American Journal of Plant Sciences
Vol.5 No.12(2014), Article ID:46299,26 pages DOI:10.4236/ajps.2014.512191
Phytoplasmas and Phytoplasma Diseases: A Severe Threat to Agriculture
Assunta Bertaccini1*, Bojan Duduk2, Samanta Paltrinieri1, Nicoletta Contaldo1
1DipSA Plant Pathology, Alma Mater Studiorum University of Bologna, Bologna, Italy
2Laboratory of Applied Phytopathology, Institute of Pesticides and Environmental Protection, Belgrade, Serbia
Email: *assunta.bertaccini@unibo.it
Copyright © 2014 by authors and Scientific Research Publishing Inc.
This work is licensed under the Creative Commons Attribution International License (CC BY).
http://creativecommons.org/licenses/by/4.0/


Received 19 March 2014; revised 18 April 2014; accepted 10 May 2014
ABSTRACT
Several economically relevant phytoplasma-associated diseases are described together with an update of phytoplasma taxonomy and major biological and molecular features of phytoplasmas. Outlook about persepectives and future work to contain spread of these diseases are also reported.
Keywords:Phytoplasma Diseases, Detection, Prevention

1. Introduction
The evidence that numerous yellows-type diseases of plants, believed to be caused by viruses, were associated with phloem colonization by prokaryotes morphologically resembling mycoplasmas (mycoplasma-like organisms: MLO) was first shown in 1967 [1] . During the last three decades ribosomal rDNA sequencing has provided a wealth of evidence that these wall-less prokaryotes that parasitize plants and insects, constitute a large monophyletic group within the class Mollicutes, and the trivial name “phytoplasma” followed by designation of ‘Candidatus Phytoplasma’, were adopted to denote this taxon of plant pathogens [2] . Indirect biological proof, such as electron microscopy observation, phytoplasma and symptoms elimination after tetracycline treatments [3] , insect and dodder transmission confirmed phytoplasma association with numerous plant diseases worldwide [4] [5] . Phytoplasmas have sizes variable from 200 to 800 nm, they are polymorphic because of the lack of cell wall (Figure 1), and survive and multiply in the isotonic environments provided by plant phloem and insect hemolymph. The phytoplasma chromosome is very small (680 - 1600 kb) and sequence analysis of 16S rDNA and other housekeeping genes suggest that they are most closely related to the Acholeplasma spp. than to spiro-
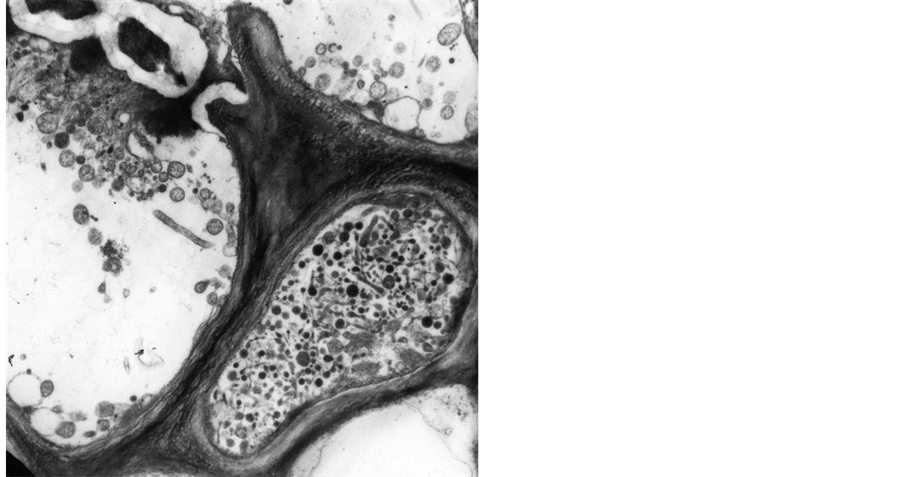
Figure 1. Elecron microscopy picture of cross section of sieve tubes with phytoplasmas (6000×).
plasmas, that are the other mollicutes reported as plant pathogen [6] [7] . The entire genome sequence has been completed for two strains of aster yellows (‘Candidatus Phytoplasma asteris’), two strains of ‘Ca. P. australiense’, and a strain of ‘Ca. P. mali’ [8] -[12] , providing the first opportunity to understand molecular mechanisms underlying pathogen-host interaction and also virulence [13] -[15] .
The application of PCR and nested-PCR assays allow to broadly detect phytoplasma presence, also in mixed infection, in field collected samples [16] [17] . The use of conserved sequences has been a major breakthrough in detection, identification, and classification of phytoplasmas [18] -[26] , and very recently a barcode system was applied also to phytoplasma detection and identification [27] . The introduction of diagnostic tests based on quantitative PCR assays (qPCR) showed to be highly sensitive, and reduced the risk of amplicon contamination, eliminating also the need for a gel-based post PCR product analysis, and making this technique a reliable alternative method to nested PCR assays in routine testing [28] -[30] . On the other hand, the possibility of phytoplasma maintenance in micropropagated shoots [31] -[33] make it possible to organize and maintain a collection of phytoplasma strains that can be provided upon request for general taxonomic identification purposes or other scientific studies worldwide.
2. Symptomatology and Economic Impact
Phytoplasmas are associated with diseases in several hundreds of plant species, including many economically important food, vegetable, and fruit crops; ornamental plants, timber and shade trees. Typical symptoms include virescence/phyllody (development of green leaf like structures instead of flowers) (Figure 2), sterility of flowers, proliferation of axillary buds resulting in a witches’ broom growth, abnormal internodes elongation and generalized stunting. Phytoplasmas may induce many other nonspecific symptoms resulting from stress to which the infected plants are subjected. Hovewer not all infections are necessarily deleterious. For example, the free branching form of poinsettia has been widely used in commercial production of this popular seasonal ornamental, as this trait results in smaller potted plants with numerous showy bracts; these plants are infected by a phytoplasma strain not found up to now in other plant species [34] [35] .
Phytoplasmas substantially undistinguishable on 16S rDNA gene can be associated with diseases inducing different symptoms and/or affecting different plant species, but different phytoplasmas can be associated with similar symptoms in the same or in different plant host(s). This oblige on one hand to search for more accurate molecular markers, ideally associated with pathogenicity features, to carry out phytoplasma identification on multi locus typing (MLT) basis and, on the other hand, to avoid assuming phytoplasma identity just from symp-
 (a)
(a) (b)
(b) (c)
(c) (d)
(d) (e)
(e) (f)
(f)
Figure 2. Symptoms associated with phytoplasma presence, (a) apple proliferation: reddening of apical leaves and witches’ broom appearance in an infected orchard in Italy; (b) European stone fruit yellows in plum: lost of apical dominance and abnormal leaf sprouting in the branches in a Japanese plum in Italy; (c) iujube witches’ broom: shoots of jujube infected tree showing small leaves and witches’ broom proliferation in a germoplams collection of jujube in China (photo by B. Duduk); (d) Sophora japonica yellows: trees showing yellows between others with normal foliage coloration located along a street in China where the phytoplasma-infected trees are graft inoculated and used as decorative varieties (photo by B. Duduk); (e) lime witches’ broom showing typical leaf symptoms of small leaves and dry branches in a lime field in Oman (photo by B. Duduk); (f) strawberry green petals in which virescent and malformed fruits are present in a field in Italy (courtesy A. Calari).
toms and host species. This situation requires molecular identification of the pathogen in order to study its host range and insect vector(s) in the different agricultural or forestry ecosystems. Most damaging and studied phytoplasma diseases are described below in some details.
2.1. Grapevine Yellows (GY)
(GY) represent a collection of widespread diseases in grapevine displaying similar symptoms that are associated with molecularly distinguishable phytoplasmas. The most important diseases in the main viticultural areas of Europe are “flavescence dorée” (FD) and “bois noir” (BN). FD is prevalent in the some of the main grapevine growing countries and it is a quarantine pathogen [36] -[38] . Molecular studies have indicated the presence of phytoplasmas belonging to 16SrV-C and 16SrV-D, respectively and that these subgroup strains differ in their geographical distributions. Both FD types are experimentally transmissible by the same leafhopper vector Scaphoideus titanus [39] . Subgroup 16SrV-D strains were detected in northern Italy [38] , France and Spain [40] [41] where severe disease outbreaks are most frequent. In other grapevine producing areas such as north-central Italy and Serbia the strains associated with FD disease outbreaks are associated with subgroup 16SrV-C strains [42] [43] . Disease symptoms mainly involve plant decline, leaf rolling, shrivelled grapes, unripened shoots and reddening or yellowing of leaves on red or white cultivars respectively. The severity and increasing presence of this disease has prompted extensive efforts for specific phytoplasma detection. One of the major problem viticulturists are facing is the variability of FD phytoplasma strains, therefore further FD strain identification is necessary and it is achieved by studying the polymorphisms in rpS3, SecY gene sequences as well as other genes [24] [40] [44] [45] . Since the majority of “new” discovered strains are very often associated with low epidemic spreading to achieve an effective disease control, with the lowest environmental impact, it is necessary to elucidate the FD strain once the phytoplasma is detected in a new areas or in infected areas after some years from an epidemic outbreak. This information is helpful to reduce and localize the pesticide application against the insect vector and can also help to decide about the need of localized or generalized uprooting of the symptomatic plants in the vineyard in which the FD is detected.
The BN disease is associated with phytoplasmas belonging to ribosomal subgroup 16SrXII-A that induce symptoms undistinguishable from FD. BN is prevalent in all viticultural areas worldwide and is transmitted to grapevine by the planthopper Hyalesthes obsoletus Signoret (Homoptera, Cixiidae) from field bindweed (Convolvulus arvensis L.) [46] and common nettle (Urtica dioica L.) as recognized sources of inoculum. In this case the disease management is very difficult since both phytoplasma and insect vectors are non-host specific, however, the usefulness of tuf gene polymorphism for rapid and cheap phytoplasma detection in epidemiological studies of BN was clearly shown [47] [48] . GY diseases occur in other areas of the world where affected plants exhibit syndromes that are seemingly indistinguishable from those of FD or BN but are associated with different phytoplasmas such as aster yellows (16SrI-B) in Italy and South Africa [49] [50] , ash yellows (16SrVII-A) in Chile [51] , and Australian grapevine yellows in Australia (16SrXII-B) [52] therefore specific local studies are necessary to devise the best management after identification of the insect vector.
2.2. Fruit Trees Declines
Economically important phytoplasma diseases of fruit trees in Europe include apple proliferation (AP), pear decline (PD) and European stone fruit yellows (ESFY). These phytoplasmas are quarantine pest and phylogenetic analyses indicate that the 16S rDNA sequences of the phytoplasmas associated with these disesases are identical or nearly identical. Psyllid vector transmission and host range specificity, clearly distinguish however the associated pathogens as separate species to which the names ‘Ca. P. mali’, ‘Ca. P. pyri’ and ‘Ca. P. prunorum’, respectively, have been assigned [53] (Table 1). Apple proliferation is present in almost all European countries and its major impact on apple (Malus domestica Borkh) cultivars is that the affected trees produce small and unmarketable fruits. The disease reduces size (by about 50%), weight (by 63% - 74%) and quality of fruit, reduces also tree vigour, increasing susceptibility to powdery mildew. Typical AP symptoms are the witches’ broom at the end of shoots, leaves are generally smaller and more dented, with unusually enlarged stipules. Fruits are smaller and flattened, and with elongated peduncles. Early leaf reddening is a good indication of the disease (Figure 2), but it can be induced also by other factors. Very recently a link was demonstrated between the diverse ‘Ca. P. mali’ strains colonizing apple trees and the severity of disease expression [54] . Although apple is the main host of ‘Ca. P. mali’, additional hosts including wild and ornamental Malus species, hazelnut (Corylus spp.), cherry (Prunus avium), apricot (P. armeniaca) and plum (P. domestica) have been identified. The
Table 1. Classification of phytoplasmas based on RFLP analyses and/or sequencing of 16S rDNA (based on 91).
-The ribosomal group has no designation since only Genbank sequences were employed for its determination.
psyllid Cacopsylla picta (Forster) is AP principal vector in northeastern Italy and in Germany, respectively, while in north-western Italy, C. melanoneura is the most abundant psyllid, and overwintering adults, are the responsible for its diffusion in apple orchards. Also the leafhopper Fieberiella florii Stal (Homoptera: Cicadellidae) has been demonstrated as vector of ‘Ca. P. mali’ at least occasionally in Germany. Pear decline (PD) was first reported after 1945 in western regions of North America, but today it is of greater economic importance in European pear orchards. Main symptoms enclose poor shoot and spur growth, dieback of shoots, premature reddening and upper rolling of leaves, reduced leaf and fruit size and fewer fruits on trees. Symptoms of pear decline on trees are a very slow decline and a progressive weakening of the tree. Symptoms similar to those of pear decline can be produced by other factors, such as rootstock-scion incompatibility, poor drainage conditions, malnutrition, and drought. In North America and UK the known vector is the psyllid Cacopsylla pyricola (Foerster), but Cacopsylla pyri (L.) has been found as the main vector in European countries. Studies have indicated that ‘Ca. P. pyri’ is able to overwinter in the body of C. pyri [55] and occurs seasonally during the vegetative stage of tree growth. While age of the tree and scion variety does not seem to influence the occurrence of the disease, differences in susceptibility among varieties and rootstocks have been reported.
European stone fruit yellows (ESFY) affects several stone fruit species inducing apricot, plum, and peach as well as several other Prunus species. Apricot (Prunus armeniaca L.) and Japanese plum (P. salicina Lindl.) are most susceptible; symptom severity on both affected species can be quite variable. Trees exhibit foliar yellowing, accompanied by leaf roll, followed by leaf reddening, reduction or suppression of dormancy with the consequent risk of frost damage, severe and progressive necroses, decline and mortality (Figure 2). European plum is generally tolerant to ESFY, some cultivars however, can show weak symptoms [56] . Wild or cultivated Prunus species, such as P. cerasifera, P. mahaleb, P. padus, P. spinosa, P. tomentosa, are tolerant to the disease, while P. avium demonstrated a high level of resistance. Cacopsylla pruni has been identified as the vector of ESFY [57] ; the species completes one generation per year and adults overwinter on shelter plants (conifers). Tolerant Prunus species can also serve as sources for pathogen spread. In contrast to phytoplasma diseases of stone fruits in North America and Europe, where ‘Ca. P. pruni’ was mainly identified [58] [59] , epidemic outbreaks on almond and stone fruits in the Middle East (Lebanon and Iran) were attributed to ‘Ca. P. phoenicium’ [60] -[63] , whereas in China and India ‘Ca. P. ziziphi’ was identified as the causal agent of severe declines of trees in both cherry and peach orchards [64] .
2.3. Citrus Witches’ Broom
Witches’ broom disease of lime (WBDL), associated with the presence of ‘Ca. P. aurantifolia’ (Table 1), is responsible for major losses of Mexican lime trees (Citrus aurantifolia L.). The disease was first observed more than 30 years ago in Oman, where the destruction of 98% of Mexican lime trees occurred and has since spread to the United Arab Emirates, Iran and India [65] . The infected Mexican lime trees in the early disease stages shows a number of witches’ brooms, bearing small new pale green leaves. In advanced stages, leaves of the older witches’ brooms dry up and eventually fall away leaving behind dead twigs, shoots and dried witches’ broom (Figure 2). At this stage, the tree is unproductive bearing very few small fruits, which die quickly. The period from the first formation of witches’ broom to the mortality of affected trees takes about five to six years. The natural host range of ‘Ca. P. aurantifolia’ includes Citrus aurantifolia, C. medica, C. limetta, C. lemon and C. jambhiri. Phytoplasmas belonging 16SrI and 16SrIX have been detected in a devastating citrus disease known as huanglongbing (HB) in Brazil mainly attributed to three species of Gram negative bacteria in the genus ‘Candidatus Liberibacter’, that is seriously threatening global citrus production [66] [67] . The pathogenic role of phytoplasmas was not clearly elucidated, but they can be detected in symptomatic plants in cases in which HB is not detected (R. Bassanezi, personal communication).
2.4. Palm Lethal Yellowing
Lethal yellowing (LY) is a quickly fatal disease that affects coconut (Cocos nucifera L.) and at least 36 other palm species in the Americas [68] . The phytoplasmas associated with devastating outbreaks of LY have been classified as members of group 16SrIV (Table 1). LY disease occurs elsewhere in the humid tropics under a variety of names depending on location, collectively, these diseases are known as “lethal yellowing-type diseases” (LYD) because they induce symptoms similar to or indistinguishable from those associated with LY. They threaten global coconut production because they kill palms quickly and are incurable. For tall-type coconuts, the onset of visible symptoms begins with a premature shedding of most or all fruit regardless of developmental stage. Aborted fruit usually develop a brownish black calyx-end rot reducing seed viability. The next symptom is inflorescence necrosis (blackening), which is most readily observed on newly opened inflorescences. Leaf discoloration (yellowing or bronzing) begins on the older, lowermost leaves, and progresses to involve successively younger leaves nearest the center of the crown. Affected leaves eventually turn brown, dessicate and hang down forming a skirt around the trunk for several days before falling to the ground. Worldwide this disease affects at least 30 species of palm, including Phoenix dactylifera (date palm), Veitchia merrilli (manila palm), Caryota rumphiana (fishtail palm), Phoenix canariensis (Canary Island date palm), and Elaeis guineensis (African oil palm) [69] -[71] . The disease is vectored by Haplaxius (Myndus) crudus (American palm cixiid) and possibly also by Cedusa species of derbid planthoppers [72] . More recently, phytoplasmas from other 16S ribosomal groups have been associated with diseases showing similar symptoms in palms in other parts of the world. In Saudi Arabia, a 16SrI group was found associated with the Al-Wijam disease of date palm [73] while in North Sudan, 16SrXIV group phytoplasmas were found (Table 1) in this species showing white tip die back and in mature palms with slow decline symptoms [70] [74] [75] . Recently, a 16SrVI phytoplasma was associated with coconut yellow decline and oil palm decline in Malaysia [70] [71] . Similarly, the Weligama wilt disease of coconut in Sri Lanka was associated with phytoplasmas belonging to the 16SrXI group (Table 1) [76] . Phytoplasmas belonging to groups 16SrXI and 16SrXIII are associated with Kalimantan wilt disease in Indonesia [77] , and aster yellows phytoplasmas could be consistently associated with a lethal wilt of oil palm in Colombia [78] .
2.5. Corn and Vegetables Phytoplasma Diseases
Many different vegetable species are seriously affected by molecularly differentiable phytoplasmas worldwide, however symptomatology is very similar for the same species regardless of the infecting phytoplasma strain. Potato witches’ broom and maize bushy stunt are among the most widespread diseases in herbaceous hosts causing severe yield losses. While symptomatic potatoes are associated with the presence of phytoplasmas belonging to diverse 16Sr groups, mainly according to different geographic areas [80] -[84] ; corn is consistently infected in America by aster yellows (16SrI-B) phytoplasmas [85] [86] , while phytoplasma producing a similar disease in Europe belong to the “stolbur” group (16SrXII) and is vectored by Repatuls panzeri and H. obsoletus [79] [87] -[89] . The majority of cultivated vegetable or annual crops were reported as infected by phytoplasma diseases and phytoplasma identification usually is mainly linked to phytoplasmas that are endemic or epidemic to the surrounding areas, indicating that phytoplasma infecting herbaceous plant hosts exhibit very little host/insect vector species specificity [90] .
3. Phytoplasma Classification and Identification
Over the last few years, phytoplasmas have been discovered at an increasingly rapid pace in emerging diseases worldwide and the 16S rDNA study by RFLP (restriction fragment lemght polymorphim) analyses [91] and sequencing has produced a detailed picture of phytoplasma diversity and of their phylogenetic relationships with other prokaryotes. The ‘Candidatus Phytoplasma’ genus was described together with 19 ‘Candidatus species’ [92] and formal descriptions for 35 ‘Ca. Phytoplasma species’ total have since been published. Still others warranting formal description have been recognized, but remain to be published: ‘Ca. P. vitis’, ‘Ca. P. palmae’, ‘Ca. P. luffae’, ‘Ca. P. cocostanzianae’ and ‘Ca. P. cocosnigeriae’.
There are many more than 1000 16S rRNA gene sequences of phytoplasmas archived in the GenBank database, and it is now clear that additional conserved DNA sequences markers should be used as supplemental tools for finer phytoplasma differentiation [93] , to better identify phytoplasma strains in order to correlate them to their differences in pathogenicity, vector relationships and host susceptibility. The rp gene sequences reveal more variation than 16S rDNA, the analyses conducted by RFLP or sequencing on tuf and/or SecY genes show also clear indications of phytoplasma strains relationships at least with their geographic distribution [20] -[23] [25] [26] [94] [95] and are now employed for epidemiologic studies in order to devise management control strategies for the main phytoplasma-associated diseases.
4. Biology and Genomics
Phytoplasmas are introduced by insect vectors during feeding activity into plant sieve tube elements, from which they spread systemically through the plants. Studies on the translocation of phytoplasmas after localized inoculation [96] or the re-colonization of trees [97] , provide evidence that the translocation of phytoplasmas cannot be explained solely by assimilate flow, although active movement by the phytoplasmas seems unlikely, considering the lack of genes coding for cytoskeleton elements or flagella [98] . Phytoplasmas are mainly spread between plants by insects in the families Cicadellidae (leafhoppers), Fulgoridae (planthoppers), and Psyllidae (psyllids), which feed on the phloem sap of infected plants, therefore their host range is strongly dependent upon feeding preferences of their insect vectors [99] . Phytoplasmas may overwinter in insect vectors or in perennial plant hosts, and interact in various ways with insect hosts, and examples of both reduced and enhanced fitness of the vectors, have been reported [98] . There is mounting evidence of transovarial transmission of phytoplasmas occurring in certain insect based on studies of several phytoplasma-vector combinations that include: Scaphoideus titanus/aster yellows [99] ; Hishimonoides sellatiformis/mulberry dwarf [100] , Matsumuratettix hiroglyphicus (Matsumura)/sugarcane white leaf [101] , and Cacopsylla melanoneura/plum [102] . The possibility of seed transmission has also been investigated following in situ detection of phytoplasma DNA in embryos from coconut palms with lethal yellowing disease [103] . Studies on in vitro growing commercial seedlings of alfalfa (Medicago sativa) firstly showed evidence of seed phytoplasma transmission, data were further confirmed by molecular testing of seedlings from phytoplasma-infected lime (Citrus aurantiaca), tomato (Lycopersicum esculentum), corn and winter oil seed rape maintained in vitro or under insect-proof conditions [104] -[106] . Phytoplasmas can also be efficiently spread via vegetative propagation such as the grafting of infected plant tissues as shoots onto healthy plants, by vegetative propagation through cuttings, and by micropropagation practices and any other method used to multiply plant material that avoids sexual reproduction.
Phytoplasmas possess the smallest genome of any plant pathogenic bacteria, only some bacterial symbionts of insects have smaller genomes, however many core housekeeping genes are present in multiple copies; they contain two rRNA operons that in several cases show sequence heterogeneity. Gene duplication and redundancy are also well represented: it was estimated that in the genome of the onion yellows (OY) phytoplasma the 18% of gene complement is represented by multiple redundant copies of only five genes, that are generally present in single copy (if they exist at all), in the other Mollicutes whose genomes have been sequenced [11] . Phytoplasma genomes contain relatively large numbers of transposon genes and insertion sequences that are unique to these bacteria, have similar compositions and contain similar genes; they are referred to either as variable mosaics (SVM) [107] -[109] or potential mobile units, (PMUs) [9] . Extrachromosomal DNA (EC-DNA) (1.7 - 7.4 kb) or plasmids of various sizes have been found in all members of the aster yellows (16SrI) and stolbur (16SrXII) phytoplasma groups as well as in some members of the X-disease (16SrIII) and clover proliferation (16SrVI) groups [110] -[114] . In the ‘Ca. P. asteris’, onion yellows strain (OY), a mildly pathogenic, insect-transmitted line (OY-M) contains two types of plasmids (EcOYM and pOYM), each of which possesses a gene encoding the putative transmembrane protein ORF3. Likewise, a non-insect-transmissible line (OY-NIM), has the corresponding plasmids (EcOYNIM and pOYNIM), but pOYNIM lacks ORF3. It was shown that in OY-M, ORF3 was transcribed from two putative promoters, while on EcOYNIM one of the promoter sequences was mutated and the other deleted. ORF3 was not expressed in the OY-NIM-infected plants. Moreover, ORF3 protein seemed more specifically expressed in OY-M-infected insects than in plants. These results suggest that ORF3 may play an integral role in the interactions of OY with the insect host of this phytoplasma [2] .
Phytoplasma genomes contain many genes encoding transporter systems, such as malate, metal-ion and amino-acid transporters, some of which are present as multiple copies. This feature suggests that phytoplasmas import many metabolites from their host cells, and this probably disturbs the metabolic balance, and causes disease symptoms in plants. Analysis of the genome sequence of the OY-M phytoplasma indicates lack of APT synthase genes, and suggests that these phytoplasmas import host ATP as yet unknown mechanisms and ATP synthesis may be strongly dependent on glycolysis. That hypothesis is supported by analysis of ca. 80-kb genomic DNA sequence derived from ‘Ca. P. asteris’ OY-W, a strain which causes severe disease symptoms [115] . Interestingly, an approximately 30-kb region was found to be duplicated in the ‘Ca. P. asteris’ OY-W genome [14] . Two sets of 5 glycolytic enzymes were encoded in this genomic region, which is a unique gene structure that has not been identified so far in any other bacterial genomes. The organization of glycolytic genes of ‘Ca. P. asteris’ AY-WB [9] is similar to that of ‘Ca. P. asteris’ OY-M, rather than ‘Ca. P. asteris’ OY-W, suggesting the duplication of glycolytic genes is unique to ‘Ca. P. asteris’ OY-W. Glycolysis seems to be an important metabolic pathway in some, but not all, phytoplasmas as these genes are completely absent in ‘Ca. P. mali’ [10] , the latter phytoplasma possesses a gene encoding 2-dehydro-3-deoxyphosphogluconate aldolase (eda) leading to he hypothesis that in this phytoplasma pyruvate is formed independently from glycolysis [116] .
5. Interaction with Hosts@NolistTemp# The occurrence of major surface epitopes, that are unique to each phytoplasma species, suggests that these proteins are key participants in specific interactions with host cells. Genes encoding these proteins have been isolated from several phytoplasma groups and are classified into three types: 1) immunodominant membrane protein (Imp), detected in the phytoplasmas associated with sweet potato witches’ broom (SPWB) [98] , apple proliferation (AP) [117] , European stone fruit yellows (ESFY), pear decline (PD), and peach yellow leaf roll (PYLR) [118] ; 2) immunodominant membrane protein A (IdpA), detected in phytoplasmas associated with western X-disease (WX) [119] ; and 3) antigenic membrane protein (Amp), detected in the aster yellows (AY), clover phyllody (CPh), and onion yellows (OY) phytoplasmas [118] [120] . A high expression of the Amp protein was confirmed in AY, CPh, and OY phytoplasmas and this protein was shown to be exported via the Sec protein secretion system, accompanied by the cleavage of its N-terminal signal sequence. The Amp protein of the OY phytoplasma forms a complex with insect microfilament composed by actin and myosin that was correlated with the phytoplasma-transmitting capacity of leafhoppers, suggesting that the interaction between Amp and microfilament complexes plays a major role in determining the transmissibility of phytoplasmas [15] . Morton et al. [118] isolated genes encoding Imps and found that the sequence identities of these genes in several phytoplasmas were not correlated with that of the 16S rDNA gene suggesting that the variability of immunodominant membrane proteins reflects factors other than evolutionary time. Although the gene encoding the immunodominant membrane protein of the WX phytoplasma is idpA, however a gene homologous to Imp was also present in the WX genomic fragment in which the idpA gene was identified [121] . This observation suggests that Imp may be a common ancestor of phytoplasma immunodominant membrane proteins. The sequence identity of Imp was found to be low among the different phytoplasma groups, however the gene organizations flanking Imp were well conserved in most phytoplasmas, and the Imp genes studied thus far are orthologous because of their similar gene organization and conserved transmembrane structure [122] . In contrast, neither the orthologue of idpA, in the complete genomic sequences of OY-M [122] , AY-WB [9] , ‘Ca. P. australiense’ [12] , or ‘Ca. P. mali’ [10] nor the orthologue of Amp, in the complete genomic sequence of ‘Ca. P. mali’ have been found. Imp may serve an important role in host-phytoplasma interactions, like many positively selected proteins. The accumulation of Amp was calculated as about 10-fold greater than that of Imp, and this level was consistent with the “immunodominant” property of Amp in AY-group phytoplasmas. Detection of Imp in phytoplasma-infected plants is also possible however, and therefore the protein amount of Imp must also be high. Western blot analysis of Imp from OY-W suggested that the signal sequence of Imp was uncleaved, and that Imp was retained in the phytoplasma cell membrane. This result agrees with previous reports that Imp is an immunodominant membrane protein in several phytoplasmas, including AP [120] and since expression of Imp in OY-W is clearly shown, antibodies against Imp could be useful for the detection of phytoplasmas as was recently shown for Amp [123] .Since phytoplasmas are intracellular parasites of both plants and insects, their ability to adapt to two diverse environments is of considerable interest. Microarray analysis of ‘Ca. P. asteris’ OY-M revealed that expression of approximately 33% of the genes changes during host switching between plant and insect, suggesting phytoplasma dramatically alters gene expression in response to its host [124] and may use transporters, secreted proteins, and metabolic enzymes in a host-specific manner. The genes encoded in the PMUs of ‘Ca. P. asteris’ AY-WB are more highly expressed in insects than in plants, most likely due to increased production of the extrachromosomal circular type of PMU during insect infection [125] . Differential gene expression between plant and insect hosts has been also reported in ‘Ca. P. asteris’ OY-M, in which TENGU, a small secreted protein identified in the phytoplasma genome, was shown to induce in transgenic Arabidopsis thaliana symptoms similar to those induced by phytoplasma presence, and is more highly expressed in plant than in insect hosts [13] .
6. Virulence and Pathogenicity
Very little is known about phytoplasma virulence. Because they lack most of the common metabolic pathways, it has been speculated that they must assimilate a wide range of materials from the host cells, probably with detrimental effects to the hosts, such as differential symptom expression correlated with differences in genome size. In onion yellows (OY) phytoplasma mild proliferation and yellowing symptoms are present in infection by mild strain (OY-M), while yellowing, stunting, proliferation and witches’ broom are induced by a severe strain (OYW) and the two phytoplasmas has different chromosome size (OY-W, 1000 kbp and OY-M chromosome, 860 kbp). Further analysis showed that five glycolytic genes were duplicated in the severe strain and the phytoplasma population of OY-W had a higher titer than OY-M. In view of these findings, the higher consumption of the carbon source (because of the duplication of the glycolytic genes) may affect the growth rate of the OY-W phytoplasma and tourn in directly or indirectly produce more severe symptoms [14] . It was also shown that a virulence factor, “tengu-su” inducer (TENGU), induces witches’ broom and dwarfism when expressed in transgenic plants of Nicotiana benthamiana and A. thaliana. Although the localization of phytoplasma is restricted to the phloem, TENGU protein was detected in the apical buds by immunohistochemical analysis, suggesting that it was transported from the phloem to other cells. Microarray analysis detected that auxin-responsive genes were down-regulated in the “tengu”-transgenic plants as compared to control plants [13] .
Since phytoplasmas have no cell wall and reside inside of host cells, their membrane proteins and secreted proteins are exposed to the cytoplasm of the host plant or insect cell, and may play important roles in host-parasite interactions and/or virulence. Thus, the identification of genes comprising a secretion system in phytoplasma genomes is important for understanding their biology. Phytoplasmas possess two known secretion systems, the YidC system for the integration of membrane proteins, and the Sec system for the integration and secretion of proteins into the host cell cytoplasm. Genes encoding SecA, SecY, and SecE have been identified in the ‘Ca. P. asteris’ OY-M genome, the SecY [126] [127] , and SecA expression has been confirmed in phytoplasma-infected plants [128] . These genes have also been identified in three other phytoplasma genomes [9] [10] [12] , and secY genes have been cloned from numerous phytoplasma strains [20] -[22] . These results strongly suggest that a functional Sec system is common to most or all phytoplasmas. Antigenic membrane protein [120] , has been reported to be a substrate of the Sec system. Amp has a Sec signal sequence at its N-terminus, which is cleaved in ‘Ca. P. asteris’ OY-M [126] , suggesting that the phytoplasma Sec system utilizes recognition and cleavage of a signal sequence, as in other bacterial Sec systems.
In the ‘Ca. P. asteris’ strain AY-WB genome, at least 56 genes encode predicted secreted proteins. One of these, SAP11, contains eukaryotic nuclear localization signals and accumulates in plant cell nuclei [9] . SAP11- expressing plants exhibit crinkled leaves and produce many stems. Moreover, the fecundity of insect vectors was increased after feeding upon SAP11-expressing plants when compared to those that fed upon nontransformed plants [129] . In addition to SAP11, SAP54 protein encoded by ‘Ca. P. asteris’ AY-WB was reported to cause morphological changes in A. thaliana floral organ development, which resembled symptoms observed on phytoplasma-infected plants [130] . Although the molecular mechanisms contributing to these changes remain unknown, it is assumed that proteins secreted by phytoplasmas may interfere with the function of genes involved in flower development [131] [132] . The identification of TENGU, SAP11, and SAP54 suggest that phytoplasma could induce symptoms by secretion of effector proteins and modification of plant-gene activity [13] [129] [133] .
Energy metabolism is certainly a key topic for understanding phytoplasma biology and pathogenesis. It has been reported that altered levels of oxygen and carbon dioxide affect phytoplasma abundance in Oenothera leaf tip cultures, and concentrations of carbohydrates in the phloem, roots and leaves of phytoplasma-infected plants differ from those in healthy plants [134] -[136] . Because of the wide array of symptoms expressed by diseased plants, nutrient depletion is probably not restricted to sugars, but also includes other compounds. A reduction in the concentration of photosynthetic pigments [137] and of total soluble proteins [137] -[139] , as well as alterations in the hormone balance [135] [140] , amino-acid transport [134] and the occurrence of folate and endopetidase gene homologues in phytoplasma genomes [141] were described as potentially affecting host plants infected with different phytoplasmas.
7. What Is New in the Field?
Many workers, mainly plant pathologists rather than mycoplasmologists, took up the challenge to culture these organisms. An early outcome of these efforts was successful axenic culture of spiroplasmas: this distinct group of organisms grew readily in media formulations whereas the phytoplasmas did not. The preliminary evidence that phytoplasmas can be grown in or on cell free laboratory media has been reported [142] -[144] . This is an important breakthrough in the study of their biology since, despite a reduced genome size in comparison to their ancestors, phytoplasmas retain an independent metabolism that allows them to survive as parasites in environments as diverse as plant phloem and insect haemolymph. This versatility is a unique property among microbes, shared only with some animalor plant-infecting viruses and with a few other microorganisms such as the causal agent of malaria.
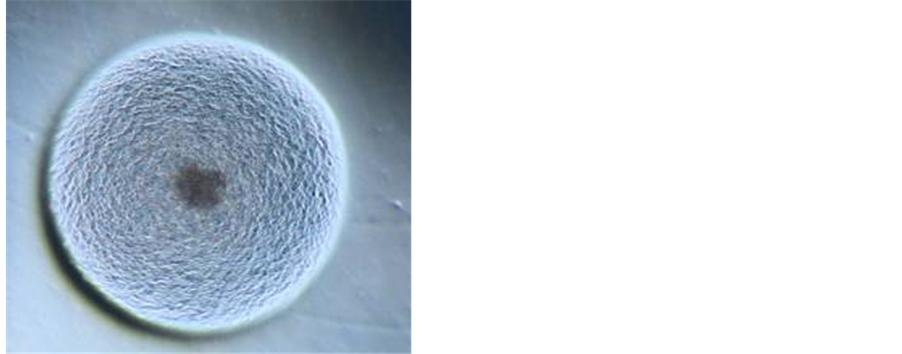
Employing specific commercially available media, it was possible to achieve growth of phytoplasmas belonging to diverse ribosomal groups from micropropagated periwinkle shoots, infected with phytoplasmas, as an initial source of inoculum. In spite of the relatively long time required for incubation in liquid medium, colony growth of phytoplasmas on agar usually occurs within two to five days, as for the majority of cultured bacteria. Although further research is needed and is in progress in order to optimize the culture system, the prospect of routine cultivation is now a real option for researchers. When observed with a binocular microscope, the phytoplasma colonies on the surface of agar medium are comparable in size and appearance to those of mycoplasmas (Figure 3). PCR assays using phytoplasma specific primers confirmed that phytoplasma DNA was present in those cultured cells used as a source of DNA template. Identification using RFLP analysis and direct sequencing of selected amplicons also confirmed phytoplasma identity [143] .
From the preliminary cultivation work a patent has been submitted to cover the commercial exploitation of the methodology, however, research carried out for scientific purposes is not restricted by the patent. A major advantage of the procedure is that it may now be possible to build a comprehensive culture collection of diverse phytoplasmas without the need to maintain these strains by micropropagation methods. One of the most immediate benefits is a genome sequencing to compare and contrast gene complement and metabolic pathways of
 (a)
(a)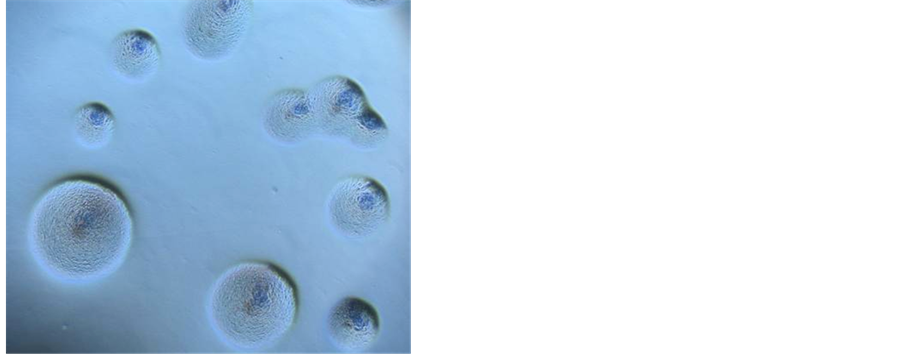 (b)
(b)
Figure 3. Phytoplasma colonies under optical microscope [(a) magnification 50×, (b) magnification 20×].
strains representative of each phytoplasma 16Sr group. Currently genome sequencing is very inefficient, largely because of the intimate association of phytoplasmas with their respective host plants and insect vectors, therefore the availability of phytoplasma colonies should help. It is likely that whole genome comparisons will emerge as the premier tool for phytoplasma identification in the near future.
As a consequence, strategies targeting treatment or prevention of phytoplasma diseases could be better defined and more effective. In addition, more detailed knowledge about basic mechanisms that regulate the survival of phytoplasmas, which are among the smallest known self replicating organisms, should be obtained. These researches are urgently needed because the use of tetracycline is forbidden in some agricultural areas (Europe) since it is perceived to endanger human health. The recognition of durable resistance to phytoplasma diseases is the only effective long term mean of controlling these diseases. Selection and screening of plants resistant to phytoplasma infection as well as the study of the modes of colonization by phytoplasmas of plant and insect vectors are other possible applications. As a consequence strategies aiming to treat and/or prevent phytoplasma related plant diseases could be better defined and more effective. The hoped-for result would be enhanced ability to reduce the socio-economic impact that phytoplasma diseases have in all agricultural areas of the world.
References
- Doi, Y., Teranaka, M., Yora, K. and Asuyama, H. (1967) Mycoplasma or PLT Grouplike Microrganisms Found in the Phloem Elements of Plants Infected with Mulberry Dwarf, Potato Witches’ Broom, Aster Yellows or Pawlownia Witches’ Broom. Japanese Journal of Phytopathology, 33, 259-266. http://dx.doi.org/10.3186/jjphytopath.33.259
- Ishii, Y., Kakizawa, S., Hoshi, A., Maejima, K., Kagiwada, S., Yamaji, Y., Oshima, K. and Namba, S. (2009) In the Non-Insect-Transmissible Line of Onion Yellows Phytoplasma (OY-NIM), the Plasmid-Encoded Transmembrane Protein ORF3 Lacks the Major Promoter Region. Microbiology, 155, 2058-2067. http://dx.doi.org/10.1099/mic.0.027409-0
- Ishiie, T., Doi, Y., Yora, K. and Asuyama, H. (1967) Suppressive Effects of Antibiotics of Tetracycline Group on Symptom Developement of Mulberry Dwarf Disease. Japanese Journal of Phytopathology, 33, 267-275. http://dx.doi.org/10.3186/jjphytopath.33.267
- Bertaccini, A. (2007) Phytoplasmas: Diversity, Taxonomy, and Epidemiology. Frontiers in Bioscience, 12, 673-689. http://dx.doi.org/10.2741/2092
- Hogenhout, S.A., Oshima, K., Ammar, E.D., Kakizawa, S., Kingdom, H.N. and Namba, S. (2008) Phytoplasmas: Bacteria that Manipulate Plants and Insects. Molecular Plant Pathology, 9, 403-423. http://dx.doi.org/10.1111/j.1364-3703.2008.00472.x
- Gasparich, G.E., Whitcom, R.F., Dodge, D., French, F.E., Glass, J. and Williamson, D.L. (2004) The Genus Spiroplasma and Its Non-Helical Descendants: Phylogenetic Classification, Correlation with Phenotype and Roots of the Mycoplasma mycoides Clade. International Journal of Systematic and Evolutionary Microbiology, 54, 893-918. http://dx.doi.org/10.1099/ijs.0.02688-0
- Lim, P.O. and Sears, B.B. (1992) Evolutionary Relationships of a Plant-Pathogenic Mycoplasmalike Organisms and Acholeplasma laidlawii Deduced from Two Ribosomal Protein Gene Sequences. Journal of Bacteriology, 174, 2606- 2611.
- Andersen, M.T., Liefting, L.W., Havukkala, I. and Beever, R.E. (2013) Comparison of the Complete Genome Sequence of Two Closely Related Isolates of ‘Candidatus Phytoplasma australiense’ Reveals Genome Plasticity. BMC Genomics, 14, 529. http://dx.doi.org/10.1186/1471-2164-14-529
- Bai, X., Zhang, J., Ewing, A., Miller, S.A., Radek, A.J., Shevchenko, D.V., Tsukerman, K., Walunas, T., Lapidus, A., Campbell, J.W. and Hogenhout, S.A. (2006) Living with Genome Instability: The Adaptation of Phytoplasmas to Diverse Environments of Their Insect and Plant Hosts. Journal of Bacteriology, 188, 3682-3696. http://dx.doi.org/10.1128/JB.188.10.3682-3696.2006
- Kube, M., Schneider, B., Kuhl, H., Dandekar, T., Heitmann, K., Migdo, A.M., Reinhardt, R. and Seemüller, E. (2008) The Linear Chromosome of the Plant-Pathogenic Mycoplasma ‘Candidatus Phytoplasma mali’. BMC Genomics, 9, 306. http://dx.doi.org/10.1186/1471-2164-9-306
- Oshima, K., Kakizawa, S., Nishigawa, H., Jung, H.Y., Wei, W., Suzuki, S., Arashida, R., Nakata, D., Miyata, S., Ugaki, M. and Namba, S. (2004) Reductive Evolution Suggested from the Complete Genome Sequence of a Plant-Pathogenic Phytoplasma. Nature Genetics, 36, 27-29. http://dx.doi.org/10.1038/ng1277
- Tran-Nguyen, L.T., Kube, M., Schneider, B., Reinhardt, R. and Gibb, K.S. (2008) Comparative Genome Analysis of ‘Candidatus Phytoplasma australiense’ (Subgroup tuf-Australia I; rp-A) and ‘Ca. Phytoplasma asteris’ Strains OY-M and AY-WB. Journal of Bacteriology, 190, 3979-3991. http://dx.doi.org/10.1128/JB.01301-07
- Hoshi, A., Oshima, K., Kakizawa, S., Ishii, Y., Ozeki, J., Hashimoto, M., Komatsu, K., Kagiwada, S., Yamaji, Y. and Namba, S. (2009) A Unique Virulence Factor for Proliferation and Dwarfism in Plants Identified from a Phytopathogenic Bacterium. Proceedings of the National Academy of Sciences of the United States of America, 106, 6416-6421. http://dx.doi.org/10.1073/pnas.0813038106
- Oshima, K., Kakizawa, S., Arashida, R., Ishii, Y., Hoshi, A., Hayashi, Y., Kawagida, S. and Namba, S. (2007) Presence of Two Glycolityc Gene Clusters in a Severe Pathogenic Line of ‘Candidatus Phytoplasma asteris’. Molecular Plant Pathology, 8, 481-489. http://dx.doi.org/10.1111/j.1364-3703.2007.00408.x
- Suzuki, S., Oshima, K., Kakizawa, S., Arashida, R., Jung, H.Y., Yamaji, Y., Nishigawa, H., Ugaki, M. and Namba, S. (2006) Interaction between the Membrane Protein of a Pathogen and Insect Microfilament Complex Determines InsectVector Specificity. Proceedings of the National Academy of Sciences of the United States of America, 103, 4252-4257. http://dx.doi.org/10.1073/pnas.0508668103
- Lee, I.-M., Gundersen, D.E., Hammond, R.W. and Davis, R.E. (1994) Use of Mycoplasmalike Organism (MLO) GroupSpecific Oligonucleotide Primers for Nested-PCR Assays to Detect Mixed-MLO Infections in a Single Host Plant. Phytopathology, 84, 559-566. http://dx.doi.org/10.1094/Phyto-84-559
- Lee, I.-M., Bertaccini, A., Vibio, M. and Gundersen, D.E. (1995) Detection of Multiple Phytoplasmas in Perennial Fruit Trees with Decline Symptoms in Italy. Phytopathology, 85, 728-735. http://dx.doi.org/10.1094/Phyto-85-728
- Cimerman, A., Pacifico, D., Salar, P., Marzachì, C. and Foissac, X. (2009) Striking Diversity of vmp1, a Variable Gene Encoding a Putative Membrane Protein of the Stolbur Phytoplasma. Applied and Environmental Microbiology, 75, 2951-2957. http://dx.doi.org/10.1128/AEM.02613-08
- Gundersen, D.E., Lee, I.-M., Schaff, D.A., Harrison, N.A., Chang, C.J., Davis, R.E. and Kinsbury, D.T. (1996) Genomic Diversity and Differentiation among Phytoplasma Strains in 16S rRNA group I (Aster Yellows and Related Phytoplasmas) and III (X-Disease and Related Phytoplasmas). International Journal of Systematic Bacteriology, 46, 64-75. http://dx.doi.org/10.1099/00207713-46-1-64
- Lee, I.-M., Zhao, Y. and Bottner, K.D. (2006) SecY Gene Sequence Analysis for Finer Differentiation of Diverse Strains in the Aster Yellows Phytoplasma Group. Molecular and Cellular Probes, 20, 87-91. http://dx.doi.org/10.1016/j.mcp.2005.10.001
- Lee, I.-M., Bottner-Parker, K.D., Zhao, Y., Davis, R.E. and Harrison, N.A. (2010) Phylogenetic Analysis and Delineation of Phytoplasmas Based on secY Gene Sequences. International Journal of Systematic and Evolutionary Microbiology, 60, 2887-2897. http://dx.doi.org/10.1099/ijs.0.019695-0
- Lee, I.-M., Bottner-Parker, K.D., Zhao, Y., Bertaccini, A. and Davis, R.E. (2012) Differentiation and Classification of Phytoplasmas in the Pigeon Pea Witches’ Broom Group (16SrIX): An Update Based on Multiple Gene Sequence Analysis. International Journal of Systematic and Evolutionary Microbiology, 62, 2279-2285. http://dx.doi.org/10.1099/ijs.0.038273-0
- Marcone, C., Lee, I.-M., Davis, R.E., Ragozzino, A. and Seemüller, E. (2000) Classification of Aster Yellows-Group Phytoplasmas Based on Combined Analyses of rRNA and Tuf Gene Sequences. International Journal of Systematic and Evolutionary Microbiology, 50, 1703-1713.
- Martini, M., Botti, S., Marcone, C., Marzachì, C., Casati, P., Bianco, P.A., Benedetti, R. and Bertaccini, A. (2002) Genetic Variability among Flavescence Dorée Phytoplasmas from Different Origins in Italy and France. Molecular and Cellular Probes, 16, 197-208. http://dx.doi.org/10.1006/mcpr.2002.0410
- Martini, M., Lee, I.-M., Bottner, K.D., Zhao, Y., Botti, S., Bertaccini, A., Harrison, N.A., Carraro, L., Marcone, C., Khan, J. and Osler, R. (2007) Ribosomal Protein Gene-Based Filogeny for Finer Differentiation and Classification of Phytoplasmas. International Journal of Systematic and Evolutionary Microbiology, 57, 2037-2051. http://dx.doi.org/10.1099/ijs.0.65013-0
- Schneider, B., Gibb, K.S. and Seemüller, E. (1997) Sequence and RFLP Analysis of the Elongation Factor Tu Gene Used in Differentiation and Classification of Phytoplasmas. Microbiology, 143, 3381-3389. http://dx.doi.org/10.1099/00221287-143-10-3381
- Makarova, O.V., Contaldo, N., Paltrinieri, S., Kawube, G., Bertaccini, A. and Nicolaisen, M. (2012) DNA Barcoding for Universal Identification of ‘Candidatus Phytoplasmas’ Using a Fragment of the Elongation Factor Tu Gene. PLOS ONE, 7, Article ID: e52092. http://dx.doi.org/10.1371/journal.pone.0052092
- Baric, S. and Dalla-Via, I. (2004) A New Approach to Apple Proliferation Detection: A Highly Sensitive Real-Time PCR Assay. Journal of Microbiological Methods, 57, 135-145. http://dx.doi.org/10.1016/j.mimet.2003.12.009
- Christensen, N.M., Nicolaisen, M., Hansen, M. and Schultz, A. (2004) Distribution of Phytoplasmas in Infected Plants as Revealed by Real Time PCR and Bioimaging. Molecular Plant-Microbe Interactions, 17, 1175-1184. http://dx.doi.org/10.1094/MPMI.2004.17.11.1175
- Marzachì, C. and Bosco, D. (2005) Relative Quantification of Chrysanthemum Yellows (16Sr I) Phytoplasma in Its Plant and Insect Host Using Real-Time Polymerase Chain Reaction. Molecular Biotechnology, 30, 117-127. http://dx.doi.org/10.1385/MB:30:2:117
- Bertaccini, A., Davis, R.E. and Lee, I.-M. (1992) In Vitro Micropropagation for Maintenance of Mycoplasmalike Organisms in Infected Plant Tissues. HortScience, 27, 1041-1043.
- Jarausch, W., Lansac, M. and Dosba, F. (1996) Long-Term Maintenance of Nonculturable Apple-Proliferation Phytoplasmas in Their Micropropagated Natural Host Plant. Plant Pathology, 45, 778-786. http://dx.doi.org/10.1046/j.1365-3059.1996.d01-171.x
- Tian, J.B., Bertaccini, A., Martini, M., Paltrinieri, S., Guo, H.P. and Pastore, M. (2000) Molecular Detection of Jujube Witches’-Broom Phytoplasmas in Micropropagated Jujube Shoots. HortScience, 35, 1274-1275.
- Bertaccini, A., Bellardi, M.G. and Vibio, M. (1996) Virus Diseases of Ornamental Shrubs. X. Euphorbia Pulcherrima Willd. Infected by Viruses and Phytoplasmas. Phytopathologia Mediterranea, 35, 129-132.
- Lee, I.M., Klopmeyer, M., Bartoszyk, I.M., Gundersen-Rindal, D.E., Chou, T., Thomson, K.L. and Eisenreich, R. (1997) Phytoplasma Induced Free-Branching in Commercial Poinsettia Cultivars. Nature Biotechnology, 15, 178-182. http://dx.doi.org/10.1038/nbt0297-178
- Bertaccini, A., Vibio, M. and Stefani, E. (1995) Detection and Molecular Characterization of Phytoplasmas Infecting Grapevine in Liguria (Italy). Phytopathologia Mediterranea, 34, 137-141.
- Daire, X., Clair, D., Reinert, W. and Boudon-Padieu, E. (1997) Detection and Differentiation of Grapevine Yellows Phytoplasmas Belonging to the Elm Yellows Group and to the Stolbur Subgroup by PCR Amplification of Nonribosomal DNA. European Journal of Plant Pathology, 103, 507-514. http://dx.doi.org/10.1023/A:1008641411025
- Martini, M., Murari, E., Mori, N. and Bertaccini, A. (1999) Identification and Epidemic Distribution of Two Flavescence Dorée-Related Phytoplasmas in Veneto (Italy). Plant Disease, 83, 925-930. http://dx.doi.org/10.1094/PDIS.1999.83.10.925
- Mori, N., Bressan, A., Martini, M., Guadagnini, M., Girolami, V. and Bertaccini, A. (2002) Experimental Transmission by Scaphoideus titanus Ball of Two Flavescence dorée-Type Phytoplasmas. Vitis, 41, 99-102.
- Angelini, E., Clair, D., Borgo, M., Bertaccini, A. and Boudon-Padieu, E. (2001) Flavescence dorée in France and Italy —Occurrence of Closely Related Phytoplasma Isolates and Their Near Relationships to Palatinate Grapevine Yellows and an Alder Yellows Phytoplasma. Vitis, 40, 79-86.
- Torres, E., Botti, S., Rahola, J., Martin, M.P. and Bertaccini, A. (2005) Grapevine Yellows Diseases in Spain: Eight Years Survey of Disease Spread and Molecular Characterization of Phytoplasmas Involved. Anales del Jardín Botánico de Madrid, 62, 127-133. http://dx.doi.org/10.3989/ajbm.2005.v62.i2.12
- Botti, S. and Bertaccini, A. (2006) Phytoplasma Infection trough Seed Transmission: Further Observations. 16th Congress IOM, Cambridge, 9-14 July 2006, 113,76.
- Duduk, B., Botti, S., Ivanović, M., Krstić, B., Dukić, N. and Bertaccini, A. (2004) Identification of Phytoplasmas Associated with Grapevine Yellows in Serbia. Journal of Phytopathology, 152, 575-579. http://dx.doi.org/10.1111/j.1439-0434.2004.00898.x
- Arnaud, G., Malembic-Maher, S., Salar, P., Maixner, M., Marcone, C., Boudon-Padieu, E. and Foissac, X. (2007) Multilocus Sequence Typing Confirms the Close Genetic Inter-Relatedness between Three Distinct Flavescence Dorée Phytoplasma Strain Clusters and Group 16SrV Phytoplasmas Infecting Grapevine and Alder in Europe. Applied and Environmental Microbiology, 73, 4001-4010. http://dx.doi.org/10.1128/AEM.02323-06
- Botti, S. and Bertaccini, A. (2007) Grapevine Yellows in Northern Italy: Molecular Identification of Flavescence Dorée Phytoplasma Strains and of Bois Noir Phytoplasmas. Journal of Applied Microbiology, 103, 2325-2330. http://dx.doi.org/10.1111/j.1365-2672.2007.03467.x
- Sforza, R., Clair, D., Daire, X., Larrue, J. and Boudon-Padieu, E. (1998) The Role of Hyalesthes obsoletus (Hemiptera: Cixiidae) in the Occurrence of Bois Noir of Grapevines in France. Journal of Phytopathology, 146, 549-556. http://dx.doi.org/10.1111/j.1439-0434.1998.tb04753.x
- Langer, M. and Maixner, M. (2004) Molecular Characterisation of Grapevine Yellows Associated Phytoplasmas of the Stolburgroup Based on RFLP-Analysis of Non-Ribosomal DNA. Vitis, 43, 191-199.
- Mori, N., Pavan, F., Bondavalli, R., Reggiani, N., Paltrinieri, S. and Bertaccini, A. (2008) Factors Affecting the Spread of “Bois Noir” Disease in North Italy Vineyards. Vitis, 47, 65-72.
- Alma, A., Davis, R.E., Vibio, M., Danielli, A., Bosco, D., Arzone, A. and Bertaccini, A. (1996) Mixed Infection of Grapevines in Northern Italy by Phytoplasmas Including 16S rRNA RFLP Subgroup 16SrI-B Strains Previously Unreported in this Host. Plant Disease, 80, 418-421. http://dx.doi.org/10.1094/PD-80-0418
- Engelbrecht, M., Joubert, J. and Burger, T.J. (2010) First Report of Aster Yellows Phytoplasma in Grapevines in South Africa. Plant Disease, 94, 373. http://dx.doi.org/10.1094/PDIS-94-3-0373A
- Gajardo, A., Fiore, N., Prodan, S., Paltrinieri, S., Botti, S., Pino, A.M., Zamorano, A., Montealegre, J. and Bertaccini, A. (2009) Phytoplasmas Associated with Grapevine Yellows Disease in Chile. Plant Disease, 93, 789-796. http://dx.doi.org/10.1094/PDIS-93-8-0789
- Padovan, A., Gibb, K. and Persley, D. (2000) Association of ‘Candidatus Phytoplasma australiense’ with Green Petal and Lethal Yellows Diseases in Strawberry. Plant Pathology, 49, 362-369. http://dx.doi.org/10.1046/j.1365-3059.2000.00461.x
- Seemüller, E. and Schneider, B. (2004) ‘Candidatus Phytoplasma mali’, ‘Candidatus Phytoplasma pyri’ and ‘Candidatus Phytoplasma prunorum’, the Causal Agents of Apple Proliferation, Pear Decline and European Stone Fruit Yellows, Respectively. International Journal of Systematic and Evolutionary Microbiology, 54, 1217-1226. http://dx.doi.org/10.1099/ijs.0.02823-0
- Seemüller, E., Kiss, E., Sule, S. and Schneider, B. (2010) Multiple Infection of Apple Trees by Distinct Strains of ‘Candidatus Phytoplasma mali’ and Its Pathological Relevance. Phytopathology, 100, 863-870. http://dx.doi.org/10.1094/PHYTO-100-9-0863
- Carraro, L., Osler, R., Loi, N., Ermacora, P. and Refatti, E. (2001) Fruit Tree Phytoplasma Diseases Diffused in Nature by Psyllids. Acta Horticulturae, 550, 345-350.
- Carraro, L., Loi, N., Ermacora, P. and Osler, R. (1998) High Tolerance of European Plum Varieties to Plum Leptonecrosis. European Journal of Plant Pathology, 104, 141-145. http://dx.doi.org/10.1023/A:1008617531529
- Carraro, L., Osler, R., Loi, N., Ermacora, P. and Refatti, E. (1998) Transmission of European Stone Fruit Yellows Phytoplasma by Cacopsylla pruni. Journal of Plant Pathology, 80, 233-239.
- Blomquist, C.L. and Kirkpatrick, B.C. (2002) Identification of Phytoplasma Taxa and Insect Vectors of Peach Yellow Leaf Roll Disease in California. Plant Disease, 86, 759-763. http://dx.doi.org/10.1094/PDIS.2002.86.7.759
- Kirkpatrick, B.C., Uyemoto, J.K. and Purcell, A.H. (1995) X-Disease. In: Compendium of Stone Fruit Diseases, American Phytopathological Society, St. Paul, 57.
- Molino Lova, M., Quaglino, F., Abou-Jawdah, Y., Choueiri, E., Sobh, H., Casati, P., Tedeschi, R., Alma, A. and Bianco, P.A. (2011) Identification of New 16SrIX Subgroups, -F and -G, among ‘Candidatus Phytoplasma phoenicium’ Strains Infecting Almond, Peach and Nectarine in Lebanon. Phytopathologia Mediterranea, 50, 273-282.
- Salehi, M., Izadpanah, K. and Heydarnejad, J. (2006) Characterization of a New Almond Witches’ Broom Phytoplasma in Iran. Journal of Phytopathology, 154, 386-391. http://dx.doi.org/10.1111/j.1439-0434.2006.01109.x
- Zirak, L., Bahar, M. and Ahoonmanesh, A. (2009) Molecular Characterization of Phytoplasmas Related to Peanut Witches Broom and Stolbur Groups Infecting Plum in Iran. Journal of Plant Pathology, 91, 713-716.
- Zirak, L., Bahar, M. and Ahoonmanesh, A. (2010) Molecular Characterization of Phytoplasmas Associated with Peach Diseases in Iran. Journal of Phytopathology, 158, 105-110. http://dx.doi.org/10.1111/j.1439-0434.2009.01585.x
- Zhu, S.F., Bertaccini, A., Lee, I.-M., Paltrinieri, S. and Hadidi, A. (2011) Cherry Lethal Yellows and Decline Phytoplasmas. In: Hadidi, A., Barba, M., Candresse, T. and Jelkmann, W., Eds., Virus and Virus-Like Diseases of Pome and Stone Fruits, APS Press, St. Paul, 255-257.
- Mardi, M., Khayam Nekouei, S.M., Karimi Farsad, L., Ehya, F., Shabani, M., Shafiee, M., Tabatabaei, M., Safarnejad, M.R., Jouzani, G.S. and Salekdeh, G.H. (2011) Witches’ Broom Disease of Mexican Lime Trees: Disaster to Be Addressed before It Will Be Too Late. Bulletin of Insectology, 64, S205-S206.
- Chen, J., Pu, X., Deng, X., Liu, S., Li, H. and Civerolo, E. (2008) A Phytoplasma Closely Related to the Pigeon Pea Witches’-Broom Phytoplasma (16SrIX) Is Associated with Citrus Huanglongbing Symptoms in the State of São Paulo, Brazil. Phytopathology, 98, 977-984. http://dx.doi.org/10.1094/PHYTO-98-9-0977
- Teixeira, D.C., Wulff, N.A., Martins, E.C., Kitajima, E.W., Bassanezi, R., Ayres, A.J., Eveillard, S., Saillard, C. and Bové, J.M. (2009) A Phytoplasma Related to ‘Candidatus Phytoplasma asteri’ Detected in Citrus Showing Huanglongbing (Yellow Shoot Disease) Symptoms in Guangdong, P. R. China. Phytopathology, 99, 236-242. http://dx.doi.org/10.1094/PHYTO-99-3-0236
- Harrison, N.A. and Jones, P. (2004) Lethal Yellowing. In: Elliott, M.L., Broschat, T.K., Uchida, J.Y. and Simone, G.W., Eds., Compendium of Ornamental Palm Diseases and Disorders, APS Press, St. Paul, 39-41.
- Mpunami, A.A., Tymon, A., Jones, P. and Dickinson, M.J. (1999) Genetic Diversity in the Coconut Lethal Yellowing Disease Phytoplasmas of East Africa. Plant Pathology, 48, 109-114. http://dx.doi.org/10.1046/j.1365-3059.1999.00314.x
- Nejat, N., Sijam, K., Abdullah, S., Vadamalai, G. and Dickinson, M. (2009) Phytoplasmas Associated with Disease of Coconut in Malaysia: Phylogenetic Groups and Host Plant Species. Plant Pathology, 58, 1152-1160. http://dx.doi.org/10.1111/j.1365-3059.2009.02153.x
- Nejat, N., Vadamalai, G., Davis, R.E., Harrison, N.A., Sijam, K., Dickinson, M., Abdullah, S.N. and Zhao, Y. (2012) ‘Candidatus Phytoplasma malaysianum’, a Novel Taxon Associated with Virescence and Phyllody of Madagascar periwinkle (Catharanthus roseus). International Journal of Systematic and Evolutionary Microbiology, 63, 540-548. http://dx.doi.org/10.1099/ijs.0.041467-0
- Brown, S.E., Been, B.O. and McLaughlin, W.A. (2006) Detection and Variability of the Lethal Yellowing Group (16SrIV) Phytoplasma in the Cedusa sp. (Hemiptera: Auchenorrhyncha: Derbidae) in Jamaica. Annals of Applied Biology, 149, 53-62. http://dx.doi.org/10.1111/j.1744-7348.2006.00072.x
- Alhudaib, K., Arocha, Y., Wilson, M. and Jones, P. (2008) First Report of a 16SrI, ‘Candidatus Phytoplasma asteris’ Group Phytoplasma Associated with a Date Palm Disease in Saudi Arabia. Plant Pathology, 57, 366. http://dx.doi.org/10.1111/j.1365-3059.2007.01667.x
- Cronjé, P., Dabek, A.J., Jones, P. and Tymon, A.M. (2000) First Report of a Phytoplasma Associated with a Disease of Date Palms in North Africa. Plant Pathology, 49, 801. http://dx.doi.org/10.1046/j.1365-3059.2000.00504.x
- Cronjé, P., Dabek, A.J., Jones, P. and Tymon, A.M. (2000) Slow Decline: A New Disease of Mature Date Palms in North Africa Associated with a Phytoplasma. Plant Pathology, 49, 804. http://dx.doi.org/10.1046/j.1365-3059.2000.00507.x
- Perera, L., Meegahakumbura, M.K., Wijesekara, H.R.T., Fernando, W.B.S. and Dickinson, M. (2012) A Phytoplasma Is Associated with Weligama Coconut Leaf Wilt Disease in Sri Lanka. Journal of Plant Pathology, 94, 205-209.
- Warokka, J. (2005) Studies on the Etiology and Epidemiology of Kalimantan Wilt Disease of Coconut in Indonesia. Dissertation, University of Nottingham, Nottingham.
- Alvarez, E., Mejia, J.F., Contaldo, N., Paltrinieri, S., Duduk, B. and Bertaccini, A. (2013) ‘Candidatus Phytoplasma asteris’ Strains Associated with Oil Palm Lethal Wilt in Colombia. Plant Disease, 98, 311-318.
- Eroglu, S., Ozbek, H. and Sahin, F. (2010) First Report of Group 16SrXII Phytoplasma Causing Stolbur Disease in Potato Plants in the Eastern and Southern Anatolia Regions of Turkey. Plant Disease, 94, 1374. http://dx.doi.org/10.1094/PDIS-06-10-0439
- Girsova, N., Bottner, K.D., Mozhaeva, K.A., Kastalyeva, T.B., Owens, R.A. and Lee, I.M. (2008) Molecular Detection and Identification of Group 16SrI and 16SrXII Phytoplasmas Associated with Diseased Potatoes in Russia. Plant Disease, 92, 654. http://dx.doi.org/10.1094/PDIS-92-4-0654A
- Hosseini, P., Bahar, M., Madani, G. and Zirak, L. (2011) Molecular Characterization of Phytoplasmas Associated with Potato Purple Top Disease in Iran. Journal of Phytopathology, 159, 241-246. http://dx.doi.org/10.1111/j.1439-0434.2010.01757.x
- Liefting, L.W., Veerakone, S., Ward, L.I. and Clover, G.R.G. (2009) First Report of ‘Candidatus Phytoplasma australiense’ in Potato. Plant Disease, 93, 969. http://dx.doi.org/10.1094/PDIS-93-9-0969A
- Mejia, J.F., Contaldo, N., Paltrinieri, S., Pardo, J.M., Rios, C.A., Alvarez, A. and Bertaccini, A. (2011) Molecular Detection and Identification of Group 16SrV and 16SrXII Phytoplasmas Associated with Potatoes in Colombia. Bulletin of Insectology, 64, S97-S98.
- Ebbert, M.A., Jeffers, D.P., Harrison, N.A. and Nault, L.R. (2001) Lack of Specificity in the Interaction between Two Maize Stunting Pathogens and Field Collected Dalbulus Leafhoppers. Entomologia Experimentalis et Applicata, 101, 49-57. http://dx.doi.org/10.1046/j.1570-7458.2001.00890.x
- Harrison, N.A., Richardson, P.A., Tsai, J.H., Ebbert, M.A. and Kramer, J.B. (1996) PCR Assay for Detection of the Phytoplasma Associated with Maize Bushy Stunt Disease. Plant Disease, 80, 263-269. http://dx.doi.org/10.1094/PD-80-0263
- Mejia, J.F., Zamora, L., Duduk, B., Quiñones, M.L., Bertaccini, A. and Contaldo, N. (2013) Caratterizzazione Molecolare di Ceppi di ‘Candidatus Phytoplasma asteris’ da Mais. Petria, 23, 135-138.
- Duduk, B. and Bertaccini, A. (2006) Corn with Symptoms of Reddening: New Host of Stolbur Phytoplasma. Plant Disease, 90, 1313-1319. http://dx.doi.org/10.1094/PD-90-1313
- Jović, J., Cvrković, T., Mitrović, M., Krnjajić, S., Petrović, A., Redinbaugh, M.G., Pratt, R.C., Hogenhout, S.A. and Toševski, I. (2009) Stolbur Phytoplasma Transmission to Maize by Reptalus Panzeri and the Disease Cycle of Maize Redness in Serbia. Phytopathology, 99, 1053-1061. http://dx.doi.org/10.1094/PHYTO-99-9-1053
- Mori, N., Mitrović, J., Smiljković, M., Duduk, N., Paltrinieri, S., Bertaccini, A. and Duduk, B. (2013) Population Dynamic and Role of Hyalesthes obsoletus Signoret (Homoptera, Cixiidae) in Corn Reddening Transmission in Serbia. Bulletin of Insectology, 66, 245-250.
- Lee, I.M., Gundersen-Rindal, D.E. and Bertaccini, A. (1998) Phytoplasma: Ecology and Genomic Diversity. Phytopathology, 88, 1359-1366. http://dx.doi.org/10.1094/PHYTO.1998.88.12.1359
- Lee, I.-M., Gundersen-Rindal, D.E., Davis, R.E. and Bartoszyk, I.M. (1998) Revised Classification Scheme of Phytoplasmas Based on RFLP Analyses of 16S rRNA and Ribosomal Protein Gene Sequences. International Journal of Systematic and Evolutionary Microbiology, 48, 1153-1169. http://dx.doi.org/10.1099/00207713-48-4-1153
- IRPCM (2004) ‘Candidatus Phytoplasma’, a Taxon for the Wall-Less, Non-Helical Prokaryotes that Colonise Plant Phloem and Insects. International Journal of Systematic and Evolutionary Microbiology, 54, 1243-1255. http://dx.doi.org/10.1099/ijs.0.02854-0
- Duduk, B. and Bertaccini, A. (2011) Phytoplasma Classification: Taxonomy Based on 16S Ribosomal Gene, Is It Enough? Phytopathogenic Mollicutes, 1, 1-13.
- Marcone, C., Neimark, H., Ragozzino, A., Lauer, U. and Seemüller, E. (1999) Chromosome Sizes of Phytoplasmas Composing Major Phylogenetic Groups and Subgroups. Phytopathology, 89, 805-810. http://dx.doi.org/10.1094/PHYTO.1999.89.9.805
- Mitrović, J., Kakizawa, S., Duduk, B., Oshima, K., Namba, S. and Bertaccini, A. (2011) The cpn60 Gene as an Additional Marker for Finer Differentiation of ‘Candidatus Phytoplasma asteris’ Related Strains. Annals of Applied Biology, 159, 41-48. http://dx.doi.org/10.1111/j.1744-7348.2011.00472.x
- Wei, W., Kakizawa, S., Suzuki, S., Jung, H.Y., Nishigawa, H., Miyata, S., Oshima, K., Ugaki, M., Hibi, T. and Namba, S. (2004) In planta Dynamic Analysis of Onion Yellows Phytoplasma Using Localized Inoculation by Insect Transmission. Phytopathology, 94, 244-250. http://dx.doi.org/10.1094/PHYTO.2004.94.3.244
- Garcia-Chapa, M., Medina, V., Viruel, M.A., Lavina, A. and Battle, A. (2003) Seasonal Detection of Pear Decline Phytoplasma by Nested-PCR in Different Pear Cultivars. Plant Pathology, 52, 513-520. http://dx.doi.org/10.1046/j.1365-3059.2003.00868.x
- Christensen, N.M., Axelsen, K.B., Nicolaisen, M. and Schultz, A. (2005) Phytoplasmas and Their Interactions with Their Hosts. Trends in Plant Science, 10, 526-535. http://dx.doi.org/10.1016/j.tplants.2005.09.008
- Alma, A., Bosco, D., Danielli, A., Bertaccini, A., Vibio, M. and Arzone, A. (1997) Identification of Phytoplasmas in Eggs, Nymphs and Adults of Scaphoideus titanus Ball Reared on Healthy Plants. Insect Molecular Biology, 6, 115-121. http://dx.doi.org/10.1111/j.1365-2583.1997.tb00079.x
- Kawakita, H., Saiki, T., Wei, W., Mitsuhashi, W., Watanabe, K. and Sato, M. (2000) Identification of Mulberry Dwarf Phytoplasmas in the Genital Organs and Eggs of Leafhopper Hishimonoides sellatiformis. Phytopathology, 90, 909- 914. http://dx.doi.org/10.1094/PHYTO.2000.90.8.909
- Hanboonsong, Y., Choosai, C., Panyim, S. and Damak, S. (2002) Transovarial Transmission of Sugarcane White Leaf Phytoplasma in the Insect Vector Matsumuratettix hiroglyphicus (Matsumura). Insect Molecular Biology, 11, 97-103. http://dx.doi.org/10.1046/j.0962-1075.2001.00314.x
- Tedeschi, R., Ferrato, V., Rossi, J. and Alma, A. (2006) Possible Phytoplasma Transovarial Transmission in the Psyllids Cacopsylla melanoneura and Cacopsylla pruni. Plant Pathology, 55, 18-24.
- Cordova, I., Jones, P. Harrison, N.A. and Oropeza, C. (2003) In Situ Detection of Phytoplasma DNA in Embryos from Coconut Palms with Lethal Yellowing Disease. Molecular Plant Pathology, 4, 99-108. http://dx.doi.org/10.1046/j.1364-3703.2003.00152.x
- Botti, S. and Bertaccini, A. (2006) FD-Related Phytoplasmas and Their Association with Epidemic and Non Epidemic Situations in Tuscany (Italy). 15th Meeting of the International Council for the Study of Virus and Virus-Like Diseases of the Grapevine (ICVG), Stellenbosch, 3-7 April 2006, 163-164.
- Calari, A., Paltrinieri, S., Contaldo, N., Sakalieva, D., Mori, N., Duduk, B. and Bertaccini, A. (2011) Molecular Evidence of Phytoplasmas in Winter Oilseed Rape, Tomato and Corn Seedlings. Bulletin of Insectology, 64, S157-S158.
- Khan, A.J., Botti, S., Paltrinieri, S., Al-Subhi, A.M. and Bertaccini, A. (2002) Phytoplasmas in Alfalfa Seedlings: Infected or Contaminated Seeds? 13th Congress IOM, Vienna, 7-12 July 2002, 6.
- Jomantiene, R. and Davis, R.E. (2006) Clusters of Diverse Genes Existing as Multiple, Sequence-Variable Mosaics in a Phytoplasma Genome. FEMS Microbiology Letters, 255, 59-65. http://dx.doi.org/10.1111/j.1574-6968.2005.00057.x
- Jomantiene, R., Zhao, Y. and Davis, R.E. (2007) Sequence-Variable Mosaics: Composites of Recurrent Transposition Characterizing the Genomes of Phylogenetically Diverse Phytoplasmas. DNA and Cell Biology, 26, 557-564.
- Wei, W., Davis, R.E., Jomantiene, R. and Zhao, Y. (2008) Ancient, Recurrent Phage Attacks and Recombination Shaped Dynamic Sequence-Variable Mosaics at the Root of Phytoplasma Genome Evolution. Proceedings of the National Academy of Sciences of the United States of America, 105, 11827-11832. http://dx.doi.org/10.1073/pnas.0805237105
- Kuboyama, T., Huang, C., Lu, X., Sawayanagi, T., Kanazawa, T., Kagami, T., Matsuda, I., Tsuchizaki, T. and Namba, S. (1998) A Plasmid Isolated from Phytopathogenic Onion Yellows Phytoplasma and Its Heterogeneity in the Pathogenic Phytoplasma Mutant. Molecular Plant-Microbe Interactions, 11, 1031-1037. http://dx.doi.org/10.1094/MPMI.1998.11.11.1031
- Nishigawa, H., Miyata, S., Oshima, K., Sawayanagi, T., Komoto, A., Kuboyama, A., Matsuda, I., Tsuchizaki, T. and Namba, S. (2001) In Planta Expression of a Protein Encoded by the Extrachromosomal DNA of a Phytoplasma and Related to Geminivirus Replication Proteins. Microbiology, 147, 507-513.
- Nishigawa, H., Oshima, K., Kakizawa, S., Jung, H., Kuboyama, T., Miyata, S., Ugaki, M. and Namba, S. (2002) Evidence of Intermolecular Recombination between Extrachromosomal DNAs in Phytoplasma: A Trigger for the Biological Diversity of Phytoplasma. Microbiology, 148, 1389-1396.
- Oshima, K., Kakizawa, S., Nishigawa, H., Kuboyama, T., Miyata, S., Ugaki, M. and Namba, S. (2001) A Plasmid of Phytoplasma Encodes a Unique Replication Protein Having both Plasmidand Virus-Like Domains: Clue to Viral Ancestry or Result of Virus/Plasmid Recombination. Virology, 285, 270-277. http://dx.doi.org/10.1006/viro.2001.0938
- Rekab, D., Carraro, L., Schneider, B., Seemüller, E., Chen, J., Chang, C.J., Locci, R. and Firrao, G. (1999) Geminivirus-Related Extrachromosomal DNAs of the X-Clade Phytoplasmas Share High Sequence Similarity. Microbiology, 145, 1453-1459. http://dx.doi.org/10.1099/13500872-145-6-1453
- Oshima, K., Shiomi, T., Kuboyama, T., Sawayanagi, T., Nishigawa, H. and Namba, S. (2001) Isolation and Characterization of Derivative Lines of the Onion Yellows Phytoplasma That Do Not Cause Stunting or Phloem Hyperplasia. Phytopathology, 91, 1024-1029. http://dx.doi.org/10.1094/PHYTO.2001.91.11.1024
- Kube, M., Mitrovic, J., Duduk, B., Rabus, R. and Seemüller, E. (2012) Current View on Phytoplasma Genomes and Encoded Metabolism. The Scientific World Journal, 2012, Article ID: 185942, 25 pages. http://dx.doi.org/10.1100/2012/185942
- Berg, M., Davies, D.L., Clark, M.F., Vetten, J., Maier, G. and Seemüller, E. (1999) Isolation of a Gene Encoding an Immunodominant Memebrane Protein Gene in the Apple Proliferation Phytoplasma and Expression and Characterization of the Gene Product. Microbiology, 145, 1937-1943. http://dx.doi.org/10.1099/13500872-145-8-1937
- Morton, A., Davies, D.L., Blomquist, C.L. and Barbara, D.J. (2003) Characterization of Homologues of the Apple Proliferation Immunodominant Membrane Protein Gene from Three Related Phytoplasmas. Molecular Plant Pathology, 4, 109-114. http://dx.doi.org/10.1046/j.1364-3703.2003.00155.x
- Blomquist, C.L., Barbara, D.J., Davies, D.L., Clark, M.F. and Kirkpatrick, B.C. (2001) An Immunodominant Membrane Protein Gene from the Western X-Disease Phytoplasma Is Distinct from Those of Other Phytoplasmas. Microbiology, 147, 571-580.
- Barbara, D.J., Morton, A., Clark, M.F. and Davies, D.L. (2002) Immunodominant Membrane Proteins from Two Phytoplasmas in the Aster Yellows Clade (Chlorante Aster Yellows and Clover Phyllody) Are Highly Divergent in the Major Hydrophilic Region. Microbiology, 148, 157-167.
- Liefting, L.W. and Kirkpatrick, B.C. (2003) Cosmid Cloning and Sample Sequencing of the Genome of the Uncultivable Mollicute, Western X-Disease Phytoplasma, Using DNA Purified by Pulsed-field Gel Electrophoresis. FEMS Microbiology Letters, 221, 203-211. http://dx.doi.org/10.1016/S0378-1097(03)00183-6
- Kakizawa, S., Oshima, K., Ishii, Y., Hoshi, A., Maejima, K., Jung, H.Y., Yamaji, Y. and Namba, S. (2009) Cloning of Immunodominant Membrane Protein Genes of Phytoplasmas and Their in Planta Expression. FEMS Microbiology Letters, 293, 91-101. http://dx.doi.org/10.1111/j.1574-6968.2009.01509.x
- Arashida, R., Kakizawa, S., Ishii, Y., Hoshi, A., Jung, H.Y., Kagiwada, S., Yamaji, Y., Oshima, K. and Namba, S. (2008) Cloning and Characterization of the Antigenic Membrane Protein (Amp) Gene and in Situ Detection of Amp from Malformed Flowers Infected with Japanese Hydrangea Phyllody Phytoplasma. Phytopathology, 98, 769-775. http://dx.doi.org/10.1094/PHYTO-98-7-0769
- Oshima, K., Ishii, Y., Kakizawa, S., Sugawara, K., Neriya, Y., Himeno, M., Minato, N., Miura, C., Shiraishi, T., Yamaji, Y. and Namba, S. (2011) Dramatic Transcriptional Changes in an Intracellular Parasite Enable Host Switching between Plant and Insect. PLoS ONE, 6, Article ID: e23242. http://dx.doi.org/10.1371/journal.pone.0023242
- Toruno, T.Y., Seruga Musić, M., Simi, S., Nicolaisen, M. and Hogenhout, S.A. (2010) Phytoplasma PMU1 Exists as Linear Chromosomal and Circular Extrachromosomal Elements and Has Enhanced Expression in Insect Vectors Compared with Plant Hosts. Molecular Microbiology, 77, 1406-1415. http://dx.doi.org/10.1111/j.1365-2958.2010.07296.x
- Kakizawa, S., Oshima, K., Kuboyama, T., Nishigawa, H., Jung, H.Y., Sawayanagi, T., Tsuchizaki, T., Miyata, S., Ugaki, M. and Namba, S. (2001) Cloning and Expression Analysis of Phytoplasma Protein Translocation Genes. Molecular Plant-Microbe Interactions, 14, 1043-1050. http://dx.doi.org/10.1094/MPMI.2001.14.9.1043
- Kakizawa, S., Oshima, K., Nishigawa, H., Jung, H.Y., Wei, W., Suzuki, S., Tanaka, M., Miyata, S., Ugaki, M. and Namba, S. (2004) Secretion Oimmunodominant Membrane Protein from Onion Yellows Phytoplasma through the Sec Protein-Translocation System in Escherichia coli. Microbiology, 150, 135-142. http://dx.doi.org/10.1099/mic.0.26521-0
- Wei, W., Kakizawa, S., Jung, H.Y., Suzuki, S., Tanaka, M., Nishigawa, H., Miyata, S., Oshima, K., Ugaki, M., Hibi, T. and Namba, S. (2004) An Antibody against the SecA Membrane Protein of One Phytoplasma Reacts with Those of Phylogenetically Different Phytoplasmas. Phytopathology, 94, 683-686. http://dx.doi.org/10.1094/PHYTO.2004.94.7.683
- Sugio, A., Kingdom, H.N., MacLean, A.M., Grieve, V.M. and Hogenhout, S.A. (2011) Phytoplasma Protein Effector SAP11 Enhances Insect Vector Reproduction by Manipulating Plant Development and Defense Hormone Biosynthesis. Proceedings of the National Academy of Sciences of the United States of America, 108, E1254-E1263. http://dx.doi.org/10.1073/pnas.1105664108
- MacLean, A.M., Sugio, A., Makarova, O.V., Findlay, K.C., Grieve, V.M., Toth, R., Nicolaisen, M. and Hogenhout, S.A. (2011) Phytoplasma Effector SAP54 Induces Indeterminate Leaf-Like Flower Development in Arabidopsis Plants. Plant Physiology, 157, 831-841. http://dx.doi.org/10.1104/pp.111.181586
- Sugio, A., MacLean, A.M., Kingdom, H.N., Grieve, V.M., Manimekalai, R. and Hogenhout, S.A. (2011) Diverse Targets of Phytoplasma Effectors: From Plant Development to Defense against Insects. Annual Review of Phytopathology, 49, 175-195. http://dx.doi.org/10.1146/annurev-phyto-072910-095323
- Sugio, A. and Hogenhout, S.A. (2012) The Genome Biology of Phytoplasma: Modulators of Plants and Insects. Current Opinion in Microbiology, 15, 247-254. http://dx.doi.org/10.1016/j.mib.2012.04.002
- Himeno, M., Neriya, Y., Minato, N., Miura, C., Sugawara, K., Ishii, Y., Yamaji, Y., Kakizawa, S., Oshima, K. and Namba, S. (2011) Unique Morphological Changes in Plant Pathogenic Phytoplasma-Infected Petunia Flowers Are Related to Transcriptional Regulation of Floral Homeotic Genes in an Organ-Specific Manner. Plant Journal, 67, 971- 979. http://dx.doi.org/10.1111/j.1365-313X.2011.04650.x
- Lepka, P., Atitt, M., Moll, E. and Seemüller, E. (1999) Effect of Phytoplasmal Infection on Concentration and Translocation of Carbohydrates and Amino Acids in Periwinkle and Tobacco. Physiological and Molecular Plant Pathology, 55, 59-68. http://dx.doi.org/10.1006/pmpp.1999.0202
- Maust, B.E., Espadas, F., Talavera, C., Aguilar, M., Santamaría, J.M. and Oropeza, C. (2003) Changes in Carbohydrate Metabolism in Coconut Palms Infected with the Lethal Yellowing Phytoplasma. Phytopathology, 93, 976-981. http://dx.doi.org/10.1094/PHYTO.2003.93.8.976
- Sears, B.B., Klomparens, K.L., Wood, J.I. and Schewe, G. (1997) Effect of Altered Levels of Oxygen and Carbon Dioxide on Phytoplasma Abundance in Oenothera Leaf Tip Cultures. Physiological and Molecular Plant Pathology, 50, 275-287. http://dx.doi.org/10.1006/pmpp.1997.0086
- Bertamini, M. and Nedunchezhian, N. (2001) Effect of Phytoplasma, Stolbur-Subgroup (Bois Noir-BN)] of Photosynthetic Pigments, Saccarides, Ribulose-1,5-Bisphosphate Carboxylase, Nitrate and Nitrite Reductases and Photosynthetic Activities in Field-Grow Grapevine (Vitis vinifera L. cv Chardonnay) Leaves. Photosynthetica, 39, 119-122. http://dx.doi.org/10.1023/A:1012412406727
- Bertamini, M., Nedunchezhian, N., Tomasi, F. and Grando, M.S. (2002) Phytoplasma [Stolbur-Subgroup (Bois Noir-BN)] Infection Inhibitis Photosynthetic Pigments, Ribulose-1,5-Biphosphate Carboxylase and Photosynthetic Activities in Field Grown Grapevine (Vitis vinifera L. cv. Chardonnay) Leaves. Physiological and Molecular Plant Pathology, 61, 357-366. http://dx.doi.org/10.1006/pmpp.2003.0449
- Musetti, R., Sanità Di Toppi, L., Martini, M., Ferrini, F., Loschi, A., Favali, M.A. and Osler, R. (2005) Hydrogen Peroxide Localization and Antioxidant Status in the recovery of Apricot Plants from European Stone Fruit Yellows. European Journal of Plant Pathology, 112, 53-61. http://dx.doi.org/10.1007/s10658-004-8233-z
- Jagoueix-Eveillard, S., Tarendau, F., Guolter, K., Danet, J.L., Bové, J.M. and Garnier, M. (2001) Catharanthus roseus Genes Regulated Differentially by Mollicute Infections. Molecular Plant-Microbe Interactions, 14, 225-233. http://dx.doi.org/10.1094/MPMI.2001.14.2.225
- Davis, R.E., Jomantiene, R., Zhao, Y. and Dally, E.L. (2003) Folate Biosynthesis Pseudogenes, ψfolP and ψfolK, and an O-Sialoglycoprotein Endopeptidase Gene Homolog in the Phytoplasma Genome. DNA and Cell Biology, 22, 697- 706. http://dx.doi.org/10.1089/104454903770946674
- Bertaccini, A., Contaldo, N., Calari, A., Paltrinieri, S., Windsor, H.M. and Windsor, D. (2010) Preliminary Results of Axenic Growth of Phytoplasmas from Micropropagated Infected Periwinkle Shoots. 18th Congress of the International Organization for Mycoplasmology (IOM), Chianciano Terme, 11-16 July 2010, 147,153.
- Contaldo, N., Bertaccini, A., Paltrinieri, S., Windsor, H.M. and Windsor, D. (2012) Axenic Culture of Plant Pathogenic Phytoplasmas. Phytopathologia Mediterranea, 51, 607-617.
- Contaldo, N., Bertaccini, A., Paltrinieri, S., Windsor, D. and Windsor, H. (2013) Cultivation of Several Phytoplasmas from a Micropropagated Plant Collection. Petria, 23, 13-18.
- Lee, I.-M., Davis, R.E., Chen, T.A., Chiykowski, L.N., Fletcher, J., Hiruki, C. and Schaff, D.A. (1992) A GenotypeBase System for Identification and Classification of Mycoplasmalike Organisms (MLOs) in the Aster Yellows MLO Strain Cluster. Phytopathology, 82, 977-986. http://dx.doi.org/10.1094/Phyto-82-977
- Lee, I.-M., Gundersen-Rindal, D.E., Davis, R.E., Bottner, K.D., Marcone, C. and Seemüller, E. (2004) ‘Candidatus Phytoplasma asteris’, a Novel Phytoplasma Taxon Associated with Aster Yellows and Related Diseases. International Journal of Systematic and Evolutionary Microbiology, 54, 1037-1048. http://dx.doi.org/10.1099/ijs.0.02843-0
- Jomantiene, R., Davis, R.E., Maas, J. and Dally, E.L. (1998) Classification of New Phytoplasmas Associated with Diseases of Strawberry in Florida, Based on Analysis of 16S rRNA and Ribosomal Protein Gene Operon Sequences. International Journal of Systematic and Evolutionary Microbiology, 48, 269-277. http://dx.doi.org/10.1099/00207713-48-1-269
- Lee, I.-M., Martini, M., Bottner, K.D., Dane, R.A., Black, M.C. and Troxclair, N. (2003) Ecological Implications from a Molecular Analysis of Phytoplasmas Involved in an Aster Yellows Epidemic in Various Crops in Texas. Phytopathology, 93, 1368-1377. http://dx.doi.org/10.1094/PHYTO.2003.93.11.1368
- Lee, M.E., Grau, C.R., Lukaesko, L.A. and Lee, I.M. (2002) Identification of Aster Yellows Phytoplasmas in Soybean in Wisconsin Based on RFLP Analysis of PCR-Amplified Products (16S rDNAs). Canadian Journal of Plant Pathology, 24, 125-130. http://dx.doi.org/10.1080/07060660309506986
- Šeruga, M., Škorić, D., Botti, S., Paltrinieri, S., Juretić, N. and Bertaccini, A. (2003) Molecular Characterization of a Phytoplasma from the Aster Yellows (16SrI) Group Naturally Infecting Populus nigra L. ‘Italica’ Trees in Croatia. Forest Pathology, 33, 113-125. http://dx.doi.org/10.1046/j.1439-0329.2003.00312.x
- Valiunas, D., Jomantiene, R. and Davis, R.E. (2005) A ‘Candidatus Phytoplasma asteris’-Related Phytoplasma Associated with Cherry Little Leaf Disease Represents a New Subgroup, 16SrI-Q. Phytopathology, 95, S106.
- Jomantiene, R., Maas, J.L., Takeda, F. and Davis, R.E. (2002) Molecular Identification and Classification of Strawberry Phylloid Fruit Phytoplasma in Group 16SrI, New Subgroup. Plant Disease, 86, 920. http://dx.doi.org/10.1094/PDIS.2002.86.8.920C
- Santos-Cervantes, M.E., Chávez-Medina, J.A., Acostapardini, J., Flores-Zamora, G.L., Méndez-Lozano, J. and Leyva- López, N.E. (2010) Genetic Diversity and Geographical Distribution of Phytoplasmas Associated with Potato Purple Top Disease in Mexico. Plant Disease, 94, 388-395. http://dx.doi.org/10.1094/PDIS-94-4-0388
- Arocha-Rosete, Y., Zunnoon-Khan, S., Krukovets, I., Crosby, W., Scott, J., Bertaccini, A. and Michelutti, R. (2011) Identification and Molecular Characterization of the Phytoplasma Associated with Peach Rosette-Like Disease at the Canadian Clonal Genebank Based on the 16S rRNA Gene Analysis. Canadian Journal of Plant Pathology, 33, 127- 134. http://dx.doi.org/10.1080/07060661.2011.558854
- Arocha, Y., Antesana, O., Montellano, E., Franco, P., Plata, G. and Jones, P. (2007) ‘Candidatus Phytoplasma lycopersici’, a Phytoplasma Associated with ‘hoja de perejil’ Disease in Bolivia. International Journal of Systematic and Evolutionary Microbiology, 57, 1704-1710. http://dx.doi.org/10.1099/ijs.0.64851-0
- Gundersen, D.E., Lee, I.-M., Rehner, S.A., Davis, R.E. and Kingsbury, D.T. (1994) Phylogeny of Mycoplasmalike Organisms (Phytoplasmas): A Basis for Their Classification. Journal of Bacteriology, 176, 5244-5254.
- Zreik, L., Carle, P., Bové, J.M. and Garnier, M. (1995) Characterization of the Mycoplasmalike Organism Associated with Witches’-Broom Disease of Lime and Proposition of a ‘Candidatus’ Taxon for the Organism, ‘Candidatus Phytoplasma aurantifolia’. International Journal of Systematic and Evolutionary Microbiology, 45, 449-453. http://dx.doi.org/10.1099/00207713-45-3-449
- Seemüller, E., Marcone, C., Lauer, U., Ragozzino, A. and Göschl, M. (1998) Current Status of Molecular Classification of the Phytoplasmas. Journal of Plant Pathology, 80, 3-26.
- White, D.T., Blackall, L.L., Scott, P.T. and Walsh, K.B. (1998) Phylogenetic Positions of Phytoplasmas Associated with Dieback, Yellow Crinkle and Mosaic Diseases of Papaya, and Their Proposed Inclusion in ‘Candidatus Phytoplasma australiense’ and a New Taxon, ‘Candidatus Phytoplasma australasia’. International Journal of Systematic and Evolutionary Microbiology, 48, 941-951. http://dx.doi.org/10.1099/00207713-48-3-941
- Davis, R.E., Zhao, Y., Dally, E.L., Lee, I.M., Jomantiene, R. and Douglas, S.M. (2013) ‘Candidatus Phytoplasma pruni’, a Novel Taxon Associated with X-Disease of Stone Fruits, Prunus spp.: Multilocus Characterization Based on 16S rRNA, secY, and Ribosomal Protein Genes. International Journal of Systematic and Evolutionary Microbiology, 63, 766-776. http://dx.doi.org/10.1099/ijs.0.041202-0
- Davis, R.E., Jomantiene, R., Dally, E.L. and Wolf, T.K. (1998) Phytoplasmas Associated with Grapevine Yellows in Virginia Belong to Group 16SrI, Subgroup A (Tomato Big Bud Phytoplasma Subgroup), and Group 16SrIII, New Subgroup I. Vitis, 37, 131-137.
- Montano, H.G., Davis, R.E., Dally, E.L., Pimentel, J.P. and Brioso, P.S.T. (2000) Identification and Phylogenetic Analysis of a New Phytoplasma from Diseased Chayote in Brazil. Plant Disease, 84, 429-436. http://dx.doi.org/10.1094/PDIS.2000.84.4.429
- Alvarez, E., Mejía, J.F., Llano, G.A., Loke, J.B., Calari, A., Duduk, B. and Bertaccini, A. (2009) Detection and Molecular Characterization of a Phytoplasma Associated with Frogskin Disease in Cassava. Plant Disease, 93, 1139-1145. http://dx.doi.org/10.1094/PDIS-93-11-1139
- Davis, R.E., Dally, E.L. and Converse, R.H. (2001) Molecular Identification of a Phytoplasma Associated with Witches’-Broom Disease of Black Raspberry in Oregon and Its Classification in Group 16SrIII, New Subgroup Q. Plant Disease, 85, 1121. http://dx.doi.org/10.1094/PDIS.2001.85.10.1121A
- Valiunas, D., Jomantiene, R., Ivanauskas, A., Abraitis, R., Staniene, G., Zhao, Y. and Davis, R.E. (2009) First Report of a New Phytoplasma Subgroup, 16SrIII-T, Associated with Decline Disease Affecting Sweet and Sour Cherry Trees in Lithuania. Plant Disease, 93, 550. http://dx.doi.org/10.1094/PDIS-93-5-0550B
- Davis, R.E., Zhao, Y., Dally, E.L., Jomantiene, R., Lee, I.M., Wei, W. and Kitajima, E.W. (2012) ‘Candidatus Phytoplasma sudamericanum’, a Novel Taxon, and Strain PassWB-Br4, a New Subgroup 16SrIII-V Phytoplasma, from Diseased Passion Fruit (Passiflora edulis f. flavicarpa Deg.) International Journal of Systematic and Evolutionary Microbiology, 62, 984-989. http://dx.doi.org/10.1099/ijs.0.033423-0
- Harrison, N.A., Womack, M. and Carpio, M.L. (2002) Detection and Characterization of a Lethal Yellowing (16SrIV) Group Phytoplasma in Canary Island Date Palms Affected by Lethal Decline in Texas. Plant Disease, 86, 676-681. http://dx.doi.org/10.1094/PDIS.2002.86.6.676
- Harrison, N.A., Richardson, P.A., Kramer, J.B. and Tsai, J.H. (1994) Detection of the Mycoplasma-Like Organism Associated with Lethal Yellowing Disease of Palms in Florida by Polymerase Chain Reaction. Plant Pathology, 43, 998-1008. http://dx.doi.org/10.1111/j.1365-3059.1994.tb01649.x
- Lee, I.-M., Martini, M., Marcone, C. and Zhu, S.F. (2004) Classification of Phytoplasma Strains in the Elm Yellows Group (16SrV) and Proposal of ‘Candidatus Phytoplasma ulmi’ for the Phytoplasma Associated with Elm Yellows. International Journal of Systematic and Evolutionary Microbiology, 54, 337-347. http://dx.doi.org/10.1099/ijs.0.02697-0
- Jung, H.Y., Sawayanagi, T., Kakizawa, S., Nishigawa, H., Wei, W., Oshima, K., Miyata, S., Ugaki, M., Hibi, T. and Namba, S. (2003) ‘Candidatus Phytoplasma ziziphi’, a Novel Phytoplasma Taxon Associated with Jujube Witches’- Broom Disease. International Journal of Systematic and Evolutionary Microbiology, 53, 1037-1041. http://dx.doi.org/10.1099/ijs.0.02393-0
- Daire, X., Boudon-Padieu, E., Berville, A., Schneider, B. and Caudwell, A. (1992) Cloned DNA Probes for Detection of Grapevine Flavescence Doree Mycoplasma-Like Organism (MLO). Annals of Applied Biology, 121, 95-103. http://dx.doi.org/10.1111/j.1744-7348.1992.tb03990.x
- Malembic-Maher, S., Salar, P., Filippin, L., Carle, P., Angelini, E. and Foissac, X. (2011) Genetic Diversity of European Phytoplasmas of the 16SrV Taxonomic Group and Proposal of ‘Candidatus Phytoplasma rubi’. International Journal of Systematic and Evolutionary Microbiology, 61, 2129-2134. http://dx.doi.org/10.1099/ijs.0.025411-0
- Win, N.K.K., Lee, S.Y., Bertaccini, A., Namba, S. and Jung, H.Y. (2013) ‘Candidatus Phytoplasma balanitae’ Associated with Witches’ Broom Disease of Balanites triflora. International Journal of Systematic and Evolutionary Microbiology, 63, 636-640. http://dx.doi.org/10.1099/ijs.0.041566-0
- Hiruki, C. and Wang, K.R. (2004) Clover Proliferation Phytoplasma: ‘Candidatus Phytoplasma trifolii’. International Journal of Systematic and Evolutionary Microbiology, 54, 1349-1353. http://dx.doi.org/10.1099/ijs.0.02842-0
- Jacobs, K.A., Lee, I.-M., Griffiths, H.M., Miller Jr., F.D. and Bottner, K.D. (2003) A New Member of the Clover Proliferation Phytoplasma Group (16SrVI) Associated with Elm Yellows in Illinois. Plant Disease, 87, 241-246. http://dx.doi.org/10.1094/PDIS.2003.87.3.241
- Siddique, A., Agrawal, G.K., Alam, N. and Krishnareddy, M. (2001) Electron Microscopy and Molecular Characterization of Phytoplasmas Associated with Little Leaf Disease of Brinjal (Solanum melongena L.) and Periwinkle (Catharanthus roseus) in Bangladesh. Journal of Phytopathology, 149, 237-244. http://dx.doi.org/10.1046/j.1439-0434.2001.00590.x
- Faggioli, F., Pasquini, G., Lumia, V., Campobasso, G., Widmer, T.L. and Quimby, P.C. (2004) Molecular Identification of a New Member of the Clover Proliferation Phytoplasma Group (16SrVI) Associated with Yellow Starthistle Virescence in Italy. European Journal of Plant Pathology, 110, 353-360. http://dx.doi.org/10.1023/B:EJPP.0000021059.85956.f2
- Samad, A., Ajayakumar, P.V., Shasany, A.K., Gupta, M.K., Alam, M. and Rastogi, S. (2008) Occurrence of a Clover Proliferation (16SrVI) Group Phytoplasma Associated with Little Leaf Disease of Portulaca grandiflora in India. Plant Disease, 92, 832. http://dx.doi.org/10.1094/PDIS-92-5-0832A
- Griffiths, H.M., Sinclair, W.A., Smart, C.D. and Davis, R.E. (1999) The Phytoplasma Associated with Ash Yellows and Lilac Witches’ Broom: ‘Candidatus Phytoplasma fraxini’. International Journal of Systematic and Evolutionary Microbiology, 49, 1605-1614. http://dx.doi.org/10.1099/00207713-49-4-1605
- Barros, T.S.L., Davis, R.E., Resende, R.O. and Dally, E.L. (2002) Erigeron Witches’-Broom Phytoplasma in Brazil Represents New Subgroup VII-B in 16S rRNA Gene Group VII, the Ash Yellows Phytoplasma Group. Plant Disease, 86, 1142-1148. http://dx.doi.org/10.1094/PDIS.2002.86.10.1142
- Conci, L., Meneguzzi, N., Galdeano, E., Torres, L., Nome, C. and Nome, S. (2005) Detection and Molecular Characterisation of an Alfalfa Phytoplasma in Argentina that Represents a New Subgroup in the 16S rDNA Ash Yellows Group (‘Candidatus Phytoplasma fraxini’). European Journal of Plant Pathology, 113, 255-265. http://dx.doi.org/10.1007/s10658-005-0298-9
- Ho, K., Tsai, C. and Chung, T. (2001) Organization of Ribosomal RNA Genes from a Loofah Witches’ Broom Phytoplasma. DNA and Cell Biology, 20, 115-122. http://dx.doi.org/10.1089/104454901750070328
- Verdin, E., Salar, P., Danet, J.L., Choueiri, E., Jreijiri, F., El Zammar, S., Gèlie, B., Bové, J. and Garnier, M. (2003) ‘Candidatus Phytoplasma phoeniceum’, a New Phytoplasma Associated with an Emerging Lethal Disease of Almond Trees in Lebanon and Iran. International Journal of Systematic and Evolutionary Microbiology, 53, 833-838. http://dx.doi.org/10.1099/ijs.0.02453-0
- Duduk, B., Mejia, J.F., Calari, A. and Bertaccini, A. (2008) Identification of 16SrIX Group Phytoplasmas Infecting Colombian Periwinkles and Molecular Characterization on Several Genes. 17th Congress of the International Organization for Mycoplasmology (IOM), Tienjin, 6-11 July 2008, 112,83.
- Davis, R.E., Dally, E., Zhao, Y., Lee, I.M., Jomantiene, R., Detweiler, A.J. and Putnam, M.L. (2010) First Report of a New Subgroup 16SrIX-E (‘Candidatus Phytoplasma phoenicium’-Related) Phytoplasma Associated with Juniper Witches Broom Disease in Oregon, USA. Plant Pathology, 59, 1161. http://dx.doi.org/10.1111/j.1365-3059.2010.02294.x
- Marcone, C., Gibb, K.G., Streten, C. and Schneider, B. (2003) ‘Candidatus Phytoplasma spartii’, ‘Candidatus Phytoplasma rhamni’ and ‘Candidatus Phytoplasma allocasuarinae’, Respectively Associated with Spartium Witches’- Broom, Buckthorn Witches’ Broom and Allocasuarina Yellows Diseases. International Journal of Systematic and Evolutionary Microbiology, 54, 1025-1029.
- Seemüller, E., Schneider, B., Maurer, R., Ahrens, U., Daire, X., Kison, H., Lorenz, K., Firrao, G., Avinent, L., Sears, B.B. and Stackebrandt, E. (1994) Phylogenetic Classification of Phytopathogenic Mollicutes by Sequence Analysis of 16S Ribosomal DNA. International Journal of Systematic and Evolutionary Microbiology, 44, 440-446. http://dx.doi.org/10.1099/00207713-44-3-440
- Jung, H.Y., Sawayanagi, T., Wongkaew, P., Kakizawa, S., Nishigawa, H., Wei, W., Oshima, K., Miyata, S., Ugaki, M., Hibi, T. and Namba, S. (2003) ‘Candidatus Phytoplasma oryzae’, a Novel Phytoplasma Taxon Associated with Rice Yellow Dwarf Disease. International Journal of Systematic and Evolutionary Microbiology, 53, 1925-1929. http://dx.doi.org/10.1099/ijs.0.02531-0
- Quaglino, F., Zhao, Y., Casati, P., Bulgari, D., Bianco, P.A., Wei, W. and Davis, R.E. (2013) ‘Candidatus Phytoplasma solani’, a Novel Taxon Associated with Stolburand Bois Noir-Related Diseases of Plants. International Journal of Systematic and Evolutionary Microbiology, 63, 2879-2894. http://dx.doi.org/10.1099/ijs.0.044750-0
- Davis, R.E., Dally, E.L., Gundersen, D.E., Lee, I.M. and Habili, N. (1997) ‘Candidatus Phytoplasma australiense’, a New Phytoplasma Taxon Associated with Australian Grapevine Yellows. International Journal of Systematic and Evolutionary Microbiology, 47, 262-269. http://dx.doi.org/10.1099/00207713-47-2-262
- Padovan, A.C., Gibb, K.S., Bertaccini, A., Vibio, M., Bonfiglioli, R.E., Magarey, P.A. and Sears, B.B. (1995) Molecular Detection of the Australian Grapevine Yellows Phytoplasma and Comparison with Grapevine Yellows Phytoplasma from Italy. Australian Journal of Grape and Wine Research, 1, 25-31. http://dx.doi.org/10.1111/j.1755-0238.1995.tb00074.x
- Sawayanagi, T., Horikoshi, N., Kanehira, T., Shinohara, M., Bertaccini, A., Cousin, M.T., Hiruki, C. and Namba, S. (1999) ‘Candidatus Phytoplasma japonicum’, a New Phytoplasma Taxon Associated with Japanese Hydrangea Phyllody. International Journal of Systematic and Evolutionary Microbiology, 49, 1275-1285. http://dx.doi.org/10.1099/00207713-49-3-1275
- Valiunas, D., Staniulis, J. and Davis, R.E. (2006) ‘Candidatus Phytoplasma fragariae’, a Novel Phytoplasma Taxon Discovered in Yellows Diseased Strawberry, Fragaria × ananassa. International Journal of Systematic and Evolutionary Microbiology, 56, 277-281. http://dx.doi.org/10.1099/ijs.0.63935-0
- Quaglino, F., Zhao, Y., Bianco, P.A., Wei, W., Casati, P., Durante, G. and Davis, R.E. (2009) New 16Sr Subgroups and Distinct Single Nucleotide Polymorphism Lineages among Grapevine Bois Noir Phytoplasma Populations. Annals of Applied Biology, 154, 279-289. http://dx.doi.org/10.1111/j.1744-7348.2008.00294.x
- Martini, M., Marcone, C., Mitrović, J., Maixner, M., Delić, D., Myrta, A., Ermacora, P., Bertaccini, A. and Duduk, B. (2012) ‘Candidatus Phytoplasma convolvuli’, a New Phytoplasma Taxon Associated with Bindweed Yellows in Four European Countries. International Journal of Systematic and Evolutionary Microbiology, 62, 2910-2915. http://dx.doi.org/10.1099/ijs.0.038612-0
- Marcone, C., Schneider, B. and Seemüller, E. (2003) ‘Candidatus Phytoplasma cynodontis’, the Phytoplasma Associated with Bermuda Grass White Leaf Disease. International Journal of Systematic and Evolutionary Microbiology, 54, 1077-1082. http://dx.doi.org/10.1099/ijs.0.02837-0
- Salehi, M., Izadpanah, K., Siampour, M. and Taghizadeh, M. (2009) Molecular Characterization and Transmission of Bermuda Grass White Leaf Phytoplasma in Iran. Journal of Plant Pathology, 91, 655-661.
- Montano, H.G., Davis, R.E., Dally, E.L., Hogenhout, S., Pimentel, J.P. and Brioso, P.S. (2001) ‘Candidatus Phytoplasma brasiliense’, a New Phytoplasma Taxon Associated with Hibiscus Witches’ Broom Disease. International Journal of Systematic and Evolutionary Microbiology, 51, 1109-1118. http://dx.doi.org/10.1099/00207713-51-3-1109
- Villalobos, W., MartiniI, M., GaritaI, L., MuñozI, M., Osler, R. and Moreira, L. (2011) Guazuma ulmifolia (Sterculiaceae), a New Natural Host of 16SrXV Phytoplasma in Costa Rica. Tropical Plant Pathology, 36. http://dx.doi.org/10.1590/S1982-56762011000200007
- Arocha, Y., Lopez, M., Pinol, B., Fernandez, M., Picornell, B., Almeida, R., Palenzuela, I., Wilson, M.R. and Jones, P. (2005) ‘Candidatus Phytoplasma graminis’ and ‘Candidatus Phytoplasma caricae’, Two Novel Phytoplasmas Associated with Diseases of Sugarcane, Weeds and Papaya in Cuba. International Journal of Systematic and Evolutionary Microbiology, 55, 2451-2463. http://dx.doi.org/10.1099/ijs.0.63797-0
- Lee, I.-M., Bottner, K.D., Secor, G. and Rivera Varas, V. (2006) ‘Candidatus Phytoplasma americanum’ a Phytoplasma Associated with a Potato Purple Top Wilt Disease Complex. International Journal of Systematic and Evolutionary Microbiology, 56, 1593-1597. http://dx.doi.org/10.1099/ijs.0.64251-0
- Jung, H.Y., Sawayanagi, T., Kakizawa, S., Nishigawa, H., Miyata, S., Oshima, K., Ugaki, M., Joon-Tak, L. and Namba, S. (2002) ‘Candidatus Phytoplasma castaneae’, a Novel Phytoplasma Taxon Associated with Chestnut Witches’ Broom Disease. International Journal of Systematic and Evolutionary Microbiology, 52, 1543-1549.
- Schneider, B., Torres, E., Martìn, M.P., Schroder, M., Behnke, H.D. and Seemüller, E. (2005) ‘Candidatus Phytoplasma pini’, a Novel Taxon from Pinus silvestris and Pinus halepensis. International Journal of Systematic and Evolutionary Microbiology, 55, 303-307. http://dx.doi.org/10.1099/ijs.0.63285-0
- Harrison, N., Davis, R.E., Oropeza, C., Helmick, E., Narvaez, M., Eden-Green, S., Dollet, M. and Dickinson, M. (2014) ‘Candidatus Phytoplasma palmicola’, a Novel Taxon Associated with a Lethal Yellowing-Type Disease (LYD) of Coconut (Cocos nucifera L.) in Mozambique. International Journal of Systematic and Evolutionary Microbiology. http://dx.doi.org/10.1099/ijs.0.060053-0
- Wei, W., Davis, R.E., Lee, I.-M. and Zhao, Y. (2007) Computer-Simulated RFLP Analysis of 16S rRNA Genes: Identification of Ten New Phytoplasma Groups. International Journal of Systematic and Evolutionary Microbiology, 57, 1855-1867. http://dx.doi.org/10.1099/ijs.0.65000-0
- Al-Saady, N.A., Khan, A.J., Calari, A., Al-Subhi, A.M. and Bertaccini, A. (2008) ‘Candidatus Phytoplasma omanense’, a Phytoplasma Associated with Witches’ Broom of Cassia italica (Mill.) Lam. in Oman. International Journal of Systematic and Evolutionary Microbiology, 58, 461-466. http://dx.doi.org/10.1099/ijs.0.65425-0
- Zhao, Y., Sun, Q., Wei, W., Davis, R.E., Wu, W. and Liu, Q. (2009) ‘Candidatus Phytoplasma tamaricis’, a Novel Taxon Discovered in Witches’ Broom-Diseased Salt Cedar (Tamarix chinensis Lour.). International Journal of Systematic and Evolutionary Microbiology, 59, 2496-2504. http://dx.doi.org/10.1099/ijs.0.010413-0
NOTES

*Corresponding author.


