Advances in Anthropology
Vol.05 No.04(2015), Article ID:60322,62 pages
10.4236/aa.2015.54019
Distribution of East Eurasian Y-Chromosome and Mitochondrial DNA Haplogroups across Eurasia: Insights into the Genetic Ancestry of Bulgarians
Sena Karachanak-Yankova1*, Desislava Nesheva1*, Angel S. Galabov2, Draga Toncheva1#
1Department of Medical Genetics, Medical University of Sofia, Sofia, Bulgaria
2The Stephan Angeloff Institute of Microbiology, Bulgarian Academy of Sciences, Sofia, Bulgaria
Email: #dragatoncheva@gmail.com
Copyright © 2015 by authors and Scientific Research Publishing Inc.
This work is licensed under the Creative Commons Attribution International License (CC BY).
http://creativecommons.org/licenses/by/4.0/



Received 18 August 2015; accepted 13 October 2015; published 16 October 2015
ABSTRACT
The modern Bulgarian mitochondrial DNA (mtDNA) and Y-chromosome gene pools predominantly consist of Western Eurasian haplogroups. In contrast, the Eastern Eurasian lineages are found at very low frequencies in Bulgarians, being represented only by mtDNA haplogroups C (0.2%), D (0.4%) and Z (0.1%) (Karachanak et al., 2012) and Y-chromosome haplogroups C, N and Q (each 0.5%) (Karachanak et al., 2013) . A similar pattern is observed in ancient mtDNA samples of proto-Bulgarian human remains, which belong exclusively to Western Eurasian mtDNA haplogroups (Nesheva et al., 2015) . In order to investigate Bulgarian ancestry from the perspective of Eastern Eurasian haplogroups, we have analyzed the distribution of Y-chromosome haplogroups C, N and Q and mtDNA haplogroups C, D and Z across Eurasia. The survey was performed using literature data for more than 15,000 individuals from different Eurasian (sub-) populations for each of these haplogroups. The collected data were used to construct Eurasian frequency maps of the considered haplogroups and to test the significance of their incidence between Bulgarians and Europeans, European neighboring populations of Bulgaria and populations, which according to some historical conceptions could have common ancestry with proto-Bulgarians, namely: Altaian, Caucasus, Siberian and Central Asian populations. The spatial distribution of mtDNA haplogroups C, D and Z and Y-chromosome haplogroups C, N and Q contrasts their high frequency among Altaic populations and their occasional appearance in Bulgarians. Furthermore, the comparison of the occurrence of these haplogroups shows no link between Bulgarians and Altaic and Caucasus populations. Based on the substantial genetic input of proto-Bulgarians to the modern Bulgarian gene pool, the present study confirms the nonexistence of a close Y-chromosomal or mtDNA link between proto-Bulgarians on the one hand and Altaic and Caucasus populations on the other.
Keywords:
Bulgarians, Y-Chromosome, mtDNA, Haplogroup

1. Introduction
The territory of Bulgaria has been involved in different migration routes ever since the modern human settlement of Europe in the Upper Paleolithic. Later, it has served as a refugium and fuelled different expansion routes of postglacial re-colonization. The area of present-day Bulgaria also played a significant role in the Neolithic spread (Calafell et al., 1996; Nitecki & Nitecki, 2013; Ray & Adams, 2001) . This prehistorical population background was overlaid by the inhabitance of the Thracians, and later by the invasions of Slavs and proto-Bulgarians, which were considered as populations playing important role in the modern Bulgarian ethnogenesis (Dobrev, 1994, 2005; Haefs, 2009) .
The most disputable issue of the Bulgarian past is the origin of proto-Bulgarians. Their ancestry has been widely debated, as there are even theories describing them as Altaic and Turko-Mongolic people (Zlatarski, 1970) . According to novel archeological, historical and linguistic evidence, the ancient (proto-) Bulgarians made a substantial contribution to the modern Bulgarian gene pool (Dimitrov, 2005; Rashev, 1993) .
The previously performed phylogenetic analysis of 13 ancient mtDNA samples retrieved from proto-Bulga- rian human remains and discovered in three necropolises in Bulgaria dating to VIII-X century AD identified mtDNA haplogroups, which were found in present-day European and Western Eurasian populations. These results pointed to a Western Eurasian matrilineal origin for proto-Bulgarians as well as to a genetic similarity between them and modern Bulgarians (Nesheva et al., 2015) .
The high-resolution phylogenetic analyses of Y-chromosome and mitochondrial DNA (mtDNA) diversity of modern Bulgarians in samples comprising 808 and 855 subjects, respectively, showed that their gene pool was predominantly composed by Western Eurasian haplogroups. On the other hand, Eastern Eurasian haplogroups are represented at negligible frequencies. Thus, mtDNA haplogroups C, D and Z account for 0.2%, 0.4% and 0.1% of the modern Bulgarian gene pool, respectively and Y-chromosome haplogroups C, N and Q are each at frequency of 0.5%. The interpopulational comparisons of Y-chromosome and mtDNA haplogroup frequencies locate modern Bulgarians among European populations in both cases (Karachanak et al., 2012, 2013) .
In order to further test the reliability of the theories of Bulgarian, and in particular of proto-Bulgarian origin, we have depicted the frequency gradients of Y-chromosome haplogroups C, N and Q and mtDNA haplogroups C, D and Z across Eurasia and we have tested the significance of their incidence between Bulgarians and Europeans, European neighboring populations of Bulgaria and populations, which according to some historical conceptions could have common ancestry with proto-Bulgarians, namely: Altaian, Caucasus, Siberian and Central Asian populations.
2. Materials and Methods
We have collected literature data for the frequencies of Y-chromosome haplogroups C, N and Q and mtDNA haplogroups C, D and Z in Eurasian populations. The gathered data for the Y-chromosome haplogroups comprise 19856 samples from 186 (sub-) populations for Hg C, 16418 from 173 (sub-) populations for Hg N and 15652 samples from 160 (sub-) populations for Hg Q (supplementary Tables S1(a)-(c), respectively). For the mtDNA haplogroups we have collected data for 35083 samples representing 226 (sub-) populations for Hg C, 38039 from 242 (sub-) populations for haplogroup D and 17716 samples from 160 (sub-) populations for Hg Z (supplementary Tables S2(a)-(c), respectively). These data were used to construct frequency maps by using the Surfer Golden software following the Kriging procedure.
The occurrences of the aforementioned haplogroups were compared between modern Bulgarians and: Europeans, European neighboring populations of Bulgaria, Altaian, Caucasus, Siberian and Central Asian populations. The comparison was performed through a chi-square test using XLSTAT.
3. Results
The frequency maps of Y-chromosome haplogroups C, N and Q are given in Figure 1.
Haplogroup C, one of the earliest Out-of-Africa founder types, has highest frequencies in the middle latitudes of Asia. It has overwhelming presence among: Oroqens (90.9%), Evens (74.2%), Evenks (68.4%) (Hammer et al., 2005) , Buryats (68.2%), Mongols (65.2%) and Kalmyks (62.6%) (Malyarchuk et al., 2010) . The frequency of haplogroup C decreases in South Asia and diminishes in Europe with only few representatives in the south of the continent.
The frequency map of Hg N, which is considered of Southern East Asia origin (Shi et al., 2013) , depicts the prevalence of this haplogroup mainly in Northern Eurasia. Thus, it is most numerous in European Russia, in particular in: Kursk (92.5%) (Mirabal et al., 2009) , Tver (78.9%), Mezen (53.7%), Krasnoborsk (39.6%), Pinega (39.5%) and Vologda (38.8%); in Siberia, among Yakuts (84.2%) (Derenko et al., 2007a; Sengupta et al., 2006) and Khanty (81.5%) (Mirabal et al., 2009) ; and among Swedish Saami (44.7%) (Karlsson et al., 2006) . Haplogroup N abruptly dissipates in lower latitudes, being virtually absent in southern Eurasia, with the exception of certain southwest Chinese ethnic groups such as Sichuan (13.2%) and Yunnan (10.7%) (Zhong et al., 2011) . Unlike the other considered haplogroups, Hg N has quite uneven distribution among European populations which is summarized in Table 1. It is apparent that Hg N is most prominent in East Slavic populations (Russians, Belarusians and Ukrainians) (Battaglia et al., 2009; Kushniarevich et al., 2013; Mirabal et al., 2009) and Swedish Saami (Karlsson et al., 2006) , but is almost lacking among other Europeans.
Despite its Asian origin (Sharma et al., 2007) , Hg Q is a minor Y-chromosome haplogroup in Eurasia having two occasional frequency peaks in Northwest Siberia among Kets and Selkups (93.8% and 66.4%, respectively) (Karafet et al., 2002) and in Altai people (17.4%) (Hammer et al., 2005) , as its frequency steeply declines outside these regions.
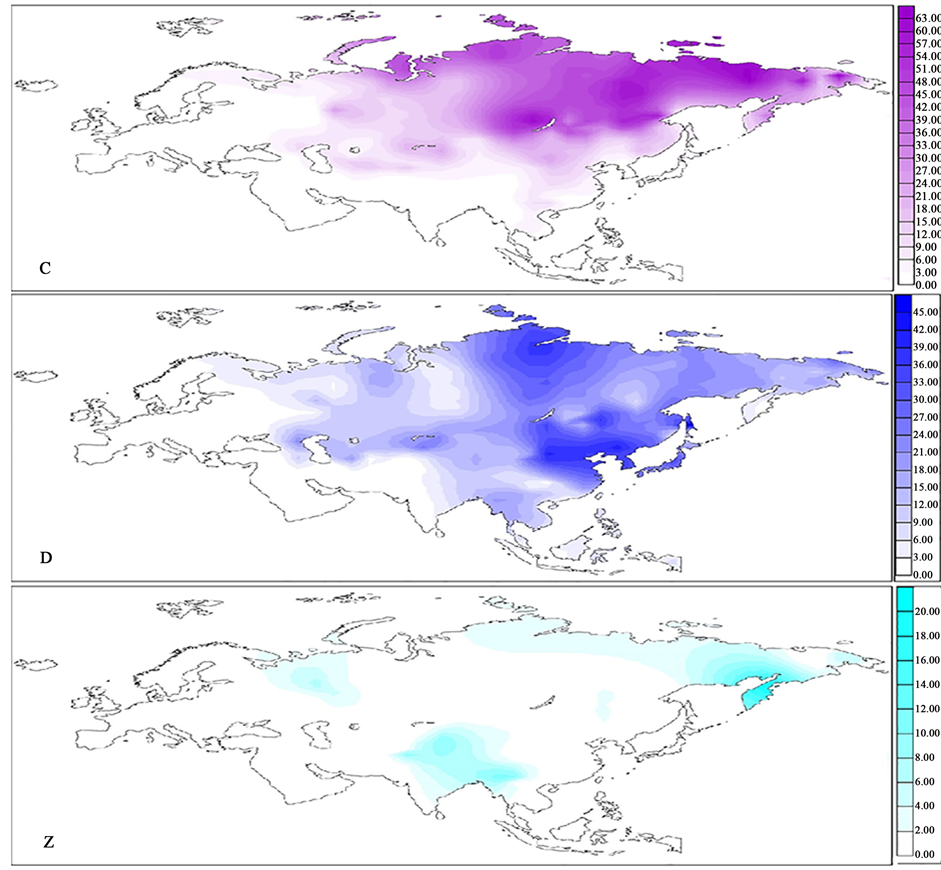
The spatial frequency maps of mtDNA haplogroups C, D and Z are presented in Figure 2.
Haplogroups C and D, which have evolved in Eastern Asia and expanded after the Last Glacial Maximum, comprise almost half of the northern and eastern Asian mtDNA gene pool (Derenko et al., 2007b; Derenko et al., 2010) .
Figure 1. Eurasian frequency distributions of Y-chromosome haplogroups C, N and Q.
Table 1. Percent frequency of Y-chromosome haplogroup N in European populations. N-number of analyzed individuals.
Figure 2. Eurasian frequency distributions of mtDNA haplogroups C, D and Z.
Haplogroup C is prevalent in Siberia. It has highest frequencies in regions inhabited by Yukaghirs (up to 72.2%) (Volodko et al., 2008) , Evenks (up to 71.8%) and Tofalars (up to 62.1%) (Derenko et al., 2010) . The occurrence of haplogroup C decreases to 5% towards the Volga-Ural region, Central Asia and the Indian subcontinent; which together delineate the diminishing border of the haplogroup.
The distributions of haplogroups C and D are almost overlapping. They slightly differ in haplogroup D frequency maximums which are found in: Oroks of Sakhalin (68.9%), Sojots from the Baikal region (46.7%), Orochens (43.2%), Manchus (42.6%) (Zhao et al., 2011) and Dolgans in North Siberia (39.5%) (Derenko et al., 2010) . Furthermore, the frequency of Hg D drops below 5% more westward and southward than Hg C, but it is also nearly inexistent in Europe and West Asia.
The minor mtDNA haplogroup Z, which is of Central Asian origin, has only a few frequency foci in Eurasia. They are scattered in: Northeastern Asia in Koryaks (21.7%); in the Volga-Ural region in Udmurts (16.7%) (Grosheva et al., 2014) and in parts of Central South Asia among Nu (16.7%) (Liu et al., 2011) , Hazara (13%) (Quintana-Murci et al., 2004) , and Kazakhs (11.3%) (Yao et al., 2004) .
The results of the chi-square test of the occurrence of Y-chromosome haplogroups C, N and Q (supplementary Table S3) and mtDNA haplogroups C and D (supplementary Table S4) are presented in Table 2 and Table 3, respectively.
Table 2. Comparison of the incidence of Y-chromosome haplogroups C, N and Q between Bulgarians and different population groups.
N-number of samples analyzed; *The populations consisting the population groups and their references are given in supplementary Table S3; **p-value population group vs Bulgarians.
Table 3. Comparison of the incidence of mtDNA haplogroups C and D between Bulgarians and different population groups.
N-number of samples analyzed; *The populations consisting the population groups and their references are given in supplementary Table S4; **p-value population group vs Bulgarians.
The occurrence of Y-chromosome haplogroups C, N and Q shows that there is statistically significant difference between Bulgarians and: Altaian, Siberian, Central Asian and European populations (p < 0.0001, p < 0.00001, p < 0.00001 and p = 0.000035, respectively). Furthermore, the chi-square test shows a Y-chromosome relation between Bulgarians and their European neighboring populations (p = 0.289988).
The absolute frequencies of mtDNA haplogroups C and D (Hg Z was excluded due to lack of data for part of the populations) point that there is significant difference between Bulgarians and: Altaian, Siberian, Central Asian and Caucasus populations (p < 0.0001, p < 0.0001, p < 0.0001 and p < 0.00001, respectively) and that Bulgarians do not statistically differ from Europeans and their European neighboring populations (p = 0.896 and p = 0.873002, respectively).
4. Discussion
The modern Bulgarian gene pool has been gradually formed from a succession of populations with different genetic impacts. According to novel historical, linguistic and archeological data, one of the strongest genetic signals in the modern Bulgarian gene pool should derive from proto-Bulgarians (Dimitrov, 2005; Rashev, 1993) .
It has been established that the modern Bulgarian mtDNA and Y-chromosome gene pools are overwhelmingly represented by Western Eurasian haplogroups. On the other hand, the Eastern Eurasian lineages among Bulgarians are represented by very low frequencies of mtDNA haplogroups C (0.2%), D (0.4%) and Z (0.1%) and Y-chromosome haplogroups C, N and Q (each 0.5%) (Karachanak et al., 2012, 2013) . Similar to these findings, ancient mtDNA analysis of proto-Bulgarian human remains has revealed the presence of only Western Eurasian haplogroups (Nesheva et al., 2015) .
In order to further genetically test the hypotheses about the origin of modern-and proto-Bulgarians, in the present study we have analyzed the distribution of Y-chromosome haplogroups C, N and Q and mtDNA haplogroups C, D and Z across Eurasia and between modern Bulgarians and different population groups.
The frequency distributions of mtDNA haplogroups C, D and Z and Y-chromosome haplogroups C, N and Q clearly depict their predominance in territories including those inhabited by Altaic (Turkic and Mongolic) populations. All of these haplogroups are underrepresented in Europe, with the exception of Y-chromosome haplogroup N, which is frequent only among East Slavic (Battaglia et al., 2009; Kushniarevich et al., 2013; Mirabal et al., 2009) populations and Swedish Saami (Karlsson et al., 2006) and is one of the features which genetically distinguish them from the remainder of the European populations.
The comparison of the incidence of Y-chromosome haplogroups C, N and Q points to a statistically significant difference between Bulgarians and Altaian, Siberian and Central Asian populations. Conversely, the chi- square test reveals a Y-chromosome link between Bulgarians and their European neighboring populations. The occurrence of mtDNA haplogroups C and D shows a similar pattern, with no association between Bulgarians and Altaian, Siberian, Central Asian and Caucasus populations and with similarity between Bulgarians and: their European neighboring populations and European populations as a whole. The discrepancy in the statistical significance of the distribution of Y-chromosome and mtDNA haplogroups in Bulgarians and Europeans is probably due to the mtDNA uniformity of Europe.
The Eurasian distribution of mtDNA haplogroups C, D and Z and Y-chromosome haplogroups C, N and Q contrasts their high frequency among Altaic populations and their occasional appearance in Bulgarians. Furthermore, the comparison of the occurrence of these haplogroups shows no link between Altaic populations and Bulgarians. Based on the substantial genetic input of proto-Bulgarians to the modern Bulgarian gene pool, our findings confirm the nonexistence of a close Y-chromosomal or mtDNA link between proto-Bulgarians on one hand and Altaic and Caucasus populations on the other.
Acknowledgements
The study was supported by the National Science Fund of Bulgaria, project: “Characterization of the anthropo-genetic identity of Bulgarians”, contract number DOO 2-110/22.05.2009.
Cite this paper
Sena Karachanak-Yankova,Desislava Nesheva,Angel S. Galabov,Draga Toncheva, (2015) Distribution of East Eurasian Y-Chromosome and Mitochondrial DNA Haplogroups across Eurasia: Insights into the Genetic Ancestry of Bulgarians. Advances in Anthropology,05,205-266. doi: 10.4236/aa.2015.54019
References
- 1. Abu-Amero, Achilli, A., Olivieri, A., Pala, M., Metspalu, E., Fornarino, S., Battaglia, V. et al. (2007). Mitochondrial DNA Variation of Modern Tuscans Supports the Near Eastern Origin of Etruscans. American Journal of Human Genetics, 80, 759-768.
http://dx.doi.org/10.1086/512822 - 2. álvarez-Iglesias, V., Mosquera-Miguel, A., Cerezo, M., Quintáns, B., Zarrabeitia, M. T., Cuscó, I. et al. (2009). New Population and Phylogenetic Features of the Internal Variation within Mitochondrial DNA Macro-Haplogroup R0. PLoS ONE, 4, e5112.
http://dx.doi.org/10.1371/journal.pone.0005112 - 3. Brandstatter, A., Klein, R., Duftner, N., Wiegand, P., & Parson, W. (2006). Application of a Quasi-Median Network Analysis for the Visualization of Character Conflicts to a Population Sample of Mitochondrial DNA Control Region Sequences from Southern Germany (Ulm). International Journal of Legal Medicine, 120, 310-314.
http://dx.doi.org/10.1007/s00414-006-0114-x - 4. Cardoso, S., Alfonso-Sanchez, M. A., Valverde, L., Odriozola, A., Perez-Miranda, A. M., Pena, J. A., & de Pancorbo, M. M. (2011). The Maternal Legacy of Basques in Northern Navarre: New Insights into the Mitochondrial DNA Diversity of the Franco-Cantabrian Area. American Journal of Physical Anthropology, 145, 480-488.
http://dx.doi.org/10.1002/ajpa.21532 - 5. Comas, D., Plaza, S., Wells, R. S., Yuldaseva, N., Lao, O., Calafell, F., & Bertranpetit, J. (2004). Admixture, Migrations, and Dispersals in Central Asia: Evidence from Maternal DNA Lineages. European Journal of Human Genetics, 12, 495- 504.
http://dx.doi.org/10.1038/sj.ejhg.5201160 - 6. Davidovic, S., Malyarchuk, B., Aleksic, J. M., Derenko, M., Topalovic, V., Litvinov, A. et al. (2015). Mitochondrial DNA Perspective of Serbian Genetic Diversity. American Journal of Physical Anthropology, 156, 449-465.
http://dx.doi.org/10.1002/ajpa.22670 - 7. Derenko, M., Malyarchuk, B., Grzybowski, T., Denisova, G., Rogalla, U., Perkova, M. et al. (2010). Origin and Post-Glacial Dispersal of Mitochondrial DNA Haplogroups C and D in Northern Asia. PLoS ONE, 5, e15214.
http://dx.doi.org/10.1371/journal.pone.0015214 - 8. Dulik, M. C., Zhadanov, S. I., Osipova, L. P., Askapuli, A., Gau, L., Gokcumen, O. et al. (2012). Mitochondrial DNA and Y Chromosome Variation Provides Evidence for a Recent Common Ancestry between Native Americans and Indigenous Altaians. American Journal of Human Genetics, 90, 229-246.
http://dx.doi.org/10.1016/j.ajhg.2011.12.014 - 9. Grzybowski, T., Malyarchuk, B. A., Derenko, M. V., Perkova, M. A., Bednarek, J., & Wozniak, M. (2007). Complex Interactions of the Eastern and Western Slavic Populations with Other European Groups as Revealed by Mitochondrial DNA Analysis. Forensic Science International: Genetics, 1, 141-147.
http://dx.doi.org/10.1016/j.fsigen.2007.01.010 - 10. Gubina, M. A., Girgol’kau, L. A., Babenko, V. N., Damba, L. D., Maksimov, V. N., & Voevoda, M. I. (2013). Mitochondrial DNA Polymorphism in Populations of Aboriginal Residents of the Far East. Russian Journal of Genetics, 49, 751-764.
http://dx.doi.org/10.1134/S1022795413070065 - 11. Hedman, M., Brandstatter, A., Pimenoff, V., Sistonen, P., Palo, J. U., Parson, W., & Sajantila, A. (2007). Finnish Mitochondrial DNA HVS-I and HVS-II Population Data. Forensic Science International, 172, 171-178.
http://dx.doi.org/10.1016/j.forsciint.2006.09.012 - 12. Irwin, J., Egyed, B., Saunier, J., Szamosi, G., O’Callaghan, J., Padar, Z., & Parsons, T. (2007). Hungarian mtDNA Population Databases from Budapest and the Baranya County Roma. International Journal of Legal Medicine, 121, 377-383.
http://dx.doi.org/10.1007/s00414-006-0128-4 - 13. Irwin, J. A., Ikramov, A., Saunier, J., Bodner, M., Amory, S., Rock, A. et al. (2010). The mtDNA Composition of Uzbekistan: A Microcosm of Central Asian Patterns. International Journal of Legal Medicine, 124, 195-204.
http://dx.doi.org/10.1007/s00414-009-0406-z - 14. Irwin, J., Saunier, J., Strouss, K., Paintner, C., Diegoli, T., Sturk, K. et al. (2008). Mitochondrial Control Region Sequences from Northern Greece and Greek Cypriots. International Journal of Legal Medicine, 122, 87-89.
http://dx.doi.org/10.1007/s00414-007-0173-7 - 15. Karachanak, S., Carossa, V., Nesheva, D., Olivieri, A., Pala, M., Hooshiar Kashani, B. et al. (2012). Bulgarians vs the Other European Populations: A Mitochondrial DNA Perspective. International Journal of Legal Medicine, 126, 497-503.
http://dx.doi.org/10.1007/s00414-011-0589-y - 16. Kushniarevich, A., Sivitskaya, L., Danilenko, N., Novogrodskii, T., Tsybovsky, I., Kiseleva, A. et al. (2013). Uniparental Genetic Heritage of Belarusians: Encounter of Rare Middle Eastern Matrilineages with a Central European Mitochondrial DNA Pool. PLoS ONE, 8, e66499.
http://dx.doi.org/10.1371/journal.pone.0066499 - 17. Lehocky, I., Baldovic, M., Kádasi, L., & Metspalu, E. (2008). A Database of Mitochondrial DNA Hypervariable Regions I and II Sequences of Individuals from Slovakia. Forensic Science International: Genetics, 2, e53-e59.
http://dx.doi.org/10.1016/j.fsigen.2007.12.008 - 18. Malyarchuk, B. A., Grzybowski, T., Derenko, M. V., Czarny, J., Drobnic, K., & Miscicka-Sliwka, D. (2003). Mitochondrial DNA Variability in Bosnians and Slovenians. Annals of Human Genetics, 67, 412-425.
http://dx.doi.org/10.1046/j.1469-1809.2003.00042.x - 19. Malyarchuk, B. A., Vanecek, T., Perkova, M. A., Derenko, M. V., & Sip, M. (2006). Mitochondrial DNA Variability in the Czech Population, with Application to the Ethnic History of Slavs. Human Biology, 78, 681-696.
http://dx.doi.org/10.1353/hub.2007.0014 - 20. Mielnik-Sikorska, M., Daca, P., Malyarchuk, B., Derenko, M., Skonieczna, K., Perkova, M. et al. (2013). The History of Slavs Inferred from Complete Mitochondrial Genome Sequences. PLoS ONE, 8, e54360.
http://dx.doi.org/10.1371/journal.pone.0054360 - 21. Mikkelsen, M., Sorensen, E., Rasmussen, E. M., & Morling, N. (2010). Mitochondrial DNA HV1 and HV2 Variation in Danes. Forensic Science International: Genetics, 4, e87-88.
http://dx.doi.org/10.1016/j.fsigen.2009.07.007 - 22. Prieto, L., Zimmermann, B., Goios, A., Rodriguez-Monge, A., Paneto, G. G., Alves, C. et al. (2011). The GHEP-EMPOP Collaboration on mtDNA Population Data—A New Resource for Forensic Casework. Forensic Science International: Genetics, 5, 146-151.
http://dx.doi.org/10.1016/j.fsigen.2010.10.013 - 23. Turchi, C., Buscemi, L., Previdere, C., Grignani, P., Brandstatter, A., Achilli, A. et al. (2008). Italian Mitochondrial DNA Database: Results of a Collaborative Exercise and Proficiency Testing. International Journal of Legal Medicine, 122, 199-204.
http://dx.doi.org/10.1007/s00414-007-0207-1 - 24. Yao, Y. G., Kong, Q. P., Wang, C. Y., Zhu, C. L., & Zhang, Y. P. (2004). Different Matrilineal Contributions to Genetic Structure of Ethnic Groups in the Silk Road Region in China. Molecular Biology and Evolution, 21, 2265-2280.
http://dx.doi.org/10.1093/molbev/msh238 - 25. Zgonjanin, D., Veselinovic, I., Kubat, M., Furac, I., Antov, M., Loncar, E., & Omorjan, R. (2010). Sequence Polymorphism of the Mitochondrial DNA Control Region in the Population of Vojvodina Province, Serbia. Legal Medicine (Tokyo, Japan), 12, 104-107. http://dx.doi.org/10.1016/j.legalmed.2009.10.007
- 26. Zheng, H.-X., Yan, S., Qin, Z.-D., & Jin, L. (2012). MtDNA Analysis of Global Populations Support That Major Population Expansions Began before Neolithic Time. Scientific Reports, 2, 745.
http://dx.doi.org/10.1038/srep00745 - 27. Zimmermann, B., Brandstatter, A., Duftner, N., Niederwieser, D., Spiroski, M., Arsov, T., & Parson, W. (2007). Mitochondrial DNA Control Region Population Data from Macedonia. Forensic Science International: Genetics, 1, e4-e9.
http://dx.doi.org/10.1016/j.fsigen.2007.03.002
Supplementary
Table S1. Absolute and percent frequencies of Y-chromosome haplogroups C (a), N (b) and Q (c) used to construct spatial frequency maps. N-number of analyzed individuals.
References S1(a)
Alonso, S., Flores, C., Cabrera, V., Alonso, A., Martin, P., Albarran, C. et al. (2005). The Place of the Basques in the European Y-Chromosome Diversity Landscape. European Journal of Human Genetics, 13, 1293-1302. http://dx.doi.org/10.1038/sj.ejhg.5201482
Al-Zahery, N., Pala, M., Battaglia, V., Grugni, V., Hamod, M. A., Kashani, B. H. et al. (2011). In Search of the Genetic Footprints of Sumerians: A Survey of Y-Chromosome and mtDNA Variation in the Marsh Arabs of Iraq. BMC Evolutionary Biology, 11, 288. http://dx.doi.org/10.1186/1471-2148-11-288
Balanovsky, O., Rootsi, S., Pshenichnov, A., Kivisild, T., Churnosov, M., Evseeva, I. et al. (2008). Two Sources of the Russian Patrilineal Heritage in Their Eurasian Context. American Journal of Human Genetics, 82, 236-250. http://dx.doi.org/10.1016/j.ajhg.2007.09.019
Battaglia, V., Fornarino, S., Al-Zahery, N., Olivieri, A., Pala, M., Myres, N. M. et al. (2009). Y-Chromosomal Evidence of the Cultural Diffusion of Agriculture in Southeast Europe. European Journal of Human Genetics, 17, 820-830. http://dx.doi.org/10.1038/ejhg.2008.249
Cadenas, A. M., Zhivotovsky, L. A., Cavalli-Sforza, L. L., Underhill, P. A., & Herrera, R. J. (2008). Y-Chromosome Diversity Characterizes the Gulf of Oman. European Journal of Human Genetics, 16, 374-386. http://dx.doi.org/10.1038/sj.ejhg.5201934
Capelli, C., Brisighelli, F., Scarnicci, F., Arredi, B., Caglia, A., Vetrugno, G. et al. (2007). Y Chromosome Genetic Variation in the Italian Peninsula Is Clinal and Supports an Admixture Model for the Mesolithic-Neolithic Encounter. Molecular Phylogenetics and Evolution, 44, 228-239. http://dx.doi.org/10.1016/j.ympev.2006.11.030
Capelli, C., Redhead, N., Abernethy, J. K., Gratrix, F., Wilson, J. F., Moen, T. et al. (2003). A Y Chromosome Census of the British Isles. Current Biology, 13, 979-984. http://dx.doi.org/10.1016/s0960-9822(03)00373-7
Cinnioğlu, C., King, R., Kivisild, T., Kalfoğlu, E., Atasoy, S., Cavalleri, G. L. et al. (2004). Excavating Y-Chromosome Haplotype Strata in Anatolia. Human Genetics, 114, 127-148. http://dx.doi.org/10.1007/s00439-003-1031-4
Dulik, M. C., Osipova, L. P., & Schurr, T. G. (2011). Y-Chromosome Variation in Altaian Kazakhs Reveals a Common Paternal Gene Pool for Kazakhs and the Influence of Mongolian Expansions. PLoS ONE, 6, e17548. http://dx.doi.org/10.1371/journal.pone.0017548
Flores, C., Maca-Meyer, N., Larruga, J. M., Cabrera, V. M., Karadsheh, N., & Gonzalez, A. M. (2005). Isolates in a Corridor of Migrations: A High-Resolution Analysis of Y-Chromosome Variation in Jordan. Journal of Human Genetics, 50, 435- 441. http://dx.doi.org/10.1007/s10038-005-0274-4
Francalacci, P., Morelli, L., Underhill, P. A., Lillie, A. S., Passarino, G., Useli, A. et al. (2003). Peopling of Three Medi- terranean Islands (Corsica, Sardinia, and Sicily) Inferred by Y-Chromosome Biallelic Variability. American Journal of Physical Anthropology, 121, 270-279. http://dx.doi.org/10.1002/ajpa.10265
Gonçalves, R., Freitas, A., Branco, M., Rosa, A., Fernandes, A. T., Zhivotovsky, L. A. et al. (2005). Y-Chromosome Lineages from Portugal, Madeira and Açores Record Elements of Sephardim and Berber Ancestry. Annals of Human Genetics, 69, 443-454. http://dx.doi.org/10.1111/j.1529-8817.2005.00161.x
Grugni, V., Battaglia, V., Hooshiar Kashani, B., Parolo, S., Al-Zahery, N., Achilli, A. et al. (2012). Ancient Migratory Events in the Middle East: New Clues from the Y-Chromosome Variation of Modern Iranians. PLoS ONE, 7, e41252. http://dx.doi.org/10.1371/journal.pone.0041252
Hammer, M. F., Karafet, T. M., Park, H., Omoto, K., Harihara, S., Stoneking, M., & Horai, S. (2005). Dual Origins of the Japanese: Common Ground for Hunter-Gatherer and Farmer Y Chromosomes. Journal of Human Genetics, 51, 47-58. http://dx.doi.org/10.1007/s10038-005-0322-0
Karachanak, S., Grugni, V., Fornarino, S., Nesheva, D., Al-Zahery, N., Battaglia, V. et al. (2013). Y-Chromosome Diversity in Modern Bulgarians: New Clues about Their Ancestry. PLoS ONE, 8, e56779. http://dx.doi.org/10.1371/journal.pone.0056779
Karlsson, A. O., Wallerström, T., Götherström, A., & Holmlund, G. (2006). Y-Chromosome Diversity in Sweden―A Long- Time Perspective. European Journal of Human Genetics, 14, 963-970. http://dx.doi.org/10.1038/sj.ejhg.5201651
King, R., Özcan, S., Carter, T., Kalfoğlu, E., Atasoy, S., Triantaphyllidis, C. et al. (2008). Differential Y-Chromosome Anatolian Influences on the Greek and Cretan Neolithic. Annals of Human Genetics, 72, 205-214. http://dx.doi.org/10.1111/j.1469-1809.2007.00414.x
Kushniarevich, A., Sivitskaya, L., Danilenko, N., Novogrodskii, T., Tsybovsky, I., Kiseleva, A. et al. (2013). Uniparental Genetic Heritage of Belarusians: Encounter of Rare Middle Eastern Matrilineages with a Central European Mitochondrial DNA Pool. PLoS ONE, 8, e66499. http://dx.doi.org/10.1371/journal.pone.0066499
Lacau, H., Gayden, T., Regueiro, M., Chennakrishnaiah, S., Bukhari, A., Underhill, P. A. et al. (2012). Afghanistan from a Y-Chromosome Perspective. European Journal of Human Genetics, 20, 1063-1070. http://dx.doi.org/10.1038/ejhg.2012.59
Larmuseau, M. H., Vanderheyden, N., Jacobs, M., Coomans, M., Larno, L., & Decorte, R. (2011). Micro-Geographic Distribution of Y-Chromosomal Variation in the Central-Western European Region Brabant. Forensic Science International: Genetics, 5, 95-99. http://dx.doi.org/10.1016/j.fsigen.2010.08.020
López-Parra, A., Gusmao, L., Tavares, L., Baeza, C., Amorim, A., Mesa, M. et al. (2009). In Search of the Pre- and Post-Neolithic Genetic Substrates in Iberia: Evidence from Y-Chromosome in Pyrenean Populations. Annals of Human Genetics, 73, 42-53. http://dx.doi.org/10.1111/j.1469-1809.2008.00478.x
Luis, J. R., Rowold, D. J., Regueiro, M., Caeiro, B., Cinnioğlu, C., Roseman, C. et al. (2004). The Levant versus the Horn of Africa: Evidence for Bidirectional Corridors of Human Migrations. The American Journal of Human Genetics, 74, 532- 544. http://dx.doi.org/10.1086/382286
Malyarchuk, B., Derenko, M., Denisova, G., Wozniak, M., Grzybowski, T., Dambueva, I., & Zakharov, I. (2010). Phylogeography of the Y-Chromosome Haplogroup C in Northern Eurasia. Annals of Human Genetics, 74, 539-546. http://dx.doi.org/10.1111/j.1469-1809.2010.00601.x
Martinez, L., Underhill, P. A., Zhivotovsky, L. A., Gayden, T., Moschonas, N. K., Chow, C.-E. T. et al. (2007). Paleolithic Y-Haplogroup Heritage Predominates in a Cretan Highland Plateau. European Journal of Human Genetics, 15, 485-493. http://www.nature.com/ejhg/journal/v15/n4/suppinfo/5201769s1.html http://dx.doi.org/10.1038/sj.ejhg.5201769
Martinez-Cruz, B., Ioana, M., Calafell, F., Arauna, L. R., Sanz, P., Ionescu, R. et al. (2012). Y-Chromosome Analysis in Individuals Bearing the Basarab Name of the First Dynasty of Wallachian Kings. PLoS ONE, 7, e41803. http://dx.doi.org/10.1371/journal.pone.0041803
Mirabal, S., Regueiro, M., Cadenas, A. M., Cavalli-Sforza, L. L., Underhill, P. A., Verbenko, D. A. et al. (2009). Y- Chromosome Distribution within the Geo-Linguistic Landscape of Northwestern Russia. European Journal of Human Genetics, 17, 1260-1273. http://www.nature.com/ejhg/journal/v17/n10/suppinfo/ejhg20096s1.html html http://dx.doi.org/10.1038/ejhg.2009.6
Noveski, P., Trivodalieva, S., Efremov, G., & Plaseska-Karanfilska, D. (2010). Y Chromosome Single Nucleotide Poly- morphisms Typing by SNaPshot Minisequencing. Balkan Journal of Medical Genetics, 13, 9-16. http://dx.doi.org/10.2478/v10034-010-0013-9
Regueiro, M., Cadenas, A., Gayden, T., Underhill, P., & Herrera, R. (2006). Iran: Tricontinental Nexus for Y-Chromosome Driven Migration. Human Heredity, 61, 132-143. http://dx.doi.org/10.1159/000093774
Regueiro, M., Rivera, L., Damnjanovic, T., Lukovic, L., Milasin, J., & Herrera, R. J. (2012). High Levels of Paleolithic Y-Chromosome Lineages Characterize Serbia. Gene, 498, 59-67. http://dx.doi.org/10.1016/j.gene.2012.01.030
Semino, O., Passarino, G., Oefner, P. J., Lin, A. A., Arbuzova, S., Beckman, L. E. et al. (2000). The Genetic Legacy of Paleolithic Homo Sapiens Sapiens in Extant Europeans: A Y Chromosome Perspective. Science, 290, 1155-1159. http://dx.doi.org/10.1126/science.290.5494.1155
Sengupta, S., Zhivotovsky, L. A., King, R., Mehdi, S., Edmonds, C. A., Chow, C.-E. T. et al. (2006). Polarity and Temporality of High-Resolution Y-Chromosome Distributions in India Identify Both Indigenous and Exogenous Expan- sions and Reveal Minor Genetic Influence of Central Asian Pastoralists. The American Journal of Human Genetics, 78, 202-221. http://dx.doi.org/10.1086/499411
Varzari, A., Kharkov, V., Nikitin, A. G., Raicu, F., Simonova, K., Stephan, W. et al. (2013). Paleo-Balkan and Slavic Contributions to the Genetic Pool of Moldavians: Insights from the Y Chromosome. PLoS ONE, 8, e53731. http://dx.doi.org/10.1371/journal.pone.0053731
Zhong, H., Shi, H., Qi, X.-B., Duan, Z.-Y., Tan, P.-P., Jin, L. et al. (2011). Extended Y Chromosome Investigation Suggests Postglacial Migrations of Modern Humans into East Asia via the Northern Route. Molecular Biology and Evolution, 28, 717-727. http://dx.doi.org/10.1093/molbev/msq247
References S1(b)
Alonso, S., Flores, C., Cabrera, V., Alonso, A., Martin, P., Albarran, C. et al. (2005). The Place of the Basques in the European Y-Chromosome Diversity Landscape. European Journal of Human Genetics, 13, 1293-1302. http://dx.doi.org/10.1038/sj.ejhg.5201482
Al-Zahery, N., Pala, M., Battaglia, V., Grugni, V., Hamod, M. A., Kashani, B. H. et al. (2011). In Search of the Genetic Footprints of Sumerians: A Survey of Y-Chromosome and mtDNA Variation in the Marsh Arabs of Iraq. BMC Evolutionary Biology, 11, 288. http://dx.doi.org/10.1186/1471-2148-11-288
Balanovsky, O., Rootsi, S., Pshenichnov, A., Kivisild, T., Churnosov, M., Evseeva, I. et al. (2008). Two Sources of the Russian Patrilineal Heritage in Their Eurasian Context. American Journal of Human Genetics, 82, 236-250. http://dx.doi.org/10.1016/j.ajhg.2007.09.019
Battaglia, V., Fornarino, S., Al-Zahery, N., Olivieri, A., Pala, M., Myres, N. M. et al.. (2009). Y-Chromosomal Evidence of the Cultural Diffusion of Agriculture in Southeast Europe. European Journal of Human Genetics, 17, 820-830. http://dx.doi.org/10.1038/ejhg.2008.249
Cadenas, A. M., Zhivotovsky, L. A., Cavalli-Sforza, L. L., Underhill, P. A., & Herrera, R. J. (2008). Y-Chromosome Diversity Characterizes the Gulf of Oman. European Journal of Human Genetics, 16, 374-386. http://dx.doi.org/10.1038/sj.ejhg.5201934
Capelli, C., Brisighelli, F., Scarnicci, F., Arredi, B., Caglia, A., Vetrugno, G. et al. (2007). Y chromosome Genetic Variation in the Italian Peninsula Is Clinal and Supports an Admixture Model for the Mesolithic-Neolithic Encounter. Molecular Phylogenetics and Evolution, 44, 228-239. http://dx.doi.org/10.1016/j.ympev.2006.11.030
Capelli, C., Redhead, N., Abernethy, J. K., Gratrix, F., Wilson, J. F., Moen, T. et al. (2003). A Y Chromosome Census of the British Isles. Current Biology, 13, 979-984. http://dx.doi.org/10.1016/s0960-9822(03)00373-7
Cinnioğlu, C., King, R., Kivisild, T., Kalfoğlu, E., Atasoy, S., Cavalleri, G. L. Et al. (2004). Excavating Y-Chromosome Haplotype Strata in Anatolia. Human genetics, 114, 127-148. http://dx.doi.org/10.1007/s00439-003-1031-4
Derenko, M., Malyarchuk, B., Denisova, G., Wozniak, M., Grzybowski, T., Dambueva, I., & Zakharov, I. (2007). Y-Chromosome Haplogroup N Dispersals from South Siberia to Europe. Journal of Human Genetics, 52, 763-770. http://dx.doi.org/10.1007/s10038-007-0179-5
Dulik, M. C., Osipova, L. P., & Schurr, T. G. (2011). Y-Chromosome Variation in Altaian Kazakhs Reveals a Common Paternal Gene Pool for Kazakhs and the Influence of Mongolian Expansions. PLoS ONE, 6, e17548. http://dx.doi.org/10.1371/journal.pone.0017548
Flores, C., Maca-Meyer, N., Larruga, J. M., Cabrera, V. M., Karadsheh, N., & Gonzalez, A. M. (2005). Isolates in a Corridor of Migrations: A High-Resolution Analysis of Y-Chromosome Variation in Jordan. Journal of Human Genetics, 50, 435- 441. http://dx.doi.org/10.1007/s10038-005-0274-4
Francalacci, P., Morelli, L., Underhill, P. A., Lillie, A. S., Passarino, G., Useli, A. et al. (2003). Peopling of Three Mediterranean Islands (Corsica, Sardinia, and Sicily) Inferred by Y-Chromosome Biallelic Variability. American Journal of Physical Anthropology, 121, 270-279. http://dx.doi.org/10.1002/ajpa.10265
Hammer, M. F., Karafet, T. M., Park, H., Omoto, K., Harihara, S., Stoneking, M., & Horai, S. (2005). Dual Origins of the Japanese: Common Ground for Hunter-Gatherer and Farmer Y Chromosomes. Journal of Human Genetics, 51, 47-58. http://dx.doi.org/10.1007/s10038-005-0322-0
Karachanak, S., Grugni, V., Fornarino, S., Nesheva, D., Al-Zahery, N., Battaglia, V. et al. (2013). Y-Chromosome Diversity in Modern Bulgarians: New Clues about Their Ancestry. PLoS ONE, 8, e56779. http://dx.doi.org/10.1371/journal.pone.0056779
Karlsson, A. O., Wallerström, T., Götherström, A., & Holmlund, G. (2006). Y-Chromosome Diversity in Sweden―A Long-Time Perspective. European Journal of Human Genetics, 14, 963-970. http://dx.doi.org/10.1038/sj.ejhg.5201651
King, R., Özcan, S., Carter, T., Kalfoğlu, E., Atasoy, S., Triantaphyllidis, C. et al. (2008). Differential Y-Chromosome Anatolian Influences on the Greek and Cretan Neolithic. Annals of Human Genetics, 72, 205-214. http://dx.doi.org/10.1111/j.1469-1809.2007.00414.x
Kushniarevich, A., Sivitskaya, L., Danilenko, N., Novogrodskii, T., Tsybovsky, I., Kiseleva, A. et al. (2013). Uniparental Genetic Heritage of Belarusians: Encounter of Rare Middle Eastern Matrilineages with a Central European Mitochondrial DNA Pool. PLoS ONE, 8, e66499. http://dx.doi.org/10.1371/journal.pone.0066499
Lacau, H., Gayden, T., Regueiro, M., Chennakrishnaiah, S., Bukhari, A., Underhill, P. A. et al. (2012). Afghanistan from a Y-Chromosome Perspective. European Journal of Human Genetics, 20, 1063-1070. http://dx.doi.org/10.1038/ejhg.2012.59
Larmuseau, M. H., Vanderheyden, N., Jacobs, M., Coomans, M., Larno, L., & Decorte, R. (2011). Micro-Geographic Distribution of Y-Chromosomal Variation in the Central-Western European Region Brabant. Forensic Science International: Genetics, 5, 95-99. http://dx.doi.org/10.1016/j.fsigen.2010.08.020
López-Parra, A., Gusmao, L., Tavares, L., Baeza, C., Amorim, A., Mesa, M. et al. (2009). In Search of the Pre- and Post-Neolithic Genetic Substrates in Iberia: Evidence from Y-Chromosome in Pyrenean Populations. Annals of Human Genetics, 73, 42-53. http://dx.doi.org/10.1111/j.1469-1809.2008.00478.x
Luis, J. R., Rowold, D. J., Regueiro, M., Caeiro, B., Cinnioğlu, C., Roseman, C. et al. (2004). The Levant versus the Horn of Africa: Evidence for Bidirectional Corridors of Human Migrations. The American Journal of Human Genetics, 74, 532- 544. http://dx.doi.org/10.1086/382286
Martinez, L., Underhill, P. A., Zhivotovsky, L. A., Gayden, T., Moschonas, N. K., Chow, C.-E. T. et al. (2007). Paleolithic Y-Haplogroup Heritage Predominates in a Cretan Highland Plateau. European Journal of Human Genetics, 15, 485-493. http://www.nature.com/ejhg/journal/v15/n4/suppinfo/5201769s1.html http://dx.doi.org/10.1038/sj.ejhg.5201769
Martinez-Cruz, B., Ioana, M., Calafell, F., Arauna, L. R., Sanz, P., Ionescu, R. et al. (2012). Y-Chromosome Analysis in Individuals Bearing the Basarab Name of the First Dynasty of Wallachian Kings. PLoS ONE, 7, e41803. http://dx.doi.org/10.1371/journal.pone.0041803
Mirabal, S., Regueiro, M., Cadenas, A. M., Cavalli-Sforza, L. L., Underhill, P. A., Verbenko, D. A. et al. (2009). Y- Chromosome Distribution within the Geo-Linguistic landscape of Northwestern Russia. European Journal of Human Genetics, 17, 1260-1273. http://www.nature.com/ejhg/journal/v17/n10/suppinfo/ejhg20096s1.html http://dx.doi.org/10.1038/ejhg.2009.6
Noveski, P., Trivodalieva, S., Efremov, G., & Plaseska-Karanfilska, D. (2010). Y Chromosome Single Nucleotide Polymorphisms Typing by SNaPshot Minisequencing. Balkan Journal of Medical Genetics, 13, 9-16. http://dx.doi.org/10.2478/v10034-010-0013-9
Regueiro, M., Cadenas, A., Gayden, T., Underhill, P., & Herrera, R. (2006). Iran: Tricontinental Nexus for Y-Chromosome Driven Migration. Human Heredity, 61, 132-143. http://dx.doi.org/10.1159/000093774
Regueiro, M., Rivera, L., Damnjanovic, T., Lukovic, L., Milasin, J., & Herrera, R. J. (2012). High Levels of Paleolithic Y-Chromosome Lineages Characterize Serbia. Gene, 498, 59-67. http://dx.doi.org/10.1016/j.gene.2012.01.030
Semino, O., Passarino, G., Oefner, P. J., Lin, A. A., Arbuzova, S., Beckman, L. E. et al. (2000). The Genetic Legacy of Paleolithic Homo Sapiens Sapiens in Extant Europeans: A Y Chromosome Perspective. Science, 290, 1155-1159. http://dx.doi.org/10.1126/science.290.5494.1155
Sengupta, S., Zhivotovsky, L. A., King, R., Mehdi, S., Edmonds, C. A., Chow, C.-E. T. et al. (2006). Polarity and Temporality of High-Resolution Y-Chromosome Distributions in India Identify Both Indigenous and Exogenous Expansions and Reveal Minor Genetic Influence of Central Asian Pastoralists. The American Journal of Human Genetics, 78, 202- 221. http://dx.doi.org/10.1086/499411
Varzari, A., Kharkov, V., Nikitin, A. G., Raicu, F., Simonova, K., Stephan, W. et al. (2013). Paleo-Balkan and Slavic Contributions to the Genetic Pool of Moldavians: Insights from the Y Chromosome. PLoS ONE, 8, e53731. http://dx.doi.org/10.1371/journal.pone.0053731
Zhong, H., Shi, H., Qi, X.-B., Duan, Z.-Y., Tan, P.-P., Jin, L. et al. (2011). Extended Y Chromosome Investigation Suggests Postglacial Migrations of Modern Humans into East Asia via the Northern Route. Molecular Biology and Evolution, 28, 717-727. http://dx.doi.org/10.1093/molbev/msq247
References S1(c)
Alonso, S., Flores, C., Cabrera, V., Alonso, A., Martin, P., Albarran, C. et al. (2005). The Place of the Basques in the European Y-Chromosome Diversity Landscape. European Journal of Human Genetics, 13, 1293-1302. http://dx.doi.org/10.1038/sj.ejhg.5201482
Al-Zahery, N., Pala, M., Battaglia, V., Grugni, V., Hamod, M. A., Kashani, B. H. et al. (2011). In Search of the Genetic Footprints of Sumerians: A Survey of Y-Chromosome and mtDNA Variation in the Marsh Arabs of Iraq. BMC Evolutionary Biology, 11, 288. http://dx.doi.org/10.1186/1471-2148-11-288
Balanovsky, O., Dibirova, K., Dybo, A., Mudrak, O., Frolova, S., Pocheshkhova, E. et al. (2011). Parallel Evolution of Genes and Languages in the Caucasus Region. Molecular Biology and Evolution, 28, 2905-2920. http://dx.doi.org/10.1093/molbev/msr126
Balanovsky, O., Rootsi, S., Pshenichnov, A., Kivisild, T., Churnosov, M., Evseeva, I. et al. (2008). Two Sources of the Russian Patrilineal Heritage in Their Eurasian Context. American Journal of Human Genetics, 82, 236-250. http://dx.doi.org/10.1016/j.ajhg.2007.09.019
Battaglia, V., Fornarino, S., Al-Zahery, N., Olivieri, A., Pala, M., Myres, N. M. et al.. (2009). Y-Chromosomal Evidence of the Cultural Diffusion of Agriculture in Southeast Europe. European Journal of Human Genetics, 17, 820-830. http://dx.doi.org/10.1038/ejhg.2008.249
Cadenas, A. M., Zhivotovsky, L. A., Cavalli-Sforza, L. L., Underhill, P. A., & Herrera, R. J. (2008). Y-Chromosome Diversity Characterizes the Gulf of Oman. European Journal of Human Genetics, 16, 374-386. http://dx.doi.org/10.1038/sj.ejhg.5201934
Capelli, C., Brisighelli, F., Scarnicci, F., Arredi, B., Caglia, A., Vetrugno, G. et al. (2007). Y Chromosome Genetic Variation in the Italian Peninsula Is Clinal and Supports an Admixture Model for the Mesolithic-Neolithic Encounter. Molecular Phylogenetics and Evolution, 44, 228-239. http://dx.doi.org/10.1016/j.ympev.2006.11.030
Cinnioğlu, C., King, R., Kivisild, T., Kalfoğlu, E., Atasoy, S., Cavalleri, G. L. et al. (2004). Excavating Y-Chromosome Haplotype Strata in Anatolia. Human genetics, 114, 127-148. http://dx.doi.org/10.1007/s00439-003-1031-4
Dulik, M. C., Osipova, L. P., & Schurr, T. G. (2011). Y-Chromosome Variation in Altaian Kazakhs Reveals a Common Paternal Gene Pool for Kazakhs and the Influence of Mongolian Expansions. PLoS ONE, 6, e17548. http://dx.doi.org/10.1371/journal.pone.0017548
Flores, C., Maca-Meyer, N., Larruga, J. M., Cabrera, V. M., Karadsheh, N., & Gonzalez, A. M. (2005). Isolates in a Corridor of Migrations: A High-Resolution Analysis of Y-Chromosome Variation in Jordan. Journal of Human Genetics, 50, 435- 441. http://dx.doi.org/10.1007/s10038-005-0274-4
Francalacci, P., Morelli, L., Underhill, P. A., Lillie, A. S., Passarino, G., Useli, A. et al. (2003). Peopling of Three Mediterranean Islands (Corsica, Sardinia, and Sicily) inferred by Y-Chromosome Biallelic Variability. American Journal of Physical Anthropology, 121, 270-279. http://dx.doi.org/10.1002/ajpa.10265
Gonçalves, R., Freitas, A., Branco, M., Rosa, A., Fernandes, A. T., Zhivotovsky, L. A. et al. (2005). Y-Chromosome Lineages from Portugal, Madeira and Açores Record Elements of Sephardim and Berber Ancestry. Annals of Human Genetics, 69, 443-454. http://dx.doi.org/10.1111/j.1529-8817.2005.00161.x
Grugni, V., Battaglia, V., Hooshiar Kashani, B., Parolo, S., Al-Zahery, N., Achilli, A. et al. (2012). Ancient Migratory Events in the Middle East: New Clues from the Y-Chromosome Variation of Modern Iranians. PLoS ONE, 7, e41252. http://dx.doi.org/10.1371/journal.pone.0041252
Hammer, M. F., Karafet, T. M., Park, H., Omoto, K., Harihara, S., Stoneking, M., & Horai, S. (2005). Dual Origins of the Japanese: Common Ground for Hunter-Gatherer and Farmer Y Chromosomes. Journal of Human Genetics, 51, 47-58. http://dx.doi.org/10.1007/s10038-005-0322-0
Karachanak, S., Grugni, V., Fornarino, S., Nesheva, D., Al-Zahery, N., Battaglia, V. et al. (2013). Y-Chromosome Diversity in Modern Bulgarians: New Clues about Their Ancestry. PLoS ONE, 8, e56779. http://dx.doi.org/10.1371/journal.pone.0056779
Karafet, T. M., Osipova, L. P., Gubina, M. A., Posukh, O. L., Zegura, S. L., & Hammer, M. F. (2002). High Levels of Y-Chromosome Differentiation among Native Siberian Populations and the Genetic Signature of a Boreal Hunter- Gatherer Way of Life. Human Biology, 74, 761-789. http://dx.doi.org/10.1353/hub.2003.0006
Karlsson, A. O., Wallerström, T., Götherström, A., & Holmlund, G. (2006). Y-Chromosome Diversity in Sweden―A Long-Time Perspective. European Journal of Human Genetics, 14, 963-970. http://dx.doi.org/10.1038/sj.ejhg.5201651
King, R., Özcan, S., Carter, T., Kalfoğlu, E., Atasoy, S., Triantaphyllidis, C. et al. (2008). Differential Y-Chromosome Anatolian Influences on the Greek and Cretan Neolithic. Annals of Human Genetics, 72, 205-214. http://dx.doi.org/10.1111/j.1469-1809.2007.00414.x
Kushniarevich, A., Sivitskaya, L., Danilenko, N., Novogrodskii, T., Tsybovsky, I., Kiseleva, A. et al. (2013). Uniparental Genetic Heritage of Belarusians: Encounter of Rare Middle Eastern Matrilineages with a Central European Mitochondrial DNA Pool. PLoS ONE, 8, e66499. http://dx.doi.org/10.1371/journal.pone.0066499
Lacau, H., Gayden, T., Regueiro, M., Chennakrishnaiah, S., Bukhari, A., Underhill, P. A. et al. (2012). Afghanistan from a Y-Chromosome Perspective. European Journal of Human Genetics, 20, 1063-1070. http://dx.doi.org/10.1038/ejhg.2012.59
Larmuseau, M. H., Vanderheyden, N., Jacobs, M., Coomans, M., Larno, L., & Decorte, R. (2011). Micro-Geographic Distribution of Y-Chromosomal Variation in the Central-Western European Region Brabant. Forensic Science International: Genetics, 5, 95-99. http://dx.doi.org/10.1016/j.fsigen.2010.08.020
López-Parra, A., Gusmao, L., Tavares, L., Baeza, C., Amorim, A., Mesa, M. et al. (2009). In Search of the Pre- and Post-Neolithic Genetic Substrates in Iberia: Evidence from Y-Chromosome in Pyrenean Populations. Annals of Human Genetics, 73, 42-53. http://dx.doi.org/10.1111/j.1469-1809.2008.00478.x
Luis, J. R., Rowold, D. J., Regueiro, M., Caeiro, B., Cinnioğlu, C., Roseman, C. et al. (2004). The Levant versus the Horn of Africa: Evidence for Bidirectional Corridors of Human Migrations. The American Journal of Human Genetics, 74, 532- 544. http://dx.doi.org/10.1086/382286
Martinez, L., Underhill, P. A., Zhivotovsky, L. A., Gayden, T., Moschonas, N. K., Chow, C.-E. T. et al. (2007). Paleolithic Y-Haplogroup Heritage Predominates in a Cretan Highland Plateau. European Journal of Human Genetics, 15, 485-493. http://www.nature.com/ejhg/journal/v15/n4/suppinfo/5201769s1.html http://dx.doi.org/10.1038/sj.ejhg.5201769
Martinez-Cruz, B., Ioana, M., Calafell, F., Arauna, L. R., Sanz, P., Ionescu, R. et al. (2012). Y-Chromosome Analysis in Individuals Bearing the Basarab Name of the First Dynasty of Wallachian Kings. PLoS ONE, 7, e41803. http://dx.doi.org/10.1371/journal.pone.0041803
Mirabal, S., Regueiro, M., Cadenas, A. M., Cavalli-Sforza, L. L., Underhill, P. A., Verbenko, D. A. et al. (2009). Y-Chromosome Distribution within the Geo-Linguistic Landscape of Northwestern Russia. European Journal of Human Genetics, 17, 1260-1273. http://www.nature.com/ejhg/journal/v17/n10/suppinfo/ejhg20096s1.html http://dx.doi.org/10.1038/ejhg.2009.6
Noveski, P., Trivodalieva, S., Efremov, G., & Plaseska-Karanfilska, D. (2010). Y Chromosome Single Nucleotide Polymorphisms Typing by SNaPshot Minisequencing. Balkan Journal of Medical Genetics, 13, 9-16. http://dx.doi.org/10.2478/v10034-010-0013-9
Regueiro, M., Cadenas, A., Gayden, T., Underhill, P., & Herrera, R. (2006). Iran: Tricontinental Nexus for Y-Chromosome Driven Migration. Human Heredity, 61, 132-143. http://dx.doi.org/10.1159/000093774
Regueiro, M., Rivera, L., Damnjanovic, T., Lukovic, L., Milasin, J., & Herrera, R. J. (2012). High Levels of Paleolithic Y-Chromosome Lineages Characterize Serbia. Gene, 498, 59-67. http://dx.doi.org/10.1016/j.gene.2012.01.030
Semino, O., Passarino, G., Oefner, P. J., Lin, A. A., Arbuzova, S., Beckman, L. E. et al. (2000). The Genetic Legacy of Paleolithic Homo Sapiens Sapiens in Extant Europeans: A Y Chromosome Perspective. Science, 290, 1155-1159. http://dx.doi.org/10.1126/science.290.5494.1155
Sengupta, S., Zhivotovsky, L. A., King, R., Mehdi, S., Edmonds, C. A., Chow, C.-E. T. et al. (2006). Polarity and Temporality of High-Resolution Y-Chromosome Distributions in India Identify Both Indigenous and Exogenous Expansions and Reveal Minor Genetic Influence of Central Asian Pastoralists. The American Journal of Human Genetics, 78, 202-221. http://dx.doi.org/10.1086/499411
Varzari, A., Kharkov, V., Nikitin, A. G., Raicu, F., Simonova, K., Stephan, W. et al. (2013). Paleo-Balkan and Slavic Contributions to the Genetic Pool of Moldavians: Insights from the Y Chromosome. PLoS ONE, 8, e53731. http://dx.doi.org/10.1371/journal.pone.0053731
Zhong, H., Shi, H., Qi, X.-B., Duan, Z.-Y., Tan, P.-P., Jin, L. et al. (2011). Extended Y Chromosome Investigation Suggests Postglacial Migrations of Modern Humans into East Asia via the Northern Route. Molecular Biology and Evolution, 28, 717-727. http://dx.doi.org/10.1093/molbev/msq247
Table S2. Absolute and percent frequencies of mtDNA haplogroup C (a), D (b) and Z (c) used to construct spatial frequency maps. N-number of analyzed individuals.
References S2(a)
Abu-Amero, K. K., Gonzalez, A. M., Larruga, J. M., Bosley, T. M., & Cabrera, V. M. (2007). Eurasian and African Mitochondrial DNA Influences in the Saudi Arabian Population. BMC Evolutionary Biology, 7, 32.
http://dx.doi.org/10.1186/1471-2148-7-32
Abu-Amero, K. K., Larruga, J. M., Cabrera, V. M., & Gonzalez, A. M. (2008). Mitochondrial DNA Structure in the Arabian Peninsula. BMC Evolutionary Biology, 8, 45. http://dx.doi.org/10.1186/1471-2148-8-45
Achilli, A., Olivieri, A., Pala, M., Metspalu, E., Fornarino, S., Battaglia, V. et al. (2007). Mitochondrial DNA Variation of Modern Tuscans Supports the Near Eastern Origin of Etruscans. American Journal of Human Genetics, 80, 759-768. http://dx.doi.org/10.1086/512822
Álvarez-Iglesias, V., Mosquera-Miguel, A., Cerezo, M., Quintáns, B., Zarrabeitia, M. T., Cuscó, I. et al. (2009). New Population and Phylogenetic Features of the Internal Variation within Mitochondrial DNA Macro-Haplogroup R0. PLoS ONE, 4, e5112. http://dx.doi.org/10.1371/journal.pone.0005112
Al-Zahery, N., Semino, O., Benuzzi, G., Magri, C., Passarino, G., Torroni, A., & Santachiara-Benerecetti, A. S. (2003). Y-Chromosome and mtDNA Polymorphisms in Iraq, a Crossroad of the Early Human Dispersal and of Post-Neolithic Migrations. Molecular Phylogenetics and Evolution, 28, 458-472. http://dx.doi.org/10.1016/S1055-7903(03)00039-3
Brandstatter, A., Klein, R., Duftner, N., Wiegand, P., & Parson, W. (2006). Application of a Quasi-Median Network Analysis for the Visualization of Character Conflicts to a Population Sample of Mitochondrial DNA Control Region Sequences from Southern Germany (Ulm). International Journal of Legal Medicine, 120, 310-314. http://dx.doi.org/10.1007/s00414-006-0114-x
Brandstatter, A., Niederstatter, H., Pavlic, M., Grubwieser, P., & Parson, W. (2007). Generating Population Data for the EMPOP Database―An Overview of the mtDNA Sequencing and Data Evaluation Processes Considering 273 Austrian Control Region Sequences as Example. Forensic Science International, 166, 164-175. http://dx.doi.org/10.1016/j.forsciint.2006.05.006
Comas, D., Plaza, S., Wells, R. S., Yuldaseva, N., Lao, O., Calafell, F., & Bertranpetit, J. (2004). Admixture, Migrations, and Dispersals in Central Asia: Evidence from Maternal DNA Lineages. European Journal of Human Genetics, 12, 495- 504. http://dx.doi.org/10.1038/sj.ejhg.5201160
Derenko, M., Malyarchuk, B., Bahmanimehr, A., Denisova, G., Perkova, M., Farjadian, S., & Yepiskoposyan, L. (2013). Complete Mitochondrial DNA Diversity in Iranians. PLoS ONE, 8, e80673. http://dx.doi.org/10.1371/journal.pone.0080673
Derenko, M., Malyarchuk, B., Denisova, G., Perkova, M., Rogalla, U., Grzybowski, T. et al. (2012). Complete Mitochondrial DNA Analysis of Eastern Eurasian Haplogroups Rarely Found in Populations of Northern Asia and Eastern Europe. PLoS ONE, 7, e32179. http://dx.doi.org/10.1371/journal.pone.0032179
Derenko, M., Malyarchuk, B., Grzybowski, T., Denisova, G., Dambueva, I., Perkova, M. et al. (2007). Phylogeographic Analysis of Mitochondrial DNA in Northern Asian Populations. The American Journal of Human Genetics, 81, 1025- 1041. http://dx.doi.org/10.1086/522933
Derenko, M., Malyarchuk, B., Grzybowski, T., Denisova, G., Rogalla, U., Perkova, M. et al. (2010). Origin and Post-Glacial Dispersal of Mitochondrial DNA Haplogroups C and D in Northern Asia. PLoS ONE, 5, e15214. http://dx.doi.org/10.1371/journal.pone.0015214
Grosheva, A., Shneider, Y. V., Zhukova, O., Morozova, I. Y., & Rychkov, S. Y. (2014). Features of the Udmurt Mitochondrial Gene Pool in Relation to Tribal Structure. Russian Journal of Genetics, 50, 975-986. http://dx.doi.org/10.1134/S1022795414090063
Grzybowski, T., Malyarchuk, B. A., Derenko, M. V., Perkova, M. A., Bednarek, J., & Wozniak, M. (2007). Complex Interactions of the Eastern and Western Slavic Populations with Other European Groups as Revealed by Mitochondrial DNA Analysis. Forensic Science International: Genetics, 1, 141-147. http://dx.doi.org/10.1016/j.fsigen.2007.01.010
Gubina, M. A., Girgol’kau, L. A., Babenko, V. N., Damba, L. D., Maksimov, V. N., & Voevoda, M. I. (2013). Mitochondrial DNA Polymorphism in Populations of Aboriginal Residents of the Far East. Russian Journal of Genetics, 49, 751-764. http://dx.doi.org/10.1134/S1022795413070065
Hedman, M., Brandstätter, A., Pimenoff, V., Sistonen, P., Palo, J. U., Parson, W., & Sajantila, A. (2007). Finnish Mitochondrial DNA HVS-I and HVS-II Population Data. Forensic Science International, 172, 171-178. http://dx.doi.org/10.1016/j.forsciint.2006.09.012
Irwin, J. A., Ikramov, A., Saunier, J., Bodner, M., Amory, S., Rock, A. et al. (2010). The mtDNA Composition of Uzbekistan: A Microcosm of Central Asian Patterns. International Journal of Legal Medicine, 124, 195-204. http://dx.doi.org/10.1007/s00414-009-0406-z
Irwin, J., Egyed, B., Saunier, J., Szamosi, G., O’Callaghan, J., Padar, Z., & Parsons, T. (2007). Hungarian mtDNA Population Databases from Budapest and the Baranya County Roma. International Journal of Legal Medicine, 121, 377- 383. http://dx.doi.org/10.1007/s00414-006-0128-4
Irwin, J., Saunier, J., Strouss, K., Paintner, C., Diegoli, T., Sturk, K. et al. (2008). Mitochondrial Control Region Sequences from Northern Greece and Greek Cypriots. International Journal of Legal Medicine, 122, 87-89. http://dx.doi.org/10.1007/s00414-007-0173-7
Karachanak, S., Carossa, V., Nesheva, D., Olivieri, A., Pala, M., Hooshiar Kashani, B. et al. (2012). Bulgarians vs the Other European Populations: A Mitochondrial DNA Perspective. International Journal of Legal Medicine, 126, 497-503. http://dx.doi.org/10.1007/s00414-011-0589-y
Kushniarevich, A., Sivitskaya, L., Danilenko, N., Novogrodskii, T., Tsybovsky, I., Kiseleva, A. et al. (2013). Uniparental Genetic Heritage of Belarusians: Encounter of Rare Middle Eastern Matrilineages with a Central European Mitochondrial DNA Pool. PLoS ONE, 8, e66499. http://dx.doi.org/10.1371/journal.pone.0066499
Lehocký, I., Baldovič, M., Kádaši, Ľ., & Metspalu, E. (2008). A Database of Mitochondrial DNA Hypervariable Regions I and II Sequences of Individuals from Slovakia. Forensic Science International: Genetics, 2, e53-e59. http://dx.doi.org/10.1016/j.fsigen.2007.12.008
Liu, C., Wang, S.-Y., Zhao, M., Xu, Z.-Y., Hu, Y.-H., Chen, F. et al. (2011). Mitochondrial DNA Polymorphisms in Gelao Ethnic Group Residing in Southwest China. Forensic Science International: Genetics, 5, e4-e10. http://dx.doi.org/10.1016/j.fsigen.2010.04.007
Malyarchuk, B. A., Grzybowski, T., Derenko, M. V., Czarny, J., Drobnič, K., & Miścicka-Śliwka, D. (2003). Mitochondrial DNA Variability in Bosnians and Slovenians. Annals of Human Genetics, 67, 412-425. http://dx.doi.org/10.1046/j.1469-1809.2003.00042.x
Malyarchuk, B. A., Vanecek, T., Perkova, M. A., Derenko, M. V., & Sip, M. (2006). Mitochondrial DNA Variability in the Czech Population, with Application to the Ethnic History of Slavs. Human Biology, 78, 681-696. http://dx.doi.org/10.1353/hub.2007.0014
Malyarchuk, B., Derenko, M., Denisova, G., & Kravtsova, O. (2010). Mitogenomic Diversity in Tatars from the Volga-Ural Region of Russia. Molecular Biology and Evolution, 27, 2220-2226. http://dx.doi.org/10.1093/molbev/msq065
Mielnik-Sikorska, M., Daca, P., Malyarchuk, B., Derenko, M., Skonieczna, K., Perkova, M. et al. (2013). The History of Slavs Inferred from Complete Mitochondrial Genome Sequences. PLoS ONE, 8, e54360. http://dx.doi.org/10.1371/journal.pone.0054360
Mikkelsen, M., Sorensen, E., Rasmussen, E. M., & Morling, N. (2010). Mitochondrial DNA HV1 and HV2 Variation in Danes. Forensic Science International: Genetics, 4, e87-88. http://dx.doi.org/10.1016/j.fsigen.2009.07.007
Pimenoff, V. N., Comas, D., Palo, J. U., Vershubsky, G., Kozlov, A., & Sajantila, A. (2008). Northwest Siberian Khanty and Mansi in the Junction of West and East Eurasian Gene Pools as Revealed by Uniparental Markers. European Journal of Human Genetics, 16, 1254-1264. http://www.nature.com/ejhg/journal/v16/n10/suppinfo/ejhg2008101s1.html
http://dx.doi.org/10.1038/ejhg.2008.101
Prieto, L., Zimmermann, B., Goios, A., Rodriguez-Monge, A., Paneto, G. G., Alves, C. et al. (2011). The GHEP-EMPOP Collaboration on mtDNA Population Data―A New Resource for Forensic Casework. Forensic Science International: Genetics, 5, 146-151. http://dx.doi.org/10.1016/j.fsigen.2010.10.013
Quintana-Murci, L., Chaix, R., Wells, R. S., Behar, D. M., Sayar, H., Scozzari, R. et al. (2004). Where West Meets East: The Complex mtDNA Landscape of the Southwest and Central Asian Corridor. American Journal of Human Genetics, 74, 827-845. http://dx.doi.org/10.1086/383236
Richard, C., Pennarun, E., Kivisild, T., Tambets, K., Tolk, H. V., Metspalu, E. et al. (2007). An mtDNA Perspective of French Genetic Variation. Annals of Human Biology, 34, 68-79. http://dx.doi.org/10.1080/03014460601076098
Simonson, T. S., Xing, J., Barrett, R., Jerah, E., Loa, P., Zhang, Y. et al. (2011). Ancestry of the Iban Is Predominantly Southeast Asian: Genetic Evidence from Autosomal, Mitochondrial, and Y Chromosomes. PLoS ONE, 6, e16338. http://dx.doi.org/10.1371/journal.pone.0016338
Turchi, C., Buscemi, L., Previdere, C., Grignani, P., Brandstatter, A., Achilli, A. et al. (2008). Italian Mitochondrial DNA Database: Results of a Collaborative Exercise and Proficiency Testing. International Journal of Legal Medicine, 122, 199-204. http://dx.doi.org/10.1007/s00414-007-0207-1
Volodko, N. V., Starikovskaya, E. B., Mazunin, I. O., Eltsov, N. P., Naidenko, P. V., Wallace, D. C., & Sukernik, R. I. (2008). Mitochondrial Genome Diversity in Arctic Siberians, with Particular Reference to the Evolutionary History of Beringia and Pleistocenic Peopling of the Americas. The American Journal of Human Genetics, 82, 1084-1100. http://dx.doi.org/10.1016/j.ajhg.2008.03.019
Yao, Y. G., Kong, Q. P., Wang, C. Y., Zhu, C. L., & Zhang, Y. P. (2004). Different Matrilineal Contributions to Genetic Structure of Ethnic Groups in the Silk Road Region in China. Molecular Biology and Evolution, 21, 2265-2280. http://dx.doi.org/10.1093/molbev/msh238
Yao, Y.-G., Kong, Q.-P., Bandelt, H.-J., Kivisild, T. et al. (2002). Phylogeographic Differentiation of Mitochondrial DNA in Han Chinese. The American Journal of Human Genetics, 70, 635-651. http://dx.doi.org/10.1086/338999
Zgonjanin, D., Veselinovic, I., Kubat, M., Furac, I., Antov, M., Loncar, E., & Omorjan, R. (2010). Sequence Polymorphism of the Mitochondrial DNA Control Region in the Population of Vojvodina Province, Serbia. Legal medicine (Tokyo, Japan), 12, 104-107. http://dx.doi.org/10.1016/j.legalmed.2009.10.007
Zhao, Y. B., Sun, W. Y., Zhan, Y., Di, W., & Yu, C. C. (2011). Mitochondrial DNA Evidence of Southward Migration of Manchus in China. Molekuliarnaia Biologiia (Moskva), 45, 825-830. http://dx.doi.org/10.1134/s0026893311050153
Zheng, H.-X., Yan, S., Qin, Z.-D., & Jin, L. (2012). MtDNA Analysis of Global Populations Support That Major Population Expansions Began before Neolithic Time. Scientific Reports, 2, 745. http://dx.doi.org/10.1038/srep00745
Zimmermann, B., Brandstatter, A., Duftner, N., Niederwieser, D., Spiroski, M., Arsov, T., & Parson, W. (2007). Mitochondrial DNA Control Region Population Data from Macedonia. Forensic Science International: Genetics, 1, e4-e9. http://dx.doi.org/10.1016/j.fsigen.2007.03.002
References S2(b)
Abu-Amero, K. K., Gonzalez, A. M., Larruga, J. M., Bosley, T. M., & Cabrera, V. M. (2007). Eurasian and African Mitochondrial DNA Influences in the Saudi Arabian Population. BMC Evolutionary Biology, 7, 32. http://dx.doi.org/10.1186/1471-2148-7-32
Abu-Amero, K. K., Larruga, J. M., Cabrera, V. M., & Gonzalez, A. M. (2008). Mitochondrial DNA Structure in the Arabian Peninsula. BMC Evolutionary Biology, 8, 45. http://dx.doi.org/10.1186/1471-2148-8-45
Achilli, A., Olivieri, A., Pala, M., Metspalu, E., Fornarino, S., Battaglia, V. et al. (2007). Mitochondrial DNA Variation of Modern Tuscans Supports the Near Eastern Origin of Etruscans. American Journal of Human Genetics, 80, 759-768. http://dx.doi.org/10.1086/512822
Álvarez-Iglesias, V., Mosquera-Miguel, A., Cerezo, M., Quintáns, B., Zarrabeitia, M. T., Cuscó, I. et al. (2009). New Population and Phylogenetic Features of the Internal Variation within Mitochondrial DNA Macro-Haplogroup R0. PLoS ONE, 4, e5112. http://dx.doi.org/10.1371/journal.pone.0005112
Al-Zahery, N., Semino, O., Benuzzi, G., Magri, C., Passarino, G., Torroni, A., & Santachiara-Benerecetti, A. S. (2003). Y- Chromosome and mtDNA Polymorphisms in Iraq, a Crossroad of the Early Human Dispersal and of Post-Neolithic Migrations. Molecular Phylogenetics and Evolution, 28, 458-472. http://dx.doi.org/10.1016/S1055-7903(03)00039-3
Brandstatter, A., Klein, R., Duftner, N., Wiegand, P., & Parson, W. (2006). Application of a Quasi-Median Network Analysis for the Visualization of Character Conflicts to a Population Sample of Mitochondrial DNA Control Region Sequences from Southern Germany (Ulm). International Journal of Legal Medicine, 120, 310-314. http://dx.doi.org/10.1007/s00414-006-0114-x
Brandstatter, A., Niederstatter, H., Pavlic, M., Grubwieser, P., & Parson, W. (2007). Generating Population Data for the EMPOP Database―An Overview of the mtDNA Sequencing and Data Evaluation Processes Considering 273 Austrian Control Region Sequences as Example. Forensic Science International, 166, 164-175. http://dx.doi.org/10.1016/j.forsciint.2006.05.006
Cardoso, S., Alfonso-Sanchez, M. A., Valverde, L., Odriozola, A., Perez-Miranda, A. M., Pena, J. A., & de Pancorbo, M. M. (2011). The Maternal Legacy of Basques in Northern Navarre: New Insights into the Mitochondrial DNA Diversity of the Franco-Cantabrian Area. American Journal of Physical Anthropology, 145, 480-488. http://dx.doi.org/10.1002/ajpa.21532
Comas, D., Plaza, S., Wells, R. S., Yuldaseva, N., Lao, O., Calafell, F., & Bertranpetit, J. (2004). Admixture, Migrations, and Dispersals in Central Asia: Evidence from Maternal DNA Lineages. European Journal of Human Genetics, 12, 495- 504. http://dx.doi.org/10.1038/sj.ejhg.5201160
Derenko, M., Malyarchuk, B., Bahmanimehr, A., Denisova, G., Perkova, M., Farjadian, S., & Yepiskoposyan, L. (2013). Complete Mitochondrial DNA Diversity in Iranians. PLoS ONE, 8, e80673. http://dx.doi.org/10.1371/journal.pone.0080673
Derenko, M., Malyarchuk, B., Denisova, G., Perkova, M., Rogalla, U., Grzybowski, T. et al. (2012). Complete Mitochondrial DNA Analysis of Eastern Eurasian Haplogroups Rarely Found in Populations of Northern Asia and Eastern Europe. PLoS ONE, 7, e32179. http://dx.doi.org/10.1371/journal.pone.0032179
Derenko, M., Malyarchuk, B., Grzybowski, T., Denisova, G., Dambueva, I., Perkova, M. et al. (2007). Phylogeographic Analysis of Mitochondrial DNA in Northern Asian Populations. The American Journal of Human Genetics, 81, 1025- 1041. http://dx.doi.org/10.1086/522933
Derenko, M., Malyarchuk, B., Grzybowski, T., Denisova, G., Rogalla, U., Perkova, M. et al. (2010). Origin and Post-Glacial Dispersal of Mitochondrial DNA Haplogroups C and D in Northern Asia. PLoS ONE, 5, e15214. http://dx.doi.org/10.1371/journal.pone.0015214
Grosheva, A., Shneider, Y. V., Zhukova, O., Morozova, I. Y., & Rychkov, S. Y. (2014). Features of the Udmurt Mitochondrial Gene Pool in Relation to Tribal Structure. Russian Journal of Genetics, 50, 975-986. http://dx.doi.org/10.1134/S1022795414090063
Grzybowski, T., Malyarchuk, B. A., Derenko, M. V., Perkova, M. A., Bednarek, J., & Wozniak, M. (2007). Complex Interactions of the Eastern and Western Slavic Populations with Other European Groups as Revealed by Mitochondrial DNA Analysis. Forensic Science International: Genetics, 1, 141-147. http://dx.doi.org/10.1016/j.fsigen.2007.01.010
Gubina, M. A., Girgol’kau, L. A., Babenko, V. N., Damba, L. D., Maksimov, V. N., & Voevoda, M. I. (2013). Mitochondrial DNA Polymorphism in Populations of Aboriginal Residents of the Far East. Russian Journal of Genetics, 49, 751-764. http://dx.doi.org/10.1134/S1022795413070065
Hedman, M., Brandstätter, A., Pimenoff, V., Sistonen, P., Palo, J. U., Parson, W., & Sajantila, A. (2007). Finnish Mitochondrial DNA HVS-I and HVS-II Population Data. Forensic Science International, 172, 171-178. http://dx.doi.org/10.1016/j.forsciint.2006.09.012
Irwin, J. A., Ikramov, A., Saunier, J., Bodner, M., Amory, S., Rock, A. et al. (2010). The mtDNA Composition of Uzbekistan: A Microcosm of Central Asian Patterns. International Journal of Legal Medicine, 124, 195-204. http://dx.doi.org/10.1007/s00414-009-0406-z
Irwin, J., Egyed, B., Saunier, J., Szamosi, G., O’Callaghan, J., Padar, Z., & Parsons, T. (2007). Hungarian mtDNA Population Databases from Budapest and the Baranya County Roma. International Journal of Legal Medicine, 121, 377- 383. http://dx.doi.org/10.1007/s00414-006-0128-4
Irwin, J., Saunier, J., Strouss, K., Paintner, C., Diegoli, T., Sturk, K. et al. (2008). Mitochondrial Control Region Sequences from Northern Greece and Greek Cypriots. International Journal of Legal Medicine, 122, 87-89. http://dx.doi.org/10.1007/s00414-007-0173-7
Karachanak, S., Carossa, V., Nesheva, D., Olivieri, A., Pala, M., Hooshiar Kashani, B. et al. (2012). Bulgarians vs the Other European Populations: A Mitochondrial DNA Perspective. International Journal of Legal Medicine, 126, 497-503. http://dx.doi.org/10.1007/s00414-011-0589-y
Kushniarevich, A., Sivitskaya, L., Danilenko, N., Novogrodskii, T., Tsybovsky, I., Kiseleva, A. et al. (2013). Uniparental Genetic Heritage of Belarusians: Encounter of Rare Middle Eastern Matrilineages with a Central European Mitochondrial DNA Pool. PLoS ONE, 8, e66499. http://dx.doi.org/10.1371/journal.pone.0066499
Lehocký, I., Baldovič, M., Kádaši, Ľ., & Metspalu, E. (2008). A Database of Mitochondrial DNA Hypervariable Regions I and II Sequences of Individuals from Slovakia. Forensic Science International: Genetics, 2, e53-e59. http://dx.doi.org/10.1016/j.fsigen.2007.12.008
Liu, C., Wang, S.-Y., Zhao, M., Xu, Z.-Y., Hu, Y.-H., Chen, F. et al. (2011). Mitochondrial DNA Polymorphisms in Gelao Ethnic Group Residing in Southwest China. Forensic Science International: Genetics, 5, e4-e10. http://dx.doi.org/10.1016/j.fsigen.2010.04.007
Malyarchuk, B. A., Grzybowski, T., Derenko, M. V., Czarny, J., Drobnič, K., & Miścicka-Śliwka, D. (2003). Mitochondrial DNA Variability in Bosnians and Slovenians. Annals of Human Genetics, 67, 412-425. http://dx.doi.org/10.1046/j.1469-1809.2003.00042.x
Malyarchuk, B. A., Vanecek, T., Perkova, M. A., Derenko, M. V., & Sip, M. (2006). Mitochondrial DNA Variability in the Czech Population, with Application to the Ethnic History of Slavs. Human Biology, 78, 681-696. http://dx.doi.org/10.1353/hub.2007.0014
Malyarchuk, B., Derenko, M., Denisova, G., & Kravtsova, O. (2010). Mitogenomic Diversity in Tatars from the Volga-Ural Region of Russia. Molecular Biology and Evolution, 27, 2220-2226. http://dx.doi.org/10.1093/molbev/msq065
Mielnik-Sikorska, M., Daca, P., Malyarchuk, B., Derenko, M., Skonieczna, K., Perkova, M. et al. (2013). The History of Slavs Inferred from Complete Mitochondrial Genome Sequences. PLoS ONE, 8, e54360. http://dx.doi.org/10.1371/journal.pone.0054360
Mikkelsen, M., Sorensen, E., Rasmussen, E. M., & Morling, N. (2010). Mitochondrial DNA HV1 and HV2 Variation in Danes. Forensic Science International: Genetics, 4, e87-88. http://dx.doi.org/10.1016/j.fsigen.2009.07.007
Pimenoff, V. N., Comas, D., Palo, J. U., Vershubsky, G., Kozlov, A., & Sajantila, A. (2008). Northwest Siberian Khanty and Mansi in the Junction of West and East Eurasian Gene Pools as Revealed by Uniparental Markers. European Journal of Human Genetics, 16, 1254-1264. http://www.nature.com/ejhg/journal/v16/n10/suppinfo/ejhg2008101s1.html http://dx.doi.org/10.1038/ejhg.2008.101
Prieto, L., Zimmermann, B., Goios, A., Rodriguez-Monge, A., Paneto, G. G., Alves, C. et al. (2011). The GHEP-EMPOP Collaboration on mtDNA Population Data―A New Resource for Forensic Casework. Forensic Science International: Genetics, 5, 146-151. http://dx.doi.org/10.1016/j.fsigen.2010.10.013
Quintana-Murci, L., Chaix, R., Wells, R. S., Behar, D. M., Sayar, H., Scozzari, R. et al. (2004). Where West Meets East: The Complex mtDNA Landscape of the Southwest and Central Asian Corridor. American Journal of Human Genetics, 74, 827-845. http://dx.doi.org/10.1086/383236
Richard, C., Pennarun, E., Kivisild, T., Tambets, K., Tolk, H. V., Metspalu, E. et al. (2007). An mtDNA Perspective of French Genetic Variation. Annals of Human Biology, 34, 68-79. http://dx.doi.org/10.1080/03014460601076098
Simonson, T. S., Xing, J., Barrett, R., Jerah, E., Loa, P., Zhang, Y. et al. (2011). Ancestry of the Iban Is Predominantly Southeast Asian: Genetic Evidence from Autosomal, Mitochondrial, and Y Chromosomes. PLoS ONE, 6, e16338. http://dx.doi.org/10.1371/journal.pone.0016338
Turchi, C., Buscemi, L., Previdere, C., Grignani, P., Brandstatter, A., Achilli, A. et al. (2008). Italian Mitochondrial DNA Database: Results of a Collaborative Exercise and Proficiency Testing. International Journal of Legal Medicine, 122, 199-204. http://dx.doi.org/10.1007/s00414-007-0207-1
Volodko, N. V., Starikovskaya, E. B., Mazunin, I. O., Eltsov, N. P., Naidenko, P. V., Wallace, D. C., & Sukernik, R. I. (2008). Mitochondrial Genome Diversity in Arctic Siberians, with Particular Reference to the Evolutionary History of Beringia and Pleistocenic Peopling of the Americas. The American Journal of Human Genetics, 82, 1084-1100. http://dx.doi.org/10.1016/j.ajhg.2008.03.019
Yao, Y. G., Kong, Q. P., Wang, C. Y., Zhu, C. L., & Zhang, Y. P. (2004). Different Matrilineal Contributions to Genetic Structure of Ethnic Groups in the Silk Road Region in China. Molecular Biology and Evolution, 21, 2265-2280. http://dx.doi.org/10.1093/molbev/msh238
Yao, Y.-G., Kong, Q.-P., Bandelt, H.-J., Kivisild, T. et al. (2002). Phylogeographic Differentiation of Mitochondrial DNA in Han Chinese. The American Journal of Human Genetics, 70, 635-651. http://dx.doi.org/10.1086/338999
Zgonjanin, D., Veselinovic, I., Kubat, M., Furac, I., Antov, M., Loncar, E., & Omorjan, R. (2010). Sequence Polymorphism of the Mitochondrial DNA Control Region in the Population of Vojvodina Province, Serbia. Legal medicine (Tokyo, Japan), 12, 104-107. http://dx.doi.org/10.1016/j.legalmed.2009.10.007
Zhao, Y. B., Sun, W. Y., Zhan, Y., Di, W., & Yu, C. C. (2011). Mitochondrial DNA Evidence of Southward Migration of Manchus in China. Molekuliarnaia Biologiia (Moskva), 45, 825-830. http://dx.doi.org/10.1134/s0026893311050153
Zheng, H.-X., Yan, S., Qin, Z.-D., & Jin, L. (2012). MtDNA Analysis of Global Populations Support That Major Population Expansions Began before Neolithic Time. Scientific Reports, 2, 745. http://dx.doi.org/10.1038/srep00745
Zimmermann, B., Brandstatter, A., Duftner, N., Niederwieser, D., Spiroski, M., Arsov, T., & Parson, W. (2007). Mitochondrial DNA Control Region Population Data from Macedonia. Forensic Science International: Genetics, 1, e4-e9. http://dx.doi.org/10.1016/j.fsigen.2007.03.002
References S2(c)
Abu-Amero, K. K., Gonzalez, A. M., Larruga, J. M., Bosley, T. M., & Cabrera, V. M. (2007). Eurasian and African Mitochondrial DNA Influences in the Saudi Arabian Population. BMC Evolutionary Biology, 7, 32. http://dx.doi.org/10.1186/1471-2148-7-32
Abu-Amero, K. K., Larruga, J. M., Cabrera, V. M., & Gonzalez, A. M. (2008). Mitochondrial DNA Structure in the Arabian Peninsula. BMC Evolutionary Biology, 8, 45. http://dx.doi.org/10.1186/1471-2148-8-45
Achilli, A., Olivieri, A., Pala, M., Metspalu, E., Fornarino, S., Battaglia, V. et al. (2007). Mitochondrial DNA Variation of Modern Tuscans Supports the Near Eastern Origin of Etruscans. American Journal of Human Genetics, 80, 759-768. http://dx.doi.org/10.1086/512822
Álvarez-Iglesias, V., Mosquera-Miguel, A., Cerezo, M., Quintáns, B., Zarrabeitia, M. T., Cuscó, I. et al. (2009). New Population and Phylogenetic Features of the Internal Variation within Mitochondrial DNA Macro-Haplogroup R0. PLoS ONE, 4, e5112. http://dx.doi.org/10.1371/journal.pone.0005112
Al-Zahery, N., Semino, O., Benuzzi, G., Magri, C., Passarino, G., Torroni, A., & Santachiara-Benerecetti, A. S. (2003). Y-Chromosome and mtDNA Polymorphisms in Iraq, a Crossroad of the Early Human Dispersal and of Post-Neolithic Migrations. Molecular Phylogenetics and Evolution, 28, 458-472. http://dx.doi.org/10.1016/S1055-7903(03)00039-3
Brandstatter, A., Klein, R., Duftner, N., Wiegand, P., & Parson, W. (2006). Application of a Quasi-Median Network Analysis for the Visualization of Character Conflicts to a Population Sample of Mitochondrial DNA Control Region Sequences from Southern Germany (Ulm). International Journal of Legal Medicine, 120, 310-314. http://dx.doi.org/10.1007/s00414-006-0114-x
Brandstatter, A., Niederstatter, H., Pavlic, M., Grubwieser, P., & Parson, W. (2007). Generating Population Data for the EMPOP Database―An Overview of the mtDNA Sequencing and Data Evaluation Processes Considering 273 Austrian Control Region Sequences as Example. Forensic Science International, 166, 164-175. http://dx.doi.org/10.1016/j.forsciint.2006.05.006
Cardoso, S., Alfonso-Sanchez, M. A., Valverde, L., Odriozola, A., Perez-Miranda, A. M., Pena, J. A., & de Pancorbo, M. M. (2011). The Maternal Legacy of Basques in Northern Navarre: New Insights into the Mitochondrial DNA Diversity of the Franco-Cantabrian Area. American Journal of Physical Anthropology, 145, 480-488. http://dx.doi.org/10.1002/ajpa.21532
Comas, D., Plaza, S., Wells, R. S., Yuldaseva, N., Lao, O., Calafell, F., & Bertranpetit, J. (2004). Admixture, Migrations, and Dispersals in Central Asia: Evidence from Maternal DNA Lineages. European Journal of Human Genetics, 12, 495- 504. http://dx.doi.org/10.1038/sj.ejhg.5201160
Derenko, M., Malyarchuk, B., Bahmanimehr, A., Denisova, G., Perkova, M., Farjadian, S., & Yepiskoposyan, L. (2013). Complete Mitochondrial DNA Diversity in Iranians. PLoS ONE, 8, e80673. http://dx.doi.org/10.1371/journal.pone.0080673
Derenko, M., Malyarchuk, B., Denisova, G., Perkova, M., Rogalla, U., Grzybowski, T. et al. (2012). Complete Mitochondrial DNA Analysis of Eastern Eurasian Haplogroups Rarely Found in Populations of Northern Asia and Eastern Europe. PLoS ONE, 7, e32179. http://dx.doi.org/10.1371/journal.pone.0032179
Derenko, M., Malyarchuk, B., Grzybowski, T., Denisova, G., Dambueva, I., Perkova, M. et al. (2007). Phylogeographic Analysis of Mitochondrial DNA in Northern Asian Populations. The American Journal of Human Genetics, 81, 1025- 1041. http://dx.doi.org/10.1086/522933
Derenko, M., Malyarchuk, B., Grzybowski, T., Denisova, G., Rogalla, U., Perkova, M. et al. (2010). Origin and Post-Glacial Dispersal of Mitochondrial DNA Haplogroups C and D in Northern Asia. PLoS ONE, 5, e15214. http://dx.doi.org/10.1371/journal.pone.0015214
Grosheva, A., Shneider, Y. V., Zhukova, O., Morozova, I. Y., & Rychkov, S. Y. (2014). Features of the Udmurt Mitochondrial Gene Pool in Relation to Tribal Structure. Russian Journal of Genetics, 50, 975-986. http://dx.doi.org/10.1134/S1022795414090063
Grzybowski, T., Malyarchuk, B. A., Derenko, M. V., Perkova, M. A., Bednarek, J., & Wozniak, M. (2007). Complex Interactions of the Eastern and Western Slavic Populations with Other European Groups as Revealed by Mitochondrial DNA Analysis. Forensic Science International: Genetics, 1, 141-147. http://dx.doi.org/10.1016/j.fsigen.2007.01.010
Gubina, M. A., Girgol’kau, L. A., Babenko, V. N., Damba, L. D., Maksimov, V. N., & Voevoda, M. I. (2013). Mitochondrial DNA Polymorphism in Populations of Aboriginal Residents of the Far East. Russian Journal of Genetics, 49, 751-764. http://dx.doi.org/10.1134/S1022795413070065
Hedman, M., Brandstätter, A., Pimenoff, V., Sistonen, P., Palo, J. U., Parson, W., & Sajantila, A. (2007). Finnish Mitochondrial DNA HVS-I and HVS-II Population Data. Forensic Science International, 172, 171-178. http://dx.doi.org/10.1016/j.forsciint.2006.09.012
Irwin, J. A., Ikramov, A., Saunier, J., Bodner, M., Amory, S., Rock, A. et al. (2010). The mtDNA Composition of Uzbekistan: A Microcosm of Central Asian Patterns. International Journal of Legal Medicine, 124, 195-204. http://dx.doi.org/10.1007/s00414-009-0406-z
Irwin, J., Egyed, B., Saunier, J., Szamosi, G., O’Callaghan, J., Padar, Z., & Parsons, T. (2007). Hungarian mtDNA Population Databases from Budapest and the Baranya County Roma. International Journal of Legal Medicine, 121, 377- 383. http://dx.doi.org/10.1007/s00414-006-0128-4
Irwin, J., Saunier, J., Strouss, K., Paintner, C., Diegoli, T., Sturk, K. et al. (2008). Mitochondrial Control Region Sequences from Northern Greece and Greek Cypriots. International Journal of Legal Medicine, 122, 87-89. http://dx.doi.org/10.1007/s00414-007-0173-7
Karachanak, S., Carossa, V., Nesheva, D., Olivieri, A., Pala, M., Hooshiar Kashani, B. et al. (2012). Bulgarians vs the Other European Populations: A Mitochondrial DNA Perspective. International Journal of Legal Medicine, 126, 497-503. http://dx.doi.org/10.1007/s00414-011-0589-y
Kushniarevich, A., Sivitskaya, L., Danilenko, N., Novogrodskii, T., Tsybovsky, I., Kiseleva, A. et al. (2013). Uniparental Genetic Heritage of Belarusians: Encounter of Rare Middle Eastern Matrilineages with a Central European Mitochondrial DNA Pool. PLoS ONE, 8, e66499. http://dx.doi.org/10.1371/journal.pone.0066499
Lehocký, I., Baldovič, M., Kádaši, Ľ., & Metspalu, E. (2008). A Database of Mitochondrial DNA Hypervariable Regions I and II Sequences of Individuals from Slovakia. Forensic Science International: Genetics, 2, e53-e59. http://dx.doi.org/10.1016/j.fsigen.2007.12.008
Liu, C., Wang, S.-Y., Zhao, M., Xu, Z.-Y., Hu, Y.-H., Chen, F. et al. (2011). Mitochondrial DNA Polymorphisms in Gelao Ethnic Group Residing in Southwest China. Forensic Science International: Genetics, 5, e4-e10. http://dx.doi.org/10.1016/j.fsigen.2010.04.007
Malyarchuk, B. A., Grzybowski, T., Derenko, M. V., Czarny, J., Drobnič, K., & Miścicka-Śliwka, D. (2003). Mitochondrial DNA Variability in Bosnians and Slovenians. Annals of Human Genetics, 67, 412-425. http://dx.doi.org/10.1046/j.1469-1809.2003.00042.x
Malyarchuk, B. A., Vanecek, T., Perkova, M. A., Derenko, M. V., & Sip, M. (2006). Mitochondrial DNA Variability in the Czech Population, with Application to the Ethnic History of Slavs. Human Biology, 78, 681-696. http://dx.doi.org/10.1353/hub.2007.0014
Malyarchuk, B., Derenko, M., Denisova, G., & Kravtsova, O. (2010). Mitogenomic Diversity in Tatars from the Volga-Ural Region of Russia. Molecular Biology and Evolution, 27, 2220-2226. http://dx.doi.org/10.1093/molbev/msq065
Mielnik-Sikorska, M., Daca, P., Malyarchuk, B., Derenko, M., Skonieczna, K., Perkova, M. et al. (2013). The History of Slavs Inferred from Complete Mitochondrial Genome Sequences. PLoS ONE, 8, e54360. http://dx.doi.org/10.1371/journal.pone.0054360
Mikkelsen, M., Sorensen, E., Rasmussen, E. M., & Morling, N. (2010). Mitochondrial DNA HV1 and HV2 Variation in Danes. Forensic Science International: Genetics, 4, e87-88. http://dx.doi.org/10.1016/j.fsigen.2009.07.007
Pimenoff, V. N., Comas, D., Palo, J. U., Vershubsky, G., Kozlov, A., & Sajantila, A. (2008). Northwest Siberian Khanty and Mansi in the junction of West and East Eurasian Gene Pools as Revealed by Uniparental Markers. European Journal of Human Genetics, 16, 1254-1264. http://www.nature.com/ejhg/journal/v16/n10/suppinfo/ejhg2008101s1.html http://dx.doi.org/10.1038/ejhg.2008.101
Prieto, L., Zimmermann, B., Goios, A., Rodriguez-Monge, A., Paneto, G. G., Alves, C. et al. (2011). The GHEP-EMPOP Collaboration on mtDNA Population Data―A New Resource for Forensic Casework. Forensic Science International: Genetics, 5, 146-151. http://dx.doi.org/10.1016/j.fsigen.2010.10.013
Quintana-Murci, L., Chaix, R., Wells, R. S., Behar, D. M., Sayar, H., Scozzari, R. et al. (2004). Where West Meets East: The Complex mtDNA Landscape of the Southwest and Central Asian Corridor. American Journal of Human Genetics, 74, 827-845. http://dx.doi.org/10.1086/383236
Richard, C., Pennarun, E., Kivisild, T., Tambets, K., Tolk, H. V., Metspalu, E. et al. (2007). An mtDNA Perspective of French Genetic Variation. Annals of Human Biology, 34, 68-79. http://dx.doi.org/10.1080/03014460601076098
Simonson, T. S., Xing, J., Barrett, R., Jerah, E., Loa, P., Zhang, Y. et al. (2011). Ancestry of the Iban Is Predominantly Southeast Asian: Genetic Evidence from Autosomal, Mitochondrial, and Y Chromosomes. PLoS ONE, 6, e16338. http://dx.doi.org/10.1371/journal.pone.0016338
Turchi, C., Buscemi, L., Previdere, C., Grignani, P., Brandstatter, A., Achilli, A. et al. (2008). Italian Mitochondrial DNA Database: Results of a Collaborative Exercise and Proficiency Testing. International Journal of Legal Medicine, 122, 199- 204. http://dx.doi.org/10.1007/s00414-007-0207-1
Volodko, N. V., Starikovskaya, E. B., Mazunin, I. O., Eltsov, N. P., Naidenko, P. V., Wallace, D. C., & Sukernik, R. I. (2008). Mitochondrial Genome Diversity in Arctic Siberians, with Particular Reference to the Evolutionary History of Beringia and Pleistocenic Peopling of the Americas. The American Journal of Human Genetics, 82, 1084-1100. http://dx.doi.org/10.1016/j.ajhg.2008.03.019
Yao, Y. G., Kong, Q. P., Wang, C. Y., Zhu, C. L., & Zhang, Y. P. (2004). Different Matrilineal Contributions to Genetic Structure of Ethnic Groups in the Silk Road Region in China. Molecular Biology and Evolution, 21, 2265-2280. http://dx.doi.org/10.1093/molbev/msh238
Yao, Y.-G., Kong, Q.-P., Bandelt, H.-J., Kivisild, T. et al. (2002). Phylogeographic Differentiation of Mitochondrial DNA in Han Chinese. The American Journal of Human Genetics, 70, 635-651. http://dx.doi.org/10.1086/338999
Zgonjanin, D., Veselinovic, I., Kubat, M., Furac, I., Antov, M., Loncar, E., & Omorjan, R. (2010). Sequence Polymorphism of the Mitochondrial DNA Control Region in the Population of Vojvodina Province, Serbia. Legal medicine (Tokyo, Japan), 12, 104-107. http://dx.doi.org/10.1016/j.legalmed.2009.10.007
Zhao, Y. B., Sun, W. Y., Zhan, Y., Di, W., & Yu, C. C. (2011). Mitochondrial DNA Evidence of Southward Migration of Manchus in China. Molekuliarnaia Biologiia (Moskva), 45, 825-830. http://dx.doi.org/10.1134/s0026893311050153
Zheng, H.-X., Yan, S., Qin, Z.-D., & Jin, L. (2012). MtDNA Analysis of Global Populations Support That Major Population Expansions Began before Neolithic Time. Scientific Reports, 2, 745. http://dx.doi.org/10.1038/srep00745
Zimmermann, B., Brandstatter, A., Duftner, N., Niederwieser, D., Spiroski, M., Arsov, T., & Parson, W. (2007). Mitochondrial DNA Control Region Population Data from Macedonia. Forensic Science International: Genetics, 1, e4-e9. http://dx.doi.org/10.1016/j.fsigen.2007.03.002
Table S3. Absolute frequencies of Y-chromosome haplogroups C, N and Q in the population groups used in the chi-square test comparison. N-number of analyzed individuals; *p-value population group vs Bulgarians.
References S3
Abu-Amero, Alonso, S., Flores, C., Cabrera, V., Alonso, A., Martin, P., Albarran, C. et al. (2005). The Place of the Basques in the European Y-Chromosome Diversity Landscape. European Journal of Human Genetics, 13, 1293-1302. http://dx.doi.org/10.1038/sj.ejhg.5201482
Battaglia, V., Fornarino, S., Al-Zahery, N., Olivieri, A., Pala, M., Myres, N. M. et al. (2009). Y-Chromosomal Evidence of the Cultural Diffusion of Agriculture in Southeast Europe. European Journal of Human Genetics, 17, 820-830. http://dx.doi.org/10.1038/ejhg.2008.249
Capelli, C., Redhead, N., Abernethy, J. K., Gratrix, F., Wilson, J. F., Moen, T. et al. (2003). A Y Chromosome Census of the British Isles. Current Biology, 13, 979-984. http://dx.doi.org/10.1016/s0960-9822(03)00373-7
Di Cristofaro, J., Pennarun, E., Mazières, S., Myres, N. M., Lin, A. A., Temori, S. A. et al. (2013). Afghan Hindu Kush: Where Eurasian Sub-Continent Gene Flows Converge. PLoS ONE, 8, e76748. http://dx.doi.org/10.1371/journal.pone.0076748
Dulik, M. C., Osipova, L. P., & Schurr, T. G. (2011). Y-Chromosome Variation in Altaian Kazakhs Reveals a Common Paternal Gene Pool for Kazakhs and the Influence of Mongolian Expansions. PLoS ONE, 6, e17548. http://dx.doi.org/10.1371/journal.pone.0017548
Dulik, M. C., Zhadanov, S. I., Osipova, L. P., Askapuli, A., Gau, L., Gokcumen, O. et al. (2012). Mitochondrial DNA and Y Chromosome Variation Provides Evidence for a Recent Common Ancestry between Native Americans and Indigenous Altaians. American Journal of Human Genetics, 90, 229-246. http://dx.doi.org/10.1016/j.ajhg.2011.12.014
Fedorova, S. A., Reidla, M., Metspalu, E., Metspalu, M., Rootsi, S., Tambets, K. et al. (2013). Autosomal and Uniparental Portraits of the Native Populations of Sakha (Yakutia): Implications for the Peopling of Northeast Eurasia. BMC Evolutionary Biology, 13, 127. http://dx.doi.org/10.1186/1471-2148-13-127
Francalacci, P., Morelli, L., Underhill, P. A., Lillie, A. S., Passarino, G., Useli, A. et al. (2003). Peopling of Three Mediterranean Islands (Corsica, Sardinia, and Sicily) Inferred by Y-Chromosome Biallelic Variability. American Journal of Physical Anthropology, 121, 270-279. http://dx.doi.org/10.1002/ajpa.10265
Gonçalves, R., Freitas, A., Branco, M., Rosa, A., Fernandes, A. T., Zhivotovsky, L. A. et al. (2005). Y-Chromosome Lineages from Portugal, Madeira and Açores Record Elements of Sephardim and Berber Ancestry. Annals of Human Genetics, 69, 443-454. http://dx.doi.org/10.1111/j.1529-8817.2005.00161.x
Gubina, M. A., Damba, L. D., Babenko, V. N., Romaschenko, A. G., & Voevoda, M. I. (2013). Haplotype Diversity in mtDNA and Y-Chromosome in Populations of Altai-Sayan Region. Russian Journal of Genetics, 49, 329-343. http://dx.doi.org/10.1134/S1022795412120034
Hammer, M. F., Karafet, T. M., Park, H., Omoto, K., Harihara, S., Stoneking, M., & Horai, S. (2005). Dual Origins of the Japanese: Common Ground for Hunter-Gatherer and Farmer Y Chromosomes. Journal of Human Genetics, 51, 47-58. http://dx.doi.org/10.1007/s10038-005-0322-0
Karachanak, S., Grugni, V., Fornarino, S., Nesheva, D., Al-Zahery, N., Battaglia, V. et al. (2013). Y-Chromosome Diversity in Modern Bulgarians: New Clues about Their Ancestry. PLoS ONE, 8, e56779. http://dx.doi.org/10.1371/journal.pone.0056779
Kharkov, V. N., Khamina, K. V., Medvedeva, O. F., Simonova, K. V., Eremina, E. R., & Stepanov, V. A. (2014). Gene Pool of Buryats: Clinal Variability and Territorial Subdivision Based on Data of Y-Chromosome Markers. Russian Journal of Genetics, 50, 180-190. http://dx.doi.org/10.1134/S1022795413110082
Kushniarevich, A., Sivitskaya, L., Danilenko, N., Novogrodskii, T., Tsybovsky, I., Kiseleva, A. et al. (2013). Uniparental Genetic Heritage of Belarusians: Encounter of Rare Middle Eastern Matrilineages with a Central European Mitochondrial DNA Pool. PLoS ONE, 8, e66499. http://dx.doi.org/10.1371/journal.pone.0066499
Lacau, H., Gayden, T., Regueiro, M., Chennakrishnaiah, S., Bukhari, A., Underhill, P. A. et al. (2012). Afghanistan from a Y-Chromosome Perspective. European Journal of Human Genetics, 20, 1063-1070. http://dx.doi.org/10.1038/ejhg.2012.59
López-Parra, A., Gusmao, L., Tavares, L., Baeza, C., Amorim, A., Mesa, M. et al. (2009). In search of the Pre- and Post-Neolithic Genetic Substrates in Iberia: Evidence from Y-Chromosome in Pyrenean Populations. Annals of Human Genetics, 73, 42-53. http://dx.doi.org/10.1111/j.1469-1809.2008.00478.x
Martinez-Cruz, B., Ioana, M., Calafell, F., Arauna, L. R., Sanz, P., Ionescu, R. et al. (2012). Y-Chromosome Analysis in Individuals Bearing the Basarab Name of the First Dynasty of Wallachian Kings. PLoS ONE, 7, e41803. http://dx.doi.org/10.1371/journal.pone.0041803
Noveski, P., Trivodalieva, S., Efremov, G., & Plaseska-Karanfilska, D. (2010). Y Chromosome Single Nucleotide Polymorphisms Typing by SNaPshot Minisequencing. Balkan Journal of Medical Genetics, 13, 9-16. http://dx.doi.org/10.2478/v10034-010-0013-9
Regueiro, M., Rivera, L., Damnjanovic, T., Lukovic, L., Milasin, J., & Herrera, R. J. (2012). High Levels of Paleolithic Y-Chromosome Lineages Characterize Serbia. Gene, 498, 59-67. http://dx.doi.org/10.1016/j.gene.2012.01.030
Semino, O., Passarino, G., Oefner, P. J., Lin, A. A., Arbuzova, S., Beckman, L. E. et al. (2000). The Genetic Legacy of Paleolithic Homo Sapiens Sapiens in Extant Europeans: A Y Chromosome Perspective. Science, 290, 1155-1159. http://dx.doi.org/10.1126/science.290.5494.1155
Table S4. Absolute frequencies of mtDNA haplogroups C and D in the population groups used in the chi-square test comparison. N-number of analyzed individuals; *p-value population group vs Bulgarians
References S4
Abu-Amero, Achilli, A., Olivieri, A., Pala, M., Metspalu, E., Fornarino, S., Battaglia, V. et al. (2007). Mitochondrial DNA Variation of Modern Tuscans Supports the Near Eastern Origin of Etruscans. American Journal of Human Genetics, 80, 759-768. http://dx.doi.org/10.1086/512822
Álvarez-Iglesias, V., Mosquera-Miguel, A., Cerezo, M., Quintáns, B., Zarrabeitia, M. T., Cuscó, I. et al. (2009). New Population and Phylogenetic Features of the Internal Variation within Mitochondrial DNA Macro-Haplogroup R0. PLoS ONE, 4, e5112. http://dx.doi.org/10.1371/journal.pone.0005112
Brandstatter, A., Klein, R., Duftner, N., Wiegand, P., & Parson, W. (2006). Application of a Quasi-Median Network Analysis for the Visualization of Character Conflicts to a Population Sample of Mitochondrial DNA Control Region Sequences from Southern Germany (Ulm). International Journal of Legal Medicine, 120, 310-314. http://dx.doi.org/10.1007/s00414-006-0114-x
Cardoso, S., Alfonso-Sanchez, M. A., Valverde, L., Odriozola, A., Perez-Miranda, A. M., Pena, J. A., & de Pancorbo, M. M. (2011). The Maternal Legacy of Basques in Northern Navarre: New Insights into the Mitochondrial DNA Diversity of the Franco-Cantabrian Area. American Journal of Physical Anthropology, 145, 480-488. http://dx.doi.org/10.1002/ajpa.21532
Comas, D., Plaza, S., Wells, R. S., Yuldaseva, N., Lao, O., Calafell, F., & Bertranpetit, J. (2004). Admixture, Migrations, and Dispersals in Central Asia: Evidence from Maternal DNA Lineages. European Journal of Human Genetics, 12, 495- 504. http://dx.doi.org/10.1038/sj.ejhg.5201160
Davidovic, S., Malyarchuk, B., Aleksic, J. M., Derenko, M., Topalovic, V., Litvinov, A. et al. (2015). Mitochondrial DNA Perspective of Serbian Genetic Diversity. American Journal of Physical Anthropology, 156, 449-465. http://dx.doi.org/10.1002/ajpa.22670
Derenko, M., Malyarchuk, B., Grzybowski, T., Denisova, G., Rogalla, U., Perkova, M. et al. (2010). Origin and Post-Glacial Dispersal of Mitochondrial DNA Haplogroups C and D in Northern Asia. PLoS ONE, 5, e15214. http://dx.doi.org/10.1371/journal.pone.0015214
Dulik, M. C., Zhadanov, S. I., Osipova, L. P., Askapuli, A., Gau, L., Gokcumen, O. et al. (2012). Mitochondrial DNA and Y Chromosome Variation Provides Evidence for a Recent Common Ancestry between Native Americans and Indigenous Altaians. American Journal of Human Genetics, 90, 229-246. http://dx.doi.org/10.1016/j.ajhg.2011.12.014
Grzybowski, T., Malyarchuk, B. A., Derenko, M. V., Perkova, M. A., Bednarek, J., & Wozniak, M. (2007). Complex Interactions of the Eastern and Western Slavic Populations with Other European Groups as Revealed by Mitochondrial DNA Analysis. Forensic Science International: Genetics, 1, 141-147. http://dx.doi.org/10.1016/j.fsigen.2007.01.010
Gubina, M. A., Girgol’kau, L. A., Babenko, V. N., Damba, L. D., Maksimov, V. N., & Voevoda, M. I. (2013). Mitochondrial DNA Polymorphism in Populations of Aboriginal Residents of the Far East. Russian Journal of Genetics, 49, 751-764. http://dx.doi.org/10.1134/S1022795413070065
Hedman, M., Brandstätter, A., Pimenoff, V., Sistonen, P., Palo, J. U., Parson, W., & Sajantila, A. (2007). Finnish Mitochondrial DNA HVS-I and HVS-II Population Data. Forensic Science International, 172, 171-178. http://dx.doi.org/10.1016/j.forsciint.2006.09.012
Irwin, J., Egyed, B., Saunier, J., Szamosi, G., O’Callaghan, J., Padar, Z., & Parsons, T. (2007). Hungarian mtDNA Population Databases from Budapest and the Baranya County Roma. International Journal of Legal Medicine, 121, 377-383. http://dx.doi.org/10.1007/s00414-006-0128-4
Irwin, J. A., Ikramov, A., Saunier, J., Bodner, M., Amory, S., Rock, A. et al. (2010). The mtDNA Composition of Uzbekistan: A Microcosm of Central Asian Patterns. International Journal of Legal Medicine, 124, 195-204. http://dx.doi.org/10.1007/s00414-009-0406-z
Irwin, J., Saunier, J., Strouss, K., Paintner, C., Diegoli, T., Sturk, K. et al. (2008). Mitochondrial Control Region Sequences from Northern Greece and Greek Cypriots. International Journal of Legal Medicine, 122, 87-89. http://dx.doi.org/10.1007/s00414-007-0173-7
Karachanak, S., Carossa, V., Nesheva, D., Olivieri, A., Pala, M., Hooshiar Kashani, B. et al. (2012). Bulgarians vs the Other European Populations: A Mitochondrial DNA Perspective. International Journal of Legal Medicine, 126, 497-503. http://dx.doi.org/10.1007/s00414-011-0589-y
Kushniarevich, A., Sivitskaya, L., Danilenko, N., Novogrodskii, T., Tsybovsky, I., Kiseleva, A. et al. (2013). Uniparental Genetic Heritage of Belarusians: Encounter of Rare Middle Eastern Matrilineages with a Central European Mitochondrial DNA Pool. PLoS ONE, 8, e66499. http://dx.doi.org/10.1371/journal.pone.0066499
Lehocký, I., Baldovič, M., Kádaši, Ľ., & Metspalu, E. (2008). A Database of Mitochondrial DNA Hypervariable Regions I and II Sequences of Individuals from Slovakia. Forensic Science International: Genetics, 2, e53-e59. http://dx.doi.org/10.1016/j.fsigen.2007.12.008
Malyarchuk, B. A., Grzybowski, T., Derenko, M. V., Czarny, J., Drobnič, K., & Miścicka-Śliwka, D. (2003). Mitochondrial DNA Variability in Bosnians and Slovenians. Annals of Human Genetics, 67, 412-425. http://dx.doi.org/10.1046/j.1469-1809.2003.00042.x
Malyarchuk, B. A., Vanecek, T., Perkova, M. A., Derenko, M. V., & Sip, M. (2006). Mitochondrial DNA Variability in the Czech Population, with Application to the Ethnic History of Slavs. Human Biology, 78, 681-696. http://dx.doi.org/10.1353/hub.2007.0014
Mielnik-Sikorska, M., Daca, P., Malyarchuk, B., Derenko, M., Skonieczna, K., Perkova, M. et al. (2013). The History of Slavs Inferred from Complete Mitochondrial Genome Sequences. PLoS ONE, 8, e54360. http://dx.doi.org/10.1371/journal.pone.0054360
Mikkelsen, M., Sorensen, E., Rasmussen, E. M., & Morling, N. (2010). Mitochondrial DNA HV1 and HV2 Variation in Danes. Forensic Science International: Genetics, 4, e87-88. http://dx.doi.org/10.1016/j.fsigen.2009.07.007
Prieto, L., Zimmermann, B., Goios, A., Rodriguez-Monge, A., Paneto, G. G., Alves, C. et al. (2011). The GHEP-EMPOP Collaboration on mtDNA Population Data―A New Resource for Forensic Casework. Forensic Science International: Genetics, 5, 146-151. http://dx.doi.org/10.1016/j.fsigen.2010.10.013
Turchi, C., Buscemi, L., Previdere, C., Grignani, P., Brandstatter, A., Achilli, A. et al. (2008). Italian Mitochondrial DNA Database: Results of a Collaborative Exercise and Proficiency Testing. International Journal of Legal Medicine, 122, 199- 204. http://dx.doi.org/10.1007/s00414-007-0207-1
Yao, Y. G., Kong, Q. P., Wang, C. Y., Zhu, C. L., & Zhang, Y. P. (2004). Different Matrilineal Contributions to Genetic Structure of Ethnic Groups in the Silk Road Region in China. Molecular Biology and Evolution, 21, 2265-2280. http://dx.doi.org/10.1093/molbev/msh238
Zgonjanin, D., Veselinovic, I., Kubat, M., Furac, I., Antov, M., Loncar, E., & Omorjan, R. (2010). Sequence Polymorphism of the Mitochondrial DNA Control Region in the Population of Vojvodina Province, Serbia. Legal Medicine (Tokyo, Japan), 12, 104-107. http://dx.doi.org/10.1016/j.legalmed.2009.10.007
Zheng, H.-X., Yan, S., Qin, Z.-D., & Jin, L. (2012). MtDNA Analysis of Global Populations Support That Major Population Expansions Began before Neolithic Time. Scientific Reports, 2, 745. http://dx.doi.org/10.1038/srep00745
Zimmermann, B., Brandstatter, A., Duftner, N., Niederwieser, D., Spiroski, M., Arsov, T., & Parson, W. (2007). Mitochondrial DNA Control Region Population Data from Macedonia. Forensic Science International: Genetics, 1, e4-e9. http://dx.doi.org/10.1016/j.fsigen.2007.03.002
NOTES
*These authors have equally contributed to this work.
#Corresponding author.