Open Journal of Genetics
Vol.2 No.1(2012), Article ID:17762,6 pages DOI:10.4236/ojgen.2012.21001
Irr1/Scc3 cohesin interacts with Rec8 in meiotic prophase of Saccharomyces cerevisiae
![]()
Institute of Biochemistry and Biophysics, Polish Academy of Sciences, Warszawa, Poland
Email: *ania218@ibb.waw.pl
Received 7 November 2011; revised 11 December 2011; accepted 21 January 2012
Keywords: Irr1/Scc3; Cohesin; Rec8; Meiosis; Saccharomyces cerevisiae
ABSTRACT
The meiotic cohesin complex of S. cerevisiae shares with the mitotic one the Irr1/Scc3, Smc1, and Smc3 subunits, while the meiosis-specific subunit Rec8 replaces mitotic subunit Scc1/Mcd1. We noticed earlier that the irr1-1 mutation (F658G) severely affected meiosis. The irr1-1/IRR1 cells were entering meiosis before having completed mitotic cell division. Using meiotic two-hybrid assay and co-immunoprecipitation we show that in cells arrested in pachytene due to a lack of a gene-regulatory factor Ndt80, the Irr1 protein interacts with Rec8p and the irr1-1 mutation abolishes this interaction. These findings indicate an important role of Irr1p in early stages of meiosis.
1. INTRODUCTION
Diploid eukaryotic cells give rise to haploid ones through meiosis. The essential components of the meiotic apparatus are highly conserved among eukaryotes, but the regulatory pathways responsible for initiating the meiotic program vary substantially [1].
The first phase of chromosome segregation in meiosis (meiosis I, a reductional segregation) requires that homologous chromosomes separate. During this phase, homologs (each consisting of two sister chromatids) must be aligned and then linked through recombination during an extended prophase. The homologs are then segregated to the opposite poles while the sister chromatids are kept together through meiosis-specific cohesion. Missegregation of chromosomes in meiosis I may occur due to defective cohesion along the chromosome arms or to reduced crossing-over [reviewed in 1-3].
Sister chromatid cohesion, established during the premeiotic DNA replication, is central to two rounds of meiotic chromosome segregation. The meiotic cohesin complex of S. cerevisiae differs from the mitotic one by having the Rec8 subunit largely replacing Scc1/Mcd1 [4]. The other subunits, Irr1/Scc3, Smc1, and Smc3 are common to the two types of cohesin complexes. Rec8 is important to hold sister chromatids together on the metaphase I and metaphase II spindles [4]. At the end of meiotic prophase I, the four homologous chromatids are linked through chiasmata resulting from recombination, and through cohesin linkages between sister chromatids. For the homologs to segregate during meiosis I, cohesins must be removed along the chromosome arms. Cohesins are maintained at centromeres, allowing the sister chromatids to continue to associate until the metaphase II-toanaphase II transition, when the remaining cohesin is removed, allowing formation of four haploids [reviewed in 1,2,5]. The release of cohesion occurs in two steps. It is released over a portion of each bivalent to allow dissolution of chiasmata and homolog segregation in meiosis I. Here, cohesin is removed from the chromosome arms by an activated protease separase which cleaves Rec8 [6,7, reviewed in 8]. Multiple phosphorylation sites within Rec8, and two different kinases, CK1delta/ epsilon and Dbf4-dependent Cdc7 are required for Rec8 cleavage and meiosis I nuclear division [9]. However, during meiosis I centromere cohesion is protected because centromeric Rec8p is not cleaved by separase, which is then protected by shugoshin/MEI-S332 proteins and dephosphorylated by PP2A phosphatase [7,8,10].
Recent data indicate that the protein Rec8 has multiple roles. It is essential to maintain sister chromatid cohesion, but it is also required for homolog pairing and participates in meiotic recombination as a prerequisite for synaptonemal complex (SC) assembly [3]. These roles of Rec8p are genetically separable from the cohesion function.
In our previous paper [11] we found that the irr1-1 point mutation (F658G) severely affected meiosis. This mutation is lethal in the haploid and and semi-dominant in heterozygous irr1-1/IRR1 diploid. A fraction of such diploid cells produced incorrect asci whose morphology suggested that they entered meiosis before having completed mitotic cell division. Here we show that in meiotc cells arrested in pachytene due to a lack of a gene-regulatory factor Ndt80 [12], the Irr1 protein interacts with Rec8p and the irr1-1 mutation compromises this interacttion. This indicates an important role of Irr1p in early stages of meiosis.
2. MATERIALS AND METHODS
2.1. Strains, Media, Plasmids
Yeast strains and plasmids used in this study are listed in Table 1. Escherichia coli XL1-Blue MRF’ (Stratagene, Saint Quentin en Yvelines, France) was used for molecular manipulations. Yeast culture media were prepared as described [13]. YPD contained 1% Bacto-yeast extract, 2% Bacto-peptone and 2% (all w/v) glucose. SD contained 0.67% yeast nitrogen base without amino acids (Difco, Detroit, MI, USA) and 2% glucose. For auxotrophic strains, the media contained appropriate supplements. Standard methods were used to genetically manipulate yeast cells [13]. Routinely, whenever possible strains were backcrossed at least twice.



Table 1. S. cerevisiae strains and plasmids used in this study.
2.2. Meiosis-Specific Two-Hybrid Assay
Interactions of Rec8p with Irr1p were investigated according to Arora et al. [14] and the following plasmids were constructed: PREC8-REC8-GAL4-AD TADH1, PADH1- LexA-BD-irr1-1ΔC435 TADH1, PADH1-LexA-BD-IRR1 ΔC435 TADH1, and PADH1-LexA-BD-IRR1 TADH1. The fusion of Rec8p to the C terminus of the Gal4 activation domain was initially prepared in YEp351 and then cloned into pYEp181, generated by amplification of GAL4-TADH1 on pGAD424 and fusing to PREC8-REC8 fragment obtained from p4052 plasmid [15]. Fusions of Irr1 to the LexA DNA binding domain were prepared in pBTM116. Haploid reporter strains SK661a or SK662α [14] were transformed with respective individual plasmids and mated in appropriate pair wise combinations. The resulting diploids were grown to saturation in minimal medium Leu– Trp–. Subsequently, cells were harvested, transferred into 1% potassium acetate and incubated for 16 hours at 30˚C with good aeration. Equivalents of 5 OD600 units were harvested, cells were lysed and β-galactosidase activity was detected out according to standard protocols (Clontech). One unit of β-galactosidase hydrolyzes 1 μmol of o-nitrophenyl β-D-galactopyranoside per minute per one OD600. All experiments were repeated ten times.
2.3. Western Blotting and Co-Immunoprecipitation
To visualize chimeric HAor c-Myc-tagged proteins on Western blots, protein samples (100 μg/lane) were subjected to 8% SDS-PAGE. Electrophoresis was followed by blotting onto Hybond-C extra membrane and probing with anti-HA (HA.11, clone 16B12) (BabCO) or anti-cMyc 9E10 (BabCO) monoclonal antibody. For co-immunoprecipitation, cells expressing chimeric Irr1p-Myc and Rec8p-HA or Irr1-1p-Myc and Rec8p-HA proteins were harvested at an OD600 = 1 and resuspended in 50 mM sodium phosphate buffer pH 7.5, 50 mM NaCl, 2 mM MgCl2, 0.2% Triton X-100, containing protease inhibitor cocktail Complete (Roche) and 25 U/μl of endonuclease Benzonase (Merck). Fresh, non-frozen cells were homogenized with glass beads in a bead-beater; the homogenate was spun down in a microfuge. The whole procedure was carried out on ice or at 4˚C. Soluble protein extracts were incubated with Protein G-agarose (Sigma) covered with anti-HA (Covance, monoclonal antibody HA.11 clone 16B12) or anti-Myc (Covance, monoclonal antibody clone 9E10) overnight at 4˚C. The immunoprecipitate was analyzed by SDS-PAGE, blotted and probed with anti-HA and anti-Myc antisera, as above. No Irr1p-Rec8p co-immunoprecipitation could be detected when cells or homogenates were frozen during the procedure.
3. RESULTS AND DISCUSSION
The exact architecture of meiotic cohesin complexes in S. cerevisiae is not well established. Rec8p (similarly to Scc1p/Mcd1p) is known to interact with Smc1p and Smc3p [16] but the role of Irr1p/Scc3p is poorly described. Our previous study indicated that mutated Irr1-1p cohesin (bearing the F658G substitution) present in the heterozygous diploid irr1-1/IRR1 causes chromosome missegregation and aberrations in meiotic divisions manifested by asci of incorrect morphology. In numerous cells meiosis was initiated without completion of mitotic divisions [11]. Our data suggested malfunctioning of the spindle assembly checkpoint, but we assumed that such effect could also be due to a disturbance of a putative Irr1-1p-Rec8p interaction in early meiosis.
The Irr1 protein is present both in mitotic and meiotic cells [6]. Very recently Katis et al. [9] detected Irr1p/ Scc3p in a multi-protein complex derived from diploid yeast cells arrested in metaphase I, in which TAP-tagged Rec8p served as a bait. Here we carried out a meiotic two-hybrid assay using N-terminal fusion of LexA-BD with Irr1p and Rec8p C-terminally fused with Gal4-AD and expressed from REC8 promoter. Irr1p was used in two versions: full-length Irr1p and Irr1ΔC435p, devoid of 435 C-terminal amino acids. The shortened protein Irr1ΔC435 of 715 amino acids was used previously in our mitotic two-hybrid assay and it was the only bait which produced an interaction [17]. According to the InterPro [18] and Pfam [19] databases, the domain STAG of Irr1 is localized between K211 and G350 and thus is preserved in Irr1ΔC435p. The role of STAG domain is not known but it is evolutionarily conserved and thus very likely to be important for Irr1p functioning. Both full-length Irr1p and Irr1ΔC435 were tested for interaction with Rec8p. The interaction was followed in a ndt80Δ/ndt80Δ diploid strain of SK1 background, equipped with the lacZ reporter system [14]. The fusion protein LexA-BD-Irr1 (whole-length Irr1p) did not interact with Rec8-Gal4-AD. Since the whole-length Irr1p used in our laboratory in a previous two-hybrid screen also did not produce results [17] we assume that LexABD is masked in such fusion protein. Moreover, there are no literature data concerning interactions between wholelength Irr1p and other proteins. Data present in databases came from global screens and do not include sufficient details.
While LexA-Irr1ΔC435 distinctly interacted with Rec8p: β-galactosidase signal (representing strength of interaction between partners) was significantly stronger than that due to the strong interaction between Spo11- LexAAD and Rec104-Gal4-BD used as a positive control [14] (Figure 1(a)). In contrast, the mutated protein Irr1-1ΔC435 (bearing the F658G substitution) hardly interacted with Rec8p: the signal was only twice that of the negative control (Spo11-LexA-AD without a partner). We checked that LexA-BD-Irr1ΔC435, LexABD-Irr1-1 ΔC435 and Rec8-Gal4-AD were present in the cell in comparable amounts and had correct molecular masses. Thus, the significantly reduced (by ca. 88%) two-hybrid reporter signal in a strain bearing LexABD-Irr1-1 compared to the control LexABD-Irr1 is not caused by a reduction of the LexABD-Irr1-1 level (Figure 1(b)).
These results supported the initial conjecture that in
 (a)
(a) (b)
(b)
Figure 1. Irr1/Scc3 protein interacts with Rec8p in early meiosis and F658G substitution abolishes this interaction. (a) Interaction between Irr1ΔC435p and Rec8p (black bar) is reflected by β-galactosidase activity, shown relative to a positive control (100%). Interaction between Irr1-1ΔC435p and Rec8p is represented by white bar. Positive control: interaction between Spo11-LexAAD and Rec104-Gal4-BD, negative controls: 1: Spo11-LexA-AD without partner; 2: LexA-BD-Irr1ΔC435 with plasmid bearing only Gal4-BD; 3: LexABD-Irr1-1 ΔC435 with plasmid bearing only Gal4-BD; 4: Rec8-Gal4-AD with plasmid bearing only LexA-BD. Error bars—standard deviation. Diploids ndt80Δ/ ndt80Δ equipped with lacZ reporter system were grown to saturation, cells were transferred into 1% potassium acetate and incubated for 16 hours at 30˚C. Cells were lysed and β-galactosidase was assayed. Experiments were repeated ten times; (b) LexA-BD-Irr1ΔC435, LexABD-Irr1-1ΔC435 and Rec8-Gal4-AD proteins are synthesized in the cell and have predicted molecular masses. For each sample, equal number of cells was subjected to trichloroacetic acid treatment and detected by Western blotting using antibodies anti-Gal4-AD or anti-LexABD. Loading control: the membrane stained with Ponceau S to visualize proteins. (-) control cells transformed with plasmids without insert. Strains expressing Irr1 ΔC435 or Irr1-1ΔC435 are denoted Irr1 and Irr1-1, respectively.
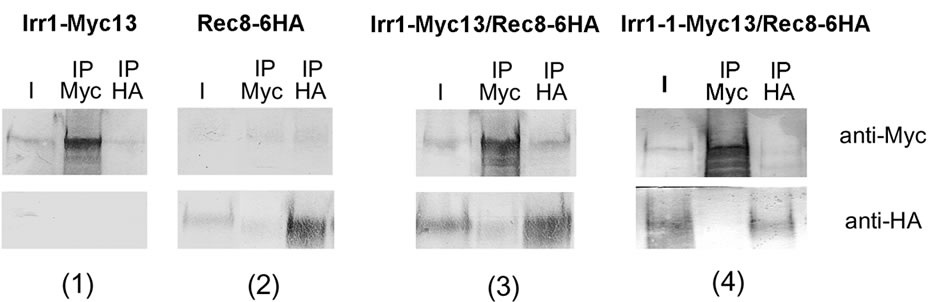
Figure 2. Immunoprecipitation confirms Irr1p-Rec8p interaction in cells arrested in early meiosis. Both copies of IRR1 were tagged with MYC and both copies of REC8 were tagged with HA. To study interaction between Irr1-1p and Rec8p, Irr1-1 allele was introduced on a centromeric plasmid. After immunoprecipitation using anti-Myc (IP Myc) or anti-HA (IP HA) antibodies, samples were separated by SDS-PAGE. Irr1p-Myc13 and Irr1-1pMyc13 were revealed using anti-Myc antibodies, Rec8p-6HA was detected with anti-HA antibodies. Panels (1) and (2) control immunoprecipitation of extracts from cells bearing only IRR1-MYC13 or REC8-6HA, respectively. Panel (3) immunoprecipitate from cells expressing Irr1p-Myc13 and Rec8pHA, panel (4) immunoprecipitate from cells expressing Irr1p-1-Myc13 and Rec8p-HA. I-input.
early meiosis Irr1p interacts with Rec8p and that F658 of Irr1p is important for this interaction. However, it has to be noticed that such interaction can be indirect and another protein(s) may act as a bridge between Irr1p and Rec8p.
To verify the two-hybrid data a co-immunoprecipitation study of whole-length Irr1p and Rec8p was performed. We constructed strain ndt80Δ/ndt80Δ bearing both copies of IRR1 tagged with MYC13 and both copies of REC8 tagged with HA6. In this strain induced into meiosis Irr1p co-precipitated with Rec8p (Figure 2, panel 3) which is consistent with the Irr1p-Rec8p interaction in early meiosis measured in the two-hybrid assay. It has to be noticed that only a fraction of Rec8p co-precipitates with Irr1p which can result from experimental conditions but also suggests that not a whole pool of Irr1p and Rec8p is involved in the interaction.
In parallel, we introduced into a ndt80Δ/ndt80Δ diploid the mutated irr1-1-MYC13 construct on a plasmid. This diploid was irr1Δ/IRR1 and had both copies of REC8 tagged with HA6. In extracts derived from this strain Irr1-1-Myc and Rec8-HA proteins do not co-precipitate (Panel 4). In that case no detectable co-immunoprecipitation of Irr1-1-Myc and Rec8-HA was observed. This is consistent with the finding described above that the F658G substitution abolishes the Irr1p-Rec8p interaction (Figure 2, panel 4, lane 3).
4. CONCLUSION
Rec8p has been found to participate in several cellular processes via its independent features [3]. Our findings indicate that participation of Rec8p in early meiosis very likely involves an interaction with Irr1p/Scc3p, although only an in vitro study using purified proteins can give a definite proof that the Irr1p-Rec8p interaction is direct. Whether the observed interaction is also part of the other processes in which Rec8p participates requires further studies.
5. ACKNOWLEDGEMENTS
We thank Drs. Kim Nasmyth, Nancy Kleckner, Frank Uhlmann, Scott Keeney and Matthew J. Neale for strains and protocols and Jan Fronk for critical reading of the manuscript. This work was supported in part by the Ministry of Science and Higher Education, grants 0038/P01/ 2009/36 and 4568/B/P01/2010/38.
REFERENCES
- Marston, A.L. and Amon, A. (2004) Meiosis: Cell-cycle controls shuffle and deal. Nature Reviews Molecular Cell Biology, 5, 983-997. doi:10.1038/nrm1526
- Lee, B. and Amon, A. (2001) Meiosis: How to create a specialized cell cycle. Current Opinion in Cell Biology, 13, 770-777. doi:10.1016/S0955-0674(00)00282-9
- Brar, G.A., Hochwagen, A. and Ee, L.S. et al. (2009) The multiple roles of cohesin in meiotic chromosome morphogenesis and pairing. Molecular Biology of the Cell, 20, 1030-1047. doi:10.1091/mbc.E08-06-0637
- Klein, F., Mahr, P., Galova, M., et al. (1999) A central role for cohesins in sister chromatid cohesion, formation of axial elements, and recombination during yeast meiosis. Cell, 98, 91-103. doi:10.1016/S0092-8674(00)80609-1
- Nasmyth, K. (2001) Disseminating the genome: joining, resolving, and separating sister chromatids during mitosis and meiosis. Annual Review of Genetics, 35, 673-745. doi:10.1146/annurev.genet.35.102401.091334
- Buonomo, S.B., Clyne, R.K., Fuchs, J., et al. (2000) Disjunction of homologous chromosomes in meiosis I depends on proteolytic cleavage of the meiotic cohesin Rec8 by separin. Cell, 103, 387-398. doi:10.1016/S0092-8674(00)00131-8
- Shonn, M.A., McCarroll, R. and Murray, A.W. (2000) Requirement of the spindle checkpoint for proper chromosome segregation in budding yeast meiosis. Science, 289, 300-303. doi:10.1126/science.289.5477.300
- Uhlmann, F. (2003) Separase regulation during mitosis. Biochemical Society Symposia, 70, 243-251.
- Katis, V.L., Lipp, J.J., Imre, R., et al. (2010) Rec8 phosphorylation by casein kinase 1 and Cdc7-Dbf4 kinase regulates cohesin cleavage by separase during meiosis. Developmental Cell, 18, 397-409. doi:10.1016/j.devcel.2010.01.014
- Brar, G.A., Kiburz, B.M., Zhang, Y., et al. (2006) Rec8 phosphorylation and recombination promote the step-wise loss of cohesin in meiosis. Nature, 441, 532-536. doi:10.1038/nature04794
- Cena A., Kozłowska E., Płochocka D., et al. (2008) The F658G substitution in Saccharomyces cerevisiae cohesin Irr1/Scc3 is semi-dominant in the diploid and disturbs mitosis, meiosis and the cell cycle. European Journal of Cell Biology, 87, 8318-8344. doi:10.1016/j.ejcb.2008.05.002
- Chu, S. and Herskowitz, I. (1998) Gametogenesis in yeast is regulated by a transcriptional cascade dependent on Ndt80. Molecular Cell, 1, 685-696. doi:10.1016/S1097-2765(00)80068-4
- Rose, M., Winston, F. and Hieter, P. (1990) Methods in yeast genetics. A cold spring harbor laboratory course. Cold Spring Harbor Laboratory Press, New York.
- Arora, C., Kee, K., Maleki, S., et al. (2004) Antiviral protein Ski8 is a direct partner of Spo11 in meiotic DNA break formation, independent of its cytoplasmic role in RNA metabolism. Molecular Cell, 13, 549-559. doi:10.1016/S1097-2765(04)00063-2
- Buonomo, S.B., Rabitsch, K.P., Fuchs, J., et al. (2003) Division of the nucleolus and its release of CDC14 during anaphase of meiosis I depends on separase, SPO12, and SLK19. Developmental Cell, 4, 727-739. doi:10.1016/S1534-5807(03)00129-1
- Gruber, S., Haering, C.H. and Nasmyth, K. (2003) Chromosomal cohesin forms a ring. Cell, 112, 765-777. doi:10.1016/S0092-8674(03)00162-4
- Bialkowska, A. and Kurlandzka, A. (2002) Proteins interacting with Lin 1p, a putative link between chromosome segregation, mRNA splicing and DNA replication in Saccharomyces cerevisiae. Yeast, 19, 1323-1333. doi:10.1002/yea.919
- Hunter, S., Apweiler, R., Attwood, T.K., et al. (2009) InterPro: The integrative protein signature database. Nucleic Acids Research, 37, D224-D228. doi:10.1093/nar/gkn785
- Finn, R.D., Mistry, J., Tate, J., et al. (2010) The Pfam protein families database. Nucleic Acids Research, 38, D211-D222. doi:10.1093/nar/gkp985
NOTES
*Corresponding author.

