Applied Mathematics
Vol.4 No.1(2013), Article ID:27196,6 pages DOI:10.4236/am.2013.41012
Optimization of Penicillin G Acylase Immobilization Process by Surface Response Methodology Using Central Composite Design
1Pilot Biotechnology Department, Pasteur Institute of Iran, Tehran, Iran
2Physiology and Pharmacology Department, Pasteur Institute of Iran, Tehran, Iran
Email: *dnsa@pasteur.ac.ir
Received August 11, 2012; revised October 24, 2012; accepted November 1, 2012
Keywords: Quadratic Model; Significant; Enzyme Concentration; Temperature of the Adsorption
ABSTRACT
Penicillin G acylase was immobilized onto superparamagnetite iron oxide nanoparticles employing response surface methodology through central composite design. Polynomial quadratic model was selected as a model. The value of the determination coefficient (R2) calculated from the quadratic regression model was 0.845, while the value of the adjusted (R2) was 0.74. The regression analysis of the data showed that the quadratic model selected were appropriate thereby enzyme concentration (A), reaction temperature (D), enzyme concentration* reaction temperature (AD), quadratics enzyme concentration (A2) and reaction temperature (D2) were found to be significant factors in immobilization process of penicillin G acylase.
1. Introduction
Penicillin G acylase is a biocatalyst produced by bacteria as well as fungi in order to acylate/deacylate penicillin G to its constituent 6-aminopenicilanic acid and phenyl acetic acid or vice versa which is termed as hydrolysis (H) and synthesis (S) respectively, (Figure 1). Due to hydrolytic and synthetic ability of penicillin G acylase and importance of semi synthetic antibiotics, the enzyme has profound uses in related industry.
Penicillin acylase in its soluble form is unstable, cannot be separated from the reaction mixture easily which will add to the production cost of the final product thus such a commercially, industrially important biocatalyst has been immobilized by different techniques employing various supports by many investigators [1-7]. Each technique or method has its own limitation and disadvantages where immobilization method can alter kinetic properties of immobilized enzyme as compared to its soluble counterpart. The changes can be brought about by the supports or the reagents employed. Investigators are trying to employ technique and reagents which will improve the kinetic properties of the enzyme under immobilization process as compared to its soluble counterpart [7-11]. In this article, we have attempted to optimize the immobilization process of penicillin G acylase onto super para magnetite iron oxide nanoparticles by considering factors such as, enzyme concentration, weight of nanoparticle, concentration of polyethyleneimine, temperature and time of reaction using surface response methodology. Response surface methodology (RSM) is a collection of statistical and mathematical techniques useful for developing, improving, and optimizing processes. The applications of RSM are in industry, biological & chemical sciences, pharmaceutical, physical and engineering sciences. Further the most extensive applications of RSM are in the particular situations where several input variables potentially influence some performance measure or quality characteristic of the process. In RSM, the main effects and interactions between various factors, each at different levels can be simultaneously studied [12].
2. Materials and Methods
2.1. Materials
Penicillin G acylase, 6-aminopenicilanic acid (6-APA), polyethyleneimine, p-dimethylaminobenzaldehyde (pDMBA), were obtained from Sigma, USA. Benzyl penicillin was procured from local market; other reagents used were of analytical grade.
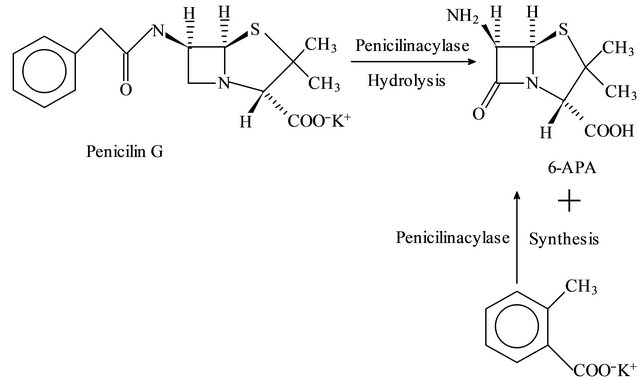
Figure 1. Penicillin G acylase catalyzed reaction (S/H).
2.2. Penicillin G Acylase Activity
Penicillin acylase activity was determined as reported by Norouzian et al. [5], measuring the amount of 6-aminopenicilanic acid (6-APA) formed at 35˚C ± 1˚C, employing 2% (w/v) benzyl penicillin prepared in 0.1 M phosphate buffer pH 7.5. The formed 6-APA was estimated with p-dimethylaminobenzaldehyde (p-DMBA). One unit of enzyme activity was defined as the amount of enzyme catalyzing the hydrolysis of µmol of penicillin G in 1 min under the above assay condition.
2.3. Immobilization of Penicillin G Acylase
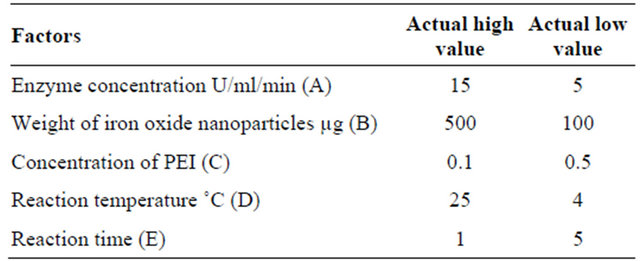
Penicillin G acylase was immobilized onto synthesized superparamagnetite iron oxide nanoparticles. The factors could influence the immobilization process were considered to be; enzyme concentration (A: U/ml/min) weight of iron oxide nanoparticles (B: µg), concentration of polyethyleneimine (C: %), reaction temperature (D: ˚C) and reaction time (E: h). The immobilization process was optimized by surface response methodology (SRM) through central composite design (CCD) employing Ease State program, version 6. Furthermore, central composite design contains an imbedded factorial or fractional factorial design with center point that is augmented with a group of start points that allow estimation of the curvature. The mathematical representation of RSM in this study is poly nominal quadratic second order model. Totally 50 experiments were designed by providing the actual low and high values by considering five factors which could influence the optimization process. The above mentioned factors were used to construct the experiment’s matrix (Tables 1 and 2).
2.4. Statistical Analysis
Data obtained through the experimental matrix are computed for the determinations of regression coefficient of
Table 1. The actual high and low values of the factors considered to immobilize penicillin G acylase.
the second order multiple regression model. The analysis of regression and variance was performed by Design Expert (Ease State Version 6.0).
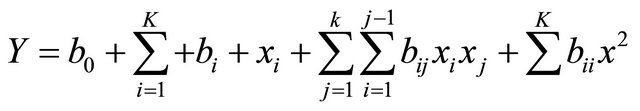
where Y is the predicted response variable, b0, bi, bii and bij are regression coefficient of the model, xi, xj represent the independent variables in the form of actual value.
3. Results and Discussion
Table 1 shows the actual value at high and low level. Through Table 1 the experimental matrix was designed employing 2 level factorial design. The response (Y) was expressed in terms of units of enzyme adsorbed onto iron oxide nanoparticles. The responses obtained revealed that experiments no. 4, 12, 17 and 38 are yielded highest immobilization rate. It is noticed that adsorption of the enzyme onto the support has taken place at 4˚C and in experiments 6 and 21, the amount of enzyme is at minimum level, but the adsorption of the enzyme has taken place at 4˚C with almost 50% adsorption equalizing to that of high value of enzyme concentration showing enzyme concentration and reaction temperature are important factors in immobilization process of penicillin G acylase (Table 3, Figures 2 and 3).
Table 2. The experimental matrix designed by Ease State version 6, bearing the actual factors at high and low level where the response is expressed in terms of unit/ml.
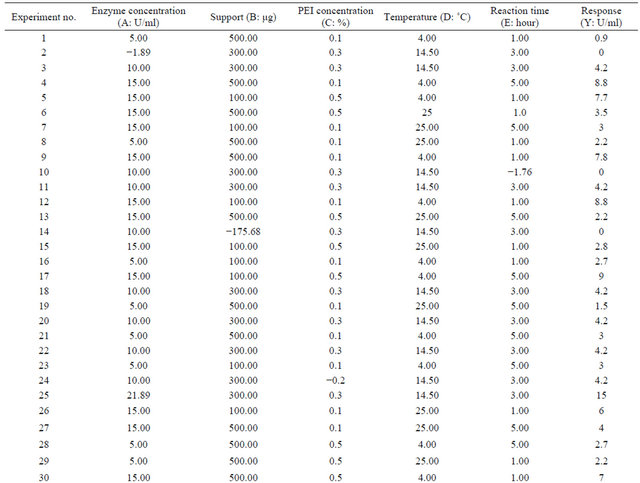
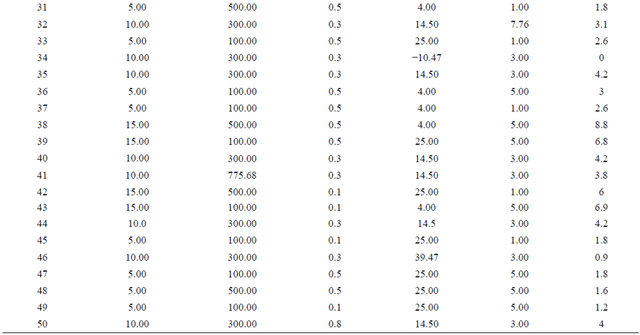
Table 3. Regression analysis of variance for RSM (ANOVA).
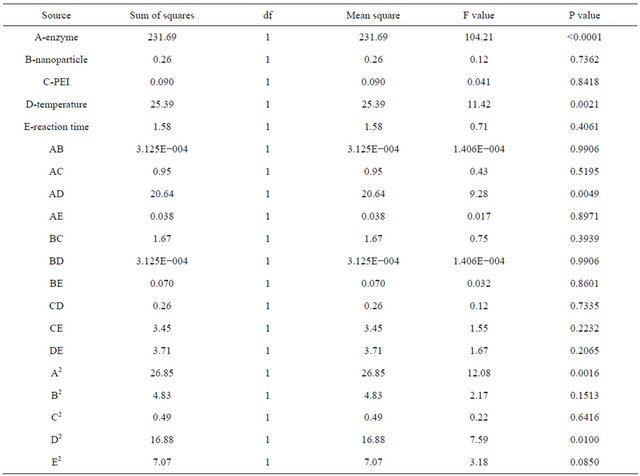
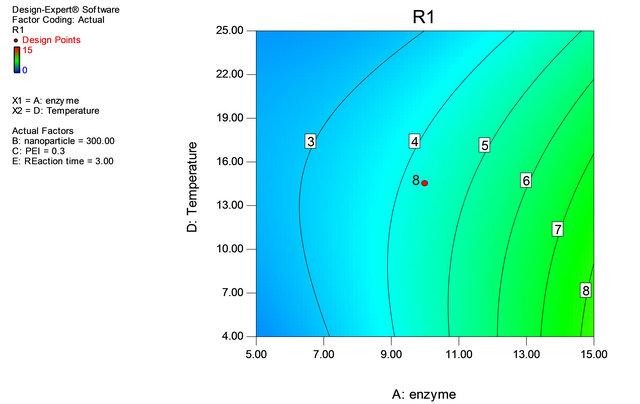
Figure 2. Contour plot of enzyme and temperature.
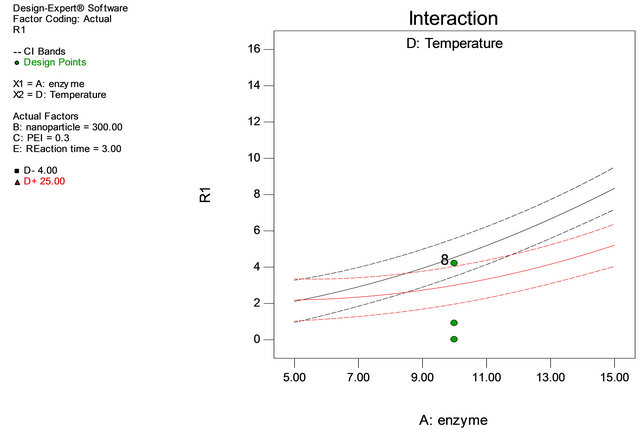
Figure 3. Interaction of enzyme and temperature.
Since response surface methodology is a statistical technique employed to optimize the processes where several factors are involved. This methodology can be employed to optimize bioprocesses and the obtained model can be validated in order to follow up the accuracy of the model [13-16]. The statistical significance of regression equation was checked by F-test, and the analysis of variance (ANOVA) for response surface quadratic polynomial model done by software (Design-Expert) is shown in Table 3. The P value of 0.0009 for response surface quadratic model shows significance of the model thus it was selected to study further. The value of the determination coefficient (R2) calculated from the quadratic regression model was 0.845, while the value of the adjusted (R2) was 0.74 that almost indicates fairly high degree of correlation between the observed and predicted values. All these results suggested that the model, as evidenced by the calculated F value (7.95) and low probability value (P < 0.0001), was adequate for the prediction of the immobilization of penicillin G acylase within the variable range employed. The lack-of-fit test measures failure of the model to represent the data in the experimental domain at points which are not included in the regression. As shown in Table 3, F value (2.93) of lack-of-fit implied that it was not significant relative to the pure error, which indicated the model equation was adequate for predicting the yield of immobilization process under any combination of values of the variables. The P value was used as a tool to check the significance of each coefficient, which in turn may indicate the pattern of the interactions among the variables. The smaller the P value, the more significant the corresponding coefficient. The whole design consisted of 50 experimental points, which included 32 factorial points, 10 axial and 8 central points. By employing multiple regression analysis on the experimental data, the response Y for the yield i.e. the amount of penicillin G acylase immobilized on to super paramagnetic iron oxide nanoparticles can be obtained by the following second-order polynomial equation.

As it can be observed from the Table 3 the P values for factors like A, D, AD, A2 and D2 are < 0.0001, 0.001, 0.0049 and 0.0016 respectively showing the significance of the factors involved in immobilization process of penicillin G acylase.
The quadratic model was validated by performing the immobilization process under optimal conditions and the yield of penicillin acylase was comparable to that of calculated by the model thereby the selected model was appropirate.In this way penicillin G acylase (industrially and commercially important biocatalyst) could be immobilized through optimization by surface response methodology
4. Acknowledgements
The corresponding author thereby acknowledges the grant given by Iran National Science Foundation: INSF to carry out this research project.
REFERENCES
- A. S. M. Chong and X. S. Zhao, “Design of Large Pore Mesoporous Material for Immobilization of Penicillin G Acylase Biocatalysts,” Catalysis Today, Vol. 93-95, No. 1, 2004, pp. 293-299. doi:10.1016/j.cattod.2004.06.064
- D. A. Cecchini, I. Sera, D. Ubiali, M. Terreni and A. M. Albertini, “New Active Site Oriented Glyoxyl-Agarose Derivatives of E. coli Penicillin G Acylase,” BMC Biotechnology, Vol. 7, No. 1, 2007, pp. 54-67. doi:10.1186/1472-6750-7-54
- F. Scaramozzino, I. Estruch, P. Rossolillo, M. Terreni and A. M. Albertini, “Improvement of Catalytic Properties of E. coli Penicillin G Acylase on Glyoxyl Agarose by Addition of a Six Amino Acid Tag,” Journal of Applied and Environmental Microbiology, Vol. 71, No. 12, 2005, pp. 8937-8940.
- W. S. Adriano, E. H. C. Filho, J. A. Silva, R. L. C. Giordano and R. B. Gonçalves, “Optimization of Penicillin G Acylase Multipoint Immobilization onto Glutaraldehyde—Chitosan Beads,” Biotechnology and Applied Biochemistry, Vol. 41, No. 3, 2005, pp. 201-207. doi:10.1042/BA20040061
- D. Norouzian, S. Javadpour, N. Moazami and A. Akbarzadeh, “Immobilization of Whole Cell Penicillin G Acylase in Gelatin Open Pore Matrix,” Enzyme and Microbial Technology, Vol. 30, No. 1, 2002, pp. 26-29.
- J. Zhao, Y. Wang, G. Luo and S. Zhu, “Immobilization of Penicillin G Acylase on Macro-Mesoporous Silica Sphers,” Bioresource Technology, Vol. 102, No. 2, 2011, pp. 529- 535. doi:10.1016/j.biortech.2010.09.076
- R. V. Bahulekar, A. A. Prabhune, H. SivaRaman and S. Ponrathnams, “Immobilization of Penicillin G Acylase on Functionalized Macroporous Polymer Beads,” Polymer, Vol. 34, No. 1, 1993, pp. 163-166. doi:10.1016/0032-3861(93)90300-Y
- J. Huang, X. Lih, Y. Zheng, Y. Zhong, R. Zhao, X. Gao and H. Yan, “Immobilization of of Penicillin G Acylase on Poly[(Glycidyl Methacrylate)-co-(Glycerol Monomethacrylate)]—Grafted Magnetic Microspheres,” Macromolecular Bioscience, Vol. 8, No. 6, 2008, pp. 508-515.
- F. N. Crespilho, R. M. Lost, S. A. Tavarian and O. N. Oliveira, “Enzyme Immobilization on Ag Nanoparticles/Polyaniline Nanocomposite,” Biosensors and Bioelectronics, Vol. 24, No. 10, 2009, pp. 3072-3077. doi:10.1016/j.bios.2009.03.026
- B. Gao, X. Wang and Y. Shen, “Studies on Characters of Immobilizing Penicillin G Acylase on a Novel Composite Support,” Biochemical Engineering Journal, Vol. 28, No. 2, 2006, pp. 140-147. doi:10.1016/j.bej.2005.10.007
- G. K. Kouassi, J. Lrudayaraj and G. McCarty, “Examination of Cholesterol Oxidase Attachment to Magnetic Nanoparticles,” Journal of Nanobiotechnology, Vol. 3, No. 1, 2005, pp. 1-9. doi:10.1186/1477-3155-3-1
- R. V. Lenth, “Response-Surface Methods in R, Using rsm,” Journal of Statistical Software, Vol. 32, No. 7, 2009, pp. 1-17.
- Y. J. Hung, C.-C. Peng, J. T. C. Tzen, M.-J. Chen and J.-R. Liu, “Immobilization of Neocallimastix patriciarum Xylanase on Artificial Oil Bodies and Statistical Optimization of Enzyme Activity,” Bioresource Technology, Vol. 99, No. 18, 2008, pp. 8662-8666. doi:10.1016/j.biortech.2008.04.017
- A. Gűvenç, N. Kapucu, H. Kapucu, Ö. Aydoğan and Ü. Mehmetoğlu, “Enzymatic Esterification of Isoamyl Alcohol Obtained from Fusel Oil: Optimization by Response Surface Methodology,” Enzyme and Microbial Technology, Vol. 40, No. 4, 2007, pp. 778-785. doi:10.1016/j.enzmictec.2006.06.010
- N. Peatciyammal, B. Balachandar, M. D. Kumar, K. Tamilarasan and C. Muthakumaran, “Stastical Optimization of Enzymatic Hydrolysis of Potato (Solanum tuberosum) Starch by Immobilized Enzyme,” International Journal of Chemical and Biological Engineering, Vol. 3, No. 3, 2010, pp. 124-128.
- Q. Zhao, J. F. Kennedy, X. Wang, X. Yuan, B. Zhao, Y. Peng and Y. Huang, “Optimization of Ultrasonic Circulation Extraction of Polysaccharide from Asparagus Officinalis Using Response Surface Methodology,” International Journal of Biological Macromolecules, Vol. 49, No. 2, 2011, pp. 181-187. doi:10.1016/j.ijbiomac.04.012
NOTES
*Corresponding author.

