 Advances in Anthropology 2012. Vol.2, No.1, 14-23 Published Online February 2012 in SciRes (http://www.SciRP.org/journal/aa) http://dx.doi.org/10.4236/aa.2012.21002 Copyright © 2012 SciRes. 14 A Time Series of Prehistoric Mitochondrial DNA Reveals Western European Genetic Diversity Was Largely Established by the Bronze Age François-Xavier Ricaut1*, Murray P. Cox2*, Marie Lacan1,3, Christine Keyser3, Francis Duranthon1, Bertrand Ludes3, Jean Guilaine4 , Eric Crubézy1 1Laboratoire d’Anthropologie Moléculaire et Imagerie de Synthèse, Centre National de la Recherche Scientifique, Université de Toulouse (Paul Sabatier), Toulouse, France 2Institute of Molecular BioScienc es, Massey University, Palmerston North, New Zealand 3Laboratoire d’Anthropologie Moléculaire, Centre National de la Rech erche Scientifique, Institute of Legal Medicine, University of Strasbourg, Strasbourg, France 4Centre de Recherche sur la Préh istoire et la Protohistoire de la Méditerra née, École des Hautes Etudes en Sciences Sociales, To ulouse, France Email: *fx.ricaut@infonie.fr, *m.p.cox@massey.ac.nz Received December 8th, 2011; revised J anu ary 1st, 2012; accepted February 1st, 2012 A major unanswered question concerns the roles of continuity versus change in prehistoric Europe. For the first time, genetic samples of reasonable size taken at multiple time points are revealing piecemeal snapshots of European prehistory at different dates and places across the continent. Here, we pull these disparate datasets together to illustrate how human genetic variation has changed spatially and temporally in Europe from the Mesolithic through to the present day. Mitochondrial DNA (mtDNA) haplogroups were determined for 532 European individuals from four major eras: the Mesolithic, Neolithic, Chalco- lithic (late Neolithic/early Bronze Age transition) and Modern periods. The Mesolithic was characterized by low mtDNA diversity. These initial European settler haplogroups declined rapidly in the Neolithic, as farmers from the east introduced a new suite of mtDNA lineages into Western Europe. For the first time, we show that the Chalcolithic was also a time of substantial genetic change in Europe. However, rather than the arrival of new mtDNA lineages, this period was characterized by major fluctuations in the fre- quencies of existing haplogroups. Besides the expansion of haplogroup H, there were few major changes in mtDNA diversity from the Chalcolithic to modern times, thus suggesting that the basic profile of mod- ern western European mtDNA diversity was largely established by the Bronze Age. Keywords: Europe; Prehistory; MtDNA; Haplogroup Diversity; Temporal Dynamics Introduction Modern European genetic diversity is increasingly well char- acterized. Researchers have detailed the geographic distribution of maternal (mtDNA) and paternal (Y chromosome) lineages, and are now delving into the spatial distribution of autosomal variants. European individuals can be assigned to modern na- tions, and even in particular cases, to areas as small as villages (Novembre et al., 2008). Yet how this genetic diversity has changed through time is much less well understood. Precise real-time information is largely restricted by advances in an- cient DNA research, a growing but challenging field. Never- theless, samples of reasonable size taken at multiple time points are now starting to provide snapshots of genetic variation dur- ing European prehistory at different times and places across the continent. It therefore seems timely to attempt a first pass at describing how human genetic variation has changed spatially and temporally in Europe from the Stone Age through to the present day. A major unanswered question concerns the roles of continu- ity versus change in prehistoric Europe. For instance, the main drivers of the agricultural expansion have long been a topic of considerable debate. What originally was perceived as a simple disti nctio n betwe en demi c vers us cultural diffusion is now gen- erally recognized as a more complex set of processes. These processes produced broad-scale demographic trends, while still allowing mosaic regional patterns (e.g., Ammerman & Cavalli- Sforza, 1984; Whittle, 1996; Whittle & Cummings, 2007). This more nuanced view emphasizes cultural and population disper- sals, as well as variable patterns of population admixture, where the relative biological contributions of Near Eastern Neolithic farmers and indigenous Mesolithic hunter-gatherers are not distributed uniformly across Europe (Sampietro et al., 2007; Bramanti et al., 2009; von Cramon-Taubadel & Pinhasi, 2011). Current understanding emphasizes two different migra- tion routes; a relatively rapid diffusion via a southern route along the Mediterranean coast (associated with the Impressed Ware and Cardial Ware culture), together with a slower north- ern route along the Danube valley into central Europe (associ- ated with the Linearbandkeramik, or LBK, culture) (Guilaine, 1997, 2003; Gronenborn, 1999; Zvelebil, 2004; Guilaine & Manen, 2007). Mosaic models such as these emphasize the complex processes by which the transition to agriculture in Europe probably took place, simultaneously explaining the seemingly conflicting results obtained from archaeological evidence (e.g., Guilaine, 2003; Bar-Yosef, 2004; Pinhasi et al., *Corresponding author.  F.-X. RICAUT ET AL. 2005; Bailey & Spikins, 2008; Bocquet-Appel et al, 2009; Rowley-Conwy, 2011), osteological data (e.g., Crubézy et al., 2002; Pinhasi & von Cramon-Taubadel, 2009; von Cramon- Taubadel & Pinhasi, 2011), modern genetic analyses (e.g., Cavalli-Sforza et al., 1994; Chikhi et al., 1998; Renfrew & Boyle, 2000; Bellwood & Renfrew, 2002; Richards, 2003; Belle et al., 2006; Balaresque et al, 2010; Palanichamy et al., 2010; Soares et al., 2010) and ancient DNA studies (e.g., Haak et al., 2005, 2010; Sampietro et al., 2007; Bramanti et al., 2009; Malmström et al., 2009; Lacan et al., 2011). Although considerable research has been focused on the spread of agriculture through Europe (as evidenced by the large number of studies cited above), the periods immediately prior to, and following, the agricultural revolution have been explored in far less detail. In this context, three subject areas would seem to warrant further examination: 1) Ancient southwestern/Mediterranean biological data are typically underrepresented, mainly due to bone preservation issues outside the colder/temperate climate regions of the north. Extant research has typically focused on ancient northern or central European populations (Haak et al., 2005, 2010; Bra- manti et al., 2009; Pinhasi & von Cramon-Taubadel 2009; von Cramon-Taubadel & Pinhasi 2011). Few studies have analyzed and compared ancient biological data from multiple sites or regions across Europe. 2) The effects of cultural differences on genetic diversity between southern and northern Europe through the Neolithic and Chalcolithic periods have not been addressed. This time frame saw two contrasting Neolithic cultures in Europe (Lin- earbandkeramik in central Europe and the Impressed/Cardial Ware culture in southern Europe), each seemingly linked to a different Chalcolithic subsistence strategy (a cattle-based, fresh milk dairying economy in northern Europe, and agropastoral cultures based on preserved milk sheep/goat farming in south- ern Europe) (Itan et al., 2009). The role of genetic connections between these two regions and cultures remains unclear. 3) Little is known about changes in genetic diversity during the important post-Neolithic transitional period. In addition to the Mesolithic/Neolithic transition, the Chalcolithic period also witnessed substantial culture change, particularly in terms of the adoption of metal tools, new agricultural techniques (yoke and ard-plough), means of transport (wheeled vehicles) and emerging long-distance exchange networks (Sherratt 1981). These important cultural processes had the potential for major demographic impacts on European populations. While studies of nuclear DNA still involve extremely small numbers of individuals (Burger et al., 2007; Haak et al., 2008, 2010; Malmström et al., 2009; Lacan et al., 2011) and are not considered further here, the accumulation of mtDNA sequences from ancient European individuals is increasing rapidly. We now have mtDNA data for >100 individuals from the Meso- lithic period (more than ~9000 years before present), the Neo- lithic period (the samples in this study are from the early Neo- lithic, ~9000 - 5500 years before present) and the Chalcolithic period (the transition from the late Neolithic to the early Bronze Age, ~5500 - 3700 years before present). This dataset, while still modest, is an extraordinary achieve- ment, and at last provides a test bed to explore simple demo- graphic scenarios that begin to move our perspective of Euro- pean genetic prehistory beyond intra-site/intra-region analyses. Using these ancient mtDNA sequences, we interrogate hap- logroup frequency distributions to resolve through-time bio- logical relationships between two different regions in western Europe (southern and northern), four different time points (Mesolithic, Neolithic, Chalcolithic and the Modern era), and two different cultural traditions (Linearbandkeramik versus the Impressed/Cardial Ware culture). To our knowledge, this is the first extended analysis of spatial and temporal genetic continu- ity through European prehistory. Methods Samples Mitochondrial DNA haplogroup classifications were deter- mined for 532 European individuals from four major eras: the Mesolithic, Neolithic, Chalcolithic and Modern periods (Table 1). This dataset includes 109 ancient samples, representing 15 individuals from the Mesolithic period (Bramanti et al., 2009), 56 individuals from the Neolithic (Haak et al., 2005, 2010; Haak, 2006; Sampietro et al., 2007; Deguilloux et al., 2011), and 38 individuals from the Chalcolithic (Itan et al., 2009; Haak et al., 2008). The Mesolithic dataset analyzed here spe- cifically excludes Mesolithic communities in close proximity to Neolithic sites, because these samples are potentially affected by gene flow from neighboring Neolithic peoples (Bramanti Table 1. Population samples use d in t hi s stu dy. Period Geographic Ar ea Location Dates n Culture Reference Mesolithic Central/North Germany, Russia, Poland, Lith u ania 13,400 - 2250 BC15 Kunda, Na rva, Zedmar, Beuronien Bramanti et al., 2009 Neolithic Central/North Austria, Hungary, Germany, 5500 - 5000 BC 47 Linearbandkeramik (LBK) Haak et al., 2005; Haak, 2006 Southwestern Spain 3500 - 3000 BC 11 ND Sampietro et al., 2007 Southwestern France 4200 BC 3 Megalithic Deguilloux et al., 2011 Chalcolithic Central/North Germany 2700 - 2400 BC 9 Corded Ware Culture (CWC) Haak et al ., 2008 Southwestern France 3030 - 2890 BC 29 ND Lacan et al., 2011 Modern Central/North Germany 2000 AD 213 - Tetzlaff e t al., 2007 Southwestern France 2000 AD 210 - Dubut et al., 2004 Abbreviations: n, number of samples; NS, not determined. Copyright © 2012 SciRes. 15  F.-X. RICAUT ET AL. et al., 2009). For purposes of comparison, we also included 423 modern Europeans from locations near the ancient DNA sam- pling sites (Dubut et al., 2004; Tetzlaff et al., 2007). Despite the few ancient DNA datasets available, we pur- posely excluded any data from studies where any doubts have been raised about ancient DNA reliability, unclear sample pro- venance, uncertain dating or cultural affiliation (particularly during the Mesolithic), or sites that fall outside the geographic range under study here (i.e., essentially central/western Europe) (Izagirre & de la Rúa, 1999; Di Benedetto et al., 2000; Chan- dler et al., 2005; Fernández Domínguez, 2005; Ermini et al., 2008; Bramanti et al., 2009; Malmström et al., 2009). Mitochondrial DNA Haplogroups The ancient and modern samples were clustered into 24 hap- logroups, 7 of which are found only in individuals living today (Table 2). We used the same haplogroup affiliations as deter- mined in the original studies wherever possible. These were checked manually against the sequence data. Due to the small number of individuals carrying some lineages, related sub- haplogroups were sometimes clustered into their root-hap- logroup to provide sufficient power for the following analyses. Table 2. Observed mtDNA haplogroup frequencies across the Mesolithic, Neo- lithic, Chalcolithic and Mode rn periods. Time Period Hg Mesolithic Neolithic Chalcolithic Modern U* 0.133 0 0.026 0.014 U2 0 0 0 0.014 U3 0 0.016 0 0.002 U4 0.133 0.033 0 0.033 U5 0 0 0.105 0.007 U5a 0.267 0.033 0 0.059 U5b 0.467 0.016 0.053 0.024 U6a1 0 0 0 0.005 U7 0 0 0 0.002 U8 0 0 0 0.007 HV 0 0.066 0.079 0.047 H 0 0.230 0.184 0.423 N 0 0.131 0 0.005 K 0 0.098 0.158 0.090 T 0 0.033 0 0.026 T1 0 0 0 0.028 T2 0 0.164 0.053 0.064 T3 0 0 0 0.002 J 0 0.033 0 0.014 J1 0 0.082 0.158 0.047 J2 0 0 0 0.012 I 0 0.016 0.026 0.021 W 0 0.033 0 0.024 X 0 0.016 0.158 0.012 ND 0 0 0 0.017 Abbreviations: Hg, haplogroup; ND, not determined. Statistics Differences in haplogroup frequencies between temporally separated European populations were determined using an in- house resampling algorithm implemented in R (http://www. r-project.org/) (code available on request). The frequency pro- bability density was inferred for each haplogroup via Monte Carlo simulation. Because sample sizes are small, ancient hap- logroup frequencies typically have large uncertainty. The fre- quency probability density explicitly accounts for this uncer- tainty. For each haplogroup, the observed frequency in the younger sample was compared with the frequency probability density of the older sample. This process yields the probability of the younger sample frequency being observed given uncer- tainty around the ancestral sample frequency. In effect, we asked whether time-adjacent haplogroup frequencies differ sig- nificantly, and therefore, whether haplogroup frequencies have changed between older and younger samples; or whether hap- logroup frequencies are statistically indistinguishable, and there- fore, show no evidence of change between adjacent time points. We emphasize that this statistical approach explicitly accounts for small sample sizes, even when these are extremely limited (e.g., Eulau). Smaller sample sizes are merely reflected by lar- ger confidence intervals. All population comparisons were performed in natural time order: Modern deriving from Chalco- lithic, Chalcolithic deriving from Neolithic, and Neolithic de- riving from Mesolithic. Results We explored mtDNA haplogroup frequencies in 532 western European individuals from four different time points: the Me- solithic, Neolithic, Chalcolithic and Modern periods. Indivi- duals carrying 24 different haplogroups were identified (Table 2). Notably, however, the distribution of these haplogroups (supplemental Tables 1-4) changed markedly across the four sampled time points (Figure 1). Figure 1. Pie charts showing mtDNA haplogroup frequencies through time. For visual clarity, haplogroups with frequencies <5% across all four samples have been collapsed into a “not determined” cate- gory. (Also see supplemental Movie 1, which illustrates dynami- cally how mtDNA haplogroup frequencies have changed through the four sampled time points). Copyright © 2012 SciRes. 16  F.-X. RICAUT ET AL. The Mesolithic/Neolithic Transition This time period is characterized by substantial change, with significant reductions in many of the U haplogroups (U*, U4, U5a and U5b), together with the sudden appearance of several new haplogroups, including H, HV, I, J, J1, K, N1, T, T2, W and X. The small size of the Mesolithic sample (n = 15) greatly reduces the statistical power of this analysis. Therefore, with the exception of haplogroup H, the sudden appearance of non- U haplogroups is not statistically significant in our analysis. However, we anticipate that the sudden arrival of these hap- logroups will be recognized as significant as additional typing increases the Mesolithic sample size and overcomes the low power of the current dataset. Indeed, while several U hap- logroups (i.e., U*, U4, U5 and U8) are traditionally associated with the Upper Paleolithic settlement of Europe, many non-U haplogroups are thought to reflect later phases of population migration and colonization (Richards et al., 2000; Bramanti et al., 2009; Malyarchuk et al., 2010; Soares et al., 2010). We anticipate that future sampling will increase the resolution of this important phase in European prehistory. The Neolithic/Bronze Age or Chalcolithic Transition This time period is much less well studied than the agricul- tural revolution. Surprisingly, we also observe considerable change in haplogroup frequencies in the lead up to the Chalco- lithic period, with significant increases in U5 and X, and statis- tically significant decreases in U3, U4, U5a, J, N1, T, T2 and W. While the Neolithic has long been recognized as a period of substantial immigration and population change, the transition to the Bronze Age is usually considered more of an indigenous affair. However, our analysis illustrates that the Chalcolithic transition was, from the perspective of haplogroup frequencies, at least as demographically disruptive as the earlier agricultural revolution. The Chalcolithic/Modern Transition Conversely, there is little evidence of change in haplogroup frequencies between the Chalcolithic and Modern periods. The only statistically significant differences are reductions in U5, J1 and X, and a substantial increase in haplogroup H, which at a frequency of 42%, dominates the western European mtDNA gene pool today. MtDNA haplogroup frequencies at these four time points are illustrated in Figure 1. However, static images like these can- not easily capture how haplogroup frequencies have changed through time. In practice, it is likely that haplogroup frequent- cies often changed quickly due to short, intense periods of mi- gration and social upheaval, rather than varying regularly through time. Nevertheless, we can see important dynamic facets of the data, including the extensive diversification of mtDNA hap- logroups between the Mesolithic and the Neolithic, and the increasing dominance of haplogroup H from the Chalcolithic to modern times. Discussion This temporal analysis suggests that two of the three transi- tional periods (Mesolithic/Neolithic and Neolithic/Bronze Age) saw substantial change in the distribution of European mtDNA haplogroup diversity. However, these changes likely had dif- ferent proximate causes. During the Mesolithic/Neolithic tran- sition, the appearance of new mtDNA haplogroups may have been caused in part by the post-glacial recolonization of Europe (e.g., H, K and T2) and/or by the expansion of Neolithic farm- ers from the east (e.g., J2, N1 and T2). In either case, there seems little doubt that Europe experienced some degree of gene flow during this time period (Haak et al., 2010; Soares et al., 2010). Conversely, the Chalcolithic transition (late Neolithic to early Bronze Age transition) seems to reflect continuity of ex- isting lineages rather than the appearance of new ones; few new haplogroups are observed in the Chalcolithic, but many existing haplogroups underwent substantial changes in allele frequency. Finally, there is surprisingly little change in haplogroup diver- sity from the Chalcolithic to the Modern period. While hap- logroup frequencies altered substantially during the Neolithic, they began to stabilize during the Chalcolithic and have re- mained relatively static ever since. These general patterns hold true when the analysis is re- peated at regional scales; e.g., diachronic analyses for southern Europe (essentially modern France, supplemental Table 2) and northern Europe (essentially modern Germany, supplemental Table 3). The main distinction being that there is little differ- ence between Neolithic and Chalcolithic samples in the south, while Neolithic and Chalcolithic populations are quite different in the north. Conversely, three haplogroups (U5b, I and X) have similar frequencies between the southern Neolithic sample and the northern Chalcolithic sample (supplemental Table 4). The northern Chalcolithic sample (Eulau, n = 9) is extremely small, which may accentuate this difference. We note, however, that the statistical procedure used here explicitly accounts for small sample sizes. No haplogroups have similar frequencies between the northern Neolithic and northern Chalcolithic sam- ples (supplemental Table 3). We would not place too much weight on this observed pattern given limitations of the avail- able data. However, this may indicate relatively more continu- ity between the southern Neolithic and northern Chalcolithic populations (i.e., a signal of northward gene flow). This would agree with some hypotheses regarding the northward spread of the Bell Beaker culture during the late Neolithic/early Chalco- lithic (Vander Linden, 2007; Guilaine et al., 2011), particularly since Eulau is one of the few sites where both cultures (Bell Beaker and Corded Ware) existed side by side (Haak et al., 2008). Conclusion This statistical analysis highlights both continuity and change in mtDNA diversity through western European prehistory. As new samples become available, we expect that this first dia- chronic synthesis of mtDNA change in Western Europe will rapidly be improved upon. Broader geographical sampling will fill in regional details, including the effects of spatially and temporally restricted cultural processes, and will likely high- light more detailed regional variability. Further, we anticipate that technological advances will soon expand this story, mov- ing it from solely a maternal perspective to instead capture paternal and biparental aspects of European genetic prehistory. For now, however, the main story is this. The Mesolithic period was characterized by low mtDNA diversity, entirely dominated by U haplogroups. The Neolithic saw the start of an ongoing decline in these haplogroups, as farmers from the east intro- duced a suite of new mtDNA haplogroups into Western Europe. Copyright © 2012 SciRes. 17  F.-X. RICAUT ET AL. Although not previously appreciated, the Chalcolithic (late Neo- lithic to early Bronze Age transition) was also a time of sub- stantial genetic change, which did more to shape current mt- DNA diversity than any subsequent time periods. However, rather than the arrival of new mtDNA lineages, the Chalcolithic is characterized by major fluctuations in the frequencies of existing haplogroups. During this period, southern Europe ex- perienced regional continuity, although there may be evidence of northward population movements as the Neolithic gave way to the Chalcolithic. Apart from the ongoing expansion of hap- logroup H, there were few major changes from the Chalcolithic period to modern times, and it appears that the basic profile of modern western European mtDNA diversity was largely estab- lished by the Bronze Age. In this context, the well-documented narratives of the historic period seem to have played out on a background of European genetic diversity that was ultimately laid down thousands of years earlier. Acknowledgements This research was supported by grants from the French De- partment of Research and the CNRS-département INNE to F.X.R., and a Rutherford Fellowship awarded to M.P.C. by the Royal Society of New Zealand (RDF-10-MAU-001). We are grateful to Patrice Gerard and Rebecca Coles for constructive comments on the manuscript. REFERENCES Ammerman, A. J., & Cavalli-Sforza, L. L. (1984). The neolithic transi- tion and the genetics of populations in Europe. Princeton, NJ: Prin- ceton University Press. Bailey, G., & Spikins, P. (2008). Mesolithic Europe. Cambridge: Cam- bridge University Press. Balaresque, P., Bowden, G. R., Adams, S. M., Leung, H. Y., King, T.E., Rosser, Z. H., Goodwin, J., Moisan, J. P., Richard, C., Millward, A., Demaine, A. G., Barbujani, G., Previdere, C., Wilson, I. J., Tyler- Smith, C., & Jobling, M. A. (2010). A p redo minantly Neolithic origin for European paternal lineages. PLoS Biology, 8, e1000285. doi:10.1371/journal.pbio.1000285 Bar-Yosef, O. (2004). East to west: Agricultural origins and dispersal into Europe. Current Anthropology, 45, S1-S3. doi:10.1086/423970 doi:10.1086/423970 Belle, E. M., Landry, P. A., & Barbujani, G. (2006). Origins and evolu- tion of the Europeans’ genome: Evidence from multiple microsatel- lite loci. Proceedings of the Royal Society B, 273, 1595-1602. doi:10.1098/rspb.2006.3494 Bellwood, R., & Renfrew, C. (2002). Examining the farming/language dispersal hypothesis. Cambridge: McDonald Institute for Archaeo- logical Research. Bramanti, B., Thomas, M. G., Haak, W., Unterlaender, M., Jores, P., Tambets, K., Antanaitis-Jacobs, I., Haidle, M. N., Jankauskas, R., Kind, C. J., Lueth, F., Terberger, T., Hiller, J., Matsumura, S., & Forster, P. (2009). Genetic discontinuity between local hunter-gath- erers and central Europe’s first farmers. Science, 326, 137-140. doi:10.1126/science.1176869 Bocquet-Appel, J. P., N aji, S., Vander Linden, M., & Kozlowski, J. K. (2009). Detection of diffusion and contact zones of early farming in Europe from the space-time distribution of 14C dates. Journal of Archaeological Science, 36, 807- 820. doi:10.1016/j.jas.2008.11.004 Burger, J., Kirchner, M., Bramanti, B., Haak, W., & Thomas, M. G. (2007). Absence of the lactase-persistence-associated allele in early Neolithic Europeans. Proceedings of the National Academy of Sci- ences USA, 104, 3736-3741. doi:10.1073/pnas.0607187104 Cavalli-Sforza, L. L., Menozzi, P., & Piazza, A. (1994). The history and geography of human genes. Princeton, NJ: Princeton University Press. Chandler, H., Sykes, B., & Zilhão, J. (2005). Using ancient DNA to examine genetic continuity at the Mesolithic-Neolithic transition in Portugal. In P. Arias, R. Ontanon, & A. Garcia-Monco (eds.), Actas del III Congreso del Neolitico en la Peninsula Iberica, 781-786. Chikhi, L., Destro-B isol, G., Berto rell e, G., Pascali, V, & Ba rbu jani, G. (1998). Clines of nuclear DNA markers suggest a largely Neolithic ancestry of the European gene pool. Proceedings of the National Academy of Sciences USA, 95, 9053-9058. doi:10.1073/pnas.95.15.9053 Crubézy, E., Bruzek, J., & Guilaine, J. (2002). The transition to agri- culture in Europe: An anthropobiological perspective. Biennial Book of the European Anthropolo gi c a l Association, 2, 93-110. Deguilloux, M. F., Soler, L., Pemonge, M. H., Scarre, C., Joussaume, R., & Laporte, L. (2011). News from the west: Ancient DNA from a French megalithic burial chamber. American Journal of Physical Anthropology, 144, 108-118. doi:10.1002/ajpa.21376 Di Benedetto, G., Nasidze, I.S., Stenico, M., Nigro, L., Krings, M., Lanzinger, M., Vigilant, L., Stoneking, M., Pääbo, S., & Barbujani, G. (2000). Mitochondrial DNA sequences in prehistoric human re- mains from the Alps. European Journal of Human Genetics, 8, 669- 677. doi:10.1038/sj.ejhg.5200514 Dubut, V., Chollet, L., Murail, P., Cartault, F., Béraud-Colomb, E., Serre, M., & Mogentale-Profizi, N. (2004). MtDNA polymorphisms in five French groups: Importance of regional sampling. European Journal of Human Genetics, 12, 293-300. doi:10.1038/sj.ejhg.5201145 Ermini, L., Olivieri, C., Rizzi, E., Corti, G., Bonnal, R., Soares, P., Luciani, S., Marota, I., De Bellis, G., Richards, M.B., & Rollo, F. (2008). The complete mitochondrial genome sequence of the Tyro- lean Iceman. Current Biology, 18, 1687-1693. doi:10.1016/j.cub.2008.09.028 Fernández Domínguez, E. (2005). Polimorfismos de DNA mitocondrial en poblaciones antiguas de la cuenca mediterránea. Ph.D. Thesis, Barcelona: University of Barcelo na. Gronenborn, D. (1999). A variation on the basic theme: The transition to farming in southern central Europe. Journal of World Prehistory, 2, 123-210. doi:10.1023/A:1022374312372 Guilaine, J. (1997). Résumé des Cours et travaux 1996-1997. Paris: Collège de France. Guilaine, J. (2003). De la Vague à la tombe. la conquête néolithique de la méditerranée (8000-2000 av. JC). Paris: Seuil Press. Guilaine, J., & Manen, C. (2007). From Mesolithic to early Neolithic in the western Mediterranean. In A. Whittle, & V. Cummings (eds.), Going over: The mesolithic-neolithic transition in north-west Europe (pp. 21-51). Lon do n: Pr oc ee di ng s o f the British Academy. Guilaine, J., Tusa, S., & Veneroso, P. (2011). La Sicile et l’Europe Campaniforme. Toulouse: Archives d’Ecologie Prehistorique. Haak, W. (2006). Populationsgenetik der ersten Bauern Mitteleuropas. Eine aDNA-Studie an neolithischem Skelettmaterial. Ph.D. Thesis, Mainz: University of Mainz. Haak, W., Balanovsky, O., Sanchez, J. J., Koshel, S., Zaporozhchenko, V., Adler, C. J., D er Sark issian, C. S., Brandt, G., Sch w arz, C., Nick- lisch, N., Dresely, V., Fritsch, B., Balanovska, E., Villems, R., Meller, H., Alt, K. W., Cooper, A., & the Genographic Consortium. (2010). Ancient DNA from European early Neolithic farmers reveals their near eastern affinities. PLoS Biology, 9, e1000536. doi:10.1371/journal.pbio.1000536 Haak, W., Brandt, G., de Jong, H. N., Meyer, C., Ganslmeier, R., Heyd, V., Hawkesworth, C., Pike, A. W., Meller, H., & Alt, K. W. (2008). Ancient DNA, strontium isotopes, and osteological analysis shed light on social and kinship organization of the Later Stone Age. Pro- ceedings of the National Academy of Sciences USA, 105, 18226- 18231. doi:10.1073/pnas.0807592105 Haak, W., Forster, P., Bramanti, B., Matsu mura, S. , Brand t, G., Tan zer, M., Villems, R., Renfrew, C., Gronenborn, D., Alt, K. W., & Burger, J. (2005). Ancient DNA from the first European farmers in 7500- year-old Neolithic sites. Science, 310, 1016-1018. Itan, Y., Powell, A., Beaumont, M. A., Burger, J., & Thomas, M. G. (2009). The origins of lactase persistence in Europe. PLoS Computa- Copyright © 2012 SciRes. 18  F.-X. RICAUT ET AL. Copyright © 2012 SciRes. 19 tional Biology, 5, e1000491. doi:10.1371/journal.pcbi.1000491 Izagirre, N., & de la Rúa, C. (1999). Absence of mtDNA haplogroup V in ancient Basques, American Journal of Human Genetics, 65, 199-207. doi:10.1086/302442 Lacan, M., Keyser, C., Ricaut, F. X., Brucato, N., Duranthon, F., Gui- laine, J., Crubézy, E., & Ludes, B. (2011). Ancient DNA revealed male diffusion through the Neolithic Mediterranean route. Proceed- ings of the National Academy of Sciences USA, 108, 9788- 9791. doi:10.1073/pnas.1100723108 Malmström, H., Gilbert, M. T. P., Thomas, M. G., Branström, M., Stora, J., Molnar, P., Andersen, P. K., Bendixen, C., Holmlund, G., Götherström, A., Willerslev, E. (2009). Ancient DNA reveals lack of continuity between Neolithic hunter-gatherers and contemporary Scandinavians. Current Biolo gy , 19, 1758-1762. doi:10.1016/j.cub.2009.09.017 Malyarchuk, B., Derenko, M., Grzybowski, T., Perkova, M., Rogalla, U., Vanecek, T., & Tsybovsky, I. (2010). The peopling of Europe from the mitochondrial haplogroup U5 perspective. PLoS ONE, 5, e10285. doi:10.1371/journal.pone.0010285 Novembre, J., Johnson, T., Bryc, K., Kutalik, Z., Boyko, A. R., Auton, A., Indap, A., King, K. S., Bergmann, S., Nelson, M. R., Stephens, M., & Bustamante, C. D. (2008). Genes mirror geography within Europe. Nature, 456, 98-101. doi:10.1038/nature07331 Palanichamy, M. G., Zhang, C. L., Mitra, B., Malyarchuk, B., Derenko, M., Chaudhuri, T. K., & Zhang, Y. P. (2010). Mitochondrial hap- logroup N1a phylogeography, with implication to the origin of Euro- pean farmers. BMC Evolutionary Biology, 10, 304. doi:10.1186/1471-2148-10-304 Pinhasi, R., Fort, J., & Ammerman, A. J. (2005). Tracing the origin and spread of agriculture in Europe. PL oS Biology, 3, e410. doi:10.1371/journal.pbio.0030410 Pinhasi, R., & von Cramon-Taubadel, N. (2009). Craniometric data supports demic diffusion model for the spread of agriculture into Europe. PLoS ONE, 4, e6747. doi:10.1371/journal.pone.0006747 Renfrew, C., & Boyle, K. (2000). Archaeogenetics: DNA and the population prehistory of Europe. Cambridge: McDonald Institute for Archaeological Research. Richards, M. (2003). The Neolithic invasion of Europe. Annual Review of Anthropology, 32, 135-162. doi:10.1146/annurev.anthro.32.061002.093207 Richards, M. V., Macauley, V., Hickey, E., Vega, E., Sykes, B., Guida, V., Rengo, C., Sellitto, D., Cruciani, F., Kivisild, T., Villems, R., Thomas, M., Rychkov, S., Rychkov, O., Rychkov, Y., Gölge, M., Dimitrov, D., Hill, E., Bradley, D., Romano, V., Calì, F., Vona, G., Demaine, A., Papiha, S., T riantaphyl lidis, C., Stefanescu, G., Hat ina, J., Belledi, M., Di Rienzo, A., Novelletto, A., Oppenheim, A., Nørby, S., Al-Zaheri, N., Santachiara-Benerecetti, S., Scozari, R., Torroni, A., & Bandelt, H. J. (2000). Tracing European founder lineages in the near eastern mtDNA pool. American Journal of Human Genetics, 67, 1251-1276. Rowley-Conwy, P. (2011). Westward Ho! The spread of agriculture from Central Eu rope to the Atlantic. Current Anthropology, 52, S431- S451. doi:10.1086/658368 Sampietro, M.L., Lao, O., Caramelli, D., Lari, M., Pou, R., Marti, M., Bertranpetit, J., Lalueza-Fox, C. (2007). Palaeogenetic evidence supports a dual model of Neolithic spreading into Europe. Proceed- ings of the Royal Society B, 274, 2161-2167. doi:10.1098/rspb.2007.0465 Sherratt, A. (1981). Plough and pastoralism: aspects of the secondary products revolution. In I. Hodder, G. Isaac, & N. Hammond (eds.), Pattern of the past: Studies in honour of David Clarke (pp. 261-305). Cambridge: Cambridge University Press. Soares, P., Achilli, A., Se mino, O., Davies, W., Macaulay, V., Bandelt, H.-J., Torroni, A., & Richards, M. B. (2010). The archaeogenetics of Europe. Current Biology, 20, R174-R183. doi:10.1016/j.cub.2009.11.054 Tetzlaff, S., Brandstätter, A., Wegener, R., Parson, W., & Weirich, V. (2007). Mitochondrial DNA population data of HVS-I and HVS-II sequences from a northeast German sample. Forensic Science Inter- national, 172, 218-224. doi:10.1016/j.forsciint.2006.12.016 Vander Linden, M. (2007). What linked the Bell Beakers in third mil- lennium BC Europe? Antiquity, 81, 343-352. Von Cramon-Taubadel, N., & Pinhasi, R. (2011). Craniometric data support a mosaic model of demic and cultural Neolithic diffusion to outlying regions of Europe. Proceedings of the Royal Society B, 278, 2874-2880. doi:10.1098/rspb.2010.2678 Whittle, A. W. R. (1996). Europe in the Neolithic: The Creation of New Worlds. Cambridge: Cambridge University Press. Whittle, A., & Cummings, V. (2007). Going over: The mesolithic-neo- lithic transition in north-west Europe. Oxford: Oxford University Press. doi:10.5871/bacad/9780197264140.001.0001 Zvelebil, M. (2004) Who were we 6000 years ago? In search of prehis- toric identities. In M. Jones (ed.), Traces of ancestry: Studies in honour of Colin Renfrew. Cambridge: McDonald Institute Mono- graphs. Supplemental Materials Supplemental Movie 1. http://youtu.be/C5nijHibX3c Dynamic illustration of mtDNA haplogroup frequencies changing through the four sampled time points. (supplemental Tables 1-4 see the following pages) 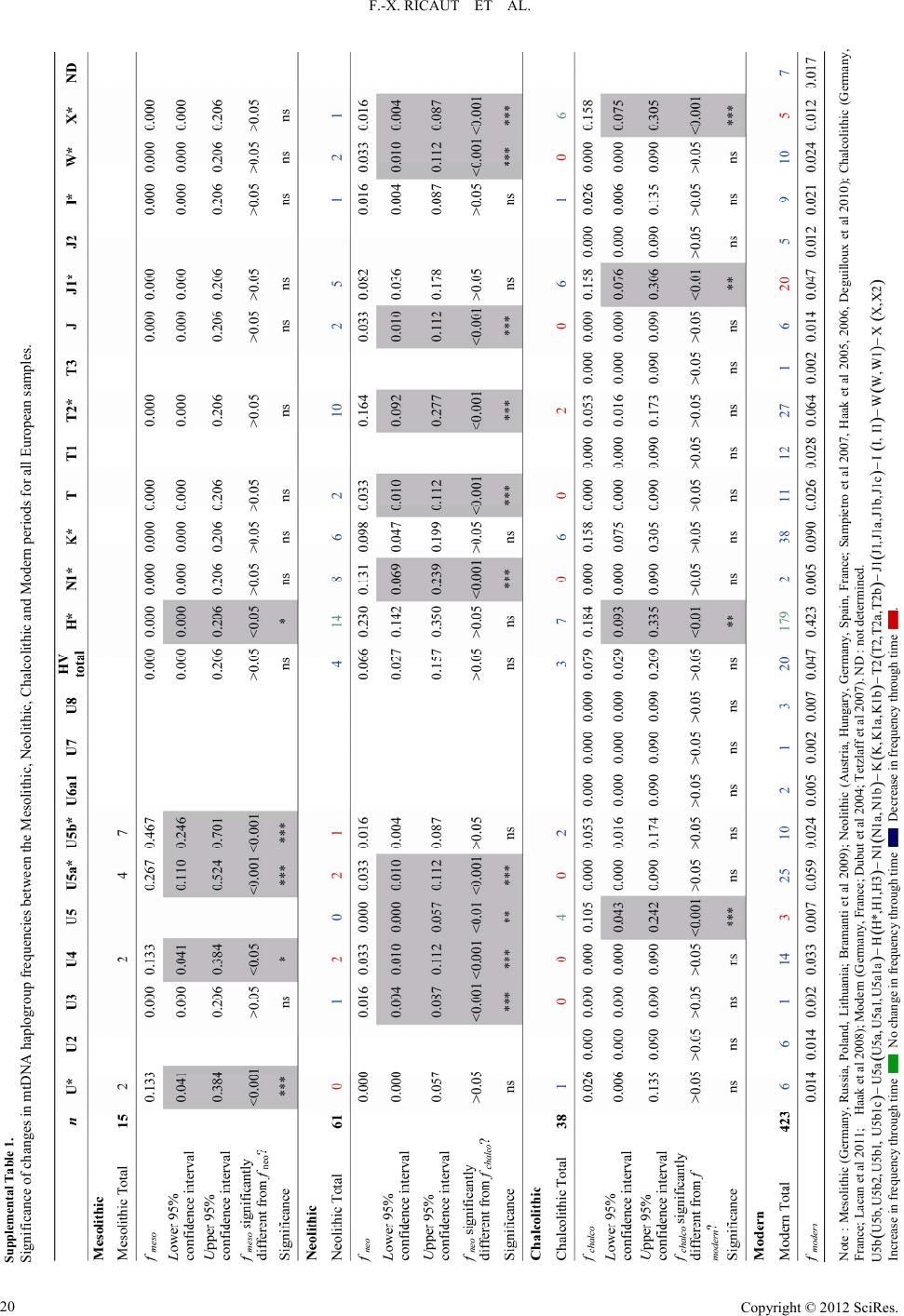 F.-X. RICAUT ET AL. Note : Mesolithic (Germany, Russia, Poland, Lithuania; Bramanti et al 2009); Neolithic (Austria, Hungary, Germany, Spain, France; Sampietro et al 2007, Haak et al 2005, 2006, Deguilloux et al 2010); Chalcolithi c (Germany, France; Lacan et al 2011; Haak e t al 2008); Modern (Germany, France; Dubut et al 2004; Tetzlaff et al 2007). ND : not determined. Increase i n frequency through tim e No change in freque ncy through time Decrease in frequency through time . Supplemental Table 1. Significance of changes in mtDNA haplogroup frequencies between the Mesolithic, Neolithic, Chalcolithic and Modern periods for all European samples. Copyright © 2012 SciRes. 20  F.-X. RICAUT ET AL. ote: Mesolithic (Germany, Russia, Poland, Lithuania; Bramanti et al 2009); Neolithic (Austria, Hungary, Germany, Spain, France; Sampietro et al 2007, Haak et al 2005, 2006, Deguilloux et al 2010); Chalcolithic (Germany, France; Lacan et al 2011; Haa k et al 2008); Modern (Germany, France; Dubut et al 2004; Tetzlaff et al 2007). ND : not determined. Increase i n frequency through tim e No change in freque ncy through time Decrease in frequency through time . Supplemental Table 2. Significance of changes in mtDNA haplogroup frequencies between the Mesolithic, Neolithic, Ch alcolithic and Modern periods in Southern Europe. Copyright © 2012 SciRes. 21  F.-X. RICAUT ET AL. Note : Mesolithic (Germany, Russia, Poland, Lithuania; Bramanti et al 2009); Neolithic (Austria, Hungary, Germany, Spain, France; Sampietro et al 2007, Haak et al 2005, 2006, Deguilloux et al 2010); Chalcolithic (G ermany, France; Lacan et al 2011; Haa k et al 2008); Modern (Germany, France; Dubut et al 2004; Tetzlaff et al 2007). ND : not determined. Increase i n frequency through tim e No change in frequency through time Decrease in frequency through time . Supplemental Table 3. Significance of changes in mtDNA haplogroup frequencies between the Mesolithic, Neolithic, Ch alcolithic and Modern periods in Northern Europe. Copyright © 2012 SciRes. 22  F.-X. RICAUT ET AL. Copyright © 2012 SciRes. 23 Note: Mesolithic (Germany, Russia, Poland, Lithuania; Bramanti et al 2009); Neolithic (Austria, Hungary, Germany, Spain, France; Sampietro et al 2007, Haak et al 2005, 2006, Deguilloux et al 2010); Chalcolithic (Germany, France; Lacan et al 2011; Haak e t al 2008); Modern (Germany, France; Dubut et al 2004; Tetzlaff et al 2007). ND : not determined. crease in frequency through time Supplemental Table 4. Significance of changes in mtDNA haplogroup frequencies between the Neolithic (Southern Europe) and the Chalcolithic (Northern Europe).
|