Advances in Biological Chemistry
Vol. 2 No. 3 (2012) , Article ID: 21806 , 5 pages DOI:10.4236/abc.2012.23030
Measurement of DNA damage by terminal deoxynucleotidyl transferase reaction
![]()
1Group of Environmental Adaptation Science, Faculty of Environmental Earth Science, Hokkaido University, Sapporo, Japan
2Environmental Adaptation Science, Division of Environmental Science Development, Graduate School of Environmental Science, Hokkaido University, Sapporo, Japan
3Laboratory of Environmental Health Sciences, Graduate School of Health Science, Hokkaido University, Sapporo, Japan
4Higher Education Research and Development, Division Institute for the Advancement of Higher Education, Hokkaido University, Sapporo, Japan
5Laboratory of Environmental Health Sciences, Faculty of Health Sciences, Hokkaido University, Sapporo, Japan
Email: *HUkura@ees.hokudai.ac.jpU
Received 12 May 2012; revised 22 June 2012; accepted 30 June 2012
Keywords: DNA Damage; Quantification; TUNEL; Apoptosis; PC12 Cells
ABSTRACT
An improved terminal deoxynucleotidyl transferasemediated dUTP-biotin nick end labeling method for the quantification of DNA damage in tissues and cultured cells was developed. Many reports have revealed that histochemistry of DNA damage can be visualized using immunohistochemistry for the terminal deoxynucleotidyl transferase reaction in tissue sections. However, few reports have described quantification of DNA damage in tissues or cells. In this study, to estimate the degree of DNA damage, the confirmed method for histochemistry using biotinylated dUTP and deoxynucleotidyl transferase was applied to label the cleaved DNA ends caused by DNA damage in tissues or cells. After end-labeling, avidin-conjugated peroxidase was reacted. A significant correlation was observed between numbers of cleaved DNA ends and peroxidase activity after the reaction. The obtained signals for presented method showed higher than those for ordinary method, and correlate with degree of DNA damage caused by serum deprivation and chemical dose. In addition, DNA damage caused by apoptosis in cells treated with 6-hydroxydopamine or Cu and in the tissues of rats administered a diet containing no Zn could be evaluated quantitatively using the present method.
1. INTRODUCTION
Many different methods have been used to evaluate DNA damage caused by apoptosis and/or necrosis. As a marker of fragmented DNA, the terminal deoxynucleotidyl transferase (TdT)-mediated dUTP-biotin nick end-labeling (TUNEL) method [1] has been employed widely to identify cells damaged by conditions such as brain ischemia [2], and in tissues treated with anti-cancer drugs [3]. However, few TUNEL signals due to DNA damage in the sections have been observed, and these data are not quantitative. In addition, to evaluate DNA damage in tissues or cells, 8-hydroxy-2’-deoxyguanosine [4], and formamide and thymine glycol lesions [5] in DNA have been measured. Few methods are available for measuring DNA damage directly.
Apoptosis is a representative morphology of programmed cell death that occurs within a developmental context in response to a definable physiologic stimulus [6]. Apoptosis differs from necrosis, which involves accidental and pathologic cell death. Several apoptotic pathways have been discovered [7]. DNA fragmentation due to apoptosis was analyzed by electrophoresis, revealing multiple nucleotide bands, known as laddering. However, little information is available concerning the quantification of DNA fragmentation due to apoptosis and DNA damage caused by necrosis.
We estimated that color development by the enzymatic reaction using ordinary TUNEL method in the sections can be applied reactions in vitro to measure the DNA damage. In this study, to develop an improved method for the quantification of DNA damage caused by apoptosis and necrosis, an ordinary method using the TdT reaction for visualization of DNA damages in tissue sections (TUNEL method) was applied to measure the numbers of cleaved DNA ends caused by DNA damage in tissues and cultured cells. This novel method is expected to be used widely for studying the mechanisms of apoptosis and DNA damage caused by chemicals.
2. MATERIALS AND METHODS
2.1. Materials
PC12 cells, a cell line of rat pheochromocytoma cell, were purchased from the American Type Culture Collection (USA). Dulbecco’s modified Eagle’s medium (DMEM), streptavidin-conjugated peroxidase, and o-phenylenediamine dihydrochloride (OPD) were obtained from SigmaAldrich (St. Louis, MO, USA). Fetal bovine serum (FBS) was from HyClone (Rockville, MD, USA). TdT, HindIII and HinfI were purchased from Toyobo (Osaka, Japan). Biotin-16-2’-dUTP, the blocking reagent and a high pure PCR template preparation kit were obtained from Roche Diagnostics (Mannheim, Germany). Other chemicals were of analytical regent grade.
2.2. Cell Culture
PC12 cells in 35 cm2 flasks or on sterilized cover glass in 6-well plates were maintained in a humidified incubator with 5% CO2 at 37˚C. The cells were incubated in DMEM with and without 10% FBS for 2 hr to 3 days. When the medium was changed to serum-free medium, cells in the flask were washed twice with serum-free DMEM.
2.3. DNA Extraction
Cultured cells were divided into 5 groups. The cells in the control group were cultured in medium containing serum without chemicals. After extraction of DNA from control cells, part of the DNA was supplied for digestion of restriction enzymes. The second group of cells was cultured in serum-free medium without chemicals for 2 - 72 hr. It is well known that apoptosis is induced by removing the serum from whole culture medium [8]. The third, fourth and fifth group constituted cells that were cultured for 72 hr in medium containing serum with 200 mM CuSO4, 200 mM 6-hydroxydopamine (6OHDA) and 0.1 to 10 ppm triethanolamine (TEA), respectively. It is well known that copper ions [9], and 6OHDA [10] can induce apoptosis in PC12 cells. High dose of TEA may induce DNA damage. DNA extraction was carried out using the high pure PCR template preparation kit.
On the other hand, genomic DNA was extracted from about 0.3 g of testis tissue from rats administered with 0 or 20 mg zinc/100 g diet (equivalent to an ordinary diet) [11].
2.4. Digestion of Restriction Enzymes
DNA (each 1 mg) was cleaved with 1 ml of HindIII or HinfI under the condition of 10 mM Tris-HCl, pH7.5, 10 mM MgCl2, 1 mM dithiothreitol and 50 mM NaCl in total 20 ml solution for 1 hr at 37˚C. After reaction, DNA was precipitated by 70% ethanol and 0.2 M sodium acetate buffer, pH 4.5.
2.5. Improved TUNEL Method to Quantify DNA Damage
To quantify the DNA damage, an equal amount of genomic DNA (0.02 to 2 mg) prepared from the cells or tissues was placed on a 96-well plate, and the plate was incubated at 4˚C overnight. The wells were washed twice with 40 mM Tris-HCl buffer, pH 7.4, containing 150 mM NaCl. Nonspecific binding sites were blocked with blocking reagent. The TdT reaction was performed in a reaction mixture consisting of 30 mM Tris-HCl buffer, pH 7.4, containing 140 mM sodium cacodylate, 1 mM cobalt chloride, 0.1 units TdT/mL, and 4 mM biotinylated dUTP at 37˚C for 1 hr. The wells were washed twice with the same buffer. After adding of streptavidin-conjugated peroxidase (diluted 1:200 with same buffer), the plates were incubated for 1 hr at 37˚C, and then washed twice with the same buffer. Then, 0.1% OPD in 50 mM phosphate-citrate buffer, pH 5.0, containing 0.03% sodium perborate was added to the wells. After 30 min, HCl was added to stop the enzyme reaction. Absorbance at 495 nm was measured with a Microplate Reader model 450 (Bio-Rad, USA).
3. RESULTS AND DISCUSSION
To examine whether DNA damage may be evaluated quantitatively using our improved TUNEL method, the level of TUNEL positivity, which is dependent on DNA cleavage by restriction enzymes, was measured (Figure 1). Following DNA damage, the number of DNA ends is usually increased. Thus, it was expected that the positive signal of the improved TUNEL method would be increased in a manner dependent on the number of DNA ends. As shown in Figure 1, although the level of TUNEL positivity in intact genomic DNA samples was almost identical, the level of TUNEL positivity in DNA cleaved with HindIII was increased as the number of DNA ends
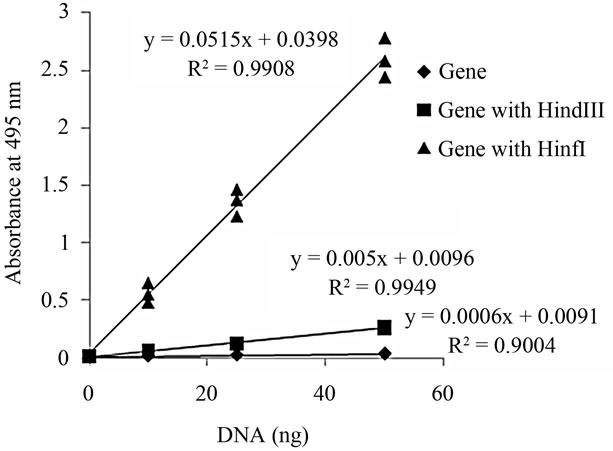
Figure 1. TUNEL-positive levels in DNA cleaved with HindIII or HinfI. In set, numerical formula obtained by the method of least squares is shown.
increased. In addition, level of TUNEL positivity in DNA cleaved with HinfI was about 10-times greater than that in DNA cleaved with HindIII. These values were as expected from the viewpoint of the appearance frequency of each restriction enzyme, because the number of DNA ends from DNA cleaved with HinfI was about 16-times greater compared with that from DNA cleaved with HindIII. A strong significant correlation was observed between TUNEL positivity levels and DNA cleavage sites (Figure 2). From these results, it can be concluded that the improved method for the quantification of DNA damage was able to measure the number of DNA ends.
To examine whether ladder fragmentation due to apoptosis is reflected by TUNEL-positive levels measured by the presented method, DNA damage in cells cultured in medium without serum was evaluated (Figure 2). Serum deprivation induced DNA damage in a time-dependent manner. A significant correlation was observed between TUNEL signals and the duration of serum deprivation (p < 0.05) (Figure 2(a)). In addition, the increasing profile for level of TUNEL positivity (Figure 2(a)) is a good agreement with that of fluorescent intensity dependent on ordinary TUNEL signal under the same condition (Figure 3). Because the cells induced apoptosis by serum deprivation were used in the study, fluorescence can be detected. However, when we use the sections, level of TUNEL positivity by ordinary method was too low (data not shown). As shown in Figure 2, TUNEL positivity by presented method was higher than that by the ordinary TUNEL method. From these results, it appears that the present method is able to evaluate DNA damage status caused by apoptosis in PC12 cells. Of course, it has previously been shown that an increase in TUNEL signal is indicative only of DNA damage, not of apoptosis in particular [12,13]. After the confirmation of DNA ladder fragmentation using agarose gel electrophoresis, it has been recognized as apoptosis [14]. As shown in Figure 2(b), DNA ladder formation in cells cultured in medium
 (a) (b)
(a) (b)
Figure 2. Relative TUNEL-positive levels of DNA damage in PC12 cells cultured in medium without FBS for 0 to 72 hr (a). In (b), electrophoresis of DNA extracted from PC12 cells cultured in the medium without FBS for 6, 48, 72 hr. Error bar means SEM (n = 4).
without serum for 48 - 72 hr was indicated by electrophoresis. Thus, the serum deprivation in PC12 cells also induces apoptosis (Figure 2). From these results using our TUNEL method, it is clear that the apoptosis may progress markedly over during 24 - 48 hr of serum deprivation (Figures 2 and 3). If the ordinary TUNEL method was used for this experiment, we would only able to determine whether apoptosis did or did not occur in each section. Moreover, DNA damage caused by chemicals can be evaluated by our TUNEL method (Figure 4). TEA has been used for consumer goods and as an industrial chemical. As shown in Figure 4(a), TEA could not induce apoptosis. However, high dose TEA induces DNA damage (Figure 4(b)). It is concluded that when the improved TUNEL method presented here is applied, the degree of apoptosis and/or degree of DNA injury can be evaluated quantitatively. From this viewpoint, our method has an advantage as compared with the ordinary TUNEL method.
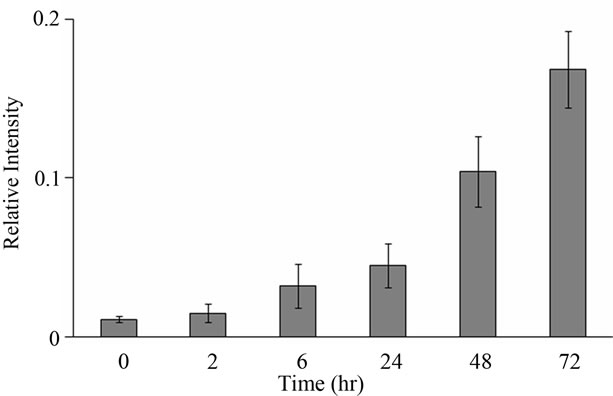
Figure 3. Relative fluorescent intensity obtained by ordinary TUNEL method under the same condition of Figure 2(a). Error bar means SEM (n = 3). The ordinary TUNEL method was performed following the descriptions of Gavrieli et al. [1] and Hosokawa et al. [3].
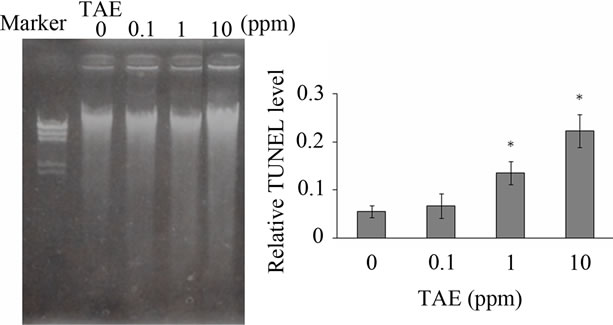 (a) (b)
(a) (b)
Figure 4. Electrophoresis of DNA extracted from PC12 cells cultured in medium containing FBS with TEA (a), and relative TUNEL-positive levels of DNA damage in PC12 cells cultured in medium containing FBS with 0 to 10 ppm TEA for 72 hr (b). Error bar means SEM (n = 4). *indicates p < 0.05.
Finally, we examined whether the present method could be applied to cell and animal models. Cu and 6OHDA are well-known inducers of apoptosis in PC12 cells [9,10]. As shown in Figure 5(a), after treatment of PC12 cells with Cu or 6OHDA, TUNEL-positive levels in DNA increased significantly as compared with those in DNA extracted from control cells. Kawakami et al. [9] and Woodgate et al. [10] described that these chemicals can induce apoptosis at concentrations above 50 mM in the medium. This finding indicates that the TUNEL-positive level reflects the exact degree of apoptosis in the cells.
On the other hand, it was reported that the functional and morphologic changes observed in the testes of rats administered a 0 mg Zn/100 g diet were caused by increased apoptosis [11]. The authors revealed this phenomenon using the ordinary TUNEL method and tissue sections. As shown in Figure 5(b), TUNEL-positive levels in DNA extracted from the testes of rats fed the 0 mg Zn/100 g diet increased significantly (p < 0.001) as compared with those fed the 20 mg Zn/100 g diet (equivalent
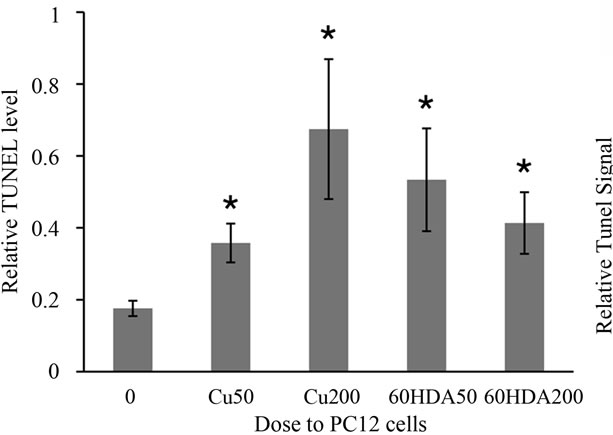 (a)
(a) (b)
(b)
Figure 5. Relative TUNEL levels of nuclei DNA extracted from cells treated with 50 and 200 mM of CuSO4 and 50 and 200 mM of 6OHDA for 72 hr (a) and relative TUNEL levels of nuclei DNA extracted from testis of rat with 0 and 20 mg Zn/100 g diet (b). *indicates p < 0.05.
to an ordinary diet). From these results, it was suggested that the present method is effective for evaluating the degree of apoptosis in tissues.
4. CONCLUSION
We have developed an improved method for the quantification of DNA damage caused by apoptosis and/or necrosis. Linde et al. also reported a new TUNEL assay using a scintillating microplate [15]. However as a radio isotope is used in their method, and the procedure is time consuming, it is not convenient. Compared with their method, our improved method is a simple and reproducible technique for quantitatively evaluating DNA damages caused by not only apoptosis but also chemicals. Many researchers are currently examining the relationship between apoptosis and numerous biologic functions in various organisms. We expect that our improved method will contribute to the progress in this field.
5. ACKNOWLEDGEMENTS
This research was supported by Grants-in-Aid from the Japan Society for the Promotion of Science (No. 20310017 and No. 23655139 for Kurasaki). The authors are indebted to Prof. H. Yanagisawa from the Department of Public Health and Environmental Medicine, Jikei University School of Medicine, Tokyo, Japan for kindly providing the tissue samples from the rats administered the 0 and 20 mg Zn/100 g diet. The authors also thank Ms. Ikumi Yanagiuchi for her technical assistance.
![]()
![]()
REFERENCES
- Gavrieli, Y., Sherman, Y. and Ben-Sasson, S.A. (1992) Identification of programmed cell death in situ via specific labeling of nuclear DNA fragmentation. Journal of Cell Biolology, 119, 493-501. doi:10.1083/jcb.119.3.493
- Tu, Y.F., Tsai, Y.S., Wang, L.W., Wu, H.C., Huang, C.C. and Ho, C.J. (2011) Overweight worsens apoptosis, neuroinflammation and blood-brain barrier damage after hypoxic ischemia in neonatal brain through JNK hyperactivation. Journal of Neuroinflammation, 8, 40. doi:10.1186/1742-2094-8-40
- Hosokawa, T., Okabe, M., Saito, S., Saito, T. and Kurasaki, M. (2000) Protective role of metallothionein on DNA damage in rat kidney caused by cis-diamminedichloroplatinum. Pharmacology and Toxicology, 86, 276- 282. doi:10.1111/j.0901-9928.2000.860606.x
- Shigenaga, M.K., Park, J.W., Cundy, K.C., Gimeno, C.J. and Ames, B.N. (1990) In vivo oxidative DNA damage: measurement of 8-hydroxy-2’-deoxyguanosine in DNA and urine by high-performance liquid chromatography with electrochemical detection. Methods in Enzymology, 186, 521-530. doi:10.1016/0076-6879(90)86146-M
- Maccubbin, A.E., Patrzyc, H.B., Ersing, N., Budzinski, E.E., Dawidzik, J.B., Wallace, J.C., Iijima, H. and Box, H.C. (1999) Assay for reactive oxygen species-induced DNA damage: Measurement of the formamido and thymine glycol lesions. Biochimica et Biophysica Acta, 1454, 80-88. doi:10.1016/S0925-4439(99)00022-8
- Spencer, S.L. and Sorger, P.K. (2011) Measuring and modeling apoptosis in single cells. Cell, 144, 926-939. doi:10.1016/j.cell.2011.03.002
- Jacobson, M.D., Weil, M. and Raff, M.C. (1997) Programmed cell death in animal development. Cell, 88, 347-354. doi:10.1016/S0092-8674(00)81873-5
- Batistatou, A. and Greene, L.A. (1991) Aurintricarboxylic acid rescues PC12 cell and sympathetic neurons from cell death caused by nerve growth factor deprivation: correlation with suppression of endonuclease activity. Journal of Cell Biology, 115, 461-471. doi:10.1083/jcb.115.2.461
- Kawakami, M., Inagawa, R., Hosokawa, T., Saito, T. and Kurasaki, M. (2008) Mechanism of apoptosis induced by copper in PC12 cells. Food and Chemical Toxicology, 46, 2157-2164. doi:10.1016/j.fct.2008.02.014
- Woodgate, A., MacGibbon, G., Walton, M. and Dragunow, M. (1999) The toxicity of 6-hydroxydopamine on PC12 and P19 cells. Brain Research Molecular Brain Research, 69, 84-92. doi:10.1016/S0169-328X(99)00103-5
- Nodera, M., Yanagisawa, H. and Wada, O. (2001) Increased apoptosis in a variety of tissues of zinc-deficient rats. Life Sciences, 69, 1639-1649. doi:10.1016/S0024-3205(01)01252-8
- Rink, A., Fung, K.M., Trojanowski, J.Q., Lee, V.M., Neugebauer, E. and McIntosh, T.K. (1995) Evidence of apoptotic cell death after experimental traumatic brain injury in the rat. American Journal of Pathology, 147, 1575-1583.
- Conti, A.C., Raghupathi, R., Trojanowski, J.Q. and McIntosh, T.K. (1998) Experimental brain injury induces regionally distinct apoptosis during the acute and delayed post-traumatic period. Journal of Neuroscience, 18, 5663- 5672.
- Walker, P.R., LeBlanc, J. and Sikorska, M. (1997) Evidence that DNA fragmentation in apoptosis is initiated and propagated by single-strand breaks. Cell Death and Differentiation, 4, 506-515. doi:10.1038/sj.cdd.4400273
- Linde, V., Flodgaard, H., Kastrup, J.S. and Bjørn, S. (2000) Measurement of apoptosis by the TUNEL method using scintillating microplates. Analytical Biochemistry, 280, 186-188. doi:10.1006/abio.2000.4512
NOTES
*Corresponding author.

