 World Journal of AIDS, 2011, 1, 208-218 doi:10.4236/wja.2011.14030 Published Online December 2011 (http://www.SciRP.org/journal/wja) Copyright © 2011 SciRes. WJA Structural Analysis of Predicted HIV-1 Secis Elements Paushali Roy*, Sayak Ganguli, Pooja Sharma, Protip Basu, Abhijit Datta DBT Centre for Bioinformatics, Presidency University, Kolkata, India. E-mail: *paushali.06@gmail.com Received August 30th, 2011; revised October 6th, 2011; accepted October 19th, 2011. ABSTRACT Incorporation of Selenocysteine into protein requires an RNA structural motif, SECIS (Selenocysteine insertion se- quence) element that, along with other factors, demarcates UGA-Sec from the UGA termination codon, for expression of Selenoproteins (in case of eukaryotes). It has been predicted that during HIV infection, several functional viral se- lenoproteins are expressed and synthesis of these viral selenoproteins deplete the selenium level of the host. It might be that even the viral genome has the SECIS elements in their Selenoprotein mRNA, and during infection, the host cellular machinery is transformed in such a way that the human Sec tRNA binds to the viral Selenoprotein mRNA, instead of binding to its own Selenoprotein mRNA, thus leading to expression of viral selenoproteins. This hypothesis was tested in this study by identifying the SECIS elements in the HIV-1 genome and further predicting their secondary and tertiary structures. We then tried to dock these tertiary structures with human Sec tRNA. Here we report putatively the presence of 3215 SECIS elements in the HIV-1 genome and that the hu man Sec tRNA sec binds to the viral SECIS elements present in the viral selenoprotein mRNA. Based on an earlier finding, it was observed that atoms of A8 and U9, which present in human Sec tRNA, are the possible key sites for binding. Keywords: Selenocystei ne , Selenium, S ECIS Element, Selenoprotein, Human Sec tRNAsec, UGA Codon 1. Introduction Selenium, an essential micronutrient, is a natural com- ponent of selenium dependent enzymes, and in most of these it occurs in the amino acid selenocysteine, that is present in the catalytic centers of the proteins [1]. These selenium dependent enzymes called selenoproteins in- clude one or more Selenocysteine residues, where sele- nium acts an antioxidant [2]. Selenium plays an impor- tant role in the proper functioning of the immune system and inhibiting the progression of HIV infection to AIDS. It is required for the activity of the enzyme glutathione peroxidase, and deficiency in selenium may cause myo- pathy, cardiomyopathy and immune dysfunction [3]. Selenoproteins such as glutathione peroxidases, thio- redoxin reductases, and iodothyronine deiodinases are involved in redox reactions [4]. At the physiological level, these enzymes are involved in diverse metabolic and physiological functions ranging from antioxidant defense to fertility, muscle development and function, thyroid hormone metabolism, and immune function [5]. Expression of the selenoproteins requires the incorpo- ration and biosynthesis of the amino acid Selenocysteine (reviewed in Atkins & Gesteland, 2000). Selenocysteine is the 21st amino acid in the genetic code and is encoded by the codon UGA that is generally a termination codon. Certain factors have been found in eukaryotes that medi- ate the biosynthesis of Selenocysteine and thus the ex- pression of selenoproteins [6,7]. One of the major factor are the SECIS elements, an RNA structural motif, that have been found in the 3’ UTR of the eukaryotic seleno- protein mRNA. The 3’ and 5’ untranslated regions of the HIV-1 ge- nome have all the RNA motifs concentrated within it, these include internal ribosome entry sites, packaging signals, pseudoknots, transfer RNA mimics, ribosomal frameshift motifs, and cis-regulatory elements [8,9]. In the human immunodeficiency virus (HIV), RNA struc- tures activate transcription, initiate reverse transcription, facilitate genomic dimerization, direct HIV packaging, manipulate reading frames, regulate RNA nuclear export, signal polyadenylation, and interact with viral and host proteins [9-13]. Most potential regulatory structures within the HIV-1 genome are uncharacterized raising the possibility of new RNA structure-mediated regulation to be identified [14].  Structural Analysis of Predicted HIV-1 Secis Elements209 It has been reported that during HIV infection the level of selenium in the host, decreases and expression of viral selenoproteins increases. Also, it has been proposed that HIV-1 may encode several selenoproteins one of which has significant sequence similarity to GPx that is a mammalian selenoprotein [15]. Selenocysteine insertion sequence (SECIS) element has not yet been identified in the HIV genome by either biologic or computational methods. Sequence analysis has identified locations in HIV-1 strain HXB2 where SECIS element could exist [16]. The aim of this study is to identify the plausible SECIS elements in the HIV-1 genome and deduce their role in the deficiency of selenium and increased expres- sion of viral selenoproteins during HIV infection. The results obtained showed that, indeed there are SECIS elements present in the HIV-1 genome and the human Sec tRNASec binds to the viral selenoprotein mRNA, wherein possibly the key residues are the atoms of A8 and U9 which are involved in stability of the binding. We hypothesize that during HIV infection when trans- lation occurs, the human Selenocysteine tRNASec binds to the viral selenoprotein mRNA that has the presence of SECIS elements. Thus Selenocysteine would get incor- porated in the growing polypeptide chain, utilizing the host’s selenium, and will lead to the expression of viral selenoproteins instead of human selenoproteins. 2. Methods We first retrieved 847 complete genome sequences of the HIV-1 genome from the NCBI database. The RNA regu- latory motifs for all these sequences were then obtained using a stacking energy thermodynamic model based on Bayesian statistics for identifying the homologs of Regu- latory RNA motifs and elements against an input mRNA sequence. The full process of a typical Bayesian analysis can be roughly described as consisting of three main steps: 1) setting up a full probability model that includes all the variables so as to capture the relation- ship among these variables; 2) summarizing the findings for particu- lar interests by appropriate posterior distributions; 3) evaluating the appropriateness of the model and suggest- ing improvements [17]. A standard procedure for carrying out step 1) is to first write down the likelihood function, i.e., the probability of the observed data given the unknowns, and multiply it by a prior distribution, i.e., a distribution for all the unob- served variables (typically unknown parameters). The joint probability is represented as joint = likelihood prior, i.e., ,pypy p where the prior distribution reveals what is known about the parameter without the knowledge of the data. Bayes- ian inference is drawn by examining the probability of all possible values of the parameter after considering the data. Accordingly, step 2) is completed by obtaining the posterior distribution: py pyp py pyp py py where the posterior distribution tells us what is known about y given knowledge of the data. Both sequence homologs and structural homologs of regulatory RNA motifs could be identified. In this work the basic focus was on the RNA structural motif named SECIS (Selenocysteine Insertion Sequence) element. Our next in-s ili c o experiment was performed on the same set of HIV-1 genome sequences to specifically identify the SECIS elements, if present, in the genome. This was done using a computational tool based on a SECIS consensus model the key feature of which is a conserved guanosine in a small apical loop of the prop- erly positioned structure [18]. Using the sequences of the SECIS elements obtained, we designed their secondary structures based on the RNA secondary structure (folding) prediction algorithm given by M.Zuker. The algorithm predicts the possible secondary structures based on minimum free energy (∆G) criterion. We arranged the secondary structures accord- ing to increasing free energies (a negative quantity), and selected the first 20 which had the least free energy val- ues. As we had hypothesized, our next experiment was to see whether the human Selenocysteine tRNA binds to the viral selenoprotein mRNA. For this the tertiary structures of the above 20 secondary structures were designed using computational tools. Human selenocysteine tRNA se- quence was obtained from NCBI and secondary structure is designed. The tertiary structure of human Selenocys- teine tRNA was obtained from PDB (PDB id 3A3A). We then tried to dock them individually, i.e. we performed twenty dockings, with the 20 tertiary structures of SECIS elements as receptor and human Selenocysteine tRNA as the ligand. In the last part of our work, we removed the residues A8 and U9 from the tertiary structure of human Seleno- cysteine tRNA and performed the dockings again. The new docking results were compared with the earlier docking results. 3. Results and Discussion The 3’ untranslated region of all the sequences showed he presence of SECIS elements (Table 1). t Copyright © 2011 SciRes. WJA  Structural Analysis of Predicted HIV-1 Secis Elements Copyright © 2011 SciRes. WJA 210 Table 1. Motifs found in 3’ UTR of HIV-1 genome sequences. Strain Secis Type-1Strain Secis Type-1 gi|217038387|gb|FJ460501.1| HIV-1 isolate HK004 from Hong Kong, complete genome ✓ gi|170878295|gb|EU541617.1| HIV-1 clone pIIIB from USA, complete genome ✓ gi|13540181|gb|AF289550.1| HIV-1 clone 96TZ-BF110 from Tanzania, complete genome ✓ gi|161334695|gb|EU220698.1| HIV-1 isolate 04CA7750 from Canada, complete genome ✓ gi|167651353|gb|EU293450.1| HIV-1 isolate 99ZALT46 from South Africa, complete genome ✓ gi|117940228|gb|DQ912823.1| HIV-1 isolate MA from Denmark, complete genome ✓ gi|213495604|gb|FJ195091.1| HIV-1 isolate BREPM1081 from Brazil, complete genome ✓ gi|168208535|gb|EU448296.1| HIV-1 strain 06FR-CRN from France, complete genome ✓ gi|212674726|gb|EU884501.1| HIV-1 isolate ES P1423 (CRF02_AG) from Spain, complete genome ✓ gi|164415926|gb|DQ020274.2| HIV-1 isolate CB134 from Cuba, complete genome ✓ gi|195409392|gb|EU697909.1| HIV-1 isolate J11456 from Saudi Arabia, complete genome ✓ gi|157885655|gb|EU031915.1| HIV-1 isolate 07MYKLD49 from Malaysia, complete genome ✓ gi|83026775|gb|DQ295192.1| HIV-1 isolate 04LSK7 from South Korea, complete genome ✓ gi|125541773|gb|EF192591.1| HIV-1 isolate CU-98-26 from Thailand, complete genome ✓ gi|197257781|gb|EU693240.1| HIV-1 isolate 06CM-BA-040 from Cameroon, complete genome ✓ gi|85035359|gb|DQ230841.1| HIV-1 isolate TW_D3 from Taiwan, complete genome ✓ gi|194500414|gb|EU861977.1| HIV-1 isolate 60000 from Italy, complete genome ✓ gi|117643970|gb|EF029069.1| HIV-1 isolate U.NL.01.H10986_C11 from Netherlands, complete genome ✓ gi|209156839|gb|FJ213780.1| HIV-1 isolate UY05_4752 from Uruguay, complete genome ✓ gi|51980229|gb|AY612637.1| HIV-1 isolate PT2695 from Portugal, complete genome ✓ gi|2944126|gb|U71182.1|HIVU71182 HIV-1 isolate RL42 from China, complete genome ✓ gi|112497950|gb|DQ676887.1| HIV-1 isolate PS4048_Day143 from Australia, complete genome ✓ gi|158967436|gb|EU110097.1| HIV-1 isolate ML1990PCR from Kenya, complete genome ✓ gi|63081177|gb|AY968312.1| HIV-1 isolate ARE195FL from Argentina, complete genome ✓ gi|6651466|gb|AF193277.1| HIV-1 isolate RU98001 from Russia, complete genome ✓ gi|18643009|gb|AY074891.1| HIV-1 isolate 00BWMO35.1 from Botswana, complete genome ✓ gi|3947925|gb|AF049337.1| HIV-1 CRF04_cpx clone 94CY032-3 from Cyprus, complete genome ✓ gi|74099684|gb|DQ083238.1| HIV-1 isolate 1579A from India, complete genome ✓ gi|62361768|gb|AY882421.1| HIV-1 isolate 9196/01 from Germany, complete genome ✓ gi|29409304|gb|AY093604.1| HIV-1 isolate 95SN7808 from Senegal, complete genome ✓ gi|18699247|gb|AF414006.1| HIV-1 isolate 98BY10443 from Belarus, complete genome ✓ gi|47118239|gb|AY536235.1| HIV-1 isolate CH12 from Chile, complete genome ✓ gi|18699185|gb|AF413987.1| HIV-1 isolate 98UA0116 from Ukraine, complete genome ✓ gi|47118229|gb|AY536236.1| HIV-1 isolate V62 from Venezuela, complete genome ✓ gi|6466838|gb|AF184155.1| HIV-1 G829 from Ghana complete genome ✓ gi|38679157|gb|AY352657.1| HIV-1 isolate UG266 from Uganda, complete genome ✓ gi|56131599|gb|AY805330.1| HIV-1 isolate HIV1084i from Zambia, complete genome ✓ gi|38679140|gb|AY352655.1| HIV-1 isolate SE9010 from Sweden, complete genome ✓ gi|6690753|gb|AF197341.1| HIV-1 isolate 90CF4071 from Central African Republic, complete genome ✓ gi|14530226|gb|AF286236.1|AF286236 HIV-1 isolate 83CD003 from Republic of the Congo, complete genome ✓ gi|17352343|gb|AY046058.1| HIV-l from Greece, complete genome ✓ gi|3779261|gb|AF064699.1|AF064699 HIV-1 isolate BFP90 from Burkina Faso, complete genome ✓ gi|13569307|gb|AF286233.1|AF286233 HIV-1 strain 98IS002 from Israel, complete genome ✓ gi|5668910|gb|AF076474.1|AF076474 HIV-1 isolate VI354 from Gabon, complete genome ✓ gi|6090965|gb|AF075703.1|AF075703 HIV-1 isolate FIN9363 subtype F1 from Finland, complete genome ✓ - - This result thus confirmed putatively, to some extent, that SECIS elements may be present in the HIV genome. Other motifs were also obtained i.e. K-Box, GY-Box, Gamma interferon activated inhibitor of Ceruloplasmin mRNA translation (GAIT element), Brd-Box, Cytoplas- mic polyadenylation element, Alcohol dehydrogenase 3’UTR down regulation control element (ADH_DRE), Mos polyadenylation response element (Mos-PRE), An- drogen receptor CU-rich element (AR_CURE) in the 3’ UTR and those in the 5’ UTR are Terminal Oligo-  Structural Analysis of Predicted HIV-1 Secis Elements211 pyrimidine Tract (TOP), Internal Ribosome Entry Site (IRES), Upstream Open Reading Frame (uORF).Exonic regulatory motifs, transcriptional regulatory motifs, miRNA target sites and RNA structural elements were also found, (see Table 2). The presence of Selenocysteine insertion sequence (SECIS) elements has been confirmed in eukaryotes (in- cluding humans). In eukaryotes, SECIS elements are required for the expression of selenoproteins. Functional selenoproteins, similar to mammalian selenoproteins, have been found in the HIV-1 genome. Based on these already proven theories it was thought that, the HIV ge- nome may contain SECIS elements and this was con- firmed by performing a search for SECIS elements on all the 847 complete genome sequences of the HIV-1 ge- nome. The number of SECIS elements obtained puta- tively was variable for each sequence. The total number of SECIS elements obtained was 3215. Since lower free energy value means a highly stable structure, so out of the 3215 structures 25 most stable predicted structures were selected (see Figure 1). The sequence of Human selenocysteine tRNA was re- trieved from PDB (>3A3A:A|PDBID|CHAIN|SEQUEN- CEGCCCGGAUGAUCCUCAGUGGUCUGGGGUGC- AGGCUUCAAACCUGUAGCUGUCUAGCGACAGA- GUGGUUCAAUUCCACCUUUCGGGCGCCA) and its corresponding secondary structure was designed by ap- plication of RNA covariance models, which are general, Table 2. Motifs found in regions other than 3’ and 5’UTR. Strain Exonic Regulatory Motifs Transcriptional Regulatory Motifs miRNA Target Sites RNA Structural Elements gi|217038387|gb|FJ460501.1| HIV-1 isolate HK004 from Hong Kong, complete genome 54 17 111 gi|13540181|gb|AF289550.1| HIV-1 clone 96TZ-BF110 from Tanzania, complete genome 53 18 89 gi|167651353|gb|EU293450.1| HIV-1 isolate 99ZALT46 from South Africa, complete genome 52 21 97 gi|213495604|gb|FJ195091.1| HIV-1 isolate BREPM1081 from Brazil, complete genome 54 19 109 1 gi|212674726|gb|EU884501.1| HIV-1 isolate ES P1423 (CRF02_AG) from Spain, complete genome 47 19 93 gi|195409392|gb|EU697909.1| HIV-1 isolate J11456 from Saudi Arabia, complete genome 42 17 104 gi|83026775|gb|DQ295192.1| HIV-1 isolate 04LSK7 from South Korea, complete genome 47 20 108 gi|197257781|gb|EU693240.1| HIV-1 isolate 06CM-BA-040 from Cameroon, complete genome 48 16 88 gi|194500414|gb|EU861977.1| HIV-1 isolate 60000 from Italy, complete genome 51 19 96 gi|209156839|gb|FJ213780.1| HIV-1 isolate UY05_4752 from Uruguay, complete genome 49 16 89 gi|2944126|gb|U71182.1|HIVU71182 HIV-1 isolate RL42 from China, complete genome 56 17 103 gi|158967436|gb|EU110097.1| HIV-1 isolate ML1990PCR from Kenya, complete genome 53 18 75 gi|6651466|gb|AF193277.1| HIV-1 isolate RU98001 from Russia, complete genome 57 16 96 gi|3947925|gb|AF049337.1| HIV-1 CRF04_cpx clone 94CY032-3 from Cyprus, complete genome 56 18 105 gi|170878295|gb|EU541617.1| HIV-1 clone pIIIB from USA, complete genome 61 21 100 gi|161334695|gb|EU220698.1| HIV-1 isolate 04CA7750 from Canada, complete genome 51 16 94 gi|117940228|gb|DQ912823.1| HIV-1 isolate MA from Denmark, complete genome 54 19 93 gi|168208535|gb|EU448296.1| HIV-1 strain 06FR-CRN from France, complete genome 52 25 88 Copyright © 2011 SciRes. WJA  Structural Analysis of Predicted HIV-1 Secis Elements 212 gi|164415926|gb|DQ020274.2| HIV-1 isolate CB134 from Cuba, complete genome 42 20 105 gi|157885655|gb|EU031915.1| HIV-1 isolate 07MYKLD49 from Malaysia, complete genome 51 17 96 gi|125541773|gb|EF192591.1| HIV-1 isolate CU-98-26 from Thailand, complete genome 50 24 95 gi|85035359|gb|DQ230841.1| HIV-1 isolate TW_D3 from Taiwan, complete genome 52 22 91 gi|117643970|gb|EF029069.1| HIV-1 isolate U.NL.01.H10986_C11 from Netherlands, complete genome 51 19 89 gi|51980229|gb|AY612637.1| HIV-1 isolate PT2695 from Portugal, complete genome 39 19 107 gi|112497950|gb|DQ676887.1| HIV-1 isolate PS4048_Day143 from Australia, complete genome 51 19 78 gi|63081177|gb|AY968312.1| HIV-1 isolate ARE195FL from Argentina, complete genome 57 24 102 gi|18643009|gb|AY074891.1| HIV-1 isolate 00BWMO35.1 from Botswana, complete genome 54 20 109 gi|23986250|gb|AY049711.1| HIV-1 isolate 01IN565.14 from India, complete genome 51 18 95 gi|46243163|gb|AY535660.1| HIV-1 isolate EE0369 from Estonia, complete genome 54 18 102 gi|62361768|gb|AY882421.1| HIV-1 isolate 9196/01 from Germany, complete genome 47 19 85 gi|18699247|gb|AF414006.1| HIV-1 isolate 98BY10443 from Belarus, complete genome 55 17 104 gi|18699185|gb|AF413987.1| HIV-1 isolate 98UA0116 from Ukraine, complete genome 47 19 99 1 gi|6466838|gb|AF184155.1| HIV-1 G829 from Ghana complete genome 47 20 87 gi|56131599|gb|AY805330.1| HIV-1 isolate HIV1084i from Zambia, complete genome 50 21 100 gi|29409304|gb|AY093604.1| HIV-1 isolate 95SN7808 from Senegal, complete genome 53 26 90 gi|47118239|gb|AY536235.1| HIV-1 isolate CH12 from Chile, complete genome 47 18 94 1 gi|47118229|gb|AY536236.1| HIV-1 isolate V62 from Venezuela, complete genome 50 17 103 gi|38679157|gb|AY352657.1| HIV-1 isolate UG266 from Uganda, complete genome 57 21 100 gi|38679131|gb|AY352654.1| HIV-1 isolate SE8646 from Sweden, complete genome 43 19 101 gi|6690753|gb|AF197341.1| HIV-1 isolate 90CF4071 from Central African Republic, complete genome 50 22 93 gi|17352343|gb|AY046058.1| HIV-l from Greece, complete genome 55 22 84 gi|14530226|gb|AF286236.1|AF286236 HIV-1 isolate 83CD003 from Republic of the Congo, complete genome 54 18 107 gi|3779261|gb|AF064699.1|AF064699 HIV-1 isolate BFP90 from Burkina Faso, complete genome 50 12 91 gi|13569307|gb|AF286233.1|AF286233 HIV-1 strain 98IS002 from Israel, complete genome 53 24 85 gi|6090965|gb|AF075703.1|AF075703 HIV-1 isolate FIN9363 subtype F1 from Finland, complete genome 45 18 99 gi|5668910|gb|AF076474.1|AF076474 HIV-1 isolate VI354 from Gabon, complete genome 50 16 94 Copyright © 2011 SciRes. WJA  Structural Analysis of Predicted HIV-1 Secis Elements Copyright © 2011 SciRes. WJA 213 Figure 1. Predicted secondary structures of SECIS elements. probabilistic secondary structure profiles based on sto- chastic context-free grammars (see Figure 2). The tertiary structures of all the 20 SECIS elements showed a similar kind of a structure, (see Figure 3). These were docked to the crystal structure of human Selenocysteine tRNA (see Figure 4). Also known is the fact that during HIV infection sele- nium pool of the host gets depleted and viral selenopro- teins increase. During translation, the tRNA binds to the mRNA (at the corresponding codon) for the expression of the protein. Human Selenocysteine tRNA (tRNASec) has an anticodon complementary to the UGA codon. If the human Selenocysteine tRNA binds to the viral mRNA that has the SECIS elements, then the viral se- lenoproteins might get expressed and this may be the probable cause of the increase in viral selenoproteins and depletion of host selenium, as it is being used up by the viral genome. So, the human Selenocysteine tRNA, in- stead of getting attached to its own selenoprotein mRNA, Figure 2. Predicted secondary structure of human seleno- ysteine tRNA. c 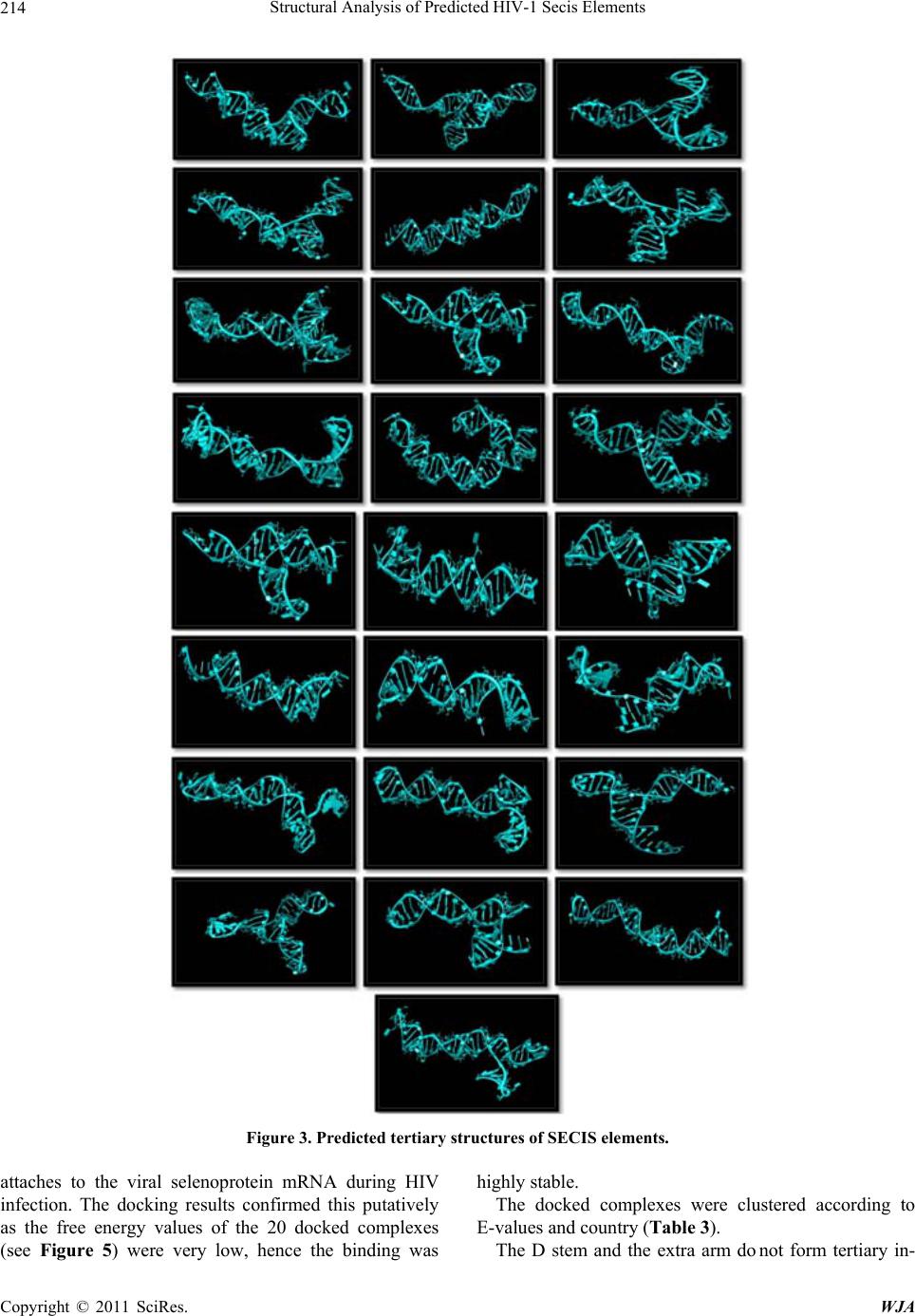 Structural Analysis of Predicted HIV-1 Secis Elements 214 Figure 3. Predicted tertiary structures of SECIS elements. attaches to the viral selenoprotein mRNA during HIV infection. The docking results confirmed this putatively as the free energy values of the 20 docked complexes (see Figure 5) were very low, hence the binding was highly stable. The docked complexes were clustered according to E-values and country (Table 3). The D stem and the extra arm do not form tertiary in- Copyright © 2011 SciRes. WJA  Structural Analysis of Predicted HIV-1 Secis Elements215 Table 3. Clustering according to E-values and country. Sequence Details E Total >gi|217038387|gb|FJ460501.1| HIV-1 isolate HK004 from Hong Kong, complete genome −26049.78 >gi|217038377|gb|FJ460500.1| HIV-1 isolate HK003 from Hong Kong, complete genome −24043.97 >gi|217038367|gb|FJ460499.1| HIV-1 isolate HK002 from Hong Kong, complete genome −27350.14 >gi|167651343|gb|EU293449.1| HIV-1 isolate 99ZALT45 from South Africa, complete genome −37608.79 >gi|149939408|gb|EF633445.1| HIV-1 isolate R1 from South Africa, complete genome −29866.54 >gi|63098379|gb|DQ011175.1| HIV-1 isolate 03ZASK005B2 from South Africa, complete genome −19193.88 >gi|63098294|gb|DQ011166.1| HIV-1 isolate 04ZASK135B1 from South Africa, complete genome −20754.28 >gi|85700643|gb|DQ351234.1| HIV-1 isolate 03ZASK233B1 from South Africa, complete genome −15853.64 >gi|85700503|gb|DQ351220.1| HIV-1 isolate 02ZAPS006MB1 from South Africa, complete genome −29420.57 >gi|68522063|gb|DQ093598.1| HIV-1 isolate 04ZAPS202B1 from South Africa, complete genome −16265.70 >gi|51572093|gb|AY703908.1| HIV-1 isolate 03ZASK040B1 from South Africa, complete genome −15903.14 >gi|46486663|gb|AY585268.1| HIV-1 isolate C.ZA.1069MB from South Africa, complete genome −29545.00 >gi|57338555|gb|AY838567.1| HIV-1 isolate 1069MB from South Africa, complete genome −16336.54 >gi|24181477|gb|AF411964.1| HIV-1 isolate 99ZACM4 from South Africa, complete genome −34332.66 >gi|29119285|gb|AY173954.1| HIV-1 isolate US3 from USA, complete genome −22281.61 >gi|37677763|gb|AY331283.1| HIV-1 isolate 1001-09 from USA, complete genome −30852.84 >gi|37677753|gb|AY331282.1| HIV-1 isolate 1001-07 from USA, complete genome −18468.71 >gi|55735993|gb|AY771593.1| HIV-1 isolate BREPM278 from Brazil, complete genome −27615.47 >gi|55735957|gb|AY771589.1| HIV-1 isolate BREPM108 from Brazil, complete genome −17908.92 >gi|157274079|gb|EF637057.1| HIV-1 isolate BREPM1023 from Brazil, complete genome −38704.25 >gi|157274021|gb|EF637051.1| HIV-1 isolate BREPM1032 from Brazil, complete genome −16073.70 >gi|157274001|gb|EF637049.1| HIV-1 isolate BREPM1035 from Brazil, complete genome −21868.21 >gi|86277616|gb|DQ358809.1| HIV-1 isolate 02BR011 from Brazil, complete genome −44666.35 >gi|221474|dbj|D10112.1|HIVCAM1 Human immunodeficiency virus 1 proviral DNA, complete genome −32795.28 Figure 4. Predicted crystal structure of human selenocys- teine tRNA. teractions in tRNASec. Rather, tRNASec has an open cavity, in place of the tertiary core of a canonical tRNA. The linker residues, A8 and U9, connecting the acceptor and D stems, are not involved in tertiary base pairing. Instead, U9 is stacked on the first base pair of the extra arm. These features might allow tRNASec to be the target of the Selenocysteine synthesis/incorporation machineries. Following this finding, the residues A8 and U9 were re- moved from the structure of human selenocysteine tRNA and this was docked with the SECIS element (one from Group-3).The atoms of the residues have been high- lighted (see Figure 6) and the bond between the SECIS lement and the residues is shown (see Figure 7). e Copyright © 2011 SciRes. WJA 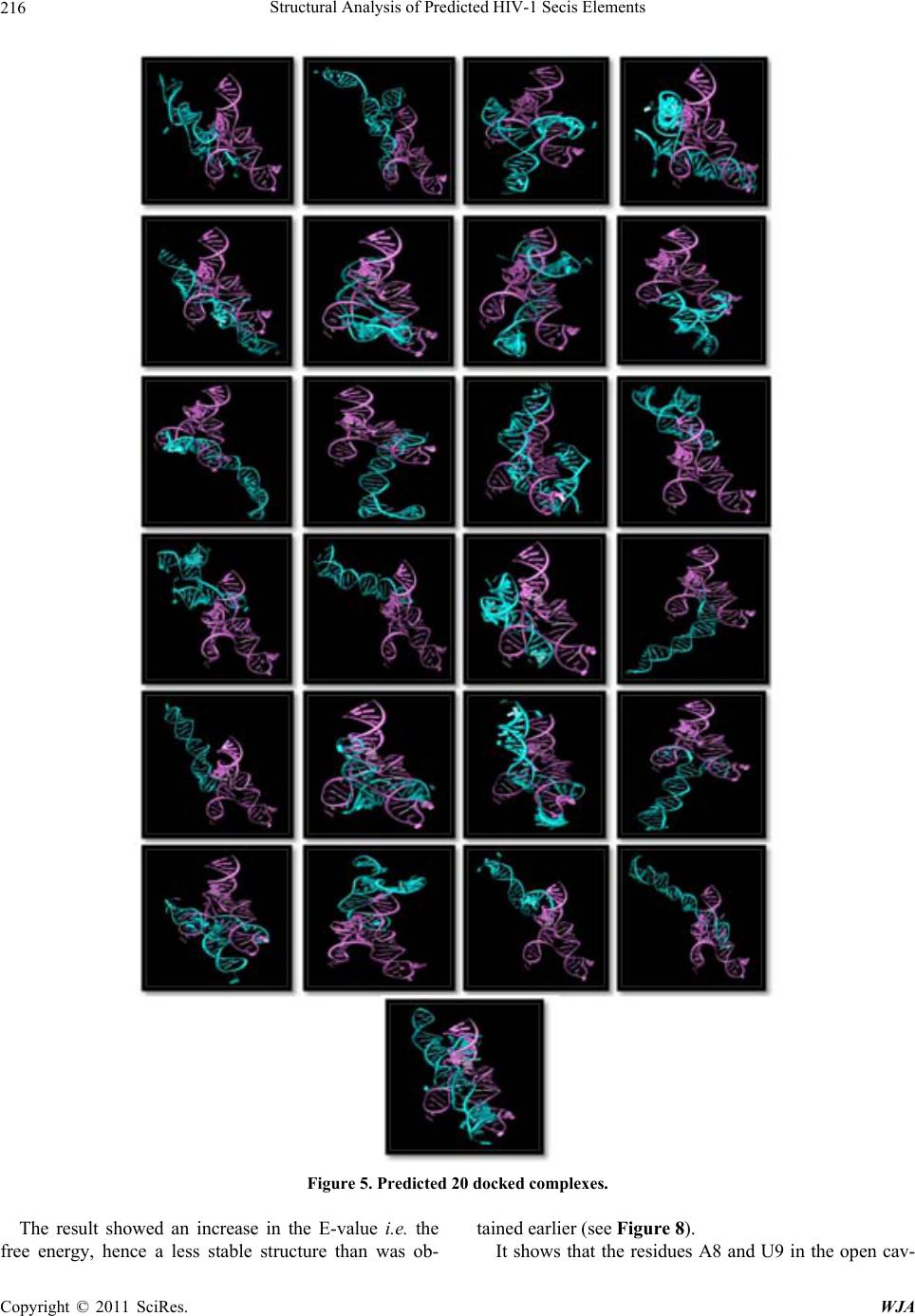 Structural Analysis of Predicted HIV-1 Secis Elements 216 Figure 5. Predicted 20 docked complexes. The result showed an increase in the E-value i.e. the free energy, hence a less stable structure than was ob- tained earlier (see Figure 8). It shows that the residues A8 and U9 in the open cav- Copyright © 2011 SciRes. WJA  Structural Analysis of Predicted HIV-1 Secis Elements217 Figure 6. Residues A8 and U9 of the predicted structures have been highlighted. Figure 7. Bond betwee n the SECIS el em ent and the residue s A8 and U9 of the predicted structures is shown. Figure 8. Showing docked complexes with and without the residues A8 and U9 of the predicted structures and the respective E-values. Copyright © 2011 SciRes. WJA  Structural Analysis of Predicted HIV-1 Secis Elements 218 ity are an important part of the stable binding of the hu- man Sec tRNASec and HIV SECIS elements. REFERENCES [1] V. N. Gladyshev, T. C. Stadtman, D. L. Hatfield and K. T. Jeang, “Levels of Major Selenoproteins in T Cells De- crease during HIV Infection and Low Molecular Mass Selenium Compounds Increase,” Proceedings of the Na- tional Academy of Sciences of the United States of Amer- ica, Vol. 96, No. 3, 1999, pp. 835-839. doi:10.1073/pnas.96.3.835 [2] B. Moghadaszadeh and A. H. Beggs, “Selenoproteins and Their Impact on Human Health through Diverse Physio- logical Pathways,” Physiology, Vol. 21, No. 5, 2006, pp. 307-315. doi:10.1152/physiol.00021.2006 [3] B. M. Dworkin, “Selenium Deficiency in HIV Infection and the Acquired Immunodeficiency Syndrome (AIDS),” Chemico-Biological Interactions, Vol. 9, No. 2-3, 1994, pp. 181-186. doi:10.1016/0009-2797(94)90038-8 [4] J. Lu and A. Holmgren, “Selenoproteins,” Journal of Biological Chemistry, Vol. 284, No. 2, 2009, pp.723-727. doi:10.1074/jbc.R800045200 [5] S. C. Gamble, A. Wiseman and P. S. Goldfarb, “Sele- nium-Dependent Glutathione Peroxidase and Other Se- lenoproteins: Their Synthesis and Biochemical Roles,” Journal of Chemical Technology and Biotechnology, Vol. 68, No.2, 1997, pp. 123-134. doi:10.1002/(SICI)1097-4660(199702)68:2<123::AID-JC TB641>3.0.CO;2-O [6] X. M. Xu, B. A. Carlson, Y. Zhang, H. Mix, G. V. Kryukov, R. S. Glass, M. J. Berry, V. N. Gladyshev and D. L. Hatfield, “New Developments in Selenium Bio- chemistry: Selenocysteine Biosynthesis in Eukaryotes and Archaea,” Biological Trace Element Research, Vol. 119, No. 3, 2007, pp. 234-241. doi:10.1007/s12011-007-8003-9 [7] J. M. Coffin, S. H. Hughes and H. E. Varmus, “Retrovi- ruses,” Cold Spring Harbor Laboratory Press, New York, 1997. [8] A. D. Frankel and J. A. Young, “HIV-1: Fifteen Proteins and an RNA,” Annual Review of Biochemistry, Vol. 67, No. 1, 1998, pp. 1-25. doi:10.1146/annurev.biochem.67.1.1 [9] C. K. Damgaard, E. S. Andersen, B. Knudsen, J. Gorod- kin and J. Kjems, “RNA Interactions in the 5’ Region of the HIV-1 Genome,” Journal of Molecular Biology, Vol. 336, No. 2, 2004, pp. 369-379. doi:10.1016/j.jmb.2003.12.010 [10] S. P. Goff, “Host Factors Exploited by Retroviruses,” Nature Reviews Microbiology, Vol. 5, No. 4, 2007, pp. 253-263. [11] K. A. Wilkinson, R. J. Gorelick, S. M. Vasa, N. Guex, A. Rein, D. H. Mathews, M. C. Giddings and K. M. Weeks, “High-Throughput SHAPE Analysis Reveals Structures in HIV-1 Genomic RNA Strongly Conserved across Dis- tinct Biological States,” PLoS Biology, Vol. 6, No. 4, 2008, pp. 883-899. doi:10.1371/journal.pbio.0060096 [12] J. M. Watts, K. K. Dang, R. J. Gorelick, C. W. Leonard, J. W. Bess, R. Swanstrom Jr., C. L. Burch and K. M. Weeks, “Architecture and Secondary Structure of an Entire HIV-1 RNA Genome,” Nature, Vol. 460, No. 7256, 2009, pp. 711-716. doi:10.1038/nature08237 [13] L. Zhao, A. G. Cox, J. A. Ruzicka, A. A. Bhat, W. Zhang and E. W. Taylor, “Molecular Modeling and in Vitro Ac- tivity of an HIV-1-Encoded Glutathione Peroxidase,” Proceedings of the National Academy of Sciences of the United States of America, Vol. 97, No. 12, 2000, pp. 6356-6361. doi:10.1073/pnas.97.12.6356 [14] G. Leslie, “Potential SECIS Elements in HIV-1 Strain HXB2,” Journal of Acquired Immune Deficiency Syn- dromes and Human Retrovirology, Vol. 17, No. 5, 1998, pp. 398-403. doi:10.1097/00042560-199804150-00003 [15] Y. Zhang and V. N. Gladyshev, “An Algorithm for Iden- tification of Bacterial Selenocysteine Insertion Sequence Elements and Selenoprotein Genes,” Bioinformatics, Vol. 21, No. 11, 2005, pp. 2580-2589. doi:10.1093/bioinformatics/bti400 [16] M. Zuker, “Mfold Web Server for Nucleic Acid Folding and Hybridization Prediction,” Nucleic Acids Research, Vol. 31, No. 13, 2003, pp. 3406-3415. doi:10.1093/nar/gkg595 [17] Y. Ding and C. E. Lawrence, “A Bayesian Statistical Algorithm for RNA Secondary Structure Prediction,” Computers & Chemistry, Vol. 23, No. 3-4, 1999, pp. 387- 400. doi:10.1016/S0097-8485(99)00010-8 [18] Y. Itoh, S. Chiba, S. Sekine and S. Yokoyama, “Crystal Structure of Human Selenocysteine tRNA,” Nucleic Acids Research, Vol. 37, No. 18, 2009, pp. 6259-6268. doi:10.1093/nar/gkp648 Copyright © 2011 SciRes. WJA
|