Open Journal of Inorganic Chemistry
Vol.04 No.04(2014), Article ID:49965,12 pages
10.4236/ojic.2014.44007
Photocleavage Properties and DNA-Binding Studies of Oxovanadium Complexes Incorporating 2-(2-Hydroxybenzylideneamino) Isoindoline-1,3-Dione and Fluoro-Phenanthroline Derivatives
Peng Ying1, Pengfei Zeng1, Jiazheng Lu1*, Ning Yang2, Hongyuan Chen3*
1School of Pharmacy, Guangdong Pharmaceutical University, Guangzhou, China
2Department of Clinical pathology, the First Affiliated Hospital, Guangdong Pharmaceutical University, Guangzhou, China
3Department of Pathogen Biology and Immunology, School of Basic Course, Guangdong Pharmaceutical University, Guangzhou, China
Email: *lujia6812@163.com, *hychen1208@126.com
Copyright © 2014 by authors and Scientific Research Publishing Inc.
This work is licensed under the Creative Commons Attribution International License (CC BY).
http://creativecommons.org/licenses/by/4.0/



Received 22 July 2014; revised 20 August 2014; accepted 15 September 2014
ABSTRACT
Three mononuclear oxovanadium complexes [VO(Hbid)(CF3PIP)] (1) (Hbid=(E)-2-(2-hydroxyben- zylideneamino)isoindoline-1,3-dione, CF3PIP=2-(2-trifluoromethyl phenyl)imidazole[4,5-f][1,10] phenanthroline), [VO(Hbid)(m-CF3PIP)]; (2) (m-CF3PIP=2-(3-trifluoromethyl phenyl)imidazole [4, 5-f][1,10]phenanthroline) and [VO(Hbid)(p-CF3PIP)]; (3) (p-CF3PIP=2-(4-trifluoromethyl phenyl) imidazole[4,5-f][1,10]phenanthroline) have been synthesized and characterized by elemental analysis, IR, molar conductance, ES-MS and 1H NMR. The DNA-binding properties of these complexes were studied by using UV-Vis absorption titration, fluorescence spectra, viscosity measurements and thermal denaturation studies. The results show that 1, 2 and 3 interact with calf thymus DNA (CT-DNA) by intercalation modes and the magnitude of their intrinsic binding constants (Kb values) follows the order: 2 < 1 < 3. Furthermore, their photocleavage properties with pBR322 plasmid DNA were investigated by agarose gel electrophoresis experiments. The DNA cleavage capacity of complex 3 is also stronger than that of 1 and 2.
Keywords:
Oxovanadium Complexes, Synthesis, DNA-Binding, Photocleavage

1. Introduction
The development of transition metal complexes which are actually to be utilized in a wide range of biochemistry and medicine as well as therapeutic medication against tumor cells has achieved fantastic step forward in past decades [1] -[3] . Oxovanadium complexes as one group of transition metal compounds have aroused great interest among biochemists and pharmacologists owing to diverse biologic and pharmacological activities such as antiproliferative [4] , antibacterial [5] , biocatalytic oxidation [6] [7] , insulin-enhancing effects [8] , apoptosis-in- ducing activity [9] [10] , potential capabilities as DNA structural probes and DNA dependent electron transfer likewise [11] [12] . In recent years, mixed-ligand oxovanadium complexes have demonstrated both in vitro antibacterial and antifungal properties [13] as well as intense interactions with DNA [14] [15] .
Complexes with Schiff base Hbid, where Hbid=(E)-2-(2-hydroxybenzylideneamino) isoindoline-1,3-dione, displayed chemical nuclease activity by partial intercalation thus ability to inhibit the growth of both Gram-pos. and Gram-neg. bacteria [16] [17] . Imidazole[4, 5-f][1, 10] phenanthroline derivatives available to inhibit c-myc gene expression in A549 cells via NF-kB pathway exhibit certain activities towards inhibiting tumor cells to some extend [18] -[20] . Although certain attention has been paid to the research on oxovanadium complexes as potential DNA intercalator, photocleavage and anticancer agents [21] -[24] , the oxovanadium compounds with imidazole[4, 5-f][1, 10] fluoro-phenanthroline derivatives has rarely been reported yet [22] [25] .
In the present article, three mononuclear oxovanadium complexes [VO(Hbid)(CF3PIP)] (1), [VO(Hbid)(m- CF3PIP)] (2), [VO(Hbid)(p-CF3PIP)] (3), (Hbid=(E)-2-(2-hydroxybenzylideneamino)isoindoline-1,3-dione, CF3PIP=-(2-trifluoromethyl phenyl)imidazole [4,5-f][1,10]phenanthroline, m-CF3PIP=2-(3-trifluoromethyl phenyl)imidazole[4,5-f][1,10]phenanthroline, p-CF3PIP=2-(4-trifluoromethyl phenyl)imidazole[4,5-f][1,10] phenanthroline) have been synthesized and characterized by elemental analysis, IR, molar conductance, ES-MS and 1H NMR. The DNA-binding properties of these three complexes were well studied by UV-Vis titration, fluorescence spectra, viscosity measurements and thermal denaturation studies. Photocleavage reactions with pBR322 supercolied plasmid DNA were investigated by agarose gel electrophoresis experiments. The compounds employed in this work are shown in Scheme 1.
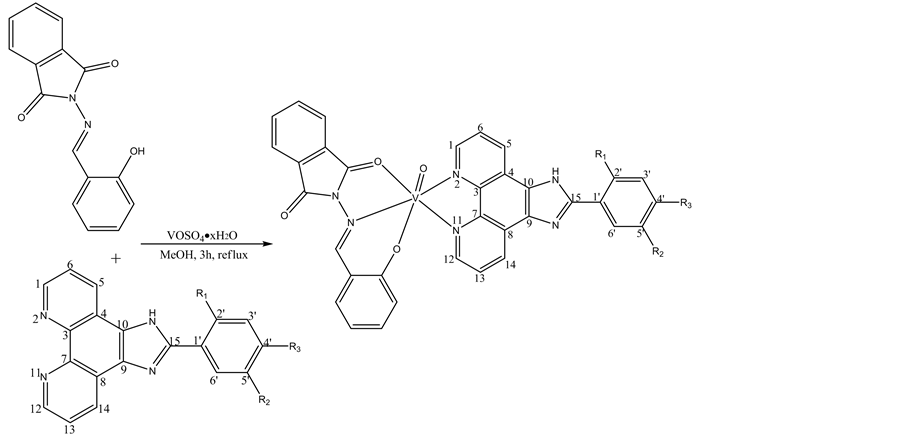
Scheme 1. Structure of VO(Hbid)(CF3PIP) (1), R1=CF3, R2=H, R3=H; VO(Hbid) (m-CF3- PIP) (2), R1=H, R2=CF3, R3=H; VO(Hbid) (p-CF3PIP) (3), R1=H, R2=H, R3=CF3.
2. Experimental
2.1. Materials and Methods
All chemicals used in the synthesis and physical measurements were analytical reagent grade or biochemical quality and without further purification unless otherwise specified. 1,10-Phenanthroline was obtained from Guangzhou Chemical Reagent Factory. VOSO4∙xH2O were purchased from Alfa Aesar, N-Aminophthalimide were purchased from TCI, CT-DNA and pBR322 supercoiled plasmid DNA were obtained from Sigma. Tris buffer 1 (Tris=tris(hydroxyl-methyl)aminomethane) containing 5 m∙mol∙L−1 Tris-HCl and 50 m∙mol∙L −1 NaCl (pH = 7.2) was used for absorption titration, fluorescence emission and viscosity measurements. Tris buffer 2 containing 50 m∙mol∙L−1 Tris-HCl and 18 m∙mol∙L−1 NaCl (pH = 7.2) was used for the gel electrophoresis experiments. A phosphoric acid buffer containing 1.5 m∙mol∙L−1 Na2HPO4, 0.5 m∙mol∙L−1 NaH2PO4 and 0.25 m∙ mol∙L−1 Na2H2EDTA (H4EDTA=N,N'-ethane-1,2-diylbis[N-(carboxymethyl) glycine]) (pH = 7.0) was used for thermal denaturation. A solution of CT-DNA in buffer 1 gave a ratio of UV absorbance at 260 and 280 nm of 1.8 - 1.9:1, indicating that the DNA was sufficiently free of protein [26] [27] . The DNA concentration per nucleotide was determined by absorption spectroscopy using the molar absorption coefficient (6600 M−1∙cm−1) at 260 nm [28] . The complexes were dissolved in DMSO and diluted with buffer solution to the required concentrations prior to use. Hbid, (E)-2-(2-hydroxybenzylideneamino) isoindoline-1,3-dione were synthesized according to a previously reported procedure [16] .
2.2. Physical Measurements
Microanalysis (C, H, and N) was carried out with a PerkinElmer 240Q elemental analyzer. Electrospray mass spectra (ES-MS) were recorded on an LCQ system (Finnigan MAT, USA) using methanol as mobile phase. 1H NMR spectra were recorded on a Varian-500 spectrometer. All chemical shifts are given relative to tetramethylsilane (TMS). Infrared spectra were recorded on a Bomem FTIR model MB102 instrument using KBr pellets method. UV-Vis spectra were recorded on a Shimadzu UV-3101 PC spectrophotometer at room temperature. Emission spectra were recorded on a Perkin-Elmer Lambda 55 spectrofluorophotometer. Molar conductivities in DMF (1 m∙mol∙L−1) solution at room temperature were measured using a DDS-307 digital direct reading conductivity meter.
2.3 Synthesis and Characterization
2.3.1. Synthesis of Phenanthroline-Based Ligands: CF3PIP, m-CF3PIP and p-CF3PIP
CF3PIP was synthesized through a modification of a previously reported procedure [29] -[31] . A mixture of 2- (Trifluoromethyl)benzaldehyde (0.69 mL, 5 m∙mol) and ammonium acetate (7.70 g, 0.1 mol) was added into a stirring solution of 1,10-Phenanthroline-5,6-dione (0.99 g, 5 m∙mol) in 60 mL of glacial acetic acid, and the mixture was continuously stirred at 60˚C for 6 hours. Then the deep red solution was cooled to room temperature and diluted with 100 mL distilled water. A grayish yellow precipitate was obtained by neutralization with ammonium hydroxide. Then the mixture was filtered and washed with water for three times. The crude solid power was purified by chromatography over 60 - 80 mesh SiO2 using absolute ethanol as eluent. The solvent was removed and the products were collected, and dried in vacuo. m-CF3PIP and p-CF3PIP were prepared by a similar procedure with the compound CF3PIP, with 3-(Trifluoromethyl)benzaldehyde (0.69 mL, 5 m∙mol) and 4-(Trifluoromethyl)benzaldehyde (0.69 ml, 5 m∙mol) in place of 2-(Trifluoromethyl)benzaldehyde (0.69 mL, 5 m∙mol) respectively, and a pink-yellow precipitate were obtained.
CF3PIP: Yield: 61%. Anal. Found: C, 65.48; H, 3.14; N, 15.35; Calcd for C20H11F3N4: C, 65.93; H, 3.04; N, 15.28. 1H NMR (DMSO-d6, 500 MHz) δ: 9.03 (s, 2H, J = 8.7 Hz, ArH), 9.02 (s, 2H, J = 8.6 Hz, ArH), 8.47 (m, 1H, J = 8.3 Hz, ArH), 8.45 (m, 1H, J = 8.2 Hz, ArH), 7.96 (m, 1H, J = 8.1 Hz, ArH), 7.82 (br, 3H, J = 7.6 Hz, ArH), 3.32 (s, 1H, -NH). 13C NMR (DMSO-d6, 500 MHz) δ: 148.9 C(15), 148.0 C(1,12), 143.7 C(5,14), 133.8 C(3ꞌ,5ꞌ), 129.7 C(1ꞌ), 129.5 C(3), 129.1 C(7), 126.6 C(2ꞌ,6ꞌ), 125.9 C(4ꞌ), 125.8 C(-CF3), 124.8 C(8), 124.7 C(4), 123.4 C(9), 123.3 C(10), 122.6 C(6,13). ES-MS: (CH3OH): m/z 365.0 ([M+H]+), 387.0 ([M+Na]+). IR (KBr disk): ν (cm−1) = 3406.8 (s, N-H), 3078.2 (m, C-H), 1606.7 (s), 1551.8 (m), 1483.0 (s, C=C), 1315.3 (vs, C-F). Conductance (Ω−1∙cm2∙mol−1): 9.2.
m-CF3PIP: Yield: 58%. Anal. Found: C, 65.67; H, 3.13; N, 15.42; Calcd for C20H11F3N4: C, 65.93; H, 3.04; N, 15.38. 1H NMR (DMSO-d6, 500 MHz) δ: 8.99 (m, 2H, J = 8.8 Hz, ArH), 8.83 (d, 2H, J = 8.7 Hz, ArH), 8.56 (m, 2H, J = 8.2 Hz, ArH), 7.78 (m, 1H, J = 8.0 Hz, ArH), 7.75 (m, 3H, J = 7.5 Hz, ArH). 3.38 (s, 1H, -NH). 13C NMR (DMSO-d6, 500 MHz) δ: 148.2 C(15), 148.0 C(1,12), 143.6 C(5,14), 135.3 C(3ꞌ,5ꞌ), 132.5 C(1ꞌ), 130.3 C(3), 129.8 C(7), 128.2 C(2ꞌ,6ꞌ), 127.9 C(4ꞌ), 127.0 C(-CF3), 126.8 C(8), 126.7 C(4), 126.0 C(9), 124.8 C(10), 123.8 C(6,13). ES-MS: (CH3OH): m/z 365.0 ([M+H]+), 387.0 ([M+Na]+). IR (KBr disk): ν (cm−1) = 3241.6 (s, N-H), 3065.8, (m, C-H), 1607.2 (m), 1566.7 (s), 1548.5 (m, C=C), 1312.2 (vs, C-F). Conductance (Ω−1∙ cm2∙mol−1): 9.6.
p-CF3PIP: Yield: 66%. Anal. Found: C, 65.78; H, 3.24; N, 15.32; Calcd for C20H11F3N4: C, 65.93; H, 3.04; N, 15.38. 1H NMR (DMSO-d6, 500 MHz) δ: 9.07 (d, 2H, J = 8.6 Hz, ArH), 8.84 (d, 2H, J = 8.4 Hz, ArH), 8.04 (d, 1H, J = 8.1 Hz, ArH), 7.97 (d, 1H, J = 7.8 Hz, ArH), 7.91 (d, 1H, J = 7.9 Hz, ArH), 7.83 (t, 3H, J = 7.7 Hz, ArH). 3.40 (s, 1H, -NH). 13C NMR (DMSO-d6, 500 MHz) δ: 149.3 C(15), 147.7 C(1,12), 143.6 C(5,14), 131.6 C(3ꞌ,5ꞌ), 131.4 C(1ꞌ), 130.1 C(3), 129.9 C(7), 129.6 C(2ꞌ,6ꞌ), 128.4 C(4ꞌ), 125.5 C(-CF3), 125.2 C(8), 123.2 C(4ꞌ), 123.1 C(4), 122.4 C(9), 122.3 C(10), 121.6 C(6,13). ES-MS: (CH3OH): m/z 365.0 ([M+H]+), 387.0 ([M+Na]+). IR (KBr disk): ν (cm−1) = 3371.2 (s, N-H), 3065.8 (w, C-H), 1608.2 (s), 1565.5 (m, C=C), 1326.5 (vs, C-F). Conductance (Ω−1∙cm2∙mol−1): 8.6.
2.3.2. Synthesis of [VO (Hbid)(CF3PIP)] (1)
[VO (Hbid) (CF3PIP)] was prepared by the synthetic steps similar to a previously reported method [24] [32] [33] . A mixture of CF3PIP (0.364 g, 1 m∙mol) and Hbid (0.266 g, 1 m∙mol) in absolute CH3OH (60 mL) was heated at 80˚C under argon for 2 h. After dissolution, an aqueous solution (5 mL) of 1.0 m mol (0.217 g) quantity of VOSO4∙xH2O was added dropwise to this mixture, and a breen precipitate was formed. The soild power was purified by recrystallization in methanol, and dried in vacuo. Yield: 36%. Anal. Found: C, 60.28; H, 2.92; N, 12.09; Calcd for C35H20F3N6O4V: C, 60.35; H, 2.89; N, 12.07. 1H-NMR (DMSO-d6, 500 MHz) δ: 9.03 (s, 1H, CH=N), 8.68 (dd, 2H, ArH, J = 8.5 Hz), 8.60 (d, 2H, ArH, J = 8.4 Hz), 8.49 (br, 6H, ArH, J = 8.2 Hz), 8.26 (d, 1H, J = 8.4 Hz), 8.14 (d, 1H, ArH, J = 7.8 Hz), 7.82 (dd, 2H, ArH, J = 7.2 Hz), 7.14 (d, 2H, ArH, J = 6.8 Hz), 6.52 (dd, 2H, ArH, J = 7.8 Hz), 5.02 (s, 1H, -NH). ES-MS (CH3OH): m/z: 695.1 ([M-H]−), 727.1 ([M+CH3OH− H]−). IR (KBr disk): ν (cm−1) = 3425.1 (N-H), 2973.3, 3077.0 (m, C=C-H), 1724.2 (vs, HC=N), 1608.0 (vs, C=O), 1314.1 (s, -CF3), 974.1 (s, V=O), 730.8 (m, V-N). Conductance (Ω−1∙cm2∙mol−1): 10.7.
2.3.3. Synthesis of [VO (Hbid)(m-CF3PIP)] (2)
This compound was synthesized by a similar procedure as for the complex 1, with m-CF3PIP (0.364 g, 1 m∙mol) in place of CF3PIP. Yield: 43%. Anal. Found: C, 60.36; H, 2.91; N, 12.06; Calcd for C35H20F3N6O4V: C, 60.35; H, 2.89; N, 12.07. 1H NMR (DMSO-d6, 500 MHz) δ: 8.87 (s, 1H, CH=N), 8.59 (dd, 2H, ArH, J = 8.5 Hz), 8.44 (d, 2H, ArH, J = 8.5 Hz), 8.36 (br, 6H, ArH, J = 8.3 Hz), 8.27 (d, 1H, J = 8.2 Hz), 8.08 (d, 1H, ArH, J = 7.7 Hz), 7.72 (dd, 2H, ArH, J = 7.8 Hz), 7.34 (d, 2H, ArH, J = 6.6 Hz), 6.48 (dd, 2H, ArH, J = 7.6 Hz), 5.20 (s, 1H, -NH). ES-MS (CH3OH): m/z: 695.1 ([M-H]−), 727.1 ([M + CH3OH − H]−). IR (KBr disk): ν (cm−1)=3428.3 (N-H), 3027.0 (m, C=C-H), 1721.9 (s, HC=N), 1607.5 (vs, C=O), 1312.2 (s, -CF3), 968.8 (s, V=O), 713.7 (m, V-N). Conductance (Ω−1∙cm2∙mol−1): 9.7.
2.3.4. Synthesis of [VO (Hbid) (p-CF3PIP)] (3)
This complex was prepared using a similar procedure described for complex 1. Yield: 36%. Anal. Found: C, 60.28; H, 2.93; N, 12.06; Calcd for C35H20F3N6O4V: C, 60.35; H, 2.89; N, 12.07. 1H-NMR (DMSO-d6, 500MHz) δ: 9.12 (s, 1H, CH=N), 8.86 (d, 2H, ArH, J = 8.5 Hz), 8.63 (d, 2H, ArH, J = 8.4 Hz), 8.44 (br, 6H, ArH, J = 8.2 Hz), 8.26 (d, 1H, J = 8.4 Hz), 8.12 (d, 1H, ArH, J = 7.8 Hz), 7.80 (d, 2H, ArH, J = 7.2 Hz), 7.34 (d, 2H, ArH, J = 6.8 Hz), 6.62 (t, 2H, ArH, J = 7.8 Hz), 5.12 (s, 1H, -NH). ES-MS (CH3OH): m/z: 695.1 ([M-H]−), 727.1 ([M + CH3OH-H]−). IR (KBr disk): ν (cm−1) = 3401.6 (N-H), 2982.9, 3077.2 (m, C=C-H), 1725.7 (vs, HC=N), 1609.4 (vs, C=O), 1314.5 (s, -CF3), 956.6 (s, V=O), 730.9 (m, V-N). Conductance (Ω−1∙cm2∙mol−1): 11.8.
2.4. DNA-Binding and Cleavage
Absorption titration of the oxovanadium complexes in buffer 1 were performed using a fixed concentration of the oxovanadium complexes (20 μ∙mol∙L−1) to which the DNA stock solutions were added. The oxovanadium-DNA solutions were incubated at room temperature for 3 mins before the absorption spectra were recorded. In order to further elucidate the binding strength of the complexes, the intrinsic binding constant Kb with CT-DNA was obtained by monitoring the change in the absorbance of the ligand transfer band with increasing amounts of DNA. Kb was then calculated using the following equation [25] [34] [35] :
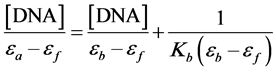
where [DNA] is the concentration of DNA in the base pairs, and εa – εf and εb refer to the corresponding apparent absorption coefficient Aobsd/[Vanadium], the extinction coefficient for the free oxovanadium complexes and the extinction coefficient for the oxovanadium complexes in the fully bound form, respectively. In plots of [DNA]/(εa – εf) versus [DNA], Kb is obtained by the ratio of the slope to the intercept.
Viscosity measurements were carried out with an Ubbelohde viscometer maintained at a constant temperature of (28.0 ± 0.1) C in a thermostatic bath. Flow time was measured with a digital stopwatch, and each sample was measured five times to obtain the average flow time. Date are presented as (h/h0)1/3 versus binding ratio [36] [37] , where h is the viscosity of DNA in the presence of complexes while h0 is the viscosity of DNA alone.
Thermal denaturation studies were carried out with Shimadzu UV-3101 PC spectrophotometer equipped with a Peltier temperature-controlling programmer (±0.1˚C). The melting curves were obtained by measuring the absorbance at 260 nm for solutions of CT-DNA (80 μ∙mol∙L−1) in the absence and presence of oxovanadium complex [20 μ∙mol∙L−1] as a function of the temperature was scanned from 50˚C to 90˚C at a speed of 5˚C min−1. The melting temperature (Tm) was taken as the mid-point of the hyperchromic transition.
The photoinduced cleavage activity of supercoiled pBR322 DNA by the oxovanadium complexes was studied by performing agarose gel electrophoresis experiment, pBR322 DNA (0.1 μg) was treated with the complexes in buffer 2 in different concentrations, and the solutions were incubated at 37˚C in the incubator for 1 h. The samples were analyzed by electrophoresis for 2 h at 90 V in tris-boric buffer containing 0.8% agarose gel. The gel was stained with 0.1 μL/ml DuRed nucleic acid gel stain and photographed under UV light on an Alpha Innotech IS-5500 fluorescence chemiluminescence and visible imaging system [38] [39] .
3. Results and Discussion
3.1. Synthesis and Characterization
The Schiff base ligand was prepared by a reaction of salicylaldehyde with N-Aminophthalimide in the appropriate mole ratios using absolute ethanol as solvent [28] . The oxovanadium complexes 1-3 were obtained by refluxing of corresponding Schiff bases, VOSO4∙xH2O and phenanthroline-based ligand in absolute methanol. The desired complexes were purified by recrystallization.
According to the IR spectra of phenanthroline-based ligands show absorption at ca. 3400 cm−1 (imidazole N-H), 3065 - 3078 cm−1 (C=C-H), 1483-1608 cm−1 (C=C), 1312-1326 cm−1 (C-F), respectively. In parallel, IR spectra absorption of complexes 1-3 observed at 3401-3425 cm−1 for (imidazole N-H), 2982 - 3077 cm−1 for (C=C-H), 1607 - 1725 cm−1 for (C=O) and (-CH=N), 1312 - 1314 cm−1 for (-CF3), 713 - 730 cm−1 for (V-N), respectively. The strong (VO) band at 956 - 974 cm−1 could be cleared identified for the formulation of the complex.
Electronic spectra of complexes 1-3 show an intense band at ca. 272 nm assignable to π―π* transition of aromatic rings of phenanthroline-based ligands [15] [25] . Bands in the UV-vis region (320 - 350 nm) are assignable to intraligand transitions of the Schiff base. Five- and six-coordination complexes of oxovanadium (IV) are usually square pyramidal/trigonal bipyramidal and distorted octahedral, respectively [40] [41] . From the above spectral data, the Schiff base bonded through the phenolate oxygen, imine nitrogen, and carbonyl oxygen leaving the carbonyl as a pendant group. This implies that these complexes bear the central V (IV) atom in a square- pyramidal geometry.
In the 1H NMR spectra of phenanthroline-based ligands showed peaks of aromatic hydrogen (ArH), imidazole secondary amine (NH) proton. However, in the 1H NMR spectra of complexes, the peaks of imine (CH=N) proton were observed, which affirmed that the Schiff base ligand was coordinated to vanadium.
In the ES-MS spectra of CF3PIP, m-CF3PIP and p-CF3PIP showed peaks at m/z 365 ([M+H]+) and 387 ([M + Na]+). The molecular ion peaks of three complexes at m/z 695 ([M-H]−) and 727 ([M+CH3OH-H]−). Elemental analysis, ES-MS, IR and 1H NMR data of all the compounds are in good agreement with the expected structures.
The molar conductance values of phenanthroline-based ligands and complexes 1-3 in DMF (1 m∙mol∙L−1) are shown in Table 1. The conductive properties indicates that both ligands L1-L3 and oxovanadium complexes 1-3 are nonelectrolytes.
Table 1. Molar conductivity data of oxovanadium complexes and the ligands.
3.2. DNA-Binding Studies
3.2.1. Electronic Absorption Titration
For the purpose of evaluating the capacity of oxovanadium complexes interacting with CT-DNA, the electronic spectra of complexes 1, 2 and 3 in the absence and presence of increasing amounts of CT-DNA are shown in Figure 1. As the CT-DNA concentration is increased, obvious hypochromism as well as a certain degree of bathochromism for the three oxovanadium complexes can be observed in the ultraviolet titration process at ca. 272 nm, which is attributed to π→π*. The hypochromism and bathochromism shift for 1 at 272 nm are 28.72% and 2 nm, respectively. In contrast, the hypochromism and bathochromism of complexes 2 and 3 are 20.93%, 1 nm and 30.36%, 2.5 nm, respectively. According to previously reported results [14] [23] -[25] , the UV-Vis spectral characteristics suggest that complexes 1-3 interact with CT-DNA most likely through a mode that involves a stacking interaction between the aromatic chromophore and the base pairs of DNA.
In order to elucidate the binding strengths of these complexes, the intrinsic binding constant Kb were calculated by monitoring the changes of absorbance in the ligand transfer bands with increasing amounts of CT-DNA. The Kb values of complexes 1-3 were calculated as 7.40 × 104 (mol∙L−1)−1, 5.42 × 104 (mol∙L−1)−1 and 1.25 × 105 (mol∙L−1)−1, respectively, indicating that they strongly interact with DNA by intercalation. And the DNA-bind- ing activities of these three complexes are higher than that of reported mononuclear oxovanadium complexes VO(SAA)(phen) (4.50 × 104 M−1) and VO(MOSAA) (phen) (2.95 × 104 M−1) [25] , under our experimental conditions. The value of Kb increases in the order 2 < 1 < 3, this may due to the substituents of trifluoromethyl phenyl introduced on the phenanthroline ring of complexes 1-3, which may intercalate into base pairs of CT-DNA in complexes 1-3 will be altered, resulting in higher Kb. Meanwhile, the presence of an -CF3 group on the aromatic moieties, which acts as an electron-drawing substituent on the intercalative ligand and thus gives stronger stacking interactions with DNA for complexes 1-3. And the electronic effects of substituted group at the different locations introduced on the aromatic ring of phenanthroline-based ligands may make differences in the DNA-binding affinities. The results also indicates an -CF3 group substituted on the para-position of the aromatic exhibits stronger DNA-binding affinity than substituted on the ortho-position and meta-position.
3.2.2. Fluorescence Spectroscopic Studies
The interaction of the complexes (20 μ∙mol∙L−1) with CT-DNA was investigated using fluorescence emission titration experiment in the Tris buffer 1 at room temperature. The emission spectra of complexes 1, 2 and 3 in the absence and presence of CT-DNA exhibit luminescence in Tris buffer, with a maxima appearing at 442 nm, 443 nm and 440 nm respectively, which are shown in Figure 2. With increasing concentrations of CT-DNA, the emission intensities of complexes 1-3 grow to around by about 0.26, 0.19, and 0.36 times larger than those in the absence of CT-DNA and saturates at a [DNA]/[V] ratio of 20:1. The enhancement of emission intensity is indicative of binding of the complexes to the hydrophobic pocket of DNA, since the hydrophobic environment inside the DNA helix reduces the accessibility of water molecules to the complex and the complex mobility is restricted at the binding site, leading to decrease of the vibrational modes of relaxation [42] [43] . In addition, it also shows that 3 interacts with DNA more strongly than both 1 and 2, consistent with the UV-vis absorption titration spectra results.
3.2.3. Viscosity Measurements
To further clarify the nature of the binding interaction between both oxovanadium complexes and DNA in the absence of X-ray structural data or NMR spectra, viscosity measurements that are regarded as the least controversial and the most rigorous means of testing the binding model of DNA in solution [44] . It is well-known that a classical intercalation model leads to an apparent increase in viscosity of DNA solution due to base-pairs are pushed apart and hence an increase in overall DNA length [45] . In contrast, a partial, non-classical intercalation of compounds could bend (or kink) the DNA helix and reduce its effective length and, concomitantly, its viscosity [46] .
The effects of complexes 1, 2 and 3 on the viscosity of CT-DNA are shown in Figure 3. As can be seen in Figure 3, with amounts of the complexes increasing, the relative viscosity of DNA increases continuously to some extend. The results thus provide strong evidence for the interaction of complexes 1, 2 and 3 with CT-DNA by intercalation modes. Moreover, the increasing degree of viscosity is 2 < 1 < 3, which is consistent with electronic absorption spectra data, supporting the conclusion that complexes 1-3 interact with DNA by intercalation modes.
3.2.4. Thermal Denaturation Studies
DNA melting experiments are generally applied to investigate the extent of intercalation, which were carried out by monitoring the intensity of DNA bases at 260 nm at different temperatures, both in the absence and presence of oxovanadium complexes. Thermal behaviors of DNA in the presence of compounds can give insight into their conformational changes when the temperature is raised and offer information about the interaction strength of complexes with DNA. As a rule, with the temperature in the solution rising, the double-stranded DNA will gradually dissociate to single strands and generate a hyperchromic effect on absorption spectra of DNA bases. Thus, the melting temperature Tm, which is defined as the temperature where half of the total base pairs are un- bounded, is usually introduced. Generally, Tm will increase considerable when intercalative binding occurs, since intercalation of the complexes into DNA base pairs causes stabilization of base stacking and hence raises the melting temperature of the double-stranded DNA [11] [24] [25] .


Figure 1. Absorption spectral of the complexes 1 (a), 2 (b) and 3 (c) in Tris-HCl buffer 1 upon increasing amounts of CT-DNA. [V] = 20 μM, [DNA] = (0 - 100) μM. Arrow shows the decreasing absorbance upon increasing CT-DNA concentrations. Insert: Plots of [DNA]/(εa − εf) versus [DNA] for the titration of [V] with CT-DNA.


Figure 2. Emission spectra of 1 (a), 2 (b) and 3 (c) in Tris-HCl buffer 1 in the absence and presence of CT-DNA. [V] = 20 μ∙mol∙L−1. Arrows show the increasing intensity with increasing concentrations of DNA.
Figure 3. Effect of increasing amounts of complexes 1 (p), 2 (▼) and 3 (<) on the relative viscosity of CT-DNA at (28.0 ± 0.1)˚C, [DNA] = 0.2 m∙mol∙L−1.
The melting curves of CT-DNA in the absence and presence of complexes 1-3 are shown in Figure 4; As shown in Figure 4, The Tm of CT-DNA in the absence of the complexes is 62.2˚C ± 0.2˚C, meanwhile, when the complexes at a concentration ratio [V]/[DNA] of 1:4, we can observe that Tm values in the presence of the complexes 1, 2 and 3 are 68.3˚C ± 0.2˚C, 65.3˚C ± 0.2˚C, 69.2˚C ± 0.2˚C, respectively. A comparison of the ΔTm values of the complexes (6.1˚C, 3.1˚C and 7.0˚C for 1, 2, and 3, respectively) was agreement with those classical intercalators [47] [48] , which provided strong evidence for their binding with DNA by intercalation.
The experimental data reveals that complexes in collaboration with phenanthroline-substituted auxiliary li-
Figure 4. Thermal denaturation of CT-DNA in the absence and presence of complexes 1, 2 and 3, [V] = 20 μM, [DNA] = 80 μM.
gands exhibit appreciable DNA intercalative activities, in addition, complex 3 showed a larger ΔTm, indicating that 3 exhibits stronger DNA-binding affinity when interacting with DNA, in conformity with the electronic absorption titration.
3.3. Photocleavage Studies
The interaction mode between these oxovanadium complexes and plasmid DNA were further investigated by agarose gel electrophoresis experiments. When circular plasmid DNA is subject to electrophoresis, relatively fast migration will be observed for the intact supercoiled form (form Ι). If scission occurs on one strand (nicking), the supercoil will relax to generate a slower moving open circular form (form II). If both strands are cleaved, a linear form (form III) migrating between form I and II will be generated [49] [50] .
As is shown both in Figure 5 and Figure 6, it is clear to figure out the cleavage activity of plasmid pBR322 DNA after incubation with different concentrations of oxovanadium complexes at 37˚C for 1 h at dark condition. The nicked form II can barely observed for H2O2 alone in the absence of the complexes (lane 2 in Figure 5 and Figure 6, respectively). Nevertheless, in the presence of complexes 1, 2 and 3 with 30 m∙mol∙L−1 H2O2, apparent cleavage of pBR322 DNA observed from the formation of form II, and cleaving activity tends to increase gradually with the concentration of the complexes increase from 15 μ∙mol∙L−1 to 60 μ∙mol∙L−1 (lane 3, 4 and 5 in Figure 5 and Figure 6). Furthermore, under the same conditions with lane 4, addition of 0.02 mol∙L−1 L-Histi- dine works as a singlet oxygen quencher (lane 6, lane 11 in Figure 5 and Figure 6, respectively) slightly inhibits the cleavage of DNA. Meanwhile, no cleavage of plasmid DNA occurred in the presence of hydroxyl radical scavenger DMSO (lane 7, lane 12 in Figure 5 and Figure 6, respectively), suggesting that ·OH radicals is likely to be the reactive species on account of the oxidation of VO2+ in the presence of H2O2 for the cleavage reaction [51] . Under comparative experimental conditions, a linear form (form III) (lane 4, lane 5 in Figure 6) can be observed when the concentration of complex 3 reaches to 30 μ∙mol∙L−1 and 60 μ∙mol∙L−1, respectively, implying the photocleavage ability of complex 3 is stronger than that of complexes 1 and 2, follows the order of 3 > 1 > 2. The results are actually consistent with the degree of their intrinsic binding constant (Kb), indicating that the electrochemical characteristics of phenanroline-based auxiliary ligands as well as the existence of substituted strong electronic-withdrawing group (-CF3) at the different locations introduced on the aromatic ring of phenanroline-based ligands plays an important role in the DNA-binding affinity.
4. Conclusion
In this paper, three novel Schiff base oxovanadium complexes, [VO(Hbid)(CF3PIP)] (1), [VO(Hbid)(m-CF3PIP)] (2), [VO(Hbid)(p-CF3PIP)] (3) have been synthesized and characterized by EA, ES-MS, IR, 1H NMR and molar conductance. Their DNA-binding activities with CT-DNA were evaluated using UV-Vis titration, viscosity measurements and thermal denaturation. These three oxovanadium complexes interact with CT-DNA by intercalation modes. Their photocleavage activities of supercoiled plasmid DNA were investigated in the presence of H2O2 probably via generating hydroxyl radical, the results also show efficiently oxidative cleavage activities, in
Figure 5. Cleavage of pBR322 DNA by complexes 1 and 2 (15 - 60 μ∙mol∙L−1) in the absence and presence of H2O2 (30 m∙mol∙L−1) in buffer 2 (pH 7.2).lane 1, DNA control; lane 2, DNA + H2O2; lane 3, DNA + 1 (15 μ∙mol∙L−1) + H2O2; lane 4, DNA + 1 (30 μ∙mol∙L−1) + H2O2; lane 5, DNA + 1 (60 μ∙mol∙L−1) + H2O2; lane 6, DNA + 1 (30 μ∙mol∙ L−1) + H2O2+ L-Histidine (0.02 mol∙L−1); lane 7, DNA + 1 (30 μ∙mol∙L−1) + H2O2+ DMSO (2 μ∙L); lane 8, DNA + 2 (15 μ∙mol∙L−1) + H2O2; lane 9, DNA + 2 (30 μ∙mol∙L−1) + H2O2; lane 10, DNA + 2 (60 μ∙mol∙L−1) + H2O2; lane 11, DNA + 2 (30 μ∙mol∙L−1) + H2O2+ L-Histidine (0.02 mol∙L−1); lane 12, DNA + 2 (30 μ∙mol∙L−1) + H2O2+ DMSO (2 μ∙L).
Figure 6. Cleavage of pBR322 DNA by complex 3 (15 - 60 μ∙mol∙L−1) in the absence and presence of H2O2 (30 m∙ mol∙L−1) in buffer 2 (pH 7.2). Lane 1, DNA control; lane 2, DNA + H2O2; lane 3, DNA + 3 (15 μ∙mol∙L−1) + H2O2; lane 4, DNA + 3 (30 μ∙mol∙L−1) + H2O2; lane 5, DNA + 3 (60 μ∙mol∙L−1) + H2O2; lane 6, DNA + 3 (30 μ∙mol∙L−1) + H2O2 + L-Histidine (0.02 mol∙L−1); lane 7, DNA + 3 (30 μ∙mol∙L−1) + H2O2+ DMSO (2 μ∙L).
conformity with DNA-binding behaviors of the complexes. In addition, complex 3 exhibits a higher DNA- binding affinity in contrast to complexes 1 and 2, indicating that the electrochemical characteristics of phenanroline plane and the existence of substituted strong electronic-withdrawing group (-CF3) at the different locations introduced on the aromatic ring of phenanroline-based ligands plays an important role in the DNA-binding affinity.
Acknowledgements
We gratefully acknowledge financial support for this work by the Science and technology Research Project of Guangdong Province (No.2012B031800431), P. R. China and Cultivation of Natural Science Joint Fund of the First Affiliated Hospital and the Scientific & Technical Department of Guangdong Pharmaceutical University (No. 2014-36).
References
- Chen, Z.F., Liu, Y.C., Huang, K.B. and Liang, H. (2013) Alkaloid-Metal Based Anticancer Agents. Current Topics in Medicinal Chemistry (Sharjah, United Arab Emirates), 13, 2104-2115. http://dx.doi.org/10.2174/15680266113139990146
- Rafique, S., Idrees, M., Nasim, A., Akbar, H. and Athar, A. (2010) Transition Metal Complexes as Potential Therapeutic Agents. Biotechnology and Molecular Biology Reviews, 5, 38-45.
- Gu, J. and Codd, R. (2012) Copper(II)-Based Metal Affinity Chromatography for the Isolation of the Anticancer Agent Bleomycin from Streptomyces verticillus Culture. Journal of Inorganic Biochemistry, 115, 198-203. http://dx.doi.org/10.1016/j.jinorgbio.2012.01.015
- Leon, I. E., Porro, V., Di, V.A.L., Naso, L.G., Williams, P.A.M., Bollati-Fogolin, M. and Etcheverry, S.B. (2014) Antiproliferative and Apoptosis-Inducing Activity of an Oxidovanadium(IV) Complex with the Flavonoid Silibinin against Osteosarcoma Cells. Journal of Biological Inorganic Chemistry, 19, 59-74. http://dx.doi.org/10.1007/s00775-013-1061-x
- Anupama, B. and Gyana Kumari, C. (2011) Synthesis, Characterization, DNA Binding and Antimicrobial Activity of 4-Amino Antipyrine Schiff Base Metal Complexes. Research Journal of Pharmaceutical, Biological and Chemical Sciences, 2, 140-159.
- Walmsley, R.S., Litwinski, C., Antunes, E., Hlangothi, P., Hosten, E., McCleland, C., Nyokong, T., Torto, N. and Tshentu, Z.R. (2013) Oxovanadium(IV)-Containing Poly(Styrene-co-4'-Ethenyl-2-Hydroxyphenylimidazole) Electrospun Nanofibers for the Catalytic Oxidation of Thioanisole. Journal of Molecular Catalysis A: Chemical, 379, 94-102. http://dx.doi.org/10.1016/j.molcata.2013.07.018
- Parihar, S., Jadeja, R.N. and Gupta, V.K. (2014) Novel Oxovanadium(IV) Complexes with 4-Acyl Pyrazolone Ligands: Synthesis, Crystal Structure and catalytic Activity towards the Oxidation of Benzylic Alcohols. RSC Advances, 4, 10295-10302. http://dx.doi.org/10.1039/c3ra46896h
- Sanna, D., Micera, G. and Garribba, E. (2013) Interaction of Insulin-Enhancing Vanadium Compounds with Human Serum holo-Transferrin. Inorganic Chemistry, 52, 11975-11985. http://dx.doi.org/10.1021/ic401716x
- Yamaguchi, T., Watanabe, S., Matsumura, Y., Tokuoka, Y. and Yokoyama, A. (2012) Synthesis of Oxovanadium Complexes and Their Apoptosis-Inducing Activity in Leukemia Cells. Chemical & Pharmaceutical Bulletin, 60, 508- 512. http://dx.doi.org/10.1248/cpb.60.508
- Yamaguchi, T., Watanabe, S., Matsumura, Y., Tokuoka, Y. and Yokoyama, A. (2012) Oxovanadium Complexes with Quinoline and Pyridinone Ligands: Syntheses of the Complexes and Effect of Alkyl Chains on Their Apoptosis-Inducing Activity in Leukemia Cells. Bioorganic & Medicinal Chemistry, 20, 3058-3064. http://dx.doi.org/10.1016/j.bmc.2012.02.063
- Liao, X.W., Pan, W.J., He, R.W., Guo, H.W., Ying, P. and Lu, J.Z. (2014) Unsymmetrical Oxovanadium Complexes Derived from Salicylaldehyde and Phenanthroline: Synthesis, DNA Interactions and Antitumor Activities. Chemical Biology & Drug Design, 83, 367-378. http://dx.doi.org/10.1111/cbdd.12248
- Zhai, S.S., Guo, Q., Dong, J.F., Xu, T. and Li, L.Z. (2014) An Oxovanadium(IV) Complex of an L-Serine Schiff Base and 1,10-Phenanthroline: Synthesis, Crystal Structure, and DNA and Albumin-Binding Properties. Transition Metal Chemistry, 39, 271-280. http://dx.doi.org/10.1007/s11243-014-9800-6
- Sumrra, S.H. and Chohan, Z.H. (2013) In Vitro Antibacterial, Antifungal and Cytotoxic Activities of Some Triazole Schiff Bases and Their Oxovanadium(IV) Complexes. Journal of Enzyme Inhibition and Medicinal Chemistry, 28, 1291-1299. http://dx.doi.org/10.3109/14756366.2012.735666
- Liao, X.W., Lu, J.Z., Ying, P., Zhao, P., Bai, Y.L., Li, W.J. and Liu, M.P. (2013) DNA Binding, Antitumor Activities and Hydroxyl Radical Scavenging Properties of Novel Oxovanadium(IV) Complexes with Substituted Isoniazid. Journal of Biological Inorganic Chemistry, 18, 975-984. http://dx.doi.org/10.1007/s00775-013-1046-9
- Jing, B.Q., Dong, J.F., Li, J.H., Xu, T. and Li, L.Z. (2013) Synthesis, Crystal Structure and DNA Interaction of an Oxovanadium(IV) Complex Containing L-Valine Schiff Base and 1,10-Phenanthroline. Journal of Coordination Chemistry, 66, 520-529. http://dx.doi.org/10.1080/00958972.2013.763939
- Shiva, K.L. and Revanasiddappa, H.D. (2011) Synthesis, Characterization, Antimicrobial, DNA Binding and Oxidative Cleavage Activities of Cu(II) and Co(II) Complexes with 2-(2-Hydroxybenzylideneamino) Isoindoline-1, 3-Dione. Journal of Coordination Chemistry, 64, 699-714. http://dx.doi.org/10.1080/00958972.2011.554544
- Sugihara, H. and Hiratani, K. (1996) 1,10-Phenanthroline Derivatives as Ionophores for Alkali Metal Ions. Coordination Chemistry Reviews, 148, 285-299. http://dx.doi.org/10.1016/0010-8545(95)01178-1
- Sun, D.D., Wang, W.Z., Mao, J.W., Mei, W.J. and Liu, J. (2012) Imidazo [4,5-f][1,10] Phenanthroline Derivatives as Inhibitor of C-Myc Gene Expression in A549 Cells via NF-κB Pathway. Bioorganic & Medicinal Chemistry Letters, 22, 102-105. http://dx.doi.org/10.1016/j.bmcl.2011.11.063
- Gomathi Sankareswari, V., Vinod, D., Mahalakshmi, A., Alamelu, M., Kumaresan, G., Ramaraj, R. and Rajagopal, S. (2014) Interaction of Oxovanadium(IV)-Salphen Complexes with Bovine Serum Albumin and Their Cytotoxicity against Cancer. Dalton Transactions, 43, 3260-3272. http://dx.doi.org/10.1039/c3dt52505h
- Naso, L.G., Lezama, L., Rojo, T., Etcheverry, S.B., Valcarcel, M., Roura, M., Salado, C., Ferrer, E.G. and Williams, P.A.M. (2013) Biological Evaluation of Morin and Its New Oxovanadium(IV) Complex as Antio-Xidant and Specific Anti-Cancer Agents. Chemico-Biological Interactions, 206, 289-301. http://dx.doi.org/10.1016/j.cbi.2013.10.006
- Zhang, Y.L., Wang, X.S., Fang, W., Cai, X.Y., Li, H.Z., Mao, J.W., Jin, X.B., Bai, Y.L. and Lu, J.Z. (2013) In Vitro Study of the Cytotoxicities of Two Mixed-Ligand Oxovanadium Complexes on Human Hepatoma Cells. Pharmazie, 68, 827-834.
- El-Asmy, A.A., Al-Gammal, O.A., Saad, D.A. and Ghazy, S.E. (2009) Synthesis, Characterization, Molecular Modeling and Eukaryotic DNA Degradation of 1-(3,4-Dihydroxybenzylidene) Thiosemicarbazide Complexes. Journal of Molecular Structure, 934, 9-22. http://dx.doi.org/10.1016/j.molstruc.2009.05.039
- Du, Y.f., Lu, J.Z., Guo, H.W., Jiang, J., Chao, P., Chen, F. and Pan, J.M. (2010) DNA Binding and Photocleavage Properties of Two Mixed-Ligand Oxovanadium Complexes. Transition Metal Chemistry, 35, 859-864. http://dx.doi.org/10.1007/s11243-010-9404-8
- Lu, J.Z., Guo, H.W., Zeng, X.D., Zhang, Y.L., Zhao, P., Jiang, J. and Zang, L.Q. (2012) Synthesis and Characterization of Unsymmetrical Oxidovanadium Complexes: DNA-Binding, Cleavage Studies and Antitumor Activities. Journal of Inorganic Biochemistry, 112, 39-48. http://dx.doi.org/10.1016/j.jinorgbio.2012.02.034
- Guo, H.W., Lu, J.Z., Ruan, Z.G., Zhang, Y.L., Liu, Y.J. Zang, L.Q. Jiang, J. and Huang, J.W. (2012) Synthesis, DNA-Binding, Cytotoxicity, and Cleavage Studies of Unsymmetrical Oxovanadium Complexes. Journal of Coordination Chemistry, 65, 191-204. http://dx.doi.org/10.1080/00958972.2011.645204
- Jeslin Kanaga Inba, P., Annaraj, B., Thalamuthu, S. and Neelakantan, M.A. (2013) Cu(II), Ni(II), and Zn(II) Complexes of Salan-Type Ligand Containing Ester Groups: Synthesis, Characterization, Electrochemical Properties and in Vitro Biological Activities. Bioinorganic Chemistry and Applications, 2013, 439848-439860. http://dx.doi.org/10.1155/2013/439848
- Liu, Y.J., Li, Z.Z., Liang, Z.H., Yao, J.H. and Huang, H.L. (2011) Cytotoxicity, Apoptosis, Cellular Uptake, Cell Cycle Arrest, Photocleavage and Antioxidant Activity of 1,10-Phenanthroline Ruthenium(II) Complexes. DNA and Cell Biology, 30, 839-848. http://dx.doi.org/10.1089/dna.2011.1243
- Reichmann, M.E., Rice, C.A., Thomas, C.A. and Doty, P. (1954) A Further Examination of the Molecular Weight and Size of Deoxypentose Nucleic Acid. Journal of the American Chemical Society, 76, 3047-3053. http://dx.doi.org/10.1021/ja01640a067
- Wu, Q., Fan, C.D., Chen, T.F., Liu, C.R., Mei, W.J., Chen, S.D., Wang, B.G., Chen, Y.Y. and Zheng, W.J. (2013) Microwave-Assisted Synthesis of Arene Ruthenium(II) Complexes That Induce S-Phase Arrest in Cancer Cells by DNA Damage-Mediated p53 Phosphorylation. European Journal of Medicinal Chemistry, 63, 57-63. http://dx.doi.org/10.1016/j.ejmech.2013.01.037
- Cai, Z.B., Liu, L.F., Hong, Y.Q. and Zhou, M. (2013) Synthesis, Characterization and Nonlinear Optical Responses of Nickel(II) Complexes with Phenanthroline-Based Ligands. Journal of Coordination Chemistry, 66, 2388-2397. http://dx.doi.org/10.1080/00958972.2013.805322
- Tang, T.T., Tang, G.D., Kou, S.S., Zhao, J.Y., Culnane, L.F. and Zhang, Y. (2014) Experimental and DFT Studies on the Vibrational and Electronic Spectra of 2-(1H-Imidazo [4,5-f][1,10] Phenanthrolin-2-yl) phenol. Spectrochimica Acta, Part A: Molecular and Biomolecular Spectroscopy, 117, 144-151. http://dx.doi.org/10.1016/j.saa.2013.07.108
- Raman, N. and Selvan, A. (2011) DNA Interaction, Enhanced DNA Photocleavage, Electrochemistry, Thermal Investigation and Biopotential Properties of New Mixed-Ligand Complexes of Cu(II)/VO(IV) Based on Schiff Bases. Journal of Molecular Structure, 985, 173-183. http://dx.doi.org/10.1016/j.molstruc.2010.10.038
- Hazari, P.P., Pandey, A.K., Chaturvedi, S., Tiwari, A.K., Chandna, S., Dwarakanath, B.S. and Mishra, A.k. (2012) Synthesis of Oxovanadium(IV) Schiff Base Complexes Derived from C-Substituted Diamines and Pyridoxal-5-Phosphate as Antitumor Agents. Chemical Biology & Drug Design, 79, 223-234. http://dx.doi.org/10.1111/j.1747-0285.2011.01265.x
- Li, C.Z., Wu, J.G., Wang, L.F., Min, R., Jia, N.Y. and Guo, J. (1999) Synthesis, Characterization and Antitumor Activity of Copper(II) Complex with Nicotinamido-4-Bis(2-Chloroethyl) Aminobenzaldimine. Journal of Inorganic Biochemistry, 73, 195-202. http://dx.doi.org/10.1016/S0162-0134(99)00005-7
- Zhao, P., Xu, L.C., Huang, J.W., Fu, B., Yu, H.C., Zhang, W.H., Chen, J., Yao, J.H. and Ji, L.N. (2008) DNA-Binding and Photocleavage Properties of Cationic Porphyrin-Anthraquinone Hybrids with Different Lengths of Links. Bioorganic Chemistry, 36, 278-287. http://dx.doi.org/10.1016/j.bioorg.2008.08.002
- Tabatabaee, M., Bordbar, M., Ghassemzadeh, M., Tahrir, M., Mehri Lighvan, Z. and Neumuller, B. (2013) Two New Neutral Copper(II) Complexes with Dipicolinic Acid and 3-Amino-1H-1,2,4-Triazole Formed under Different Reaction Conditions: Synthesis, Characterization, Molecular Structures and DNA-Binding Studies. European Journal of Medicinal Chemistry, 70, 364-371. http://dx.doi.org/10.1016/j.ejmech.2013.10.009
- Gabra, N.M., Mustafa, B., Kumar, Y.P., Shobha Devi, C., Shilpa, M., Reddy, K.L. and Satyanarayana, S. (2014) DNA-Binding and Cleavage, Cytotoxicity Properties of Ru(II) Complexes with 2-(4’-Chloro-Phenyl) Imidazo[4,5-f][1,10] Phenanthroline, Ligand and Their “Light Switch” on-off Effect. Medicinal Chemistry Research, 23, 224-235. http://dx.doi.org/10.1007/s00044-013-0617-1
- Hazari, P.P., Pandey, A.K., Chaturvedi, S., Tiwari, A.K., Chandna, S., Dwarakanath, B.S. and Mishra, A.K. (2012) Synthesis of Oxovanadium(IV) Schiff Base Complexes Derived from C-Substituted Diamines and Pyridoxal-5-Phos- phate as Antitumor Agents. Chemical Biology & Drug Design, 79, 223-234. http://dx.doi.org/10.1111/j.1747-0285.2011.01265.x
- Zhou, C.Q., Lin, Y.L., Chen, J.X. and Chen, W.H. (2012) Synergetic DNA-Cleaving Activities of the Metal Complexes of a Polyether-Tethered Pyrrole-Polyamide Dimer. Chemistry & biodiversity, 9, 1125-1132. http://dx.doi.org/10.1002/cbdv.201100183
- Chen, C., Bai, F.Y., Zhang, R., Song, G., Shan, H., Xing, N. and Xing, Y.H. (2013) Synthesis, Structure and Catalytic Bromination of Supramolecular Oxovanadium Complexes Containing Oxalate. Journal of Coordination Chemistry, 66, 671-688. http://dx.doi.org/10.1080/00958972.2013.765561
- Bian, L., Li, L.Z., Zhang, Q.F., Dong, J.F., Xu, T., Li, J.H. and Kong, J.M. (2012) Synthesis, Crystal Structures, DNA Binding and Cleavage Studies of Two Oxovanadium(IV) Complexes of 1,10-Phenanthroline and Schiff Bases Derived from Tryptophan. Transition Metal Chemistry, 37, 783-790. http://dx.doi.org/10.1007/s11243-012-9653-9
- Lewis, N.A., Liu, F., Seymour, L., Magnusen, A. and Erves, T.R. (2012) Synthesis, Characterisation and Preliminary In Vitro Studies of Vanadium(IV) Complexes with a Schiff Base and Thiosemicarbazones as Mixed Ligands. European Journal of Inorganic Chemistry, 2012, 664-677. http://dx.doi.org/10.1002/ejic.201100898
- Benitez, J., Becco, L., Correia, I., Leal, S.M. and Guiset, H. (2011) Vanadium Polypyridyl Compounds as Potential Antiparasitic and Antitumoral Agents: New Achievements. Journal of Inorganic Biochemistry, 105, 303-312. http://dx.doi.org/10.1016/j.jinorgbio.2010.11.001
- Mezo, G., Herenyi, L., Habdas, J., Majer, Z., Mysliwa-Kurdziel, B., Toth, K. and Csik, G. (2011) Syntheses and DNA Binding of New Cationic Porphyrin-Tetrapeptide Conjugates. Biophysical Chemistry, 155, 36-44. http://dx.doi.org/10.1016/j.bpc.2011.02.007
- Jang, Y.J., Yeo, G.Y., Park, B. and Kim, S.K. (2011) The Relationship between the Structures of Periphery Ligands and the DNA Binding Mode of [Ru(II) (1,10-Phenanthroline) (L1L2) Dipyrido[3,2-a:2’,3’-c] Phenazine] n+ (L1 = Cl or Pyridine and L2 = Pyridine, n = 1, 2). Biophysical Chemistry, 158, 38-45. http://dx.doi.org/10.1016/j.bpc.2011.04.014
- Tan, D.J., He, Y., Qiu, L. and Zhang, Z.T. (2007) Studies on the Interactions of Sodium Chrysin-6-Sulfonate and Its Dinuclear Zinc, Tetra-Nuclear Calcium Complex with DNA. Chemical Research and Application, 19, 502-506.
- McCrate, A., Carlone, M., Nielsen, M. and Swavey, S. (2010) Anaerobic Photocleavage of Supercoiled DNA by a Ruthenium(II) Substituted Fluorinated Porphyrin. Inorganic Chemistry Communications, 13, 537-539. http://dx.doi.org/10.1016/j.inoche.2010.01.031
- Surendra, D.C., Siva Kumar, V., John Venison, S., Vetha Potheher, I. and Rajalaxmi Subahashini, D. (2013) Synthesis, Structural Characterisation, Bio-Potential Efficiency and DNA Cleavage Applications of Nicotinamide Metal Complexes. Journal of Molecular Structure, 1040, 192-205. http://dx.doi.org/10.1016/j.molstruc.2013.02.019
- Li, H.J., Bo, H.B., Wang, J.Q., Shao, H.W. and Huang, S.L. (2011) Separation of Supercoiled from Open Circular Forms of Plasmid DNA and Biological Activity Detection. Cytotechnology, 63, 7-12. http://dx.doi.org/10.1007/s10616-010-9322-9
- Nejedly, K., Sykorova, E., Diekmann, S. and Paleeek, E. (1998) Analysis of a Curved DNA Constructed from Alternating dAn:dTn-Tracts in Linear and Supercoiled Form by High Resolution Chemical Probing. Biophysical Chemistry, 73, 205-216. http://dx.doi.org/10.1016/S0301-4622(98)00145-8
- Benitez, J., Guggeri, L., Tomaz, I., Pessoa, J.C., Moreno, V., Lorenzo, J., Aviles, F.X., Garat, B. and Gambino, D. (2009) A Novel Vanadyl Complex with a Polypyridyl DNA Intercalator as Ligand: A Potential Anti-Protozoa and Anti-Tumor Agent. Journal of Inorganic Biochemistry, 103, 1386-1394. http://dx.doi.org/10.1016/j.jinorgbio.2009.07.013
NOTES

*Corresponding authors.






