Computational Molecular Bioscience
Vol.3 No.4(2013), Article ID:40941,13 pages DOI:10.4236/cmb.2013.34010
Interaction of Cationic and Anionic Phthalocyanines with Adenosine Deaminase, Molecular Dynamics Simulation and Docking Studies
School of Chemistry, Damghan University, Damghan, Iran
Email: *ajloo@du.ac.ir
Copyright © 2013 Davood Ajloo et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. In accordance of the Creative Commons Attribution License all Copyrights © 2013 are reserved for SCIRP and the owner of the intellectual property Davood Ajloo et al. All Copyright © 2013 are guarded by law and by SCIRP as a guardian.
Received September 28, 2013; revised October 28, 2013; accepted November 5, 2013
Keywords: Molecular Dynamics; Molecular Docking; Phthalocyanine; Adenosine Deaminase; Denaturation
ABSTRACT
Interactions of anionic, cationic and metal phthalocyanine with adenosine deaminase were studied by molecular dynamics and docking simulation. Structural parameters such as solvent accessible surface area (SAS), mid-point of transition temperature (Tm), radial distribution function (RDF) and hydrogen bond, helix, coil, beta percentage and other physical parameters were obtained. The denaturation of adenosine deaminase (ADA) by heat, anionic and cationic phthalocyanines was compared. A series of 20 ns simulation performed at temperatures ranging from 275 to 450 K, starting from the ADA native structure. Results of radial distribution functions (RDFs) showed that metallic derivative at low concentration behaves the same as osmolytes that increases the beta form and increases the enzyme stability. Molecular docking studies have been carried out to confirm the simulation results. Investigation of binding site and free energy confirmed that the efficiency of interaction with adenosine deaminase depends on metal core. Binding energy of non-metallic form is more negative than metallic form and it significantly decreases for phthalocyanine. Self-aggregation of anionic phthalocyanine decreases in comparison with cationic derivative, therefore enzyme denaturation in the presence of anionic form is higher than the other. Furthermore, thermal stability of the enzyme also depends on temperature in presence of phthalocyanine. Binding site of phthalocyanine on the enzyme has been identified by docking analysis.
1. Introduction
Adenosine deaminase (ADA) is a cytosolic enzyme, which has been the object of notable interest, mainly because in humans a genital defect in the enzyme causes severe combined immunodeficiency disease (SCID). ADA is an aminohydrolase (EC 3.5.4.4) which shares in the purine metabolism where it degrades either adenosine or 2’- deoxyadenosine producing inosine or 2’-deoxyinosine, respectively. The enzyme is widely distributed and many of its biochemical properties have been studied in different species [1]. ADA is a glycoprotein that consists of a single polypeptide chain of 311 amino acids. It was sequenced in 1984 [2]. The primary amino acids sequence of ADA is conserved across species [3]. The crystal structure also revealed that ADA is a metalloenzyme that complexes one mole of Zn2+ per mole of protein [4]. The product of human ADA gene consists of 363 amino acids (41 kDa) and there is a high degree of amino acids sequence conservation among species. The enzyme contains a parallel α/β barrel motif with eight central β strands and eight α helices, which is a common structure, found in 1/10 of known enzymes [5]. It also contains five added helices. In humans, the highest ADA activity is found in thymus and other lymphoid tissues whereas the lowest activity can be viewed in erythrocytes [6]. Adenosine deaminase, an enzyme distributed in the human tissues [7] is considered a good marker of cell arbitrated immunity [8]. It plays an important role in lymphocyte spread and differentiation, [9] and shows its highest activity in T-lymphocytes [10]. The human enzyme is genetically polymorphic and is found in 2 electrophoretically distinct forms termed ADA-1 and ADA-2 [11] disorders, especially in pancreatic cancer. It may be a serum marker for the diagnosis of pancreatic cancer [12]. ADA is involved in some diseases such as tuberculosis [13], brain tumor [14], lung cancer and mesothelioma [15]. In this study we have investigated the interaction between two phthalocyanines and ADA.
Metal phthalocyanines (MPc) are interesting species that have been considered for many applications in industry [16,17]. The likeness in structure between phthalocyanine and the biological molecules chlorophyll and haemoglobin adds to their interest, and a huge number of different MPcs have been produced over the years [18]. As a class of macrocyclic planar aromatic compounds, MPcs show special physical and chemical properties, and there have been many experimental studies of their optical, magnetic, and electrical properties. To interpret the electronic spectra of MPcs has been the subject of theoretical investigations [17,19]. Phthalocyanine is an organic compound which forms stable combinations with many metal atoms. These are incorporated at the center of the planar phthalocyanine molecule. The role of the metal atom is of interest in several fields where phthalocyanines find applications, for example, in photovoltaic energy change [20-23] and as catalysts in electrochemical reduction [24]. The primary PSs used for PDT have an important tendency to aggregate under their large planar aromatic ring systems, leading to strong Pc-Pc interactions and non-covalent complexes with proteins and other potential targeting compounds [25]. The important role of metalloenzymes in the biological processes has led to the synthesis and study of their model analogs. One of the most important classes of metalloenzymes constitutes the heme proteins. They are formed by conjugation of proteins with iron porphyrin which plays the role of the prosthetic group. Interaction between iron and cobalt tetra sulphonated phthalocyanines and globin results in the formation of the green complexes as has been proved by difference spectroscopy. Nowadays, phthalocyanines are widely used in the dyeing industry. Nearly a quarter of all pigments of organic origin are related to this class of compounds. Furthermore, they are used for the fabrication of high-speed and high-resolution optical media [26], as light harvesters in photovoltaic applications [27], and as experimental catalysts in redox reactions [28]. These dyes absorb strongly in the red and near infrared (NIR) part of the visible spectrum providing them with their characteristic blue or greenish color. Pcs that absorb in the NIR are especially interesting for photomedical applications such as fluorescence imaging, Photochemical internalisation (PCI), and Photodynamic Therapy (PDT) [29].
On the other hand simulation methods are relatively more available and less expensive than experimental methods for investigation of drug and receptor interaction. Molecular dynamics and docking are widely used in macromolecular structure and drug design [30-36].
In this work we study the influence of phthalocyanine on the structural parameters ADA enzyme by molecular dynamics and docking methods.
2. Methods
2.1. Molecular Dynamics Simulation
The structures of phthalocyanines with and without Ni metal were drawn using (Hyperchem 7 software). These structures were optimized by Molecular Mechanics Force Field (MM+) subsequently using the semi-empirical AM1 method (Scheme 1). For molecular dynamics simulation of phthalocyanines, force field parameters and geometries were generated Using PRODRG serve http://davapc1.bioch.dundee.ac.uk/cgi-bin/prodrg). The starting structure of ADA was set up based on the X-ray crystal structure of ADA (PDB code 1VFL). A cubic simulation box of the volume 9.2 × 9.2 × 8.7 nm3 was made with ADA and 0, 6, 12, and 18 phthalocyanine were placed randomly in this box respectively. The system was equilibrated for 20 ns at constant pressure (1 atm) and temperature 300, 325, 335, 350, 365, 375, 390, 425, and 450 K using the Parilleno-Rahman procedure. The resulting system of MD models contains a protein and various concentrations of phthalocyanine. All MD simulations were carried out using the GROMACS 4.5.4 Package [37]. The simple point charge (SPC) model was used to describe water [38]. A different time step was used to integrate the equations of motion with the Verlet algorithm [39]. The long-range electrostatic interactions were treated with the particle mesh Ewald method [40]. Temperatures and pressures were controlled by a NoseHoover thermostat [41,42] and Parrinello-Rahman barostate [43] with coupling constants of 0.1 and 0.5, respectively. For all simulations, the atomic coordinates were saved every 20 ps for analysis. The computer applied the Rocks cluster networking and Centos running systems and the repeated trajectory showed similar result. The structure of two phthalocyanines was shown in Scheme 1.
2.2. Molecular Dynamics Data Analyses
The conformational changes of the protein during MD simulations were checked by the root-mean-square derivations (RMSD) with its X-ray structure as a reference. The RMSD value, a measure of molecular mobility, is calculated by translating and rotating the coordinates of the instantaneous structure to superimpose the reference with a maximum overlap. The RMSD is defined as:
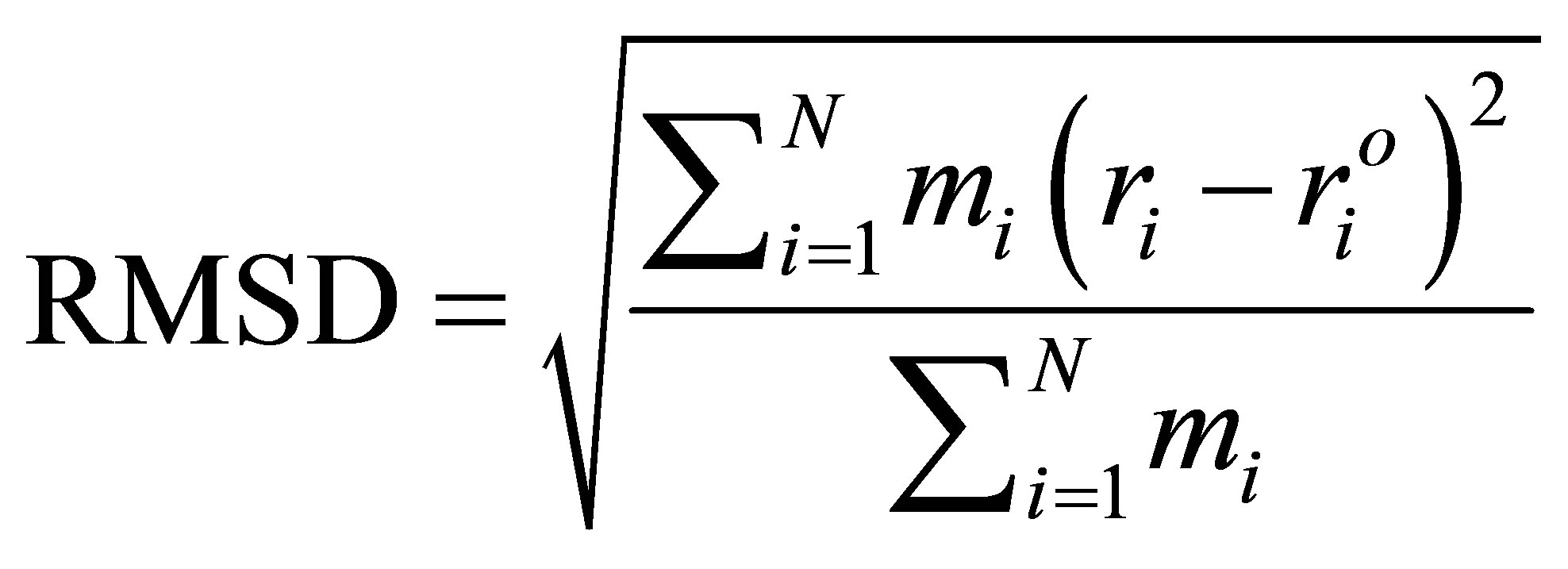 (1)
(1)
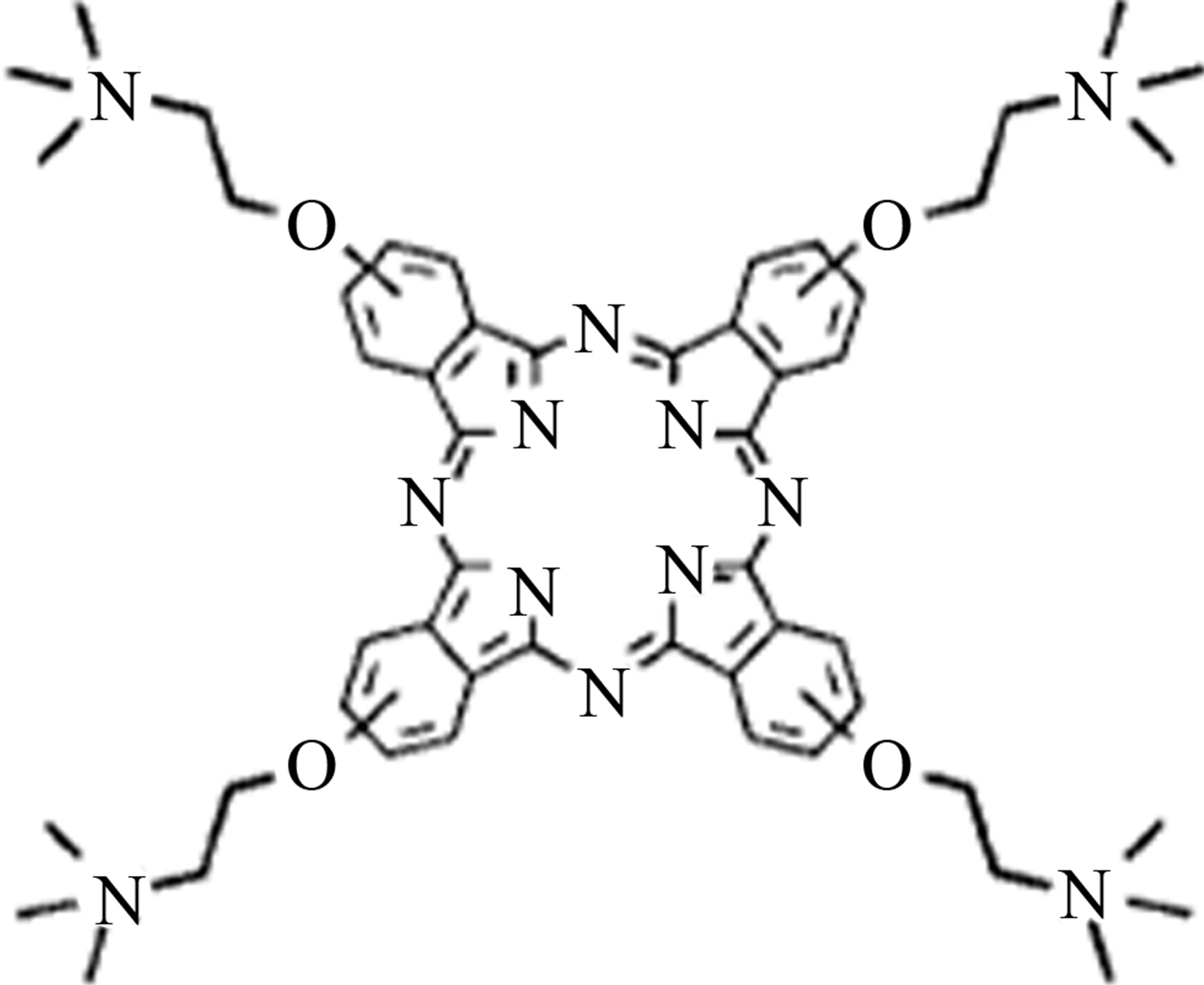
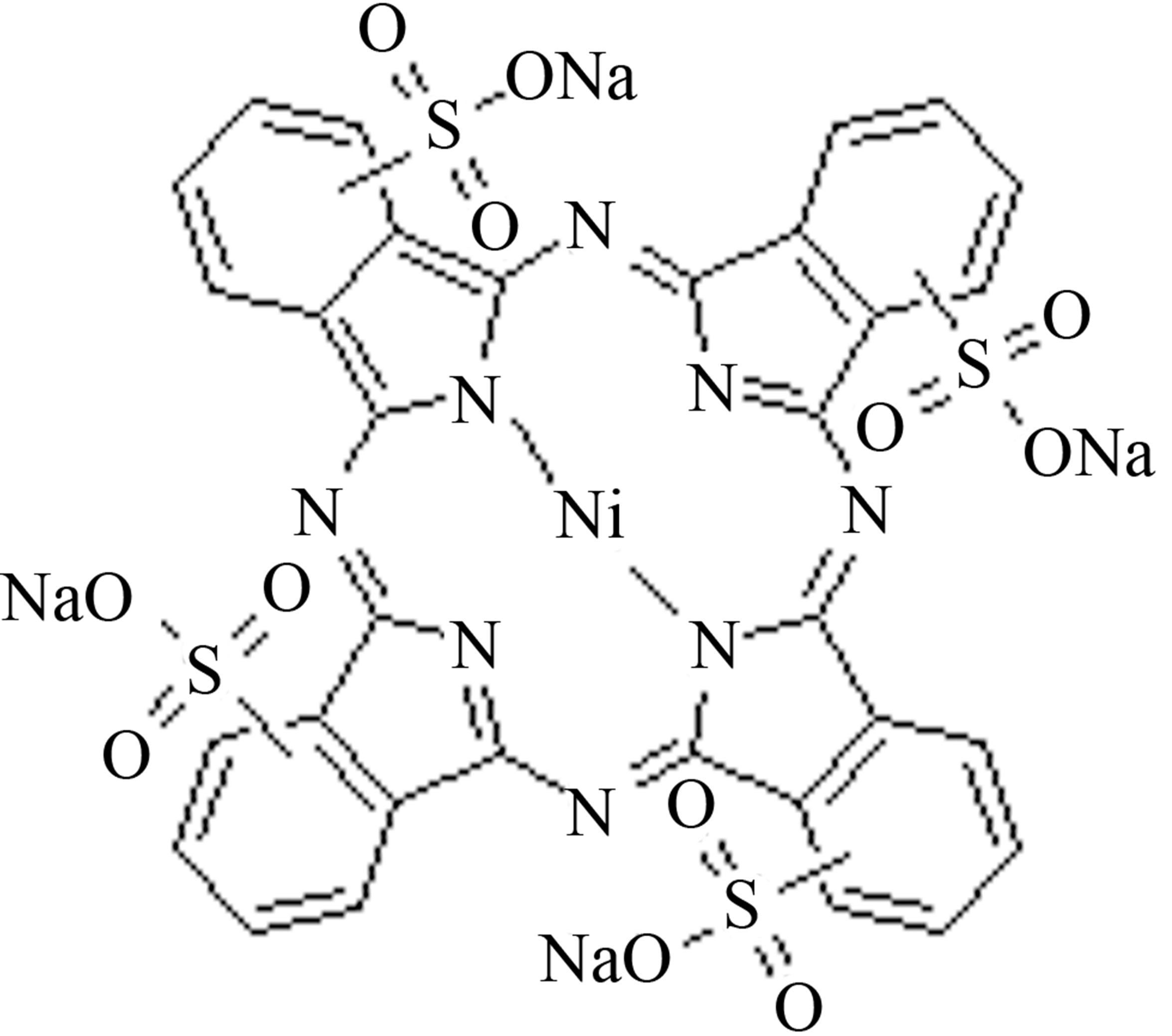
Scheme 1. Chemical structure of anionic (NiTNasPc) and cationic (PcTtme) phthalocyanines.
where mi is the mass of atom i. ri and 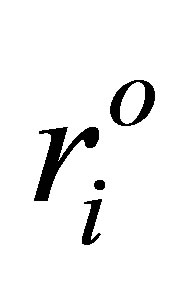 are the coordinates of atom i at a certain instance during MD simulations and at its reference state, respectively. RMSDs were calculated for the trajectories, from the starting structures of ADA as a function of time. In all systems, RMSDs reach a stable value within the first nanosecond of all the analyses.
are the coordinates of atom i at a certain instance during MD simulations and at its reference state, respectively. RMSDs were calculated for the trajectories, from the starting structures of ADA as a function of time. In all systems, RMSDs reach a stable value within the first nanosecond of all the analyses.
The simulation trajectories were analyzed using several auxiliary programs provided with the GROMACS 4.5.4 package. The programs include g_hbond for the Hydrogen-bond (H-bond) between hydrogen donors and acceptors. H-bonds are considered to be intact if the donor-to-acceptor distance is fewer than 0.35 nm and the donor-hydrogen-acceptor angle is within 30˚ of linearity.
g_gyrate compute a rough measure for radius of gyration (Rg) the compactness of a structure. The radius of gyration (Rg) for a protein is defined as the mass-weighted geometric mean of the distance of each atom from the protein’s center of mass. The radius of gyration for the ADA was computed using all the peptide atoms in the standard formula.
 (2)
(2)
where ri is the distance of atom i from the center of mass of the enzyme, and mi is its mass.
The radial distribution function (RDF) is the probability density of finding a particle at distance r from the reference particle. RDF between particles of type A and B is defined in the following term:
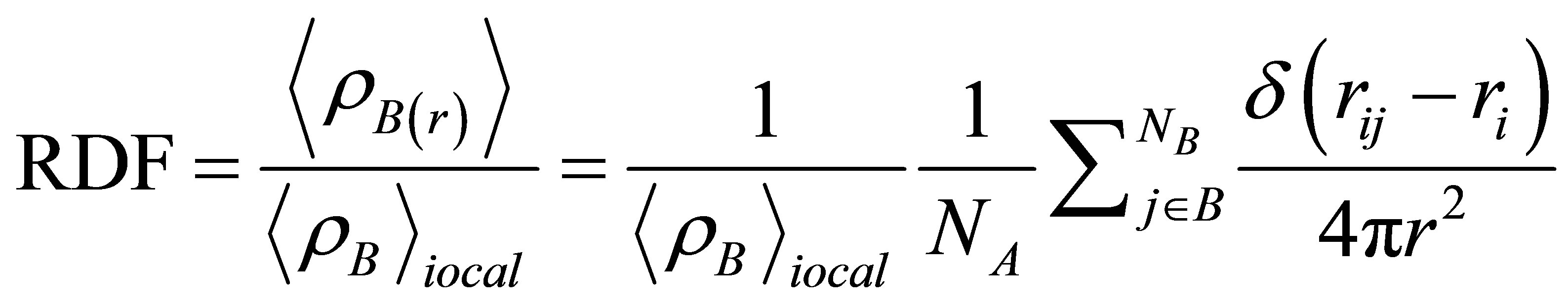 (3)
(3)
Where 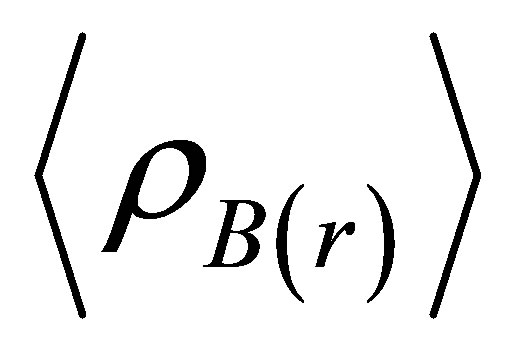 is the particle density of type B at a distance r around particles A, and
is the particle density of type B at a distance r around particles A, and 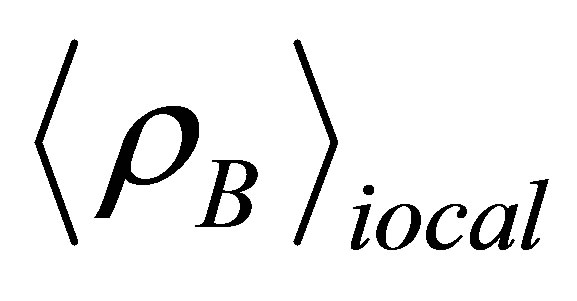 is the particle density of type B averaged over all spheres around particles A with radius rmax. Usually the value of rmax is half of the box length.
is the particle density of type B averaged over all spheres around particles A with radius rmax. Usually the value of rmax is half of the box length.
g_sas computes hydrophobic, hydrophilic and total solvent accessible surface area.
Percentages of helix, coil and beta were obtained by web server VADAR software [44].
2.3. Ligand Docking
To predict the binding energy of phthalocyanines to protein, Autodock software was used. The PDB files of phthalocyanine and phthalocyanine with best geometries extracted from Hyperchem were loaded into Auto Dock Tool (ADT) to calculate free energy of interaction. Polar hydrogen’s were added, and Gasteiger charges were computed for the ligand, the rigid root and the rotatable bonds were defined by the AutoTors tool of ADT. For the protein, all water molecules were removed, Kollman charges and solvation parameters were added (producing a pdbqs file). Based on the atom types, the suitable maps were calculated. Nonpolar hydrogens were merged for each atom. Grid maps of 126 × 126 × 126 points with a grid-point spacing of 0.375 were produced using the Auto Grid tool of ADT. The 250 genetic algorithm (GA) runs were performed with the following parameters: Population size of 150, maximum number of 2.5 × 105 energy evaluations, maximum number of 27,000 generations, an elitism of 1, a mutation rate of 0.02, and a crossover rate of 0.8. The resulting conformations were clustered using a root-mean-square deviation (RMSD) of 0.5 and the clusters were ranked in order of increasing energy. We want to find out computationally whether phthalocyanine and phthalocyanine will interact or bind to protein, and if so, we would like to compare the binding energy of them, as well as the affinity of the binding or interaction and Gibbs energy of interaction. AutoDock (3.0.5) software was used for calculation.
3. Results and Discussion
3.1. Effect of Temperature and Concentration
Here, we describe simulation of ADA at various temperatures and concentration, focusing on the unfolding process. Increasing temperature accelerates protein unfolding. Temperature is believed to alter the structure of the network of hydrogen bonds of protein in water and increase the SAS and protein size as well as decrease the intermolecular hydrogen bond, electrostatic and hydrophobic interactions of proteins. Structure parameters were obtained from MD simulation for each temperature and results were averaged. The structure information such as intermolecular hydrogen bonding (HB) between ADA and solvent molecules, and RMSD were obtained and averaged at each temperature.
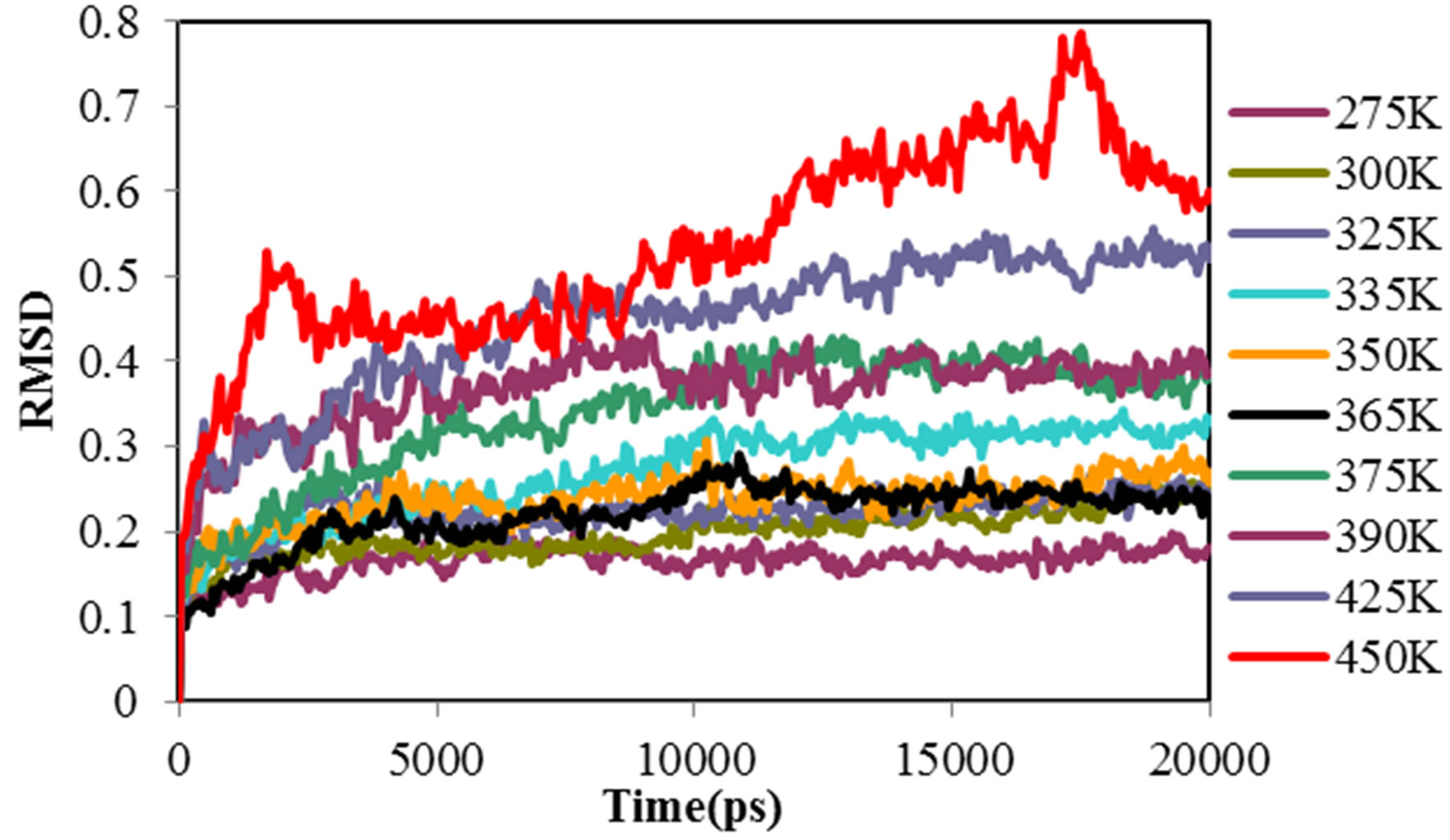
Root mean square deviation (RMSD) of the ADA for all temperatures and concentration were obtained. Figures 1(a) and (b) show the ADA RMSD in the 20 ns time interval in all temperatures and concentration. The
 (a)
(a)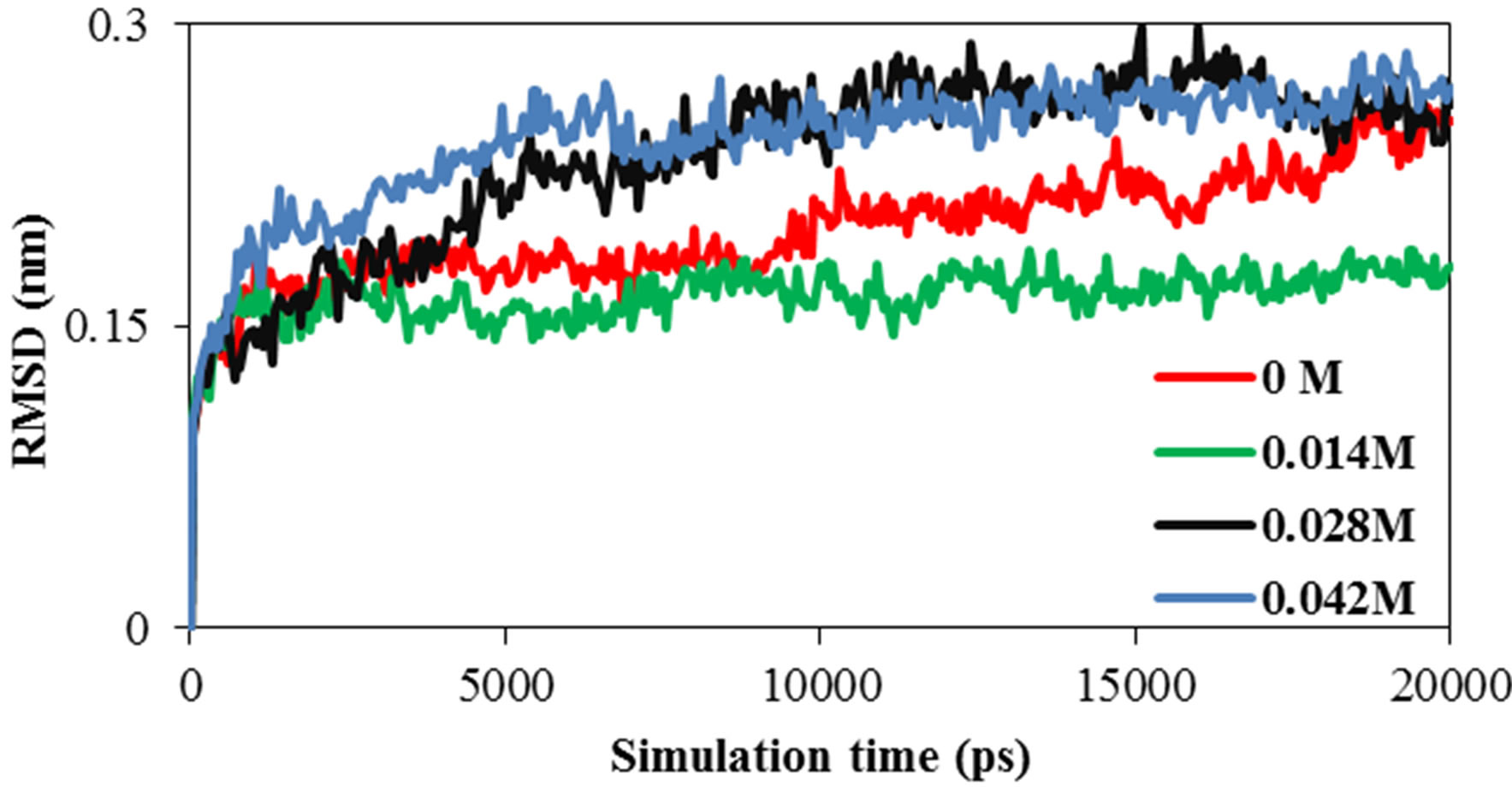 (b)
(b)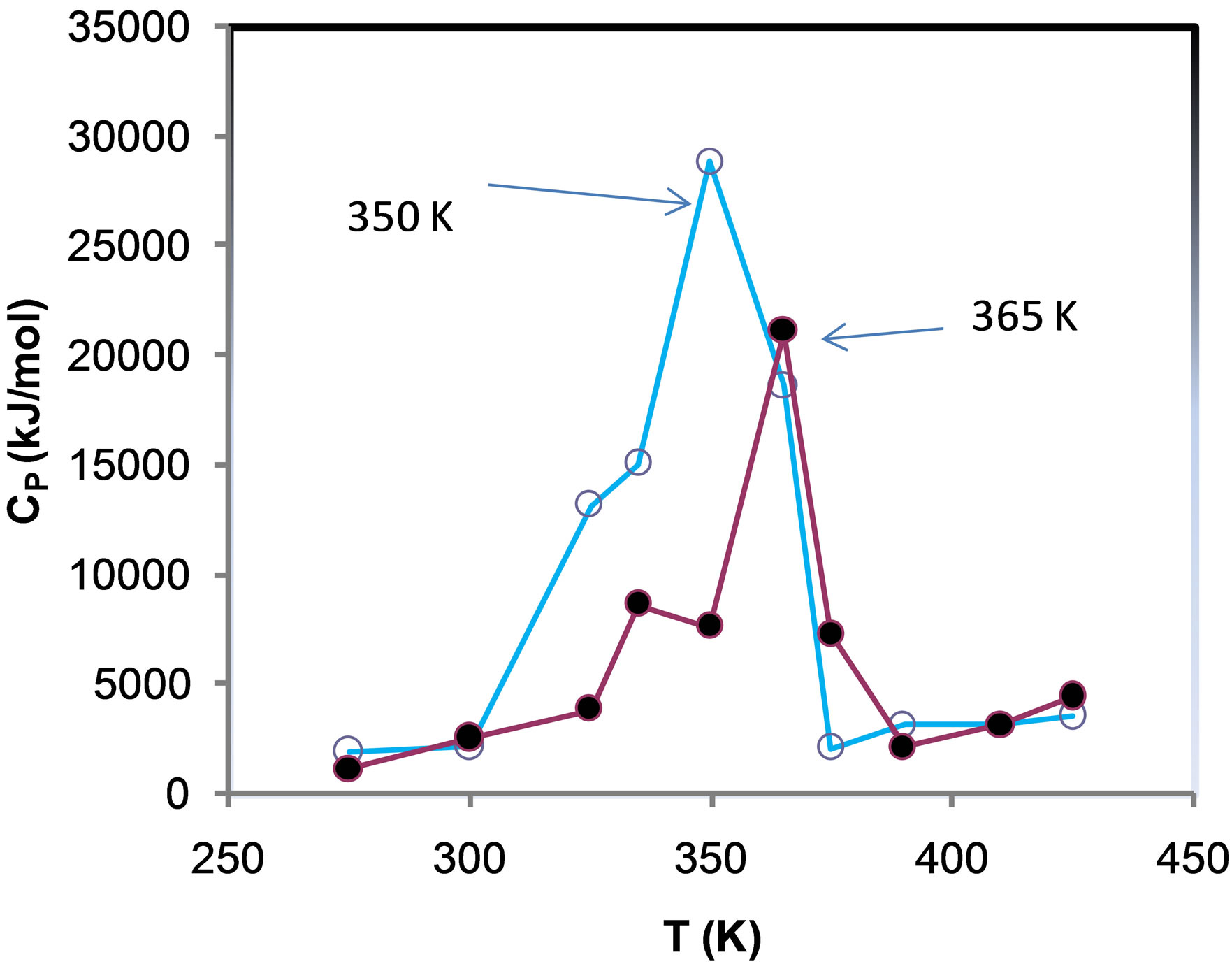 (c)
(c)
Figure 1. Root mean square deviation of ADA structure (a) in different temperature and (b) different concentration of NiTNasPc (c) DSC profile of ADA in the absence (open circle) and the presence (fill circle) of 0.014 M of phthalocyanine.
figure shows that ADA has more structural changes (RMSD) at higher temperature and concentration except at lower concentration which has lower RMSD. Selected concentration of Ni (II) tetra sodium sulphonated phthalocyanine (NiTNasPc) was 0 to 0.042 M. The structure information such as surface area, solvent accessible surface area, hydrogen bonding between ADA and solvent molecules, gyrate radius, RMSD and RDF were obtained at 300 K.
Root mean square deviation (RMSD) of the ADA in the presence of various concentrations of phthalocyanines was obtained. Figure 1(b) shows the RMSD of ADA in the 20 ns time interval in the presence of phthalocyanines. It shows that enzyme reaches a stable state after about 20 ns, (RMSD) and structural changes of system in the higher concentration of phthalocyanines is more than the lower concentrations.
Heat capacity in constant pressure (Cp) and different temperature were obtained from energy data. This data is same as differential scanning calorimetry (DSC).
The melting temperature of ADA in temperature range from 275 to 450 was about 350 K. Figure 1(c) shows DSC profile or representative thermogram in several temperatures for unfolding of ADA enzyme in the absence (a) and preence of 6 molecules (0.014 M) phthalocyanine. Maximum of peak is called the midpoint of temperature transition (Tm) that is 350 K and 390 K in the absence and presence of (0.014 M) phthalocyanine, respectively. So the phthalocyanine increases the ADA stability in this concentration.
Figure 2 shows the average values of RMSD, solvent accessible surface area (SAS), interaprotein hydrogen bond HB (p-p), protein-solvent hydrogen bond HB (p-sol), radius of gyration (Rg), helix, coil and beta percentage in different temperatures (left column) and different concentration (right column).
RMSD, Hydrophobic and total surface area, intraprotein HB, coil percentage (obtained by VADAR) increases, while protein-solvent HB, helix percentage decreases by temperature. It is due to denaturation and predominately unfolding of enzyme. Other parameters have irregular trends that may be due to existence of an intermediate or semi-stable structure.
Variation of solvent accessible surface area for ADA was computed by g_sas program. Variation of surface area in the 20 ns time evolution was significant and gained. Figure 2 shows increase of surface in the presence of higher concentration of NiTNasPc and decrease at low concentration.
Variation of solvent accessible surface area for ADA was computed by g_sas key. Variation of surface area in the 20 ns time evolution was significant and gained. Figure 2 shows the average surface area of ADA in the absence and presence of 0.014 to 0.042 M NiTNasPc. This
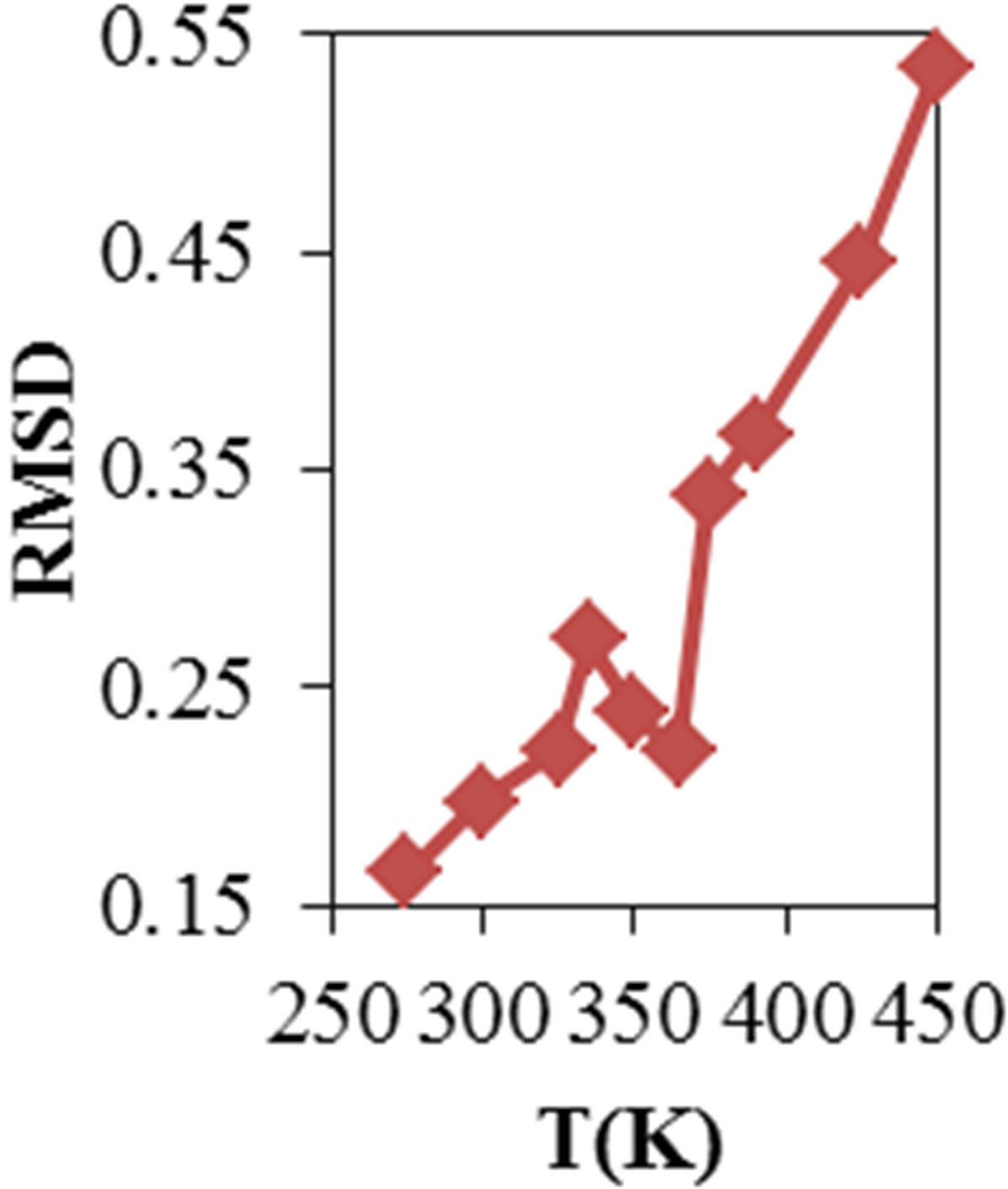
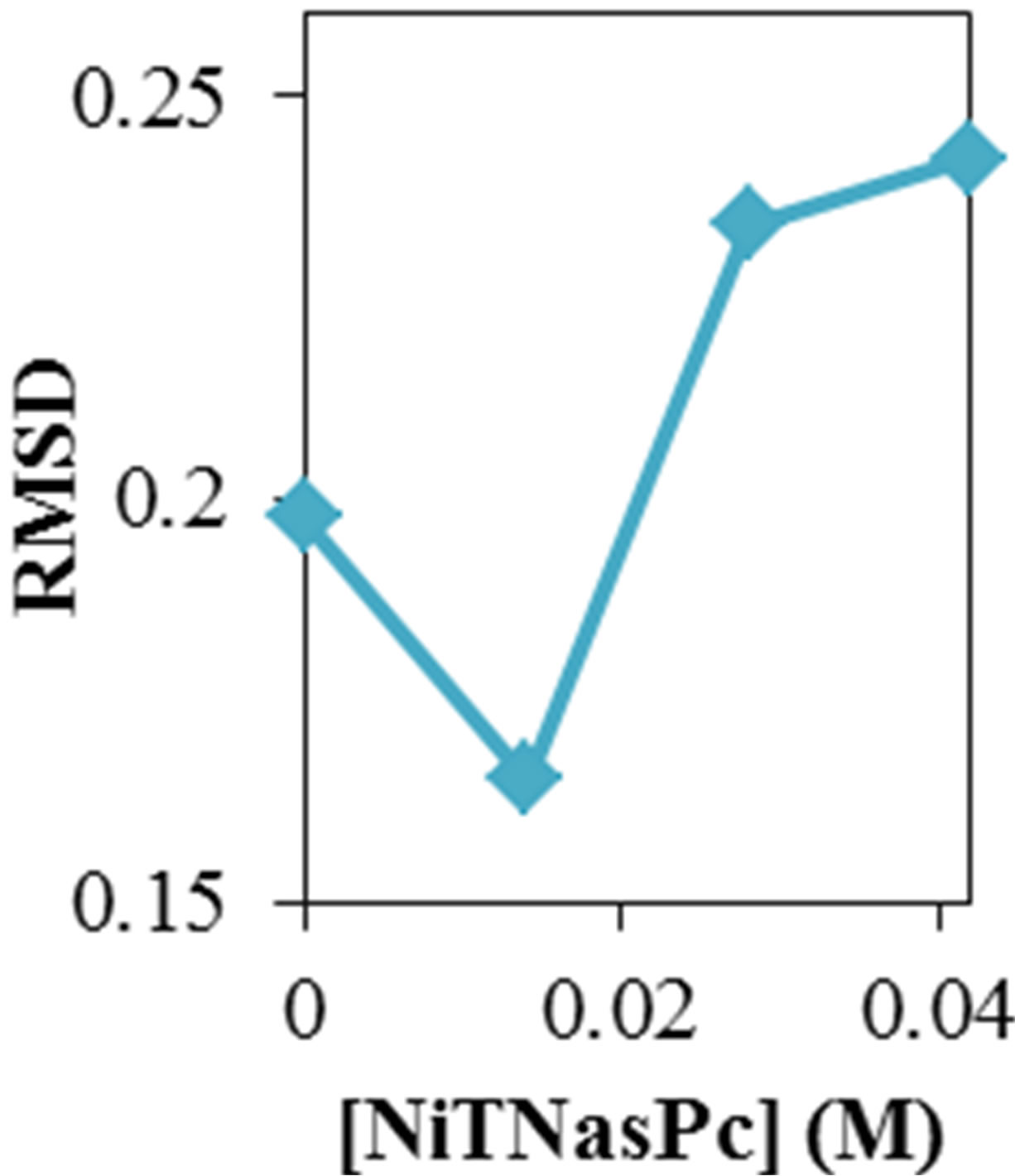
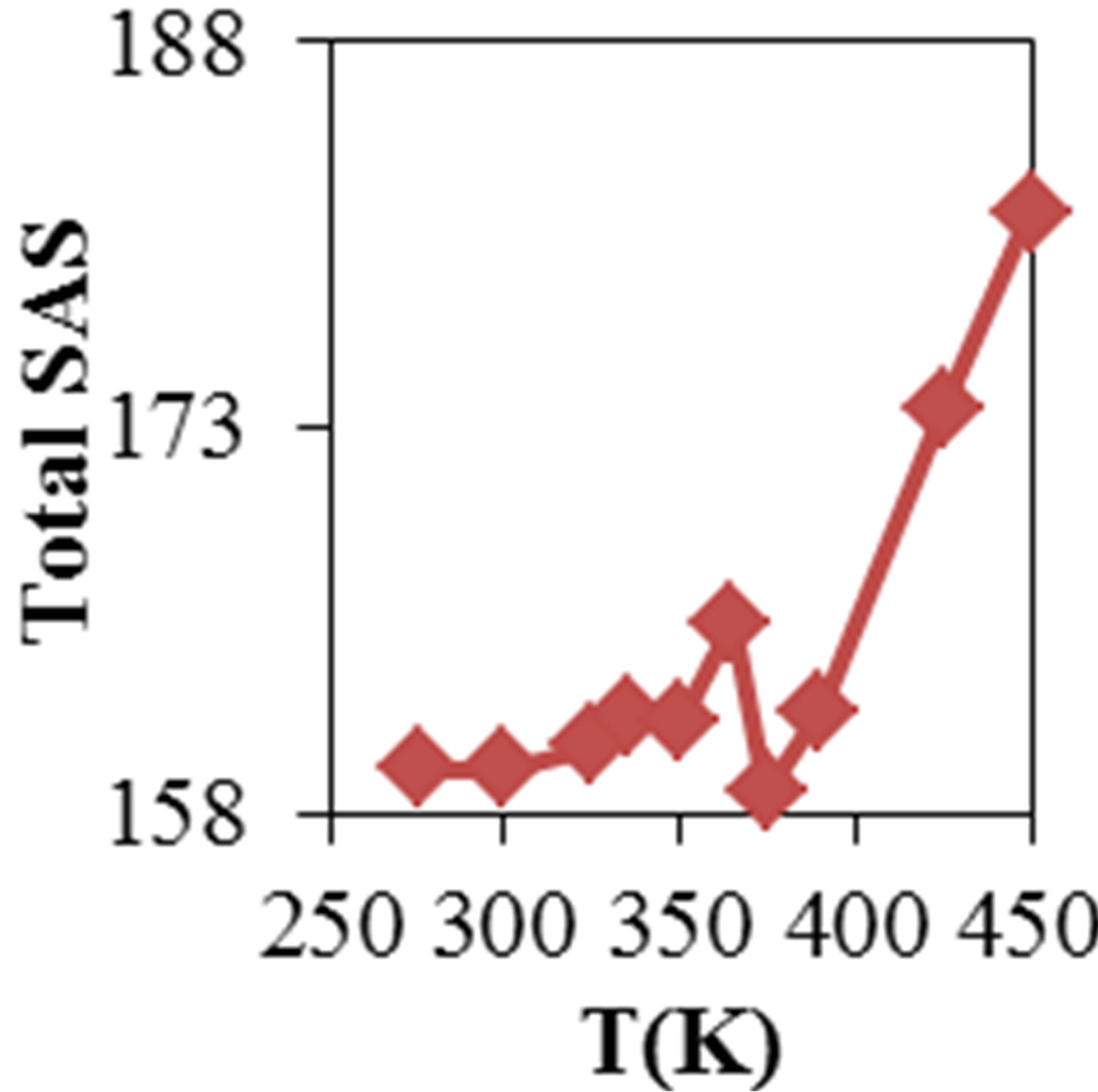
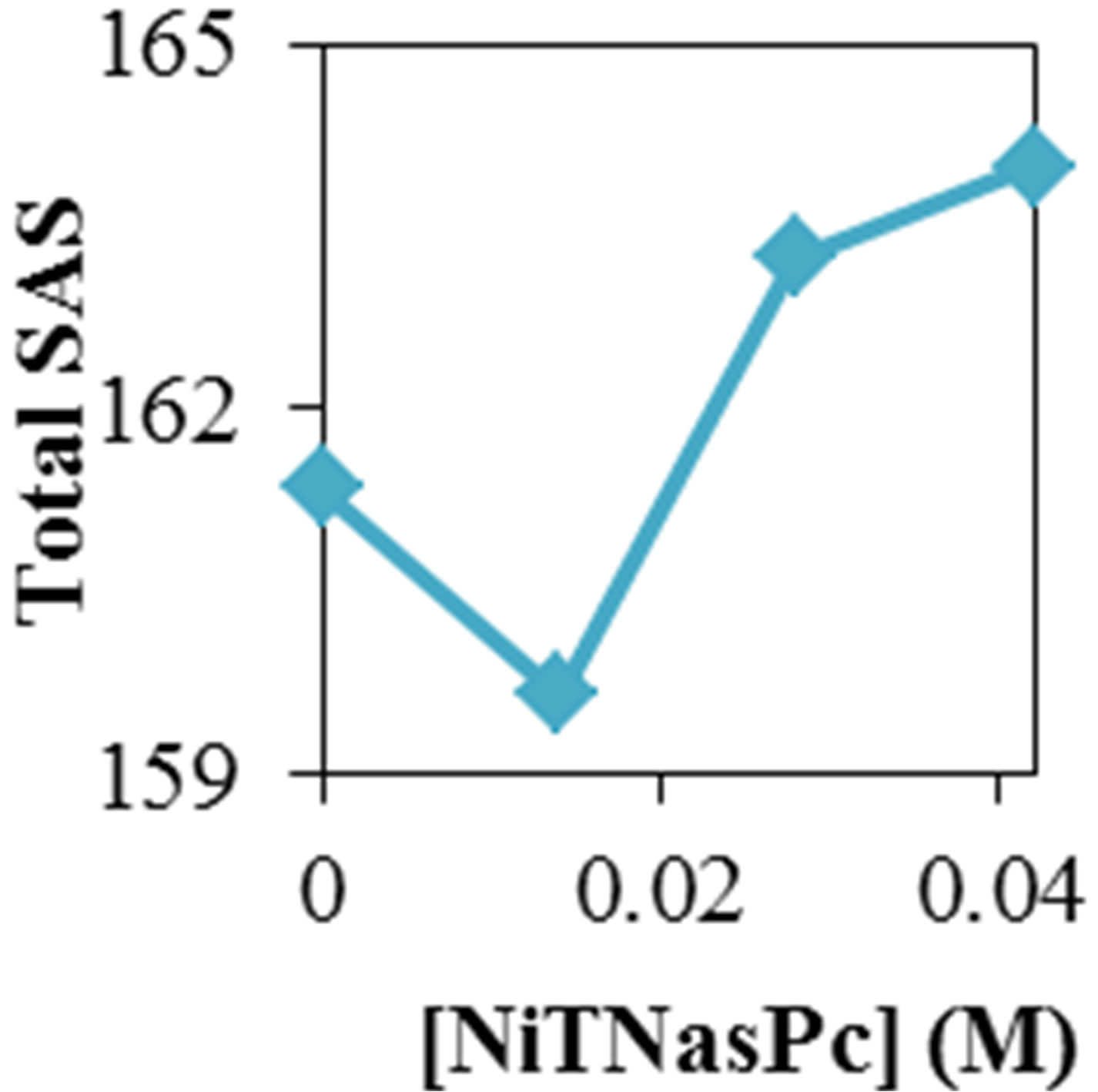
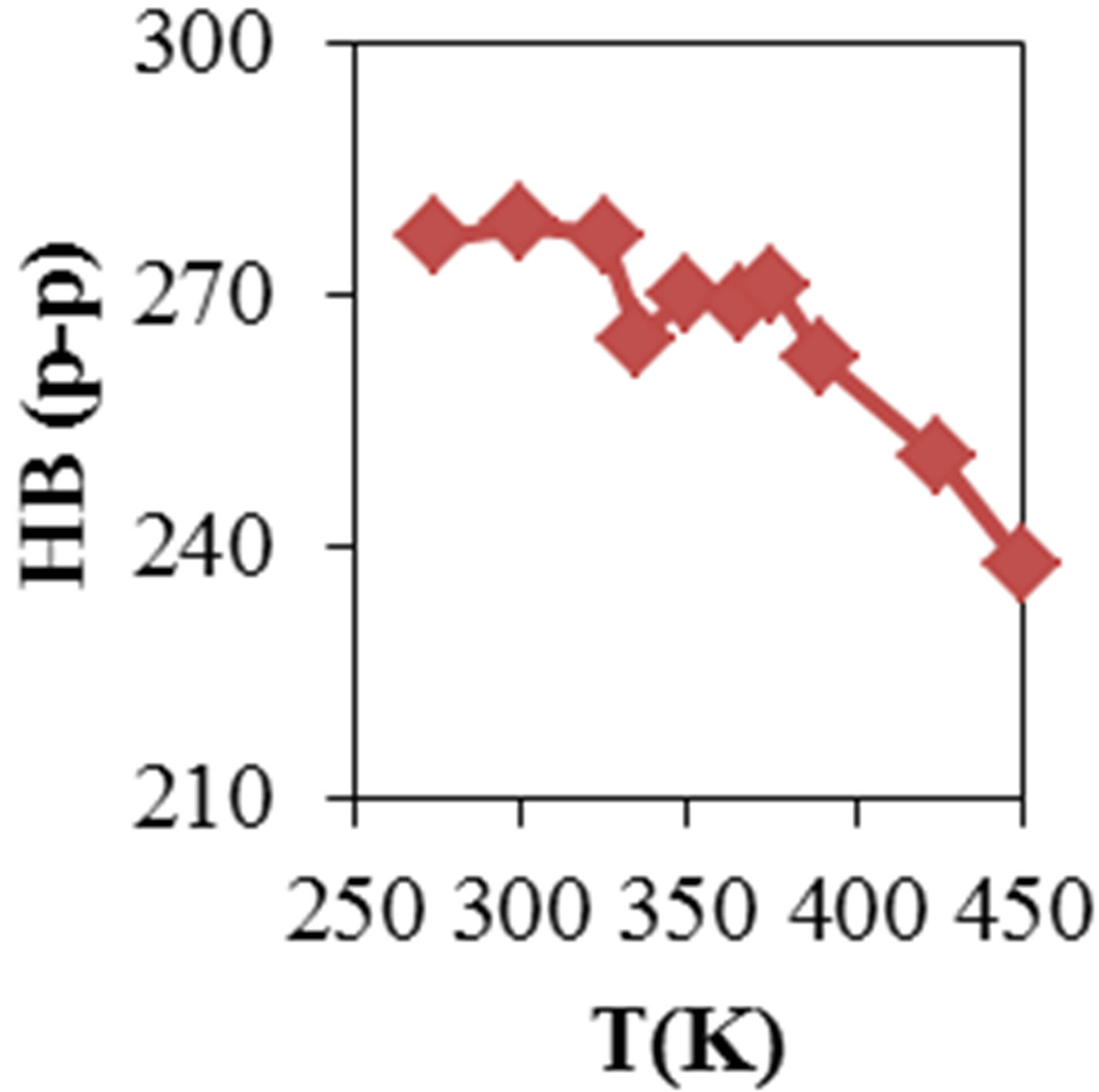
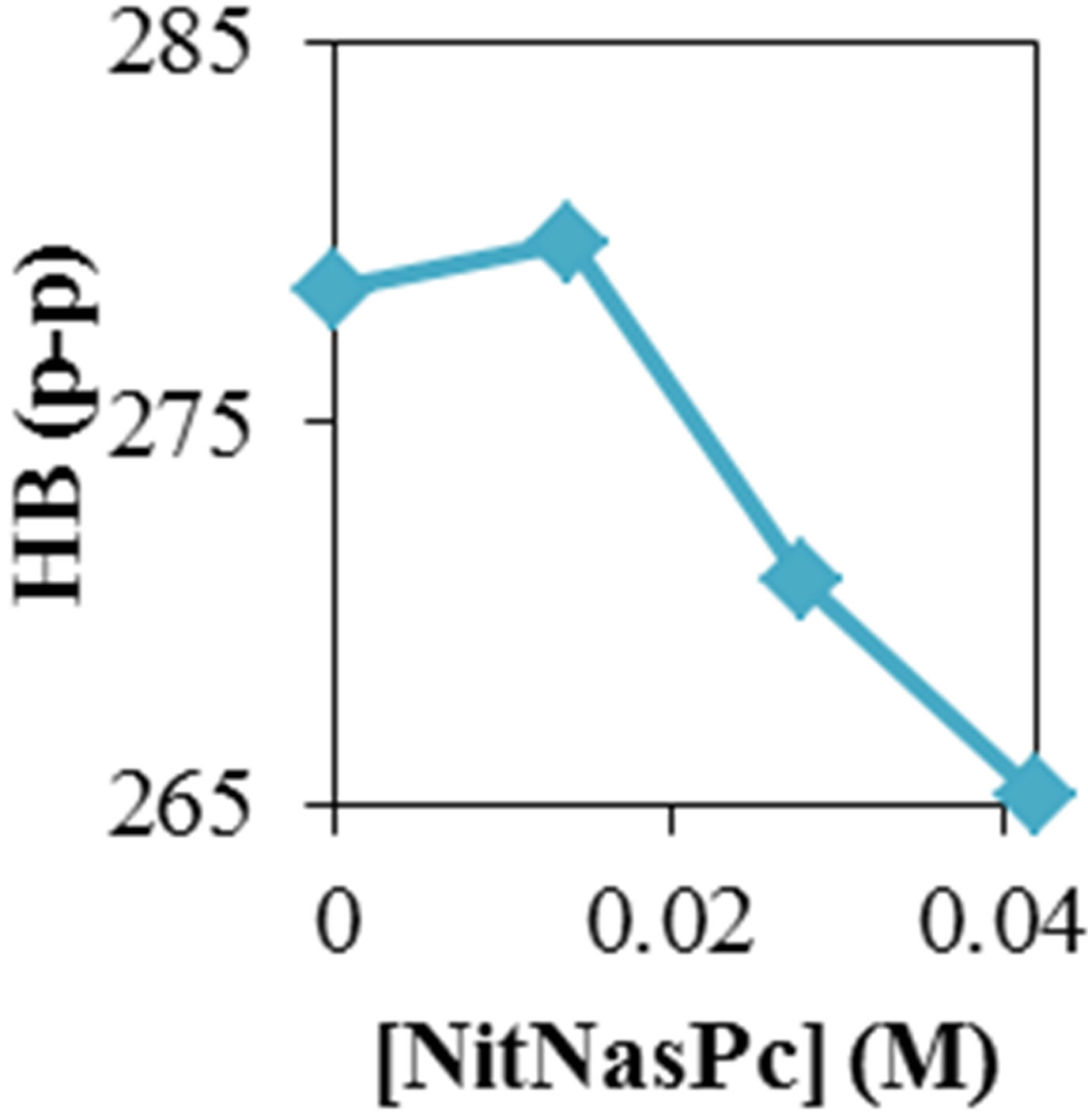
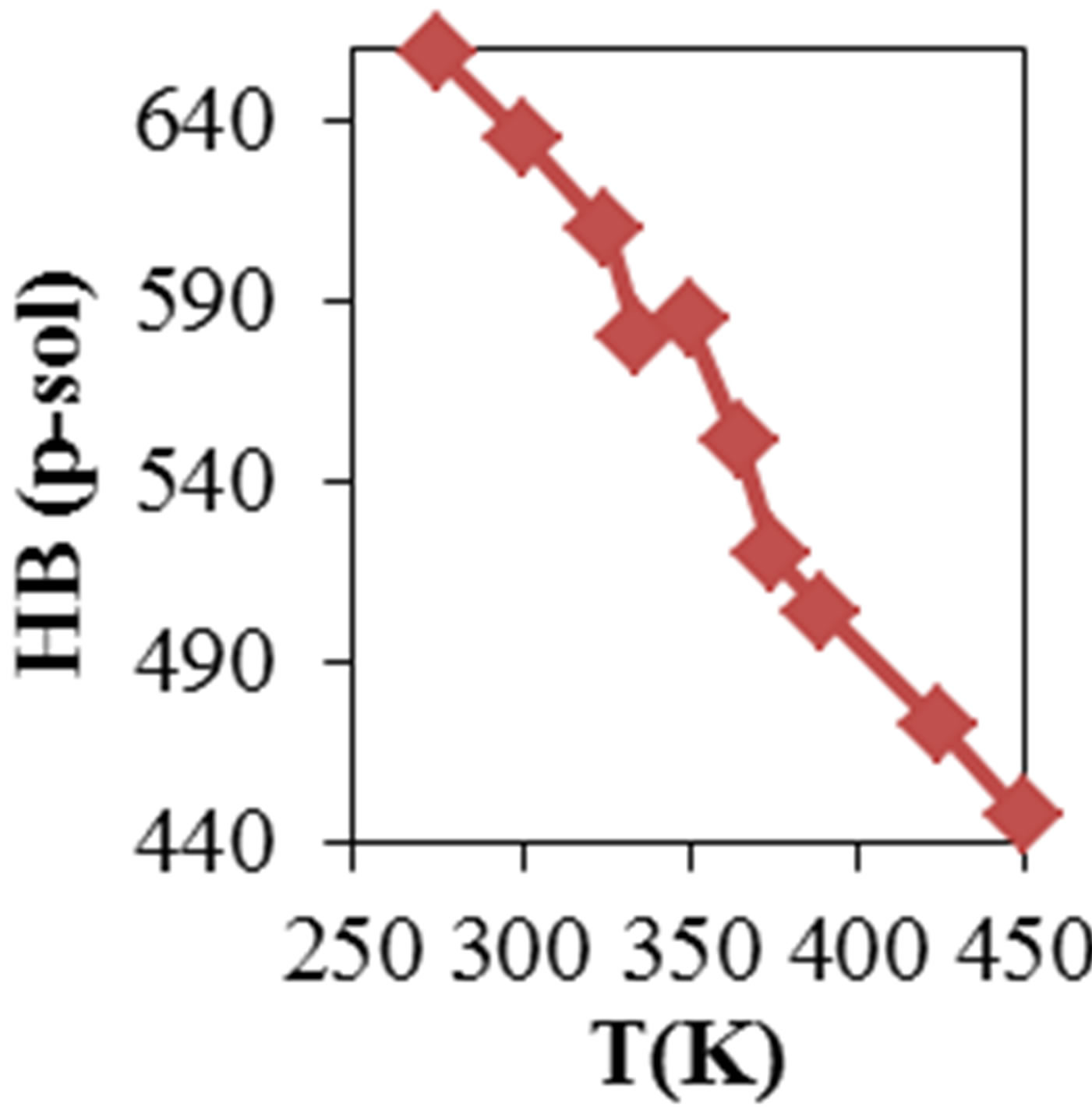
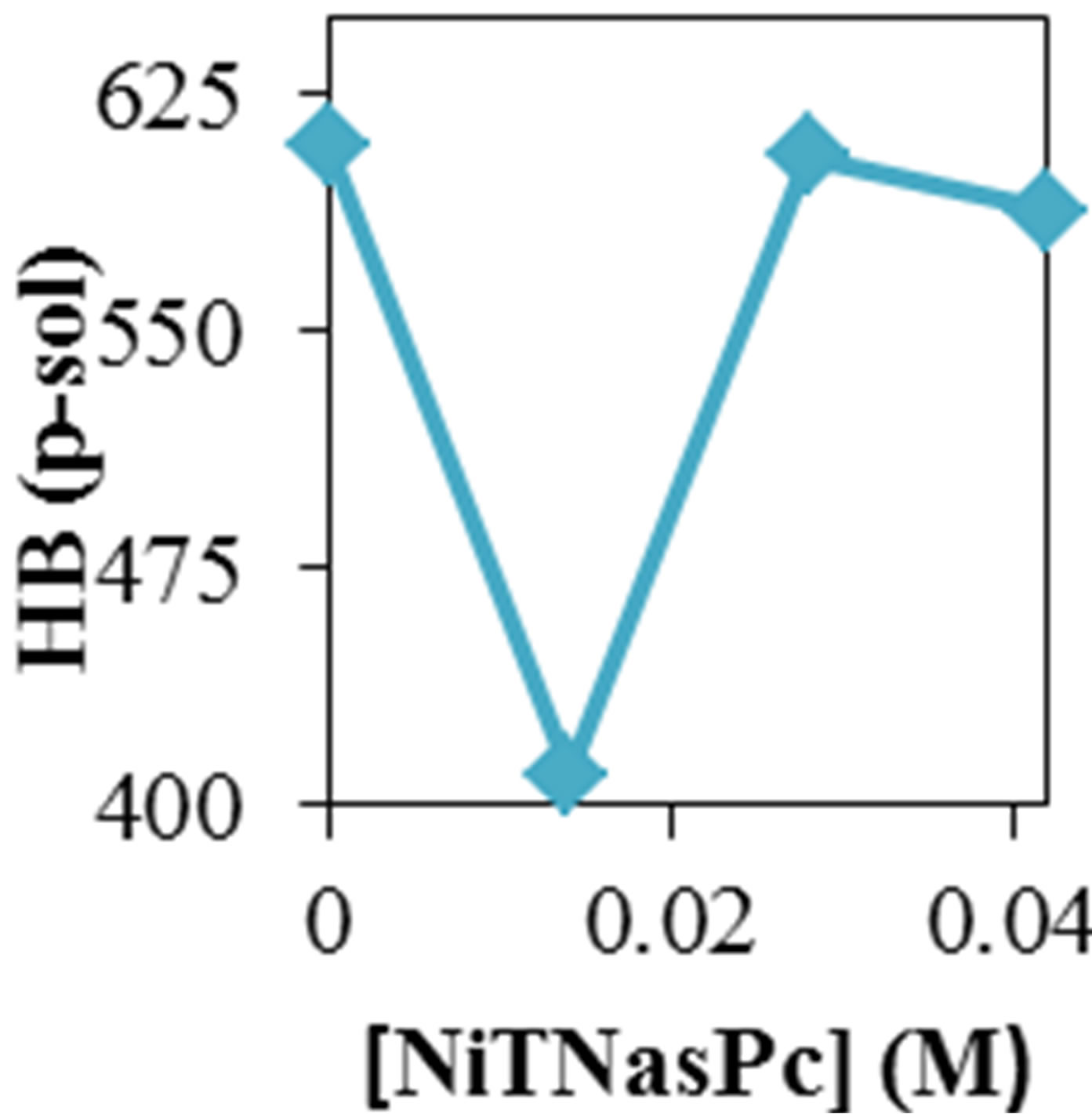
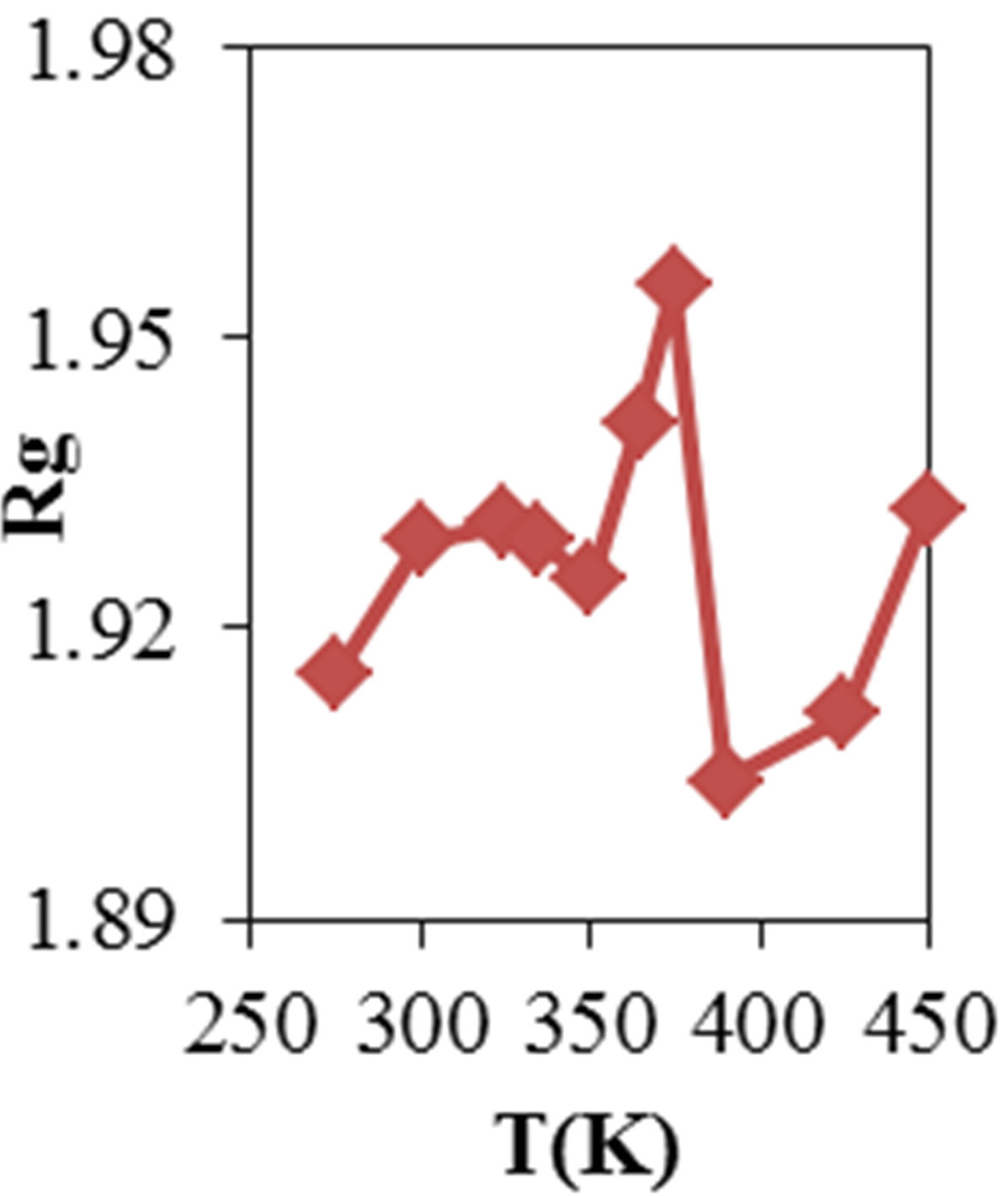
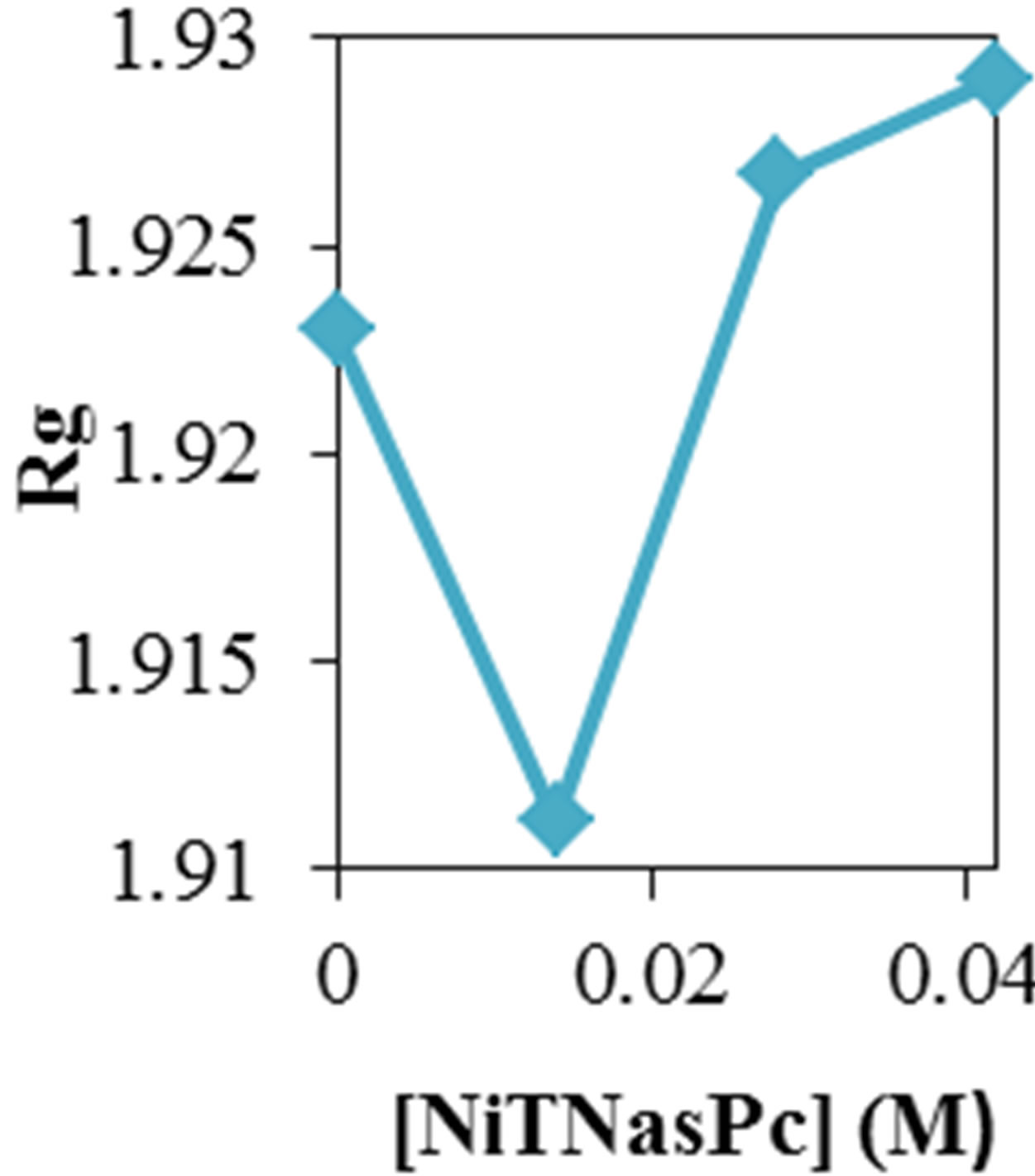
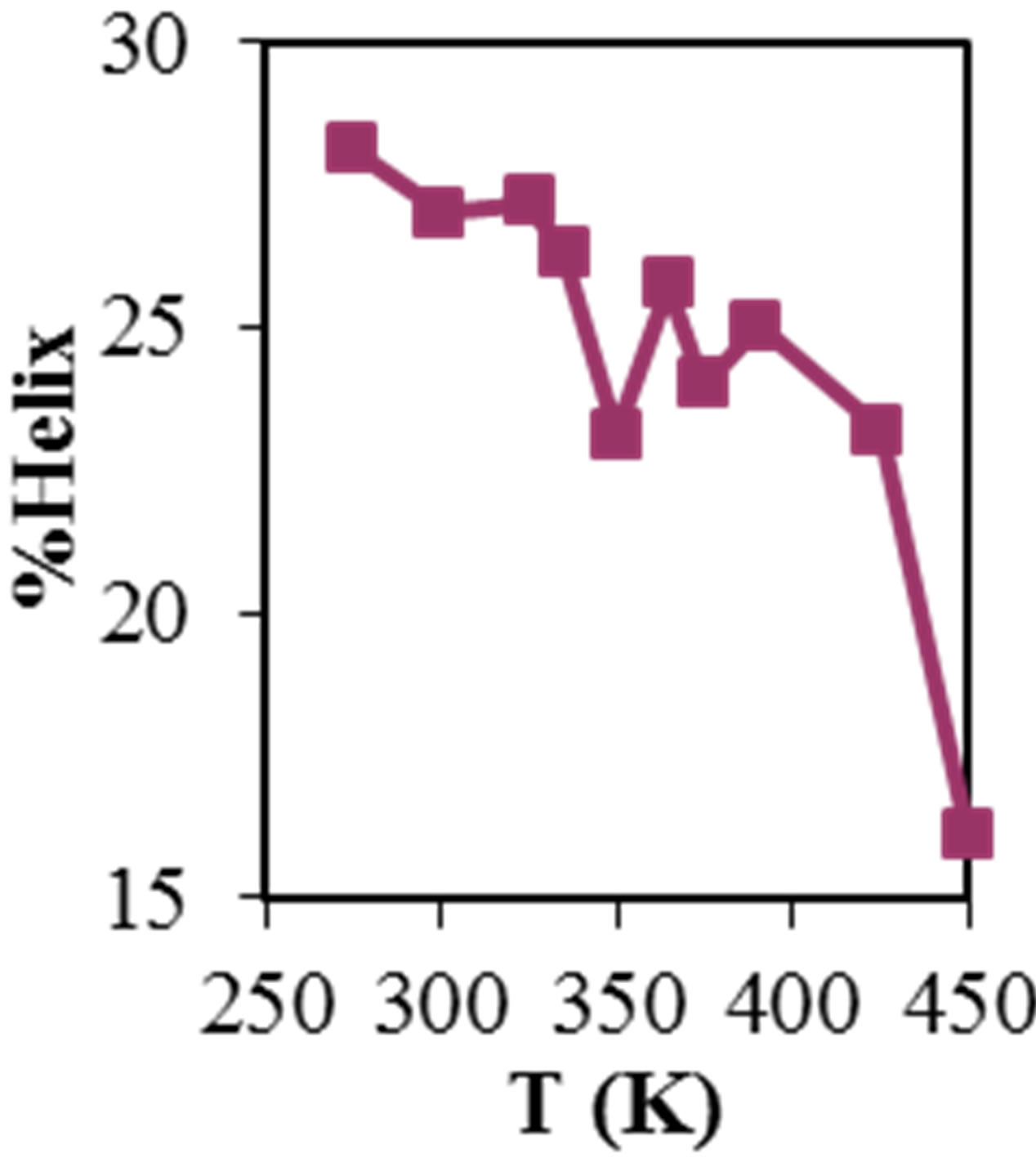
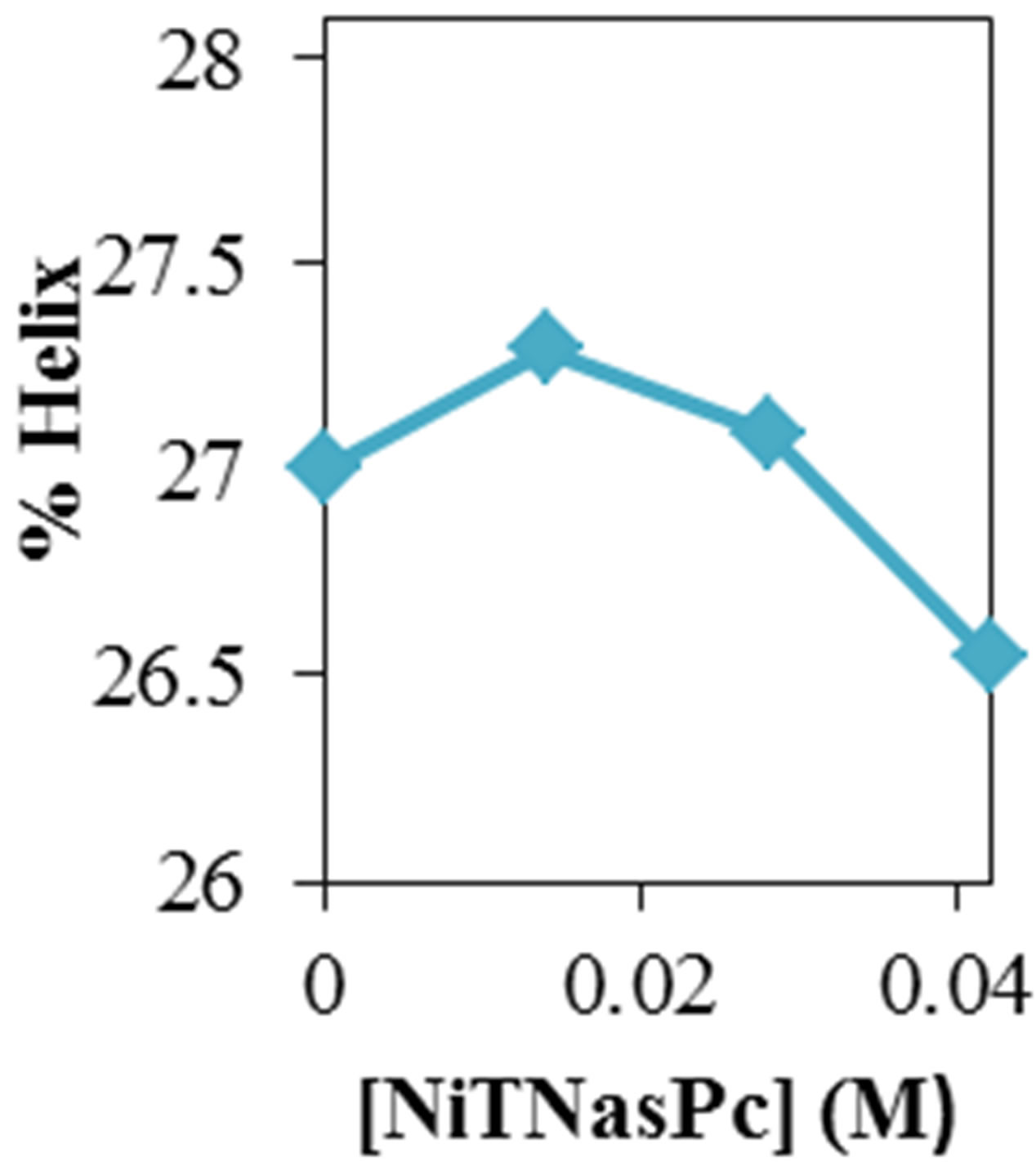
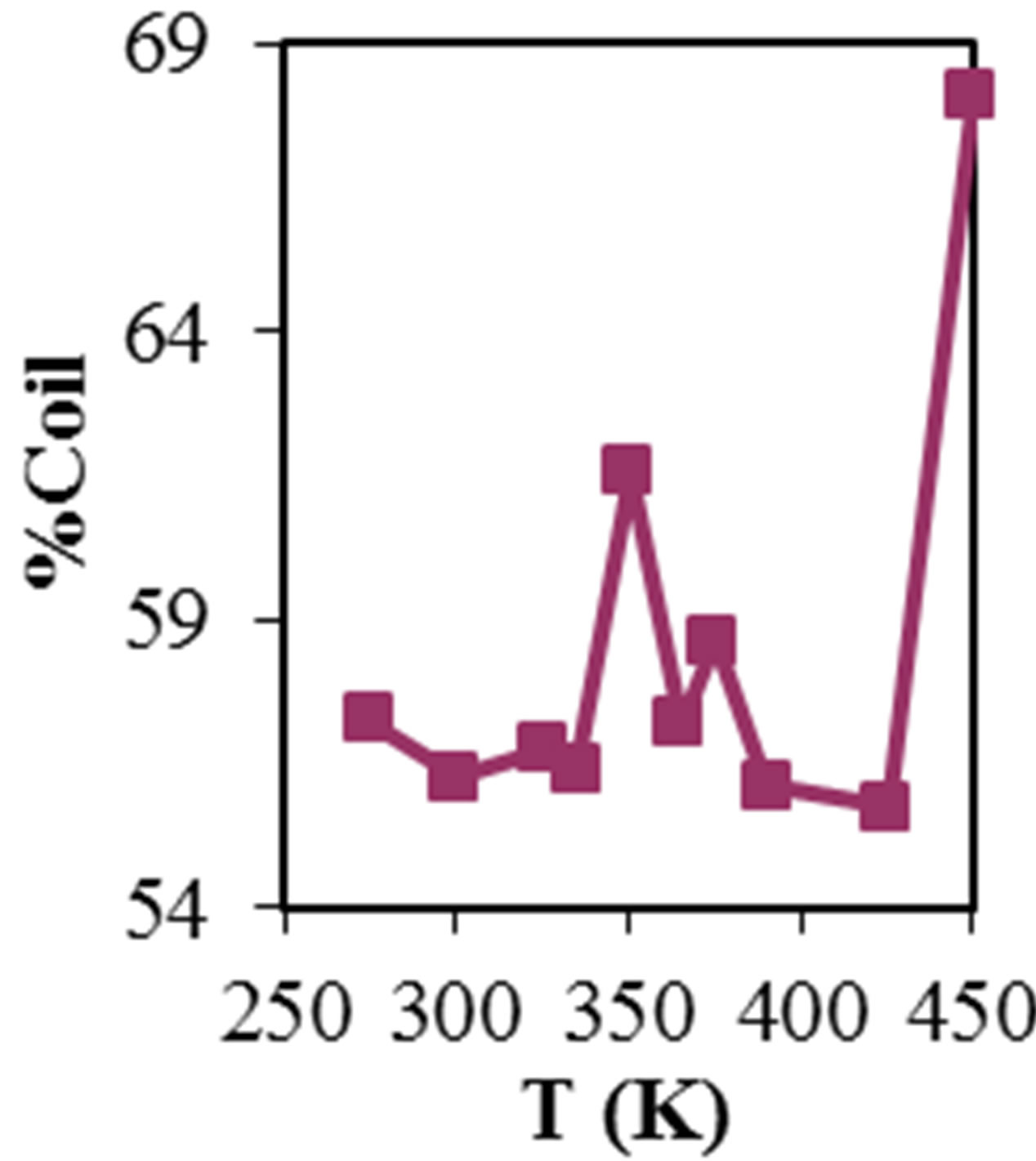
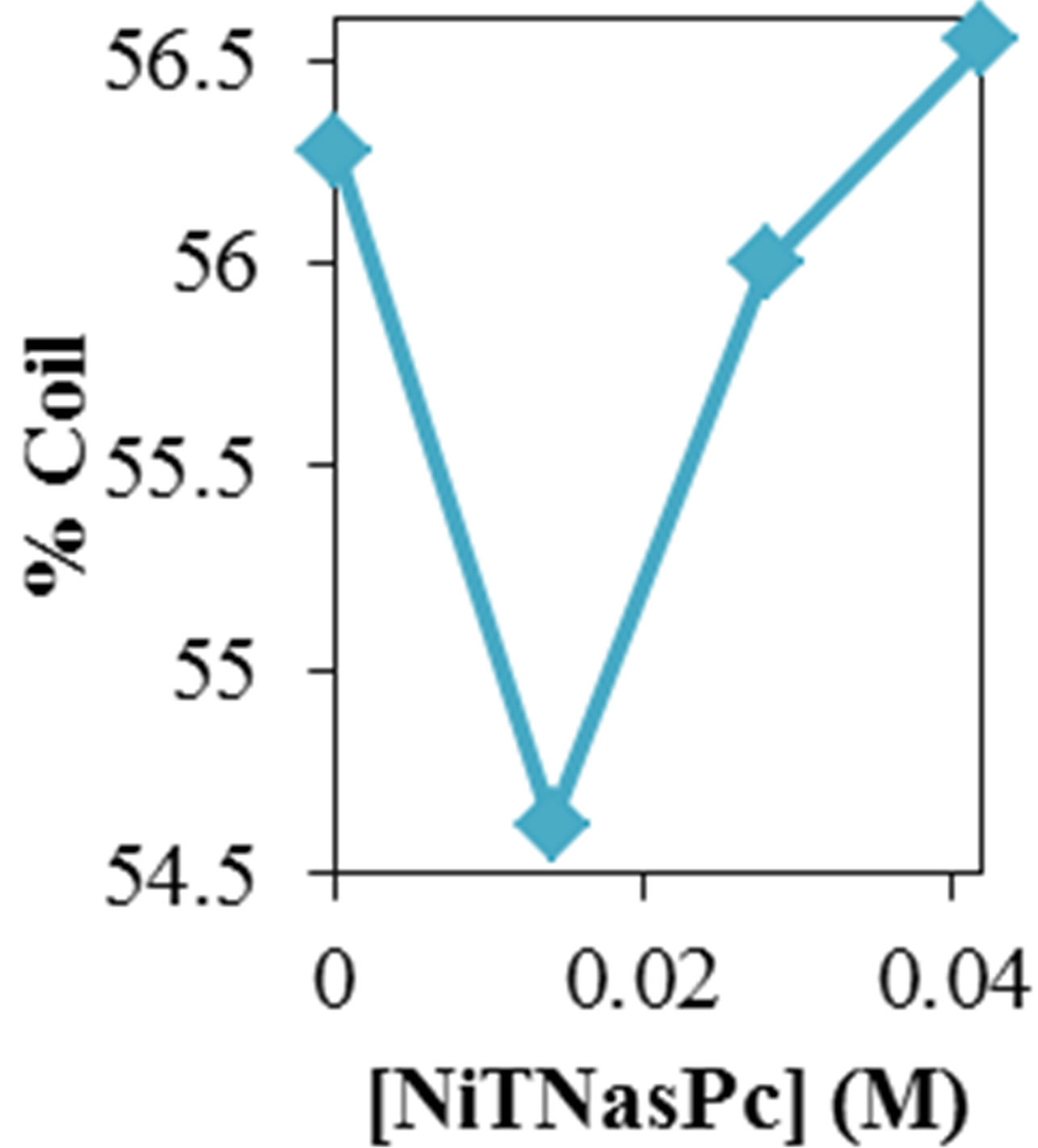
Figure 2. ADA structural changes during 20 ns at different temperatures (left) and in the presence of different concentration of NiTNasPc at 300 K.
figure shows increase of surface in the presence of higher concentration of NiTNasPc. This result is in good accordance with reduction of intermolecular hydrogen bond between ADA and proves the ADA has been unfolded and obviously surface area of ADA in system with high concentration of NiTNasPc is increased more because of more reduction of intermolecular hydrogen bond of ADA.
Figure 2 also shows the radius of gyration (Rg) for ADA in the 20 ns time interval in the absence and presence of 0.014, 0.028 and 0.042 M NiTNasPc respectively. This figure shows increase of Rg for ADA in the presence of high concentration of NiTNasPc and decrease in the low concentration. This proves that ADA has been unfolded and obviously surface area and therefore Rg for ADA in system with high concentration of NiTNasPc is increased more because of more increase of surface area of ADA.
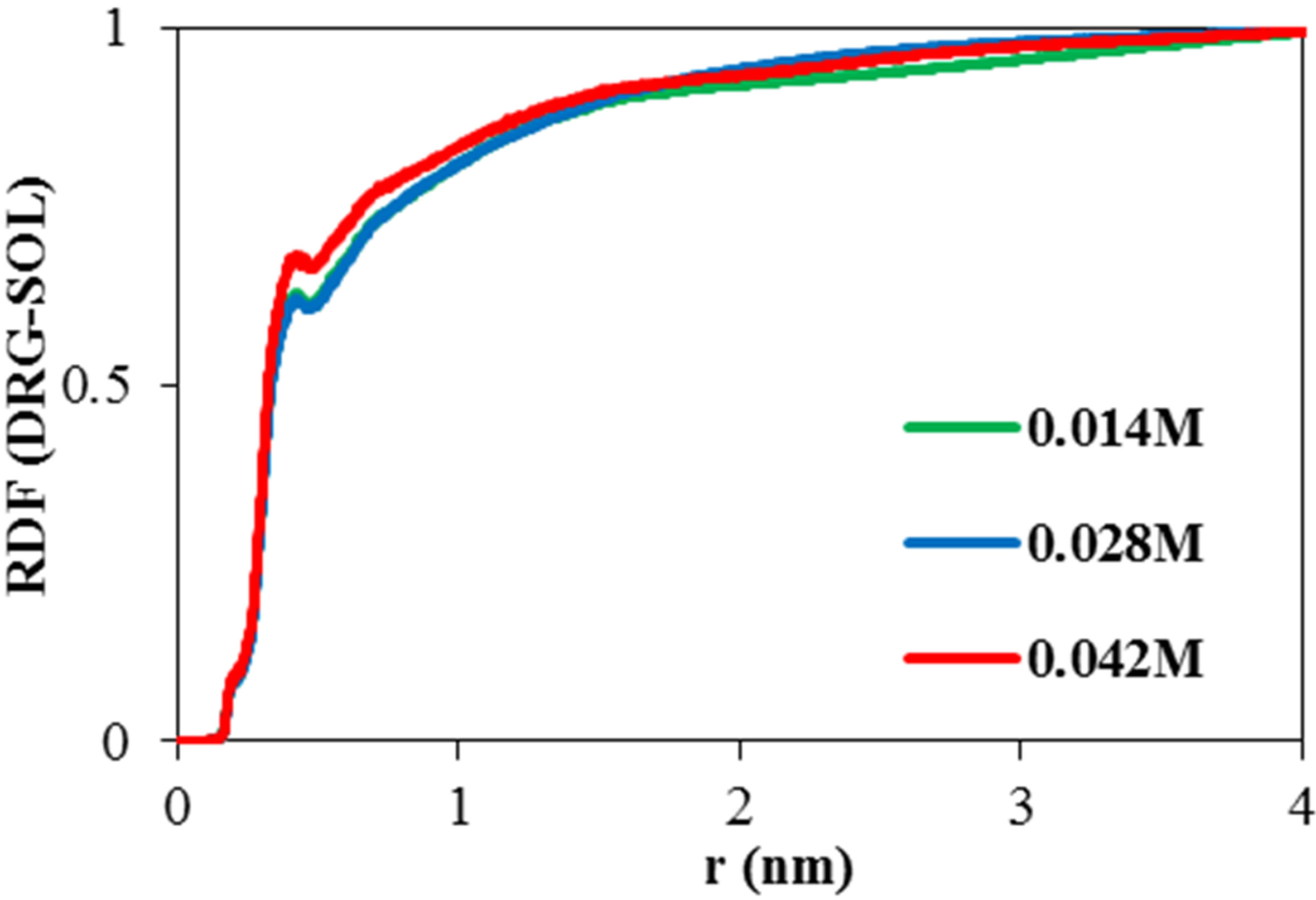
Radial distribution function (RDF) is a criterion for distributed atoms, molecules or other species around target specie. The variation of solvent RDF around ADA versus distance is shown in Figure 3(a) in various concentrations of NiTNasPc. It can be seen that RDF is decreased for the phthalocyanine by increasing the concentration. At high concentration, solvent becomes far away from ADA and this result is in good agreement with hydrogen bond results in Figure 2. An increase of RDF in aqueous solution with a decrease in self-aggregation is usually viewed. Existence probability (RDF) of NiTNasPc around each other is depicted in Figure 3(b). This figure shows that by increasing NiTNasPc concentration, aggregation decreases too.
 (a)
(a)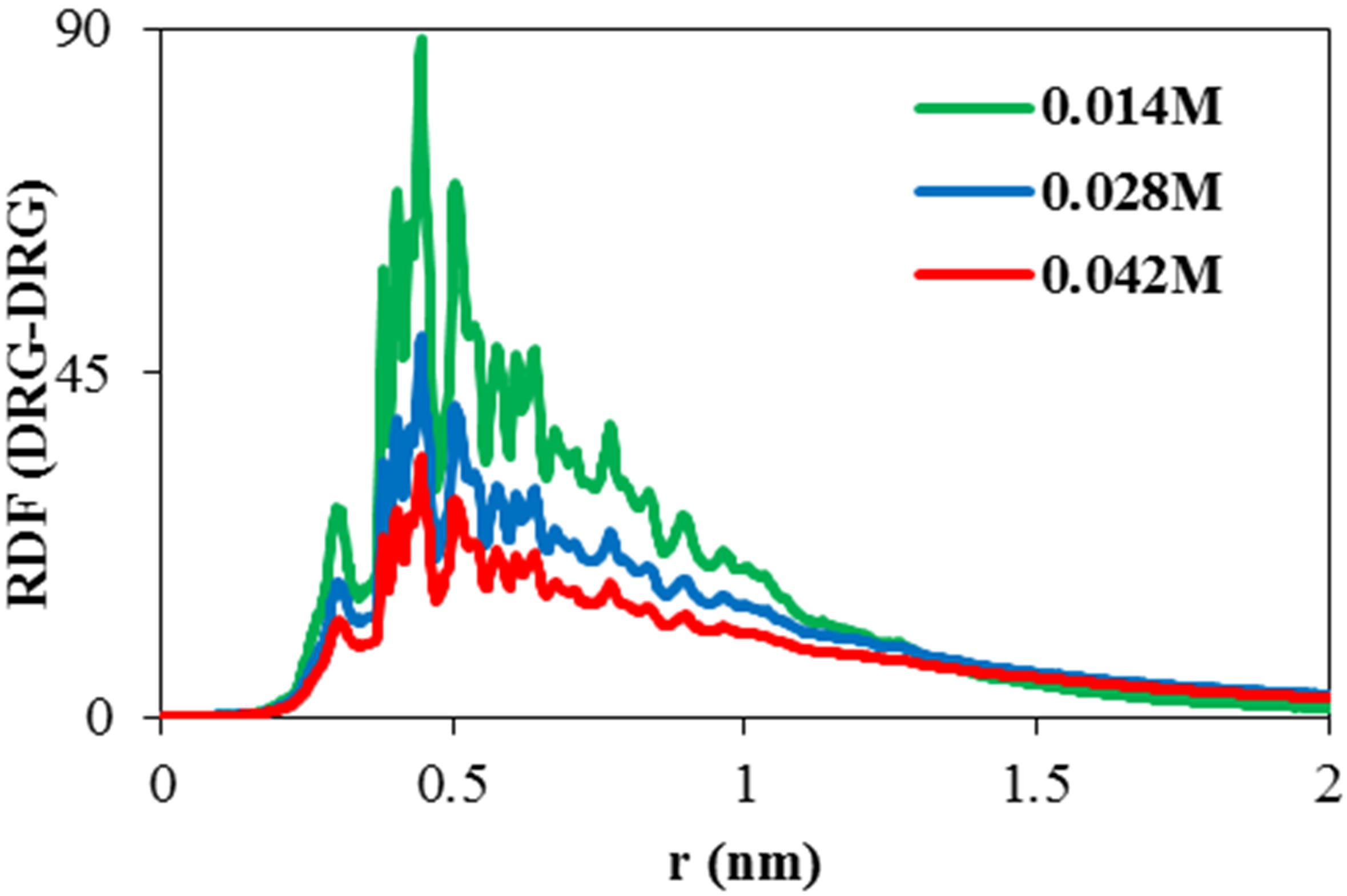 (b)
(b)
Figure 3. (a) Calculated RDF for NiTNasPc-Solvent (b) RDF of NiTNasPc-NiTNasPc by molecular dynamics in the presence of 0.014, 0.028 and 0.042 M NiTNasPc, (DRG = NiTNasPc).
3.2. Comparison of Metal and Non-Metal Phthalocyanines
Ni(ІІ) in the aromatic rings of phthalocyanine plays an important role in Pcs effect on ADA and its self aggregation. Metal causes circulation of charge in the rings and so increasing the phthalocyanines aggregation. Structural parameters such as variation of SAS, hydrogen bond, Rg and other physical parameters from analyzing trajectory of molecular dynamics were compared.
Variation of SAS for hydrophobic part of ADA was computed by g_sas program. Variation of surface area in the 20 ns time evolution was significant and obtained. Figure 4(a) shows the surface area of ADA in the 20 ns time interval in the presence of TNasPc and NiTNasPc. This figure shows increase of surface in the presence of TNasPc. This proves the ADA has been unfolded more in system with TNasPc. In conclusion ADA is more stable in the presence of NiTNasPc. Metal can increase enzyme stability. Figure 4(b) shows the variation of hydrogen bond of ADA in the 20 ns time interval in the presence of NiTNasPc and TNasPc. This figure shows decrease of interamolecular hydrogen bond of ADA in the presence of TNasPc and this result is in good accordance with increase of hydrophobic surface area of ADA. It means that TNasPc unfold enzyme more than NiTNasPc, therefore Ni causes more enzyme stability.
Figure 4(c) shows the Rg of ADA in the 20 ns time interval in the presence of TNasPc and NiTNasPc. This figure shows decrease of Rg for ADA in the presence of NiTNasPc and this result is in good accordance with decrease of SAS and increase of inter-molecular hydrogen bond of ADA. This proves the ADA has been unfolded more in the presence of TNasPc and therefore radius gyration of ADA in system with TNasPc is increased more than the system with NiTNasPc.
The radial distribution function of TNasPc around TNasPc and NiTNasPc around NiTNasPc is shown in Figure 4(d). It can be seen that RDF is increased for NiTNasPc. This result is in good agreement with hydrogen bond results. An increase of RDF in NiTNasPcs with increase of self aggregations is usually observed. It means that NiTNasPc tend more to self aggregate.
3.3. Comparison of Cationic and Anionic Phthalocyanine
In order to study effect of cationic and anionic Pcs on ADA denaturation, TNasPc and tetrakis [2-(trimethylammonium)ethoxy] phthalocyanine was used to calculate the stability of the protein.
The structure of ADA which has been optimized during 20 ns was optimized separately in the presence of 6 (0.014 M) molecules of anionic and cationic phthalocyanines in a period of 30 ns. Figures 5(a) and (c) show the
 (a)
(a)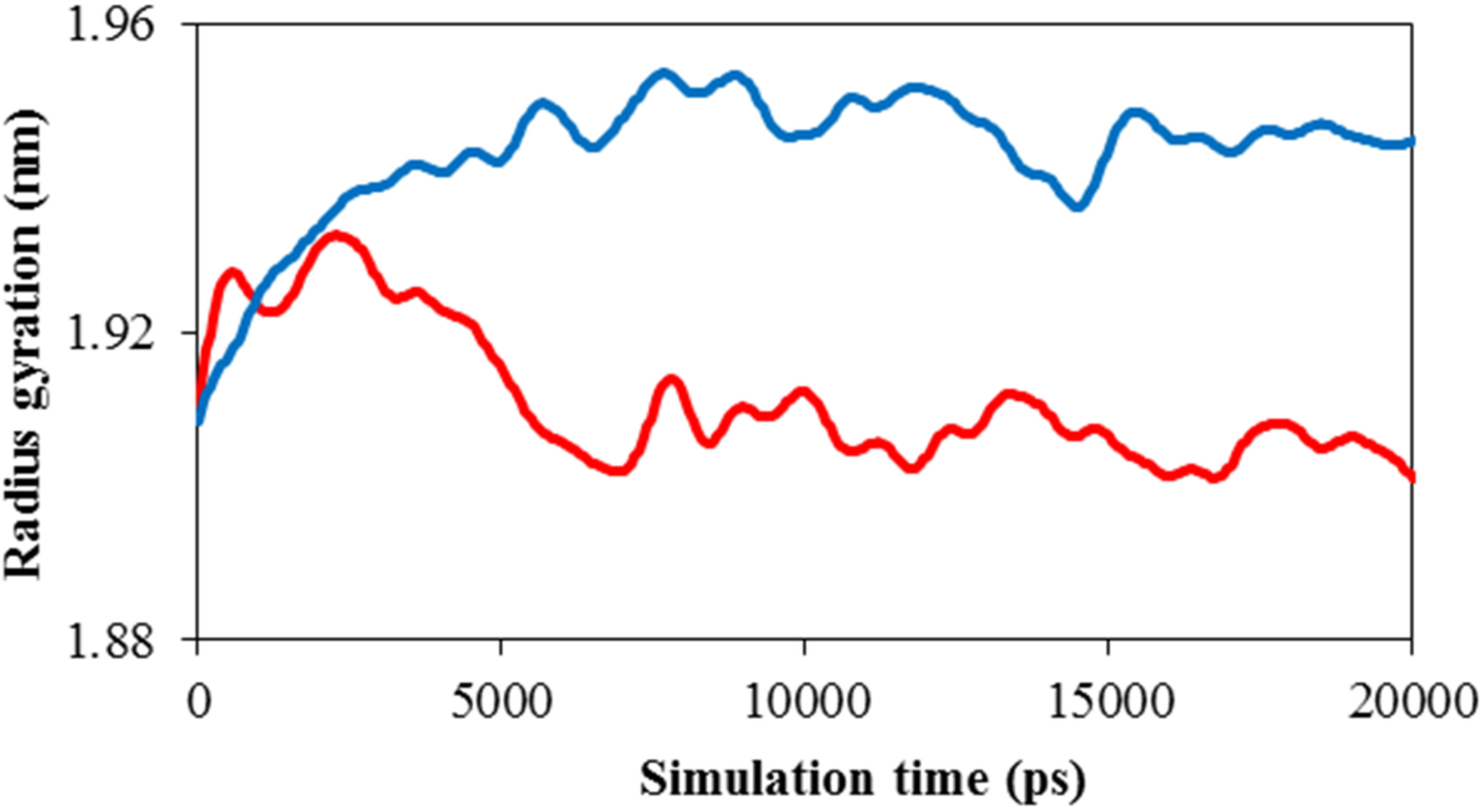 (b)
(b)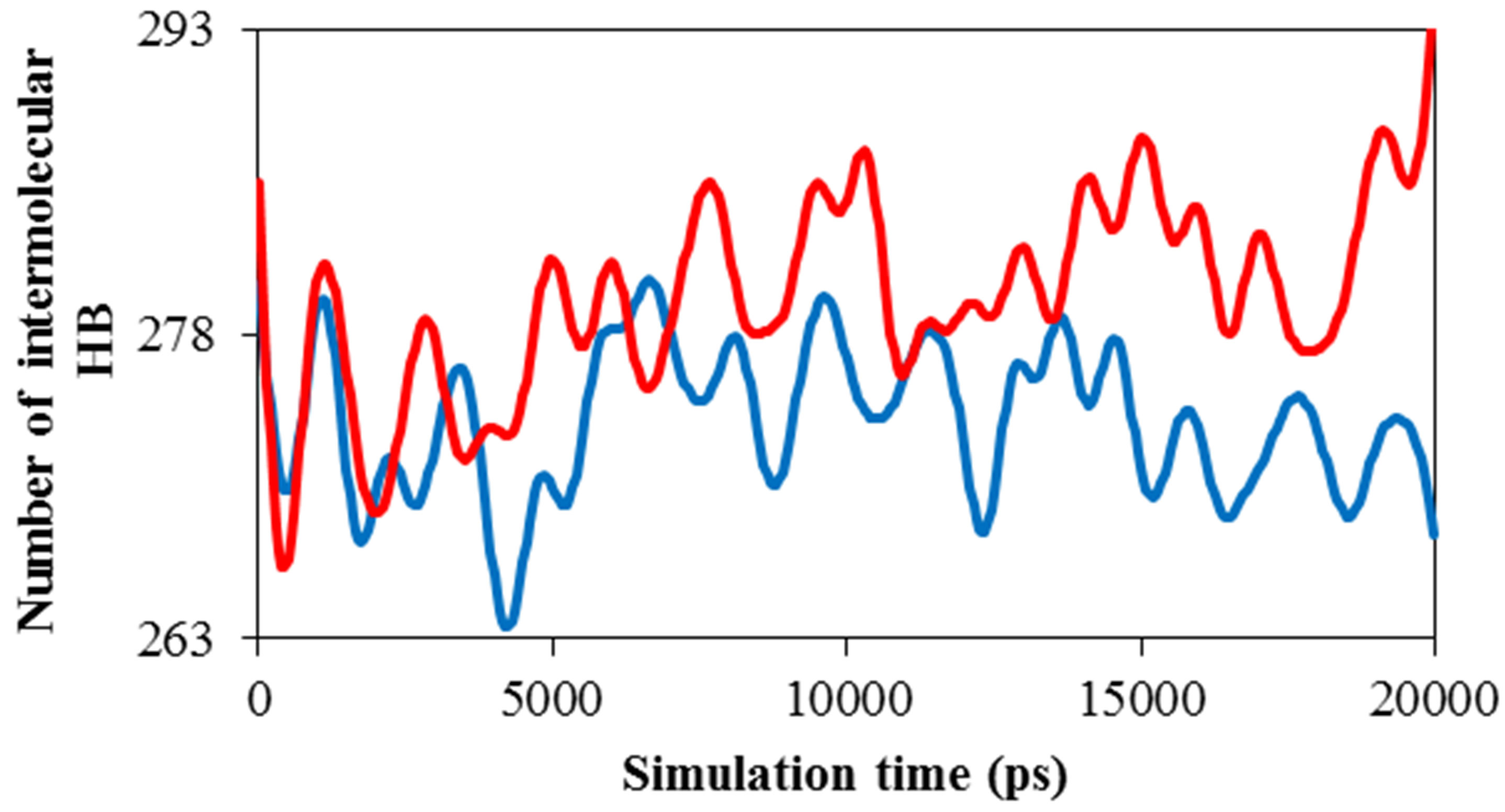 (c)
(c)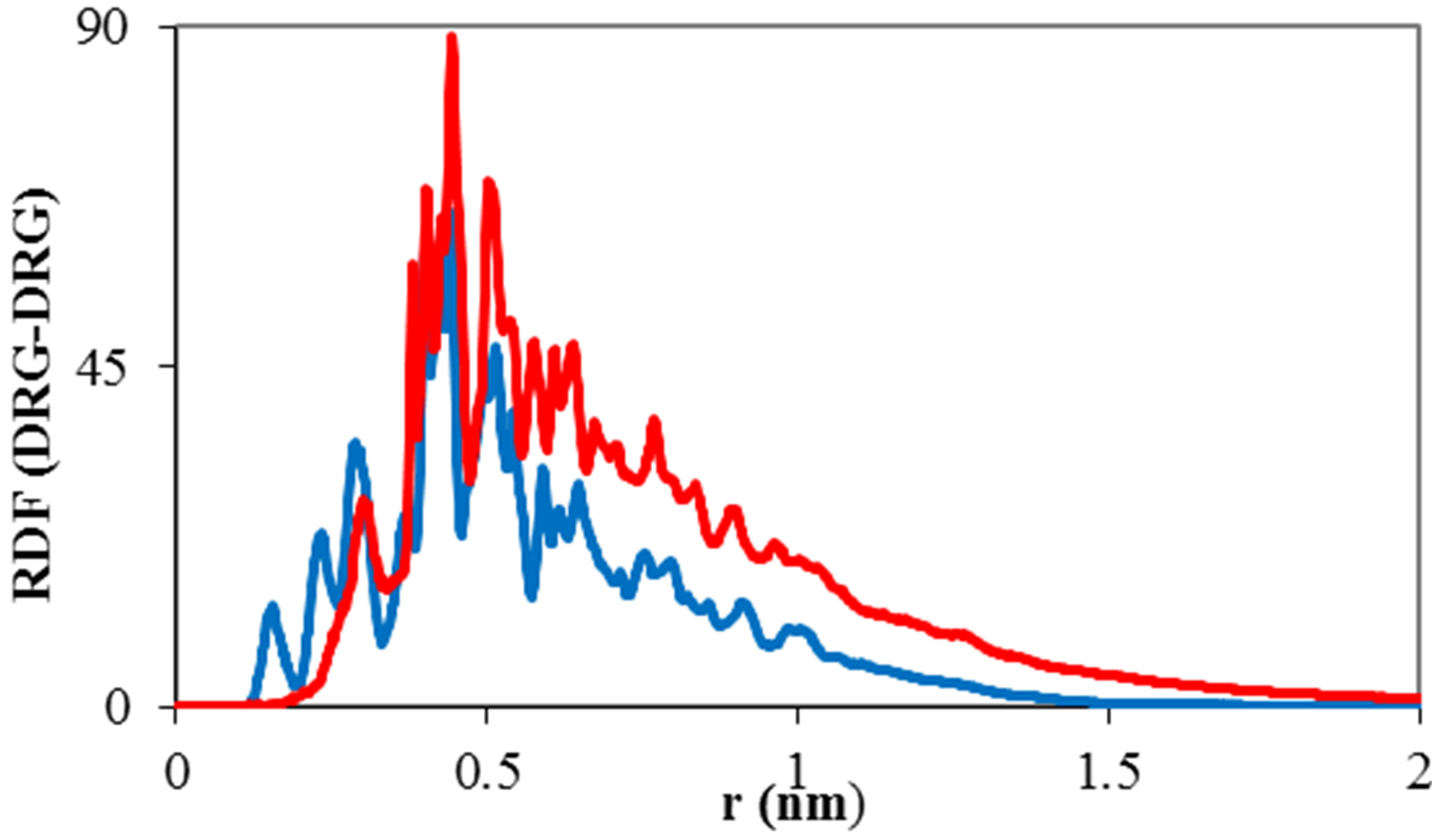 (d)
(d)
Figure 4. Calculated (a) solvent accessible surface area of hydrophobic part of ADA (b) ADA hydrogen bond by (c) Radius gyration of ADA (d) RDF of NiTNasPc-NiTNasPc and TNasPc-TNasPc by molecular dynamics in 0.014 M NiTNasPc (Red) and TNasPc (Blue).
starting point of the MD simulation for both phthalocyanines. The software automatically distributes ligands around the ADA. The final structure, after 20 ns, is shown in Figures 5(b) and (d). It shows that ligands tend to relocate to the specific site.
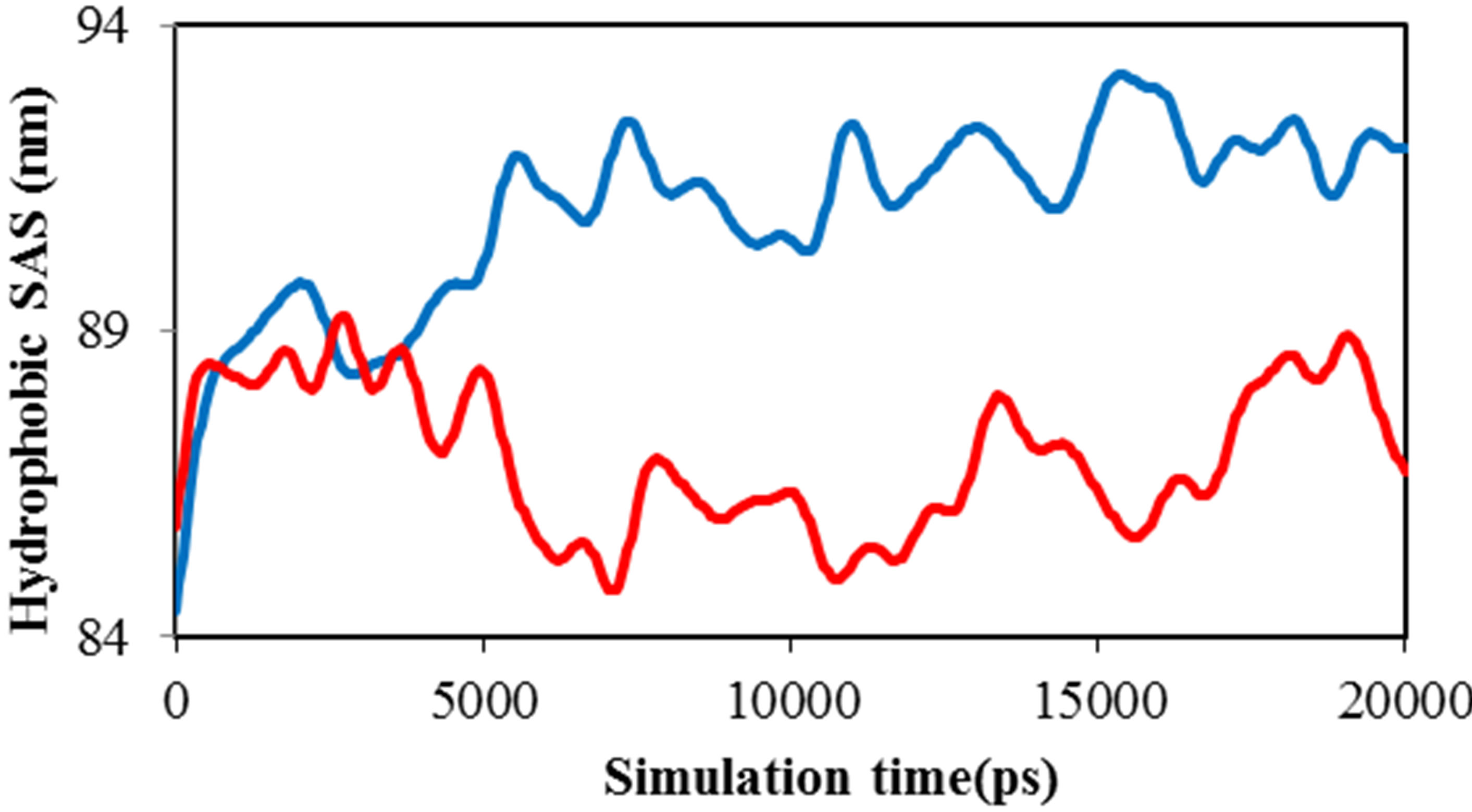
Solvent accessible surface area for ADA was computed by g_sas program. Variation of area in the 20 ns time evolution was significant and obtained. Figure 6(a) shows the surface area of ADA in the presence of 0.014 M TNasPc (anionic) and PcTtme (cationic). This figure shows increase of ADA surface in the presence of TNasPc.
 (a)
(a)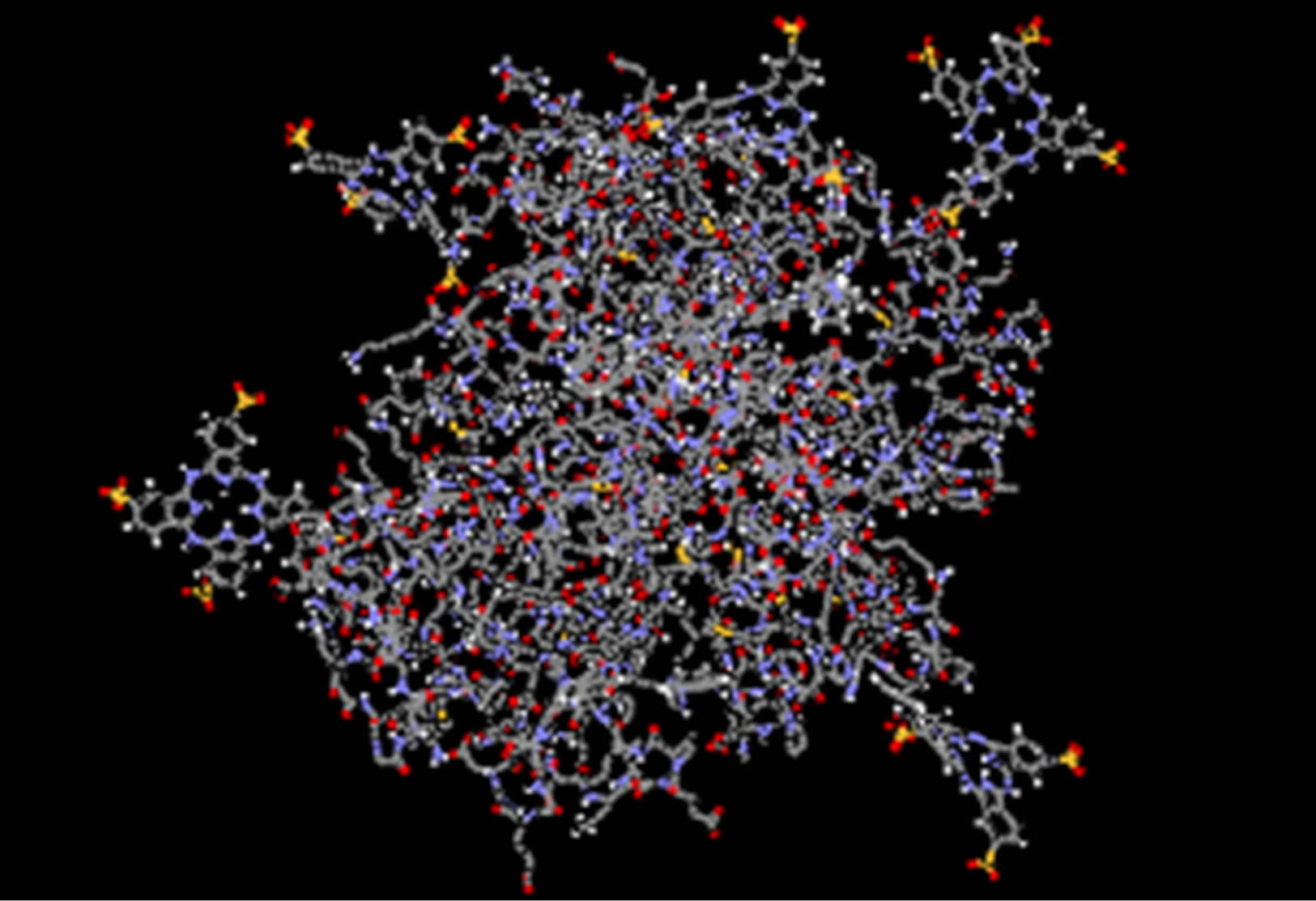 (b)
(b)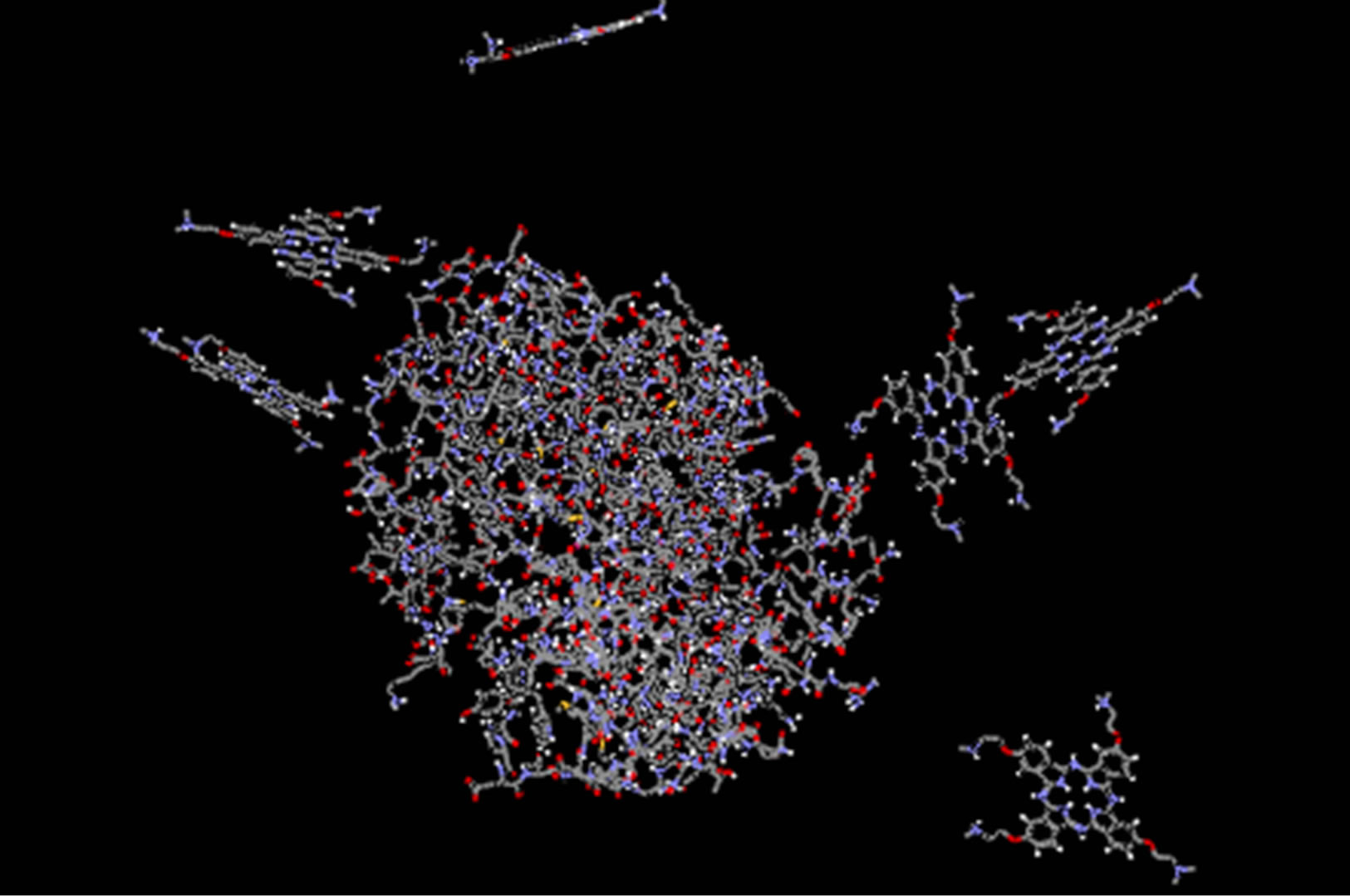 (c)
(c) (d)
(d)
Figure 5. Structure of ADA at the initial time of 30 ns MD in the presence of 6 molecules of anionic phthalocyanine at initial (a) and final (b) and cationic phthalocyanine at initial (c) and final time (d).
This proves the ADA has been unfolded more in the presence of TNasPc.
Figure 6(b) shows the Rg for ADA in the 20 ns time interval in the presence of TNasPc and PcTtme. This figure shows decrease of Rg for ADA in the presence of cationic phthalocyanine. This proves the ADA has been unfolded more in the presence of TNasPc and therefore radius gyration of ADA in system with TNasPc is more than PcTtme.
The radial distribution function of TNasPc around TNasPc and PcTtme around PcTtme versus distance is shown in Figure 6(c). It can be seen that RDF is increased for PcTtme. Increase of RDF with increase of self-aggregations in PcTtme is usually observed. It confirmed that cationic tendency to self aggregate is more than anionic Pcs.
Distance diagram of TNasPc-TNasPc and PcTtmePcTtme (DRG-DRG) by molecular dynamics in the presence 0.014 M TNasPc and PcTtme showed that distance between cationic phthalocyanine is lower than anionic derivative which is depicted in Figure 6(d). It means that cationic phthalocyanine aggregate more than anionic derivative which is in good accordance with RDF diagrams.
3.4. Comparison of Docking Energy
The interaction of phthalocyanine and metallic derivatives with enzyme and with each other was studied by AutoDock 3.0.5. To calculate binding energy of phthalocyanine and metallic derivatives to enzyme, autodock tool was used. This software reports 250 sites which some of these sites have equal energy and so form a cluster.
Tables 1 and 2 show the results of free energy of docking for 10 ranks belong to phthalocyanine and metallic derivatives, respectively. These tables at the first show that free energy of binding for non-metallic phthalocyanine is more negative than metallic derivatives; the first rank is the most probable docking site because of their higher cluster rank. We can see in second column of table the free energy of docking and negative value find at higher position, third column in each calculation run show number of sites with similar energy that are in one cluster, it means the number of randomly occupied sites which are selected was repeated three times. Docking results show that, each phthalocyanines bind to different site having different free energies of docking. We selected the grid box in two different sizes. In one of them, the box surrounded the entire enzyme while in the second; it only surrounded the active site. The free energy of docking was calculated and sorted so the highest negative free energy appeared in the first rank. To compare the different ligand in different conditions, we only considered the free energy of the first rank. When we look at the results for the whole enzyme, the trend of docking
 (a)
(a) (b)
(b)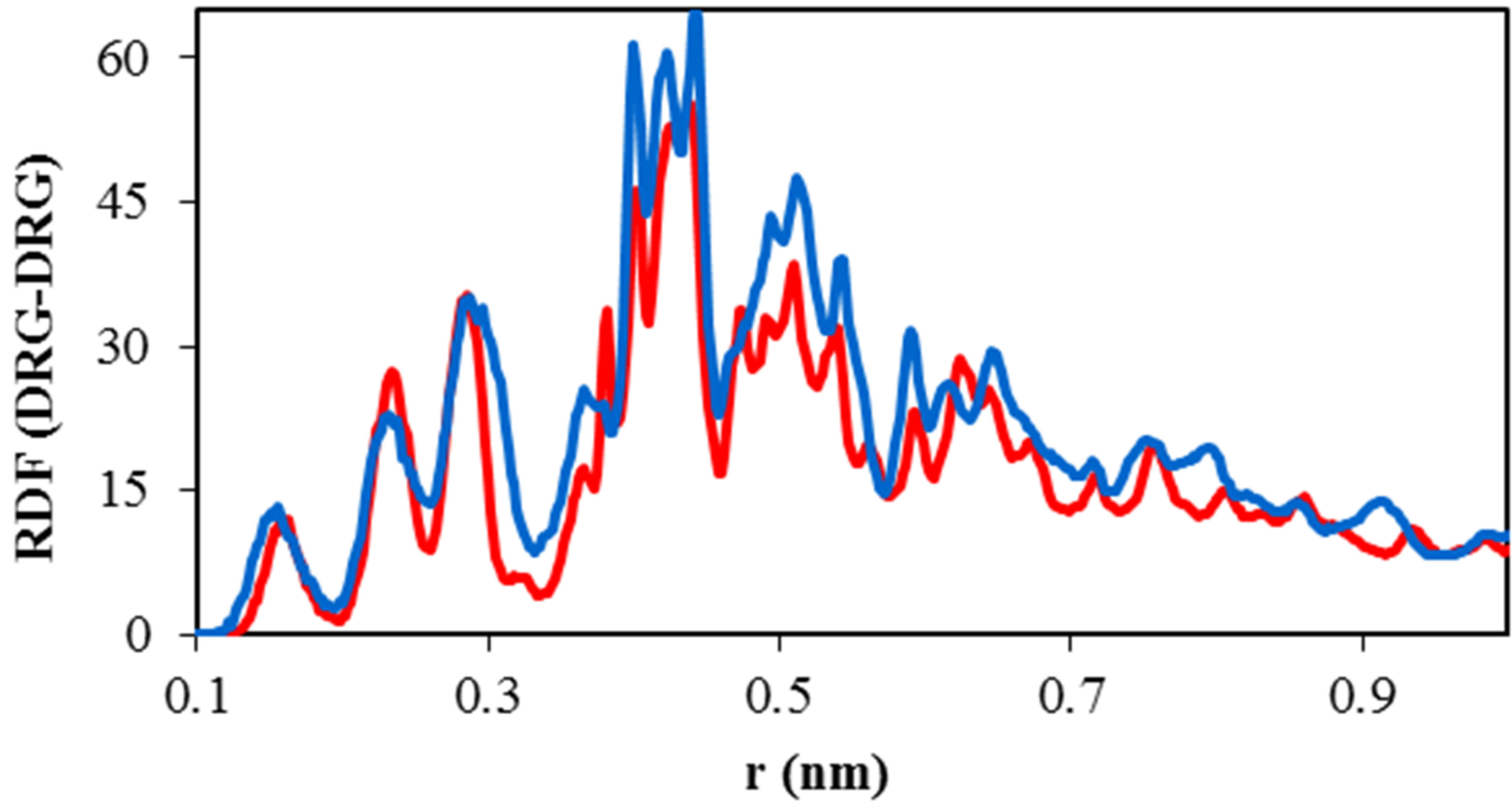 (c)
(c)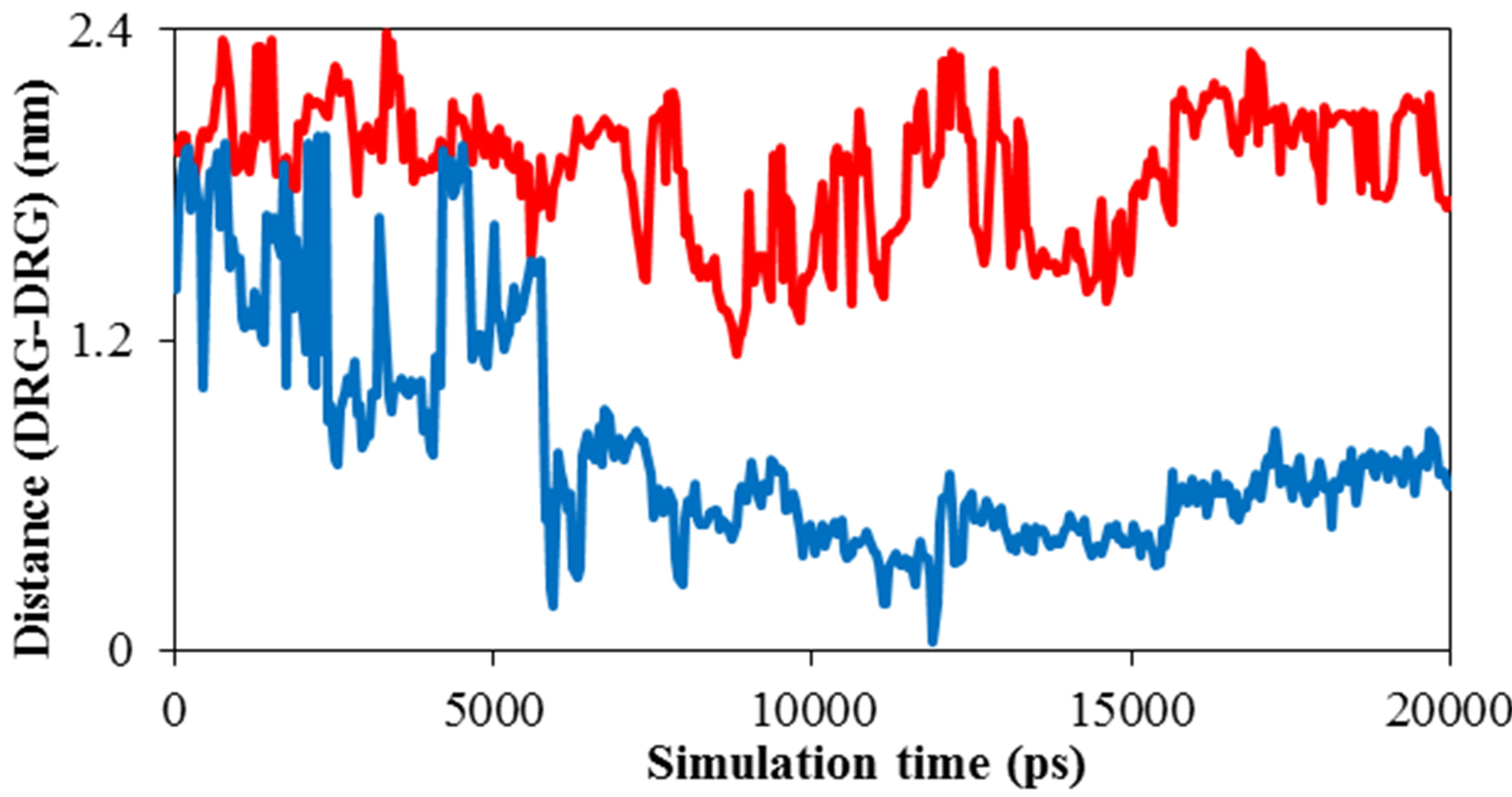 (d)
(d)
Figure 6. Calculated (a) solvent accessible surface area of ADA (b) Radius of gyration of ADA (c). RDF of TNasPcTNasPc and PcTtme-PcTtme (d) Distance (DRG-DRG) of TNasPc-TNasPc and PcTtme-PcTtme by molecular dynamics in 0.014 M TNasPc (Red) and PcTtme (Blue).
free energy in both ligands in different states is metallic derivatives lower than non-metallic phthalocyanine. Since, this value is negative in all three conditions; docking results are now compatible with simulation data.
Values of lowest docking energy for interaction of NiTNasPc for whole and active site of ADA was obtained −3.99 and −2.50 kcal/mol respectively which show more negative value for whole enzyme due to limiting the box to active site cause difficulty in ligands that cannot move freely. Values of lowest docking energy for interaction of TNasPc for whole and active site of ADA was also obtained −4.89 and −1.88 kcal/mol respectively which again show more negative value for whole enzyme due to steric caused by limiting box. Comparison of Tables 1 and 2 shows more binding energy of TNasPc with ADA which is in good agreement with simulation results. As a result Ni metal can stabilize enzyme structure.
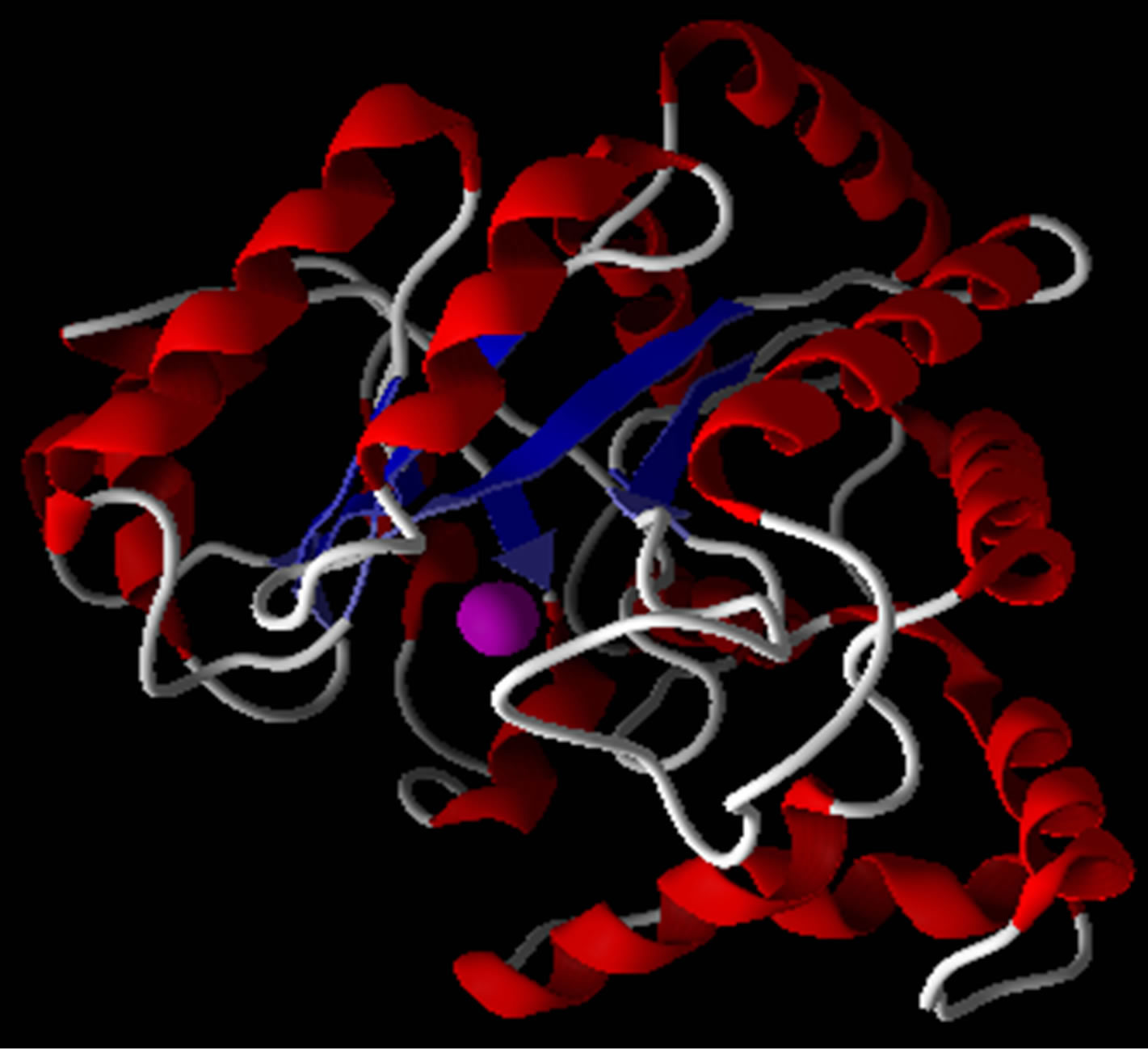
Lowest interaction energy of NiTNasPc with NiTNasPc and TNasPc with TNasPc has been reported in Table 3. Interaction free energy (ΔGint) in kcal/mol for NiTNasPc and TNasPc calculated −2.99 and −1.96 kcal/mol respectively. Averaged value of free energy for phthalocyanine and metallic derivative show more negative value of non-metallic phthalocyanine than metallic derivative. It means that non-metallic phthalocyanine bind better to enzyme. Figure 7(a) shows enzyme (1VFL) taken from protein data bank www.RCSB.org. Figure 7(b) Shows the binding sites for the all ranks negative clusters of non-metallic phthalocyanine near the ADA. Figure 7(c) shows the binding sites for the lower negative clusters of non-metallic phthalocyanine near the ADA. Figure 7(d) shows expended active site of part c which depicts the negative part of non-metallic phthalocyanine located near positive amino acids (Lys203, 229 and Arg232).
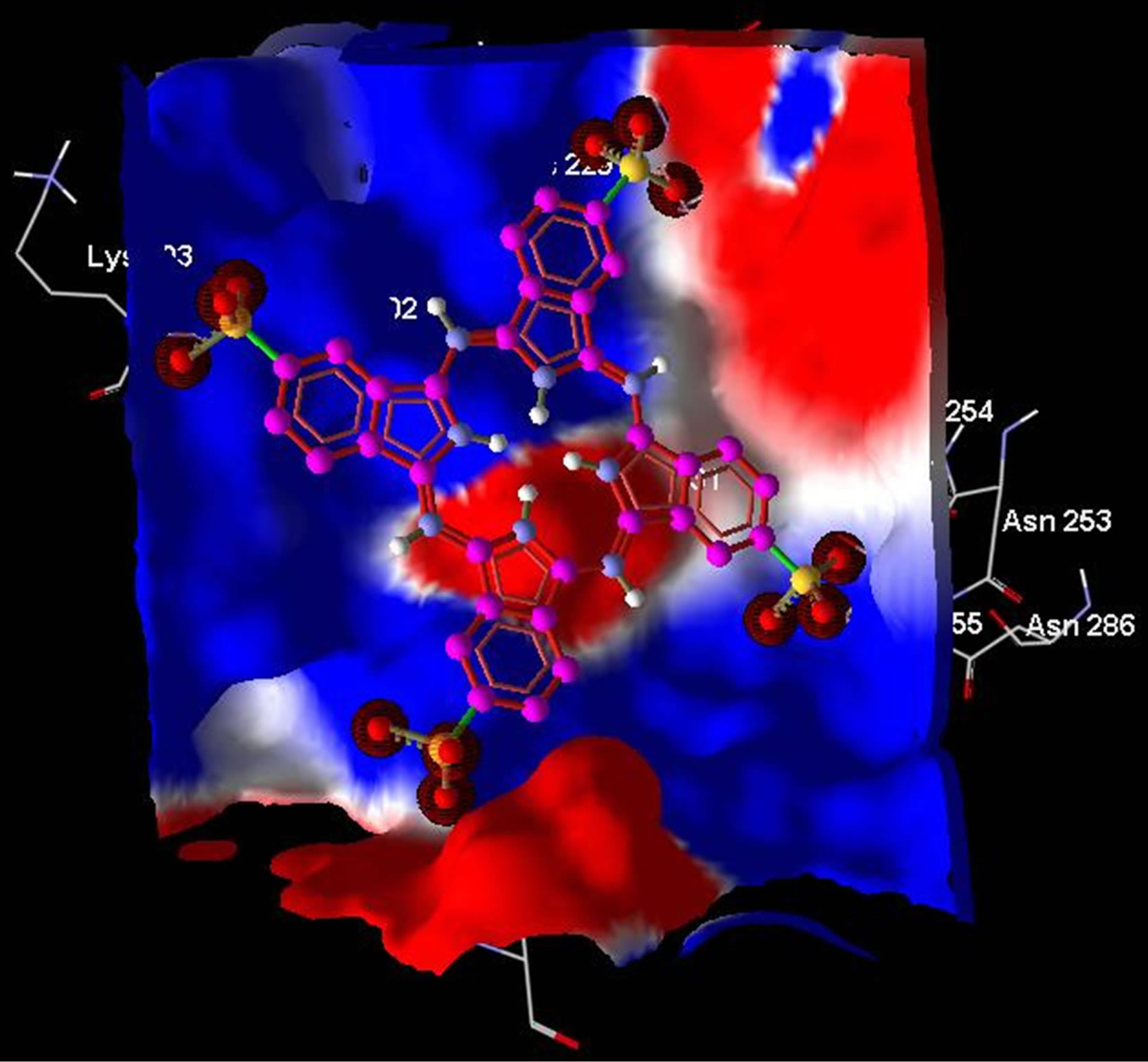
On the other hand, electrostatic surfaces correspond to the most negative docking sites (first negative rank) for interaction of non-metallic and metallic phthalocyanines with ADA were shown in Figure 8. Amino acid residues were explicitly shown for cationic and anionic phthalocyanines, so that the red color represents the negative charge and blue color represents the positive charges, respectively. As we see in this figure, red color surfaces
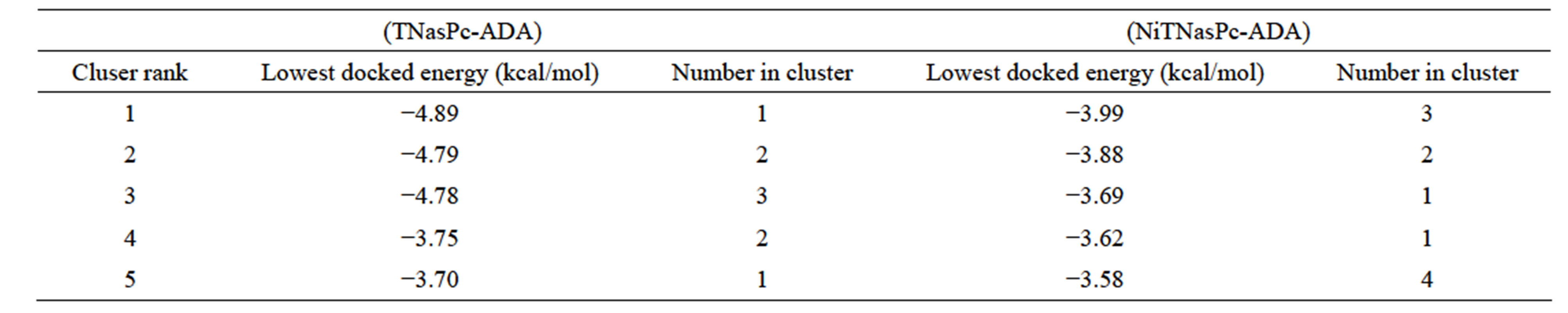
Table 1. Binding free energy (ΔGbind) in kcal/mol for tetra-sulphonated phthalocyanine (TNasPc) and nickel (II) tetra sulfunated phthalocyanine (NiTNasPc) with whole ADA calculated by AutoDock.

Table 2. Binding free energy (ΔGbind) in kcal/mol for Tetra sulfunated phthalocyanine (TNasPc) and Tetra sulfunated nickel (II) phthalocyanine (NiTNasPc) with active site ADA calculated by AutoDock.
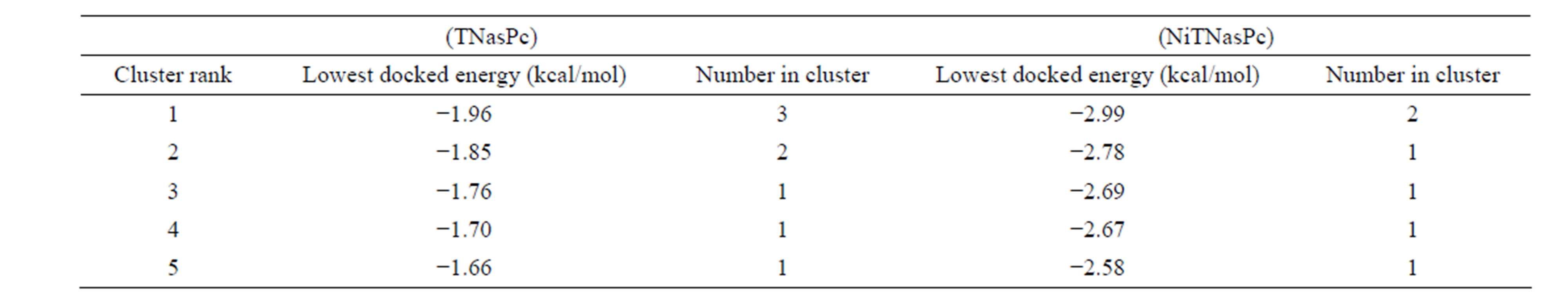
Table 3. Self interaction free energy (ΔGint) in kcal/mol for Tetra sulphonated nickel (II) phthalocyanine (NiTNasPc) and Tetra sulphonated phthalocyanine (TNasPc) calculated by AutoDock.
 (a)
(a)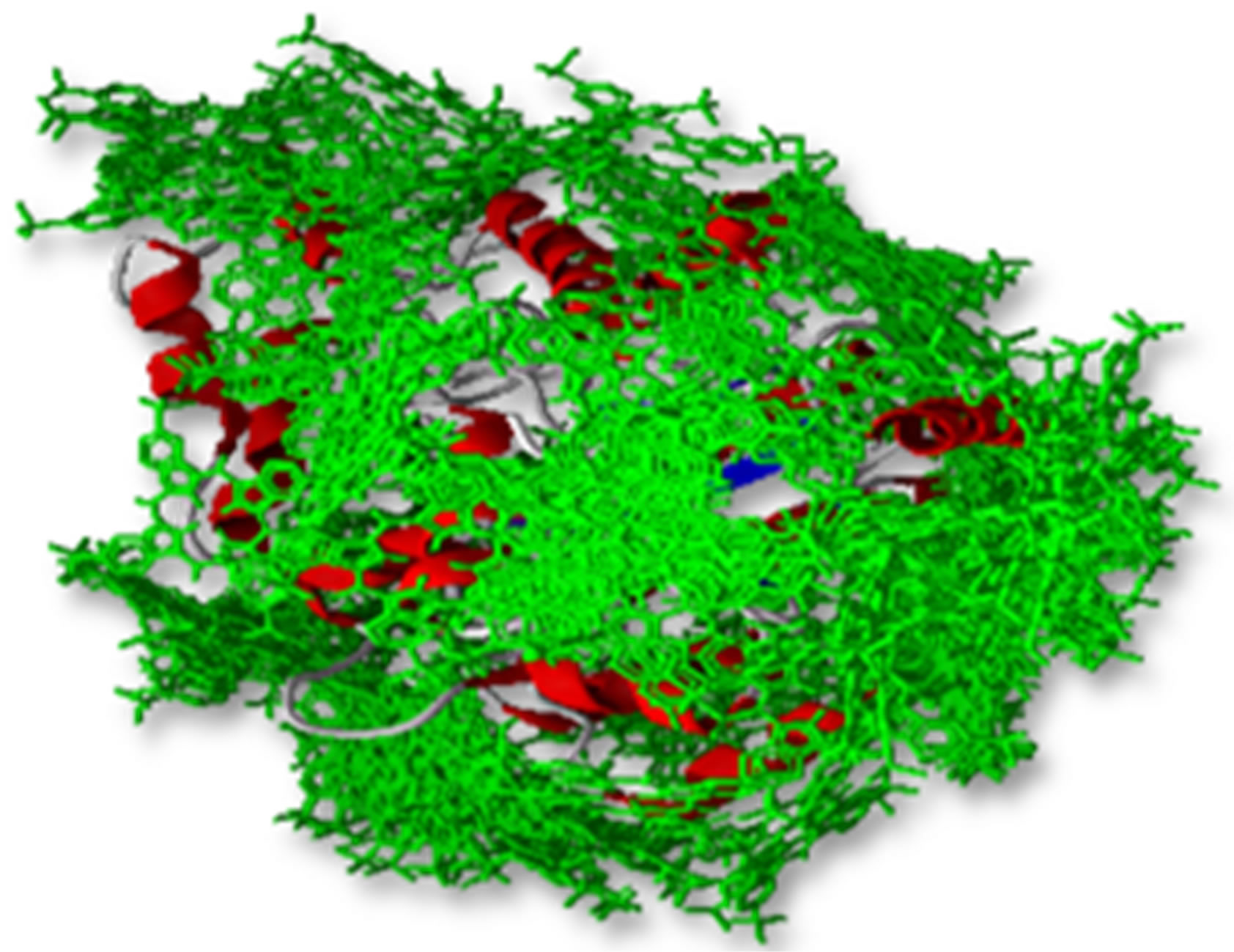 (b)
(b)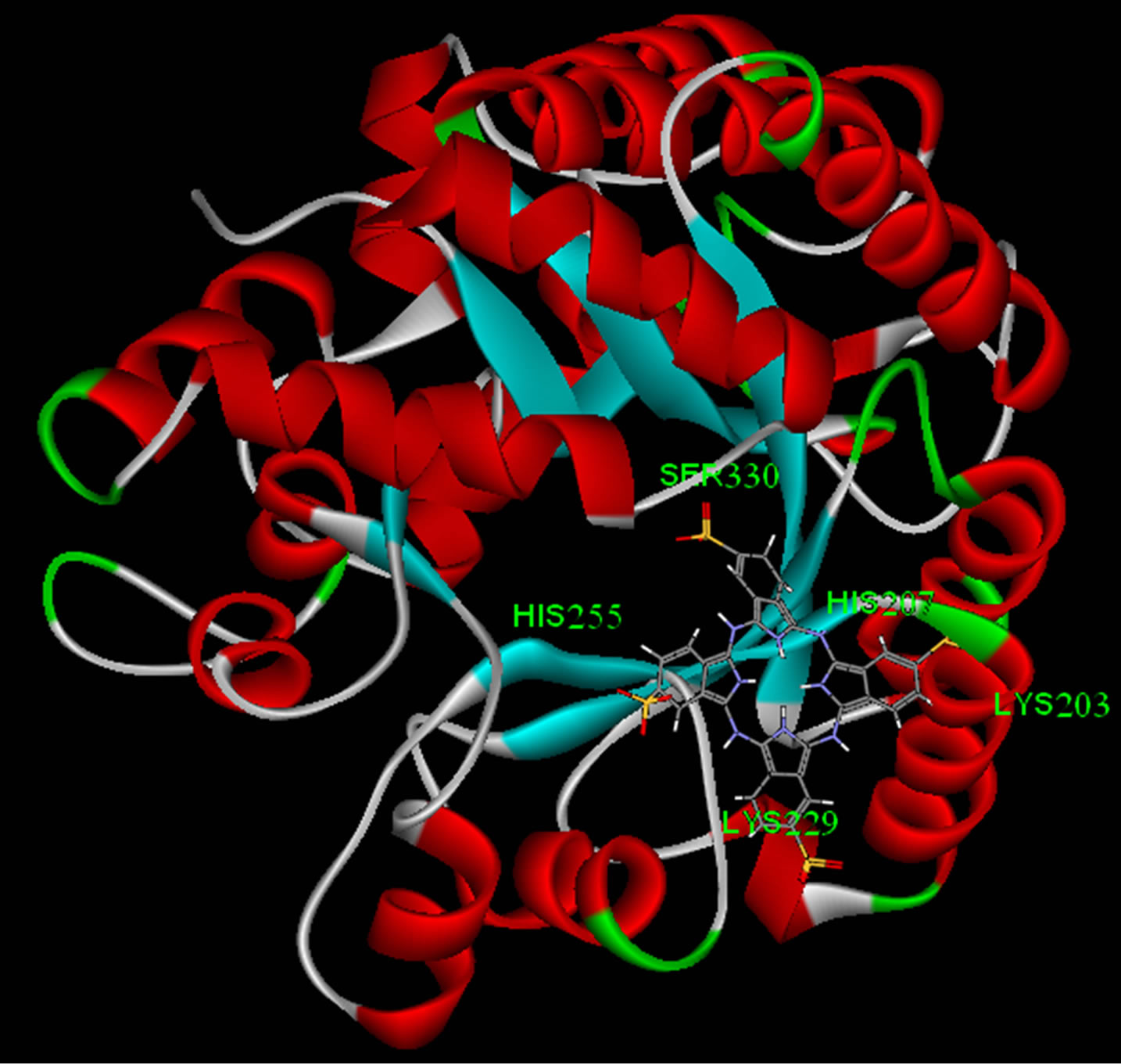 (c)
(c)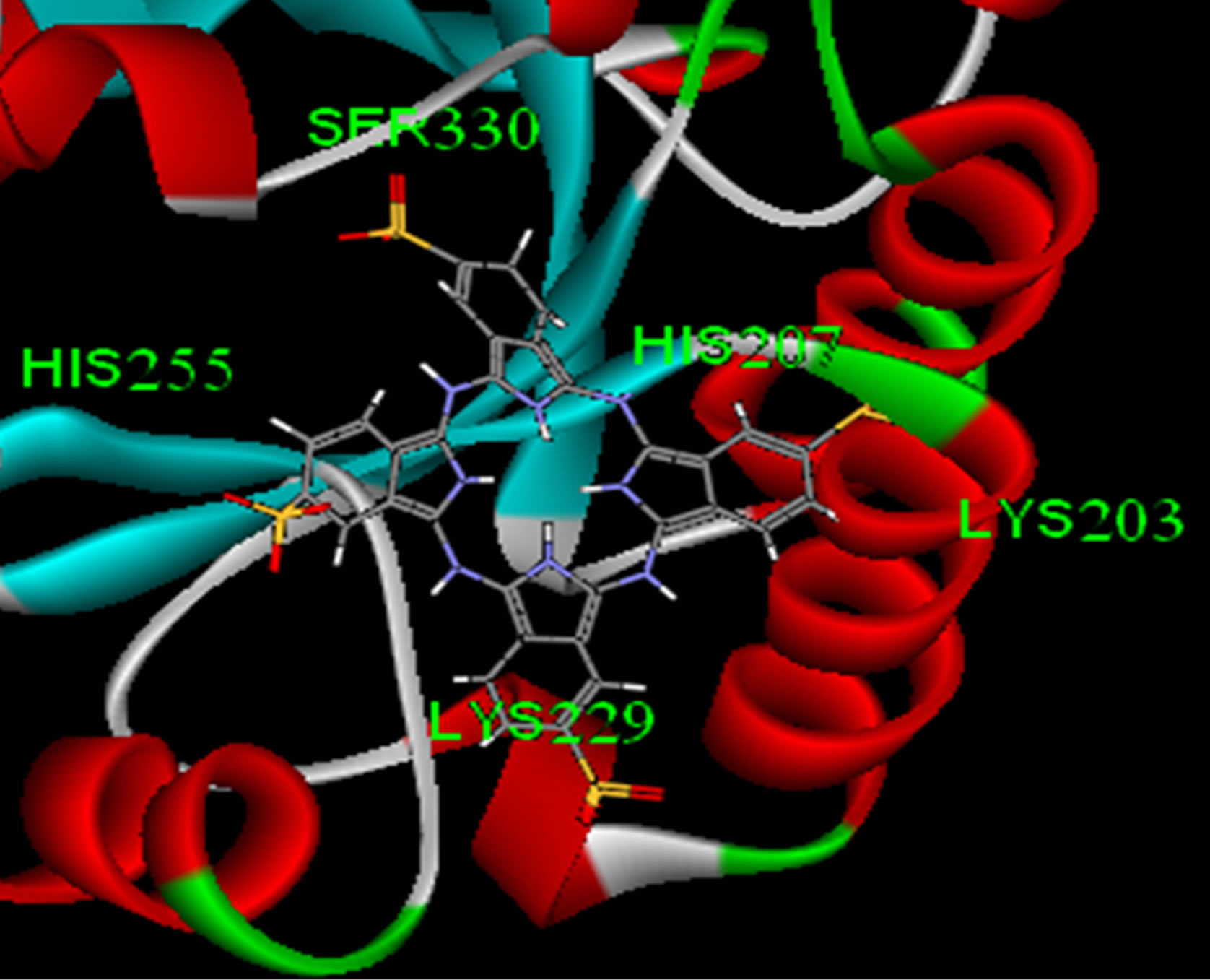 (d)
(d)
Figure 7. (a) Enzyme structure (1VLF) taken from protein data bank www.RCSB.org; (b) The binding sites for the all ranks negative clusters of tetra sulphonated phthalocyanine near the ADA (c) The binding sites for the most negative clusters of tetra sulphonated phthalocyanine near the ADA; (d) Expanded part c, denoting positive amino acids in the binding site.
(representative of negative charges) are higher in the positive parts phthalocyanine (Figure 8(a)) while the blue color (representative of negative charges) are higher in negative parts phthalocyanine (Figure 8(b)). Figures 8(c) and (d) show the amino acids belong to the most negative docking sites that are different for non-metallic and metallic phthalocyanies.
In order to confirm simulation results, we have compared docking energy between cationic and anionic Pcs with ADA and each other by auto dock tool. To better understand which derivative aggregate more or which derivative has more effect on ADA stability, we have calculated their free energies. Tables 4 and 5 show results of binding free energies of PcTtme and TNasPc with ADA and with each other respectively. The binding energy of PcTtme and TNasPc with ADA were −4.26 and −4.89 kcal/mol respectively which shows more interaction of anionic Pc with ADA.
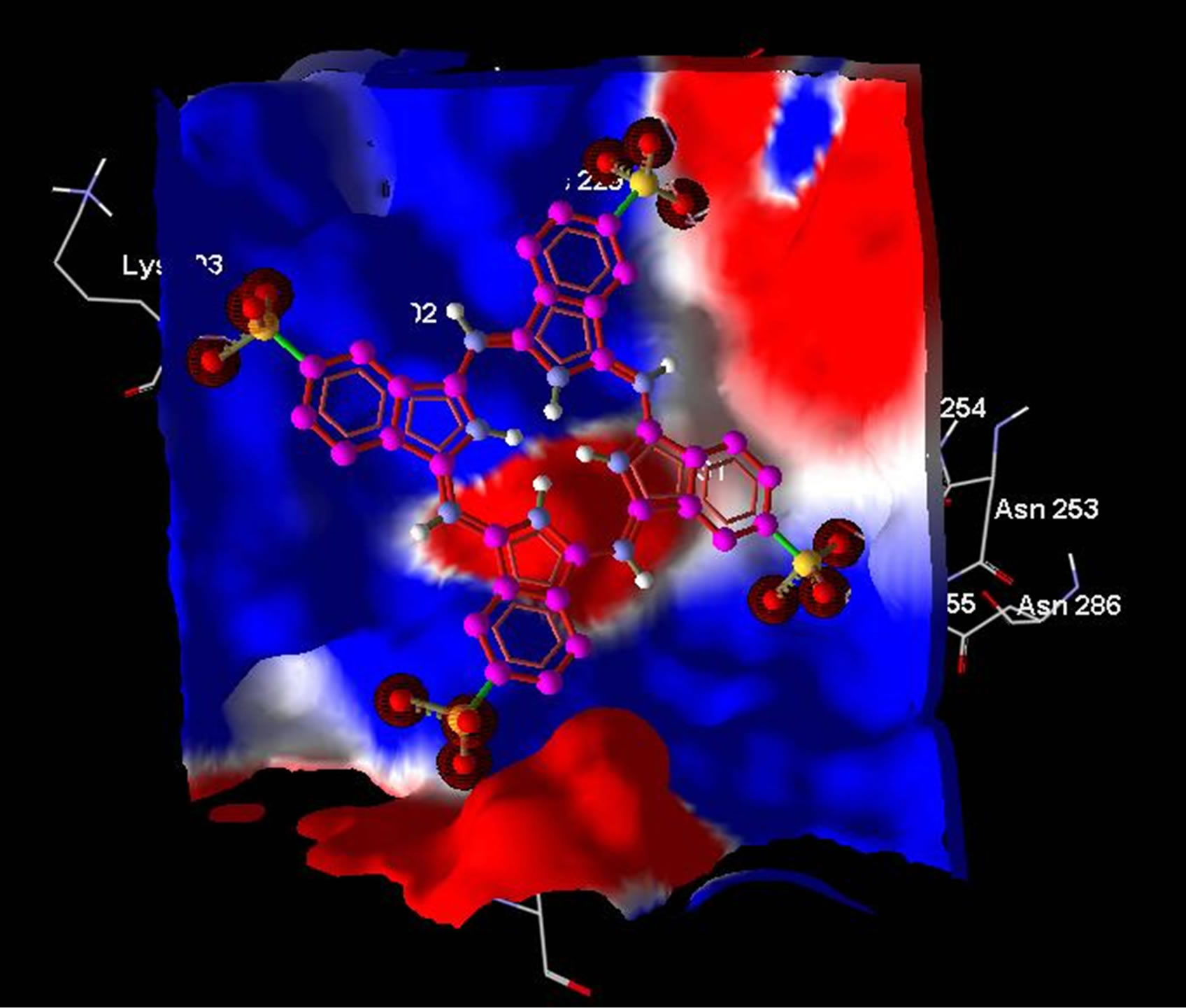
Averaged value of interaction free energy for cationic phthalocyanine and anionic derivative were −6.82 and −1.96 kcal/mol respectively which show more negative value of cationic phthalocyanine. In conclusion anionic phthalocyanine bind better to enzyme and cationic Pcs have more self-aggregation. These calculations are carried out without substrate and are in good agreement with simulation results. On the other hand, electrostatic surfaces correspond to the most negative docking sites (first negative rank) for interaction of cationic and anionic phthalocyanines with ADA were shown in Figure 9. Amino acid residues were explicitly shown for cationic and anionic phthalocyanines, so that the red color represents the negative charge and blue color represents the positive charges, respectively. As we see in this figure, red color surfaces (representative of negative charges) are higher in the positive parts (Figure 9(a) while the blue color (representative of negative charges) is higher in negative parts phthalocyanine (Figure 9(b)). Figure 9(c) and (d) show the amino acids belong to most negative docking sites that are different for cationic and anionic phthalocyanins.
 (a)
(a)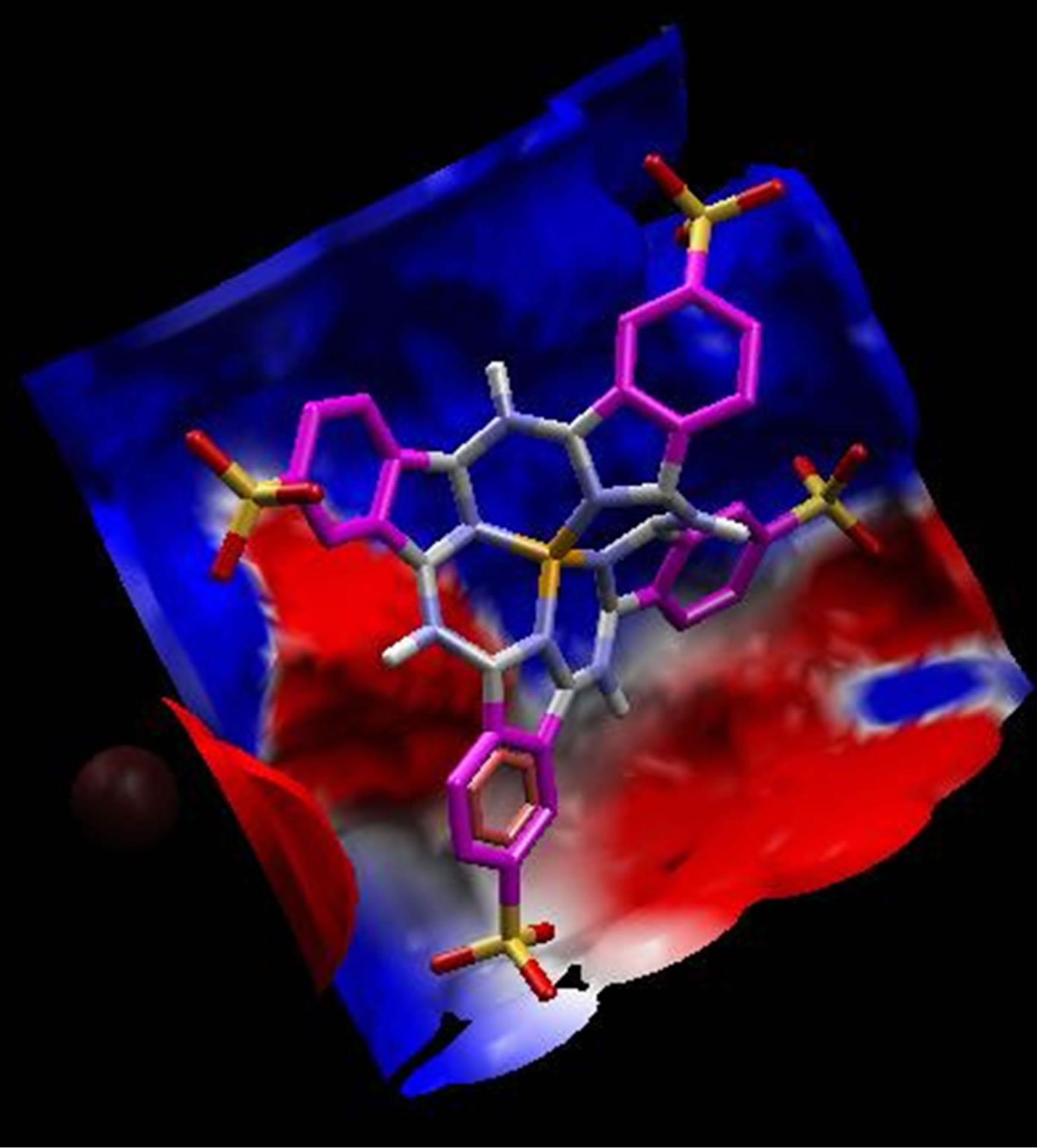 (b)
(b)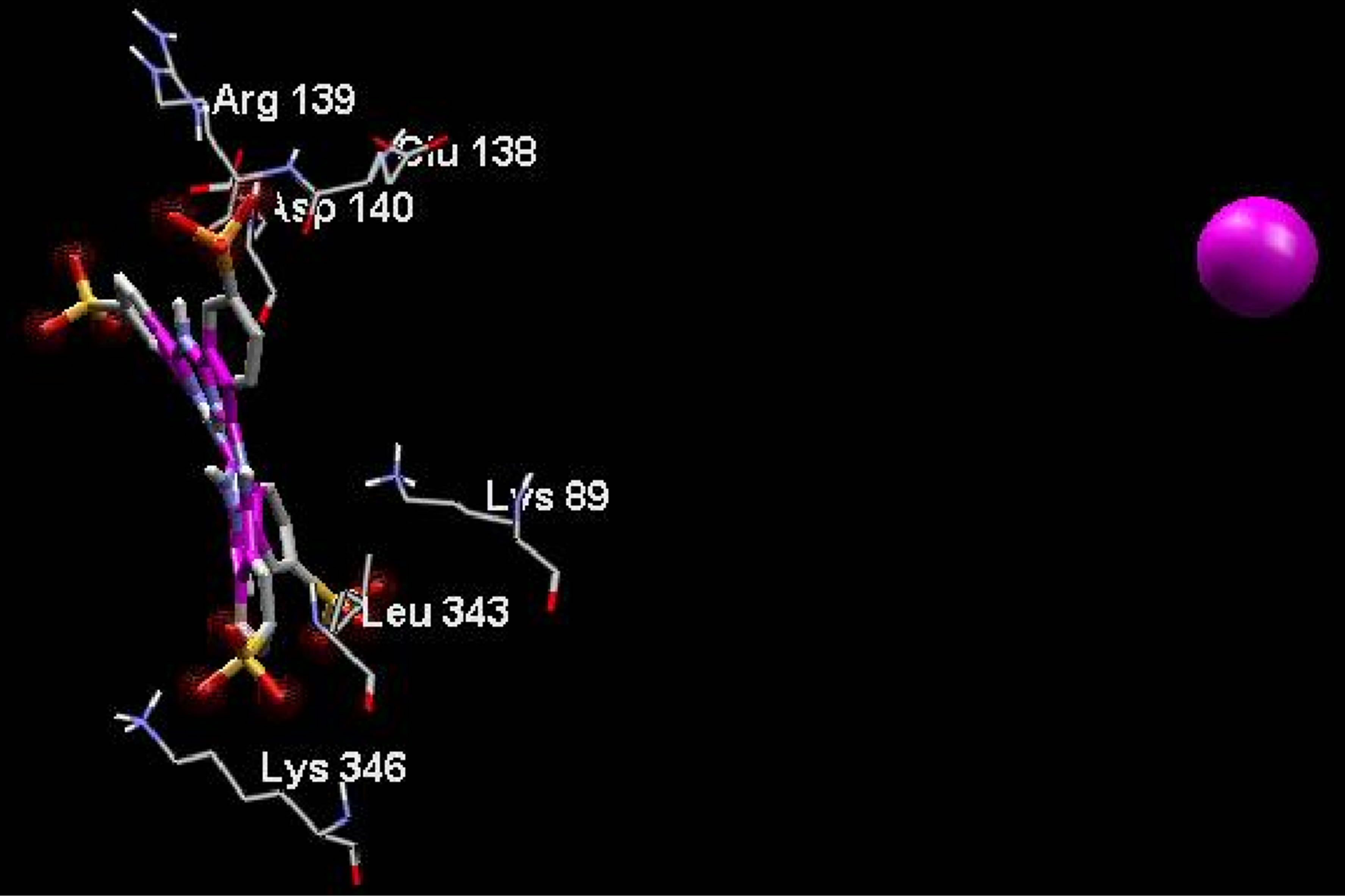 (c)
(c)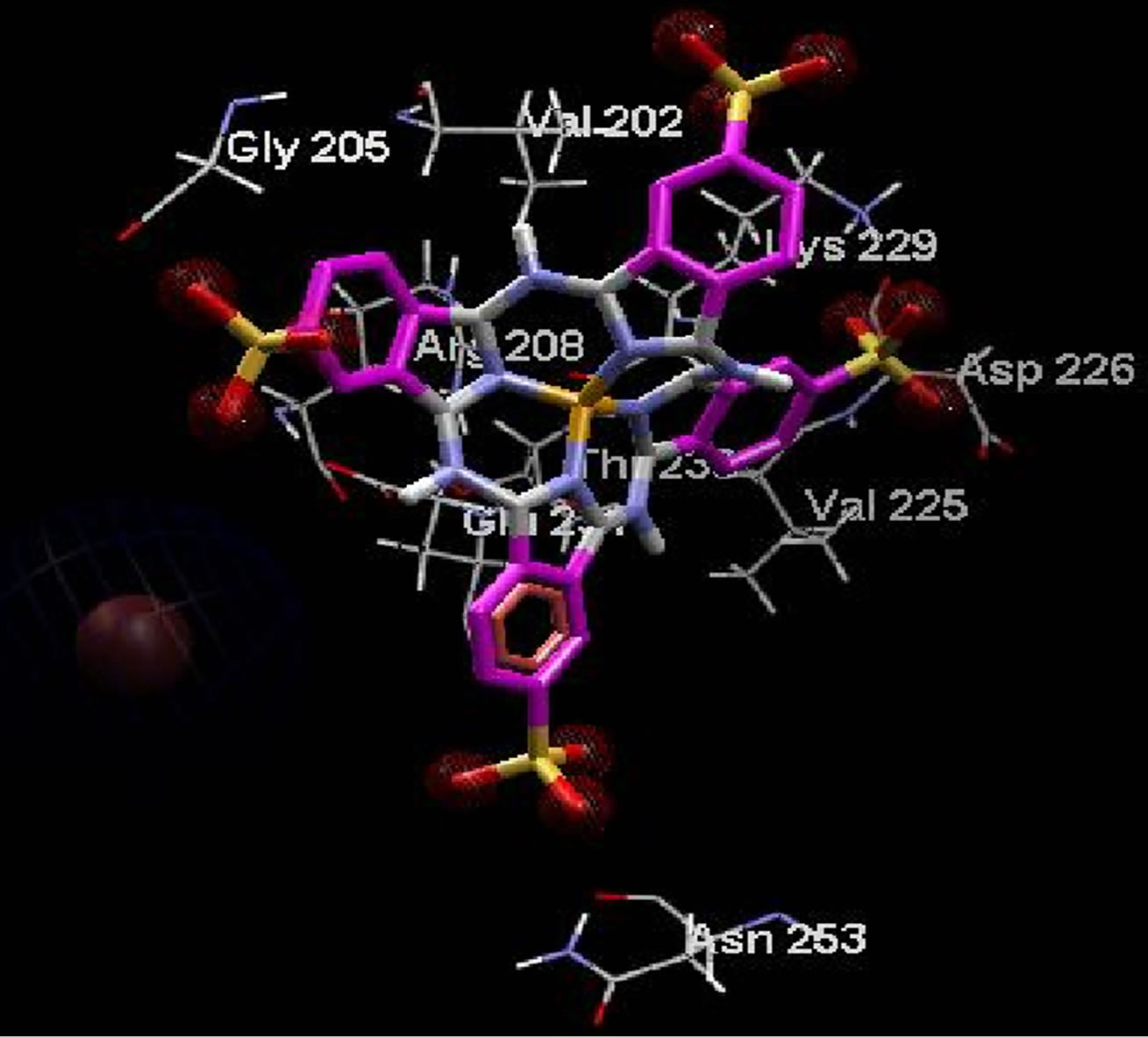 (d)
(d)
Figure 8. Electrostatic surfaces correspond to the most negative docking sites (first negative rank) for interaction of nonmetallic (a) and metallic phthalocyanines (b) with ADA. Amino acid residues were explicitly shown for non-metallic (c) and metallic phthalocyanines (d). Red and blue colors represent the negative and positive charges, respectively.
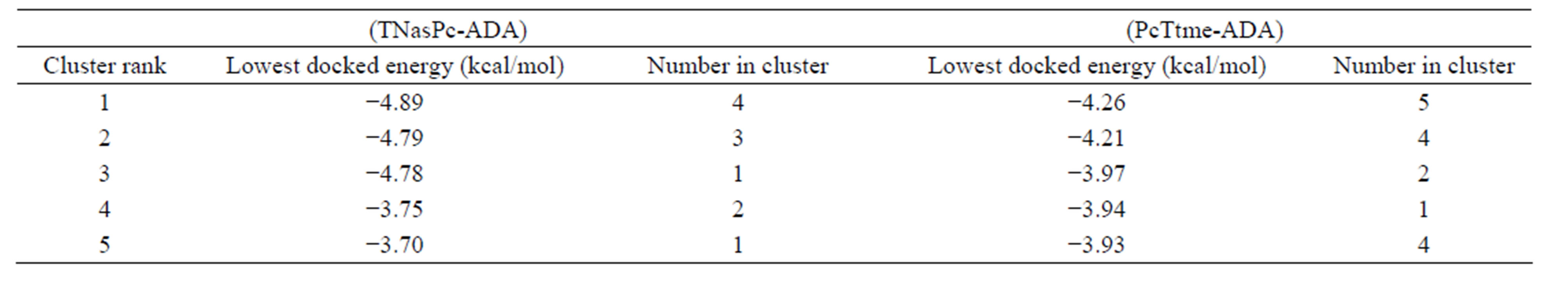
Table 4. Docking free energy (ΔGdoc) in kcal/mol for cationic (PcTtme) and anionic (TNasPc) with ADA calculated by AutoDock.
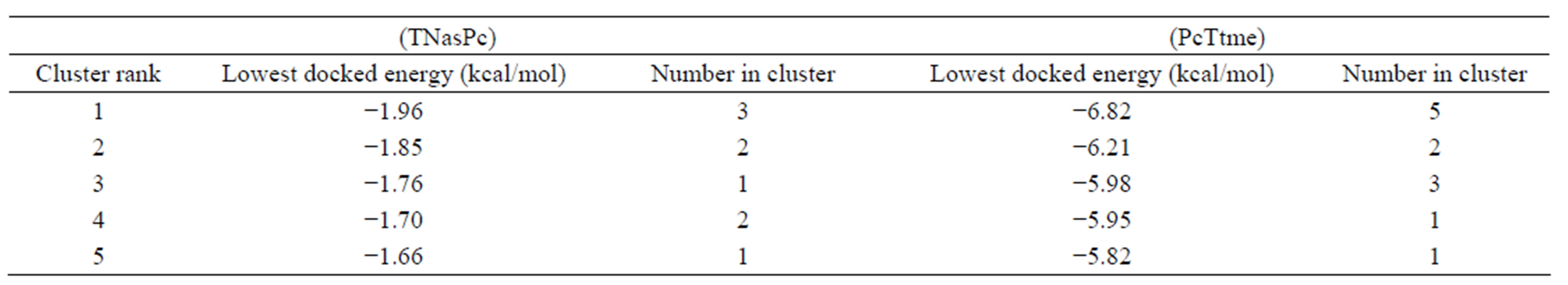
Table 5. Self interaction free energy (ΔGint) in kcal/mol for (PcTtme) with (PcTtme) and (TNasPc) with (TNasPc) calculated by AutoDock.
 (a)
(a)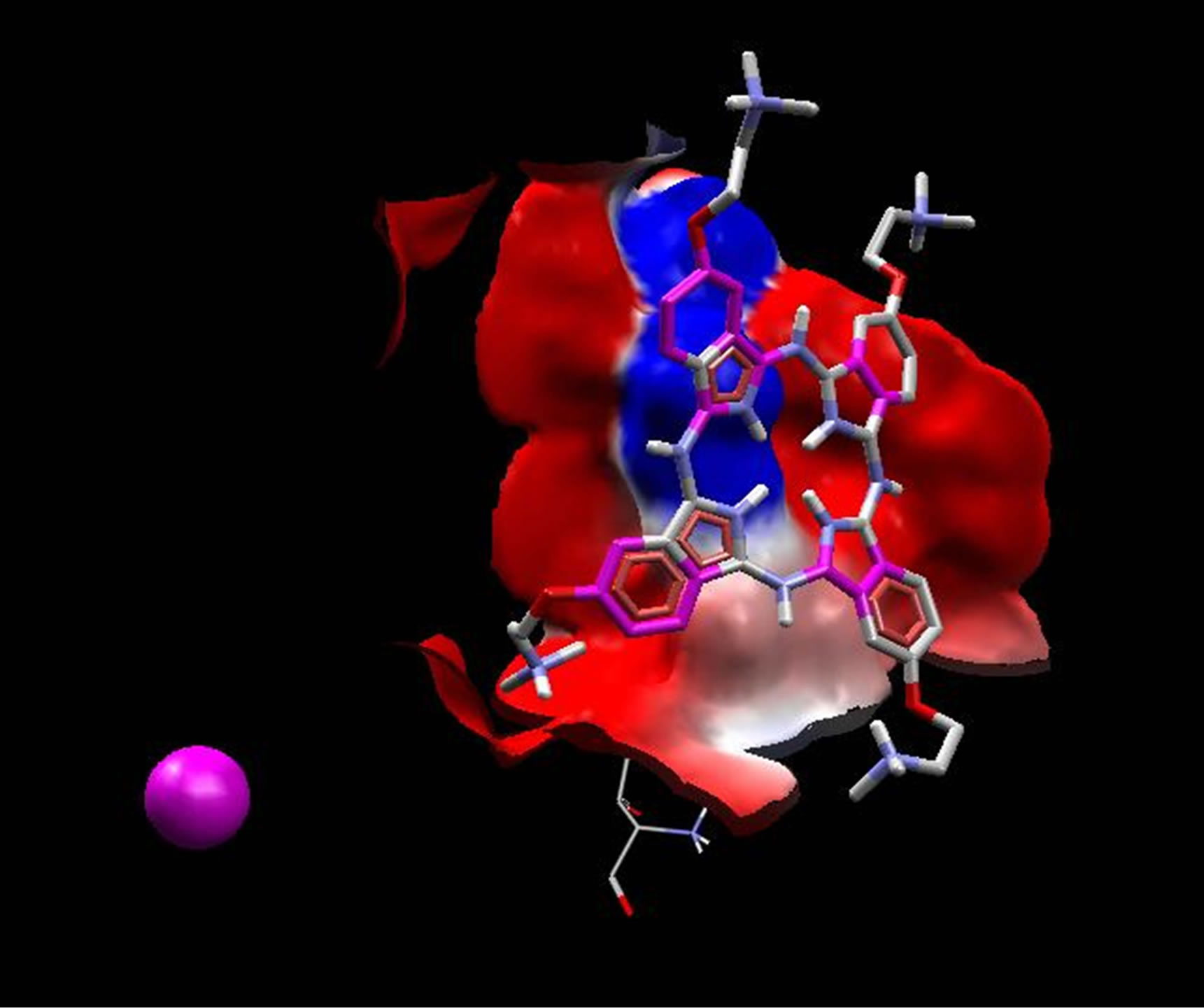 (b)
(b)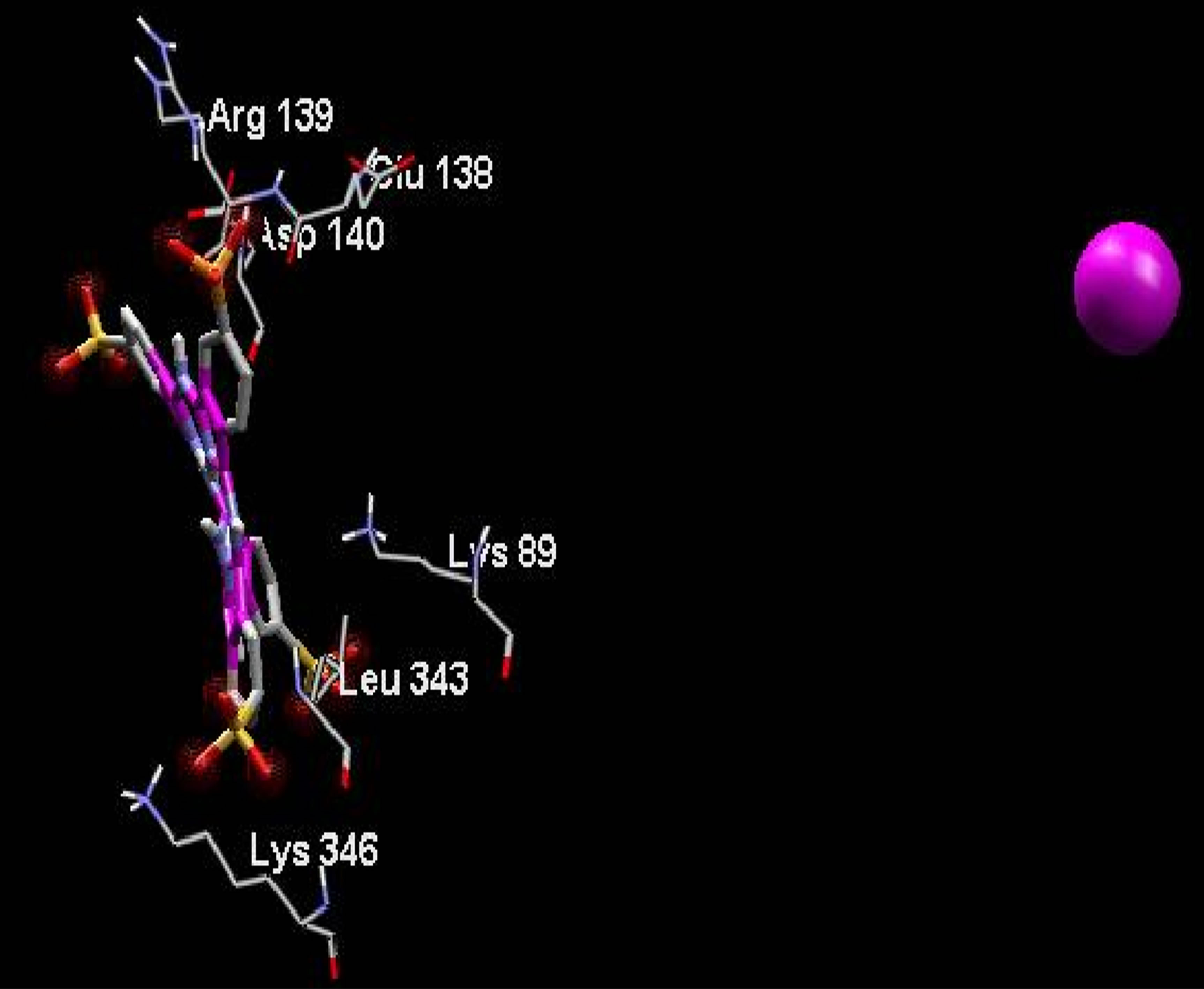 (c)
(c)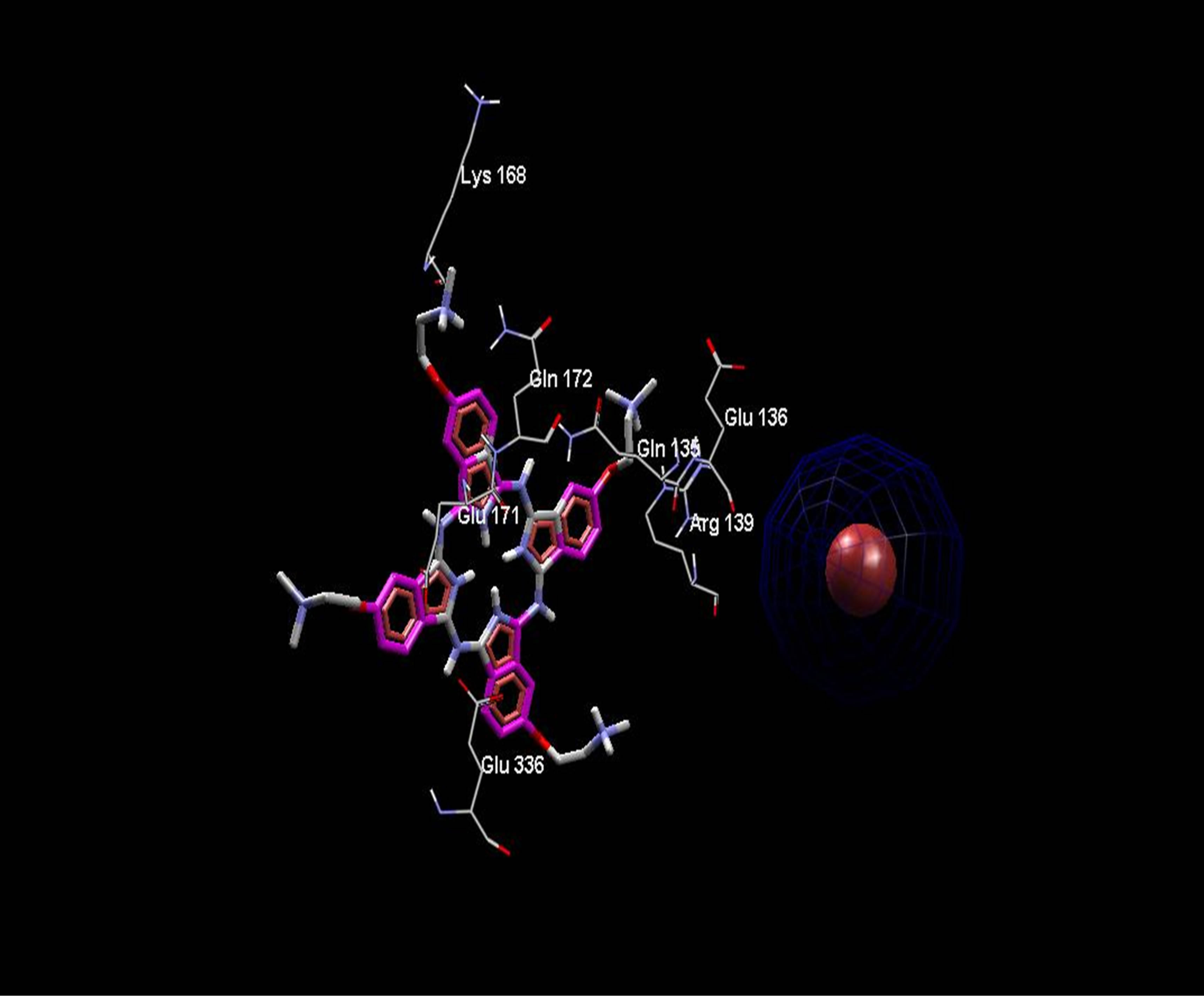 (d)
(d)
Figure 9. Electrostatic surfaces correspond to the most negative docking sites (first negative rank) for interactionof cationic (a) and anionic phthalocyanines (b) with ADA. Amino acid residues were explicitly shown for cationic (c) and anionic phthalocyanines (d). Red and blue colors represent the negative and positive charges, respectively.
4. Conclusion
Metal phthalocyanine decreases the enzyme structure at high concentration and increases it at low concentration. Binding energy of anionic to ADA is more negative than cationic phthalocyanine and dimerization free energy for cationic is negative than anionic. Binding energy to ADA for non-metallic is more negative than metallic phthalocyanine and dimerization free energy for metal anionic Pc is more negative than nonmetal Pc. Therefore interaction of anionic Pc with ADA enzyme is more than cationic form. Protein denaturation is more in the presence of anionic form. This work provides additional information on the effect of a denaturant, Pc, with different concentration, temperature, with Ni metal and non-metallic form and cationic and anionic derivative on its rings on the stability of ADA. It demonstrates that non-metallic Pc and anionic derivative reduce protein stability and more unfolds enzyme structure.
5. Acknowledgements
Financial supporter of Damghan University is acknowledged.
REFERENCES
- N. Spencer, D. Hopkinson and H. Harris, “Adenosine Deaminase Polymorphism in Man,” Annals of Human Genetics, Vol. 32, No. 1, 1968, pp. 9-14. http://dx.doi.org/10.1111/j.1469-1809.1968.tb00044.x
- P. E. Doddona, D. S. Schewach, E. N. Kelly, P. Argos, A. F. Markham and S. H. Orkin, “Human Adenosine Deaminase,” The Journal of Biological Chemistry, Vol. 259, No. 19, 1984, pp. 12101-12106.
- Z. Chang, P. Nygaard, A. C. Chinualt and R. E. Kellems, “Deduced Amino Acid Sequence of Escherichia coli Adenosine Deaminase Reveals Evolutionary Conserved Amino Acid Residues Implication for Catalytic Function,” Biochemistry, Vol. 30, No. 8, 1991, pp. 2273-2280. http://dx.doi.org/10.1021/bi00222a033
- D. K. Wilson, F. B. Rudolph and F. A. Quiocho, “Atomic Structure of Adenosine Deaminase Complexed with a Transition-State Analog: Understanding Catalysis and Immunodeficiency Mutations,” Science, Vol. 252, No. 5010, 1991, pp. 1278-1284. http://dx.doi.org/10.1126/science.1925539
- G. K. Farber and G. A. Petsko, “The Evolution of Alpha/Beta Barrel Enzymes,” Trends in Biochemical Sciences, Vol. 15, No. 6, 1990, pp. 228-234. http://dx.doi.org/10.1016/0968-0004(90)90035-A
- M. Hershfield, B. Mitchell, C. R. Scriver, A. L. Beaudet, W. Sly and D. Valle, “The Metabolic and Molecular Basis of Inherited Disease,” 17th Edition, McGraw-Hill, New York, 1995, pp. 1725-1768.
- M. van Der Weyden and W. Kelley, “Human Adenosine Deaminase. Distribution and Properties,” The Journal of Biological Chemistry, Vol. 251, No. 18, 1976, pp. 5448- 5456.
- M. F. Baghanha, A. Pego and M. A. Lima, “Serum and Pleural Adenosine Deaminase Correlation with Lymphocyte Populations,” Chest, Vol. 97, No. 3, 1990, pp. 605- 610. http://dx.doi.org/10.1378/chest.97.3.605
- T. Hovi, J. F. Smyth, A. C. Allison and S. C. Williams, “Role of Adenosine Deaminase in Lymphocyte Proliferation,” Clinical & Experimental Immunology, Vol. 23, No. 3, 1976, pp. 395-403.
- J. L. Sullivan and R. J. Wedgewood, “Adenosine Deaminase Activity in Lymphocytes,” British Journal of Haematology, Vol. 37, No. 1, 1977, pp. 157-158.
- Y. Edwards, D. Hopkinson and H. Harris, “Adenosine Deaminase Isozymes in Human,” The Jornal Brasileiro de Pneumologia, Vol. 35, 1971, pp. 207-219.
- M. K. Bi, S. Klü, Y. Meriç, F. Ylmaz, O. Baar, G. Ylmaz, O. Yüksel, E. Yldrm and Z. Ztürk, “Serum Adenosine Deaminase Levels in Pancreatic Diseases,” Pancerat, Vol. 7, No. 5-6, 2007, pp. 5-6.
- M. Kaisemann, A. Kritski, M. Pereira and A. Trajman, “Pleural Fluid Adenosine Deaminase Detection for the Diagnosis of Pleural Tubercnlosis,” The Jornal Brasileiro de Pneumologia, Vol. 30, No. 6, 2004, pp. 549-556.
- S. Manjula, A. Raja, S. Rao, A. Aroor and A. Rao, “Serum Adenosine Deaminase Activity in Brain Tumours,” Acta Neurochirurgica, Vol. 121, No. 3-4, 1993, pp. 149- 151. http://dx.doi.org/10.1007/BF01809267
- O. Z. Yildirim, H. Hasanoglu, O. Akyol, M. Gokirmak and N. Koksal, “Serum Adenosine Deaminase Activities in Lung Cancer and Mesothelioma,” Clinical Biochemistry, Vol. 32, No. 4, 1999, pp. 283-285. http://dx.doi.org/10.1016/S0009-9120(99)00013-2
- A. B. P. Lever, M. R. Hempstead, C. C. Leznoff, W. Lin, M. Melnik, W. A. Nevin and P. Seymour, “Recent Studies in Phthalocyanine Chemistry,” Pure and Applied Chemistry, Vol. 58, No. 11, 1986, pp. 1467-1476. http://dx.doi.org/10.1351/pac198658111467
- E. Orti and J. L. Bredas, “Photoelectron Spectra of Phthalocyanine Thin Films: A Theoretical Interpretation,” Journal of the American Chemical Society, Vol. 114, No. 22, 1992, pp. 8669-8675. http://dx.doi.org/10.1021/ja00048a046
- H. Ali and J. E. van Lier, “Porphyrins and Phthalocyanines as Photosensitizers and Radiosensitizers,” In: K. M. Kadish, K. M. Smith and R. Guilard, Eds., Handbook of Porphyrin Science with Applications to Chemistry, Physics, Materials Science, Engineering, Biology and Medecine, World Scientific: Singapore, 2010, pp. 1-119.
- A. Henriksson and M. Sundbom, “Semiempirical Molecular Orbital Studies of Phthalocyanines,” Theoretica Chimica Acta, Vol. 27, No. 3, 1972, pp. 213-222. http://dx.doi.org/10.1007/BF01046366
- A. K. Gosh, D. L. Morel, T. Feng and R. F. Shaw, “Three Dimensional Solar Cells Based on Optical Confinement Geometries,” Journal of Applied Physics, Vol. 45, 1974, pp. 230-236.
- A. Ghosh, P. G. Gassman and J. Almlof, “Substituent Effects in Porphyrazines and Phthalocyanines,” Journal of the American Chemical Society, Vol. 116, No. 5, 1994, pp. 1932-1940. http://dx.doi.org/10.1021/ja00084a038
- R. D. Loutfy and J. H. Sharp, “Photovoltaic Properties of Metal-Free Phthalocyanines,” Journal of Chemical Physics, Vol. 71, No. 3, 1979, pp. 1211-1217. http://dx.doi.org/10.1063/1.438476
- F. R. Fan and L. R. Faulkner, “Photovoltaic Effects of Metalfree and Zinc Phthalocyanines. I. Dark Electrical Properties of Rectifying Cells,” Journal of Chemical Physics, Vol. 69, No. 7, 1978, pp. 3334-3341. http://dx.doi.org/10.1063/1.436987
- H. Jahnke, M. Schonborn and G. Zimmerman, “Synthesis and Characterization of ‘Face-to-Face’ Porphyrins,” Topics in Current Chemistry, Vol. 61, 1976, pp. 133-181. http://dx.doi.org/10.1007/BFb0046059
- W. M. Sharmana, J. E van Liera and C. M. Allen, “Targeted Photodynamic Therapy by Receptor Mediated Delivery Systems,” Advanced Drug Delivery Reviews, Vol. 56, No. 1, 2004, pp. 53-76. http://dx.doi.org/10.1016/j.addr.2003.08.015
- Y. Y. Geng, D. H. Gu, Y. Q. Wu and F. X. Gan, “High Speed Recording Property of Phthalocyanine Thin Film for Compact Disc Recordable,” SPIE—The International Society for Optics and Photonics, Vol. 63, 2003, pp. 63-66.
- K. Petritsch, R. H. Friend, A. Lux, G. Rozenberg, S. C. Moratti and A. B. Holmes, “Liquid Crystalline Phthalocyanines in Organic Solar Cells,” Synthetic Metals, Vol. 102, No. 1-3, 1999, pp. 1776-1777. http://dx.doi.org/10.1016/S0379-6779(98)01035-2
- T. Nyokong and S. Vilakazi, “Phthalocyanines and Related Complexes as Electrocatalysts for the Detection of Nitric Oxide,” Talanta, Vol. 61, No. 1, 2003, pp. 27-35. http://dx.doi.org/10.1016/S0039-9140(03)00356-4
- M. Ethirajan, N. J. Patel and R. K. Pandey, “PorphyrinBased Multifunctional Agents for Tumorimaging and Photodynamic Therapy (PDT),” In: K. M. Kadish, K. M. Smith and R. Guilard, Eds., Handbook of Porphyrin Science with Applications to Chemistry, Physics, Materials Science, Engineering, Biology and Medecine, World Scientific, Singapore. 2010, pp. 249-323.
- X. Yu, C. Z. Qing, Y. Y. Li and X. Z. Hong, “A Molecular Dynamics and Computational Study of Human KAT3 Involved in KYN Pathway,” Science China Chemistry, Vol. 56, No. 4, 2013, pp. 514-523. http://dx.doi.org/10.1007/s11426-012-4802-8
- D. S. Zhao, Y. X. Chen and Q. Liu, “Exploring the Binding Mechanism of Thioflavin-T to the β-Amyloid Peptide by Blind Docking Method,” Science China Chemistry, Vol. 42, 2012, pp. 226-228.
- Y. X. Yu and S. Fujimoto, “Molecular Dynamics Simulation of the A-DNA to B-DNA Transition in Aqueous RbCl Solution,” Science China Chemistry, Vol. 56, No. 4, 2013, pp. 524-532. http://dx.doi.org/10.1007/s11426-012-4825-1
- D. Ajloo, S. Hajipour, A. A. Saboury and S. Zakavi, “Effect of Cationic and Anionic Porphyrins on the Structure and Activity of Adenosine Deaminase,” Bulletin of the Korean Chemical Society, Vol. 32, No. 9, 2011, pp. 3411-3420. http://dx.doi.org/10.5012/bkcs.2011.32.9.3411
- D. Ajloo, M. Sangian, M. Ghadamgahi, M. Evini and A. A. Sabouri, “Effect of Two Imidazolium Derivatives of Ionic Liquids on the Structure and Activity of Adenosine Deaminase,” International Journal of Biological Macromolecules, Vol. 55, 2013, pp. 47-61. http://dx.doi.org/10.1016/j.ijbiomac.2012.12.042
- M. Ghadamghahi, D. Ajloo and M. Moalem, “Kinetic Studies on the Self-Aggregation of a Non Ionic Porphyrin in the Presence and Absence of Ionic Liquid by Molecular Dynamics Simulation,” Journal of Porphyrins and Phthalocyanines, Vol. 16, No. 10, 2012, pp. 1082-1093. http://dx.doi.org/10.1142/S1088424612500915
- M. Ghadamghahi and D. Ajloo, “Calculation and Prediction of Rate and Equilibrium Constants for Aggregation of Porphyrin by Molecular Dynamics, Docking and QSPR,” Journal of Porphyrins and Phthalocyanines, Vol. 15, No. 4, 2011, pp. 240-256. http://dx.doi.org/10.1142/S1088424611003215
- E. van Der Spoel, B. Lindahl, G. Hess, A. E. Groenhof, H. J. Mark and H. J. C. Berendsen, “GROMACS: Fast, Flexible and Free,” Journal of Computational Chemistry, Vol. 26, No. 16, 2005, pp. 1701-1718. http://dx.doi.org/10.1002/jcc.20291
- H. J. C. Berendsen, J. P. M. Postma, W. F. Hermans and J. van Gunsteren, In: B. Pullman, Ed., Intermolecular Forces, Dordecht, Holland, Reidel, 1981, 2005, pp. 331- 342.
- L. Verlet, “Computer ‘Experiments’ on Classical Fluids. I. Thermodynamical Properties of Lennard-Jones Molecules,” Physical Review Letters, Vol. 159, 1967, pp. 98- 103.
- T. A. Darden, D. York and L. G. Pedersen, “A LongRange Electrostatic Potential Based on the Wolf Method Charge-Neutral Condition,” Chemical Physics, Vol. 98, No. 12, 1993, pp. 10089-10092. http://dx.doi.org/10.1063/1.464397
- S. A. Nosé, “A Molecular Dynamics Method for Simulations in the Canonical Ensemble,” Molecular Physics, Vol. 52, No. 2, 1984, pp. 255-268. http://dx.doi.org/10.1080/00268978400101201
- W. G. Hoover, “Canonical Dynamics: Equilibrium PhaseSpace Distributions,” Physical Review Letters, Vol. 31, 1985, pp. 1695-1697.
- M. Parrinello and A. Rahman, “Crystal Structure and Pair Potentials: A Molecular-Dynamics Study,” Physical Review Letters, Vol. 45, No. 14, 1980, pp. 1196-1199. http://dx.doi.org/10.1103/PhysRevLett.45.1196
- L. Ranjan, H. Zhang, H. Monzavi, R. F. Boyko, B. D. Sykes and D. S. Wishart, “VADAR: A Web Server for Quantitative Evaluation of Protein Structure Quality,” Nucleic Acids Research, Vol. 31, No. 13, 2003, pp. 3316- 3319. http://dx.doi.org/10.1093/nar/gkg565
NOTES
*Corresponding author.

