillin, ampicilline and tetracycline) resistance. Thus these isolates could potentially be used in the treatment of Industrial effluents. They could also serve as models for studies on heavy metal and antibiotic resistance in marine bacteria.
5. Acknowledgements
This work was supported by grants 04-01-02-SF014 and UKM GUP BTK 07-75-198.
REFERENCES
- F. Grimont and P. A. D. Grimont, “The Genus Serratia,” Prokaryotes, Vol. 6, 2006, pp. 219-244. doi:10.1007/0-387-30746-X_11
- I. Chopra and M. Roberts, “Tetracycline Antibiotics: Mode of Action, Applications, Molecular Biology, and Epidemiology of Bacterial Resistance,” Microbiology and Molecular Biology Reviews, Vol. 65, No. 2, 2001, pp. 232- 260. doi:10.1128/MMBR.65.2.232-260.2001
- E. B. Winshell and H. C. Neu, “Relation of Cell Wall Lipid Content of Serratia marcescens to Resistance to Antimicrobial Agents,” Antimicrobial Agents and Chemotherapy, Vol. 6, No. 1, 1974, pp. 73-75. doi:10.1128/AAC.6.1.73
- C. L. Mitchelmore, E. A. Verde, A. H. Ringwood and V. M. Weis, “Differential Accumulation of Heavy Metals in the Sea Anemone Anthopleura elegantissima as a Function of Symbiotic State,” Aquatic Toxicology, Vol. 64, No. 3, 2003, pp. 317-329. doi:10.1016/S0166-445X(03)00055-9
- M. H. Al-Yousuf, El-Shahawi and S. M. Al-Chais, “Trace Metals in Liver, Skin and Muscle of Lethrinus lentjan Fish Species in Relation to Body Length and Sex,” Science of the Total Environment, Vol. 256, No. 2-3, 2000, pp. 87-94. doi:10.1016/S0048-9697(99)00363-0
- R. C. Weast, “CRC Handbook of Chemistry and Physics 1983-1984,” 64th Edition, CRC, Boca Raton, 1983.
- D. H. Nies, “Microbial Heavy-Metal Resistance,” Applied Microbiology and Biotechnology, Vol. 51, No. 6, 1999, pp. 730-750. doi:10.1007/s002530051457
- A. W. Bauer, W. M. M. Kirby, J. C. Sherris and M. Turck, “Antibiotic Susceptibility Testing by a Standardized Single Disk Method,” The American Journal of Clinical Pathology, Vol. 45, No. 4, 1966, pp. 493-496.
- A. Malik and R. Jaiswal, “Metal Resistance in Pseudomonas Strains Isolated from Soil Treated with Industrial Wastewater,” World Journal of Microbiology and Biotechnology, Vol. 16, No. 2, 2000, pp. 177-182. doi:10.1023/A:1008905902282
- B. Ajithkumar, V. P. Ajithkumar, R. Iriye, Y. Doi and T. Sakai, “Spore-Forming Serratia marcescens subsp. sakuensis subsp. nov., Isolated from a Domestic Wastewater Treatment Tank,” International Journal of Systematic and Evolutionary Microbiology, Vol. 53, No. 1, 2003, pp. 253- 258. doi:10.1099/ijs.0.02158-0
- C. X. Zhang, S. Y. Yang, M. X. Xu, J. Sun, H. Liu, J. R. Liu, H. Liu, F. Kan, J. Sun, R. Lai and K. Y. Zhang, “ Serratia nematodiphila sp. nov., Associated Symbiotically with the Entomopathogenic Nematode Heterorhabditidoides chongmingensis (Rhabditida: Rhabditidae),” International Journal of Systematic and Evolutionary Microbiology, Vol. 59, 2009, pp. 1603-1608.
- G. L. Button, M. A. Miller and J. C. Tsang, “Antibiogram and Lipid Analysis of a Pigmented Strain of Serratia marcescens and Its Nonpigmented Variants ANTrmcROBnL,” Antimicrobial Agents and Chemotherapy, Vol. 7, No. 2, 1975, pp. 219-222. doi:10.1128/AAC.7.2.219
- M. J. Ding and S. J. Sung, “Drug Resistance, R. Plasmids and Pigmentation of Serratia marcescens Isolated in Taiwan,” Chinese Journal of Microbiology and Immunology, Vol. 20, No. 1, 1987, pp. 69-79.
- S. Y. Koo and K. S. Cho, “Characterization of Serratia sp. K1RP-49 for Application to the Rhizoremediation of Heavy Metals,” Survival and Sustainability, 2011, pp. 3-13. doi:10.1007/978-3-540-95991-5_1
- J. Marrero, G. Auling, O. Coto and D. H. Nies, “HighLevel Resistance to Cobalt and Nickel but Probably No Transenvelope Efflux: Metal Resistance in the Cuban Serratia marcescens Strain C-1,” Microbial Ecology, Vol. 53, No. 1, 2007, pp. 123-133. doi:10.1007/s00248-006-9152-7
- D. Cruz-Vega, E. Cervantes-González, D. Ammons, L. I. Rojas-Avellzapa, J. García-Mena, R. C. Pless and N. G. Rojasavelizapa, “Tolerance and Removal of Metals by Microorganisms Isolated from a Pitch Lake,” Haz Waste Management: B 1.1, 2008.
- S. M. Yunus, H. M. Hamim, O. M. Anas, S. N. Aripin and S. M. Arif, “Mo (VI) Reduction to Molybdenum Blue by Serratia marcescens Strain Dr. Y9,” Polish Journal of Microbiology, Vol. 58, No. 2, 2009, pp. 141-147.
- M. Y. Shukor, S. H. Habib, M. F. Rahman, H. Jirangon, M. P. Abdullah, N. A. Shamaan and M. A. Syed, “Hexavalent Molybdenum Reduction to Molybdenum Blue by S. marcescens Strain Dr. Y6,” Applied Biochemistry and Biotechnology, Vol. 149, No. 1, 2008, pp. 33-43. doi:10.1007/s12010-008-8137-z
- V. L. Campos, R. Moraqa, J. Yánez, C. A. Zaror and M. A. Mondaca, “Chromate Reduction by Serratia marcescens Isolated from Tannery Effluent,” Bulletin of Environmental Contamination and Toxicology, Vol. 75, No. 2, 2005, pp. 400-406. doi:10.1007/s00128-005-0767-z
NOTES
*Corresponding author.

