 Advances in Anthropology 2013. Vol.3, No.4, 240-248 Published Online November 2013 in SciRes (http://www.scirp.org/journal/aa) http://dx.doi.org/10.4236/aa.2013.34034 Open Access 240 Identification of Plant Remains in Underwater Archaeological Areas by Morphological Analysis and DNA Barcoding Angelo Gismondi1, Donatella Leonardi1, Flavio Enei2, Antonella Canini1* 1Department of Biology, University of Rome “Tor Vergata”, Via della Ricerca Scientifica snc, Rome, Italy 2Civic Museum and Castle of Santa Severa, Via del Castello snc, Santa Severa, Rome, Italy Email: *canini@uniroma2.it Received May 28th, 2013; revised June 27th, 2013; accepted July 29th, 2013 Copyright © 2013 Angelo Gismondi et al. This is an open access article distributed under the Creative Com- mons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, pro- vided the original work is properly cited. DNA barcode technique has only recently been applied to archaeobotanical studies. In fact, in association with morphological, scanning electron and optical microscopic analyses, these specific methods allow re- searcher to scientifically classify antique flora samples. Therefore, this project wants to improve, to en- courage and spread further use of this protocol and to highlight the potentialities of the molecular biology and microscopy related to botanical fossils. In conclusion, ancient Olea europaea L. and Crataegus monogyna Jacq. seeds, a Pinus sp. pollen cone, a Quercus petraea (Mattuschka) Liebl. acorn, animal fi- bers and gymnosperm woody fragments, found in a 1st Century BC sunken Dressel 1B amphora, have clearly been identified, in order to enhance knowledge about Central Italy past human activity and envi- ronment. This research has also demonstrated the applicability of this scientific approach on specimens derived from underwater archaeological site. Keywords: Barcoding; Optical Microscopy; SEM Analysis; Archaeobotany; Plant Remains Introduction Sea level variations, due to the effects of post-glacial ages and earth’s structure modifications, caused the submersion, or the sinking, of prehistoric and archaeological sites in water (Bailey & Flemming, 2008). The detection of these areas and their great historical interest induced the development of the “underwater archaeology”. On the other hand, this discipline was also encouraged by the discovery of thousands of sunken wrecks and their content. In literature, only a few works are pre- sented about this science, although it produces useful elements to clarify, suppose and predict different aspects of human an- cestors’ activities and ecosystem (Willcox, 1977; Edge & Gib- bins, 1988; Gorham & Bryant, 2001; Hansson & Foley, 2008; Claesson, 2011). Since the beginning of the last century, flora ancient remains have captured the attention of scientists be- cause of the large amount of new information they could pro- vide about the past (Banning, 2002). A lot of vegetal remains were found in archaeological sites; in fact, flowers, leaves, fruits and woods were regularly used as foods, drugs or funeral offers by ancient populations (Grasso & Fiorentino, 2009). The detection of botanical elements in caves, archaeological areas and hypogean structures and their taxonomic identification allowed scientists to increase knowledge on plant evolution, ethnobotany and reconstruction of past environments (Marota et al., 2002; Liepelt et al., 2006). Morphological observation is the principal approach applied by archaeobotanists to classify plant remains; however, this method sometimes showed wrong and contradictory results (Gismondi et al., 2012). On the other hand, the introduction and the application of molecular analyses on ancient remains, especially after the development of next-generation sequencing techniques, allowed researchers to obtain more accurate data, even in presence of scarce amounts of template (Manen et al., 2003). In particular, DNA barcoding, a method that uses standard nucleotide sequence analysis for species classification (Kress & Erickson, 2008), was recently and successfully associated to archaeobotany (Gould et al., 2010; Gismondi et al., 2012). In 1994, at seven meters under- water, a Dressel 1B amphora, dated back to the 1st Century BC, was found on the sandy seabed of the Pyrgi seaport-canal ar- chaeological site (Santa Severa, Rome) (Enei, 2008). Aim of this work was the botanical identification of flora remains, found in Pyrgi archaeological find, applying, but also encour- aging and improving, the combined use of innovative and clas- sical scientific approaches, in order to increase knowledge about past human populations and ecology. Materials and Methods Detection and Cleaning of Plant Remains Vegetal remains were collected from a sealed amphora (type Dressel B1) found underwater in Pyrgi seaport-canal (Santa Severa, Rome) (Enei, 2008). Samples were preserved water- logged until the analyses. Then, they were washed in H2Odd, dried at room temperature for 48 hours, decontaminated by UV light for 30 minutes and stored at −20˚C to reduce further an- cient DNA (aDNA) degradation (Poinar, 2002; Pruvost et al., 2007). *Corresponding author.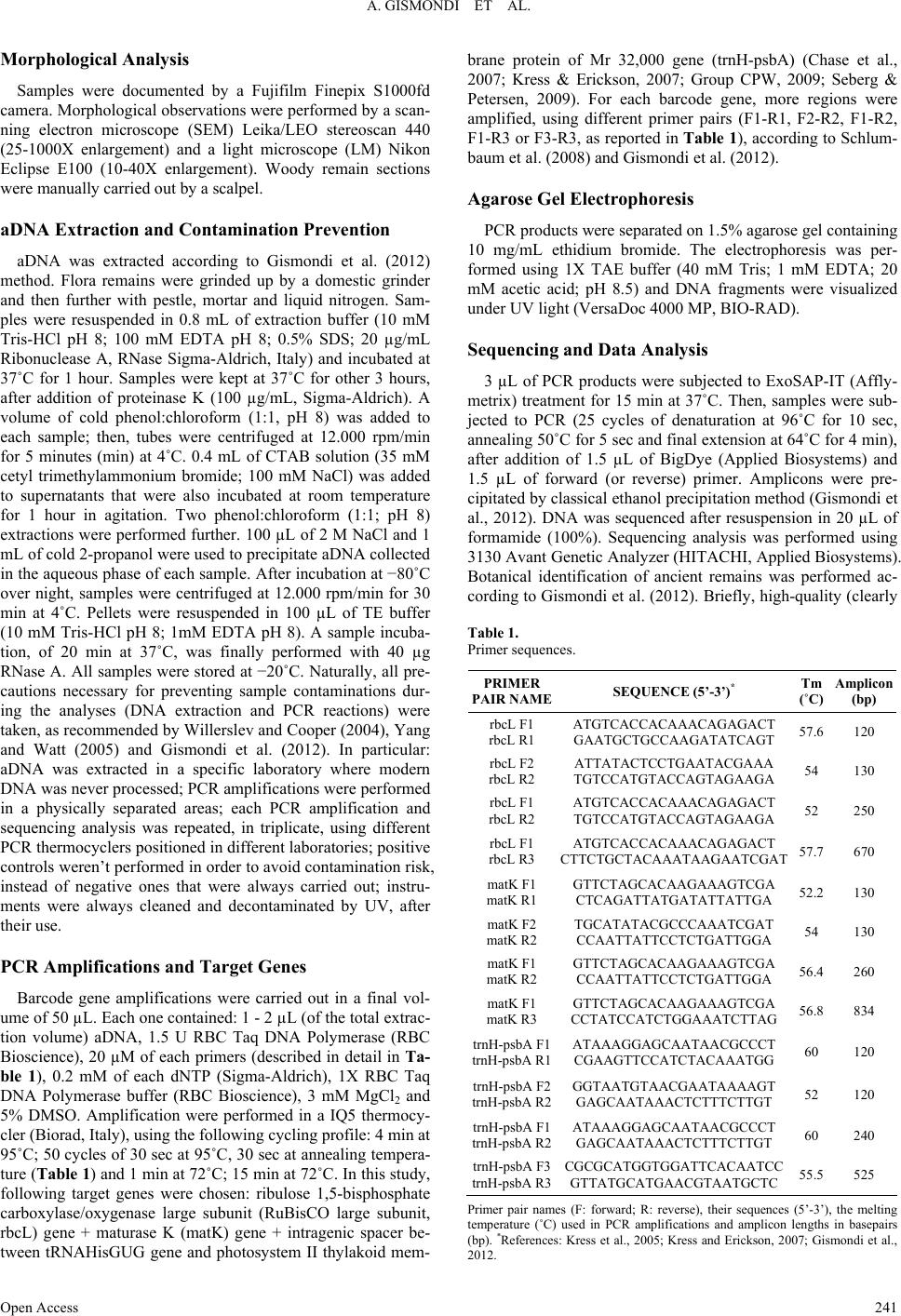  A. GISMONDI ET AL. Morphological Analysis Samples were documented by a Fujifilm Finepix S1000fd camera. Morphological observations were performed by a scan- ning electron microscope (SEM) Leika/LEO stereoscan 440 (25-1000X enlargement) and a light microscope (LM) Nikon Eclipse E100 (10-40X enlargement). Woody remain sections were manually carried out by a scalpel. aDNA Extraction and Contamination Prevention aDNA was extracted according to Gismondi et al. (2012) method. Flora remains were grinded up by a domestic grinder and then further with pestle, mortar and liquid nitrogen. Sam- ples were resuspended in 0.8 mL of extraction buffer (10 mM Tris-HCl pH 8; 100 mM EDTA pH 8; 0.5% SDS; 20 µg/mL Ribonuclease A, RNase Sigma-Aldrich, Italy) and incubated at 37˚C for 1 hour. Samples were kept at 37˚C for other 3 hours, after addition of proteinase K (100 µg/mL, Sigma-Aldrich). A volume of cold phenol:chloroform (1:1, pH 8) was added to each sample; then, tubes were centrifuged at 12.000 rpm/min for 5 minutes (min) at 4˚C. 0.4 mL of CTAB solution (35 mM cetyl trimethylammonium bromide; 100 mM NaCl) was added to supernatants that were also incubated at room temperature for 1 hour in agitation. Two phenol:chloroform (1:1; pH 8) extractions were performed further. 100 µL of 2 M NaCl and 1 mL of cold 2-propanol were used to precipitate aDNA collected in the aqueous phase of each sample. After incubation at −80˚C over night, samples were centrifuged at 12.000 rpm/min for 30 min at 4˚C. Pellets were resuspended in 100 µL of TE buffer (10 mM Tris-HCl pH 8; 1mM EDTA pH 8). A sample incuba- tion, of 20 min at 37˚C, was finally performed with 40 µg RNase A. All samples were stored at −20˚C. Naturally, all pre- cautions necessary for preventing sample contaminations dur- ing the analyses (DNA extraction and PCR reactions) were taken, as recommended by Willerslev and Cooper (2004), Yang and Watt (2005) and Gismondi et al. (2012). In particular: aDNA was extracted in a specific laboratory where modern DNA was never processed; PCR amplifications were performed in a physically separated areas; each PCR amplification and sequencing analysis was repeated, in triplicate, using different PCR thermocyclers positioned in different laboratories; positive controls weren’t performed in order to avoid contamination risk, instead of negative ones that were always carried out; instru- ments were always cleaned and decontaminated by UV, after their use. PCR Amplifications and Target Genes Barcode gene amplifications were carried out in a final vol- ume of 50 µL. Each one contained: 1 - 2 µL (of the total extrac- tion volume) aDNA, 1.5 U RBC Taq DNA Polymerase (RBC Bioscience), 20 µM of each primers (described in detail in Ta- ble 1), 0.2 mM of each dNTP (Sigma-Aldrich), 1X RBC Taq DNA Polymerase buffer (RBC Bioscience), 3 mM MgCl2 and 5% DMSO. Amplification were performed in a IQ5 thermocy- cler (Biorad, Italy), using the following cycling profile: 4 min at 95˚C; 50 cycles of 30 sec at 95˚C, 30 sec at annealing tempera- ture (Table 1) and 1 min at 72˚C; 15 min at 72˚C. In this study, following target genes were chosen: ribulose 1,5-bisphosphate carboxylase/oxygenase large subunit (RuBisCO large subunit, rbcL) gene + maturase K (matK) gene + intragenic spacer be- tween tRNAHisGUG gene and photosystem II thylakoid mem- brane protein of Mr 32,000 gene (trnH-psbA) (Chase et al., 2007; Kress & Erickson, 2007; Group CPW, 2009; Seberg & Petersen, 2009). For each barcode gene, more regions were amplified, using different primer pairs (F1-R1, F2-R2, F1-R2, F1-R3 or F3-R3, as reported in Table 1), according to Schlum- baum et al. (2008) and Gismondi et al. (2012). Agarose Gel Electrophoresis PCR products were separated on 1.5% agarose gel containing 10 mg/mL ethidium bromide. The electrophoresis was per- formed using 1X TAE buffer (40 mM Tris; 1 mM EDTA; 20 mM acetic acid; pH 8.5) and DNA fragments were visualized under UV light (VersaDoc 4000 MP, BIO-RAD). Sequencing and D ata Analysis 3 µL of PCR products were subjected to ExoSAP-IT (Affly- metrix) treatment for 15 min at 37˚C. Then, samples were sub- jected to PCR (25 cycles of denaturation at 96˚C for 10 sec, annealing 50˚C for 5 sec and final extension at 64˚C for 4 min), after addition of 1.5 µL of BigDye (Applied Biosystems) and 1.5 µL of forward (or reverse) primer. Amplicons were pre- cipitated by classical ethanol precipitation method (Gismondi et al., 2012). DNA was sequenced after resuspension in 20 µL of formamide (100%). Sequencing analysis was performed using 3130 Avant Genetic Analyzer (HITACHI, Applied Biosystems). Botanical identification of ancient remains was performed ac- cording to Gismondi et al. (2012). Briefly, high-quality (clearly Table 1. Primer sequences. PRIMER PAIR NAMESEQUENCE (5’-3 ’)* Tm (˚C) Amplicon (bp) rbcL F1 rbcL R1 ATGTCACCACAAACAGAGACT GAATGCTGCCAAGATATCAGT 57.6120 rbcL F2 rbcL R2 ATTATACTCCTGAATACGAAA TGTCCATGTACCAGTAGAAGA 54130 rbcL F1 rbcL R2 ATGTCACCACAAACAGAGACT TGTCCATGTACCAGTAGAAGA 52250 rbcL F1 rbcL R3 ATGTCACCACAAACAGAGACT CTTCTGCTACAAATAAGAATCGAT 57.7670 matK F1 matK R1 GTTCTAGCACAAGAAAGTCGA CTCAGATTATGATATTATTGA 52.2130 matK F2 matK R2 TGCATATACGCCCAAATCGAT CCAATTATTCCTCTGATTGGA 54130 matK F1 matK R2 GTTCTAGCACAAGAAAGTCGA CCAATTATTCCTCTGATTGGA 56.4260 matK F1 matK R3 GTTCTAGCACAAGAAAGTCGA CCTATCCATCTGGAAATCTTAG 56.8834 trnH-psbA F1 trnH-psbA R1 ATAAAGGAGCAATAACGCCCT CGAAGTTCCATCTACAAATGG 60120 trnH-psbA F2 trnH-psbA R2 GGTAATGTAACGAATAAAAGT GAGCAATAAACTCTTTCTTGT 52120 trnH-psbA F1 trnH-psbA R2 ATAAAGGAGCAATAACGCCCT GAGCAATAAACTCTTTCTTGT 60240 trnH-psbA F3 trnH-psbA R3 CGCGCATGGTGGATTCACAATCC GTTATGCATGAACGTAATGCTC 55.5525 Primer pair names (F: forward; R: reverse), their sequences (5’-3’), the melting temperature (˚C) used in PCR amplifications and amplicon lengths in basepairs (bp). *References: Kress et al., 2005; Kress and Erickson, 2007; Gismondi et al., 2012. Open Access 241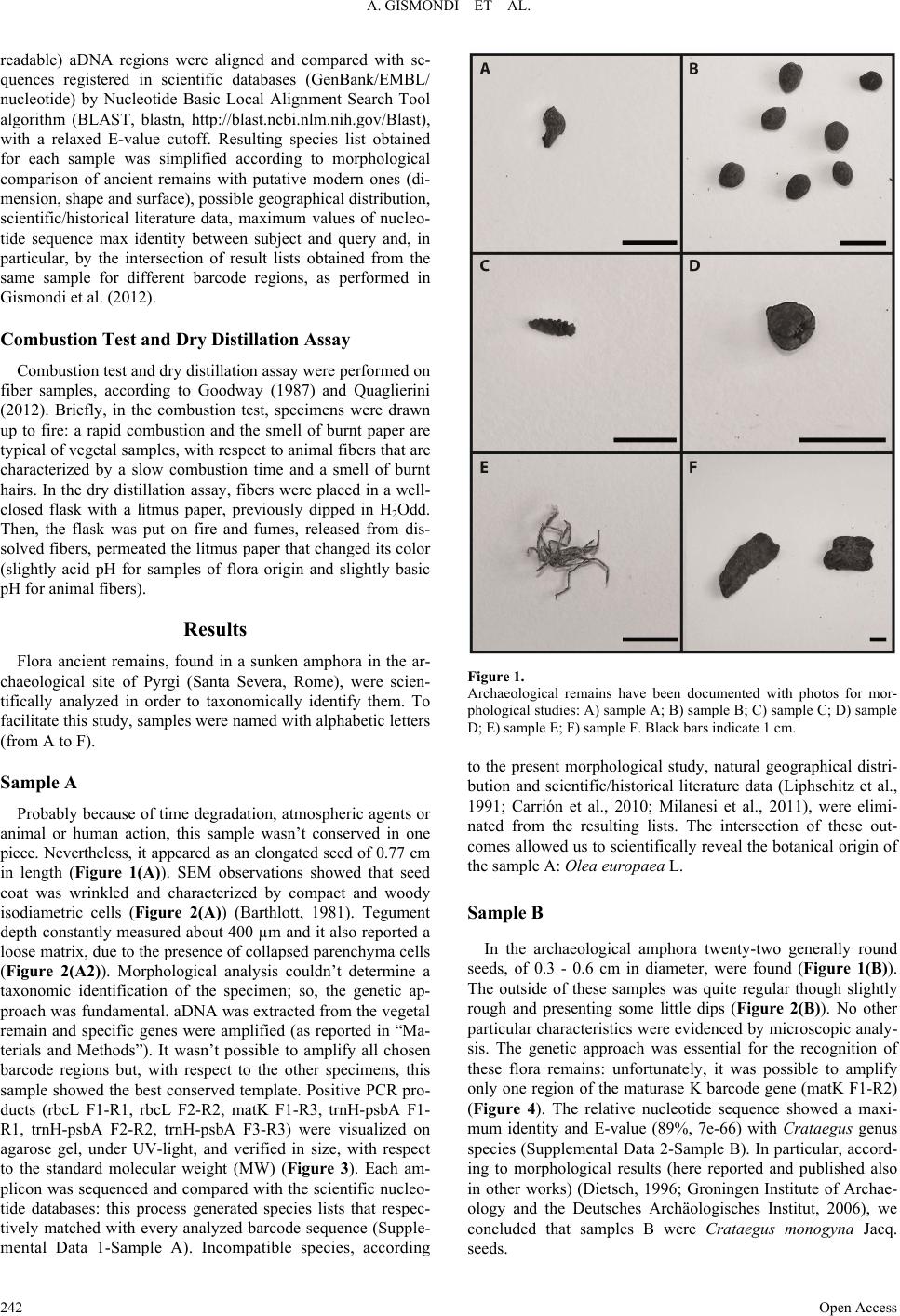  A. GISMONDI ET AL. readable) aDNA regions were aligned and compared with se- quences registered in scientific databases (GenBank/EMBL/ nucleotide) by Nucleotide Basic Local Alignment Search Tool algorithm (BLAST, blastn, http://blast.ncbi.nlm.nih.gov/Blast), with a relaxed E-value cutoff. Resulting species list obtained for each sample was simplified according to morphological comparison of ancient remains with putative modern ones (di- mension, shape and surface), possible geographical distribution, scientific/historical literature data, maximum values of nucleo- tide sequence max identity between subject and query and, in particular, by the intersection of result lists obtained from the same sample for different barcode regions, as performed in Gismondi et al. (2012). Combustion Test and Dry Distillation Assay Combustion test and dry distillation assay were performed on fiber samples, according to Goodway (1987) and Quaglierini (2012). Briefly, in the combustion test, specimens were drawn up to fire: a rapid combustion and the smell of burnt paper are typical of vegetal samples, with respect to animal fibers that are characterized by a slow combustion time and a smell of burnt hairs. In the dry distillation assay, fibers were placed in a well- closed flask with a litmus paper, previously dipped in H2Odd. Then, the flask was put on fire and fumes, released from dis- solved fibers, permeated the litmus paper that changed its color (slightly acid pH for samples of flora origin and slightly basic pH for animal fibers). Results Flora ancient remains, found in a sunken amphora in the ar- chaeological site of Pyrgi (Santa Severa, Rome), were scien- tifically analyzed in order to taxonomically identify them. To facilitate this study, samples were named with alphabetic letters (from A to F). Sample A Probably because of time degradation, atmospheric agents or animal or human action, this sample wasn’t conserved in one piece. Nevertheless, it appeared as an elongated seed of 0.77 cm in length (Figure 1(A)). SEM observations showed that seed coat was wrinkled and characterized by compact and woody isodiametric cells (Figure 2(A)) (Barthlott, 1981). Tegument depth constantly measured about 400 µm and it also reported a loose matrix, due to the presence of collapsed parenchyma cells (Figure 2(A2)). Morphological analysis couldn’t determine a taxonomic identification of the specimen; so, the genetic ap- proach was fundamental. aDNA was extracted from the vegetal remain and specific genes were amplified (as reported in “Ma- terials and Methods”). It wasn’t possible to amplify all chosen barcode regions but, with respect to the other specimens, this sample showed the best conserved template. Positive PCR pro- ducts (rbcL F1-R1, rbcL F2-R2, matK F1-R3, trnH-psbA F1- R1, trnH-psbA F2-R2, trnH-psbA F3-R3) were visualized on agarose gel, under UV-light, and verified in size, with respect to the standard molecular weight (MW) (Figure 3). Each am- plicon was sequenced and compared with the scientific nucleo- tide databases: this process generated species lists that respec- tively matched with every analyzed barcode sequence (Supple- mental Data 1-Sample A). Incompatible species, according Figure 1. Archaeological remains have been documented with photos for mor- phological studies: A) sample A; B) sample B; C) sample C; D) sample D; E) sample E; F) sample F. Black bars indicate 1 cm. to the present morphological study, natural geographical distri- bution and scientific/historical literature data (Liphschitz et al., 1991; Carrión et al., 2010; Milanesi et al., 2011), were elimi- nated from the resulting lists. The intersection of these out- comes allowed us to scientifically reveal the botanical origin of the sample A: Olea europaea L. Sample B In the archaeological amphora twenty-two generally round seeds, of 0.3 - 0.6 cm in diameter, were found (Figure 1(B)). The outside of these samples was quite regular though slightly rough and presenting some little dips (Figure 2(B)). No other particular characteristics were evidenced by microscopic analy- sis. The genetic approach was essential for the recognition of these flora remains: unfortunately, it was possible to amplify only one region of the maturase K barcode gene (matK F1-R2) (Figure 4). The relative nucleotide sequence showed a maxi- mum identity and E-value (89%, 7e-66) with Crataegus genus species (Supplemental Data 2-Sample B). In particular, accord- ing to morphological results (here reported and published also in other works) (Dietsch, 1996; Groningen Institute of Archae- ology and the Deutsches Archäologisches Institut, 2006), we concluded that samples B were Crataegus monogyna Jacq. eeds. s Open Access 242 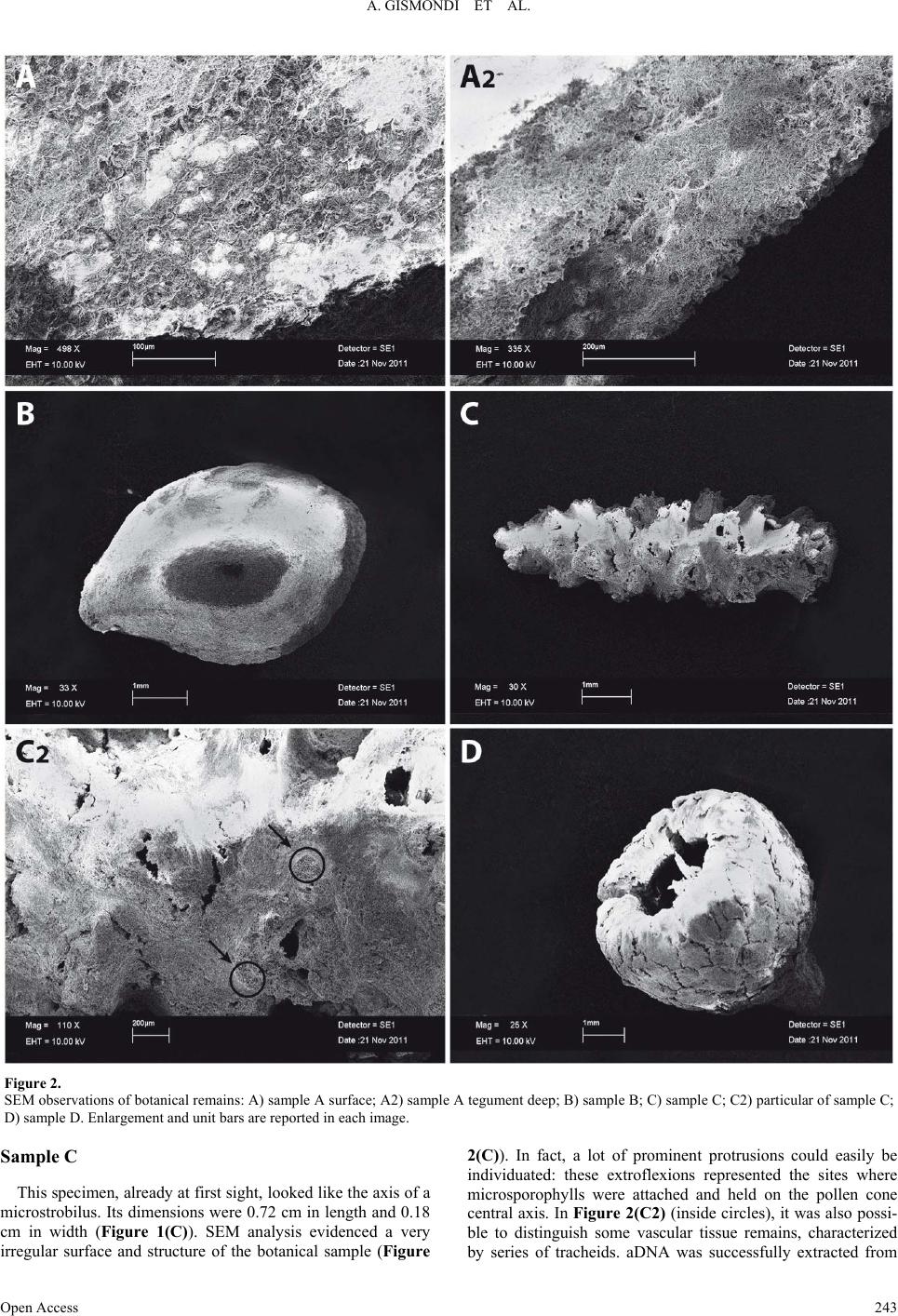 A. GISMONDI ET AL. Open Access 243 Figure 2. SEM observations of botanical remains: A) sample A surface; A2) sample A tegument deep; B) sample B; C) sample C; C2) particular of sample C; D) sample D. Enlargement and unit bars are reported in each image. 2(C)). In fact, a lot of prominent protrusions could easily be individuated: these extroflexions represented the sites where microsporophylls were attached and held on the pollen cone central axis. In Figure 2(C2) (inside circles), it was also possi- ble to distinguish some vascular tissue remains, characterized by series of tracheids. aDNA was successfully extracted from Sample C This specimen, already at first sight, looked like the axis of a microstrobilus. Its dimensions were 0.72 cm in length and 0.18 cm in width (Figure 1(C)). SEM analysis evidenced a very irregular surface and structure of the botanical sample (Figure  A. GISMONDI ET AL. the remain but, maybe because of its excessive degradation, only one PCR amplification (trnH-psbA barcode gene, region F3-R3) was obtained (Figure 4). Results, reported in Supple- mental Data 3-Sample C, confirmed that sample was the fossil- ized remains of a Pinus sp. pollen cone. Sample D Last seed remain, found in the underwater archaeological site, was clearly an acorn. The pericarp was very reduced and com- pletely protected by a thick cupule. The whole structure was rounded, sub-spherical and measured about 0.4 cm (Figure 1(D)). Cupule scales were lanceolated and embricated (Figure 2(D)). Molecular study was performed on the oak nut and the single obtained amplicon (rbcL F1-R2) was fractionized on agarose gel (Figure 4). Positive PCR product was analyzed and matched up with nucleotide sequence databases. Outcomes, reported in Supplemental Data 4-Sample D, showed high se- quence identity (95%) of the sample with Quercus genus. In particular, according to cupule specific morphological features (Pignatti, 1982; Heinz and Barbaza, 1998), this specimen was identified as Quercus petraea (Mattuschka) Liebl. fruit. Sample E No aDNA trace was detected in sample E. Therefore, for its taxonomic identification, our research was essentially based on morphological and microscopic observations. This specimen presented a filamentous structure (Figure 1(E)). At first, we considered the possibility that it could be a vegetal fiber (cot- ton-Gossypium sp. L., linen-Linum usitatissimum L., hemp- Cannabis sativa L., jute-Corchorus sp. L. or ramie-Boehmeria nivea L.). Light microscope image (Figure 5(E1)) showed that, in reality, the sample was made up of several united filaments. Each of these filaments appeared quite uniform, internally full and not characterized, on its surface, by the presence of cell walls (Figures 5(E2), (E3), (E4) and (E5)). According to these preliminary results, the ancient remain could be neither fossil cotton nor linen remains: in fact, Gossipium sp. L. product is Figure 3. PCR amplifications of sample A aDNA have been detected by UV-light after migration on agarose gel 1.5%. MW (molecular weight; 100, 500 and 800 base pairs are indicated); lane 1 (rbcL barcode gene, region F1-R1, 130bp); lane 2 (rbcL barcode gene, region F2-R2, 130 bp); lane 3 (matK barcode gene, region F1-R3, 834bp); lane 4 (trnH-psbA barcode gene, region F1-R1, 120bp); lane 5 (trnH-psbA bar- code gene, region F2-R2, 120bp); lane 6 (trnH- psbA barcode gene, region F3-R3, 525bp). Figure 4. Positive PCR products visualized on aga- rose gel 1% by UV-light. MW (Molecular weight; 200 and 600 base pairs are indi- cated), lane D (sample D aDNA, rbcL bar- code gene, region F1-R2, 201bp), lane B (sample B aDNA, matK barcode gene, re- gion F1-R2, 200bp); lane C (sample C aDNA, trnH-psbA barcode gene, region F3- R3, 525bp). flat, ribbon-like and helix packaged (Han et al., 1998; Krakh- malev & Paiziev, 2006) whilst L. usitati ssimum L. fiber is hollow and characterized by horizontal rings that regularly recur along the filament (Florian et al., 1990). Hemp, jute and ramie fila- ments were also observed by optical microscopy (Figures 5(H), (J) and (R)), in order to discover some similarities between them and the ancient specimen. Unfortunately, all these fibers appeared very different with respect to the archaeological sam- ple. Ramie fiber only presented a remote likeness with the sam- ple E. So, to clarify this doubt, SEM analysis was performed on the ancient remain (Figures 6(E1), (E2) and (E3)) and ramie filaments (Figures 6(R1), (R2) and (R3)). Both fibers meas- ured about 30 - 35 µm in thickness but, outwardly, they were very dissimilar: sample E showed a uniform and quite smooth surface with respect to ramie one that was characterized by filamentous micro-subunits. All these data suggested that sam- ple E was not a vegetal fiber. Thus, we have wanted to demon- strate that it had an animal origin. Apart from the slow burning time of the sample and the smell of burnt hairs that was pro- duced during the empiric combustion test, we also observed that, in the dry distillation assay, litmus paper slightly changing its color towards basic pH (Figure 7), confirming the animal nature of the ancient remain. Probably it was raw silk, accord- ing to Goodway (1987) observations. Sample F Last samples were represented by woody fragments (Figure 1(F)). DNA barcoding technique wasn’t applicable, in this case, because of the absence or too low levels of aDNA. Conse- quently, only microscopic approach allowed us to obtain some information about them. Remain sections clearly showed the Open Access 244  A. GISMONDI ET AL. Open Access 245 lar and observational approaches, in order to carefully recog- nize it. In particular, for the genetic characterization, the best presence of vascular bundles between intrafascicular paren- chyma (Figure 8(F1)). By the enlargement of these fascicles (Figures 8(F2) and (F3)), lignified cell walls of xylem tissue were easily detectable. In particular, as tracheids but not vessel elements could be distinguished in the section, we concluded that ancient woods belonged to gymnosperm plants (Minnis, 1987; Yaman, 2011). Discussion Ancient DNA (aDNA) is the most important and informative biological component that scientists can find in archaeological areas (Gugerli et al., 2005). In fact, its analysis allowed re- searchers to explain, understand and know a lot of sides of hu- man past and habitat. The presence of aDNA was also detected in underwater conditions and sunken ancient structures (Coolen & Gibson, 2009; Schlumbaum et al., 2012). These data en- couraged the development of this project whose object was the taxonomic classification of botanical remains found in the un- derwater archaeological site of Pyrgi (Santa Marinella, Rome). Moreover, this study also shot for both promote further the use of DNA barcode scientific method, only lately applied to ar- chaeobotany (Gould et al., 2010; Gismondi et al., 2012), and reassess the application of different microscopic techniques, in the cases where aDNA was too much damaged or not informa- tive. In particular, in our previous work (Gismondi et al., 2012), we demonstrated that morphological observations of macro- remains, also if well preserved, could sometimes induce to sample misclassifications. For this fact, the present work wants to underline the importance of the synergy of different tech- niques to scientifically identify past plant species. At first, as genetic investigation would have required specimen destruction, all flora ancient remains (6 different typologies, named from A to F) were photographed for macroscopic and metric documen- tation (Figure 1). Each sample was analyzed, by using molecu- Figure 5. Light microscope images. E1, E2, E3, E4 and E5) Sample E at different enlargement (4X-25X), H) hemp (25X), J) jute (10X); R) ramie (25X). Mag = 202 X EHT = 10.00 kV 100 m Detector = SE1 Date: 30 Nov = 2007 Detector = SE1 Date: 8 Nov 2011 Mag = 1.00 KX EHT = 10.00 kV 30 m 100 m Mag = 300 X EHT = 10.00 kV Mag = 1.00 KX EHT = 10.00 kV 10 m Detector = SE1 Date: 8 Jan 2009 Detec to r = SE1 Date: 21 Nov = 2011 Mag = 1.00 KX EHT = 10.00 kV 30 m Detector = SE1 Date: 8 Nov 2011 Mag = 1.00 KX EHT = 10.00 kV 20 m Detector = SE1 Date: 8 Jan 2009 Figure 6. SEM images of sample E (E1, E2, E3) and ramie (R1, R2, R3). Enlargement and unit bars are reported in each image.  A. GISMONDI ET AL. Figure 7. Litmus paper (on the right), used in the dry distil- lation assay, slightly changing its color towards basic pH with respect to control litmus paper (on the left), only imbued with H2Odd. Figure 8. Light microscope images of sample F sections. Enlargement: 10× for F1, 40× for F2 and F3. plant barcode genes, reported in literature (Kress and Erickson, 2008), were chosen. As aDNA could easily present double strand breaks, to obtain at least one positive amplification, for each target gene, we tried to amplify more regions. So, primers were designed to produce very short (80 - 300 bp) and overlap- ping or consequent amplicons, according to Schlumbaum et al. (2008) and Gismondi et al. (2012). However, we also subjected aDNA to PCR amplifications of longer sequences (>500 bp) in order to determine if the template hadn’t an excessive deg- radation level. Optimal preservation conditions of the samples can represent, in archaeogenetics, a crucial element: undam- aged aDNAs, usually, are extracted from remains that were found frozen, desiccated or conserved in oxygen-free environ- ments (Palmer et al., 2012). Probably because Pyrgi flora re- mains were preserved in a hermetically sealed amphora, al- though submerged, no external contamination or excessive phy- sical and chemical degradation was produced on samples. In fact, in spite of the adverse environmental situation, specimen morphological and genetic studies were successfully complet- ed. In particular, with great surprise and fortunately, aDNA ex- tracted from A and C samples allow us to amplify even long nucleotide sequences (Figures 3 line A3 and A6, Figure 4 lane C), demonstrating a good preservation degree of ancient nucleic acids. In addition to anoxic conditions, in fact, remains were maybe highly preserved from the degradation by the presence in the archaeological vessel of vegetables rich in antioxidant molecules (Hansson and Foley, 2008). However, the production of long amplicons starting from ancient or degraded DNA is not so uncommon (Willerslev & Cooper, 2004; Yoccoz et al., 2012). Specimen aDNA amplifications (Figures 3 and 4), their sequencing and matching with nucleotide databases (Supple- mentary information files), in association with microscopic ana- lyses (Figure 2), accurately allowed the identification of the botanical origin of sample A, B, C and D. In particular, for sample B classification the genetic analysis was essential. Therefore, aDNA techniques, even if time-consuming and ex- pensive methods, often represent an accurate and rigorous ap- proach able to support and balance morphological analysis lacks. Unfortunately, aDNA wasn’t detectable from E and F an- cient remains. In these cases, light microscopy (Figures 5 and 8), SEM analysis (Figure 6) and other experimental assays (Figure 7) were necessary. Since the archaeological amphora was identified by Enei et al. (2008) as an ancient replaced of garbage can, vegetal re- mains, identified and classified in the present research (Olea europaea L. and Crataegus monogyna Jacq. seeds, a Pinus sp. pollen cone, a Quercus petraea (Mattuschka) Liebl. acorn, ani- mal fibers and gymnosperm woody fragments), surely could be considered as food leftovers or remainders of antique tools. Therefore, this work increased insights about Central Italy past human activity and habitat of the 1st Century BC: in fact, it in- dicated the possible vegetation history and the plant species that were used, grown or traded by people of that period. On the other hand, we improved and encouraged further the complemen- tary application of microscopic techniques and DNA barcoding to archaeobotany, underlining the innovation of its relevance also on samples derived from underwater archeological areas. REFERENCES Bailey, G. N., & Flemming, N. C. (2008). Archaeology of the conti- nental shelf: Marine resources, submerged landscapes and underwa- ter archaeology. Quaternary Scie n c e Reviews, 27, 2153-2165. http://dx.doi.org/10.1016/j.quascirev.2008.08.012 Banning, E. B. (2002). Analysing plant remains. The archaeologist’s laboratory. The analysis of archaeological data. Interdisciplinary contributions to archaeology. Springer. http://dx.doi.org/10.1007/0-306-47654-1_11 Barthlott, W. (1981). Epidermal and seed surface characters of plants: Systematic applicability and some evolutionary aspects. Nordic Journal of Botany, 1, 345-355. http://dx.doi.org/10.1111/j.1756-1051.1981.tb00704.x Carrión, Y., Ntinoub, M., & Badalc, E. (2010). Olea europaea L. in the north Mediterranean basin during the Pleniglacial and the Early- Middle Holocene. Quaternary Science Reviews, 29 , 952-968. http://dx.doi.org/10.1016/j.quascirev.2009.12.015 Chase, M. W., Cowan, R. S., Hollingsworth, P. M., et al. (2007). A pro- posal for a standardized protocol to barcode all land plants. Taxon, 56, 295-299. Claesson, S. (2011). The value and valuation of maritime cultural heri- tage. International Journa l of Cultural Property, 18, 61-80. Open Access 246  A. GISMONDI ET AL. http://dx.doi.org/10.1017/S0940739111000051 Coolen, M. J. L., & Gibson, A. E. (2009). Ancient DNA in lake sedi- ment records. Science Highlights: Paleolimnology, 17, 104-106. Dietsch, M. F. (1996). Gathered fruits and cultivated plants at Bercy (Paris), a Neolithic village in a fluvial context. Vegetation History and Archaeobotany, 5 , 89-97. http://dx.doi.org/10.1007/BF00189438 Edge, C., & Gibbins, D. (1988). Underwater discovery of Roman sur- gical equipment. British Medical Journal, 297, 1645-1646. http://dx.doi.org/10.1136/bmj.297.6664.1645 Enei, F. (2008). Pyrgi sommersa. Ricognizioni archeologiche subac- quee nel porto dell’antica Caere, S. Marinella. Edito dal Comune di S. Marinella/Museo Civico, Italy. Florian, M. L. E., Kronkright, D. P., & Norton, R. E. (1990). Identifica- tion of plant and animal materials in artifacts. Greenville, NC: Ed- ward Brothers, Inc. 49 (J. Paul Getty Trust). Gismondi, A., Rolfo, M. F., Leonardi, D., Rickards, O., & Canini, A. (2012). Identification of ancient Olea europaea L. and Cornus mas L. seeds by DNA barcode. Comptes Rendus Biologies, 335, 472-479. http://dx.doi.org/10.1016/j.crvi.2012.05.004 Goodway, M. (1987). Fiber identification in practice. Journal of the American Institute for Conservation, 26, 27-44. http://dx.doi.org/10.2307/3179659 Gorham, L. D., & Bryant, V. M. (2001). Pollen, phytoliths, and other microscopic plant remains in underwater archaeology. International Journal of Nautical Archaeology, 30, 282-298. Gould, B. A., León, B., Buffen, A. M., & Thompson, L. G. (2010). Evi- dence of a high-Andean, mid-Holocene plant community: An ancient DNA analysis of glacially preserved remains. American Journal of Botany, 97, 1579-1584. http://dx.doi.org/10.3732/ajb.1000058 Grasso, A. M., & Fiorentino, G. (2009). Studi archeobotanici per l’Ita- lia medievale: Una sintesi. In G. Volpe, P. Favia (a cura di), & V. Atti del Congresso Nazionale di Archeologia Medievale, 30 settem- bre-3 ottobre 2009 Foggia-Manfredonia (pp. 120-126). All’insegna del Giglio: Firenze, Groningen Institute of Archaeology (University of Groningen) & The Deutsches Archäologisches Institut (Berlin) (2006). The digital plant atlas. http:// depa.eldoc.ub.rug.nl/? page= browse&family = Rosaceae Group CPW (2009). A DNA barcode for land plants. Proceedings of the National Academy o f Sciences USA, 106, 12794-12797. http://dx.doi.org/10.1073/pnas.0905845106 Gugerli, F., Parducci, L., & Petit, R. J. (2005). Ancient plant DNA: Review and prospects. New Phytologist, 166, 409-418. http://dx.doi.org/10.1111/j.1469-8137.2005.01360.x Han, Y. J., Cho, Y. J., Lambert, W. E., & Bragg, C. K. (1998). Identifi- cation and measurement of convolutions in cotton fiber using image analysis. Artificial Intelligence Review, 12, 201-211. http://dx.doi.org/10.1023/A:1006521329471 Hansson, M. C., & Foley, B. P. (2008). Ancient DNA fragments inside classical Greek amphoras reveal cargo of 2400-year-old shipwreck. Journal of Archaeological Science, 35, 1169-1176. http://dx.doi.org/10.1016/j.jas.2007.08.009 Heinz, C., & Barbaza, M. (1998). Environmental changes during the late glacial and post-glacial in the central Pyrenees (France): New charcoal analysis and archaeological data. Review of Palaeobotany and Palynology, 104, 1-17. http://dx.doi.org/10.1016/S0034-6667(98)00050-5 Krakhmalev, V. A., & Paiziev, A. A. (2006). Spiral structures of cotton fiber. Cellulose, 13, 45-52. http://dx.doi.org/10.1007/s10570-005-9023-2 Kress, W. J., & Erickson, D. L. (2007). A two-locus global DNA bar- code for land plants: the coding rbcL gene complements the non- coding trnH-psbA spacer region. PLoS One, 6, e508. http://dx.doi.org/10.1371/journal.pone.0000508 Kress, W. J., & Erickson, D. L. (2008). DNA barcodes: Genes, genom- ics and bioinformatics. Proceedings of the National Academy of Sci- ences USA, 105, 2761-2762. http://dx.doi.org/10.1073/pnas.0800476105 Kress, W. J., Wurdack, K. J., Zimmer, E. A., Weigt, L. A., & Janzen, D. H. (2005). Use of DNA barcodes to identify flowering plants. Pro- ceedings of the National Academy of Sc i e n c e s U SA , 1 0 2 , 8369-8374. http://dx.doi.org/10.1073/pnas.0503123102 Liepelt, S., Sperisen, C., Deguilloux, M. F., Petit, R. J., Kissling, R., Spencer, M., De Beaulieu, J. L., Taberlet, P., Gielly, L., & Ziegenha- gen, B. (2006). Authenticated DNA from ancient wood remains. An- nals of Botany, 98, 1107-1111. http://dx.doi.org/10.1093/aob/mcl188 Liphschitz, N., Gophna, R., Hartman, M., & Biger, G. (1991). The be- ginning of olive (Olea europaea) cultivation in the old world: A re- assessment. Journal of Archaeological Science, 18, 441-453. http://dx.doi.org/10.1016/0305-4403(91)90037-P Manen, J. F., Boubyb, L., Dalnokic, O., Marinvalb, P., Turgayd, M., & Schlumbaumd, A. (2003). Microsatellites from archaeological Vitis vinifera seeds allow a tentative assignment of the geographical origin of ancient cultivars. Journal of Archaeological Science, 30, 721-729. http://dx.doi.org/10.1016/S0305-4403(02)00244-3 Marota, I., Basile, C., Ubaldi, M., & Rollo, F. (2002). DNA decay rate in papyri and human remains from Egyptian archaeological sites. American Journal of Physica l Anthropology, 117, 310-318. http://dx.doi.org/10.1002/ajpa.10045 Milanesi, C., Sorbi, A., Paolucci, E., Antonucci, F., Menesatti, P., Costa, C., Pallottino, F., Vignani, R., Cimato, A., Ciacci, A., & Mauro, C. (2011). Pomology observations, morphometric analysis, ultrastructu- ral study and allelic profiles of “olivastra Seggianese” endocarps from ancient olive trees (Olea europaea L.). Comptes Rendus Biolo- gies, 334, 39-49. http://dx.doi.org/10.1016/j.crvi.2010.11.006 Minnis, P. E. (1987). Identification of wood from archaeological sites in the american southwest. I. Keys for gymnosperms. Journal of Ar- chaeological Science, 14, 121-131. http://dx.doi.org/10.1016/0305-4403(87)90002-1 Palmer, S. A., Smith, O., & Allaby, R. G. (2012). The blossoming of plant archaeogenetics. An na ls of Anatomy, 194, 146-156. http://dx.doi.org/10.1016/j.aanat.2011.03.012 Pignatti, S. (1982). Flora d’Italia. Edagricole Editore, I, 113-120. Poinar, H. N. (2002). The genetic secrets some fossils hold. Accounts of Chemical Research, 35, 676-684. http://dx.doi.org/10.1021/ar000207x Pruvost, M., Schwarz, R., Correia, V. B., Champlot, S., Braguier, S., Morel, N., Fernandez-Jalvo, Y., Grange, T., & Geigl, E. M. (2007). Freshly excavated fossil bones are best for amplification of ancient DNA. Proceedings of the National Academy of Sciences USA, 104, 739-744. http://dx.doi.org/10.1073/pnas.0610257104 Quaglierini, C. (2012). Chimica delle fibre tessili. Seconda Edizione. Scienze Zanichelli Editore. Schlumbaum, A., Tensen, M., & Jaenicke-Després, V. (2008). Ancient plant DNA in archaeobotany. Vegetation History and Archaeobotany, 17, 233-244. http://dx.doi.org/10.1007/s00334-007-0125-7 Schlumbaum, A., Van Glabeke, S., & Roland-Ruiz, I. (2012). Towards the onset of fruit tree growing north of the Alps: Ancient DNA from waterlogged apple (Malus sp.) seed fragments. Annals of Anatomy, 194, 157-162. http://dx.doi.org/10.1016/j.aanat.2011.03.004 Seberg, O., & Petersen, G. (2009). How many loci does it take to DNA barcode a crocus? PLoS One, 4, e4598. http://dx.doi.org/10.1371/journal.pone.0004598 Willcox, G. H. (1977). Exotic plants from roman waterlogged sites in London. Journal o f Archaeological Science, 4, 269-282. http://dx.doi.org/10.1016/0305-4403(77)90094-2 Willerslev, E., & Cooper, A. (2004). Ancient DNA. Proceedings of the Royal Society B, 272, 3-16. http://dx.doi.org/10.1098/rspb.2004.2813 Yaman, B. (2011). Anatomy of archaeological wood charcoals from Yenibademli Mound (Imbros), Western Turkey. Mediterranean Ar- chaeology and Archaeometry, 11, 33-39. Yang, Y. D., & Watt, K. (2005). Contamination controls when prepar- ing archaeological remains for ancient DNA analysis. Journal of Ar- chaeological Science, 32, 331-336. http://dx.doi.org/10.1016/j.jas.2004.09.008 Yoccoz, N. G., Bråthen, K. A., Gielly, L., Haile, J., Edwards, M. E., GOSLAR, T., et al. (2012). DNA from soil mirrors plant taxonomic and growth form diversity. Molecular Ecology , 21, 3647-3655. http://dx.doi.org/10.1111/j.1365-294X.2012.05545.x Open Access 247  A. GISMONDI ET AL. Supplemental data files (1, 2, 3 and 4) are available on demand contacting the corresponding author (canini@uniroma2.it). Open Access 248
|