Journal of Biophysical Chemistry
Vol. 3 No. 4 (2012) , Article ID: 25063 , 7 pages DOI:10.4236/jbpc.2012.34042
A single base permutation in any loop of a folded intramolecular quadruplex influences its structure and stability
![]()
1Department of Biology, Texas Woman’s University, Denton, USA
2Department of Chemistry and Biochemistry, Texas Woman’s University, Denton, USA; *Corresponding Author: rsheardy@twu.edu
Received 16 August 2012; revised 21 September 2012; accepted 29 September 2012
Keywords: Cell DNA Quadruplexes; Sequence Context; Circular Dichroism; Structure; Stability
ABSTRACT
The human telomere sequence (TTAGGG)4 folds into an unusual conformation possessing three G-tetrads linked by TTA loops. The first loop is a propeller loop while the second and third loops are transverse loops. Using Circular Dichroism (CD) spectroscopy, we have investigated the effect of sequence context on the structures and stabilities of intramolecular G-quadruplexes related to the human telomere sequence by considering all permutations of T and A within the loops. The results indicate that changing only one base in any one loop can have a dramatic effect on the conformation of the quadruplex as well as its melting temperature, Tm. Thus, each sequence studied has a unique CD spectrum and Tm. In general, variants with a modified second loop are the most stable while the wild type sequence is the least stable. The observed difference in CD spectra and melting temperature are discussed in terms of base stacking within the loop and stacking of the loop bases with adjacent G-tetrads.
1. INTRODUCTION
Our research group has been investigating the structure and stability of unusual DNA conformations for many years. Whether we are looking at B - Z junctions [1], DNA hairpins [2,3] or DNA quadruplexes [4], one observation always presents itself—DNA conformation is regulated by sequence and environment. For a DNA oligomer such as (XXXYYY)z, where X is A, C and/or T, Y is G or T and z = 1, 2, 4 or more, the conformation of any secondary structure formed is highly dependent on the identities of X and Y, the number of repeats z and the conditions under which the DNA is prepared (temperature, pH, counterions present and their concentrations, etc). For example, consider the complex equilibria indicated in Scheme 1 for (TTAGGG)4 and its Watson-Crick complement (CCCTAA)4. At 95˚C in either Na+ or K+, (TTAGGG)4 exists as a single stranded structure. Upon cooling, it can form an intramolecular quadruplex whose conformation depends upon whether the counterion is Na+ or K+ [5-8]. Further, the Na+ form can be converted to the K+ by titration with KCl and vice versa [7]. Similarly, (CCCTAA)4 at 95˚C, pH 7.0 in the presence of Na+ or K+ will form the so called i-motif upon cooling to 25˚C and lowering the pH to 5.5 [9-13]. Finally, cooling an equimolar mixture of (TTAGGG)4 and (CCCTAA)4 from 95˚C to 25˚C, in either Na+ or K+, will result in the formation of a normal Watson-Crick double helical conformation.
In consideration of the unfolding of the K+ form of the quadruplex formed from (TTAGGG)4, the human telomere sequence, we have previously demonstrated that the unfolding proceeds via a three state mechanism and is reversible [14]. The proposed conformation of the folded structure is depicted in Figure 1" target="_self"> Figure 1. This structure is
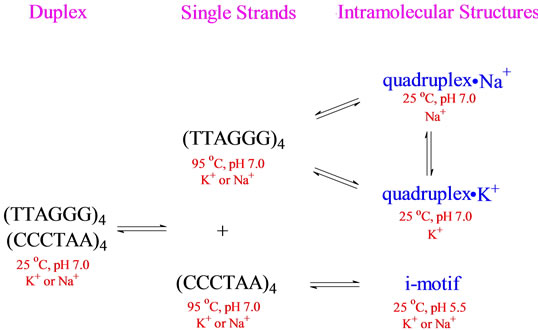
Scheme 1.Schematic representation of the equilibria involved with the human telomere sequence and its Watson-Crick complement.
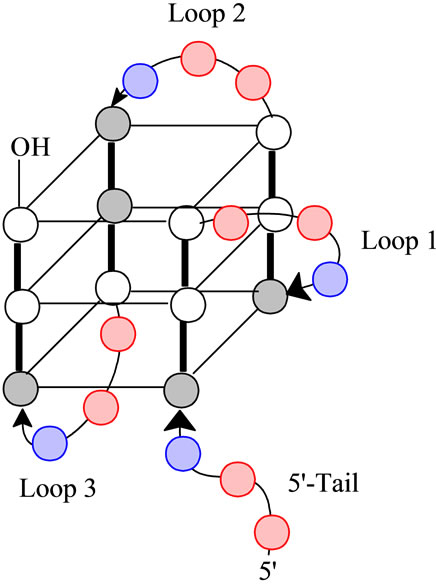
Figure 1. A schematic representation of the folded structure for (TTAGGG)4. The gray circles are anti-guanines, the white circles are syn-guanines, the red circles are thymidine and the blue circles are adenine. Note the designations of Loops 1 through 3 all of which have the sequence of -(TTA)- as well as the 5’-Tail. Loop 1 is described as a propeller loop while Loops 2 and 3 are lateral loops [17].
characterized by having three G-tetrads linked by Loop 1 (a propeller loop) and Loops 2 and 3 (transverse loops). Further, we should note the presence of a 5’-Tail. For the wild type sequence, i.e., the human telomere sequence, all loops and the tail have a sequence of 5’- TTA-3’. Further, the tetrads have a mixture of both syn and antiguanines. Ultimately, the loops determine the strand orientations, which for this conformation, is a mixture of parallel and antiparallel. One possible mechanism for the unfolding is through opening of the quadruplex at Loop 2; i.e., Loop 2 acts as a hinge. This would require breaking six hydrogen bonds on the front face and six hydrogen bonds on the back face of the quadruplex with concomitant release of the sequestered K+ ions leading to a double hairpin which unfolds in the second step of the transition [4].
It is well established that the folding motif for any DNA intramolecular quadruplex is highly regulated by the types of loops present [15-20]. The types of loops present will determine: 1) the syn or anti conformations of the guanine bases; 2) the strand orientations (i.e., parallel, antiparallel or mixed parallel-antiparallel); and 3) the orientation of the hydrogen bonds within one G-tetrad relative to the G-tetrad above and below [21]. All of these variations will ultimately impact the thermodynamics of the unfolding process.
With our awareness of the relationships between sequence context, conformation and stability, it is quite reasonable to assume that sequence context variations in the loops of intramolecular quadruplexes will influence their confirmations and stabilities. Further, can we use sequence context effects to delineate the unfolding mechanism? We have recently shown that changing the sequence of all loops and the tail systematically influences conformation and stability [4]. Here, we report the effects of changing the sequence context of any one loop, one loop at a time. We demonstrate that changing just one base in any one loop can dramatically influence conformation and stability as determined by circular dichroism (CD) studies.
2. METHODS AND MATERIALS
2.1. Buffer Preparation
Standard potassium phosphate buffer (150 mM K+, 10 mM 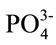 and 0.1 mM EDTA) at pH 7 was prepared, filtered through a 0.45 µm Millipore filter and degassed before being stored for use.
and 0.1 mM EDTA) at pH 7 was prepared, filtered through a 0.45 µm Millipore filter and degassed before being stored for use.
2.2. Sample Preparation
HPLC purified oligonucleotides were purchased from Biosynthesis Inc (Lewisville, TX) and used without further purification. All sequences were reconstituted in 1 ml filtered buffer followed by heating to 95˚C. After cooling to room temperature, the sample was stored for 24 hrs at 5˚C.
2.3. Determination of DNA Concentration
DNA concentrations solutions were determined by UV/V are using a Varian Cary 100 Bio model (Varian Associates, Palo Alto, CA) spectrometer. The reconstituted DNA sample was diluted 10 fold in 150 mM K+ buffer and scanned between 320 nm and 220 nm at 25˚C and 95˚C with subtraction of the buffer baseline. Concentrations were calculated using the molar extinction coefficients provided by the supplier of the DNA oligomers and the observed absorption at 260 nm.
2.4. Circular Dichroism Spectrophotometry
Circular dichroism spectra were collected with DESA Rapid Scanning Monochromator Spectrophotometer (Model Olis RSM 1000) attached with a temperature controller (Julabo CF31) and a nitrogen purging unit. CD spectra were collected from 320 nm to 220 nm at 1 nm intervals with an integration time of 3 seconds at 25˚C or 95˚C. For optical melting profiles, spectra were collected from 25˚C to 95˚C at an interval of 5˚C with an integration time of 3 seconds. After equilibration for 5 minutes at each temperature, the spectrum from 320 to 220 nm was collected. A baseline of the 150 mM K+ buffer was obtained from 320 - 220 nm using an integration time of 3 seconds and was subtracted from each spectrum. Samples were run in a 1 mm circular quartz cuvette. Data were analyzed using Olis Global Works and SigmaPlot version 11.
3. RESULTS AND DISCUSSION
Keeping in mind that the wild type sequence is (TTAGGG)4, we examined the conformation and stability of all permutations of the A and T bases in each loop via CD optical spectroscopy. The CD spectrum of each oligomer is unique. Figures 2 and 3 compare spec-

Figure 2. CD spectra of the loop variants with just one base different from the wild type, (TTAGGG)4, as a function of sequence: TTA (black); TTT (red); TAA (blue); ATA (green). CD spectra were collected at 25˚C in 10 mM potassium phosphate buffer, pH 7.0, with a total of 150 mM K+.
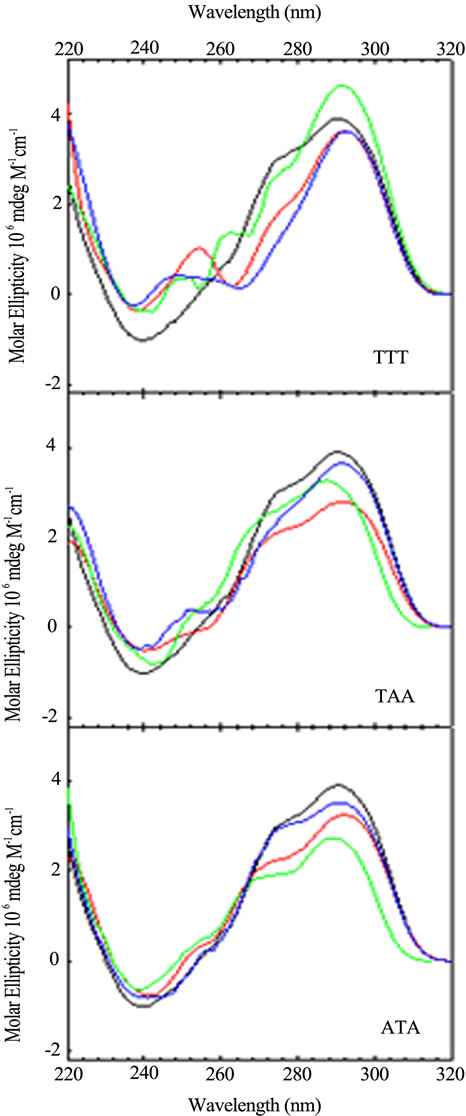
Figure 3. CD spectra of sequence variants with just one base different from the wild type, (TTAGGG)4, as a function of loop location: Wild Type (black); Loop 1 (red); Loop 2 (blue); Loop 3 (green). CD spectra were collected at 25˚C in 10 mM potassium phosphate buffer, pH 7.0, with a total of 150 mM K+.
tra, arranged by loop number or loop sequence context, respectively, for the quadruplex forming DNA sequences possessing a single base permutation of the TTA loop sequence found in each loop of the wild type structure. As can be seen, changing just one base in any one loop gives rise to a CD spectrum different from the wild type and all other sequences. The differences appear in peak or trough wavelength, peak or trough intensity and the presence or an absence of a shoulder in the 260 - 280 nm range. Similar trends are observed for the double and triple variants (data not shown).
These spectral differences are partly due to the changing of the chromophore from T to A or A to T, each of which contribute differently to the total CD spectrum [22, 23]. The other contribution to spectral changes is due to alteration of stacking of the loop bases within the loops. Hence, changing a single base in a loop changes its conformation due to differences in stacking. Sequence context plays a role in the structure and stability of DNA duplexes due to differences in base stacking for DNAs of identical GC content [24-28] and also plays a role in stacking of bases in single strand loops in duplex DNA [29,30]. The folding motifs of quadruplexes are dependent upon loop formation for the non G-rich segments and theses short loops are conformational restricted. Changing the conformation of one loop can lead to changing the conformations of the stacked tetrads and the other loops. So it should not be surprising that a single base change can influence loop conformation and overall structure.
Since base stacking influences conformation, it also influences stability. We carried out CD optical melting studies on all DNA sequences. Experimentally, we recorded the CD spectrum from 320 to 220 nm every 5˚C from 25˚C to 95˚C. To obtain Tm values (the melting temperature determined at the midpoint of the quadruplex to single strand transition), we plotted the fractional change of the molar ellipticity at 292 nm as a function of temperature. Figure 4" target="_self"> Figure 4 displays these optical melting curves grouped by loop variant. Tm values were obtained at the midpoint of the transition (i.e. fractional change = 0.5). The ΔTm values listed in Table 1 were determined by subtracting the Tm of the wild type sequence (71.4˚C) from that of the particular oligomer. As can be seen from Figure 4, not all quadruplexes are fully denatured at 95˚C so the ΔTm values in Table 1 are estimated for five of the quadruplexes as indicated by a >in front of the temperature.
It is quite remarkable that a single base change can influence the melting temperature in sometimes a dramatic fashion. For example, changing TTA to TTT increases the Tm by 8˚C to over 11˚C, depending upon the loop location of the permutation. It should be noted that we are using Tm as a basis for comparison of relative quadruplex stability. Our previous work demonstrated that the wild type sequence (TTAGGG)4 unfolds via a three state mechanism. None the less, we can still use Tm values to compare relative stabilities. Data for the double base variants and triple base variants are also included in Table 1. In general, these quadruplexes are more stable than the wild type as well.
Figure 5 graphically displays the Tm values for all permutations studied. Some generalizations can be made from these data. First, any loop triplet with a T in the first and third position gives rise to the most stable variant. The variant with all A bases in Loop 3 is the least
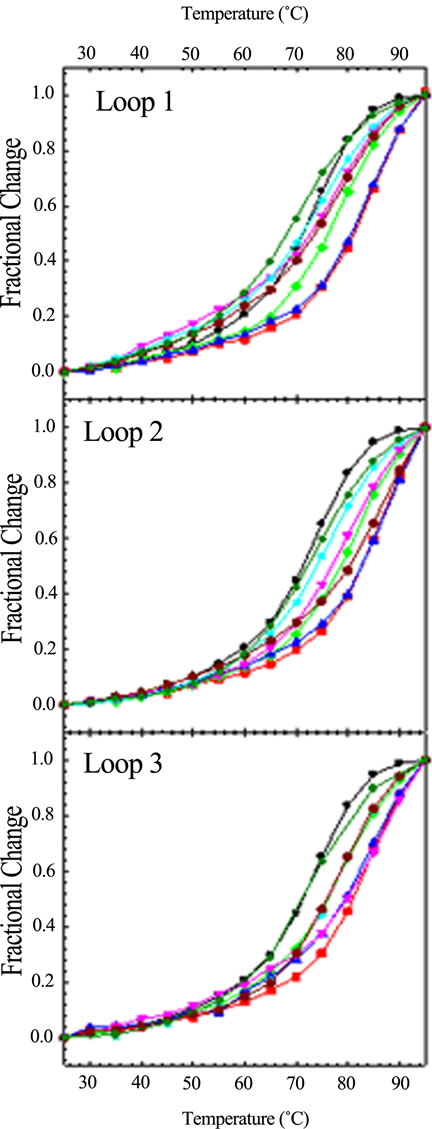
Figure 4. CD optical melting profiles of all loop variants: TTA (black circle); TAT (red square); ATT (green diamond); TTT (blue up triangle); AAT (pink down traingle); ATA (cyan star); TAA (dark red hexagon); AAA (dark green cross). CD spectra were collected in 10 mM potassium phosphate buffer, pH 7.0, with a total of 150 mM K+, every 5˚C from 25 to 95˚C with a five minute equilibration time at each temperature.
stable of all sequences. In general, the most stable variants, with respect to loop, are the Loop 2 variants.
Finally, with the exception of Loop 3 AAA, all other sequences are as stable as or more stable than the wild type sequence of TTA. In other words, the wild type se quence forms one of the least stable quadruplexes.
Table 1 also includes ΔTm values for DNA sequences previously studied in which all loops and the tail possess all permutations of A and T. As previously reported, these
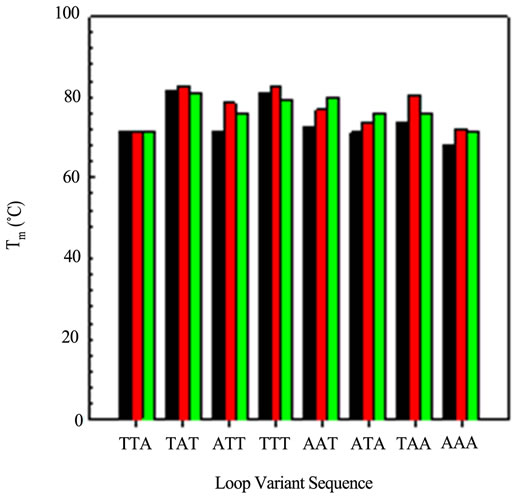
Figure 5. A comparison of the Tm values obtained from the CD optical melting profiles: Loop 1 variants (black); Loop 2 variants (red) and Loop 3 variants (green).
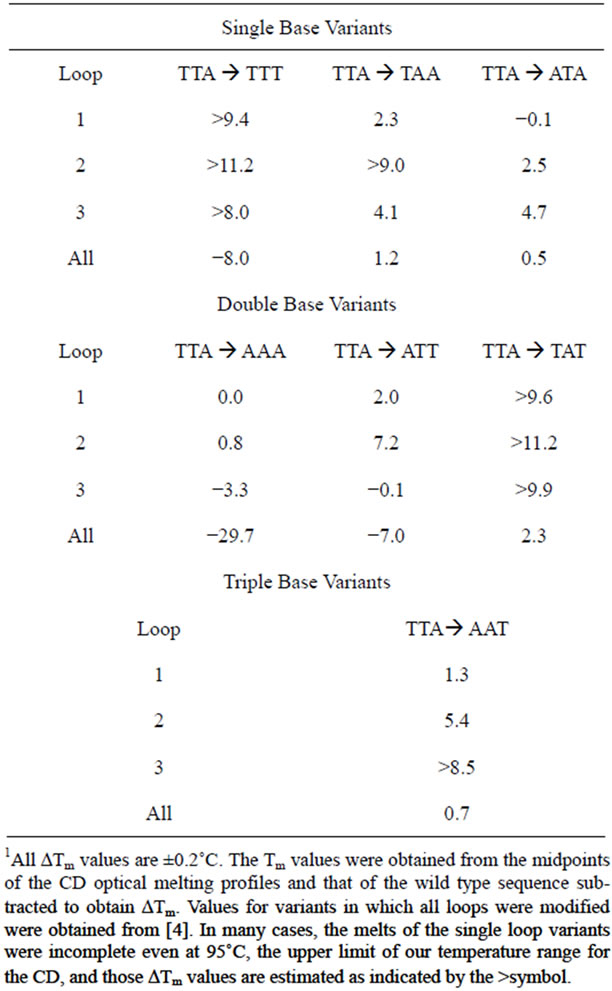
Table 1. Tabulate ΔTm values (˚C) for the various DNA oligomers.1
permutations can have dramatic effects on the overall stability and structure of the folded quadruplex [4]. However, the effect of changing one base in the loop is not additive when all loops are changed. For example, permuting TTA to TTT stabilizes the quadruplex if the permutation is only in one loop. However, changing all the loops and the tail leads to dramatic destabilization. Thus, the sequence context and length of the tail may have an effect as well on the overall stability. We are currently investigating that question.
Just as base-pair stacking in duplex DNA contributes to the total free energy of duplex formation in a sequence specific fashion [24-28], the stacking of the tetrads in an intramolecular quadruplex will also contribute to the total free energy of its folding. Additional contributions to the total free energy of quadruplex folding also comes from the base-stacking within the tail, within the loops and the stacking of third base of the tail with the first G-tetrad and the stacking of the first and third bases of the loops with the G-tetrads to which they are connected. There may also be stacking of the second base of the loops with the G-tetrads.
The stacking of unpaired bases to DNA or RNA duplexes has been thoroughly studied [31-36]. For duplex RNA, the presence of a dangling end (an overhang of unpaired bases) stabilizes the duplex relative to the blunt ended duplex with a 3’overhang more stabilizing than a 5’overhang [31,32]. For duplex DNA, the 5’overhang stabilizes more than the 3’overhang and in a sequence specific fashion of purine > T > C [33-36].
All quadruplexes studied here have a 5’-TTA tail giving rise to the stacking of the A base with the first G-tetrad. The contribution of that stack plus the stacking within the tail should be the same for all sequences studied. We have also investigated the effects of dangling ends on the human telomere quadruplex using (GGGTTA)3GGG with no overhangs, (TTAGGG)4 with a 5’-TTA overhang, (GGGTTA)4 with a 3’-TTA overhang and (TTAGGG)TTA with both a 5’ and 3’ TTA overhang via CD optical melting studies. Surprisingly, the stability order based on Tm is: no overhang > 3’-overhang > 5’- overhang > 5’- and 3’-overhang in both sodium and potassium solutions [37]. This observation is consistent to that recently reported for a single quadruplex with a 5’-TTA and a 3’-TT overhang having a lower Tm than the quadruplex with just the 5’-TTA overhang [38]. The tetrads will stack with either a 5’-T or 5’-A going into the loop and a 3’-T or 3’-A coming out of the loop. This stacking may also depend upon the type of loop (propeller vs transverse). Hence, the difference in observed Tm values also are due to changes in stacking of the tetrads with the bases in the loops and may also be due to difference in stacking of the tetrads induced by the conformational differences of the loops. A calorimetric study is currently underway to address the issues of stacking.
4. CONCLUSION
These studies have demonstrated the sensitivity of DNA G-quadruplexes to subtle sequence context changes in the loops that link the tetrads together. This sensitivity arises from the stacking interactions of the loop bases with themselves and/or with the G-tetrads. Since basebase stacking is dependent upon the nature of the bases, the preferred stacking would give rise to the lowest energy conformation of the loop. A conformational change in the loop will be compensated by an overall conformational change in the quadruplex. In general, the Tm values are both sequence specific as well as loop specific. Further, most variants have higher Tms than that of the wild type. Since unfolding of the quadruplexes located at the 3 termini of human chromosomes is the first step in DNA replication, it should not be surprising that the least stable structure evolved.
5. ACKNOWLEDGEMENTS
The authors wish to thank the Robert A Welch Foundation for the departmental research grant (# m-0200) supporting this research and the National Science Foundation (NSF# 0820845) for the acquisition of the CD spectrometer.
REFERENCES
- Otokiti, E.O. and Sheardy, R.D. (1997) Sequence effects on the relative thermodynamic stabilities of B-Z junction forming DNA oligomers. Biophysical Journal, 73, 3135-3141. doi:10.1016/S0006-3495(97)78339-5
- Paiva, A.M. and Sheardy, R.D. (2004) The influence of sequence context and length on the structure and stability of triplet repeat DNA oligomers. Biochemistry, 43, 14218-14227. doi:10.1021/bi0494368
- Paiva, A.M. and Sheardy, R.D. (2005) The influence of sequence context and length on the kinetics of duplex formation from complementary hairpins possessing (CNG) repeats. Journal of the American Chemical Society, 127, 5581-5585. doi:10.1021/ja043783n
- Tucker, B.A., Gabriel, S. and Sheardy, R.D. (2012) A CD Spectroscopic Investigation of Interand Intramolecular DNA Quadruplexes. In: Sheardy, R.D. and Winkle, S.A., Eds, Frontiers in Nucleic Acids, ACS Symposium Books, Washington DC.
- Sen, D. and Gilbert, W. (1990) Sodium-potassium switch in the formation of four-stranded G4-DNA. Nature, 344, 410-414. doi:10.1038/344410a0
- Hardin, C.C., Henderson, E., Watson, T. and Prosser, J.K. (1991) Monovalent cation induced structural transitions in telomeric DNAs: G-DNA folding intermediates. Biochemistry, 30, 4460-4472. doi:10.1021/bi00232a013
- Gray, R.D., Li, J. and Chaires, J.B. (2009) Energetics and kinetics of a conformational switch in G-quadruplex DNA. Journal of Physical Chemistry B, 113, 2676-2683. doi:10.1021/jp809578f
- Gray, R.D. and Chaires, J.B. (2011) Linkage of cation binding and folding in human telomere quadruplex DNA. Biophysical Chemistry, 159, 205-209. doi:10.1016/j.bpc.2011.06.012
- Phan, A.T., Gueron, M. and Leroy, J.L. (2000) The solution structure and internal motions of a fragment of the cytidine-rich strand of the human telomere. Journal of Molecular Biology, 299, 123-144. doi:10.1006/jmbi.2000.3613
- Phan, A.T. and Mergny, J.-L. (2002) Human telomeric DNA: G-quadruplex, i-motif and Watson-Crick double helix. Nucleic Acids Research, 30, 4618-4625. doi:10.1093/nar/gkf597
- Kaushik, M., Suehl, N. and Marky, L.A. (2007) Calorimetric unfolding of the bimolecular and i-motif complexes of the human telomere complementary strand, d(C3TA2)4. Biophysical Chemistry, 126, 154-164. doi:10.1016/j.bpc.2006.05.031
- Choi, J., Kim, S., Tachikawa, T., Fujitsuka, M. and Majima. T. (2011) pH-induced intramolecular folding dynamics of i-motif DNA. Journal of the American Chemical Society, 133, 16146-16153. doi:10.1021/ja2061984
- Jin, K.S., Shin, S.R., Ahn, B., Rho, Y., Kim. S.J. and Ree, M. (2009) pH-dependent structures of an i-motif DNA in solution. Journal of Physical Chemistry, 113, 1852-1856. doi:10.1021/jp808186z
- Antonacci, C., Chaires, J.B. and Sheardy, R.D. (2007) Biophysical characterization of the human telomeric repeat (TTAGGG)4 in potassium solution. Biochemistry, 47, 4654-4660. doi:10.1021/bi602511p
- Hazel, P., Huppert, J., Balasubramanian, S. and Neidle, S. (2004) Loop-length-dependent folding of G-quadrukplexes. Journal of the American Chemical Society, 126, 16405-16415. doi:10.1021/ja045154j
- Risitano, A. and Fox, K.R. (2004) Influence of loop size on the stability of intramolecular DNA quadruplexes. Nucleic Acids Research, 32, 2598-2606. doi:10.1093/nar/gkh598
- Rujan, I.N., Meleny, J.C. and Bolton, P.H. (2005) Vertebrate telomere repeat DNAs favor external loop propeller quadruplex structures in the presence of high concentration of potassium. Nucleic Acids Research, 33, 2022-2031. doi:10.1093/nar/gki345
- Kumar, N., Sahoo, B., Varun, K.A.S., Maiti, S. and Maiti, S. (2008) Effect of loop variation on quadruplex-Watson Crick duplex competition. Nucleic Acids Research, 36, 4433-4442. doi:10.1093/nar/gkn402
- Balkwill, G.D., Garner, T.P., Williams, H.E.L. and Searle, M.S. (2009) Folding topology of a bimolecular quadruplex containing a stable mini-hairpin motif within the diagonal loop. Journal of Molecular Biology, 385, 1600-1615. doi:10.1016/j.jmb.2008.11.050
- Fugimoto, T., Miyoshi, D., Tateishi-Karimata, H. and Sugimoto, N. (2009) Thermal stability and hydration state of DNA G-quadruplex regulated by loop regions. Nucleic Acids Symposium Series, 53, 237-238. doi:10.1093/nass/nrp119
- Gray, D.M., Wen, J.-D., Gray, C.W., Repges, R., Repges, C., Raabe, G. and Fleischhauer, J. (2008) Measured and calculated CD spectra of G-quartets stacked with the same or opposite polarities. Chirality, 20, 431-440. doi:10.1002/chir.20455
- Johnson, W.C. Jr. (1992) Analysis of circular dichroism spectra. Methods in Enzymology, 210, 426-447. doi:10.1016/0076-6879(92)10022-6
- Johnson, W.C. Jr. (2000) CD of nucleic acids. In: Borova, N., Naknishi, K. and Woody, R.W., Eds., Circular Dichroism, 2nd Edition, Wiley-VCH, New York, 703-718.
- Breslauer, K.J., Frank, R., Blocker, H. and Marky, L.A. (1986) Predicting DNA duplex stability from base sequence. Proceedings of the National Academy of Science USA, 83, 3746-3750. doi:10.1073/pnas.83.11.3746
- Doktycz, M.J., Goldstein, R.F., Paner, T.M., Gallo, F.J. and Benight, A.S. (1992) Studies of DNA dumbbells. I. melting curves of 17 DNA dumbbells with different duplex stem sequences linked by T4 endloops: Evaluation of the nearest-neighbor stacking interactions in DNA. Biopolymers, 32, 849-864. doi:10.1002/bip.360320712
- Hunter, C.A. (1993) Sequence-dependent DNA structure: The role of base stacking interactions. Journal of Molecular Biology, 230, 1025-1054. doi:10.1006/jmbi.1993.1217
- Freidman, R.A. and Honig, B. (1995) A free energy analysis of nucleic acid base stacking in aqueous solution. Biophysical Journal, 69, 1528-1535. doi:10.1016/S0006-3495(95)80023-8
- SantaLucia, J. Jr., Allawi, H.T. and Seneviratne, P.A. (1996) Improved nearest-neighbor parameters for predicting DNA duplex stability. Biochemistry, 35, 3555-3562. doi:10.1021/bi951907q
- Senior, M.M., Jones, R.A. and Breslauer, K.J. (1988) Influence of loop residues on the relative stabilites of DNA hairpin structures. Proceedings of the National Academy of Science USA, 85, 6242-6246. doi:10.1073/pnas.85.17.6242
- Paner, T.M., Amaratunga, M., Doktycz, M.J. and Benight, A.S. (1990) Analysis of melting transitions of the DNA hairpins formed from oligomer sequences d[GGATA- (X)4GTATCC] (X = A, T, G, C). Biopolymers, 29, 1715-1734. doi:10.1002/bip.360291405
- Petersheim, M. and Turner, D.H. (1983) Base-stacking and base-pairing contributions to helix stability: Thermodynamics of double-helix formation with CCGG, CCGGp, CCGGAp, ACCGGp, CCGGUp and ACCGGUp. Biochemistry, 22, 256-263. doi:10.1021/bi00271a004
- Freier, S.M., Alkema, D., Sinclair, A. and Turner, D.H. (1985) Contributions of dangling end stacking and terminal base-pair formationto the stabilities of GGCCp, XCCGGp, XGGCCp and XCCGGY. Biochemistry, 24, 4533-4539. doi:10.1021/bi00338a008
- Senior, M.M., Jones, R.A. and Breslauer, K.J. (1988) Influence of dangling thymine residues on the stability and structure of two DNA duplexes. Biochemistry, 28, 720-725. doi:10.1002/bip.360300718
- Doktycz, M.J., Paner, T.M., Amaratunga, M. and Benight, A.S. (1990) Thermodynamic stability of the 5’ danglingended DNA hairpins formed from sequences 5’-(XY)2 GGATAC(T)4GTATCC-3’ where X,Y = A, T, G, C. Biopolymers, 30, 829-845.
- Marotta, S.P. and Sheardy, R.D. (1996) Conformational properties of Z-forming DNA oligomers bearing terminal unpaired bases. Biophysical Journal, 71, 3361-3369. doi:10.1016/S0006-3495(96)79529-2
- Bommarito, S., Peyret, N. and SantaLucia, J. Jr. (2000) Thermodynamic parameters for DNA sequences with dangling ends. Nucleic Acids Research, 28, 1929-1934. doi:10.1093/nar/28.9.1929
- Agatep, A., Fagbohan, O. and Sheardy, R.D. (2012) DNA Quadruplexes with Overhangs. Chancellor’s Symposium for Creative Arts and Research, Texas Woman’s University, Denton.
- Petraccone, L., Spink, C., Trent, J.O., Garbett, N.C., Mekmaysy, C.S., Giancola, C. and Chaires, J.B. (2011) Structure and stability of higher-order human telomeric quadruplexes. Journal of the American Chemical Society, 133, 20951-20961. doi:10.1021/ja209192a

