Advances in Lung Cancer
Vol.2 No.1(2013), Article ID:29206,5 pages DOI:10.4236/alc.2013.21004
The bisulfite genomic sequencing protocol
![]()
1Department of Pharmacology and Therapeutics, McGill University, Montreal, Canada; *Corresponding Author: jane.pappas@mcgill.ca
2Department of Anatomy and Neuroscience, University College Cork, Cork, Ireland
3Department of Medicine, Université de Montréal, Montreal, Canada
Received 23 December 2012; revised 25 January 2013; accepted 6 February 2013
Keywords: Bisulfite Genomic Sequencing; DNA Methylation; Epigenetics; Cancer
ABSTRACT
The bisulfite genomic sequencing (BGS) protocol has gained worldwide popularity as the method of choice for analyzing DNA methylation. It is this popular because it is a powerful protocol and it may be coupled with many other applications. However, users often run into a slew of problems, including incomplete conversion, overly degraded DNA, sub-optimal PCR amplifications, false positives, uninformative results, or altogether failed experiments. We pinpoint the reasons why these problems arise and carefully explain the critical steps toward accomplishing a successful experiment step-by-step. This protocol has worked successfully (>99.9% conversion) on as little as 100 ng of DNA derived from nearly 10-year-old DNA samples extracted from whole blood stored at −80˚C and resulted in enough converted DNA for more than 50 PCR reactions. The aim of this article is to make learning and usage of BGS easier, more efficient and standardized for all users.
1. INTRODUCTION
The bisulfite genomic sequencing protocol (BGS) has gained worldwide popularity as the method of choice to analyze DNA methylation. DNA methylation was the first epigenetic mark to be discovered in mammalian cells [1], and while many other epigenetic marks are known and even more are currently being discovered [2], the role that DNA methylation plays in the regulation of gene expression is now widely accepted.
Bisulfite was first used to convert 5-methylcytosines to uracils in 1970 [3], but the BGS protocol used today to determine the methylation status of CpG dinucleotides in the genome was first published in 1992 [4]. Sodium bisulfite and metabisulfite ions are used to convert unmethylated cytosines to uracils in three steps (Figure 1).
Conversion can only occur in single-stranded DNA (ssDNA). Importantly, bisulfite does not affect 5-methylcytosines: the methyl group in the 5 position of the cytosine base causes 5-methylcytosines to be non-reactive to bisulfite ions. The end result following PCR amplification of bisulfite-converted DNA is that 5-methylcytosines in the original sample are read as C, whereas cytosines are read as T (Figure 2).
Optimal bisulfite conversion conditions are those that ensure that target molecules remain single-stranded throughout the chemical transformation steps. Paying particular attention to solvent molarities, pH, temperature and timing of denaturing steps is critical to the successful experiment. Unfortunately, false positives, uninformative results, or altogether failed experiments are often encountered, and this is due to three main reasons: 1) The

Figure 1. The chemistry of the bisulfite conversion reaction. Bisulfite conversion consists of three sequential chemical reactions: 1. cytosine → cytosine-6- sulfonate; 2. cytosine-6-sulfonate → uracil-6-sulfonate; and 3. uracil-6-sul-fonate → uracil. The methyl group in the 5 position of the cytosine base causes 5-methylcytosines to be non-reactive to bisulfite ions and these therefore remain unchanged.

Figure 2. Schematic representation of an arbitrary sequence following bisulfite conversion. Note that only one CG-cytosine is methylated (uppercase) whereas the other CG-cytosine and the two non-CG-cytosines are not. Following PCR amplification, 5-methylcytosines in the original DNA sample will be read as C, whereas cytosines will be read as T.
deamination of all unmethylated non-CpG-cytosine residues in the target sequence under investigation is not achieved; 2) Conditions for successful conversion are in themselves highly degradative to DNA; 3) Frequently, the target sequence has inherent characteristics, such as unusually CG-rich regions, that make it difficult to sequence. This article carefully describes the key steps of the BGS protocol and has worked successfully on as little as 100 ng of DNA (starting material) resulting in enough converted DNA for more than 50 PCR reactions. The aim of this article is to make learning and usage of BGS easier, more efficient and standardized for all users.
2. PROTOCOL
2.1. Prepare the DNA for Alkaline Denaturation and Bisulfite Conversion
STEP 1. Extract the DNA from the sample using a standard technique appropriate for the sample type, such as the phenol-chloroform technique [5] followed by Proteinase K treatment to ensure that all nuclear proteins have been removed. Protocols and kits using chaotropic salts (such as the All Prep DNA and RNA kit from QIAGEN) do not necessarily require Proteinase K treatment. See Table 1 for suggested reagents and equipment required.
STEP 2. Digest between 0.1 to 5 µg DNA with a suitable restriction enzyme such as PstI, BamHI or other frequent cutter according to the supplier’s protocol in order to arbitrarily fragment the DNA into smaller, more manageable sizes. Use care to ensure that the restriction enzyme chosen does not have a restriction site in the sequence of interest. Sonication, a somewhat less reliable method as it may vary from experiment to experiment, may also be used. Test efficiency and uniformity of fragmentation across samples by loading a fraction of the fragmented DNA (200 to 500 ng) onto an agarose gel, if possible.
Note: Fragmentation is an important step that reduces the possibility of double-strandedness, which prevents conversion. Using more DNA (e.g. up to 5 µg) is better as the highly oxidative conditions of the bisulfite conver
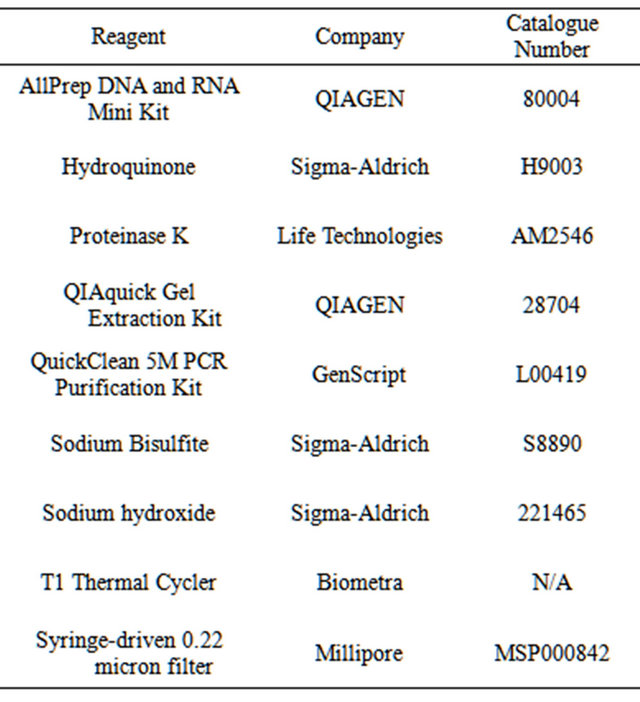
Table 1. Suggested reagents and equipment required.
sion reaction have high rates of DNA degradation. If using between 0.1 µg and 1.0 µg, make sure to increase all following precipitation times to 8h or more (overnight) and spin times to 1h or more at 13,000 xg at 4˚C.
STEP 3. Clean-up and re-concentrate the DNA using a DNA purification column, such as provided in the QuickClean 5M PCR Purification Kit (GenScript) and elute in 60 µl 2 mM Tris, pH 8.5. Use 1-2 µl to evaluate the concentration of starting material by standard spectrophotometry or nanodrop before proceeding to bisulfite conversion. Store DNA at −20˚C to 4˚C for further use.
Note: While distilled water may be used, we have found that using a common DNA solvent such as 2 mM Tris, pH 8.5 gives more consistent results. This may be explained by the fact that de-ionized or distilled water can vary in pH.
STEP 4. Before beginning the protocol, ensure that all required reagents are on hand, including 3N NaOH, freshly prepared 3.6 M sodium bisulfite, pH 5.0, and 10 mM hydroquinone.
1) To make 3N NaOH, weigh out 6.67 g NaOH pellets and dissolve in 50 ml water.
Note: NaOH solutions are stable at room temperature indefinitely.
2) To make 3.6 M sodium bisulfite, pH 5.0, weigh out 7.49 g sodium bisulfite in 15 ml water and mix for 10 minutes on a nutator protected from light. Then adjust the acidity to pH 5.0 using about 12 to 13 drops or so of 10N NaOH. Use a long glass pipette to slowly add the 10N NaOH. Remove potential contaminants by passing the sodium bisulfite solution through a 0.22 micron syringe-driven filter.
Note: Sodium bisulfite solutions should always be freshly prepared.
3) To make 10 mM hydroquinone, weigh out 22 mg hydroquinone in 10 ml water and invert until dissolved. Remove possible contaminants by passing the hydroquinone solution through a 0.22 micron pore filter.
Note: Hydroquinone solutions may be preserved in 1 ml aliquots in foil-wrapped microcentrifuge tubes at −20˚C for up to 2 months.
STEP 5. Add 1 ml hydroquinone to 15 ml sodium bisulfite and adjust to 20 ml total with distilled water. This solution may be stored at 4˚C for up to 2 days in a foilwrapped 50 ml polypropylene tube.
2.2. Denature the DNA Using NaOH and Heat
STEP 6. Proceed by denaturing 54 µl of the digested DNA with 6 µl of 3N NaOH at 37˚C for 15 min (for a final concentration of 0.3N NaOH). Place sample on ice immediately following denaturation if the next step, STEP 3, is not performed immediately.
Note: Ensure that the volume of 3N NaOH added is exactly 1/10th the final volume.
2.3. Convert the DNA Using Bisulfite
STEP 7. Sulfonate unmethylated cytosines in ssDNA.
1) Add 430 µl of the freshly prepared sodium bisulfitehydroquinone solution to 0.1 to 5 µg of the digested DNA from STEP 3 and carefully mix 5 to 10 times by inversion for minimal aeration.
2) Transfer the mix to 3 microcentrifuge tubes, 200 µl capacities.
Note: If you are planning on sequencing a sample several times to assess the various copies or alleles present in a same sample for example, it will be necessary to use several tubes per sample. Alternatively, several PCR reactions may be set up from as many aliquots of bisulfite-treated DNA. These precautions ensure that sequencings are not derived from a same template molecule in the PCR amplification step.
3) Incubate in a thermal cycler using the following program: (95˚C for 4 minutes and then at 55˚C for 4 hours) × 2 cycles; (95˚C for 4 minutes and then 55˚C for 2 hours) × 1 cycle. Store temporarily at 4˚C. Using a thermal cycler rather than a water-bath saves time, lowers the risk of contamination, and ensures that reactions will be kept in the dark without any hassles. Incubations at 95˚C ensure single-strandedness.
Note: It is not necessary to cover the solutions with mineral oil as there is very little aeration in 200 µl capacity PCR tubes. However, overlaying solutions with a drop of mineral oil is highly recommended when conversions are performed in 1.5 ml capacity microcentrifuge tubes when using other types of incubators such as waterbaths.
4) Desalt the DNA using DNA purification columns such as those from Gen Script used before in STEP 3.
5) Elute the DNA using 54 µl 2 mM Tris pH 8.0 to 8.5.
STEP 8. Deaminate cytosine-6-sulfonates to uracil-6- sulfonates.
Add 6 µl 3N NaOH to 54 µl bisulfite-treated DNA from the previous step and incubate for 15 min at 37˚C. Immediately place tube on ice.
Note: Ensure that the volume of 3N NaOH is exactly 1/10th the final volume.
STEP 9. Desulfonate uracil-6-sulfonates to uracils.
1) Precipitate and desulfonate DNA simultaneously by adding 2 µl glycogen (1 µg/µl) as a carrier, 26 µl of 10M ammonium acetate, pH 7.8 and 500 µl of 95% ice cold ethanol in this order to each reaction. Place tube in an ice-water bath for 10 minutes or at −20˚C for 1 hour (or at −20˚C overnight if using less than 1 µg starting material DNA).
2) Spin the DNA in a benchtop centrifuge at 13,000 xg for 30 minutes at 4˚C (or at 13,000 xg for >1 h at 4˚C if using less than 1 µg starting material DNA).
3) Wash the precipitate with 200 µl 70% ice-cold ethanol and spin at 13,000 xg for 5 minutes at 4˚C and air-dry. Ensure that all traces of ethanol have completely evaporated.
4) Resuspend the bisulfite-converted DNA in enough TE, pH 8.0 to make a final concentration of 25ng/µl, ideal for subsequent PCR amplification steps.
Note: Calculate the volume of TE, pH 8.0 required based on the starting amount of pre-digested DNA used for bisulfite conversion (see STEP 3). Bisulfite-converted DNA concentration cannot be accurately assessed. If using less than 1 µg genomic DNA, use 50 - 100 µl elution buffer to ensure optimal elution conditions but upwardly adjust the number of PCR cycles accordingly.
5) Make multiple aliquots to prevent too many freezethaw cycles. Store aliquots at −20˚C for short term use or at −80˚C for up to 6 months.
2.4. PCR Amplify the Bisulfite-Converted DNA
STEP 10. Design primers to the bisulfite-converted DNA sequence such that each primer is 25 to 30 bases long, has a melting temperature (Tm) of approximately 60˚C, and importantly, does not hybridize to any CpGcytosines, if possible. Aim for an amplicon length of no more than 500 bp with as few bases in the primers hybridizing to non-CpG-cytosines in the target sequence as possible. Softwares such as MethPrimer [6] work well for most sequences.
Note: Re-amplifying a smaller region within the first amplicon using nested primers is a trick often used to overcome poor amplification due to the highly oxidative conditions of BGS. However, it has recently been suggested that detection of non-CpG methylation may be impaired by two rounds of PCR [7].
STEP 11. Optimize PCR conditions, including annealing temperature (Ta) and extension time using a gradient thermal cycler. Recommended PCR conditions: 1 cycle at 95˚C for 5 min; 35 cycles at: 94˚C for 1 minute; the target annealing temperature (Ta) for 2.5 minutes, 72˚C for 1 minute; and finally 1 cycle at 72˚C for 5 minutes.
Note: Ensure that temperatures 1 to 5 degrees below and 1 to 5 degrees above the Ta are tested in a gradient thermal cycler in order to determine the temperature at which a single, specific PCR product is amplified. Using more cycles may be necessary if starting with less than 1 µg of genomic DNA.
STEP 12. Load a fraction of the PCR amplification onto an agarose gel of adequate concentration (i.e. use a 1.5% agarose gel for products between 200 and 500 bp), allow electrophoretic migration and view using standard viewing methods. If the amplified product is of the correct molecular weight and free of non-specific by-products, proceed to traditional cloning and sequencing, pyrosequencing, or other paired application.
Determine the total number of non-CpG-Cs in the target sequence and set a successful conversion threshold. For example, a threshold of 98% conversion in a sequence containing 50 non-CpG-cytosines must have 49 of its non-CpG-C converted to T in the PCR product for the data from that sample to be included in analyses.
3. DISCUSSION
The three main problems that users of the BGS protocol run into are incomplete conversion, degradation of modified DNA and sub-optimal PCR amplification.
3.1. Incomplete Conversion
Incomplete conversion arises mainly due to inadequate solution alkalinization at steps 2.1 and 3.2.1, thereby preventing conversion due to the presence of double stranded regions in the target DNA molecules at step 2.1 or due to sub-optimal desulphonation at step 3.2.1. This challenge can most often be resolved by measuring the pH directly, or by adding one mock sample in order to measure it indirectly (and not waste precious sample material). A less frequent cause for incomplete conversion is insufficient incubation at 55˚C or lack of a melting step at 95˚C at step 3.1.3. We find it necessary to allow the reaction to proceed for a minimum of 6 - 8 hours and to melt the DNA prior to the last 2 - 3 hours of the reaction.
3.2. Degradation of Modified DNA
Degradation of the modified end-product results in low yield and cannot be overcome unless more starting material is used. Bisulfite conversion reaction conditions are very harsh, causing depurination and degradation of 75% - 90% of the DNA being modified. We have found it necessary to start with at least 2 to 4 µg of genomic DNA when possible, or to spike the starting material with 5 µg of salmon sperm DNA or tRNA when not, in order to buffer the depurinating conditions mentioned above. These conditions ensure that there is enough bisulfite-converted DNA for multiple PCR reactions and for long term use.
3.3. Sub-Optimal PCR Amplification
Sub-optimal PCR results, including lack of signal, poor signal or poor signal-to-noise ratio, are often encountered following a single bisulfite DNA PCR experiment. This is most often due to insufficient yield (i.e. low modified target concentration) and/or many other homologous modified products exist in solution along with it. A practical way to get around this problem is to use a fraction (e.g. 1/50th) of the first PCR product reaction volume to amplify a smaller region within the first amplicon using nested or semi-nested primers, however, this may not always lead to the desired outcome. In fact, it has been shown that detection of non-CpG methylation may even be impaired by two rounds of PCR [7]. Lack of signal even after a second round of PCR suggests complete degradation of starting material while a smear of non-specific products suggests sub-optimal primer design. Designing longer primers (up to 35 - 40 bp) or designing different primers (i.e. to another stretch of DNA within the target region) can often overcome this challenge.
4. CONCLUSIONS
Following these steps using 2 µg of starting DNA material (before conversion) will lead to sufficient converted DNA for at least 40 successful PCR reactions and related experiments such as pyrosequencing (Figure 3), assuming that 2 µl of 25 ng/µl bisulfite-converted DNA is used per PCR reaction.
The BGS protocol is a powerful protocol that has gained worldwide popularity for analyzing DNA methyllation. It can be used alone, in clonal bisulfite sequencing [4] and direct bisulfite sequencing and GENESCAN [8], or it can be matched with many other techniques, including methylation-specific PCR [9], pyrosequencing [10], HRM [11], MS-snuPE [12], EpiTYPER MassARRAY [13] and Illumina Methylation based arrays [14]. Particular care must therefore be used to ensure efficient conversion.

Figure 3. Example of a successful bisulfite conversion followed by pyrosequencing. Note that the control C testing for presence of unconverted cytidine (yellow bar) has no peak as expected.
5. ACKNOWLEDGEMENTS
JJP would like to thank the McGill University Faculty of Medicine for their support.
REFERENCES
- Wyatt, G.R. (1951) Recognition and estimation of 5-methylcytosine in nucleic acids. Biochemical Journal, 48, 581-584.
- Tan, M., Luo, H., Lee, S., Jin, F., Yang, J.S., Montellier, E., Buchou, T., Cheng, Z., Rousseaux, S., Rajagopal, N., Lu, Z., Ye, Z., Zhu, Q., Wysocka, J., Ye, Y., Khochbin, S., Ren, B. and Zhao, Y. (2011) Identification of 67 histone marks and histone lysine croton-ylation as a new type of histone modification. Cell, 16, 1016-1028. doi:10.1016/j.cell.2011.08.008
- Hayatsu, H., Wataya, Y., Kai, K. and Lida, S. (1970) Reaction of sodium bisulfite with uracil, cytosine, and their derivatives. Biochemistry, 9, 2858-2865. doi:10.1021/bi00816a016
- Frommer, M., McDonald, L.E., Millar, D.S., Collis, C.M., Watt, F., Grigg, G.W., Molloy, P.L. and Paul, C.L. (1992) A genomic sequencing protocol that yields a positive display of 5-methylcytosine residues in individual DNA strands. Proceedings of the National Academy of Sciences of the USA, 89, 1827-1831. doi:10.1073/pnas.89.5.1827
- Sambrook, J., Fritsch, E.F. and Maniatis, T. (1989) in Molecular Cloning: A Laboratory Manual. Cold Spring Harbor Laboratory Press, New York, 1, 191-195.
- Li, L.C. and Dahiya, R. (2002) MethPrimer: Designing primers for methylation PCRs. Bioinformatics, 18, 1427- 1431. doi:10.1093/bioinformatics/18.11.1427
- Yan, J., Zierath, J.R. and Barrès, R. (2011) Evidence for non-CpG methylation in mammals. Experimental Cell Research, 317, 2555-2561. doi:10.1016/j.yexcr.2011.08.019
- Paul, C.L. and Clark, S.J. (1996) Cytosine Methylation: Quantitation by Automated Genomic Sequencing and GENESCAN Analysis. BioTechniques, 21, 126-133.
- Herman, J.G., Graff, J.R., Myohanen, S., Nelkin, B.D. and Baylin, S.B. (1996) Methylation-specific PCR: A novel PCR assay for methylation status of CpG islands. Proceedings of the National Academy of Sciences of the USA, 93, 9821-9826. doi:10.1073/pnas.93.18.9821
- Uhlmann, K., Brinckmann, A., Toliat, M.R., Ritter, H. and Nurnberg, P. (2002) Evaluation of a potential epigenetic biomarker by quantitative methyl-single nucleotide polymorphism analysis Electrophoresis, 23, 4072-4079. doi:10.1002/elps.200290023
- Wojdacz, T.K. and Dobrovic, A. (2007) Methylation-sensitive high resolution melting (MSHRM): A new approach for sensitive and high-throughput assessment of methylation. Nucleic Acids Research, 35, e41. doi:10.1093/nar/gkm013
- Gonzalgo, M.L. and Jones, P.A. (1997) Rapid quantitation of methylation differences at specific sites using methylation-sensitive single nucleotide primer extension (Ms-SNuPE), Nucleic Acids Research, 25, 2529-2531. doi:10.1093/nar/25.12.2529
- Ehrich, M., Nelson, M.R., Stanssens, P., Zabeau, M., Liloglou, T., Xinarianos, G., Cantor, C.R., Field, J.K. and van den Boom, D. (2005) Quantitative high-throughput analysis of DNA methylation patterns by base-specific cleavage and mass spectrometry. Proceedings of the National Academy of Sciences of the USA, 102, 15785- 15790. doi:10.1073/pnas.0507816102
- Adorjan, P., Distler, J., Lipscher, E., Model, F., Muller, J., Pelet, C., Braun, A., Florl, A.R., Gutig, D., Grabs, G., et al., (2002) Tumour class prediction and discovery by microarray-based DNA me-thylation analysis. Nucleic Acids Research, 30, e21. doi:10.1093/nar/30.5.e21

