American Journal of Plant Sciences
Vol.08 No.08(2017), Article ID:77374,16 pages
10.4236/ajps.2017.88121
Emergence of Plastidial Intergenic Spacers as Suitable DNA Barcodes for Arid Medicinal Plant Rhazya stricta
Samia A. Khan1, Mohammed N. Baeshen2, Hassan A. Ramadan1,3,4, Nabih A. Baeshen1
1Department of Biological Sciences, Faculty of Science, King Abdulaziz University, Jeddah, KSA
2Department of Biological Sciences, Faculty of Science, University of Jeddah, Jeddah, KSA
3Department of Marine Biology, Faculty of Marine Science, King Abdulaziz University, Jeddah, KSA
4Department of Cell Biology, National Research Center, Cairo, Egypt

Copyright © 2017 by authors and Scientific Research Publishing Inc.
This work is licensed under the Creative Commons Attribution International License (CC BY 4.0).
http://creativecommons.org/licenses/by/4.0/



Received: May 18, 2017; Accepted: June 30, 2017; Published: July 3, 2017
ABSTRACT
The desert plant Rhazya stricta has anticancer and antimicrobial properties, and is widely used in indigenous medicines of Saudi Arabia. However, the therapeutic benefits rely on an accurate identification of this species. The authenticity of R. stricta and other medicinal plants and herbs procured from local markets can be questionable due to a lack of clear phenotypic traits. DNA barcoding is an emerging technology for rapid and accurate species identification. In this study, six candidate chloroplastid barcodes were investigated for the authentication of R. stricta. We compared the DNA sequences from fifty locally collected and five market samples of R. stricta with database sequences of R. stricta and seven closely related species. We found that the coding regions matK, rbcL, rpoB, and rpoC1 were highly similar among the taxa. By contrast, the intergenic spacers psbK-psbI and atpF-atpH were variable loci distinct for the medicinal plant R. stricta. psbK-psbI clearly discriminated R. stricta samples as an efficient single locus marker, whereas a two-locus marker combination comprising psbK-psbI + atpF-atpH was also promising according to results from the Basic Local Alignment Search Tool and a maximum likelihood gene tree generated using PHyML. Two-dimen- sional DNA barcodes (i.e., QR codes) for the psbK-psbI and psbK-psbI + atpF-atpH regions were created for the validation of fresh or dried R. stricta samples.
Keywords:
Rhazya Stricta, Medicinal Plant, DNA Barcoding, matK, rbcl, rpoB, rpoC1, atpF-atpH, psbK-psbI, Two-Dimensional DNA Barcode, QR Code

1. Introduction
Rhazya stricta of the Apocynaceae family is an important folkloric medicinal plant of Arabia [1] . The genus Rhazya includes only two species, Rhazya stricta and Rhazya orientalis (syn. Amsonia orientalis) [2] and both of these are unambiguously distributed, as the natural habitats of R. stricta are the coastlines and arid regions of Arabian Peninsula and the Indian subcontinent [3] while those of R. orientalis are northwest Turkey and northeast Greece [4] . But these two sister species exhibit similarity in possessing therapeutically significant secondary metabolites. About 100s of alkaloids [2] and flavonoids from R. stricta contain innumerable pharmacological properties [5] [6] [7] . R. stricta is considered as a potential chemopreventive [8] [9] , antifungal [10] , and antidiabetic [11] agent and also produces analgesic and sedative effects [7] . Likewise the metabolites originating from R. orientalis have various anti-cancer and anti-tumor properties [12] . This study targeted the locally available samples of R. stricta species hence its sole sister species R. orientalis was not included in this study.
Like other medicinal plants, R. stricta is procured either from the vast deserts or from herbal markets, where it goes by the vernacular name “Harmal”, and is found in a dried or powdered form. Although authenticity is among the most significant aspects related to pharmacological products [13] the absence of distinct phenotypic traits for R. stricta impedes the identification of this plant. Unfortunately, contamination and adulteration of medicinal plant products are quite common in the herbal markets, which have even caused severe diseases and death in some cases [14] . It is essential to carry out molecular level identification of medicinal herbs before they are used as a therapeutic agent [15] . Thus, there is a dire need for accurately identifying R. stricta at the molecular level in addition to the traditional taxonomical examination to protect consumers.
Within the past two decades, we have witnessed advances in molecular level techniques for accurately discerning specimens, even in the absence of diagnostic morphological characteristics. These techniques have proved unambiguous for the authentication of medicinal plants [16] . One such molecular tool is DNA barcoding, which can offer quick authentication compared with traditional taxonomy [17] . This technology identifies unknown species by using standardized DNA segments as universal product codes. It targets highly conserved DNA sequences in which minor nucleotide polymorphisms have evolved [18] . These nucleotide variations aid in the creation of unique DNA barcode markers, which can be used to validate a sample from a given species [19] .
Whereas the 5' end of the mitochondrial gene for cytochrome c oxidase 1 is used for the standard barcode for animals [20] , this region is unsuitable for use in identifying plants owing to a low substitution rate [21] . A variety of complex evolutionary processes in plants (e.g., hybridization and polyploidy) complicate the distinctions for defining species boundaries [22] . Consequently, after years of tremendous efforts, not one single region has worked as an identifier for all of the plant species tested [23] [24] , and it is unlikely that a single universal plant DNA barcoding marker exists [25] .
Various regions of plant DNA have been studied for barcoding [26] [27] [28] [29] , including the coding and noncoding regions of plastid genomes (rbcL, psbA-trnH, trnL-trnF, and matK) and nuclear regions (5S, 16S, 18S, and ITS) [30] . These efforts have led to six plastid DNA regions as leading candidates for a suitable plant barcode, namely, matK, rbcL, rpoB, rpoC1, and psbK-psbI and atpF-atpH intergenic spacers [29] . Indeed, some of the loci from chloroplastid DNA, including matK, rbcL, atpF-atpH, psbK-psbI, rpoB, and rpoC1, have shown promising results for identifying medicinal plants. For example, the matK region, which was proposed as a standard barcode for flowering plants after studying 1,084 plant species [28] , produced positive results for the medicinal plant Rauvolfioideae (a subfamily of Apocynaceae) [31] . rbcL was recommended for species discrimination [29] , owing to the successes for its amplification and sequencing [27] . Moreover, taxonomic confusions among Cupressaceae, Cornaceae, Ericaceae, and Geraniaceae specimens were resolved via rbcL sequences [32] . Sequences for rpoB and rpoC1 were informative in identifying Ochradenus arabicus [19] , and Chase et al. [26] suggested they be used in combination with matK for DNA barcoding.
Promising results have also been obtained with the noncoding intergenic spacers, which have evolved comparatively rapidly and exhibit sequence divergence and high rates of insertion/deletion [27] . The intergenic spacer atpF-atpH could be used for validating medicinal plant material [33] and for distinguishing all three species in the genera Landoltia and Spirodela [34] . The intergenic spacer psbK-psbI shows potential for barcoding of the flora of Kruger National Park, South Africa [28] and its use has been endorsed over that of matK and other loci by the CBOL Plant Working Group, 2009 [29] due to its capability for discriminating species.
Nevertheless, a single locus DNA barcode has been proposed to lack sufficient variation for discriminating closely related taxa [35] . Hence, to enhance the identification capabilities of barcode markers, we investigated a multi-locus DNA barcode [27] [28] . We also determined the abilities of the six markers proposed by the CBOL Plant Working Group for discriminating R. stricta at the molecular level.
2. Materials and Methods
2.1. Ethics Testimony
The R. stricta plant is indigenous to the region and was found growing in the surrounding deserts and on roadsides; hence, no permission was required for sample collection.
2.2. Plant Material
Ten fresh leaf samples of R. stricta were collected from each of five different locations in the Western province of Saudi Arabia (at 21˚21.042', 39˚33.054'; 21˚24.849', 39˚44.044'; 21˚29.622', 39˚35.823'; 21˚26.817', 39˚31.109'; and 21˚26.423', 39˚31.853') to exclude the presence of any subspecies. We sampled plants during the winter season, when they are actively growing. Plants were identified at King Abdul Aziz University on the basis of morphological markers. Samples were immersed in liquid nitrogen and stored at -80˚C until DNA extraction. Five raw dried and powdered samples of R. stricta, were obtained from the local herbal market to check the efficiency of the proposed DNA barcode.
2.3. DNA Extraction, Amplification, and Sequencing
R. stricta is a poisonous medicinal plant with high levels of alkaloids and flavonoids that can make DNA extraction challenging. To overcome this, genomic DNA was extracted from young leaf tissues [36] using a DNeasy plant mini kit (Qiagen, Germany) according to the manufacturer’s instructions with minor modifications (elution buffer AE was heated to 65˚C and elution times were increased by 30 min). Following the same modification DNA was extracted from the R. stricta market samples. All of the extracts were stored at −20˚C until their use as templates for PCR. The concentrations and purities of the extracted DNA were quantified using a Nano Drop 2000 spectrophotometer (Thermo Scientific, Wilmington, DE, USA). The extracted genomic DNA was also examined by electrophoresis on a 1% agarose gel in 1× TBE buffer and stained with ethidium bromide (0.5 mg∙mL−1).
Target loci sequences were amplified in a total volume of 30 μL, consisting of 15 μL of GoTaq green master mix (for final concentrations of 200 μM for each deoxynucleoside triphosphate and 1.5 mM MgCl2), 1 μM each of forward and reverse primers, 25 - 250 ng of sample genomic DNA, and deionized distilled water. The primers (Bioneer Corp., South Korea) and parameters used for amplifying chloroplast loci are shown in Table 1. As initial amplifications failed for some loci (e.g., matK), which has been reported for samples in the order Gentianales [37] , the thermocycling conditions were optimized using touchdown PCR (Table 1). This resulted in successful amplification of target regions.
The amplicons werThe amplicons were analyzed on a 1.5% agarose gel as described above for genomic DNA analysis using a 100-bp DNA ladder (Promega Corp., USA) as a molecular marker. Gel images were analyzed on a gel documentation system (Biospectrum 410; UVP). PCR products were sent to Macrogen, Inc. (Seoul, South Korea) for bidirectional sequencing using the same primers to resolve ambiguities [38] .
2.4. Data Analysis
Sequence chromatograms were viewed via Flinch TV 1.4.0 (Geospiza, Inc., Seattle, WA, USA) to analyze base calls and quality values. Further analysis and alignments were carried out using Geneious 9.1 software [39] . Sequence similarities were identified using the BLAST algorithm
(http://blast.ncbi.nlm.nih.gov/blast.cgi) to judge the identification capability of the six-barcode regions. High Bit scores, Grades (a weighted score for the hit comprising the E-value, pairwise identity, and coverage), and a cutoff E-value of 1e20 were taken into account. Sequences of the barcode regions for R. stricta and
Table 1. Primers and amplification conditions for chloroplastid regions.
for seven highly similar species (related genera from the Apocynaceae family) that performed the best were downloaded from the GenBank database to assess their discriminatory efficiency (Table 2). We studied species from different genera, as no data from the chloroplast regions of other species in the genus Rhazya (other than R. stricta) appeared in our GenBank database search. These sequences were edited and aligned in Geneious Pro 9.1 [39] , using the multiple alignments tool MUSCLE plug-in [40] . For alignment, sequence orders from the Blast search were preserved. GC contents and the percentages of identical nucleotide sites were analyzed with the same software.
We used the tree-base method of identification with phylogenetic distance (via gene tree) to study the divergences of different barcode markers. Here, a query was allocated to the species with whom it clustered. In this study, phylogenetic distances were analyzed using PHyML 3.0 software [41] . On the basis of the HKY85 model of nucleotide substitution, this software generated a maximum likelihood gene tree with a 1000-replicate bootstrap test to authenticate R. stricta. After these analyses, all the sequences were deposited into the GenBank nucleotide database (Table 3).
In addition to the single locus barcodes, a double locus combination was investigated by concatenating the candidate barcode regions using Geneious Pro 9.1 software. The mean intra-and interspecific genetic distances were calculated using MEGA 6.0 [42] with the Kimura 2-parameter model. The proposed DNA barcodes were validated on five dried powered market samples ofR. stricta.
2.5. Two-Dimensional DNA Barcoding
The nucleotide sequences of the candidate barcode markers, which validated R. stricta species efficiently, were converted to two-dimensional DNA barcode images using an open-source PHP QR coding method [43] .
Table 2. Sequence sources for atpF-atpH and psbK-psbI markers from GenBank.
Table 3. Accession numbers for Rhazya stricta sequences submitted to GenBank.
3. Results
3.1. Success of DNA Isolation and Amplification
Although the extraction of DNA was challenging from some samples, DNA was successfully extracted using the modifications described in the Methods. All six of the chloroplast barcode loci were successfully amplified after optimizing the thermocycling conditions. The properties of these six loci are given in “Table 4”.
3.2. BLAST Analysis and Multiple Alignments
The BLAST analysis of the six barcode chloroplast loci (matK, rbcL, rpoB, rpoC1, atpF-atpH, and psbK-psbI) showed 100% identities and query covers with the R. stricta sequence (GenBank no. KJ485849). In addition, the sequences from the loci of R. stricta, viz., matK, rbcL, rpoB, and rpoC1, showed maximum (95%, 98%, 97%, and 98%, respectively) similarities with the other taxa examined (of the Apocynaceae family). However, atpF-atpH and psbK-psbI sequences showed ≤90.6% and ≤87.5% similarities, respectively, to the other taxa sequences (given in Table 5). They were found to be distinct for R. stricta.
There were no variations in the intraspecific alignments of sequencing products for the six loci (from fifty R. stricta samples). The mean intraspecific distances (calculated by MEGA 6) were zero, verifying that all of the fifty R. stricta samples belong to the same species with no genotypic variation.
The interspecific divergences (investigated after carrying out a multiple
Table 4. Properties of six candidate DNA barcoding loci in R. stricta sequences.
Table 5. Results of BLAST analysis for R. stricta psbK-psbI sample sequences, showing level of similarity with other neighboring species from the Apocynaceae family.
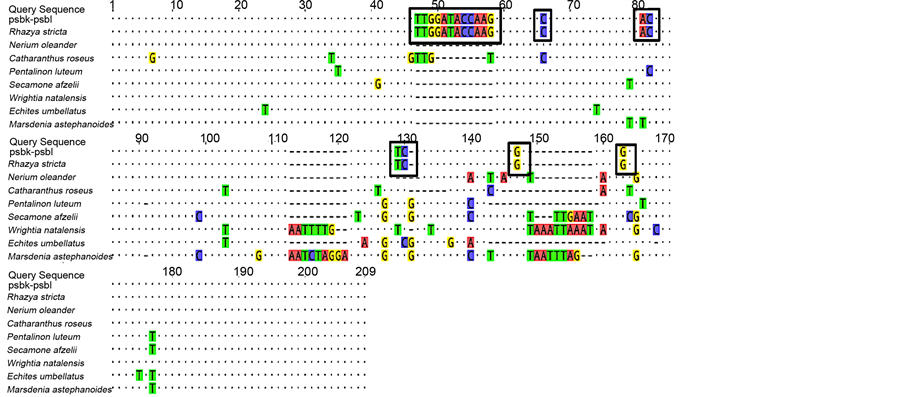
alignment between R. stricta and the seven species from Apocynaceae) for matK, rbcL, rpoB, and rpoC1 regions were low due to the high percentages of identical sites (Table 4). Nevertheless, interspecific divergences were high for atpF-atpH and psbK-psbI loci due to variations in R. stricta sample sequences (Table 4). The identical sites percentages (calculated using Geneious Pro 9.1) for these two loci were much lower than for the coding loci (Table 4). “Figure 1” highlights the variable informative sites in the psbK-psbI locus from R. stricta sample sequences. The mean interspecific distance was highest for psbK-psbI (7.80%) compared with those of the other loci.
3.3. Maximum Likelihood Tree Identification
The maximum likelihood tree [41] for the psbK-psbI region clustered sample sequences of R. stricta (query sequences) with the GenBank R. stricta sequences into a single independent clade that was highly supported (Figure 2(a)). The case was similar for the double locus (psbK-psbI + atpF-atpH) sample sequences
Figure 1. Multiple-sequence alignments highlighting the variable informative sites in the psbK-psbI locus from R. stricta sample sequences.
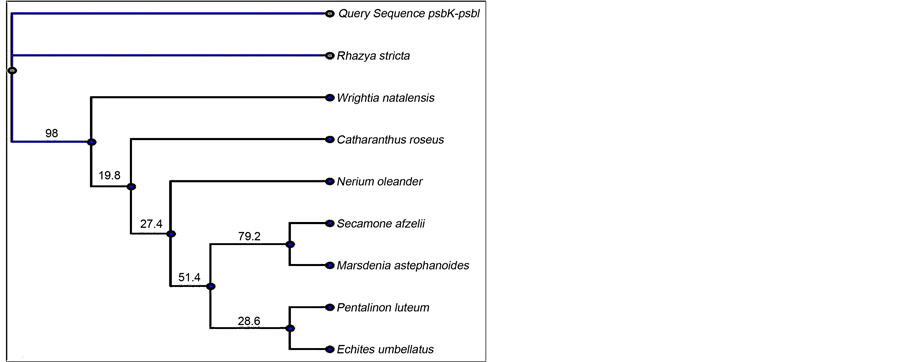
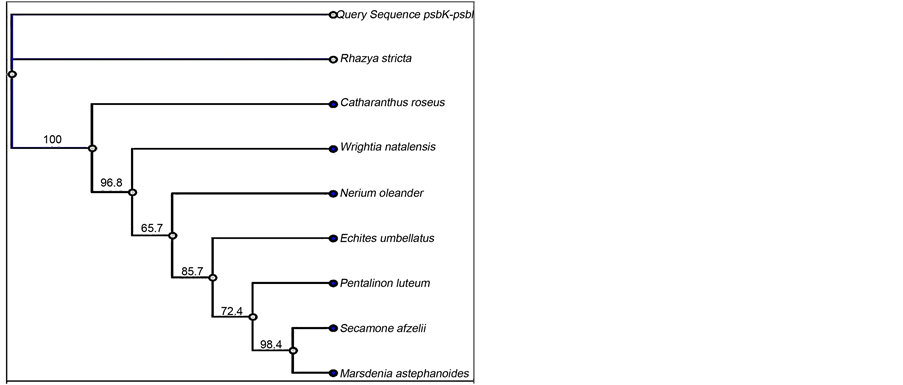
Figure 2. Maximum likelihood trees of psbK-psbI (a) and psbK-psbI + atpF-atpH (b) sequences for R. stricta. The branch labels display maximum likelihood bootstrap values.
(Figure 2(b)). Hence, the similarity-based method and tree-based identification indicate that psbK-psbI is an appropriate DNA barcode region for identifying R. stricta. The psbK-psbI intergenic region aided by atpF-atpH also displayed promising results, supporting its use as a double locus barcode for R. stricta. These two proposed DNA barcodes (psbK-psbI and psbK-psbI + atpF-atpH) clearly identified dried powdered R. stricta samples acquired from the market.
3.4. Generation of Two-Dimensional Barcodes
The purpose of DNA barcoding is to accurately and quickly identify medicinal plants and their products at the molecular level to benefit the consumer. With this aim, the nucleotide sequences of the DNA barcode markers (psbK-psbI and psbK-psbI + atpF-atpH) were considered analogous to the products’ barcodes in a supermarket and were converted to two-dimensional barcodes (i.e., QR codes) (Figure 3). The colors in the bars represent nucleotides of the marker, and the numbers at the ends denote the lengths of the markers. After scanning the two- dimensional barcode on a mobile terminal, a message can be sent to a DNA barcode database, where the authenticity of a plant or specimen, such as R. stricta, can be checked. This will aid the ability of consumers with no prior knowledge of DNA barcoding to accurately identify medicinal plants and their products.
4. Discussion
Approximately 80% of the world’s population are inclined to use herbal medicines for their primary care [44] . Similarly, the Saudi population exhibits a keen interest in the medicinal species of the local flora [45] . For example, the numerous pharmacological properties of R. stricta make this a significant medicinal plant in this part of the world [46] . Although it is easily available from its local natural habitat, identifying this plant in herbal markets is difficult due to the lack
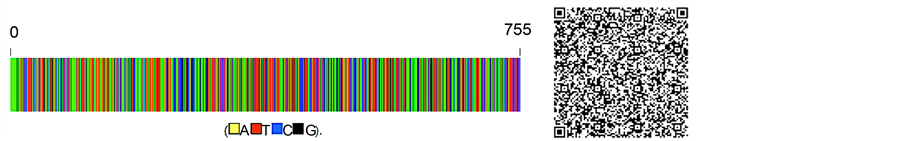


Figure 3. DNA barcodes and two-dimensional DNA barcodes of psbK-psbI (a) and psbK-psbI + atpF-atpH (b) sequences for R. stricta.
of distinct morphological traits and the variability in storage conditions and product ages. In an effort to safeguard consumers’ health, we investigated a method of DNA barcoding for validating R. stricta at the molecular level, irrespective of its physical state.
DNA barcoding is considered the renaissance of taxonomy [47] . This molecular tool emerged rapidly over the past two decades for species discrimination [25] , and it has outperformed other diagnostic tools for identifying and authenticating species [48] . Numerous studies have not only affirmed its competency for species identification, but also highlighted its strengths in defining species boundaries and flagging new species [20] [49] . Owing to its ease and swiftness, this tool has been used extensively for distinguishing medicinal plants [50] [51] [52] . It is predicted to play a significant role in identifying medicinal plant products in the future [53] .
It was proposed that an ideal DNA barcode (i) can be easily amplified and sequenced, (ii) does not exceed 1 kb in size, and (iii) has higher interspecific than intraspecific variation [54] . We selected six barcoding loci to examine with these criteria in mind. The rbcL, rpoB, rpoC1, psbK-psbI, and atpF-atpH loci were easily amplified and sequenced. The amplification of matK was challenging initially, which has been found in medicinal Uncaria species [55] and temperate flora comprising 436 species in 269 genera of land plants [56] . By contrast, the psbK-psbI region amplified easily, similar to that for Panax species (Ginsengs) [57] . The successful amplification of intergenic spacers (psbK-psbI and atpF- atpH) results from the ability to use universal primers for the highly conserved coding sequences on either side of the locus [58] .
In this study, the sequences of R. stricta for matK, rbcL, rpoC1, and rpoB were similar to those for the other taxa. Accordingly, matK did not aid in species identification in other studies, suggesting that it may be nonfunctional in some taxa [29] [17] . The rpoC1, rpoB [59] [60] , and rbcL [61] loci similarly have shown very low species discrimination in other investigations. The noncoding intergenic spacer regions, psbK-psbI and atpF-atpH, are promising barcode markers for R. stricta due to their faster divergence than genes in otherwise slow-evolving plant species [62] [27] . Indeed, the psbK-psbI region exhibited efficient barcode recovery and species discrimination in a number of studies [29] [63] . Zuo et al. [58] endorsed psbK-psbI over coding regions, as it is a more informative and powerful marker for species identification. This locus discriminates the Kruger National Park flora [28] and taxa of Orchidaceae in Korea [64] . Similarly, the atpF-atpH intergenic spacer has been endorsed as a supplementary locus [65] as it was found to successfully authenticate medicinal plant materials in a study involving 17 barcode regions [33] . Another option that was proposed was to combine psbK-psbI with other loci, such as matK and atpF-atpH [66] [28] [65] .
DNA barcoding takes advantage of polymorphisms that lead to gaps in sequences. Polymorphisms in psbK-psbI and atpF-atpH regions were observed as gaps upon amplicon alignment that were found to be distinct to R. stricta and might serve for barcoding this species. In addition, the psbK-psbI and concatenated psbK-psbI + atpF-atpH regions displayed 64.1% and 72.4% similarities (on the basis of nucleotide sites), respectively, to the other species sequences on alignment. Hence, these loci are considerably unique with regard to R. stricta. Indeed, a 100% monophyletic differentiation of the R. stricta species was found for the single locus (psbK-psbI) and double locus combination (psbK-psbI + atpF-atpH) as shown by the maximum likelihood trees. Moreover, the psbK-psbI and psbK-psbI + atpF-atpH loci successfully identified market samples of R. stricta.
5. Conclusion
In this study, we found that matK, rbcL, rpoC1, and rpoB coding regions are not effective markers for R. stricta, as they were similar to the sequences of other species. By contrast, intergenic spacer regions psbK-psbI and atpF-atpH contained sequence polymorphisms that are unique to R. stricta. We propose the use of psbK-psbI as an optimal single locus barcode and psbK-psbI + atpF-atpH as a complementary double locus combination for creating DNA barcodes to identify the medicinal plant R. stricta. We generated two-dimensional barcodes (QR codes) for these loci to facilitate molecular authentication of R. stricta via an easy and inexpensive method for consumers. This method would also greatly benefit market supervision of other medicinal plants of Saudi Arabia and its neighboring countries. It can be a preliminary step towards generating a databank of barcode markers for economically significant flora of this arid region.
Acknowledgements
King Abdullah City of Science and Technology (KACST) supported this study under the grant GSP-34-0119.
Cite this paper
Khan, S.A., Baeshen, M.N., Ramadan, H.A. and Baeshen, N.A. (2017) Emergence of Plastidial Intergenic Spacers as Suitable DNA Barcodes for Arid Medicinal Plant Rhazya stricta. American Journal of Plant Sciences, 8, 1774-1789. https://doi.org/10.4236/ajps.2017.88121
References
- 1. Mandaville, J.P. (2011) Bedouin Ethnobotany: Plant Concepts and Uses in a Desert Pastoral World. University of Arizona.
- 2. Marwat, S.K., Usman, K., Shah, S.S., Anwar, N., and Ullah, I. (2012) A Review of Phytochemistry, Bioactivities and Ethno Medicinal Uses of Rhazya Stricta Decsne (Apocynaceae). African Journal of Microbiology Research, 6, 1629-1641.
- 3. Zahran, M.A. (2010) Climate-Vegetation: Afro-Asian Mediterranean and Red Sea Coastal Lands. Springer Science & Business Media.
https://doi.org/10.1007/978-90-481-8595-5 - 4. ACEMI, A., TüRKER-KAYA, S., and Fazil, OZEN (2016) FT-IR Spectroscopy Based Evaluation of Changes in Primary Metabolites of Amsonia Orientalis after In Vitro 6-Benzylaminopurine Treatment. Notulae Botanicae Horti Agrobotanici Cluj-Napoca, 44, 209-214.
https://doi.org/10.15835/nbha44110194 - 5. Elkady, A.I., Hussein, R.A.E.H., and Abu-Zinadah, O.A. (2014) Differential Control of Growth, Apoptotic Activity and Gene Expression in Human Colon Cancer Cells by Extracts Derived from Medicinal Herbs, Rhazya Stricta and Zingiber Officinale and Their Combination. World Journal of Gastroenterology, 20, 15275-15288.
https://doi.org/10.3748/wjg.v20.i41.15275 - 6. Said, S.A., Al-Saadi, S.H.A., Al-Abri, A.R., Akhtar, M.S., Weli, A.M., and Al-Riyami, Q. (2014) Cytotoxic Properties of Some Herbal Plants in Oman. Journal of Taibah University for Science, 8, 71-74.
https://doi.org/10.1016/j.jtusci.2014.01.004 - 7. Ahmed, A., Asad, M.J., Ahmad, M.S., Qureshi, R., Shah, S.I., Gul, H., and Gulfraz, M. (2015) Antidiabetic and Hypolipidemic Potential of Rhazya Stricta Decne Extract and Its Fractions. International Current Pharmaceutical Journal, 4, 353-361.
https://doi.org/10.3329/icpj.v4i2.21484 - 8. Baeshen, N.A., Elkady, A.I., Abuzinadah, O.A., and Mutwakil, M.H. (2012) Potential Anticancer Activity of the Medicinal Herb, Rhazya Stricta, against Human Breast Cancer. African Journal of Biotechnology, 11, 8960-8972.
https://doi.org/10.5897/AJB12.570 - 9. Elkady, A.I. (2013) Crude Alkaloid Extract of Rhazya Stricta Inhibits Cell Growth and Sensitizes Human Lung Cancer Cells to Cisplatin through Induction of Apoptosis. Genetics and molecular biology, 36, 12-21.
https://doi.org/10.1590/S1415-47572013005000009 - 10. Emad, A.M., and Gamal, E.G.E. (2013) Screening for Antimicrobial Activity of Some Plants from Saudi Folk Medicine. Global Journal of Research on Medicinal Plants & Indigenous Medicine, 2, 189-197.
- 11. Baeshen, N., Lari, S., Al Doghaither, H.A. and Ramadan, H.A. (2010) Effect of Rhazya stricta Extract on Rat Adiponectin Gene and Insulin Resistance. Journal of American Science, 6, 1237-1245.
- 12. Itoh, A., Kumashiro, T., Tanahashi, T., Nagakura, N. and Nishi, T. (2002) Flavonoid Glycosides from Rhazya orientalis. Journal of Natural Products, 65, 352-357.
https://doi.org/10.1021/np010518w - 13. Veldman, S., Otieno, J., Gravendeel, B., van Andel, T. and de Boer, H. (2014) Conservation of Endangered Wild Harvested Medicinal Plants: Use of DNA Barcoding. In: Gurib-Fakim, A., Novel Plant Bioresources: Applications in Food, Medicine and Cosmetics, John Wiley & Sons, Ltd., Chichester, 81-88.
https://doi.org/10.1002/9781118460566.ch6 - 14. Moraes, D.F.C., Still, D.W., Lum, M.R. and Hirsch, A.M. (2015) DNA-Based Authentication of Botanicals and Plant-Derived Dietary Supplements: Where Have We Been and Where Are We Going? Planta Medica, 81, 687-695.
https://doi.org/10.1055/s-0035-1545843 - 15. Vassou, S.L., Kusuma, G. and Parani, M. (2015) DNA Barcoding for Species Identification from Dried and Powdered Plant Parts: A Case Study with Authentication of the Raw Drug Market Samples of Sida cordifolia. Gene, 559, 86-93.
- 16. Uncu, A.T., Frary, A. and Doganlar, S. (2015) Cultivar Origin and Admixture Detection in Turkish Olive Oils by SNP-Based CAPS Assays. Journal of Agricultural and Food Chemistry, 63, 2284-2295.
https://doi.org/10.1021/acs.jafc.5b00090 - 17. Mishra, P., Kumar, A., Nagireddy, A., Mani, D.N., Shukla, A.K., Tiwari, R. and Sundaresan, V. (2016) DNA Barcoding: An Efficient Tool to Overcome Authentication Challenges in the Herbal Market. Plant Biotechnology Journal, 14, 8-21.
https://doi.org/10.1111/pbi.12419 - 18. Ali, M.A., Gyulai, G., Hidvegi, N., Kerti, B., Al Hemaid, F.M., Pandey, A.K. and Lee, J. (2014) The Changing Epitome of Species Identification-DNA Barcoding. Saudi Journal of Biological Sciences, 21, 204-231.
- 19. Khan, S., Al-Qurainy, F., Nadeem, M. and Tarroum, M. (2012) Development of Genetic Markers for Ochradenus arabicus (Resedaceae), an Endemic Medicinal Plant of Saudi Arabia. Genetics and Molecular Research, 11, 1300-1308.
- 20. Hebert, P.D., Cywinska, A. and Ball, S.L. (2003) Biological Identifications through DNA Barcodes. Proceedings of the Royal Society of London B: Biological Sciences, 270, 313-321.
https://doi.org/10.1098/rspb.2002.2218 - 21. Wolfe, K.H., Li, W.H. and Sharp, P.M. (1987) Rates of Nucleotide Substitution Vary Greatly among Plant Mitochondrial, Chloroplast, and Nuclear DNAs. Proceedings of the National Academy of Sciences of the United States of America, 84, 9054-9058.
https://doi.org/10.1073/pnas.84.24.9054 - 22. Fazekas, A.J., Kesanakurti, P.R., Burgess, K.S., Percy, D.M., Graham, S.W., Barrett, S.C., et al. (2009) Are Plant Species Inherently Harder to Discriminate than Animal Species Using DNA Barcoding Markers? Molecular Ecology Resources, 9, 130-139.
https://doi.org/10.1111/j.1755-0998.2009.02652.x - 23. Chase, M.W. and Fay, M.F. (2009) Barcoding of Plants and Fungi. Science, 325, 682-683.
https://doi.org/10.1126/science.1176906 - 24. Chen, S., Yao, H., Han, J., Liu, C., Song, J., Shi, L., et al. (2010) Validation of the ITS2 Region as a Novel DNA Barcode for Identifying Medicinal Plant Species. PLoS ONE, 5, e8613.
https://doi.org/10.1371/journal.pone.0008613 - 25. Pecnikar, Z.F. and Buzan, E.V. (2014) 20 Years Since the Introduction of DNA Barcoding: From Theory to Application. Journal of Applied Genetics, 55, 43-52.
https://doi.org/10.1007/s13353-013-0180-y - 26. Chase, M.W., Salamin, N., Wilkinson, M., Dunwell, J.M., Kesanakurthi, R.P., Haidar, N. and Savolainen, V. (2005) Land Plants and DNA Barcodes: Short-Term and Long-Term Goals. Philosophical Transactions of the Royal Society of London B: Biological Sciences, 360, 1889-1895.
https://doi.org/10.1098/rstb.2005.1720 - 27. Kress, W.J. and Erickson, D.L. (2007) A Two-Locus Global DNA Barcode for Land Plants: The Coding rbcL Gene Complements the Non-Coding trnH-psbA Spacer Region. PLoS ONE, 2, e508.
https://doi.org/10.1371/journal.pone.0000508 - 28. Lahaye, R., Van der Bank, M., Bogarin, D., Warner, J., Pupulin, F., Gigot, G., et al. (2008) DNA Barcoding the Floras of Biodiversity Hotspots. Proceedings of the National Academy of Sciences of the United States of America, 105, 2923-2928.
https://doi.org/10.1073/pnas.0709936105 - 29. Group, C.P.W., Hollingsworth, P.M., Forrest, L.L., Spouge, J.L., Hajibabaei, M., Ratnasingham, S., et al. (2009) A DNA Barcode for Land Plants. Proceedings of the National Academy of Sciences of the United States of America, 106, 12794-12797.
https://doi.org/10.1073/pnas.0905845106 - 30. Agarwal, M., Shrivastava, N. and Padh, H. (2008) Advances in Molecular Marker Techniques and Their Applications in Plant Sciences. Plant Cell Reports, 27, 617-631.
https://doi.org/10.1007/s00299-008-0507-z - 31. Mahadani, P., Sharma, G.D. and Ghosh, S.K. (2013) Identification of Ethnomedicinal Plants (Rauvolfioideae: Apocynaceae) through DNA Barcoding from Northeast India. Pharmacognosy Magazine, 9, 255-263.
https://doi.org/10.4103/0973-1296.113284 - 32. Khan, M.Q., Khalil, A.T., Rehman, K. and Shinwari, Z.K. (2015) Searching for DNA Barcodes in Plants. Global Journal of Biotechnology & Biochemistry, 10, 1-10.
- 33. Techen, N., Parveen, I., Pan, Z. and Khan, I.A. (2014) DNA Barcoding of Medicinal Plant Material for Identification. Current Opinion in Biotechnology, 25, 103-110.
https://doi.org/10.1016/j.copbio.2013.09.010 - 34. Bog, M., Lautenschlager, U., Landrock, M.F., Landolt, E., Fuchs, J., Sree, K.S., et al. (2015) Genetic Characterization and Barcoding of Taxa in the Genera Landoltia and Spirodela (Lemnaceae) by Three Plastidic Markers and Amplified Fragment Length Polymorphism (AFLP). Hydrobiologia, 749, 169-182.
https://doi.org/10.1007/s10750-014-2163-3 - 35. Li, X., Yang, Y., Henry, R.J., Rossetto, M., Wang, Y. and Chen, S. (2015) Plant DNA Barcoding: From Gene to Genome. Biological Reviews, 90, 157-166.
https://doi.org/10.1111/brv.12104 - 36. Fazekas, A.J., Burgess, K.S., Kesanakurti, P.R., Graham, S.W., Newmaster, S.G., Husband, B.C., et al. (2008) Multiple Multilocus DNA Barcodes from the Plastid Genome Discriminate Plant Species Equally Well. PLoS ONE, 3, e2802.
https://doi.org/10.1371/journal.pone.0002802 - 37. Dunning, L.T. and Savolainen, V. (2010) Broad-Scale Amplification of matK for DNA Barcoding Plants, a Technical Note. Botanical Journal of the Linnean Society, 164, 1-9.
https://doi.org/10.1111/j.1095-8339.2010.01071.x - 38. Middendorf, L.R., Humphrey, P.G., Narayanan, N. and Roemer, S.C. (2001) Sequencing Technology. 2nd Edition, Biotechnology Set, 193-226.
- 39. Kearse, M., Moir, R., Wilson, A., Stones-Havas, S., Cheung, M., Sturrock, S., et al. (2012) Geneious Basic: An Integrated and Extendable Desktop Software Platform for the Organization and Analysis of Sequence Data. Bioinformatics, 28, 1647-1649.
https://doi.org/10.1093/bioinformatics/bts199 - 40. Edgar, R.C. (2004) MUSCLE: Multiple Sequence Alignment with High Accuracy and High Throughput. Nucleic Acids Research, 32, 1792-1797.
https://doi.org/10.1093/nar/gkh340 - 41. Guindon, S., Dufayard, J.F., Lefort, V., Anisimova, M., Hordijk, W. and Gascuel, O. (2010) New Algorithms and Methods to Estimate Maximum-Likelihood Phylogenies: Assessing the Performance of PhyML 3.0. Systematic Biology, 59, 307-321.
https://doi.org/10.1093/sysbio/syq010 - 42. Tamura, K., Stecher, G., Peterson, D., Filipski, A. and Kumar, S. (2013) MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Molecular Biology and Evolution, 30, 2725-2729.
https://doi.org/10.1093/molbev/mst197 - 43. Yu, N., Gu, H., Wei, Y., Zhu, N., Wang, Y., Zhang, H., et al. (2016) Suitable DNA Barcoding for Identification and Supervision of Piper kadsura in Chinese Medicine Markets. Molecules, 21, 1221.
https://doi.org/10.3390/molecules21091221 - 44. Choudhary, M., Kumar, V., Malhotra, H. and Singh, S. (2015) Medicinal Plants with Potential Anti-Arthritic Activity. Journal of Intercultural Ethnopharmacology, 4, 147-179.
https://doi.org/10.5455/jice.20150313021918 - 45. Al-Essa, M.A., Al-Mehaidib, A. and Al-Gain, S. (1998) Parental Awareness of Liver Disease among Children in Saudi Arabia. Annals of Saudi Medicine, 18, 79-81.
https://doi.org/10.5144/0256-4947.1998.79 - 46. Gilani, S.A., Kikuchi, A., Shinwari, Z.K., Khattak, Z.I. and Watanabe, K.N. (2007) Phytochemical, Pharmacological and Ethnobotanical Studies of Rhazya stricta Decne. Phytotherapy Research, 21, 301-307.
https://doi.org/10.1002/ptr.2064 - 47. Miller, S.E. (2007) DNA Barcoding and the Renaissance of Taxonomy. Proceedings of the National Academy of Sciences of the United States of America, 104, 4775-4776.
https://doi.org/10.1073/pnas.0700466104 - 48. Schmidt, S., Schmid-Egger, C., Morinière, J., Haszprunar, G. and Hebert, P.D. (2015) DNA Barcoding Largely Supports 250 Years of Classical Taxonomy: Identifications for Central European Bees (Hymenoptera, Apoidea partim). Molecular Ecology Resources, 15, 985-1000.
https://doi.org/10.1111/1755-0998.12363 - 49. Hebert, P.D.N., Penton, E.H., Burns, J.M., Janzen, D.H. and Hallwachs, W. (2004) Ten Species in One: DNA Barcoding Reveals Cryptic Species in the Neotropical Skipper Butterfly Astraptes fulgerator. Proceedings of the National Academy of Sciences of the United States of America, 101, 14812-14817.
https://doi.org/10.1073/pnas.0406166101 - 50. Li, M., Au, K.Y., Lam, H., Cheng, L., But, P.P.H. and Shaw, P.C. (2014) Molecular Identification and Cytotoxicity Study of Herbal Medicinal Materials That Are Confused by Aristolochia Herbs. Food Chemistry, 147, 332-339.
- 51. Zheng, S.H., Ren, W.G., Wang, Z.H. and Huang, L.F. (2015) Use of Chloroplast DNA Barcodes to Identify Osmunda japonica and Its Adulterants. Plant Systematics and Evolution, 301, 1843-1850.
https://doi.org/10.1007/s00606-015-1197-y - 52. Ganie, S.H., Upadhyay, P., Das, S. and Sharma, M.P. (2015) Authentication of Medicinal Plants by DNA Markers. Plant Gene, 4, 83-99.
- 53. Han, J., Pang, X., Liao, B., Yao, H., Song, J. and Chen, S. (2016) An Authenticity Survey of Herbal Medicines from Markets in China Using DNA Barcoding. Scientific Reports, 6, Article No. 18723.
https://doi.org/10.1038/srep18723 - 54. Wong, K.L., But, P.H. and Shaw, P.C. (2013) Evaluation of Seven DNA Barcodes for Differentiating Closely Related Medicinal Gentiana Species and Their Adulterants. Chinese Medicine, 8, 16.
https://doi.org/10.1186/1749-8546-8-16 - 55. Zhang, J.Q., Meng, S.Y., Wen, J. and Rao, G.Y. (2015) DNA Barcoding of Rhodiola (Crassulaceae): A Case Study on a Group of Recently Diversified Medicinal Plants from the Qinghai-Tibetan Plateau. PLoS ONE, 10, e0119921.
https://doi.org/10.1371/journal.pone.0119921 - 56. Burgess, K.S., Fazekas, A.J., Kesanakurti, P.R., Graham, S.W., Husband, B.C., Newmaster, S.G., et al. (2011) Discriminating Plant Species in a Local Temperate Flora Using the rbcL+matK DNA Barcode. Methods in Ecology and Evolution, 2, 333-340.
https://doi.org/10.1111/j.2041-210X.2011.00092.x - 57. Zuo, Y., Chen, Z., Kondo, K., Funamoto, T., Wen, J. and Zhou, S. (2011) DNA Barcoding of Panax Species. Planta Medica, 77, 182-187.
https://doi.org/10.1055/s-0030-1250166 - 58. Shaw, J., Lickey, E.B., Beck, J.T., Farmer, S.B., Liu, W., Miller, J., et al. (2005) The Tortoise and the Hare II: Relative Utility of 21 Noncoding Chloroplast DNA Sequences for Phylogenetic Analysis. American Journal of Botany, 92, 142-166.
https://doi.org/10.3732/ajb.92.1.142 - 59. Starr, J.R., Naczi, R.F. and Chouinard, B.N. (2009) Plant DNA Barcodes and Species Resolution in Sedges (Carex, Cyperaceae). Molecular Ecology Resources, 9, 151-163.
https://doi.org/10.1111/j.1755-0998.2009.02640.x - 60. Yu, H., Wu, K., Song, J., Zhu, Y., Yao, H., Luo, K., et al. (2014) Expedient Identification of Magnoliaceae Species by DNA Barcoding. Plant Omics, 7, 47-53.
- 61. Asahina, H., Shinozaki, J., Masuda, K., Morimitsu, Y. and Satake, M. (2010) Identification of Medicinal Dendrobium Species by Phylogenetic Analyses Using matK and rbcL Sequences. Journal of Natural Medicines, 64, 133-138.
- 62. Erickson, D.L., Spouge, J., Resch, A., Weigt, L.A. and Kress, J.W. (2008) DNA Barcoding in Land Plants: Developing Standards to Quantify and Maximize Success. Taxon, 57, 1304-1316.
- 63. Hollingsworth, M.L., Andra Clark, A.L.E.X., Forrest, L.L., Richardson, J., Pennington, R., Long, D.G., et al. (2009) Selecting Barcoding Loci for Plants: Evaluation of Seven Candidate Loci with Species-Level Sampling in Three Divergent Groups of Land Plants. Molecular Ecology Resources, 9, 439-457.
https://doi.org/10.1111/j.1755-0998.2008.02439.x - 64. Kim, H.M., Oh, S.H., Bhandari, G.S., Kim, C.S. and Park, C.W. (2014) DNA Barcoding of Orchidaceae in Korea. Molecular Ecology Resources, 14, 499-507.
https://doi.org/10.1111/1755-0998.12207 - 65. Vijayan, K. and Tsou, C.H. (2010) DNA Barcoding in Plants: Taxonomy in a New Perspective. Current Science (Bangalore), 99, 1530-1541.
- 66. Lee, H.L., Yi, D.K., Kim, J.S. and Kim, K.J. (2007) Development of Plant DNA Barcoding Markers from the Variable Noncoding Regions of Chloroplast Genome. In Abstract Presented at the 2nd International Barcode of Life Conference, Academia Sinica, Taipei, 18 September 2007.