Open Journal of Medicinal Chemistry
Vol.3 No.4(2013), Article ID:40387,7 pages DOI:10.4236/ojmc.2013.34012
Piericidins, Novel Quorum-Sensing Inhibitors against Chromobacterium violaceum CV026, from Streptomyces sp. TOHO-Y209 and TOHO-O348
Department of Microbiology, School of Pharmaceutical Sciences, Toho University, Chiba, Japan
Email: *3011001@nc.toho-u.ac.jp
Copyright © 2013 Kazuhiro Ooka et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Received October 3, 2013; revised November 4, 2013; accepted November 12, 2013
Keywords: Piericidin A1; 3’-Rhamnopiericidin A1; Piericidin E; Quorum-Sensing Inhibitor; Chromobacterium violaceum CV026; N-Acylhomoserinlactone
ABSTRACT
Piericidin A1, 3’-rhamnopiericidin A1, and a novel compound piericidin E, a new quorum-sensing (QS) inhibitor against Chromobacterium violaceum CV026, were isolated from the culture broth of Streptomyces sp. QS is well known as a microbial signaling system and controls certain types of gene expression resulting in bioluminescence, biofilm formation, swarming motility, antibiotic biosynthesis, and virulence factor production. C. violaceum CV026 is commonly used to determine qualitative and quantitative QS activity. The structures of piericidin derivatives were characterized, and their QS activities were determined.
1. Introduction
Quorum sensing (QS) is a process for cell-to-cell communication in bacteria using intermediary substances, which are excreted from bacterial cells into the environment. When the concentration of such an excreted metabolite reaches its threshold level, certain types of gene expression resulting in bioluminescence [1], biofilm formation [2,3], swarming motility [4], antibiotic biosynthesis [5,6] virulence factor production [7,8], etc., are triggered by the metabolite (autoinducer). Disruption of the QS system in pathogenic Burkholderia cepacia and B. pseudomallei resulted in reduced pathogenicity in murine and hamster infection [9,10], and erythromycin inhibits biofilm formation of Pseudomonas aeruginosa below the minimum inhibitory concentration (MIC) [11]. Therefore, compounds that inhibit QS have great potential for use in the treatment of bacterial infectious diseases.
We screened metabolites in the culture broth of about 1000 strains of Actinomycetes isolated from soils for their capacity to inhibit the production of violacein in Chromobacterium violaceum CV026 [12,13] and found that the culture broth from 103 strains produced QS inhibitors (QSIs). In this study, the structures of piericidin A1 and its derivatives isolated as QSIs were characterized, and the QS inhibitory activity and taxonomical properties of the strains that produced these QSIs were analyzed.
2. Materials and Methods
2.1. General Experimental Procedures for Screening
Streptomyces sp., isolated from soil, was cultured in a YM medium at 27˚C for 4 - 7 days under vigorous shaking. The culture broth was applied to QSI screening. C. violaceum CV026, kindly provided by Dr. Tsukasa Ikeda, Utsunomiya University, was used for the screening. CV026 was inoculated into 5 mL of Luria-Bertani broth (LB) containing 50 μg∙mL-1 of kanamycin (KM) and incubated at 27˚C for about 18 h under shaking (240 rpm). A culture of C. violaceum CV026 (200 µL) and 20 µL of 10 mM N-hexanoyl-L-homoserine lactone HHL (Santa Cruz Biotechnology Inc.) were added to 20 mL of LuriaBertani soft agar (LSA), pre-warmed at 45˚C, and mixed well, and then immediately poured onto an Luria-Bertani agar (LA) plate. The overlaid plate was cooled (10˚C) until the soft agar layer was solidified. Holes were bored in the plate using a sterilized cork borer (4 mm diameter), and then, a small amount of LSA was added to createan agar pocket. To the pocket, 50 µL of a culture broth of Streptomyces sp. was added, and the plate was incubated at 27˚C until the purple pigment developed. When a circle (white background) around agar pocket was observed, the culture was considered to contain metabolites with QS inhibitory activity.
2.2. General Experimental Procedures for QS Inhibitory Assay
Serial dilutions of piericidin and its analogs were made with methanol, and the dilutions were added to wells in microtiter plate and air-dried over a clean bench until methanol was completely evaporated. To each well, 200 µL of LSA containing C. violaceum CV026 and HHL (the same preparation as QSIs) were added, and then, the microtiter plate was cultured at 27˚C until the purple pigment in the control was well developed. The plate was dried at 60˚C until the water was completely evaporated, and 200 µL of DMSO was added and shaken for 2 h to extract violacein. The OD570 of the DMSO extract was measured.
2.3. Taxonomic Studies
An ISP medium recommended by Shirling and Gottlieb [14] and media recommended by Waksman [15] were used to investigate the cultural and physiological characteristics. The cultures were routinely observed after two weeks of incubation at 27˚C. The color names and hue numbers were determined according to the Color Harmony Manual [16]. The carbon sources were tested by their growth on a Pridham and Gottlieb’s medium containing 1.0% carbon at 27˚C [17]. The type of DAP isomers was determined by the method reported by Becker et al. [18]. The morphological properties were observed using a scanning electron microscope. 16S rDNA fragments of TOHO-Y209 and TOHO-O348 were amplified by polymerase chain reaction (PCR) using the general bacterial 16S rDNA primers 10F (5’- AGTTTGATCCTGGCTC-3’) and 1541R (5’-AAGGAGGTGATCCAGCC-3’). The DNA sequence of the amplified fragments were determined using Genetic Analyzer 3500 (Applied Biosystems, USA) by cycle sequencing with the chain termination technique using dyelabeled dideoxynucleotides.
3. Results
3.1. Taxonomical Properties of the Strains That Produced QSIs
The cultural and physiological characteristicsand utilization of carbon source are shown in Table 1. The isomer of diaminopimelic acid (DAP) in whole-cell hydrolysates of both the strains was determined as the LL-form. The scanning electron micrograph of these strains showed that the spore chains of TOHO-Y209 were spiral, whereas those of TOHO-O348 were straight (Figure 1). The spores of both the strains were cylindrical in shape, 1.0 straight µm in long and had a smooth surface. Whirls, sclerotic granules, sporangia, and flagellate spores were not observed.

Table 1. Comparison of cultural characteristics of strain TOHO-O348 and TOHO-Y209.
 (a)
(a)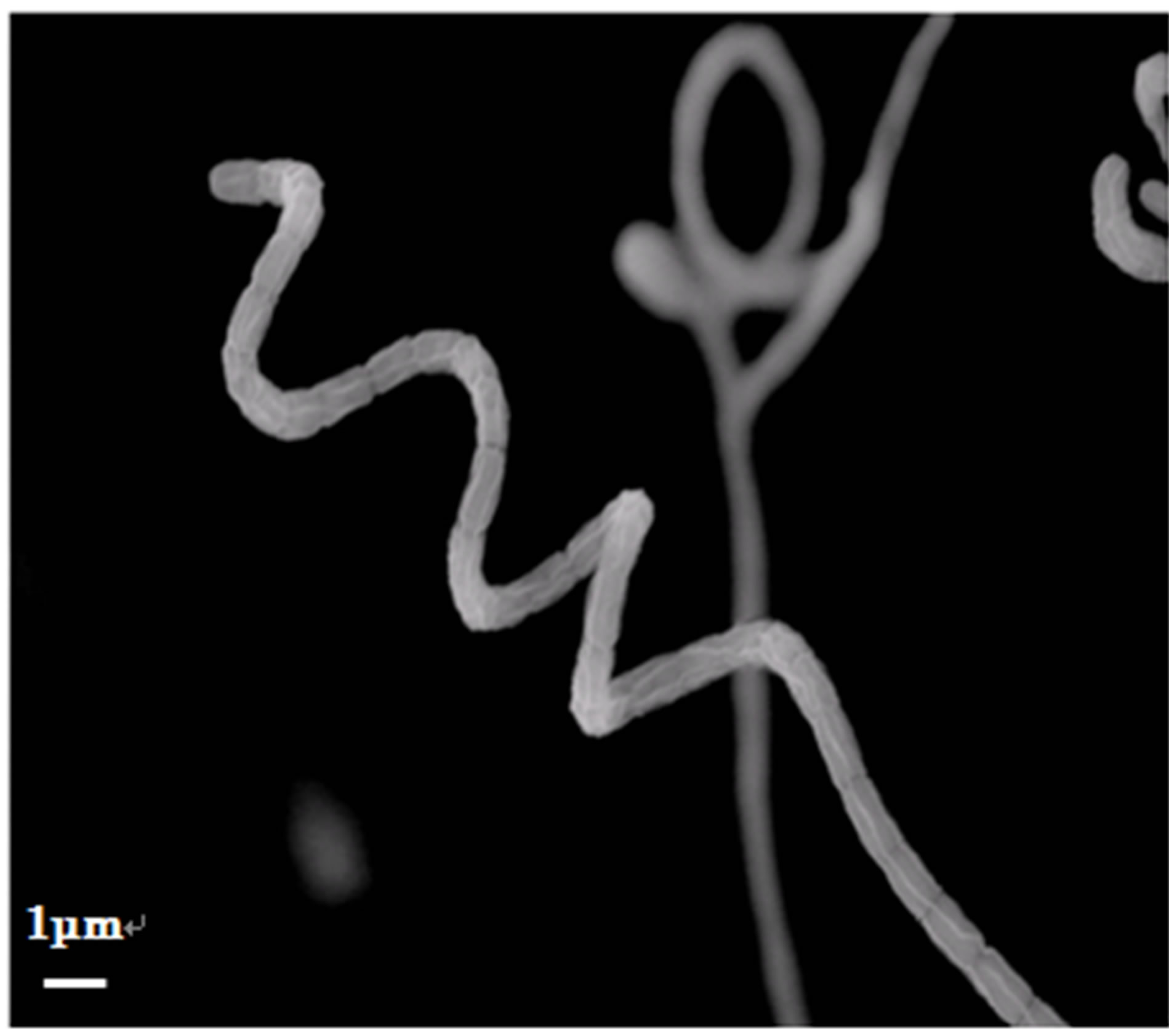 (b)
(b)
Figure 1. Scanning electron micrograph of strains TOHOY209 (a) and TOHO-O348 (b).
From the 16S rDNA gene sequence analysis, TOHOY209 was found to be closely related to Streptomyces phaeofaciens [19] (100.0%), and TOHO-O348 was found to be closely related to S. aburaviensis [20] (99.7%). Based on the above results, TOHO-Y209 (AB849954) and TOHO-O348 (AB849955) strains were confirmed to belong to genus Streptomyces.
3.2. Fermentation and Isolation
Based on the inhibitory activity in the screening, two strains, Streptomyces sp. TOHO-Y209 and TOHO-O348, were selected to identify the metabolites with anti-QS activity.
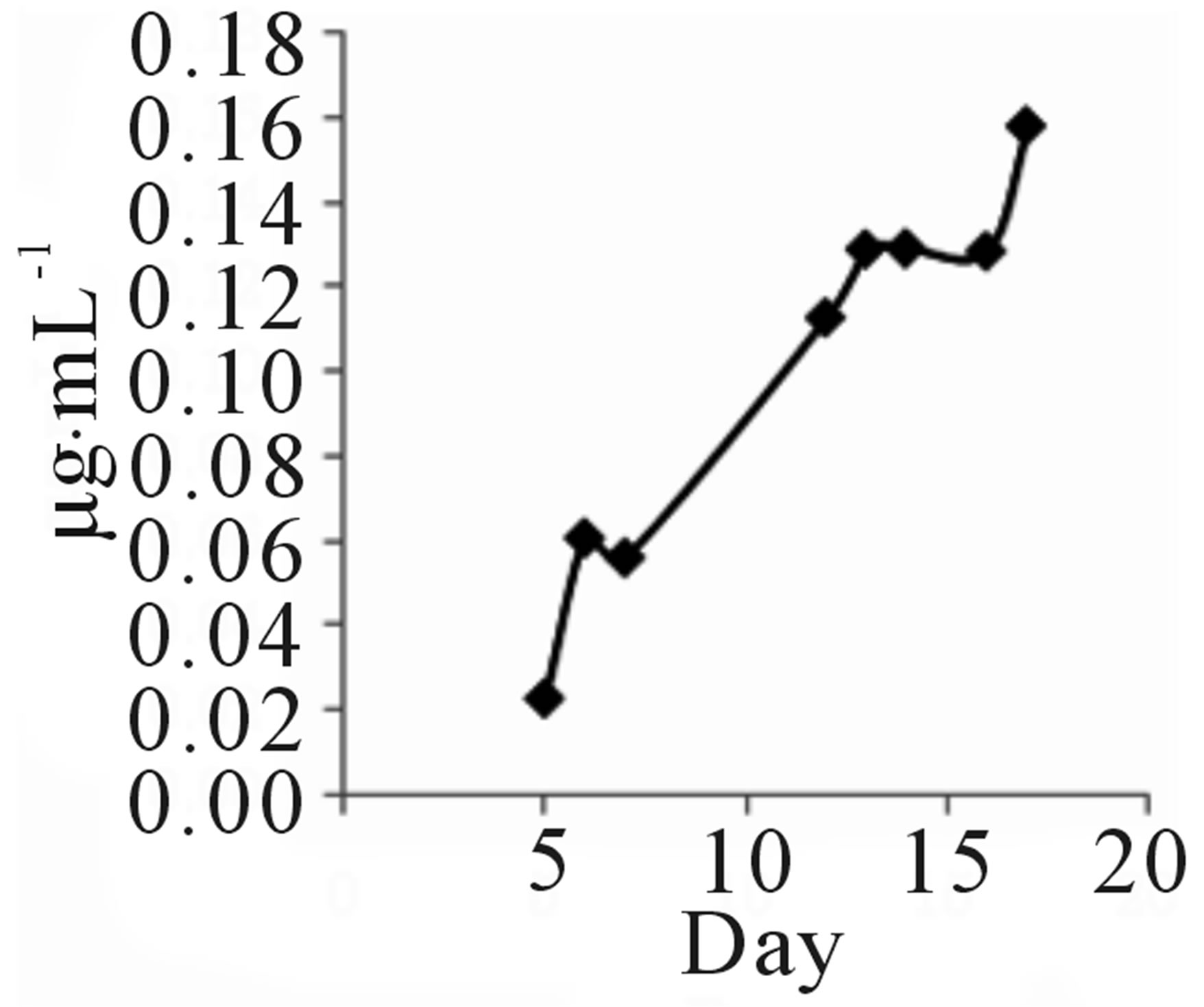
Strain TOHO-Y209 was cultivated in a yeast malt (YM) medium (10 L) at 27˚C for 14 days. The ethyl acetate extract from the culture broth was concentrated in vacuo. The residue was dissolved in a small amount of chloroform (CHCl3) and subjected to silica gel column chromatography (20 × 107 mm). The column was eluted successively with 100 mL each of CHCl3, CHCl3-1% methanol (MeOH), CHCl3-2% MeOH, and CH3OH. The QSIs were found in the CHCl3-1% MeOH eluate. This eluate was concentrated, and the brown residue (203.2 mg) obtained was subjected to octadecylsilyl (ODS) column (10 × 175 mm) chromatography and eluted successively with 30%, 60%, and 100% CH3CN. The active fractions (40 mL each) were concentrated and were subjected to high-performance liquid chromatography (HPLC) (Shim-pack PREP-ODS column, Shimadzu Corporation, 10 × 250 mm; mobile phase, 75% acetonitrile (CH3CN) containing 0.06% TFA; flow rate, 8.0 mL/min; detection, UV at 210 nm). QSI activity was found in the peaks at 7 (Compound 209-1) and 13 (Compound 209-2) min. Finally, 4.3 mg of 209-1 and 7.2 mg of 209-2 were obtained. The incubation period was decided based on the findings that 209-1 was produced after incubation for 5 days and reached a plateau after 16 days of incubation, while 209-2 production reached a plateau after 5 days (Figure 2).
TOHO-Y209 was incubated in a YM medium, and 5 mL of culture broth was evacuated at the time designated in the figure and extracted with 5 mL of ethyl acetate. The extract (100 µL) was evaporated and subjected to
 (a)
(a)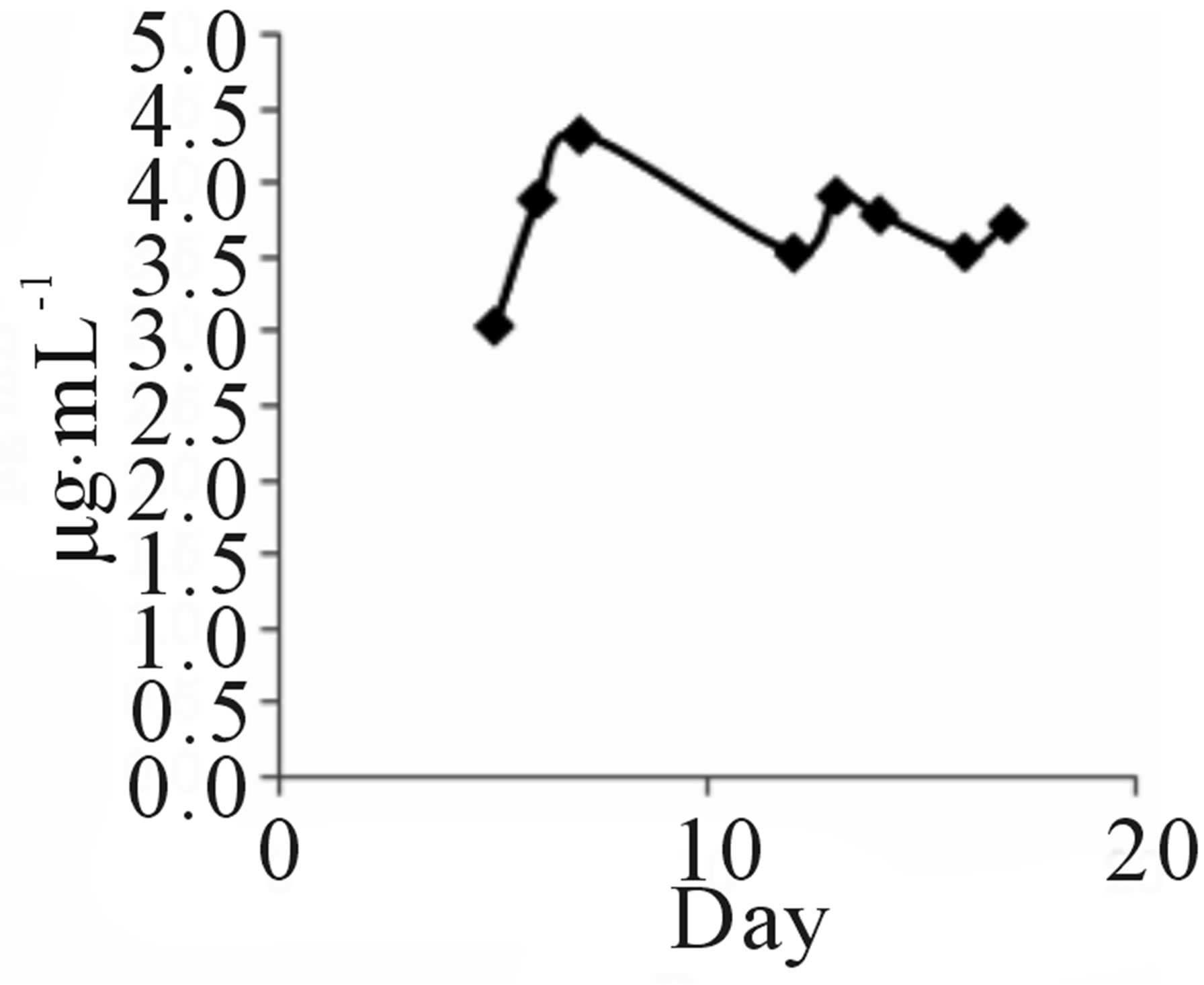 (b)
(b)
Figure 2. Time course of 209-1 (a) and 209-2 (b) productions.
HPLC to determine the amounts of 209-1 and 209-2.
Strain TOHO-Y348 was cultivated in a YM medium (10 L) at 27˚C for 7 days because the production of both QSIs reached to plateau after 5 days of incubation. The QSI produced by strain TOHO-O348 was extracted with ethyl acetate from 10 L of the culture broth and concentrated. The residue was dissolved in a small amount of CHCl3 and subjected to silica gel column chromatogramphy (25 × 300 mm) and eluted successively with 300 mL each of CHCl3, CHCl3-3% MeOH, CHCl3-5% MeOH, and CH3OH. The active fractions were combined, concentrated, and subjected to HPLC (Shim-pack PREPODS column, Shimadzu Corporation, 10 × 250 mm; mobile phase, 80% CH3CN; flow rate, 8.0 mL/min; UV detection at 210 nm). The QSI activity was observed in the peaks at 7 min (Compound 348-1) and 9 min (Compound 348-2). Finally, 8 mg of 348-1 and 23 mg of 348-2 were obtained.
3.3. Structure Elucidation
The structures of compounds 209-2, 348-1, and 348-2 were established on the basis of their UV, 1H-NMR, and 13C-NMR spectra. Compounds 209-2 and 348-2 were found to possess the same structure and were characterized as piericidin A1 (Figure 3) [21,22]. Compound 348- 1 was found to be the derivative of piericidin A1and was characterized as 3’-rhamnopiericidin A1 (Figure 4) [23].
The UV, IR, 1H-NMR, and 13C-NMR spectra of 209-1 resembled those of piericidin A1; however, the chemical shifts of certain C and H atoms were different from those for209-2, 348-1, and 348-2, as shown in Table 2. The molecular formula of 209-1 was determined to be C25H37NO5 by high-resolution mass spectroscopy (HRMS). By analyzing the1H-NMR, 13C-NMR, and various two-dimensional (2D)-NMR spectra of 209-1, 35 proton
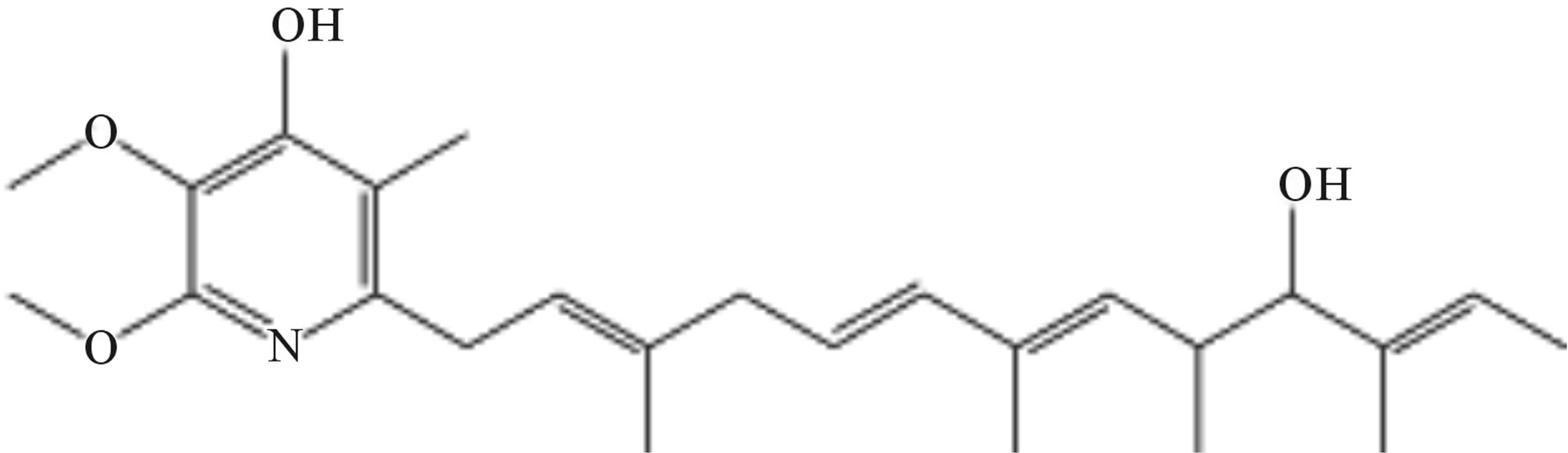
Figure 3. Structure of compounds 209-2 and 348-2.
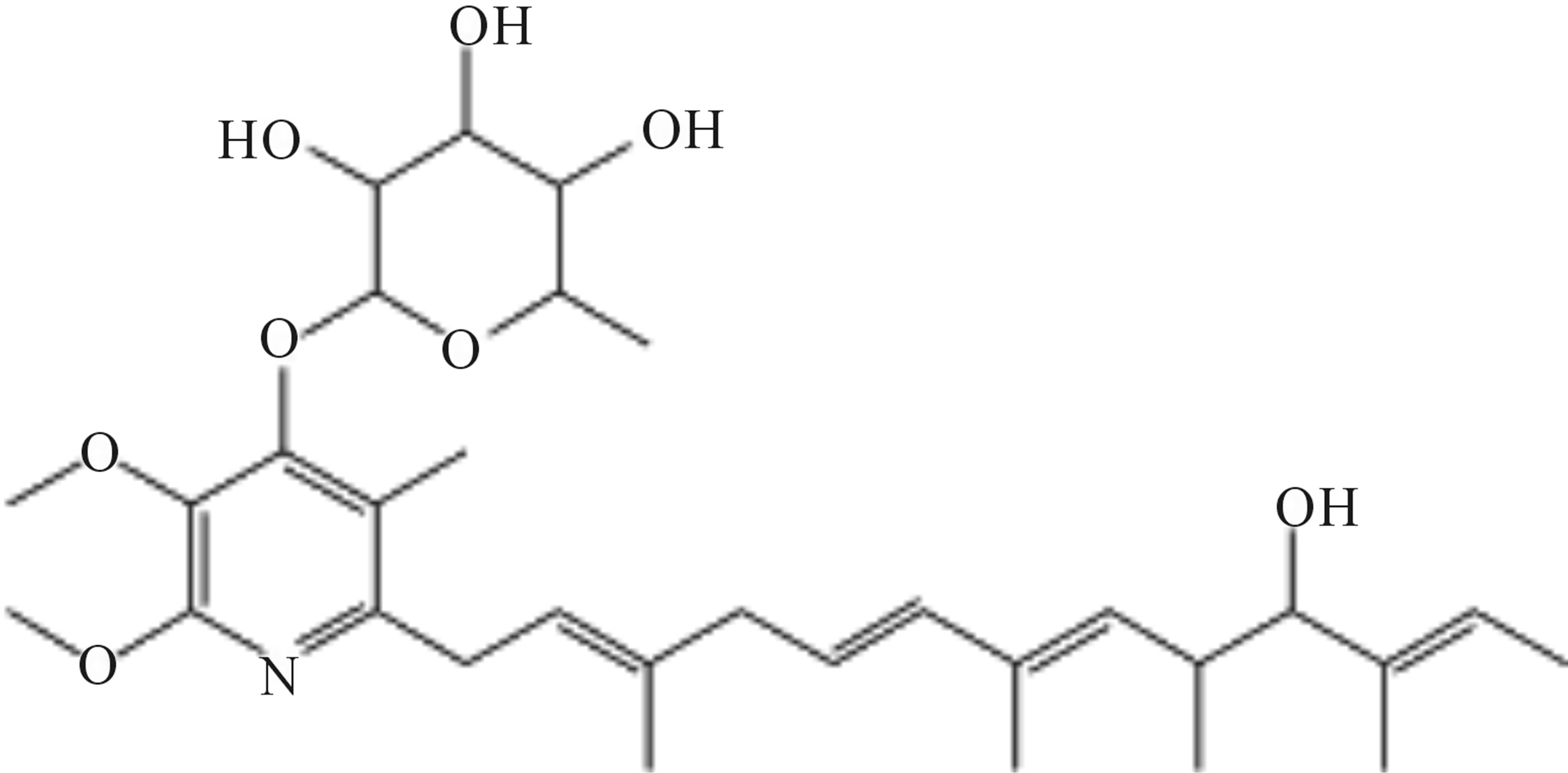
Figure 4. Structure of compound 348-1.

Table 2. Physicochemical properties of 209-1.
and 25 carbon signals were confirmed, and by analyzing the heteronuclear multiple-quantum correlation (HMQC) spectroscopy, C-H correlations were determined, as shown in Table 3.
1H-1H COSY spectral analysis of 209-1 led to the identification of partial structural units (I, II, III, and IV), as shown in Figure 5. Next, the connection between these units and the remaining functional groups was determined. The1H-13Cheteronuclear multiple-bond correlation (HMBC) correlations from 3-CH3 to C-2, C-3, and C-4; and from 4-H to C-2, C-3, and 3-CH3 established the connection between partial structures I and II. The 1H-13CHMBC correlations from 6-H to C-7 and C-8; from 7-CH3 to C-6, C-7, and C-8; and from 8-H to C-6 established the connection between partial structures II and III. The 1H-13CHMBC correlations from 10-H to 11-CH3; from 11-CH3 to C-10, C-11, and C-12; and from 12-H to C-10 established the connection between partial structures III and IV. Further, HMBC correlations from 1-H to C-2’ and C-3’; from 3’-CH3 to C-2’, C-3’, and C-4’; from 5’-OCH3 to C-5’; and from 6’-OCH3 to C-6’ were observed. The 1H-13CHMBC correlations are summarized in Figure 6.
Furthermore, the existence of eleven sp2 carbons suggested the existence of one double bond between the heteroatom and carbon atom, and five carbon-carbon double bonds. Because the index of hydrogen deficiency calculated from molecular formula C25H37NO5 was eight, the partial structure analysis described above indicated the existence of one carbon-carbon double bond and one heteronuclear double bond. The 13C-NMR spectrum confirmed the presence of a pyridine ring. The chemical shifts of C-7 and C-10 suggested that they are connected via an ether linkage, indicating the presence of an oxacyclopentane ring. Further, the butylene substituent at C-10 and hydroxyl groups at C-8 and C-4’ were characterized. The planner structure of 209-1 was supported by MS fragment analysis (Figure 7).
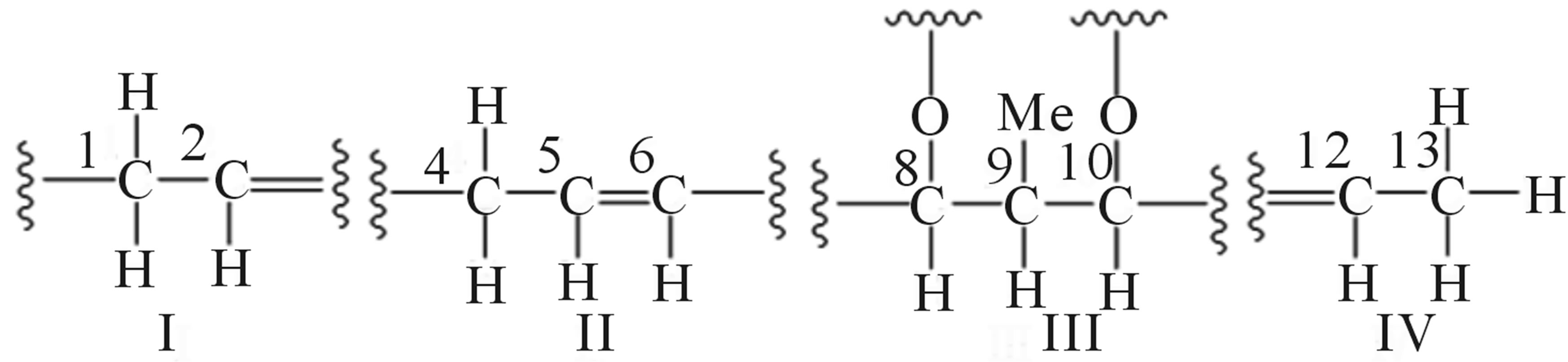
Figure 5. Partial structural units of 209-1.
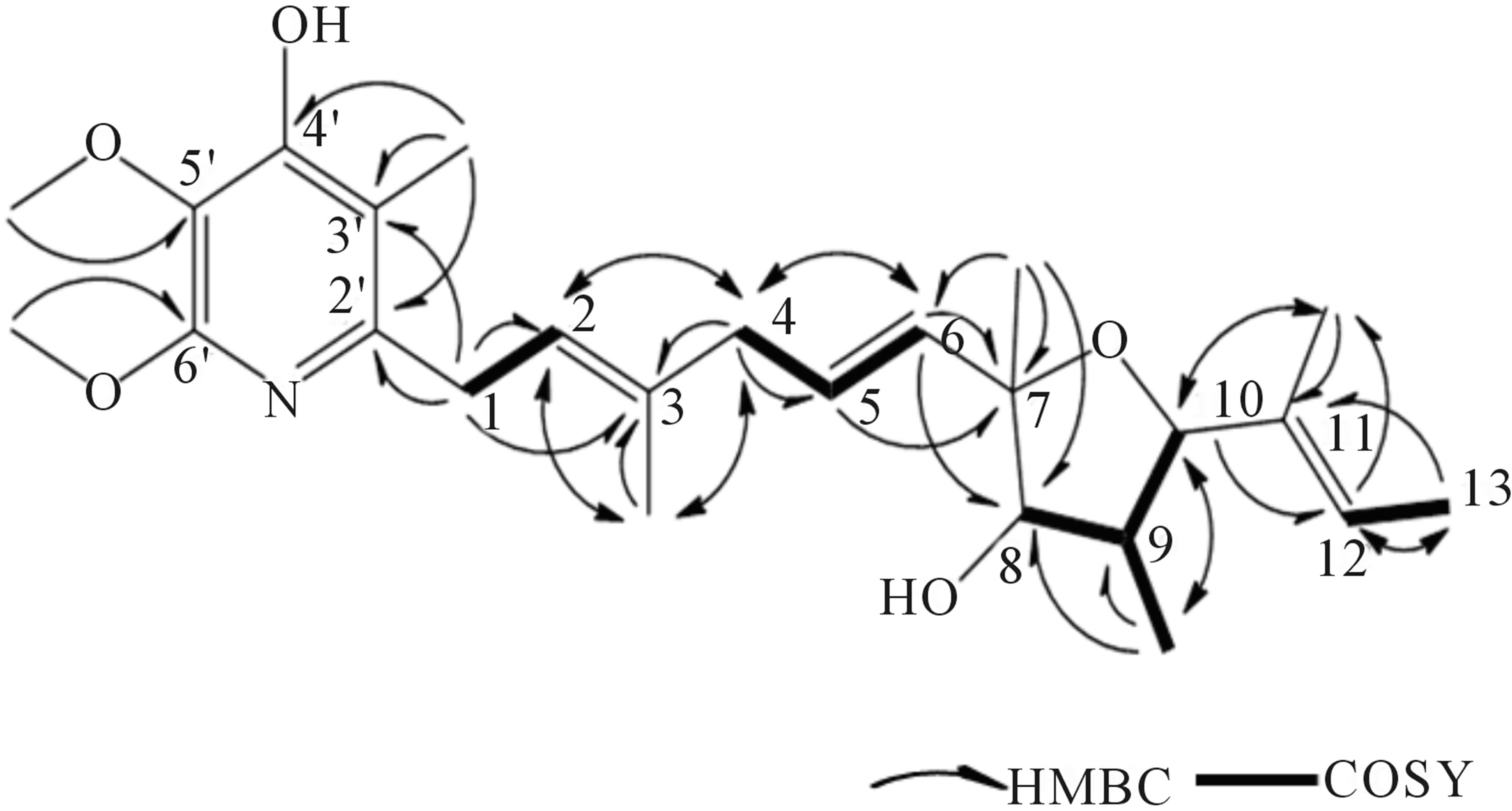
Figure 6. HMBC correlations of 209-1.
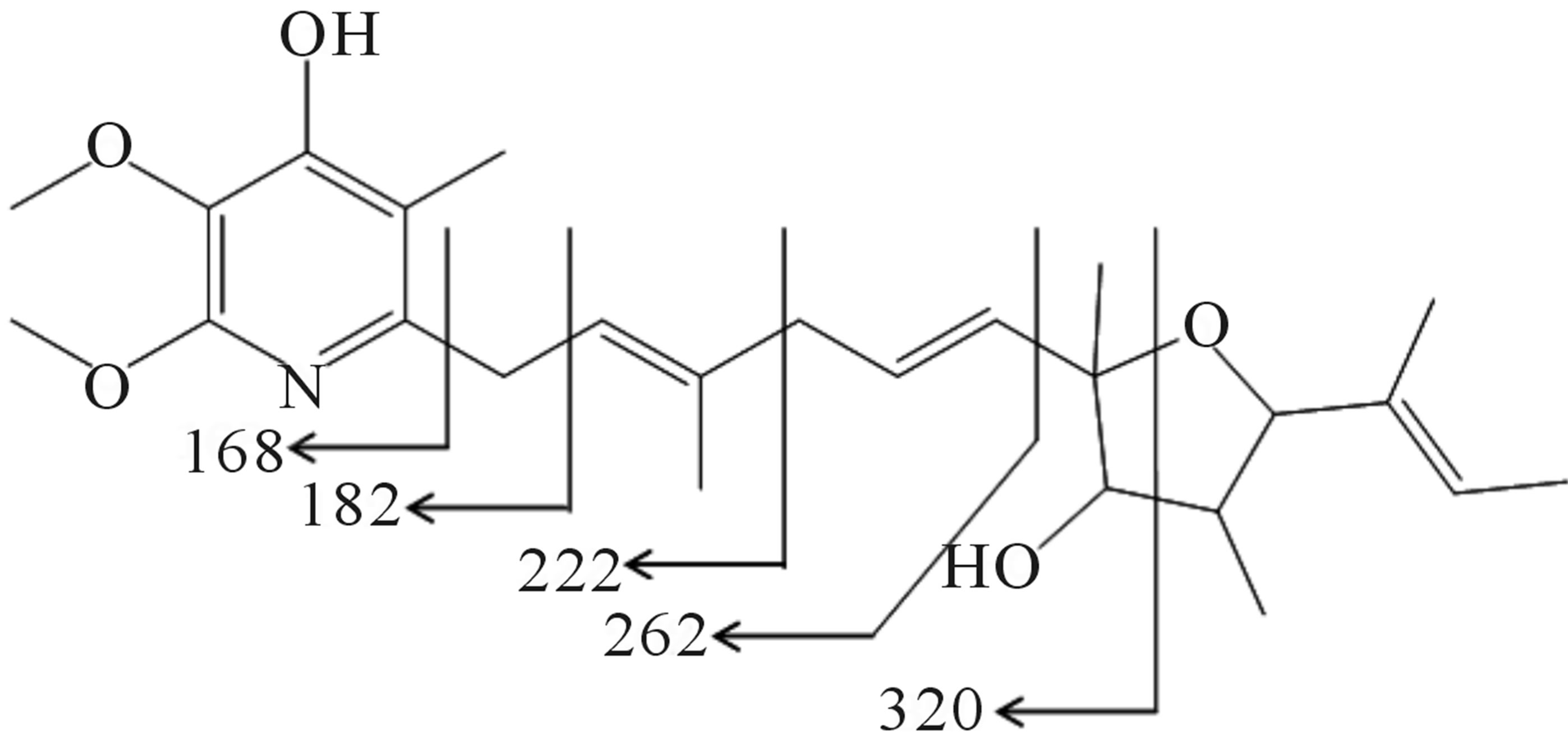
Figure 7. MS fragment analysis of compound 209-1.
The stereochemistry of 209-1 was deduced from nuclear overhauser effect (NOE) and nuclear overhauser effect spectroscopy (NOESY) experiments. The NOE correlations from 4-H to 2-H, from 4-H to 6-H, and from 10-H to 12-H indicated that the double bonds at positions 3, 6, and 11 exist in trans-conformations, and the relative stereochemistry at the oxacyclopentane ring were determined as shown in Figure 8. Thus, the structure of 209-1 was determined and designated as piericidin E.
3.4. Effect of Piericidin Derivatives on C. violaceum CV026
We investigated the QS inhibitory effects of piericidin A1, 3’-rhamnopiericidin A1, and piericidin E against C. violaceum CV026 (Figure 9). The three piericidins inhibited the purple pigment (violacein) synthesis controlled by QS, and the inhibitory effect was found to be dose dependent in the range 1to 100 µg∙mL-1. Piericidin A1 showed the strongest inhibitory activity, and the half maximal inhibitory concentration (IC50) was 10 µg∙mL-1. The antimicrobial activities (MICs) of the metabolites against C. violaceum CV026, Micrococcus luteus ATCC- 9341, Staphylococcus aureus ATCC25923, and P. aeruginosa ATCC27853 were more than 100 µg∙mL-1.
4. Discussion
In this study, certain metabolites isolated from Actino-
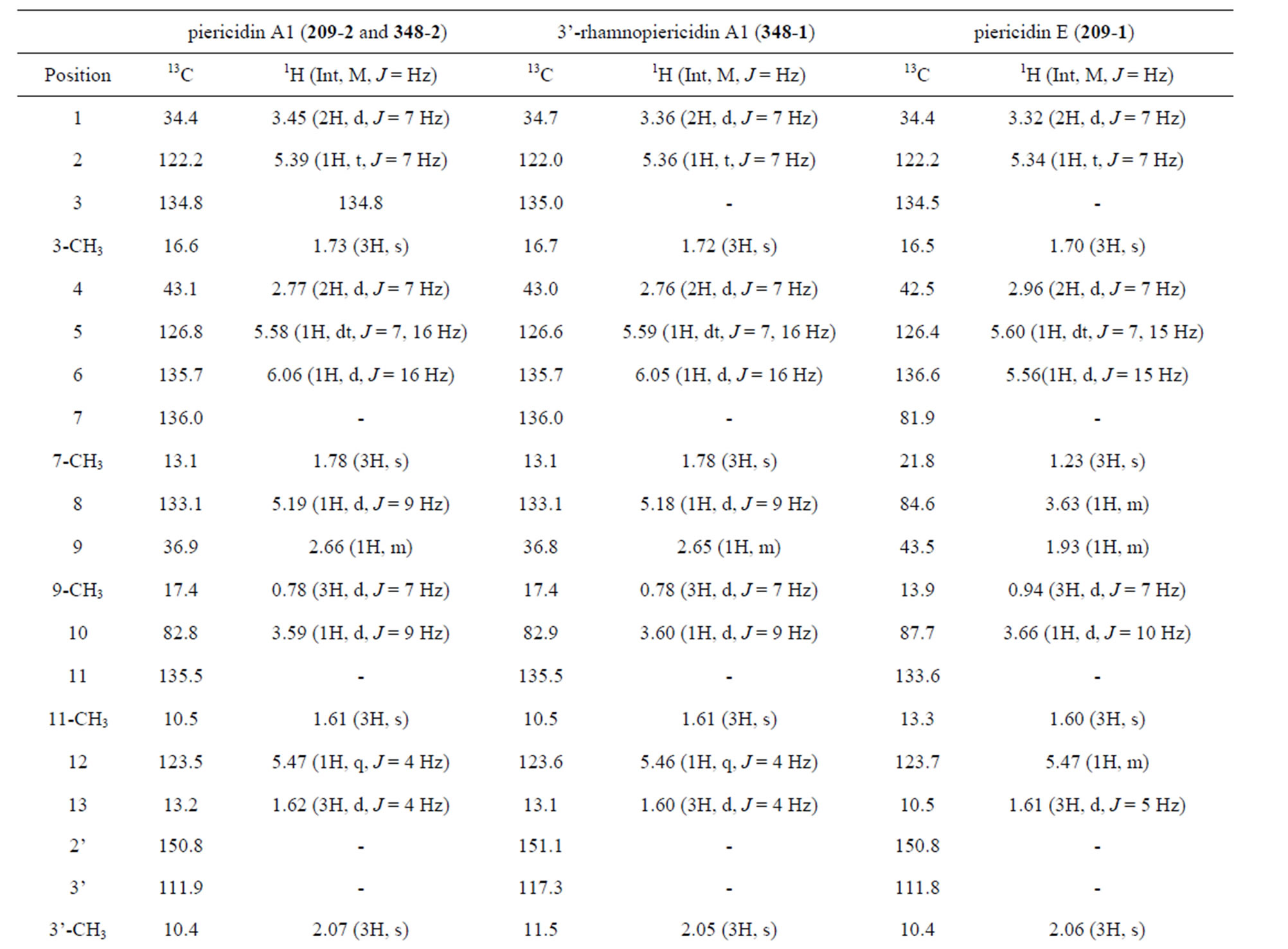
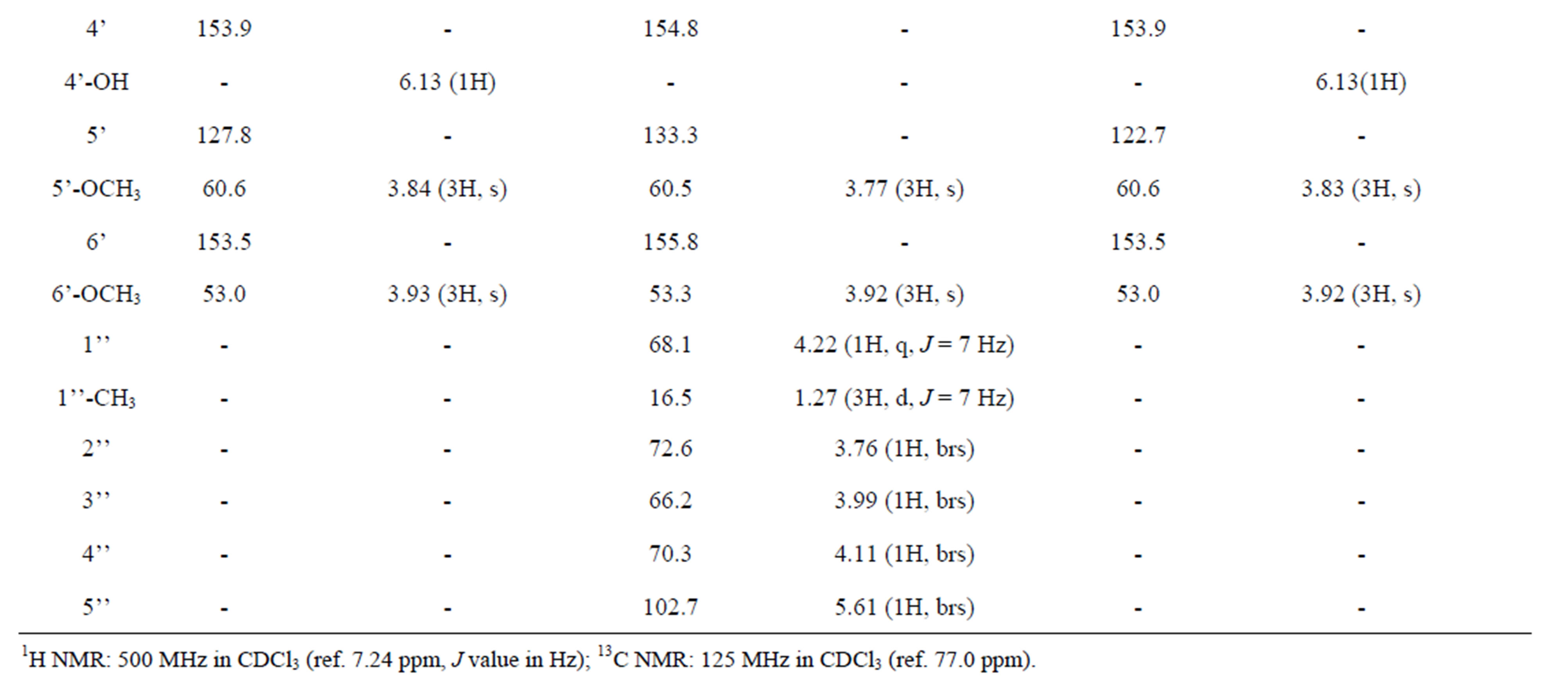
Table 3. 1H and 13C NMR chemical shifts of compounds.
myces culture broths demonstrated QS inhibiting activity against C. violaceum CV026. The major metabolite isolated from Streptomyces sp. TOHO-Y209 and TOHOO348 was piericidin A1. Compound 3’-rhamnopiericidin Awas isolated from TOHO-O348 as a minor component. Moreover, a novel piericidin derivative, designated as
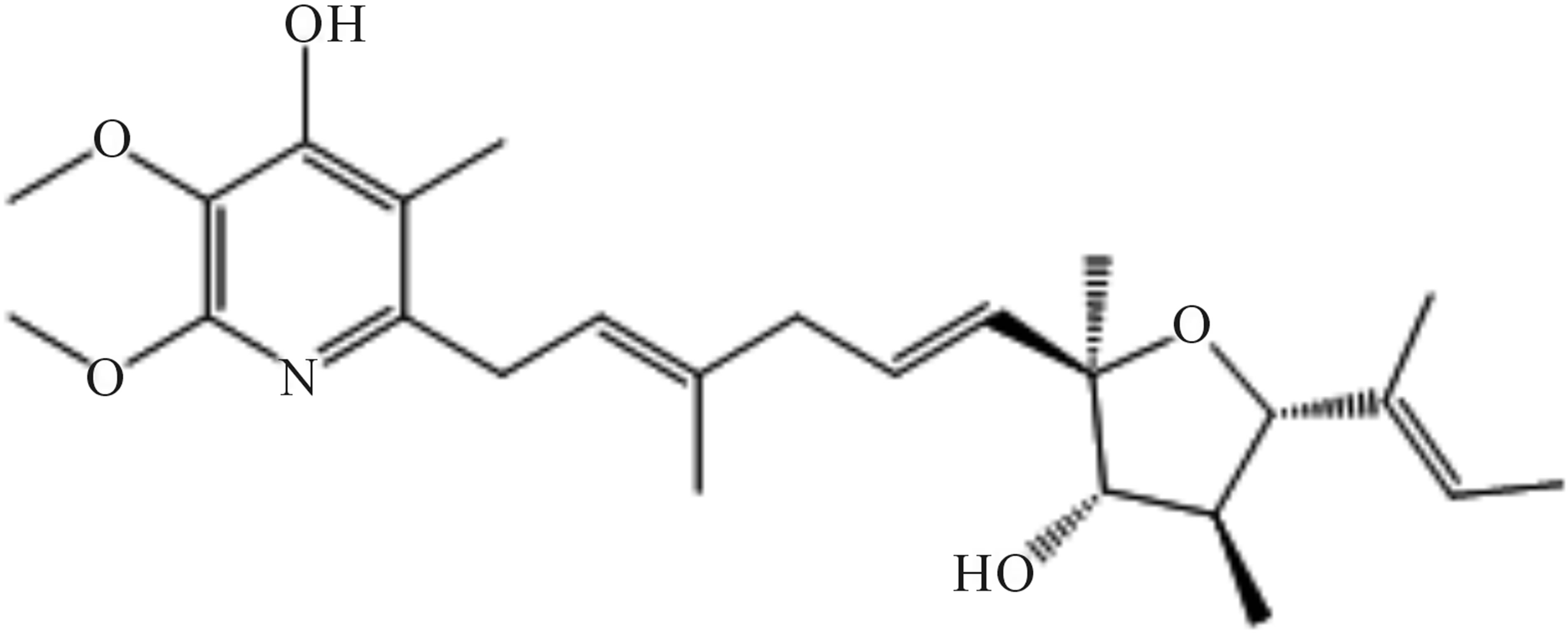
Figure 8. Structure of piericidin E (209-1).
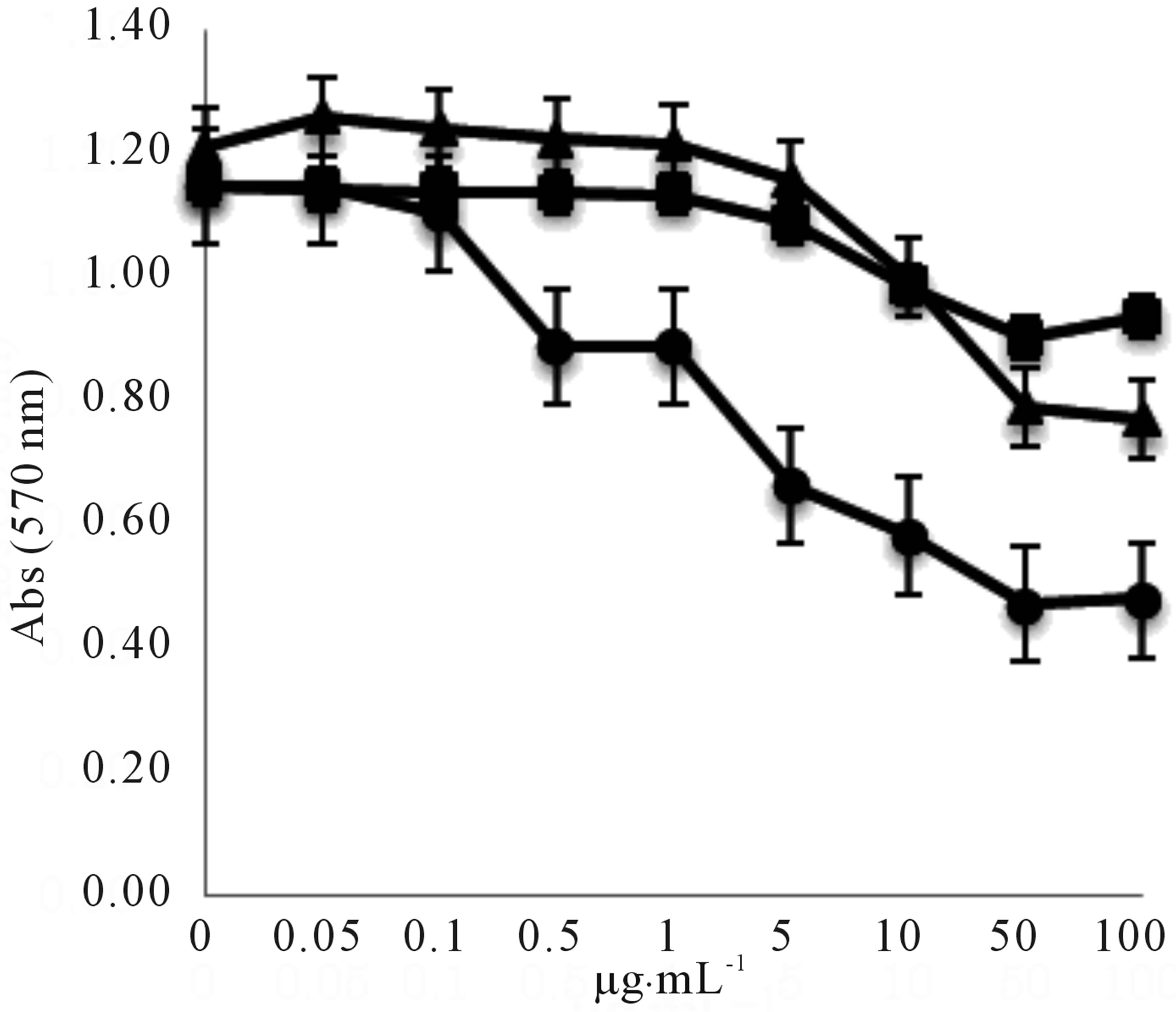
Figure 9. Effect of serial dilutions of piericidin A1 (○), 3’- rhamno piericidin A1(□), and piericidin E (∆) on violacein production in C. violaceum CV026. Violacein was extracted as described in the experimental section and quantified by measuring the optical density at a wavelength of 570 nm (OD570).
piericidin E, was isolated from TOHO-Y209.
Piericidin and its derivatives are known as potent inhibitors of NADH-ubiquinone oxidoreductase, and piericidin A1 inhibits both mitochondrial and bacterial NADHubiquinone oxidoreductases [21]. The antimicrobial activities (MICs) of piericidin A1 and 3’-rhamnopiericidin A1 previously reported on certain strains (Staphylococcus aureus FDA 209P, Bacillus subtilis ATCC 6633, Escherichia coli NIHJ, Klebsiella pneumoniae ATCC 29655, P. aeruginosa IFO 13725, and Aspergillus oryzae IFO 4221) [22] were more than 100 µg∙mL-1. Piericidin A1 and its derivatives isolated in this paper showed similar antibacterial activity as mentioned above; therefore, the QS inhibitory activity of the metabolites rather than the antibacterial activity resulted in the inhibition of violacein synthesis of C. violaceum CV026.
Until now, the QS inhibitory activity of piericidin derivatives have not been reported. The inhibitory effect was found to be dose dependent. Piericidin A1 showed the strongest QS inhibitory activity among the above mentioned compounds. The QS inhibitory activity of piericidin E was almost the same as that of 3’-rhamnopiericidin A1.
Piericidin and its derivatives were produced by Streptomyces sp. [24-27]. Although S. aburaviensis and S. phaeofaciens are not reported to produce piericidin A1, 3’-rhamnopiericidin A1, or piericidin E, Streptomyces sp. TOHO-Y209 was considered to be closely related to S. phaeofaciens and Streptomyces sp. TOHO-O348 was considered to be closely related to S. aburaviensis based on taxonomical studies.
In this study, piericidin A1 and its derivatives including a novel metabolite, piericidin E produced by Streptomyces sp. were found to be new QSIs. In this screening project, we found that 103 strains out of 1000 strains isolated from soil showed anti-QS activity in their culture broths; therefore, the strains of Actinomycetes are still a good source for screening the QSIs. The QSIs can be evaluated as drugs for antimicrobial chemotherapy.
5. Acknowledgements
The authors would like to thank Dr. Tsukasa Ikeda Utsunomiya University for kindly providing C. violaceum CV026, which was used for screening and quantitative assay.
REFERENCES
- K. H. Nealson, “Autoinduction of Bacterial Luciferase. Occurrence, Mechanism and Significance,” Archives of Microbiology, Vol. 112, No. 1, 1977, pp. 73-79. http://dx.doi.org/10.1007/BF00446657
- B. L. Bassler and R. Losick, “Bacterially Speaking,” Cell, Vol. 125, No. 2, 2006, pp. 237-246. http://dx.doi.org/10.1016/j.cell.2006.04.001
- R. J. C. McLean, M. Whiteley, D. J. Stickler and W. C. Fuqua, “Evidence of Autoinducer Activity in Naturally Occurring Biofilms,” FEMS Microbiology Letters, Vol. 154, No. 2, 1997, pp. 259-263. http://dx.doi.org/10.1111/j.1574-6968.1997.tb12653.x
- L. Eberl, M. K. Winson, C. Sternberg, G. S. Stewart, G. Christiansen, S. R. Chhabra, B. Bycroft, P. Williams, S. Molin and M. Givskov, “Involvement of N-acyl-L-homoserine Lactone Autoinducers in Controlling the Multicellular Behaviour of Serratialiquefaciens,” Molecular Microbiology, Vol. 20, No. 1, 1996, pp. 127-136. http://dx.doi.org/10.1111/j.1365-2958.1996.tb02495.x
- H. Slater, M. Crow, L. Everson and G. P. Salmond, “Phosphate Availability Regulates Biosynthesis of Two Antibiotics, Prodigiosin and Carbapenem, in Serratia via both Quorum-Sensing-Dependent and -Independent Pathways,” Molecular Microbiology, Vol. 47, No. 2, 2003, pp. 303-320. http://dx.doi.org/10.1046/j.1365-2958.2003.03295.x
- P. C. Fineran, H. Slater, L. Everson, K. Hughes and G. P. Salmond, “Biosynthesis of Tripyrrole and Beta-Lactam Secondary Metabolites in Serratia: Integration of Quorum Sensing with Multiple New Regulatory Components in the Control of Prodigiosin and Carbapenem Antibiotic Production,” Molecular Microbiology, Vol. 56, No. 6, 2005, pp. 1495-1517. http://dx.doi.org/10.1111/j.1365-2958.2005.04660.x
- J. P. Falcao, F. Sharp and S. V. perandio, “Cell-to-Cell Signaling in Intestinal Pathogens,” Current Issues in Intestinal Microbiology, Vol. 5, No. 1, 2004, pp. 9-17.
- L. C. Antunes, R. B. Ferreira, M. M. Buckner and B. B. Finlay, “Quorum Sensing in Bacterial Virulence,” Microbiology, Vol. 156, No. 8, 2010, pp. 2271-2282. http://dx.doi.org/10.1099/mic.0.038794-0
- A. Baldwin, P. A. Sokol, J. Parkhill and E. Maherthiralingam, “The Burkholderiacepacia Epidemic Strain Marker Is Part of a Novel Genomic Island Encoding both Virulence and Metabolism-Associated Genes in Burkholderiacenocepacia,” Infection and Immunity, Vol. 72, No. 3, 2004, pp. 1537-1547. http://dx.doi.org/10.1128/IAI.72.3.1537-1547.2004
- E. Valade, F. M Thibault, Y. P. Gauthier, M. Palencia, Y. Popoff and D. R. Vidal, “The PmlI-PmlR Quorum-Sensing System in Burkholderiapseudomallei Plays a Key Role in Virulence and Modulates Production of the MprA Protease,” Journal of Bacteriology, Vol. 186, No. 8, 2004, pp. 2288-2294. http://dx.doi.org/10.1128/JB.186.8.2288-2294.2004
- N. Towako, M. Hiroshi, K. Junichi, H. Tomayoshi, F. Takeshi, K. Misuzu, S. Ryo, Y. Katsunori, T. Kazunori, K. Takehiko and K. Shigeru, “Effect of Erythromycin on Chronic Respiratory Infection Caused by Pseudomonas aeruginosa with Biofilm Formation in an Experimental Murine Model,” Antimicrobial Agents and Chemotherapy, Vol. 48, No. 6, 2004, pp. 2251-2259. http://dx.doi.org/10.1128/AAC.48.6.2251-2259.2004
- D. Martinelli, G. Grossmann, U. Séquin, H. Brandl and R. Bachofen, “Effects of Natural and Chemically Synthesized Furanones on Quorum Sensing in Chromobacteriumviolaceum,” BMC Microbiology, Vol. 4, 2004, p. 25. http://dx.doi.org/10.1186/1471-2180-4-25
- K. H. McClean, M. K. Winson, L. Fish, A. Taylor, S. R. Chhabra, M. Camara, M. Daykin, J. H. Lamb, et al., “Quorum Sensing and Chromobacteriumviolaceum: Exploitation of Violacein Production and Inhibition for the Detection of N-Acylhomoserine Lactones,” Microbiology, Vol. 143, No. 12, 1997, pp. 3703-3711. http://dx.doi.org/10.1099/00221287-143-12-3703
- E. B. Shiring and D. Gottlieb, “Methods for Characterization of Streptomyces Species,” International Journal of Systematic Bacteriology, Vol. 16, No. 3, 1966, pp. 313- 340. http://dx.doi.org/10.1099/00207713-16-3-313
- S. A. Waksman, “Classification, Identification and Description of Genera and Specie,” Williams and Wilkins Co., Baltimore, 1961.
- “Color Harmony Manual,” 4th Edition, Container Corporation of America, Chicago, 1958.
- T. G. Pridham and D. Gottlieb, “The Utilization of Carbon Compounds by Some Actinomycetales as an Aid for Species Determination,” Journal of Bacteriology, Vol. 56, No. 1, 1948, pp. 107-114.
- B. Becker, M. P. Lechevalier and H. A. Lechevalier, “Chemical Composition of Cell-Wall Preparation from Strains of Various Form-Genera of Aerobic Actinomycetes,” Applied Microbiology and Biotechnology, Vol. 13, 1965, pp. 236-243.
- M. Okamoto, K. Yoshida, M. Nishikawa, T. Ando, M. Iwami, M. Kohsaka and H. Aoki, “FR-900452, a Specific Antagonist of Platelet Activating Factor (PAF) Produced by Streptomyces phaeofaciens. I. Taxonomy, Fermentation, Isolation, and Physic-Chemical and Biological Characteristics,” The Journal of Antibiotics, Vol. 39, No. 2, 1986, pp. 198-204. doi: 0.7164/antibiotics.39.198
- M. Iwami, S. Kiyoto, H. Terano, M. Kohsaka, H. Aoki and H. Imanaka, “A New Antitumore Antibiotic, FR- 900482. I. Taxonomic Studies on the Producing Strain: A New Species of the Genus Streptomyces,” The Journal of Antibiotics, Vol. 40, No. 5, 1987, pp. 589-593.
- M. Jeng, C. Hall, F. L. Crane, N. Takahashi, S. Tamura and K. Folkers, “Inhibition of Mitochondrial Electron Transport by Piericidin A and Related Compounds,” Biochemistry, Vol. 7, No. 4, 1968, pp. 1311-1322. http://dx.doi.org/10.1021/bi00844a010
- A. Urakawa, T. Sasaki, K. Yoshida, T. Otani, Y. Lei and W. Yun, “IT-143-A and B, Novel Piericidin-Group Antibiotics Produced by Streptomyces sp.,” The Journal of Antibiotics, Vol. 49, No. 10, 1996, pp. 1052-1055. http://dx.doi.org/10.7164/antibiotics.49.1052
- K. Kimura, S. Nakayama, N. Nakajima, M. Yoshihama, N. Miyata and G. Kawanishi, “A New Piericidinrhamnoside, 3’-Rhamnopiericidin A1,” The Journal of Antibiotics, Vol. 43, No. 10, 1990, pp. 1341-1343. http://dx.doi.org/10.7164/antibiotics.43.1341
- K. A. Shaaban, E. Helmke, G. Kelter, H. H. Fiebig and H. Laatsch, “Glicopiericidin C: A Cytotoxic Piericidinglucoside Antibiotic Produced by a Marine Strepromyces Isolate,” The Journal of Antibiotics, Vol. 64, No. 2, 2011, pp. 205-209. http://dx.doi.org/10.1038%2Fja.2010.125
- K. Kimura, H. Takahashi, N. Miyata, M. Yoshihama and M. Uramoto, “New Piericidin Antibiotics, 7-Demethylpiericidin A1,” The Journal of Antibiotics, Vol. 49, No. 7, 1996, pp. 697-699. http://dx.doi.org/10.7164/antibiotics.49.697
- Y. Hayakawa, S. Shirasaki, S. Shiba, T. Kawasaki, Y. Matsuo, K. Adachi and Y. Shizuri, “Piericidin C7 and C8, New Cytotoxic Antibiotics Produced by a Merine Streptomyces sp.,” The Journal of Antibiotics, Vol. 60, No. 3, 2007, pp. 196-200. http://dx.doi.org/10.1038/ja.2007.22
- J. Ueda, T. Togashi, S. Matukura, A. Nagai, T. Nakashima, H. Komaki, K. Anzai, S. Harayama, T. Doi, T. Takahashi, T. Natsume, Y. Kisu, N. Goshima, N. Nomura, M. Takagi and K. Shin-Ya, “A Novel Nuclear Export Inhibitor JBIR-02, a New Piericidin Discovered from Streptomyces sp. ML55,” The Journal of Antibiotics, Vol. 60, No. 7, 2007, pp. 459-462. http://dx.doi.org/10.1038/ja.2007.59
NOTES
*Corresponding authors.

