Open Journal of Medicinal Chemistry
Vol.2 No.2(2012), Article ID:19673,9 pages DOI:10.4236/ojmc.2012.22003
Relevant Enzymes, Genes and Regulation Mechanisms in Biosynthesis Pathway of Stilbenes
Department of Pharmacy, General Hospital of Guangzhou Military Command, Guangzhou, China
Email: *dolloway@163.com
Received February 22, 2012; revised April 23, 2012; accepted May 4, 2012
Keywords: Stilbenes; Relevant Enzyme; Regulation Mechanism; Biosynthesis Pathway; Systems Biology
ABSTRACT
Stilbenes are natural phenolic compounds which function as antimicrobial phytoalexins in plants and affect human health as cardioprotective, antibaceteria, antioxidative and antineoplastic agents. In this review, the progresses of study on relevant enzymes, genes, and regulation mechanism in biosynthesis pathway of stilbenes are described. Here we introduce a holistic and systematic method of researching relevant enzymes, genes and other regulatory factors in bio-synthesis pathway of stilbenes—Systems biology. The application of knowledge of relative enzymes, genes and regulation mechanisms in stilbenes biosynthesis in metabolic engineering which is used as a tool of improving the disease resistance of plants and health caring quality of crops is also discussed.
1. Introduction
Stilbenes, a kind of phytoalexins, are low-molecular-weight defensive substances produced by plants in response to infection after exposure to microorganisms. They are widely distributed in the plant kingdom, being reported from bryophytes and pteridophytes through gymnosperms and angiosperms. Stilbenes display a wide range of biological activitivies, such as antibacterial, antifungal, estrogenic, antitumoral [1], cardioprotective [2] and tyrosinase inhibitory activity [3]. There is great interest in their potential health benefits and capacity to improve the disease resistance of plants [4]. Much effort has been directed at the Stilbenes’ extraction, structure determination, biological activity over the past decades. In recent years, some success also has been achieved in the metabolic regulation and gene engineering of stilbenes. However, their detailed biosynthesis pathways and metabolic regulation, especially complicated regulation mechanism and expressing of genes and enzymes are unknown. So it is significant to shift focus from previous research priorities to search relevant enzymes, genes, abiotic stress and biotic signals so as to elucidate their detailed biosynthesis pathway and understand metabolic regulation networks. The elucidating of stilbenes’ biosynthesis pathway and regulation mechanism is believed to contribute to improve the disease resistance of plants and health caring quality of crops and also provide an opportunity to know more about global regulation networks and coordination between each pathway of secondary metabolism.
2. The Relevant Enzymes and Regulation in Biosynthesis Pathway of Stilbenes
2.1. The Relevant Enzymes and Regulation in Phenylpropanoid Pathway
The phenylpropanoid pathway is one of the most important plant secondary metabolism pathways and it is involved in the synthesis of a wide variety of important natural products from plants including flavonoids, lignins, coumarins, and stilbenes [5]. Phenylalanine ammonialyase (PAL), cinnamic acid 4-hydroxylase (C4H) and 4- coumarate: CoA ligase (4CL) are key enzymes in this pathway [6]. PAL, the first and key enzyme of the phenylpropanoid sequence, is the bridge between primary metabolism and secondary metabolism. PAL catalyzes the formation of transcinnamicacid by nonoxidative deamination of L-phenylalanine, which could be the rate-limiting step in the phenylalanine metabolism pathway. It produces precursors for a variety of secondary metabolites such as flavonoids, lignins, coumarins, and stilbenes. PAL genes are transcriptionally activated after microbial infection or treatment of plant cells with microbial elicittors [7]. The second step in the phenylpropanoid pathway is the hydroxylation of trans cinnamic acid to 4-coumaric acid, which is catalyzed by C4H, a cytochrome P450 monooxygenase [8,9]. C4H is induced by light, elicitors, and wounding [8,10-12]. Its induction often is closely coordinated with PAL induction [13]. The 4-coumaric acid is then activated to its CoA thioester by 4CL. 4- coumaroyl CoA is funneled into branched pathways leading to a wide array of phenolic metabolites, including lignin, flavonoids [5].
Phenylpropanoid biosynthesis comprises reactions through which metabolic channeling may occur. Metabolic channeling offers unique opportunities for enhancing and regulating cellular biochemistry and major advantage of such spatial organization is the transfer of biosynthetic intermediates between catalytic sites without diffusion into the bulk phase of the cell [14]. This phenomenon involves the physical organization of successive pathway enzymes into complexes through which metabolic intermediates are channeled [15].
Studies demonstrate phenylpropanoid pathway and flavonoid metabolism branch are assembled as a linear array of sequential enzymes loosely anchored to the cytoplasmic face of endoplasmic reticulum membranes [7,16-17]. For example, Cytochrome P450 enzymes, such as C4H, Flavanone-3-hydroxide transketolase, the ferulic acyl-5-hydroxylation enzyme are anchored to the external surface of the endoplasmic reticulum [18-20]. PAL and C4H activities are colocalized on membranes of the endoplasmic reticulum. This organization regulate the partitioning of intermediates among competing pathways and determine the intracellular deposition of end products. PAL, CHS, STS, isoflavonids synthase are structure specific enzymes, Flavanone-7-O-methyltransferase, isoflavones-4-O-methyltransferase and isoflavones (isoflavanone) dimethylallyltransferase are modification enzymes in the phenylalanine metabolism pathway.
2.2. The Relevant Enzymes and Regulation in Biosynthesis Pathway of Stilbenes
Stilbene phytoalexin is derived from phenylalanine via the general phenylpropanoid pathway [21]. The last step is catalysed by Stilbene synthase (STS) which is the key enzyme of the biosynthesis pathway. STS provides the first committed step by catalyzing the sequential decarboxylative addition of three acetate units from malonyl-CoA to a p-coumaroyl-CoA starter molecule derived from phenylalanine via the general phenylpropanoid pathway (Figure 1). For example, Resveratrol synthase (STS, EC 2.3.1.95) condenses three molecules of malonyl-CoA and one molecule of cumaryl-CoA to form resveratrol. In the same active site, chalcone synthase (CHS) can catalyse the formation of chalcone by p-coumaroyl-CoA and malonyl-CoA via the intramolecular cyclization and aromatization of the resulting linear phenylpropanoid tetraketide [22]. Downstream modification enzymes in branching pathways produce a number of biologically important compounds.
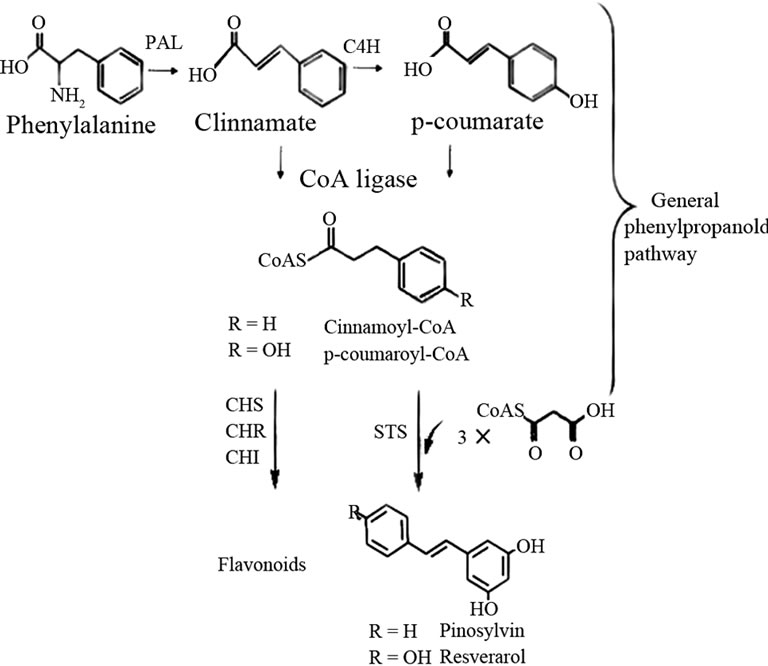
Figure 1. The biosynthesis pathway of stilbenes.
Different kinds of enzymes are involved in the biosynthesis pathway of stilbenes, which include highly speciesand substrateas well as stereospecific enzymes, modifycation enzymes and others act as regulators. Some well studied enzymes, PAL, 4CL, C4H, Pinosylvin methoxy transferase (PMT) in combination with STS are responsible for the regulation of biosynthesis of stilbenes. There are also other enzymes which are involved in the in the biosynthesis pathway and metabolic regulation of stilbenes are not definite. However, all the enzymes in the biosynthesis pathway of stilbenes can be classified according to their substrate specificity: stereochemical specific enzymes catalyse the formation of backbone of stilbenes and enzymes catalyse the modification reaction of the products which the first kind of enzymes synthesize. The first kind of enzymes include PAL, CHS, STS of phenylpropanoid pathway. The stilbene backbone is synthesized from cinnamoyl-CoA and malonyl-CoA by STS. The study on the capacity for building novel and unusual polyketides from alternative substrates of STS has shown that minor modifications can be used to direct the enzyme reaction to form a variety of different and new products [23]. The second kind of enzymes include hydroxylation enzyme, dehydrogenase, oxidase and glycosyltransferase. The specificity of these enzymes on its substrate group permutation is not high, this may be one of the main reasons why plant tissues can synthesized a series of relevant secondary metabolites [24].
The activity of key enzymes is not the only factor that affects the accumulation of stilbenes in the plants, all the enzymes involved in the formation and regulation of stilbenes as well as the supply of substrates are also associated with the accumulation of end product. The regulation mechanism of metabolic channeling also play an important role in the induction and accumulation of end product in the branch pathway of stilbenes. The activating of metabolic channeling and biosynthesis pathways of homologous synthesis is determined by the expression of the key enzymes, but the accumulation is most determined by the expression of rate-limiting enzymes in plants. Rate-limiting enzymes which usually can be found at the branch point or the downstream of biosynthesis pathways of secondary metabolites in plants are responsible for the synthesis of precursors of many secondary metabolites [25]. Interactions between transcription factors and coordinate expression of different enzymes in the metabolic channeling have synergistic effects on the accumulation of the stilbenes. Relevant enzymes in metabolic channeling can form multienzyme complexes and coordinated express in different parts of cell. Enhanced coordinate expression of the enzyme complex can lead to a dramatically accumulation of end products. For example, induced coordinate expression of PAL and STS in the biosynthesis pathways of stilbenes can affect the synthesis of the stilbenes [26].
The biosynthesis pathway of the stilbenes in plants is closed in general conditions, which is only activated in response to microbial infections and other environmental inducers. Environmental factors including biotic and abiotic stimuli, carbon-nutrition balance, genotype and ontogenesis usually control and regulate the biosynthesis of secondary metabolites in plants [27-29]. The enzymes involved in the biosynthesis pathway of stilbenes express after the expression of corresponding genes, and the expression level of the enzyme genes is under strict regulation in plant cells due to coordinate control of the biosynthetic genes by transcription factors [30]. Coordinate transcriptional control of biosynthetic genes emerges as a major mechanism dictating the final levels of secondary metabolites in plant cells [31]. This regulation of biosynthesis pathways is achieved by specific transcription factors encoded by genes unlinked to the biosynthetic gene clusters which regulate multiple physiological processes and generally respond to environmental cues such as pH, temperature, nutrition. Transcription factor activety itself is regulated by internal signals, for example plant hormones, or external signals such as microbial elicitors or UV light. Stress hormones, such as ethylene, jasmonic acid, and salicylic acid, induce STS mRNA accumulation in leaves of mature peanut plants. The expression of resveratrol synthase (RS) genes is induced by biotic and abiotic factors in peanut cell cultures [32]. Formation of pinosylvin (PS) and pinosylvin 3-O-monomethyl ether (PSM), as well as the activities of STS and Sadenosyl-l-methionine (SAM): pinosylvin O-methyltransferase (PMT), were induced strongly in needles of Scots pine seedlings upon ozone treatment, as well as in cell suspension cultures of Scots pine upon fungal elicitation [33]. A modeling method for the induction of resveratrol synthesis by UV irradiation pulses in Napoleon table grapes is proposed. Cantos etc. use the controlled UV irradiation pulses as a simple postharvest treatment to obtain possible “functional” grapes with enhanced health-promoting properties high resveratrol content [34].
3. The Application of Systems Biology in the Research on Relevant Enzymes and Genes in Biosynthesis Pathway of Stilbenes
Systems biology is a new science which makes us be able to understand biological systems grounded in the molecular level as a consistent framework of knowledge for the first time after the genomics, proteomics etc. were put forward [35]. It is such a rapidly evolving discipline endeavours to study the detailed coordinated workings of entire organisms with the ultimate goal to understand the dynamic networks of regulation and interactions that allows cells and organisms to live in a highly interactive environment [36]. The well studied molecular biology only care about individual gene and protein. However, systems biology is the study of cell signaling and gene regulatory networks and components and functions of the biological system, can also be understood as the study of all components in a biological system (genes, mRNA, protein etc.) and the interactions between these components in a certain circumstances [37]. Systems biology is a powerful tool to comprehensively explore the biological system, the application of it’s thinking model in secondary metabolites of medicinal plants brings us into a new era of understanding how to connect genes to metabolites by a systems biology approach [38].
3.1. Systems Biology Research Methods
The classical molecular biology research is to search for specific genes at the DNA level, and then to study gene functions by gene mutation, gene knockout and other means, it also can be described to study individual genes and proteins by using a variety of means. Genomics, proteomics, transcriptomics etc. are used as single means to research multiple genes or proteins at the same time. But using one of them alone provides only part of information of system without any details of interactions between components of system. systems biology integrate genomics, proteomics etc. and molecular biology in order to provide complementary datas. It is enabled by recent advances in multidisciplinary scientific disciplines and highthroughput approaches that allow for the parallel largescale measurement of biomolecules, such as mRNA, proteins and metabolites [39].
Functional genomics with the goal of characterization functions of genes has become an important method of systems biology. It provides comprehensive analysis of gene functions at the genome or system level, which shift focus from research of single gene or protein to multiple genes or proteins using high-throughput experimental methods in combination with mass datas of statistical calculation method. The technology of T-DNA insertion, transposon technology, classic subtraction hybridization, differential screening, cDNA difference analysis, mRNA differential display of gene, and serial analysis of gene expression (SAGE) of systematic analysis, cDNA microarrays and DNA chip are used to analyse information of genome sequence and elucidate the gene functions. Functional genomics is a powerful tool to reveal the biosynthesis pathways of secondary metabolites, and it will provide a solid theoretical basis for the production of secondary metabolites using metabolic engineering of medicinal plants, as well as the cell or tissue culture combined with metabolic engineering [40].
The methods of transcriptomics include differential display, gene chip, expressed sequence tags (EST) analysis, massively parallel signature sequencing (MPSS), amplified fragment length polymorphism (AFLP) [41,42]. Twodimensional electrophoresis, mass spectrometry technology bioinformatics analysis are primary methods of screening and identifying proteomics. Yeast two—hybrid system (Y2H), tandem affinity purification (TAP) can be used to study protein—protein interactions and green fluorescent protein (GFP) as maker to study subcellular localization. Metabonomics is a very important tool to study medicinal plants and promote modernization of traditional Chinese medicine [43,44], includeing nuclear magnetic resonance (NMR), gas chromatography-mass spectrometry (GC-MS), liquid chromatography-mass spectrometry (LC-MS), combined application of fourier transformation mass spectrometry (FTMS) and capillary electrophoresis-mass spectrometry (CE-MS) [45].
Technologies of Genomics, proteomics, transcriptomics and metabolomics detect the various molecules and study their functions at DNA, mRNA, protein and metabolite levels.
3.2. The Application of Systems Biology Approaches in Research on Related Enzymes and Genes in Biosynthesis Pathway of Stilbenes
When we study stilbenes with thinking model of systems biology, various levels of information including the DNA, mRNA, small molecules, proteins and protein interaction networks should to be integrated in order to obtain a series of relevent enzymes, genes or regulatory factors in it’s biosynthesis pathway. All the information can be used to construct a reasonable model in order to elucidate the biosynthesis pathway, regulation mechanisms of stilbenes [46]. For example, the study of relative enzymes and genes in biosynthetic pathway of tanshinone using thinking model of systems biology. Groups of materials with phenotypic differences are analysed in order to get datas of metabolomics, proteomics, transcriptomics which can be gained with gene chips. Systemic results about genes and enzymes revolved in the biosynthesis pathway of tanshinone were obtained [47-50]. As a result, an unique new branch of two terpene biosynthesis of tanshinone biosynthesis was found.
Our group focused the research on biosynthesis pathway of a kind of stilbenes, stilbene glucoside (2,3,5, 4’-tetra-hydroxy-stilbene-2-O-β-D-glucoside) which is the major bioactive principles in Polygonum multiflorum (Figure 2). The application of Systems biology in the research on relevant genes and enzymes in biosynthesis pathway of stilbene glucoside can be described as follows. Firstly, the possible relevant genes or enzymes should be identified from Polygonum multiflorum. The clone of a type Ш polyketide synthase gene (FmPKS) using oligonucleotide primers designed for regions conserved amongst STS (with special attention given to the closely-related species of the Polygonaceae family) in Fallopia multiflora have be conducted by Shujing Sheng [51]. The FmPKS has been identified strongly correlates with the accumulation of stilbene glucoside by using Northern blotting, RNA inference and over expression, suggesting that FmPKS might play an important role in its biosynthesis. Resveratrol (Figure 1), another kind of stilbenes, is similar to stilbene glucoside in structure. Zhongyu Liu has investigated the overexpression of a resveratrol synthase gene (PcRS) from Polygonum cuspidatum which is closely related to the Fallopia multiflora in transgenic Arabidopsis. As a result, it causes the accumulation of trans-piceid with antifungal activity [52]. The foregoing research may be summed up to search for individual possible genes or enzymes involved in biosynthesis pathway of stilbene glucoside and study gene functions using the classical molecular biology.
Secondly, search for the possible relevent genes, enzymes through approaches of functional genomics, transcriptomics, proteomics. At present, suppression subtracttive hybridization (SSH) was performed to search for genomic differences. A subtractive cDNA library was constructed by using cDNA from Polygonum multiforum root tubers with high content of stilbene glucoside as tester and low content as driver for the subtractive hybridization. As a result, 11 clones were obtained as the differentially expressed candidates which play an important roles in further validation of genes involved in the
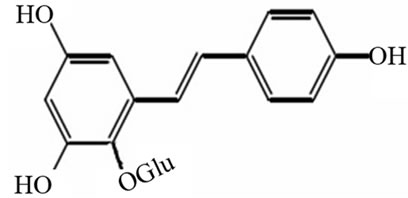
Figure 2. 2,3,5,4’-tetra-hydroxy-stilbene-2-O-β-D-glucoside.
biosynthesis pathway of stilbene glucoside in Polygonum multiflorum. Rapid-amplification of cDNA ends (RACE) was carried out to gain the full-length cDNAs. The application of RNA interference and over expression of these genes will reveal the specific functions of these genes and screen the genes involved in the biosynthesis pathway of stilbenes.
Thirdly, internal components (such as genetic mutation, interference) or external conditions of the Polygonum multiflorum is to be changed. The accumulation of stilbene glucoside, gene expression and enzyme expression in these cases should be detected. Next all the relevant information should be integrated to derivate how they regulate the biosynthesis of stilbene glucoside. The second step and third step ought to be repeated to revise and refine the model through a large number of experimental results, in order to finally gain all the relevent genes, enzymes and regulatory factors in the biosynthesis pathway of stilbene glucoside. These researches are from the thinking model of systems biology at different levels, which has important significance in revealing the biosynthesis pathway and regulation mechanism of stilbene glucoside in Polygonum multiflorum. The discussion and related diagrams give a good idea of the application of systems biology in research on relevent genes and enzymes in biosynthesis pathway of stilbene glucoside (Figure 3).
4. The Application of the Research on Enzymes and Genes in Biosynthesis Pathway of Stilbenes
The elucidating of stilbenes’ biosynthesis pathway and
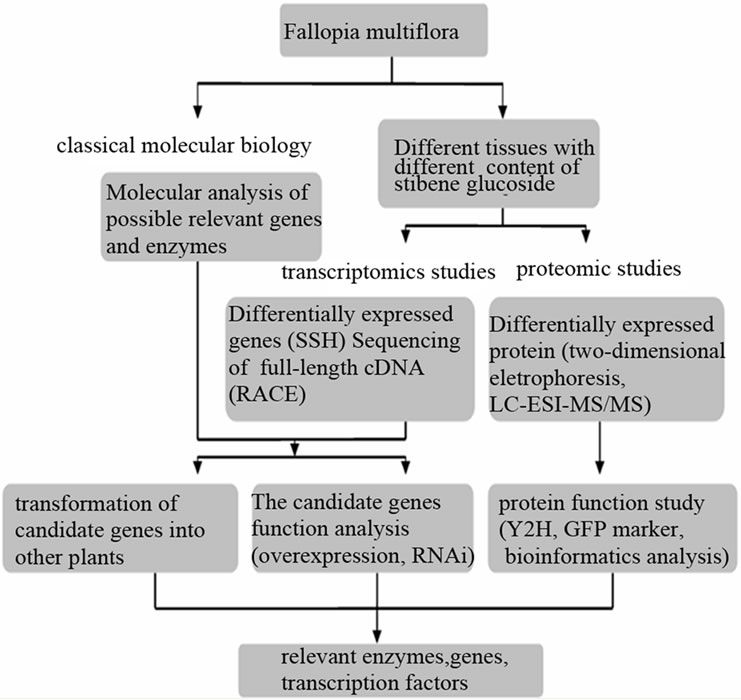
Figure 3. The application of systems biology in research on relevent genes and enzymes in biosynthesis pathway of stilbene glucoside.
regulation mechanism can accelerate the course of metabolic engineering as a tool for plant disease control and hunman health promotion. The increasing maturity of the plant genetic engineering technology promote the research on the biosynthesis pathway of stilbenes, and the two well studied field are stilbene synthase gene engineering and its transcription factors or regulation engineering.
4.1. The Genetic Engineering Research of Metabolic Key Enzymesin Biosynthesis Pathway of Stilbenes
STS plays an important regulatory role in biosynthesis pathway of resveratrol and other stilbene phytoalexin. An important goal of STS gene engineering is the genetic improvement of plants in increasing resistance against diseases. Namely transformate target plant genomes with chimeric strong promoter and STS gene through transgenic technology, which makes transgenic plants express STS in order to start the stilbene biosynthesis pathway, and change the plant traits and enhance plant defense against external violation [53].
Grape STS gene was introduced into Nicotiana tabacum, which results in transgenic tabacum express the STS and synthesize resveratrol and then dramatically increase its resistance against diseases [54,55]. The transformation of rice, wheat and other crops with grape STS gene also increase the resistance level against disease significantly of transgenic crops [56]. At present, the transgenic plants with STS gene include apple, poplar, papaya, white poplar, rape, wheat, peas, tomatoes, lettuce, hops, Arabidopsis thaliana [57-67]. Through introducing STS gene, most of transgenic plants can exhibit STS activity and synthesize exogenous stilbene phytoalexin.
4.2. The Gene Engineering of Regulating Gene or Transcription Factor in Biosynthesis Pathway of Stilbenes
Cotransformation of two or several related enzyme genes in downstream of biosynthesis pathway can be used when study stilbenes’ biosynthesis involves multiple genes expression. In this way, the new secondary biosynthesis branch can be introduced into the plant then to increase the content of stilbene secondary metabolites in the transgenic plant or synthesize exogenous stilbenes. Coordinated expression of enzyme genes in secondary metabolism channel and the same or similar cisacting elements of regulatory sequences of these genes are regulated by the same transcription factors or regulation genes. Therefore, enhancing the expression of transcription factor genes and regulation genes of these important enzyme genes is a feasible way to achieve enhanced coordinated expression of multiple genes. This requires more understanding about identification and regulation of enzymes gene involved in biosynthesis pathway of stilbenes.
5. Summary
Now we have only a rudimentary grasp of the basic framework of the main plant secondary metabolic pathways such as alkaloids synthesis pathway, phenylpropanoid biosynthesis pathway and isoprenoid biosynthesis pathway, but lack of research on rate-limiting steps, related isozymes and specific biosynthesis pathways of specific secondary metabolite. Plant secondary metabolism is a complicated dynamic process which is regulated by plant genetic background and growth process, also affected by ecological environment and pathogen infection, insect feeding and stimulation of various elicitor. The induction expression of related genes and enzymes also need be further studied. Because of disease resistance and medicinal health functions of resveratrol, stilbene glucoside and other inducible stilbenes, secondary metabolic engineering can be used to increase the content of exogenous stilbenes in transgenic plants. However, the complexity of enzymes in biosynthesis pathway and regulation of gene expression increase the difficulty in researching metabolism genetic engineering of stilbenes. The thinking model of systems biology can be used to research the biosynthesis pathway of secondary metabolites and understand sequence of events of intermediate products and final products. The enzymes and their genes expression and regulation in each reaction step as well as the interaction of each biosynthesis branch can be definited from the view of metabolic channeling. Then after continuous integration and analysis, the detailed metabolic pathways and regulation mechanism of stilbenes will be finally elucidated. This will be the future research emphasis and direction for people to understand about regulations of secondary metabolic pathways in plants and coordination between the secondary metabolic pathways.
6. Acknowledgements
We thank Dr. Shujing Sheng and Ping Yan for helpful discussions and all those who helped us obtain and identify plants: Prof. Fuwu Xing, South China Botanical Garden; and Prof. Liping Tang, Kunming Medical University.
REFERENCES
- B. Delaunois, S. Cordelier and A. Conreux1, “Molecular Engineering of Resveratrol in Plants,” Plant Biotechnology Journal, Vol. 7, No. 1, 2009, pp. 2-12. doi:10.1111/j.1467-7652.2008.00377.x
- J. K. Lin and S. H. Tsai, “Chemoprevention of Cancer and Cardiovascular Disease by Resveratrol,” Proceedings of the National Science Council, Republic of China (Part B), Vol. 23, No. 3, 1999, pp. 99-106.
- K. Likhitwitayawuid and B. Sritularak, “A New Dimeric Stilbene with Tyrosinase Inhibitory Activity from Artocarpus Gomezianus,” Journal of Natural Products, Vol. 64, 2001, pp. 1457-1459. doi:10.1021/np0101806
- S. L. He, J. C. Zheng and M. Lin, “Advances of Biological Function, Regulatory Mechanism of Biosynthesis and Genetic Engineering of in Stilbenes Plant,” Journal of Agricultural Biotechnology, Vol. 12, No. 1, 2004, pp. 102-108.
- H. C. Zhang, J. M. Liu and H. M. Chen, “Up-Regulation of Licochalcone a Biosynthesis and Secretiony tween 80 in Hairy Root Cultures of Glycyrrhiza uralensis Fisch,” Molecular Biotechnology, Vol. 47, No. 1, 2011, pp. 50- 56. doi:10.1007/s12033-010-9311-4
- L. Li, Y. Zhao and J. l. Ma, “Recent Progress on Key Enzymes: Pal, C4h, 4cl of Phenylalanine Metabolism Pathway,” China Journal of Bioinformatics, Vol. 4, 2007, pp. 187-189.
- S. Rasmussen and R. A. Dixon, “Transgene-Mediated and Elicitor-Induced Perturbation of Metabolic Channeling at the Entry Point into the Phenylpropanoid Pathway,” Plant Cell, Vol. 11, 1999, pp. 1537-1551. doi:10.2307/3870981
- T. Fahrendorf and R. A. Dixon, “Stress Responses in Alfalfa (Medicago Sativa L). XVШ: Molecular Cloning and Expression of the Elicitor-Inducible Cinnamic Acid 4- Hydroxylase Cytochrome P450,” Archives of Biochemistry and Biophysics, Vol. 305, No. 2, 1993, p. 509. doi:10.1006/abbi.1993.1454
- H. G. Teutsch, M. P. Hasenfratz and A. Lesot, “Isolation and Sequence of a cDNA Encoding the Jerusalem Artichoke Cinnamate 4-Hydroxylase, a Major Plant Cytochrome P450 Involved in the General Phenylpropanoid Pathway,” Proceedings of the National Academy of Sciences of the United States of America, Vol. 90, No. 9, 1993, p. 4102. doi:10.1073/pnas.90.9.4102
- C. R. Buell and S. C. Somerville, “Expression of Defense-Related and Putative Signaling Genes during Tolerant and Susceptible Interactions of Arabidopsis with Xanthomonas Campestris pv. Campestris,” Molecular Plant-Microbe Interactions, Vol. 8, No. 3, 1995, pp. 435- 443. doi:10.1094/MPMI-8-0435
- Y. Batard, M. Schalk and M. A. Pierrel, “Regulation of the Cinnamate 4-Hydroxylase (CYP73A1) in Jerusalem Artichoke Tubers in Response to Wounding and Chemical Treatments,” Plant Physiology, Vol. 113, No. 3, 1997, pp. 951-959.
- D. A. Bell-Lelong, J. C. Cusumano and K. Meyer, “Cinnamate 4-Hydroxylase Expression in Arabidopsis,” Plant Physiology, Vol. 113, No. 3, 1997, pp. 729-738. doi:10.1104/pp.113.3.729
- M. Mizutani, D. Ohta and R. Sato, “Isolation of cDNA and a Genomic Clone Encoding Cinnamate 4-Hydroxylase from Arabidopsis and Its Expression Manner in Planta,” Plant Physiology, Vol. 113, No. 3, 1997, pp. 755-763. doi:10.1104/pp.113.3.755
- B. S. Winkel, “Metabolic Channeling in Plant,” Annual Review of Plant Biology, Vol. 55, 2004, pp. 85-107. doi:10.1146/annurev.arplant.55.031903.141714
- X. Y. Dong, E. L. Braun and E. Grotewold, “Functional Conservation of Plant Secondary Metabolic Enzymes Revealed by Comp Lamentation of a Rabidopsis Flavonoid Mutants with Maize Genes,” Plant Physiology, Vol. 127, 2001, pp. 46-57. doi:10.1104/pp.127.1.46
- I. E. Burbulis and B. Winkel-Shirley, “Interactions among Enzymes of the Arabidopsis Flavonoid Biosynthestic Pathway,” Proceedings of the National Academy of Sciences of the United States of America, Vol. 96, No. 22, 1999, pp. 12929-12934. doi:10.1073/pnas.96.22.12929
- X. Z. He and R. A. Dixon, “Genetic Manipulation of Isoflvone 7-O-Methyltransferase Enhances Biosynthesis of 4’-O-Methylated Isoflavonoid Phytoalexins and Disease Resistance in Alfalfa,” Plant Cell, Vol. 12, 2001, pp. 1689-1702. doi:10.2307/3871183
- C. Chapple, “Molecular-Genetic Analysis of Plant Cytochrome P450-Dependent Monooxygenases,” Annual Review of Plant Biology, Vol. 49, 1998, pp. 311-343. doi:10.1146/annurev.arplant.49.1.311
- B. Winkel-Shirley, “Flavonoids Biosynthesis: A Colorful Model for Genetics, Biochemistry, Cell Biology, and Biotechnology,” Plant Physiology, Vol. 126, 2001, pp. 485- 493. doi:10.1104/pp.126.2.485
- R. A. Dixon, “Natural Products and Plant Disease Resistance,” Nature, Vol. 411, No. 14, 2001, pp. 843-847. doi:10.1038/35081178
- M. B. Austin and J. P. Noel, “The Chalcone Synthase Su-Perfamily of Type Ш Polyketide Synthases,” Natural Product Reports, Vol. 20, No. 1, 2003, pp. 79-110. doi:10.1039/b100917f
- K. Hanhineva, H. Kokko and H. Siljanen, “Stilbene Synthase Gene Transfer Caused Alterations in the Phenylpropanoid Metabolism of Transgenic Strawberry,” Journal of Experimental Botany, Vol. 60, No. 7, 2009, pp. 2093-2106. doi:10.1093/jxb/erp085
- H. Morita, H. Noguchi and J. Schroder, “Novel Polyketides Synthesized with a Higher Plant Stilbene Synthase,” European Journal of Biochemistry, Vol. 268, No. 13, 2001, pp. 3759-3766. doi:10.1046/j.1432-1327.2001.02289.x
- S. Rasmussen and R. A. Dixon, “Transgene-Mediated and Elicitor-Induced Perturbation of Metabolic Channeling at the Entry Point into the Phenylpropanoid Pathway,” Plant Cell, Vol. 11, 1999, pp. 1537-1551. doi:10.2307/3870981
- H. E. Shuilin, J. g. Zheng and X. f. Wang, “Plant Secondary Metabolism: Function, Regulation and Gene Engineering,” Chinese Journal of Applied & Environmental Biology, Vol. 8, No. 5, 2002, pp. 558-563.
- F. Melchior and H. Kindl, “Coordinateand Elicitor- Dependent Expression of Stilbene Synthase and Phenylalanine Ammonialyase Genes in Vitis cv. Optima,” Archives of Biochemistry and Biophysics, Vol. 288, No. 2, 1991, pp. 552-557. doi:10.1016/0003-9861(91)90234-A
- D. J. Kliebenstein, “Secondary Metabolites and Plant/ Environment Interactions: A View through Arabidopsis Thaliana Tinged Glasses,” Plant Cell and Environment, Vol. 27, No. 6, 2004, pp. 675-684. doi:10.1111/j.1365-3040.2004.01180.x
- M. Lerdau, “Benefits of the Carbon-Nutrient Balance Hypothesis,” Oikos, Vol. 98, No. 3, 2011, pp. 534-536. doi:10.1034/j.1600-0706.2002.980318.x
- M. A. Lila, “The Nature-Versus-Nurture Debate on Bioactive Phytochemicals: The Genome Versus Terroir,” Journal of the Science of Food and Agriculture, Vol. 86, No. 15, 2006, pp. 2510-2515. doi:10.1002/jsfa.2677
- Z. l. Yuan, C. c. Dai and L. q. Chen, “Regulation and Accumulation of Secondary Metabolites in Plant-Fungus Symbiotic System,” African Journal of Biotechnology, Vol. 6, No. 11, 2007, pp. 1266-1271.
- D. V. Endt, J. W. Kijne and J. Memelink, “Transcription Factors Controlling Plant Secondary Metabolism: What Regulates the Regulators?” Phytochemistry, Vol. 61, No. 2, 2002, pp. 107-114. doi:10.1016/S0031-9422(02)00185-1
- i. M. Chung, M. R. Park and S. Rehman, “Tissue Specific and Inducible Expression of Resveratrol Synthase Gene in Peanut Plants,” Molecules and Cells, Vol. 12, No. 3, 2001, pp. 353-359.
- H. Chiron, A. Drouet and A. C. Claudot, “Molecular Cloning and Functional Expression of a Stress-Induced Multifunctional O-Methyltransferase with Pinosylvin Methyltransferase Activity from Scots Pine (Pines sylvestris L.),” Plant Molecular Biology, Vol. 44, No. 6, 2000, pp. 733-745. doi:10.1023/A:1026507707186
- E. Cantos, J. C. Espin and F. A. Tomas-Barberan, “Postharvest Induction Modeling Method Using UV Irradiation Pulses for Obtaining Resveratrol-Enriched Table Grapes: A New Functional Fruit?” Journal of Agricultural and Food Chemistry, Vol. 49, No. 10, 2001, pp. 5052-5058. doi:10.1021/jf010366a
- H. Kitano, “Systems Biology: Toward System-Level Understanding of Biological Systems,” MIT Press, Cambridge, 2002.
- M. Latterich, “Molecular Systems Biology at the Crossroads: To Know Less about More, or to Know More about Less?” Proteome Science, Vol. 3, 2005, pp. 8-11. doi:10.1186/1477-5956-3-8
- L. Hood, J. R. Heath and M. E. Phelps, “Systems Biology and New Technologies Enable Predictive and Preventative Medicine,” Science, Vol. 305, No. 5696, 2004, pp. 640-643. doi:10.1126/science.1104635
- K.-M. Oksman-Caldentey, D. Inze and M. Oresic, “Connecting Genes Tometabolites by a Systems Biology Approach,” Proceedings of the National Academy of Sciences of the United States of America, Vol. 101, No. 27, 2004, pp. 9949-9950. doi:10.1073/pnas.0403636101
- A. Aderem, “Systems Biology: Its Practice and Challenges,” Cell, Vol. 121, No. 4, 2005, pp. 511-513. doi:10.1016/j.cell.2005.04.020
- J. x. Wang, Y. Sun and P. l. Xu, “Research Progress in Functional Plant Genomics,” Biotechnology Bulletin, Vol. 1, 2004, pp. 18-24.
- W. A. Rensink and C. R. Buell, “Microarray Expression Profiling Resources for Plant Genomics,” Trends in Plant Science, Vol. 10, No. 12, 2005, pp. 603-612. doi:10.1016/j.tplants.2005.10.003
- V. Marnik, D. P. Johan and M. J. T. van Eijk, “AFLPBased Transcript profiling (cDNA-AFLP) for genomeWide Expression Analysis,” Nature Protocols, Vol. 2, No. 6, 2007, pp. 1399-1413. doi:10.1038/nprot.2007.174
- W. Jia, P. Liu and J. Jiang, “Application of Metabonomics in Complicated Theory System Research of Traditional Chinese Medicine,” China Journal of Chinese Materia Medica, Vol. 31, No. 8, 2006, pp. 621-625.
- L. w. Qi, P. Li and J. Zhao, “Metabonomics and Modernization of TMC Research,” World Science and Technology, Vol. 8, No. 6, 2006, pp. 79-87.
- L. W. Sumner, P. Mendes and R. A. Dixon, “Plant metabolomics: Large-Scale Phytochemistry in the Functional Genomics Era,” Phytochemistry, Vol. 62, No. 6, 2003, pp. 817-836. doi:10.1016/S0031-9422(02)00708-2
- L. q. Huang, W. Gao and J. Zhou, “Systems Biology Applications to Explore Secondary metabolites in Medicinal Plants,” China Journal of Chinese Materia Medica, Vol. 35, No. 1, 2010, pp. 8-12.
- G. h. Cui, L. q. Huang and X. j. Tang, “Functional Genomics Studies of Salvia Miltiorrhiza І Establish cDNA Microarray of S. miltiorrhiza,” China Journal of Chinese Materia Medica, Vol. 32, No. 12, 2007, pp. 1137-1142.
- W. Gao, G. h. Cui and J. q. Kong, “Optimizing Expression and Purification of Recombinant Salvia Miltiorrhiza Copalyl Diphosphate Synthase Protein in E. coli and Preparation of Rabbit Antiserum against SmCPS,” Acta Pharmaceutica Sinica, Vol. 43, No. 7, 2008, p. 766.
- X. y. Wang, G. h. Cui and L. q. Huang, “A Full Length cDNA of 4-(Cytidine 5’-Diphospho)-2-C-Methyl-D-Erythritol Kinase Cloning and Analysis of Introduced Gene Expressing in Sava Miltiorrhiza,” Acta Pharmaceutica Sinica, Vol. 43, No. 12, 2008, pp. 1251-1259.
- W. Gao, M. L. Hillwig and L. q. Huang, “A Functional Genomics Approach to Tanshinone Biosynthesis Provides Stereo-Chemical Insights,” Organic Letters, Vol. 11, No. 22, 2009, pp. 5170-5173. doi:10.1021/ol902051v
- S. J. Sheng, Z. Y. Liu and W. Zhao, “Molecular Analysis of a Type Ш Polyketide Synthase Gene in Fallopia Multiflora,” Section Cellular and Molecular Biology, Vol. 65, No. 6, 2010, pp. 939-946.
- Z. y. Liu, C. x. Zhuang and S. j. Sheng, “Overexpression of a Resveratrol Synthase Gene (PcRS) from Polygonum Cuspidatum in Transgenic Arabidopsis Causes the Accumulation of Trans-Piceid with Antifungal Activity,” Plant Cell Reports, Vol. 30, No. 11, 2011, pp. 2027-2036. doi:10.1007/s00299-011-1110-2
- S. Raiber, G. Schroder and J. Schroder, “Molecular and Enzymatic Characterization of Two Stilbene Synthases from Eastern White Pine (Pinus strobus). a Single Arg/ His Difference Determines the Activity and the pH Dependence of the Enzymes,” FEBS Letters, Vol. 361, No. 2-3, 1995, pp. 229-302. doi:10.1016/0014-5793(95)00199-J
- R. Hain, B. Bleseler, H. Kindl, G. Schroder and R. Stocker, “Expression of a Stilbene Synthase Gene in Nicotiana Tabacumresults in Synthesis of the Phytoalexin Resveratrol,” Plant Molecular Biology, Vol. 15, No. 2, 1990, pp. 325-335. doi:10.1007/BF00036918
- R. Hain, H. J. Reif and E. Krause, “Disease Resistance Results from Foreign Phytoalexin Expression in a Novel Plant,” Nature, Vol. 361, No. 6408, 1993, pp. 153-156. doi:10.1038/361153a0
- W. Z. Tian, L. Ding and S. Y. Cao, “Rice Transformation with a Phytoalexin Gene and Bioassay of the Transgenic Plants,” Acta Botanica Sinica, Vol. 40, No. 9, 1998, pp. 803-808.
- L. Serazetdinova, K. H. Oldach and H. Lörz, “Expression of Transgenic Stilbene Synthases in Wheat Causes the Accumulation of Unknown Stilbene Derivatives with Antifungal Activity,” Plant Physiology, Vol. 162, No. 9, 2005, pp. 985-1002. doi:10.1016/j.jplph.2004.11.005
- I. Szankowski, K. Briviba and J. Fleschhut, “Transformation of Apple (Malus domestica Borkh.) with the Stilbene Synthase Gene from Grapevine (Vitis vinifera L.) and a PGIP Gene from Kiwi (Actinidia deliciosa),” Plant Cell Reports, Vol. 22, No. 2, 2003, pp. 141-150. doi:10.1007/s00299-003-0668-8
- S. K. Seppänen, L. SyrjäläL, et al., “Antifungal Activity of Stilbenes in in Vitro Bioassays and in Transgenic Populus Expressing a Gene Encoding Pinosylvin Synthase,” Plant Cell Reports, Vol. 22, No. 8, 2004, pp. 584- 593.
- [61] Y. J. Zhu, R. Agbayani and M. C. Jackson, “Expression of the Grapevine Stilbene Synthase Gene VST1 in Papaya Provides Increased Resistance against Diseases Caused by Phytophthora Palmivora,” Planta, Vol. 220, No. 2, 2004, pp. 241-250. doi:10.1007/s00425-004-1343-1
- [62] A. Giorcelli, F. Sparvoli and F. Mattivi, “Expression of the Stilbene Synthase (StSy) Gene from Grapevine in Transgenic White Poplar Results in High Accumulation of the Antioxidant Resveratrol Glucosides,” Transgenic Research, Vol. 13, No. 3, 2004, pp. 203-214. doi:10.1023/B:TRAG.0000034658.64990.7f
- [63] A. Hüsken, A. Baumert and C. Milkowski, “Resveratrol Gluco-Side (Piceid) Synthesis in Seeds of Transgenic Oilseed Rape (Brassica napus L.),” Theoretical and Applied Genetics, Vol. 111, No. 8, 2005, pp. 1553-1562. doi:10.1007/s00122-005-0085-1
- [64] R. Morelli, S. Das and A. Bertelli, “The Introduction of the Stilbene Synthase Gene Enhances the Natural Antiradical Activity of Lycopersicon Esculentum Mill,” Molecular and Cellular Biochemistry, Vol. 82, No. 1-2, 2006, pp. 65-73. doi:10.1007/s11010-006-1260-7
- [65] A. Richter, H. J. Jacobsen and A. de Kathen, “Transgenic Peas (Pisum sativum) Expressing Polygalacturonase Inhibiting Protein from Raspberry (Rubus idaeus) and Stilbene Synthase from Grape (Vitis vinifera),” Plant Cell Reports, Vol. 25, No. 11, 2006, pp. 1166-1173. doi:10.1007/s00299-006-0172-z
- [66] S. Liu, Y. Hu and X. Wang, “High Content of Resveratrol in Lettuce Transformed with a Stilbene Synthase Gene of Parthenocissus Henryana,” Journal of Agricultural and Food Chemistry, Vol. 54, No. 21, 2006, pp. 8082-8825. doi:10.1021/jf061462k
- [67] A. Schwekendiek, O. Spring and A. Heyerick, “Constitutive Expression of a Grapevine Stilbene Synthase Gene in Transgenic Hop (Humulus lupulus L.) Yields Resveratrol and Its Derivatives in Substantial Quantities,” Journal of Agricultural and Food Chemistry, Vol. 55, No. 17, 2007, pp. 7002-7009. doi:10.1021/jf070509e
- [68] C. Lo, J. C. Le Blanc, et al., “Detection, Characterization, and Quantification of Resveratrol Glycosides in Transgenic Arabidopsis over—Expressing a Sorghum Stilbene Synthase Gene by Liquid Chromatography/Tandem Mass Spectrometry,” Rapid Communications in Mass Spectrometry, Vol. 21, No. 24, 2007, pp. 4101-4108. doi:10.1002/rcm.3316
NOTES
*Corresponding author.

