Health
Vol.06 No.10(2014), Article ID:44460,5 pages
10.4236/health.2014.610108
Biotransformation of Carmoisine and Reactive Black 5 Dyes Using Saccharomyces cerevisiae
Abbas Sadeghi1*, Mir Aboutaleb Kazemi Bazardehi2, Shahrbanoo Raffe2, Batoul Zarif2
1Department of Environmental and Occupational Health Engineering, School of Health, Mashhad University of Medical Sciences, Mashhad, Iran
2Health Science Research Center, School of Health, Mashhad University of Medical Sciences, Mashhad, Iran
Email: *Sadeghia@mums.ac.ir
Copyright © 2014 by authors and Scientific Research Publishing Inc.
This work is licensed under the Creative Commons Attribution International License (CC BY).
http://creativecommons.org/licenses/by/4.0/



Received 16 February 2014; revised 20 March 2014; accepted 29 March 2014
ABSTRACT
Saccharomyces cerevisiae (baker’s yeast) is the most important industrial microorganisms. This yeast is commonly used as a leavening agent in baking bread and bakery products, where it produces carbon dioxide from converting of the fermentable sugars present in the dough. Nowadays, industrial and chemical activities led to produce new compounds with new kinds of contamination in the environment. Discharge of untreated or partially treated industrial sewage has created the contamination problems of rivers and lakes such as drugs, oil, heavy metals, paints, pesticides and various chemical compounds in them. Hence, it is necessary to control and reduce the levels of these compounds in wastewater and bring them to permissible values. This study aims to study the bioconversion potential of commonly available Saccharomyces cerevisiae for the two textile dyes of Carmoisine and Reactive Black 5. Reaction mixtures for biotransformation of dyes included 50 mg/l Carmoisine or 25 mg/l Reactive Black 5 and 1% dried harvested cells of S. cerevisiae (bread’s yeast) were tested. Harvested dry and wet yeast were studied for this purpose. The results show that harvested cells of Saccharomyces cerevisiae are able to bioconvert Carmoisine and Reactive Black 5. Reactive Black 5, Carmoisine are degraded by biotransformation 85% and 53% within 24 hours in water at the room temperature.
Keywords:
Saccharomyces cerevisiae, Carmoisine, Reactive Black 5, Biotransformation, Dyes, Decolourization

1. Introduction
Large numbers of synthetic dyes are used for various industrial applications and significant proportion appears in the form of wastewater and is spilled into the environment. These contaminated effluents which mainly are from dyeing and also finishing processes and are associated with the water pollution. Wastewater resulting from these shows improper impacts in terms of Biological Oxygen Demand (BOD), Chemical Oxygen Demand (COD), Total Organic Carbon (TOC), suspended solids, color, affect on pH and the organic compounds, [1] -[3] . Azo dyes may be classified as toxic and carcinogenic [4] . Synthetic dyes cannot be efficiently decolorized by traditional biological processes [5] . Thus, a number of biological and chemical methods have been developed for the efficient removal of industrial azo dyes [6] [7] .
Azo dyes are electron-deficient xenobiotic components because of their azo linkage, and also other electron- withdrawing groups, which generate an electron deficiency and make the dye less susceptible to biodegradation [8] [9] . However under the appropriate conditions, they can be degraded by reductases [10] - [12] . Azoreductases work only in the presence of reducing equivalents, e.g., FADH, NADH and NADPH [13] [14] . The available evidence indicates that azoreductase activity can be associated with more than one reductase [15] . Azoreductases are present in microorganisms, such as bacteria [16] - [18] , algae [19] and yeast [20] .
The use of microorganisms for the biodegradation of dyes is an attractive alternative to the development of bioremediation processes for the treatment of textile wastewater. Biological methods are environmentally friendly, produce less sludge than physical and chemical systems, and are relatively inexpensive, as the running cost is low. Microbial discoloration can occur via biosorption, enzymatic degradation or a combination of both [21] .
Yeast has long been known to be capable of bioaccumulation of heavy metal from solution and recently some reports for accumulation of dyes [22] - [30] . But little work has been carried out investigating the ability of yeast to act a biocatalyst for textile dyes especially using harvested cells.
This study aims to study the bioconversion potential of commonly available Saccharomyces cerevisiae yeast for the two textile dyes of Carmoisine and Reactive Black 5 at batch-scale level.
2. Materials and Methods
2.1. Chemicals
All chemicals used in the experiments were reagent grade. All solutions were prepared with distilled water. Carmoisine and Reactive Black 5 were obtained from a local company (Alvan Sabet, Tehran, Iran).
2.2. Microorganism
Harvested cells of S. cerevisiae or baker’s yeast were locally purchased from Razavi Yeast Company, Mashhad, Iran.
In this experiment, yeast was prepared at a concentration of 1%. For this purpose, 1 g of yeast was suspended in 100 ml of toxic substance solution.
2.3. Preparation of Reaction Mixtures
200 ml reaction mixtures were prepared by mixture of dye Carmoisine (50 mg/l) or Reactive Black (25 mg/l) and 2 grams Harvested cells of S. cerevisiae. The experiments were performed at room temperature (28˚C ± 2˚C).
2.4. Analytical Methods
Five milliliters of sample was taken from each beaker at definite time intervals. Samples were centrifuged to remove suspended biomass and the concentration of dye in the supernatant was determined by reading absorbance at 590 nm for Reactive Black 5 and 515 nm for Carmoisine. Absorbance measurements were carried out by using a PG Instruments, T80+UV/VIS model spectrometer.
3. Results
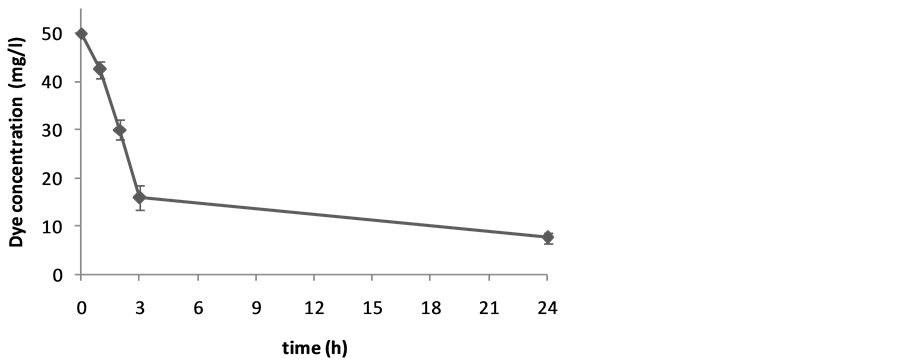
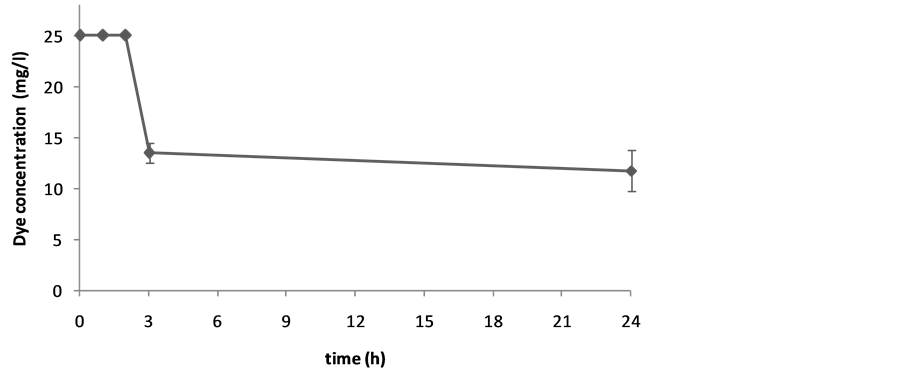
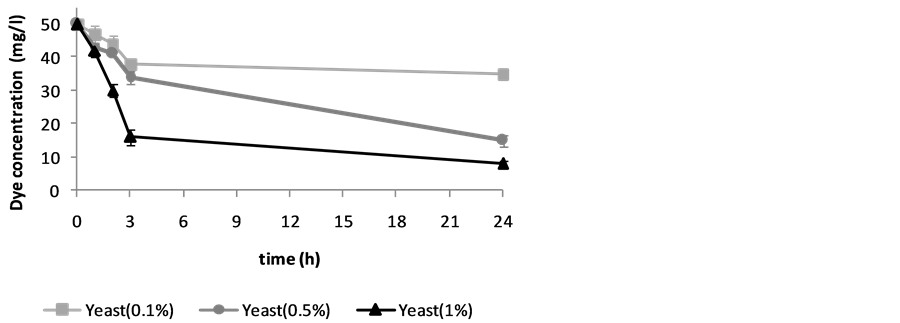
Harvested cells of S. cerevisiae were investigated in the reaction mixtures to study the ability for biotransformation of two synthetic dyes. A few dye bioconversions have been reported by this yeast. The results are given asthe units of percentage of biotransformation in Table 1. Figure 1 and Figure 2 show the decolorization of Carmoisine by S. cerevisiae. Figure 3 shows the decolorization of Reactive Black 5 (25 mg/l) and Figure 4 shows biotransformation of Carmoisine (50 mg/l) using different concentration of S. cerevisiae.
Initially different concentrations of cells (0.05% to 2.5%) were studied and it was found that by increasing the cell mass, more biotransformation happened. For main experiments the concentration of 1% was used. Decrease in the absorption indicates that decolourization of this dye occurred by degradation. Reactive Black 5 and Carmoisine are degraded by biotransformation 85% and 53% within 24 hours in water at the room temperature.
S. cerevisiae is capable of utilizing a variety of carbon and nitrogen sources. In the absence of natural carbon and nitrogen sources the yeast is able to use some other synthetic chemicals. In this investigation, an experiment protocol was designed and used to check the ability of Saccharomyces cerevisiae to utilize two textile dyes of
Table 1. Percent of the dye Bioconversion using S. cerevisiae at the different time. Values are the mean of three experiments ± SD.
Figure 1. Bioconversion of Carmoisine (50 mg/l) by S. cerevisiae (1%) during 24 hours. Values are the mean of three experiments ± SD.
Figure 2. Biotransformation of Carmoisine using wet cells of S. cerevisiae (1%) with different concentration of Carmoisine dye (1: 2.5 mg/l, 2: 2.5 mg/l, 3: 5 mg/l, 4: 10 mg/l, 5: 20 mg/l, 6: 50 mg/l).
Figure 3. The decolorization of Reactive Black 5 (25 mg/l) using S. cerevisiae (1%) during 24 hours. Values are the mean of three experiments ± SD.
Figure 4. Biotransformation of Carmoisine (50 mg/l) using different concentration of S. cerevisiae (0.1%, 0.5% and 1%) during 24 hours. Values are the mean of three experiments ± SD.
Carmoisine and Reactive Black 5 at batch-scale level. Microscopic and macroscopic observations showed that the dye decolourizations are due to microbial biotransformation and not due to biosorption.
4. Discussion
Although some reports have mentioned that S. cerevisiae in cultures is able to accumulate some dyes in several days. Biotransformation of these dyes in this study proves that the harvested cells of the S. cerevisiae can be promising for further research and practical usage in the field of dye biotransformation for example in chemical, biological sciences and industries. As well hold promise in providing a low cost and efficient means to treat the textile effluent.
References
- Solís, M., Solís, A., Pérez, H.I., Manjarrez, N. and Flores, M. (2012) Microbial Decolouration of Azo Dyes: A Review. Process Biochemistry, 47, 1723-1748. http://dx.doi.org/10.1016/j.procbio.2012.08.014
- Savin, I.I. and Butnaru, R. (2008) Wastewater Characteristics in Textile Finishing Mills. Environmental Engineering and Management Journal, 7, 859-864.
- Kuberan, T., Anburaj, J., Sundaravadivelan, C. and Kumar, P. (2011) Biodegradation of Azodye by Listeria sp. International Journal of Environmental Sciences, 1, 1760-1770.
- Puvaneswari, N., Muthukrishnan, J. and Gunasekaran, P. (2006) Toxicity Assessment Andmicrobial Degradation of Azo Dyes. Indian Journal of Experimental Biology, 44, 618-626.
- Ghoreishi, M. and Haghighi, R. (2003) Chemical Catalytic Reaction and Biological Oxidationfor Treatment of Non- Biodegradable Textile Effluent. Chemical Engineering Journal, 95, 163-169. http://dx.doi.org/10.1016/S1385-8947(03)00100-1
- Forgacs, E., Cserhati, T. and Oros, G. (2004) Removal of Synthetic Dyes from Wastewaters: A Review. Environment International, 30, 953-971. http://dx.doi.org/10.1016/j.envint.2004.02.001
- Chacko, J.T. and Subramaniam, K. (2011) Enzymatic Degradation of Azo Dyes: A Review. International Journal of Environmental Sciences, 1, 1250-1260.
- Hsueh, C.C., Chen, B.Y. and Yen, C.Y. (2009) Understanding Effects of Chemical Structure on Azo Dye Decolorization Characteristics by Aeromonas Hydrophila. Journal of Hazardous Materials, 167, 995-1001. http://dx.doi.org/10.1016/j.jhazmat.2009.01.077
- Enayatizamir, N., Tabandeh, F., Rodríguez Couto, S., Yakhchali, B., Alikhani, H.A. and Mohammadi, L. (2011) Biodegradation Pathway and Detoxification of the Diazodye Reactive Black 5 by Phanerochaete Chrysosporium. Bioresource Technology, 102, 10359-10362. http://dx.doi.org/10.1016/j.biortech.2011.08.130
- Misal, S.A., Lingojwar, D.P., Shinde, R.M. and Gawai, K.R. (2011) Purification and Characterization of Azoreductase from Alkaliphilic Strain Bacillus Badius. Process Biochemistry, 46, 1264-1269. http://dx.doi.org/10.1016/j.procbio.2011.02.013
- Stolz, A. (2001) Basic and Applied Aspects in the Microbial Degradation of Azo Dyes. Applied Microbiology and Biotechnology, 56, 69-80. http://dx.doi.org/10.1007/s002530100686
- Sugumar, S. and Thangam, B. (2012) BiodEnz: A Database of Biodegrading Enzymes. Bioinformation, 8, 40-42. http://dx.doi.org/10.6026/97320630008040
- Van der Zee, F.P. and Cervantes, F.J. (2009) Impact and Application of Electron Shuttles on the Redox (bio) Transformation of Contaminants: A Review. Biotechnology Advances, 27, 256-277. http://dx.doi.org/10.1016/j.biotechadv.2009.01.004
- Dos Santos, A.B., Cervantes, F.J. and Van Lier, J.B. (2007) Review Paper on Current Technologies for Decolourisation of Textile Wastewaters: Perspectives for Anaerobicbiotechnology. Bioresource Technology, 98, 2369-2385. http://dx.doi.org/10.1016/j.biortech.2006.11.013
- Majeau, J.A., Brar, S.K. and Tyagi, R.D. (2010) Laccases for Removal of Recalcitrant and Emerging Pollutants. Bioresource Technology, 101, 2331-2350. http://dx.doi.org/10.1016/j.biortech.2009.10.087
- Dhanve, R.S., Shedbalkar, U.U. and Jadhav, J.P. (2008) Biodegradation of Diazo Reactive Dye Navy Blue HE2R (Reactive Blue 172) by an Isolated Exiguobacterium sp RD3. Biotechnology and Bioprocess Engineering, 13, 53-60. http://dx.doi.org/10.1007/s12257-007-0165-y
- Jadhav, J.P., Phugare, S.S., Dhanve, R.S. and Jadhav, S.B. (2009) Rapid Biodegradation and Decolorization of Direct Orange 39 (Orange TGLL) by an Isolated Bacterium Pseudomonas Aeruginosa Strain BCH. Biodegradation, 21, 453- 463. http://dx.doi.org/10.1007/s10532-009-9315-6
- Patil, P.S., Phugare, S.S., Jadhav, S.B. and Jadhav, J.P. (2010) Communal Action of Microbialcultures for Red HE3B Degradation. Journal of Hazardous Materials, 181, 263-270. http://dx.doi.org/10.1016/j.jhazmat.2010.05.006
- El-Sheekh, M.M., Gharieb, M.M. and Abou-El-Souod, G.W. (2009) Biodegradation of Dyes by Some Green Algae and Cyanobacteria. International Biodeterioration and Biodegradation, 63, 699-704. http://dx.doi.org/10.1016/j.ibiod.2009.04.010
- Jadhav, J.P., Parshetti, G.K., Kalme, S.D. and Govindwar, S.P. (2007) Decolourization of Azodye Methyl Red by Saccharomyces cerevisiae MTCC 463. Chemosphere, 68, 394-400. http://dx.doi.org/10.1016/j.chemosphere.2006.12.087
- Wu, Y., Li, T. and Yang, L. (2012) Mechanisms of Removing Pollutants from Aqueous Solutions by Microorganisms and Their Aggregates: A Review. Bioresource Technology, 107, 10-18. http://dx.doi.org/10.1016/j.biortech.2011.12.088
- Murthy, Ch.V.R., Kishore Kumar, K., Jayaraju, K. and Silas, S. (2007) Biosorption Studies for Removal of Cadmium from Wastewater Using Immobilized Saccharomyces cerevisiae. Asian Journal of Chemistry, 19, 3502-3510.
- Ferreira, I., Pinho, O., Vieira, E. and Tavarela, J.G. (2010) Brewer’s Saccharomyces Yeast Biomass: Characteristics and Potential Applications. Trends in Food Science and Technology, 21, 77-84. http://dx.doi.org/10.1016/j.tifs.2009.10.008
- Machado, M.D., Soares, E.V. and Soares, H.M.V.M. (2010) Removal of Heavy Metals Using a Brewer’s Yeast Strain of Saccharomyces cerevisiae: Chemical Speciation as a Tool in the Prediction and Improving of Treatment Efficiency of Real Electroplating Effluents. Journal of Hazardous Materials, 180, 347-353. http://dx.doi.org/10.1016/j.jhazmat.2010.04.037
- Yavuz, H., Denali, A., Güngüneş, H., Safarikova, M. and Safarik, I. (2006) Biosorption of Mercury on Magnetically Modified Yeast Cells. Separation and Purification Technology, 52, 253-260. http://dx.doi.org/10.1016/j.seppur.2006.05.001
- Zhang, Y., Fan, C., Meng, Q., Diao, Z., Dong, L., Peng, X., Ma, S. and Zhou, Q. (2009) Biosorption of Pb2+ by Sac- charomyces cerevisiae in Static and Dynamic Adsorption Tests. Bulletin of Environmental Contamination and Toxi- cology, 83, 708-712. http://dx.doi.org/10.1007/s00128-009-9847-9
- Ruta, L., Paraschivescu, C., Matache, M., Avramescu, S. and Farcasanu, I.C. (2010) Removing Heavy Metals from Synthetic Effluents Using “Kamikaze” Saccharomyces cerevisiae Cells. Applied Microbiology and Biotechnology, 85, 763-771. http://dx.doi.org/10.1007/s00253-009-2266-3
- El-Sharouny, E. and El-Sersy, N.A. (2005) Biosorption of Textile Dyes by Brewer’s Yeast Biomass. Fresenius Environmental Bulletin, 14, 928-933.
- Jadhav, J.P. and Govindwar, S.P. (2006) Biotransformation of Malachite Green by Saccharomyces cerevisiae MTCC 463. Yeast, 23, 315-323. http://dx.doi.org/10.1002/yea.1356
- Kumari, K. and Abraham, T.E. (2007) Biosorption of Anionic Textile Dyes by Nonviable Biomass of Fungi and Yeast. Bioresource Technology, 98, 1704-1710. http://dx.doi.org/10.1016/j.biortech.2006.07.030
NOTES

*Corresponding author.