Advances in Biological Chemistry
Vol.3 No.3A(2013), Article ID:33604,8 pages DOI:10.4236/abc.2013.33A005
Calyculin A induces prematurely condensed chromosomes without histone H1 phosphorylation in mammalian G1-phase cells
![]()
Department of Chemistry, University of Wisconsin-Oshkosh, Oshkosh, USA
Email: paulson@uwosh.edu
Copyright © 2013 James R. Paulson, Erica R. Vander Mause. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Received 25 April 2013; revised 2 June 2013; accepted 12 June 2013
Keywords: Mitosis; Chromosome Condensation; Histones; Protein Phosphatases
ABSTRACT
It is shown here that one can induce prematurely condensed chromosomes (PCCs) in G1-phase human (HeLa) and mouse (FT210) cells by treating them with the protein phosphatase inhibitor calyculin A. However, histone H1 is not phosphorylated in these G1-PCCs. It has previously been proposed that histone H1 phosphorylation is responsible for mitotic chromosome condensation, but our results suggest that this is not the case. They indicate instead that phosphorylation of histone H1 is not required for chromosome condensation. It is known that the Cdk1 protein kinase, which triggers mitosis and also phosphorylates histone H1, cannot be activated in G1-phase because mitotic cyclins are not present. Since calyculin A induces PCCs in G1-phase in the absence of active Cdk1, our results suggest that inactivation of protein phosphatases may be just as important for the onset of chromosome condensation and other mitotic events as the activation of protein kinases.
1. INTRODUCTION
It is well known that mitosis is triggered by activation of the protein kinase Cdk1/Cyclin B, also known as Mphase Promoting Factor or MPF [1]. Many proteins are phosphorylated at mitosis either by Cdk1 itself or by secondary protein kinases, and this presumably changes the functions of those proteins so as to produce the characteristic events of mitosis.
In some cases, the roles played by particular mitotic phosphoproteins are known. For example, nuclear envelope breakdown at mitosis involves the phosphorylation of nuclear lamins, leading to depolymerization of lamin filaments and consequent disassembly of the nuclear lamina and nuclear envelope [2-4]. As another example, cessation of transcription at mitosis is due at least in part to phosphorylation of certain transcription factors by Cdk1 [5]. For some mitotic phosphoproteins, however, the function of their phosphorylation is not known. This is the case for the histones.
Histones are the major proteins associated with DNA in eukaryotic chromosomes, packaging the DNA into nucleosomes [6,7] and 30 nm chromatin fibers [8-10]. Histones H1 and H3 are extensively phosphorylated at mitosis. In fact, histone H1 was the first mitotic phosphoprotein to be identified [11]. Histone H1 is phosphorylated at several sites during mitosis, primarily but not exclusively by Cdk1 [12,13], whereas histone H3 is phosphorylated at Ser10 and Ser28, mainly by Aurora B kinase [14].
It was proposed more than 30 years ago that histone H1 phosphorylation is involved in chromosome condensation [15-17] and numerous subsequent studies have also pointed out this possibility (reviewed in [12]). However, these proposals have generally been based on the temporal correlation between H1 phosphorylation and chromosome condensation, without clear evidence for a mechanism. It must therefore be said that at this point the function of histone H1 phosphorylation at mitosis remains unknown.
Prematurely condensed chromosomes (PCCs) are the result of triggering interphase cells to enter a mitosis-like state. This phenomenon was first studied by Johnson and Rao [18] who showed that nuclear envelope breakdown and chromosome condensation could be induced in a G1-, Sor G2-phase cell by fusing it with a metaphase-arrested cell. The premature mitosis-like state presumably occurs because the interphase nucleus is exposed to active Cdk1/cyclin B provided by the mitotic cell.
In the course of our studies on protein dephosphorylation during exit from mitosis [19], we observed that cellpermeable protein phosphatase inhibitors such as calyculin A and cantharidin are able to induce prematurely condensed chromosomes (PCCs) in interphase mammalian cells. The ability of calyculin A to induce PCCs has also been observed by others [20,21] and it has been put to use in the study of chromosome dynamics, breakage and repair, and for cytogenetic analysis (e.g., [22-24]).
In the work reported here, we have used PCCs to explore the possible involvement of histone H1 phosphorylation in chromosome condensation. We show that calyculin A induces chromosome condensation in G1- phase HeLa and FT210 cells, but that histone H1 is not phosphorylated in these condensed chromosomes. This result suggests that H1 phosphorylation is not required for chromosome condensation.
2. MATERIALS AND METHODS
2.1. Chemicals and Media
Media and components were obtained from Invitrogen and Atlanta Biologicals. All other reagents were obtained from Sigma-Aldrich unless otherwise noted. Calyculin A (LC Laboratories, Woburn, MA) was prepared as a 50 μM stock solution in dimethylsulfoxide (DMSO) and stored at –20˚C. Nocodazole was prepared as a 5 mg/mL solution in DMSO and ZM447439 (Santa Cruz Biotechnology) was prepared as a 10 mM solution in DMSO. Both were also stored at –20˚C. Thymidine was dissolved to 100 mM in 0.9% NaCl, filter sterilized and stored at 2˚C.
2.2. Cell Culture and Metaphase Arrest
Mouse FT210 cells [25] were grown in suspension at 32˚C in RPMI-1640 medium and arrested in metaphase with nocodazole as previously described [26]. HeLa S3 cells were grown in suspension at 37˚C in RPMI-1640 medium supplemented with penicillin/streptomycin and 10% fetal bovine serum and diluted daily to 2.0 - 2.5 × 105/mL [27]. For metaphase arrest, HeLa cells were first synchronized in S-phase by treatment with 2.5 mM thymidine for 20 - 24 hrs [28], then released from the thymidine block and arrested with nocodazole as previously described [29]. Mitotic indices for both FT210 and HeLa cells were determined by the method used previously [26] and were typically 80% - 85% for FT210 and 80% - 95% for HeLa S3. The same method was used to determine the percentage of cells exhibiting PCCs. For each determination, at least 200 cells were counted.
2.3. Histone Extraction and Polyacrylamide Gel Electrophoresis
Metaphase chromosomes and/or interphase nuclei were prepared using the procedure for isolating metaphase chromosome clusters [19,30]. Lysis solutions contained either 2 mM p-chloromercuriphenyl sulfonate (PCMPS) or 5 mM Ellman’s reagent (5,5’-dithiobis-(2-nitrobenzoic acid), or DTNB) to block histone dephosphorylation [31]. Histones were extracted from pelleted chromosomes or nuclei with 0.2 M H2SO4, precipitated with ethanol, washed with acetone, dried and redissolved in 3 mM HCl [31]. For analysis of histone H1 phosphorylation by acid-urea gel electrophoresis, aliquots of acidextracted histones were dried using a CentriVap (Labconco) and redissolved in 4 μL of sample buffer [32]. Gels 16 cm long containing 15% acrylamide, 0.1% N,N’- methylene-bis-acrylamide, 2.5 M urea and 5.4% acetic acid were overlain with a polyethylene glycol solution during pre-electrophoresis overnight to prevent distortion of the wells [33]. After samples were loaded, electrophoresis was run at 300 V for 5 - 6 hrs until the blue component of the methyl green marker reached the bottom of the gel. All gels were stained with 0.1% Coomassie Brilliant Blue R250 (BioRad) in 50% methanol, 10% acetic acid and destained in 5% methanol, 10% acetic acid.
2.4. Light Microscopy
For microphotography, cells in a 5 mL culture aliquot were pelleted and resuspended in 2 mL 75 mM KCl. After incubation for 15 min at 37˚C they were fixed by addition of 200 µL of fresh fixative (3 volumes methanol: 1 volume acetic acid). Fixed cells were then pelleted and gently resuspended in fresh fixative three times. Droplets of the final fixed cell suspension were then dropped from a height of 20 cm onto cold, wet microscope slides to spread the chromosomes and the slides were air dried [34]. Chromosomes were stained using a solution of 40 µg/mL Hoechst 33342 in water, and viewed and photographed by epifluorescence in a Nikon Labophot microscope equipped with a Nikon Coolpix 990 camera.
3. RESULTS
3.1. Calyculin A Induces PCCs without Histone H1 Phosphorylation in G1-Phase Mouse FT210 Cells
For a first test of the ability of calyculin A to induce prematurely condensed chromosomes (PCCs) in G1-phase, we used mouse FT210 cells. It is known that Cdk1 is temperature-sensitive in these cells [35] and we have previously shown that metaphase-arrested FT210 cells can be induced to exit mitosis and return to interphase by treating them at their non-permissive temperature to inactivate Cdk1 [26]. After this treatment, the cells appear to be biochemically normal in the sense that they can complete another cell cycle, traversing G1-, Sand G2-phases and eventually reaching the next mitosis. However, when arrested at the next metaphase they are found to have twice the normal number of chromosomes because the heat treatment caused them to leave mitosis without undergoing cytokinesis [26].
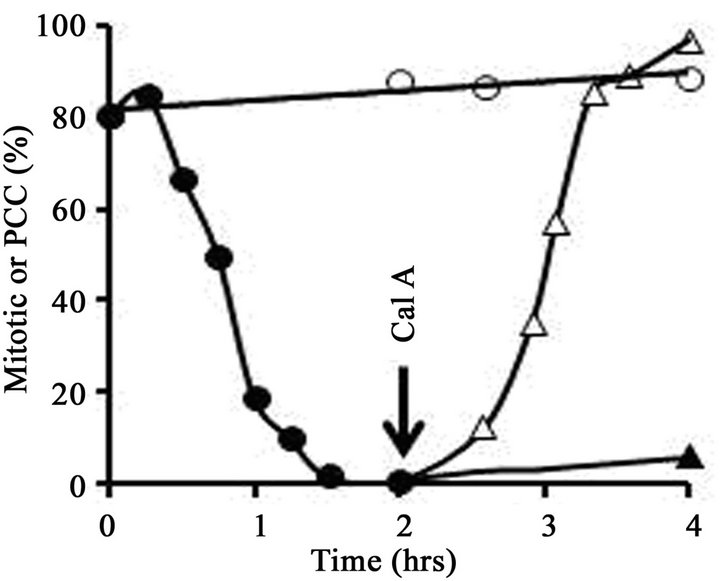
To obtain early G1-phase cells, therefore, FT210 cultures were first arrested in metaphase and then treated with heat to induce exit from mitosis. A typical experiment is shown in Figure 1. FT210 cells growing at their permissive temperature (32˚C) were blocked in metaphase (mitotic index, 81%) with 0.25 µg/mL nocodazole. An aliquot of the culture was then shifted to 41˚C and the mitotic index was followed as a function of time. As shown in Figure 1(A), the mitotic index dropped significantly during the 1.5 hr heat treatment, from 81% to less than 2%, as cells reassembled nuclear envelopes and decondensed their chromosomes. In an aliquot of the culture that was not heat-treated, the mitotic index remained above 80%. Cell viability remained high (above 95%) in all samples throughout the 4 hrs of the experiment.
After 1.5 hr, the portion of the culture that had been treated at 41˚C was shifted back to 32˚C and incubation was continued. At T = 2 hr, half of these cells were treated with 100 nM calyculin A while the other half received no further treatment. In the untreated aliquot, the mitotic index remained low (less than 5%) throughout the rest of the incubation, whereas in the aliquot treated with calyculin A, condensed chromosomes were seen in virtually all cells within 2 hr (Figure 1(A)). In short, calyculin A induced G1-phase PCCs.
Eventually more than 97% of the cells displayed condensed chromosomes in the calyculin A-treated culture aliquot (Figure 1(A)). However, histone H1 was not phosphorylated, as shown by the acid-urea polyacrylamide gel in Figure 1(B). Acid-urea gels, unlike the more familiar SDS polyacrylamide gels, separate basic proteins largely on the basis of charge. Histones are highly positively charged and migrate toward the cathode, but at the pH of the gel each phosphate group contributes one negative charge, reducing the overall positive charge on the protein and decreasing its electrophoretic mobility. Since mitotic histone H1 is phosphorylated at several sites, it migrates significantly more slowly than unphosphorylated (interphase) H1 in acid-urea gels [31,36].
The gel in Figure 1(B) shows that H1 extracted from metaphase-arrested cells at the start of the experiment (lane 1) runs with the lower mobility characteristic of hyperphosphorylated (mitotic) histone H1 (H1M). H1 extracted at the end of the experiment (T = 4 hr) from cells that never received the heat treatment is also hyperphosphorylated (lane 3). On the other hand, cells that were treated for 1.5 hrs at 41˚C and exited mitosis (Figure 1(A)) contain histone H1 that runs with the mobility
 (a)
(a) 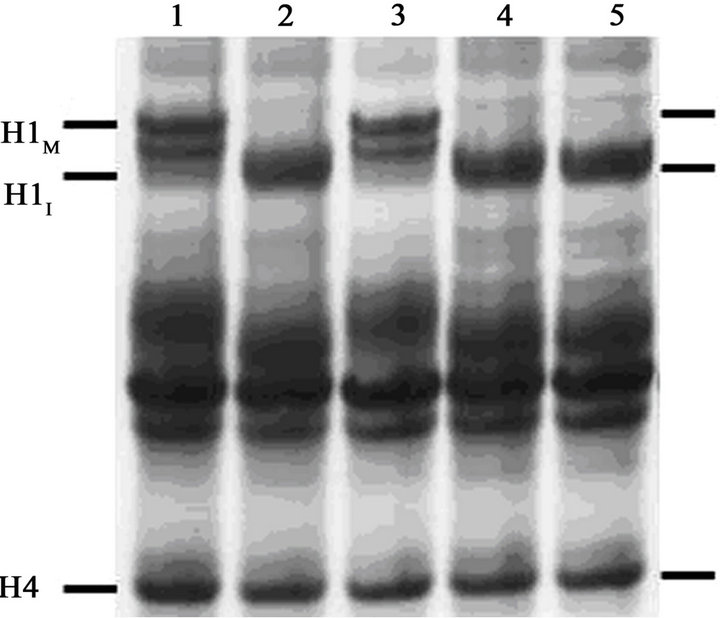 (b)
(b)
Figure 1. Induction of PCC without histone H1 phosphorylation in early G1-phase mouse FT210 cells. (A) FT210 cells, which have temperaturesensitive Cdk1, were arrested in metaphase at 32˚C (mitotic index, 81%). Aliquots were then either shifted to 41˚C (●-●) to induce exit from mitosis [26] or continued at 32˚C (○-○). At T = 1.5 hr after the start of the heat treatment, the cells at 41ºC were shifted back to 32˚C and at T = 2 hr (“Cal A”) aliquots were either treated with 100 nM calyculin A (∆-∆) or not (▲-▲). The percentage of mitotic or PCC cells is shown as a function of time. (B) Samples were taken at various times and histones analyzed on an acid-urea gel. Lane 1, metaphasearrested cells taken at T = 0 hr. Lane 2, cells treated at 41˚C for 1.5 hr, sample taken at T = 2 hr. Note that histone H1 has become dephosphorylated. Lane 3, control cells left at 32˚C, sample taken at T = 2 hr. Lane 4, cells treated at 41˚C but not treated with calyculin A, sample taken at T = 4 hr. Lane 5, cells treated at 41˚C, then treated with 100 nM calyculin A for 2 hr, sample taken at T = 4 hr. The positions of mitotic (phosphorylated) histone H1 (H1M), interphase histone H1 (H1I) and histone H4 are indicated.
characteristic of interphase H1 (H1I), both at the 2 hr (lane 2) and 4 hr (lane 4) time points.
Most importantly, cells that were induced to leave mitosis and afterwards treated with 100 nM calyculin A also contain histone H1 that runs in the interphase (unphosphorylated) position (Figure 1(B), lane 5), even though nearly all the cells in this sample contain PCCs. We conclude that calyculin A can induce prematurely condensed chromosomes in G1-phase FT210 cells, but that histone H1 is not phosphorylated in these condensed chromosomes.
3.2. PCCs Induced by Calyculin A in G1-Phase HeLa Cells Also Lack Phosphorylated Histone H1
An experiment similar to the one described above was carried out with HeLa cells, but in this case G1-phase cells were obtained by first arresting the cells in metaphase with nocodazole and then releasing them from the nocodazole block. For this experiment we used the minimum concentration of nocodazole sufficient for a good metaphase arrest (60 ng/mL) and exposed the cells to nocodazole only until a point 16 hrs after thymidine had been removed. This procedure resulted in a lower mitotic index (80% - 85%) than that attained with a longer block, but reversal of the block and exit from mitosis were more reliably achieved. To reverse the nocodazole block, metaphase-arrested cells were pelleted in the centrifuge, washed twice by resuspending in isotonic saline and pelleting again, and finally resuspended in fresh medium.
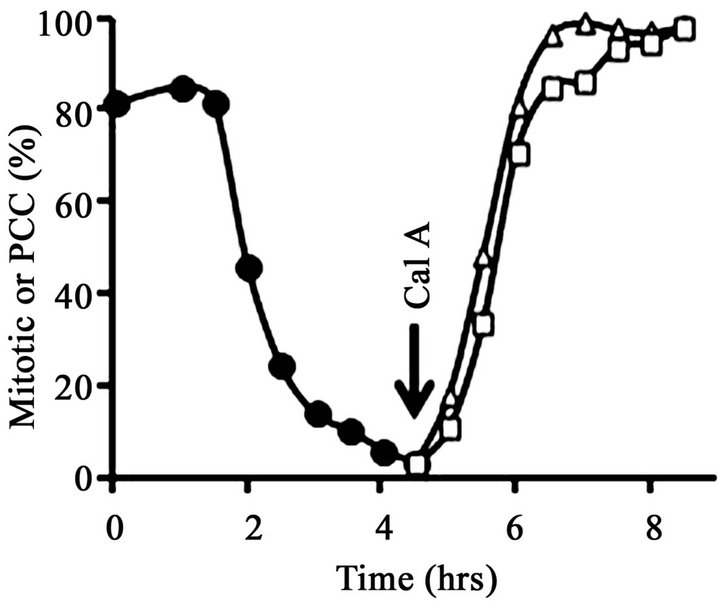
After removal of nocodazole, the metaphase-arrested cells completed mitosis and underwent cytokinesis gradually and asynchronously over the course of about 4 hours. At T = 0, the mitotic index was 81% (Figure 2(A)), most of the cells displayed condensed mitotic chromosomes (Figure 3(A)) and histone H1 was mainly in its mitotic, hyperphosphorylated form (H1M) (Figure 2(B), lane 1). However, by T = 4.5 hr the mitotic index had fallen to about 5% (Figure 2(A)), nearly all cells contained interphase nuclei (Figure 3(B)) and histone H1 was mainly in its dephosphorylated, interphase form (H1I) (Figure 2(B), lane 2). Note that in Figure 2, lane 2, a minor band remains at the H1M position. This is because at T = 4.5 hr about 5% of the cells remained in mitosis. With a mitotic index of 5%, about 10% of the histone H1 should be H1M because mitotic cells have twice the amount of DNA and histones as G1 cells.
At T = 4.5 hr, a portion of the now mainly G1-phase culture was treated with 100 nM calyculin A. After exposure to calyculin A, the percentage of cells with condensed chromosomes increased to more than 95% by T = 8.5 hr (Figure 2(A); Figure 3(C)). However, histone H1 remained primarily in its dephosphorylated, interphase form, H1I (Figure 2(B), lane 3), with no noticeable change from Figure 2(B), lane 2. This result shows that G1- phase PCCs which lack histone H1 phosphorylation can
 (a)
(a) 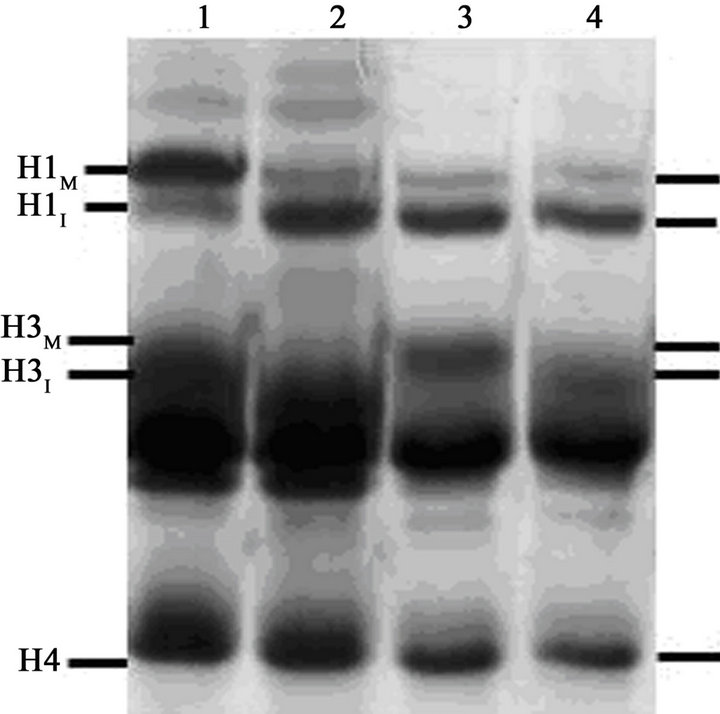 (b)
(b)
Figure 2. Induction of PCC without histone H1 phosphorylation in G1-phase HeLa S3 cells. (A) At T = 0, cells blocked with 60 ng/mL nocodazole (mitotic index, 81%) were released from metaphase arrest by pelleting, washing with saline, and then continuing incubation in fresh medium (●-●). After 4.5 hr (“Cal A”), when the mitotic index had dropped to 5%, one aliquot of the culture was treated with 100 nM calyculin A (∆-∆) while another aliquot was treated with both 100 nM calyculin A and 10 µM ZM447439 (□-□). The percentage of mitotic or PCC cells is shown as a function of time. (B) Histones were extracted from the various samples and separated on an acid-urea gel. Lane 1, metaphase-arrested cells at T = 0. Lane 2, a sample taken at 4.5 hr after removal of nocodazole. Lane 3, cells released from nocodazole (as in lane 2) and then treated with 100 nM calyculin A for 4 hr. Lane 4, cells released from nocodazole and then treated with both 100 nM calyculin A and 10 µM ZM447439 for 4 hr. The positions of mitotic (phosphorylated) histone H1 and H3 (H1M and H3M), interphase histone H1 and H3 (H1I and H3I), and histone H4 are indicated.
be induced in HeLa cells as well as in FT210 cells.
Another portion of the G1-phase culture was treated at T = 4.5 hr with 100 nM calyculin A and also with 10 µM ZM447439. In this case, too, PCCs were induced in more
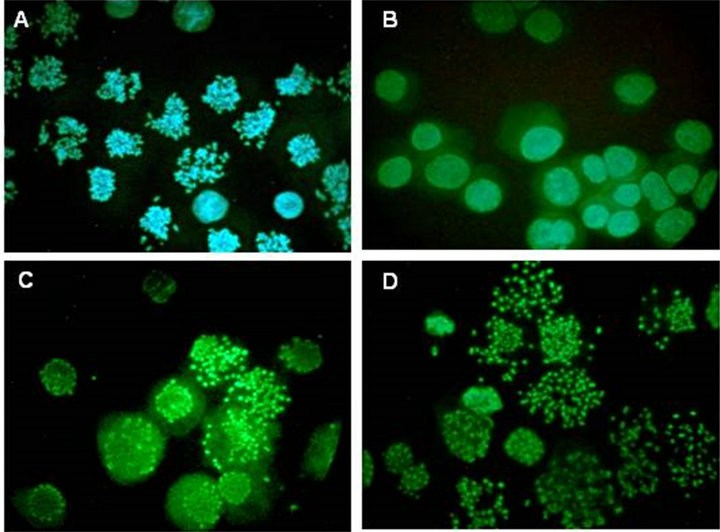
Figure 3. Microscopy of G1-phase PCC induced in HeLa cells by calyculin A. Cells from the experiment shown in Figure 2 were spread on slides, stained with Hoechst 33342 and viewed and photographed by epifluorescence. (A) The initial metaphase-arrested culture (mitotic index, 81%) at T = 0. (B) At 4.5 hr after removal of nocodazole. (C) After release from nocodazole for 4.5 hr and treatment with 100 nM calyculin A for 4 hr. (D) After release from nocodazole and treatment with both 100 nM calyculin A and 10 µM ZM447439 for 4 hr.
than 95% of cells (Figure 2(A); Figure 3(D)) but histone H1 was not phosphorylated (Figure 2(B), lane 4). One difference seen here, however, is that histone H3 appears to run with higher mobility than in Figure 2(B), lane 3. This suggests that although H3 may be phosphorylated in calyculin A-induced G1-PCCs (Figure 2(B), lane 3), it is not phosphorylated when the PCCs are induced in the presence of ZM447439 (Figure 2(B), lane 4). This is in fact expected since ZM447439 inhibits Aurora B [37], the protein kinase that phosphorylates histone H3 at Ser10 in mitosis [14]. Further analysis is needed to verify that H3 is not phosphorylated in this situation, but if this can be confirmed it would lend weight to the notion that neither H1 nor H3 phosphorylation is required for chrosome condensation.
4. DISCUSSION
It has previously been shown that calyculin A, a cellermeable inhibitor of Protein Phosphatases 1 and 2A [38], can induce prematurely condensed chromosomes (PCCs) in interphase mammalian cells [20]. The work we have presented here confirms this and shows further that treatment of HeLa or mouse FT210 cells with calyculin A during G1 phase leads to PCCs that lack phosphorylated histone H1.
These results clearly indicate that histone H1 phosphorylation is not required for formation of G1-PCCs. Since premature chromosome condensation is, as far as we know, very similar or identical to mitotic chromosome condensation, our results suggest that H1 phosphorylation may not be essential in mitosis. The same may also be true of histone H3, since calyculin A can induce G1-PCCs in HeLa cells even in the presence of ZM447439 (Figure 2; Figure 3(D)), an inhibitor of Aurora B, the protein kinase that phosphorylates histone H3 at serine 10 [14,37]. However, further investigation is necessary regarding the phosphorylation of H3 or the lack thereof in G1-PCCs induced in the presence of ZM447439.
Numerous studies have proposed that histone H1 phosphorylation is involved in mitotic chromosome condensation (e.g., [15-17,39]) and our results may seem to be at odds with them. However, these proposals were based only on the correlation between the two events. As noted by Banerjee and Chakravarti in 2011 [40], it was still unclear how H1 phosphorylation could contribute to chromosome condensation. The situation is similar in the case of histone H3, where temporal correlation between H3 phosphorylation and mitotic chromosome condensation has been demonstrated [41,42]. It has also been shown in tetrahymena micronuclei that mutation of the serine 10 phosphorylation sites in H3 to alanine disrupts chromosome condensation [43]. Nevertheless, the role of H3 phosphorylation at mitosis is not clear [44].
Our results do not necessarily contradict those of earlier studies. Although they suggest that H1 phosphorylation is not essential for chromosome condensation, we cannot rule out that it plays some role. It may be, for example, that there are multiple, redundant mechanisms of chromosome condensation and that histone H1 phosphorylation, when it is present, does make a contribution, even though it is dispensable. Perhaps H1 phosphorylation contributes to changes in chromosomes that initiate or facilitate condensation, for example chromatin remodeling [45]. If those changes persist into early G1, then H1 phosphorylation may not be required for G1-PCC. Alternatively, H1 phosphorylation could serve as a “label” [46] for the recruitment of other factors, and perhaps those factors are still present in the chromosomes in early G1. Finally, we cannot rule out the possibility that there may be significant differences between condensed mitotic chromosomes and G1-PCCs that are not evident at the level of resolution provided by the light microscope.
An intriguing aspect of our results is that PCCs and a premature mitosis-like state can be induced in G1-phase in the absence of Cdk1, the protein kinase associated with initiation of mitosis. It should not be possible to activate Cdk1 in G1-phase because the mitotic cyclins necessary for its activation should have been degraded during exit from the previous mitosis and will not be synthesized again until the next S-phase. Since Cdk1/ cyclin B phosphorylates histone H1 [12], the absence of phosphorylated H1 in the calyculin A-induced G1-PCCs confirms that active Cdk1/cyclin B is not present. This suggests that many of the effects of Cdk1/cyclin B at the onset of mitosis may be mediated through the inactivation of protein phosphatases and that protein phosphatase inhibitors such as calyculin A can mimic those effects. This in turn suggests that the inactivation of protein phosphatases may be as important a part of the initiation of mitosis as the activation of protein kinases.
5. ACKNOWLEDGEMENTS
We are grateful to the University of Wisconsin-Oshkosh Faculty Development Board for support of this work. The work was also aided by grants R15 GM39915 and R15 GM46040 from the National Institutes of Health, U.S. Public Health Service.
REFERENCES
- Norbury, C. and Nurse, P. (1992) Animal cell cycles and their control. Annual Review of Biochemistry, 61, 441- 470. doi:10.1146/annurev.bi.61.070192.002301
- Ward, G.E. and Kirschner, M.W. (1990) Identification of cell cycle-regulated phosphorylation sites on nuclear lamin C. Cell, 61, 561-577. doi:10.1016/0092-8674(90)90469-U
- Heald, R. and McKeon, F. (1990) Mutations of phosphorylation sites in lamin A that prevent nuclear lamin disassembly in mitosis. Cell, 61, 579-589. doi:10.1016/0092-8674(90)90470-Y
- Peter, M., Heitlinger, E., Häner, M., Aebi, U. and Nigg E.A. (1991) Disassembly of in vitro formed lamin headto-tail polymers by CDC2 kinase. EMBO Journal, 10, 1535-1544.
- Gottesfeld, J.M. and Forbes, D.J. (1997) Mitotic repression of the transcriptional machinery. Trends in Biochemical Sciences, 22, 197-202. doi:10.1016/S0968-0004(97)01045-1
- Luger, K., Mäder, A.W., Richmond, R.K., Sargent, D.F. and Richmond, T.J. (1997) Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature, 389, 2251-260.
- Kornberg, R.D. and Lorch, Y. (1999) Twenty-five years of the nucleosome, fundamental particle of the eukaryotic chromosome. Cell, 98, 285-294. doi:10.1016/S0092-8674(00)81958-3
- Thoma, F., Koller, T. and Klug, A. (1979) Involvement of histone H1 in the organization of the nucleosome and of the salt-dependent superstructures of chromatin. Journal of Cell Biology, 83, 403-427. doi:10.1083/jcb.83.2.403
- Langmore, J.P. and Paulson, J.R. (1983) Low angle x-ray diffraction studies of chromatin structure in vivo and in isolated nuclei and metaphase chromosomes. Journal of Cell Biology, 96, 1120-1131. doi:10.1083/jcb.96.4.1120
- Robinson, P.J., An, W., Routh, A., et al. (2008) 30 nm chromatin fibre decompaction requires both H4-K16 acetylation and linker histone eviction. Journal of Molecular Biology, 381, 816-825. doi:10.1016/j.jmb.2008.04.050
- Lake, R.S., Goidl, J.A. and Salzman, N.P. (1972) F1-histone modification at metaphase in Chinese hamster cells. Experimental Cell Research, 73, 113-121. doi:10.1016/0014-4827(72)90108-5
- Swank, R.A., Th’ng, J.P.H., Guo, X.-W., Valdez, J., Bradbury, E.M. and Gurley, L.R. (1997) Four distinct cyclin-dependent kinases phosphorylate histone H1 at all of its growth-related phosphorylation sites. Biochemistry, 36, 13761-13768. doi:10.1021/bi9714363
- Happel, N., Stoldt, S., Schmidt, B. and Doenecke, D. (2009) M phase-specific phosphorylation of histone H1.5 at threonine 10 by GSK-3. Journal of Molecular Biology, 386, 339-350. doi:10.1016/j.jmb.2008.12.047
- Hsu, J.-Y., Sun, Z.-W., Li, X., et al. (2000) Mitotic phosphorylation of histone H3 is governed by Ipl1/aurora kinase and Glc7/PP1 phosphatase in budding yeast and nematodes. Cell, 102, 279-291. doi:10.1016/S0092-8674(00)00034-9
- Bradbury, E.M., Inglis, R.J., Matthews, H.R. and Sarner, N. (1973) Phosphorylation of very-lysine-rich histone in Physarum polycephalum. Correlation with chromosome condensation. European Journal of Biochemistry, 33, 131-139. doi:10.1111/j.1432-1033.1973.tb02664.x
- Bradbury, E.M., Inglis, R.J. and Matthews, H.R. (1974) Control of cell division by very lysine rich histone (F1) phosphorylation. Nature, 241, 257-261. doi:10.1038/247257a0
- Matsumoto, Y., Yasuda, H., Mita, S., Marunouchi, T. and Yamada, M. (1980) Evidence for involvement of H1 histone phosphorylation in chromosome condensation. Nature, 284, 181-183. doi:10.1038/284181a0
- Johnson, R.T. and Rao, P.N. (1970) Mammalian cell fusion: Induction of premature chromosome condensation in interphase nuclei. Nature, 226, 717-722. doi:10.1038/226717a0
- Paulson, J.R., Patzlaff, J.S. and Vallis, A.J. (1996) Evidence that the endogenous histone H1 phosphatase in HeLa mitotic chromosomes is protein phosphatase 1, not protein phosphatase 2A. Journal of Cell Science, 109, 1437-1447.
- Gotoh, E., Asakawa, Y. and Kosaka, H. (1995) Inhibition of protein serine/threonine phosphatases directly induces premature chromosome condensation in mammalian somatic cells. Biomedical Research, 16, 63-68.
- Alsbeih, G. and Raaphorst, G.P. (1999) Differential induction of premature chromosome condensation by calyculin A in human fibroblast and tumor cell lines. Anticancer Research, 19, 903-908.
- Suzuki, M., Piao, C.Q., Zhao, Y.L. and Hei, T.K. (2001) Karyotype analysis of tumorigenic human bronchial epithelial cells transformed by chrysolite asbestos using chemically induced premature chromosome condensation technique. International Journal of Molecular Medicine, 8, 43-47.
- Bezrookove, V., Smits, R., Moeslein, G., et al. (2003) Premature chromosome condensation revisited: A novel chemical approach permits efficient cytogenetic analysis of cancers. Genes Chromosomes and Cancer, 38, 177- 186. doi:10.1002/gcc.10268
- Gotoh, E. (2007) Visualizing the dynamics of chromosome structure formation coupled with DNA replication. Chromosoma, 116, 453-462. doi:10.1007/s00412-007-0109-5
- Mineo, C., Murakami, Y., Ishimi, Y., Hanaoka, F. and Yamada, M. (1986) Isolation and analysis of a mammalian temperature-sensitive mutant defective in G2 functions. Experimental Cell Research, 167, 53-62. doi:10.1016/0014-4827(86)90203-X
- Paulson, J.R. (2007) Inactivation of Cdk1/cyclin B in metaphase-arrested mouse FT210 cells induces exit from mitosis without chromosome segregation or cytokinesis and allows passage through another cell cycle. Chromosoma, 116, 215-225. doi:10.1007/s00412-006-0093-1
- Patzlaff, J.S., Terrenoire, E., Turner, B.M., Earnshaw, W.C. and Paulson, J.R. (2010) Acetylation of core histones in response to HDAC inhibitors is diminished in mitotic HeLa cells. Experimental Cell Research, 316, 2123-2135. doi:10.1016/j.yexcr.2010.05.003
- Xeros, N. (1962) Deoxyriboside control and synchronization of mitosis. Nature, 194, 682-683. doi:10.1038/194682a0
- Paulson, J.R., Ciesielski, W.A., Schram, B.R. and Mesner, P.W. (1994) Okadaic acid induces dephosphorylation of histone H1 in metaphase-arrested HeLa cells. Journal of Cell Science, 107, 267-273.
- Paulson, J.R. (1982) Isolation of chromosome clusters from metaphase-arrested HeLa cells. Chromosoma, 85, 571-581. doi:10.1007/BF00327351
- Paulson, J.R. (1980) Sulfhydryl reagents prevent dephosphorylation and proteolysis of histones in isolated HeLa metaphase chromosomes. European Journal of Biochemistry, 111, 189-197. doi:10.1111/j.1432-1033.1980.tb06092.x
- Panyim, S. and Chalkley, R. (1969) High resolution acrylamide gel electrophoresis of histones. Archives of Biochemistry and Biophysics, 130, 337-346. doi:10.1016/0003-9861(69)90042-3
- Paulson, J.R. and Higley, L.L. (1999) Acid-urea polyacrylamide slab gel electrophoresis of proteins: Preventing distortion of gel wells during preelectrophoresis. Analytical Biochemistry, 268, 157-159. doi:10.1006/abio.1998.3026
- Musio, A., Mariani, T., Frediani, C., Ascoli, C. and Sbrana, I. (1997) Atomic force microscope imaging of chromosome structure during G-banding treatments. Genome, 40, 127-131. doi:10.1139/g97-018
- Th’ng, J.P.H., Wright, P.S., Hamaguchi, J., et al. (1990) The FT210 cell line is a mouse G2 phase mutant with a temperature-sensitive cdc2 gene product. Cell, 63, 313- 324. doi:10.1016/0092-8674(90)90164-A
- Gurley, L.R., D’Anna, J.A., Barham, S.S., Deaven, L.L. and Tobey, R.A. (1978) Histone phosphorylation and chromatin structure during mitosis in Chinese hamster cells. European Journal of Biochemistry, 84, 1-15. doi:10.1111/j.1432-1033.1978.tb12135.x
- Ditchfield, C., Keen, N. and Taylor, S.S. (2005) The Ipl1/ Aurora kinase family: Methods of inhibition and functional analysis in mammalian cells. Methods in Molecular Biology, 296, 371-381.
- Ishihara, H., Martin, B.L., Brautigan, D.L., et al. (1989) Calyculin A and okadaic acid: Inhibitors of protein phosphatase activity. Biochemical and Biophysical Research Communications, 159, 871-877. doi:10.1016/0006-291X(89)92189-X
- Th’ng, J.P.H., Guo, X.W., Swank, R.A., Crissman, H.A. and Bradbury, E.M. (1994) Inhibition of histone phosphorylation by staurosporine leads to chromosome decondensation. Journal of Biological Chemistry, 269, 9568- 9573.
- Banerjee, T. and Chakravarti, D. (2011) A peek into the complex realm of histone phosphorylation. Molecular and Cellular Biology, 31, 4858-4873. doi:10.1128/MCB.05631-11
- Hendzel, M.J., Wei, Y., Mancini, M.A., et al. (1997) Mitosis-specific phosphorylation of histone H3 initiates primarily within pericentromeric heterochromatin during G2 and spreads in an ordered fashion coincident with mitotic chromosome condensation. Chromosoma, 106, 348- 360. doi:10.1007/s004120050256
- Wei, Y., Mizzen, C.A., Cook, R.G., Gorovsky, M.A. and Allis, C.D. (1998) Phosphorylation of histone H3 at serine 10 is correlated with chromosome condensation during mitosis and meiosis in Tetrahymena. Proceedings of the National Academy of Sciences of the United States of America, 95, 7480-7484. doi:10.1073/pnas.95.13.7480
- Wei, Y., Yu, L., Bowen, J., Gorovsky, M.A. and Allis, C.D. (1999) Phosphorylation of histone H3 is required for proper chromosome condensation and segregation. Cell, 97, 99-109. doi:10.1016/S0092-8674(00)80718-7
- Pérez-Cadahía, B., Drobic, B. and Davie, J.R. (2009) H3 phosphorylation: Dual role in mitosis and interphase. Biochemistry and Cell Biology, 87, 695-709. doi:10.1139/O09-053
- Horn, P.J., Carruthers, L.M., Logie, C., et al. (2002) Phosphorylation of linker histones regulates ATP-dependent chromatin remodeling enzymes. Nature Structural Biology, 9, 263-267. doi:10.1038/nsb776
- Prigent, C. and Dimitrov, S. (2003) Phosphorylation of serine 10 in histone H3, what for? Journal of Cell Science, 116, 3677-3685. doi:10.1242/jcs.00735

